Abstract
Nurr1 is a transcription factor critical for the development of midbrain dopaminergic (DA) neurons. This study modified mouse embryonic stem (ES) cells to constitutively express Nurr1 under the elongation factor-1α promoter. The Nurr1-expression in ES cells lead to up-regulation of all DA neuronal markers tested, resulting in about a 4- to 5-fold increase in the proportion of DA neurons. In contrast, other neuronal and glial markers were not significantly changed by Nurr1 expression. It was also observed that there was an additional 4-fold increase in the number of DA neurons in Nurr1-expressing clones following treatment with Shh, FGF8 and ascorbic acid. Several lines of evidence suggest that these neurons may represent midbrain DA neuronal phenotypes; firstly, they coexpress midbrain DA markers such as aromatic l-amino acid decarboxylase, calretinin, and dopamine transporter, in addition to tyrosine hydroxylase and secondly, they do not coexpress other neurotransmitters such as GABA or serotonin. Finally, consistent with an increased number of DA neurons, the Nurr1 transduction enhanced the ability of these neurons to produce and release DA in response to membrane depolarization. This study demonstrates an efficient genetic manipulation of ES cells that facilitates differentiation to midbrain DA neurons, and it will serve as a framework of genetic engineering of ES cells by key transcription factor to regulate their cell fate.
Keywords: differentiation, dopaminergic neuron, ES cell, Nurr1, Parkinson’s disease, transcription factor
Introduction
This study examines the hypothesis that exogenous expression of a critical transcription factor may facilitate the differentiation of embryonic stem (ES) cells toward certain cell lineages. ES cells are derived from the inner cell mass of preimplantation mouse embryos (Evans & Kaufman, 1981; Martin, 1981) and represent pluripotent cells that can give rise to most cell types in the embryo (Nagy et al., 1990; Nagy et al., 1993). ES cells can be maintained in vitro as pluripotent cells, or differentiated into different cell lineages (Smith, 1991; Desbaillets et al., 2000). These unique properties make ES cells a useful tool to analyse critical steps of cell development, using both animal models as well as in vitro differentiation culture systems (Hooper et al., 1987; Thomas & Capecchi, 1987; Nagy et al., 1990; Nagy et al., 1993; Dinsmore et al., 1996; Wutz & Jaenisch, 2000). In addition, the capacity of ES cells to generate terminally differentiated cell types provides a potentially unlimited resource for cell replacement therapy (Dinsmore et al., 1996; Brustle et al., 1999; Lumelsky et al., 2001; Björklund et al., 2002).
Parkinson’s disease (PD) primarily involves degenerative loss of midbrain dopaminergic (DA) neurons. For therapeutic application of stem cells in PD, it is therefore critical to identify the signalling molecules that specify the cell fate of midbrain DA neurons. In addition, dysregulation of DA neurotransmission has been implicated in other brain disorders such as drug addiction and schizophrenia. During the last several years, signalling molecules, e.g. sonic hedgehog (Shh), fibroblast growth factors (FGF) 4 and 8, that influence the cell fate determination of DA neurons have been identified (Hynes & Rosenthal, 1999). It is thought that such signals lead to induction of specific transcription factors which in turn regulate expression of target molecules necessary for specification of these particular cell types (Goridis & Brunet, 1999). Therefore, one strategy for directing stem cells to a desired neuronal phenotype is to exogenously express key transcription factors.
Nurr1 belongs to the orphan nuclear receptor family (Law et al., 1992), and this transcription factor is relatively enriched in the developing and adult substantia nigra (Zetterstrom et al., 1996; Backman et al., 1999). As Nurr1 has been implicated in the phenotypic specification of midbrain DA neurons in vivo (Zetterstrom et al., 1997; Castillo et al., 1998; Saucedo-Cardenas et al., 1998), it is reasonable to speculate that exogenous expression of this transcription factor would facilitate DA differentiation. Recently the function of Nurr1 has been studied in recombinant neural precursor cells such as adult hippocampus precursors (Sakurada et al., 1999) and the cerebellum-derived immortalized cell line C17.2 (Wagner et al., 1999). Both studies showed that exogenous expression of Nurr1 leads to induction of tyrosine hydroxylase (TH), although by seemingly different mechanisms. Sakurada et al., (1999) demonstrated that Nurr1 binds directly to the TH gene promoter, resulting in induction of TH, in the absence of neuronal differentiation and without the expression of other DA markers. In contrast, Wagner et al., (1999) showed that Nurr1 can induce TH expression in C17.2 cells only when cocultured with ventral mesencephalic type 1 astrocytes. As in both studies partially committed precursors had been used, the observed difference may be due to the specific characteristics of those cells. Therefore, it is of great interest to explore how exogenous Nurr1 expression may affect the differentiation potential of uncommitted pluripotent ES cells. Here, we generated ES cell lines that express Nurr1 and demonstrate that the Nurr1-transduction remarkably facilitates ES cells to differentiate into the midbrain DA neuronal lineage in vitro.
Materials and methods
ES cell culture and in vitro differentiation
The mouse blastocyst-derived ES cell line D3 was obtained from A.T.C.C. (Rockland, MD), propagated and maintained as described previously (Deacon et al., 1998). In brief, undifferentiated ES cells were cultured on gelatin-coated dishes in Dulbecco’s modified minimal essential medium (DMEM, Life Technologies, Rockville, MD) supplemented with 2 mm glutamine (Life Technologies), 0.001% β-mercaptoethanol (Life Technologies), 1 × nonessential amino acids (Life Technologies), 10% donor horse serum (Sigma, St. Louis, MO), and 2000 U/mL human recombinant leukaemia inhibitory factor (LIF; R & D Systems, Minneapolis, MN).
ES cells were differentiated into embryoid bodies (EBs) on nonadherent bacterial dishes (Fisher Scientific, Pittsburgh, PA) for four days in EB medium, as described above except for removing LIF and exchanging horse serum with 10% fetal bovine serum (Hyclone, Logan, Utah). The EBs were then plated onto adhesive tissue culture surface (Fisher Scientific). After 24 h in culture, selection of neuronal precursor cells was initiated in serum-free ITSFn medium (Okabe et al., 1996). After 6–10 days of selection, cells were trypsinized and nestin+ neuronal precursors were plated on polyornithine (15 μg/mL; Sigma) and fibronectin (1 μg/mL; Sigma) coated coverslips in N2 medium (Johe et al., 1996) supplemented with 1 μg/mL laminin (Sigma) and 10 ng/mL bFGF (R & D Systems). After expansion for four days, bFGF was removed to induce differentiation to neuronal phenotypes. In some cases, 500 ng/mL Shh-N (R & D Systems) and 100 ng/mL FGF-8 (R & D Systems) was added during NP (neuronal precursor) expansion stage and 200 μm ascorbic acid (Sigma) was added during ND (neuronal differentiation) stage for further induction of DA phenotype (Lee et al., 2000). Cells were eventually fixed 15 days after starting neuronal differentiation.
Plasmid construction
To generate the pEF/Nurr1/IRES/hrGFP plasmid, mouse Nurr1 cDNA was PCR-amplified and inserted into the Sal I and BstE II sites of the pEF/IRES/hrGFP vector (Chung et al., 2002). The insertion of Nurr1 cDNA into the vector was confirmed by restriction digestion and sequence analysis. The Nurr1 construct was further confirmed by a transient cotransfection assay, where it activated a reporter construct containing Nurr1-binding sites (Murphy et al., 1996) (data not shown). A DNA cassette containing the neomycin resistant gene (EC-Neo, Stratagene, La Jolla, CA) was inserted into both pEF/IRES/hrGFP and pEF/Nurr1/IRES/hrGFP using cre recombinase according to the manufacturer’s instruction. Each construct with the neomycin resistant cassette was confirmed by restriction digestion and then used for stable transfection of ES cells.
Stable transfection and isolation of Nurr1-expressing ES cell lines
D3 cells were transfected with either pEF/IRES/hrGFP or pEF/Nurr1/IRES/hrGFP using the lipofectamine plus reagent (Life Technologies). Stably transfected cells were selected in ES medium containing 500 μg/mL Neomycin (G418 Sulphate, Clontech, Palo Alto, CA). Neomycin-resistant clones were isolated from individual colonies, expanded, and total RNAs were prepared from each clone using TriReagent (Sigma). After DNaseI treatment (DNA-free, Ambion, Austin, TX), expression of hrGFP or Nurr1 was tested by RT-PCR using the rTth RT-PCR kit (Applied Biosystems, Foster City, CA) according to the manufacturer’s instruction. The primer sets used for Nurr1 were Nurr517 (5′-AGTACCTTTATGGACAACTACAGCA-3′) and Nurr816R (5′-CGTAGTGGCCACGTAGTTCTGGT-3′). For hrGFP, hrGFP-5 (5′-CAGATCCGCGTGACCAAGGGCGCC-3′) and hrGFP-3 (5′-AGGCCGCCGTCCTCGTAGCGCAGG-3′). The cDNA quality was measured with the exon 3 specific actin primers (5′-GGTGATGACCTGGCCGTCAGGCAGCTCGTA-3′) and (5′-AACCCCAAGGCCAACCGCGAGAAGATGACC-3′).
Immunocytochemistry
For immunofluorescence staining, cells were fixed in 4% formaldehyde (Electron Microscopy Sciences, Ft. Washington, PA) for 30 min, rinsed with PBS and then incubated with blocking buffer (PBS, 10% normal donkey serum; NDS) for 10 min. Cells were then incubated overnight at 4 °C with primary antibodies diluted in PBS containing 2% NDS. The following primary antibodies were used: mouse TuJ1 (Covance, Richmond, CA; 1 : 500), rabbit antiβ-tubulin (Covance; 1: 2000), sheep anti-TH (Pel-Freez, Rogers, Arkansas; 1 : 200), rabbit anti-TH (Pel-Freez; 1 : 200), mouse anti-TH (Chemicon, Temecula, CA; 1 : 200), rabbit antiserotonin; 5HT (DiaSorin, Stillwater, MN; 1 : 2500), sheep antiaromatic l-amino acid decarboxylase; AADC (Chemicon; 1 : 200), rat antidopamine transporter; DAT (Chemicon; 1 : 2000), goat anticalretinin (Swant, Bellinzona, Switzerland; 1 : 1000), rabbit anticholine acetyltransferase; ChAT (Chemicon; 1 : 500), rabbit antigamma aminobutyric acid; GABA (Sigma; 1 : 1000), rabbit antiglutamate; Glu (Sigma; 1 : 200), rabbit antiglial fibrillary acidic protein; GFAP (DAKO, Denmark; 1 : 500), mouse antigalactocerebrosidase C; GalC (Chemicon; 1 : 500). After additional rinsing in PBS, the coverslips were incubated in fluorescent-labelled secondary antibodies (Cy2-,Rhodamine Red-X-, or Cy5-labelled donkey IgG; Jackson Immunoresearch Laboratory) in PBS with 2% NDS for 30 min at room temperature. After rinsing for 3 × 10 min in PBS, sections were mounted onto slides in Gel/Mount (Biømeda Corp., Foster City, CA). Coverslips were examined using a Leica TCS/NT confocal microscope equipped with krypton, krypton/argon and helium lasers.
Cell counting and statistical analysis
Cell density of DA neurons was determined by counting the numbers of TH+ cells and the numbers of β-tubulin+ cells per field at 63 × magnification using a Zeiss Axioplan I fluorescent microscope. Counting was performed on blind coded samples (cultures grown on the coverslips) by two independent investigators per sample. Ten visual fields were randomly selected and counted for each sample, and cell densities were calculated by dividing the numbers of TH+ cells by that of β-tubulin+ cells. Numbers presented in figures represent the average percentage and standard error of mean (SEM) of TH+ cells over β-tubulin+ cells from five samples per ES cell clone.
For statistical analysis, we used Statview software and performed analysis of variance (anova) with an alpha level of 0.01 to determine possible statistical differences between group means. When significant differences were found, posthoc analysis was performed using Fisher’s PLSD (α = 0.05)
RNA preparation and semiquantitative RT-PCR
Total RNA from plated cells at different stages in the differentiation protocol was prepared using the TriReagent (Sigma) followed by treatment with DNase I (Ambion). For RT-PCR analysis, 5 μg RNA was transcribed into cDNA with the SuperScript™ Preamplification Kit (Life Technologies) and oligo (dT) primers. The cDNA was then analysed in a PCR assay using the following primers.
TH: 5′-TCCTGCACTCCCTGTCAGAG-3′, 5′-CCAAGAGCAGCCCATCAAAGG-3′, 423 bp;
AADC: 5′-CCTACTGGCTGCTCGGACTAA-3′, 5′-GCGTACCAGGGACTCAAACTC-3′, 715 bp;
DAT: 5′-CAGAGAGGTGGAGCTCATC-3′, 5′-GGCAGATCTTCCAGACACC-3′, 328 bp;
Ptx3: 5′- CTCTCTGAAGAAGAAGCAGCG-3′, 5′-CCGAGGGCACCATGGAGGCAGC-3′, 491 bp;
AHD2: 5′-CTGCAAGTGAGGAGGTCATC-3′, 5′-CTGCTGGCTTGACAACCAC-3′, 458 bp;
Calbindin: 5′-GCAGTCATCTCTGATCACAGC-3′, 5′-GAGGTCTGTGTACTCTGCTAG-3′, 428 bp;
DBH: 5′-CTGGACAGGCATAAATGGCAG-3′, 5′-GTCTTGCTGGGAATCCAGATG-3′, 449 bp;
GluT: 5′-AAAGTTCAGAGCCTCACCAAG-3′, 5′-GATCAAATCCAGGAAGGCATC-3′, 429 bp.
PCR reactions were carried out with 1 × IN Reaction Buffer (Epicentre Technologies, Madison, WI), 1.4 nm of each primer, and 2.5 units of Taq I DNA polymerase (Promega, Madison, WI). Samples were amplified in an Eppendorf Thermocycler (Brinkmann Instruments, Westbury, NY) under the following conditions: denaturing step at 95 °C, 40 s; annealing step at 60 °C, 30 s; amplification step at 72 °C, 1 min for 20–25 cycles and a final amplification step at 72 °C, 10 min. For the semiquantitative PCR, cDNA templates were normalized by amplifying actin-specific transcripts and levels of gene transcription were detected by adjusting PCR cycling and primer design in such a way that each primer set amplified its corresponding gene product at its detection threshold to avoid saturation effects.
Analysis of catecholamines
Differentiated ES cells in 6 well plates were treated with 200 μL N3 medium supplemented with 50 mm KCl and 0.1 mm Pargyline and the media were collected after 30 min and concentrated solutions of perchloric acid (PCA) were added for a final concentration of 0.1 m PCA/0.1 mm EDTA. For measurement of catecholamine cell contents, cells were harvested in 0.1 m PCA/0.1 mm EDTA. These deproteinated samples were centrifuged and the supernatants were kept at −80 °C until further analysis. Samples were further purified by using a nylon filter 0.22 μm (Osmonics Inc.) then analysed for their catecholamine content by reverse-phase HPLC using a Velosep RP-18 column (100 × 3.2 mm, Brownlee Laboratories) and an ESA Coulochem II electrochemical detector equipped with a 5014 analytical cell as described previously (Wachtel et al., 1997). The mobile phase was composed of a 0.1 m sodium phosphate buffer (pH 2.65), 0.1 mm EDTA, 0.4 mm sodium octyl sulphate, and 9% (v/v) methanol. The flow rate of the mobile phase through the system was 0.8 mL/min. The potential of the guard cell was set at 330 mV. The potential of the first electrode in the analytical cell was set at 0 mV, the second at 310 mV. L-DOPA, dopamine, dihydroxyphenyl acetic acid (DOPAC) and homovanillic acid (HVA) were identified by retention time and quantified based on peak height using an EZChrom Chromatography Data System. The limit of detection for all compounds was < 1 pg.
DA content of each sample was normalized with the amount of total cellular proteins. For protein measurement, after harvesting cells in 0.1 m PCA/0.1 mm EDTA, precipitates were resuspended in 10 mm potassium phosphate buffer with 0.2% triton-X, pH 7 and sonicated. The protein content was measured by Bradford method (Bio-Rad Assay, Bio-Rad Laboratories, Hercules, CA) (Bradford, 1976).
Results
Exogenous expression of Nurr1 increases the differentiation of ES cells to TH+ neurons
Using the expression vector pIRES/hrGFP (Stratagene), it has been previously shown that transgene expression in ES cells is most efficient when linked to the cellular elongation factor-1α (EF) promoter (Chung et al., 2002). The expression vector pEF/Nurr1/IRES/hrGFP was thus constructed to generate Nurr1-overexpressing D3 ES cell lines. From 18 G418-resistant clones, six were found to express Nurr1 (Fig. 1) and two (D3-N2 and D3-N5) were selected for the in vitro differentiation experiments. After expansion of these clones, hrGFP expression was no longer detected either by fluorescent microscopy or RT-PCR, in spite of continuous Nurr1 expression (see below), possibly due to genetic instability of the IRES element. In addition, D3 cells were also transfected with the empty vector pEF/IRES/hrGFP as a control. Out of eight G418-resistant clones, four clones expressed hrGFP (Fig. 1) and clone D3-E5 was selected as a control for the experiments presented in this report.
Fig. 1.
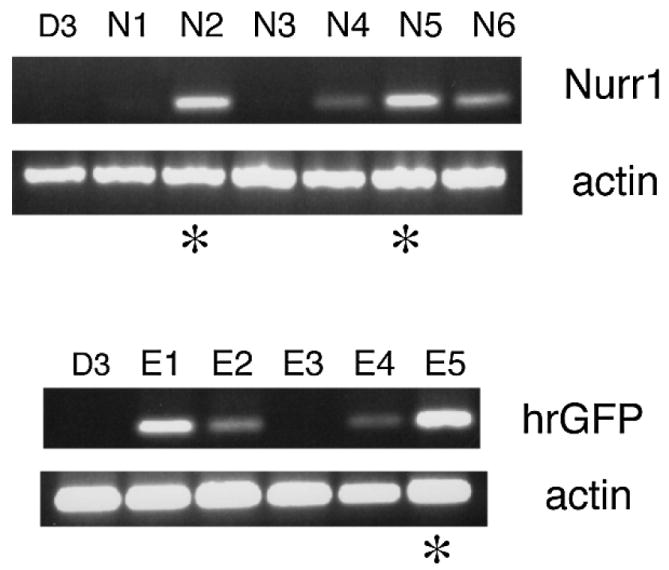
Isolation of ES cell clones that express Nurr1. D3 ES cells were transfected with either the Nurr1-expressing vector pEF/Nurr1/IRES/hrGFP or with empty vector pEF/IRES/hrGFP as a negative control, followed by selection of neomycin-resistant clones and screening for either Nurr1 (N1–N6, upper panel) or hrGFP expression (empty vector-transfected clones, E1–E5, lower panel) by RT-PCR (shown are representative samples). *The clones that were selected for further analysis.
To address whether exogenous Nurr1 expression may directly influence in vitro differentiation of D3 ES cells to the DA phenotype, induction of the TH protein, the first and rate-limiting step of DA synthesis, was analysed using a modification of a previously reported differentiation protocol (Lee et al., 2000). Briefly, cells were first grown to form embryoid bodies (EB), followed by selection and expansion of Nestin+ neural precursors, and differentiation into neural subtypes (see Materials and Methods). Both of the Nurr1-expressing clones, D3-N2 and D3-N5 showed significantly increased proportions of TH+ neurons as compared to parental D3 and control D3-E5 cells (Fig. 2a–f).
Fig. 2.
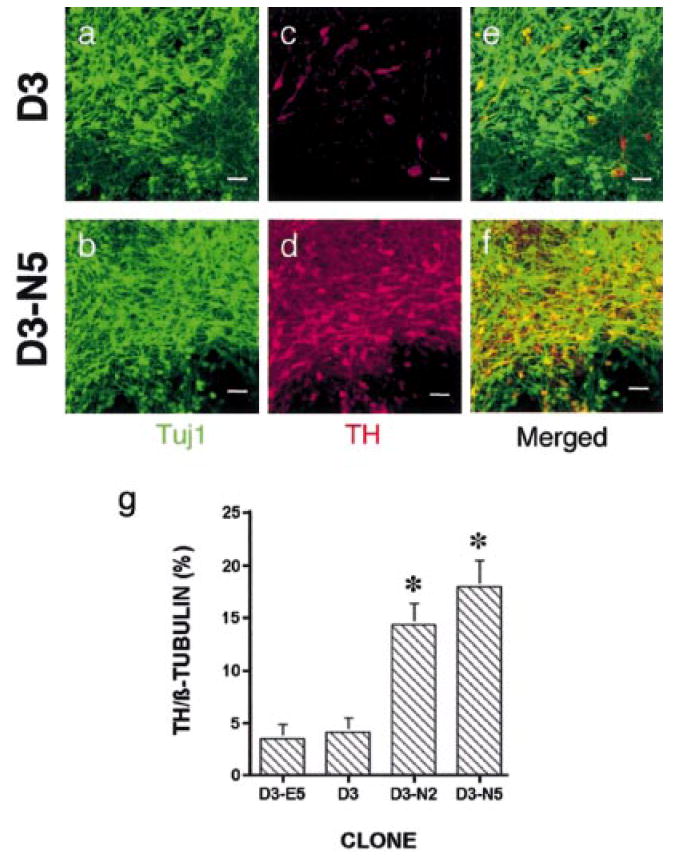
Exogenous expression of Nurr1 in ES cells increases the number of TH+ neurons after in vitro differentiation. (a–f). Detection of β-tubulin (rabbit antiβ-tubulin and Cy2-antirabbit IgG, green) and TH (sheep anti-TH and Rhodamine Red-X-antisheep IgG, red) in in vitro differentiated D3 and D3-N5 cells by immunocytochemistry. Shown are representative fields using confocal microscopy at a 40× magnification. Scale bar, 25 μm. (g) Quantification of TH+ neurons. β-tubulin+ and TH+ cells from in vitro differentiated D3, D3-E5, D3-N2 and D3-N5 ES cells were counted on 10 random fields per each sample, and cell densities of TH+ neurons were calculated by dividing the numbers of TH+ cells by that of β-tubulin+ cells. Each group represents an average of 5 samples from 2 to 3 independent experiments. anova revealed F = 8.95, P < 0.01. Fisher’s PLSD posthoc analysis was performed with a significance level of 0.05. *P < 0.05 compared with D3 and D3-E5 cells.
In both naïve and Nurr1-overexpressing D3 cells, many TH+ neurons were present in clusters, with more TH+ clusters apparent in the latter cell population. Thus, to ensure a more quantitative analysis, a cell counting method was designed using a grid of 10 random fields, which was applied to five coverslips per analysis for appropriate statistical sampling (see Materials and methods for details). All samples were blind-coded and two independent investigators counted the same sample to ensure an unbiased analysis. This counting method revealed the presence of 3.6 ± 1.3% and 4.2 ± 1.4% TH+/β-tubulin+ neurons for naïve D3 and the control D3-E5 cell line, respectively, and 14.4 ± 2.0% and 18.0 ± 2.6% for the two Nurr1-transgenic ES cell clones (the average percentage ± SEM; Fig. 2g), thus demonstrating that exogenous expression of Nurr1 in ES cells leads to a 4–5-fold increase in TH+ cell differentiation under these experimental conditions. As TH+ cells were counted after 15 days of neuronal differentiation (ND), it was next investigated whether TH is induced at an earlier period in the Nurr1-expressing clones. Whereas very limited numbers of TH+ neurons were detected at day 6 of ND, it was observed that there was a marked increase of TH+ neurons in D3-N5 cells at day 10, and the same high levels of TH+ neurons were maintained until day 15 (data not shown). This was also confirmed by reverse phase high performance liquid chromatography (HPLC) analysis of DA production (see below). These results indicate that the DA phenotype was induced by Nurr1 expression in a stage-specific manner in this ES cell system.
Exogenous expression of Nurr1 directs the ES cells to express other midbrain DA markers
It was next determined whether these TH+ neurons expressed other midbrain DA markers. To this end, the in vitro differentiated D3 or D3-N5 cells were double-stained using antibodies against TH and AADC. AADC is involved in the last step of DA biosynthesis. As shown in Fig. 3a–f, all of the TH+ neurons were also AADC+, however, some of the cells had an AADC+/TH− phenotype (Fig. 3d–f), suggesting the presence of serotonergic neurons (see below). It was also examined whether the TH+ neurons coexpressed DAT. As shown in Fig. 3g–i, some of the TH+ cells were also DAT+, indicating that these cells have acquired a mature DA phenotype. In addition, most of the TH+ cells were also positive for calretinin (Fig. 3j–l), which is expressed in many midbrain and olfactory DA neurons (Nemoto et al., 1999). Most of TH+ cells did not express gamma aminobutyric acid (GABA, Fig. 3m–o), suggesting that these TH+/GABA− neurons have the midbrain, but not olfactory, DA phenotype. Furthermore, double-immunocytochemistry showed that these TH+ cells did not coexpress 5HT (Fig. 3p–r) or dopamine β-hydroxylase (DBH; data not shown).
Fig. 3.
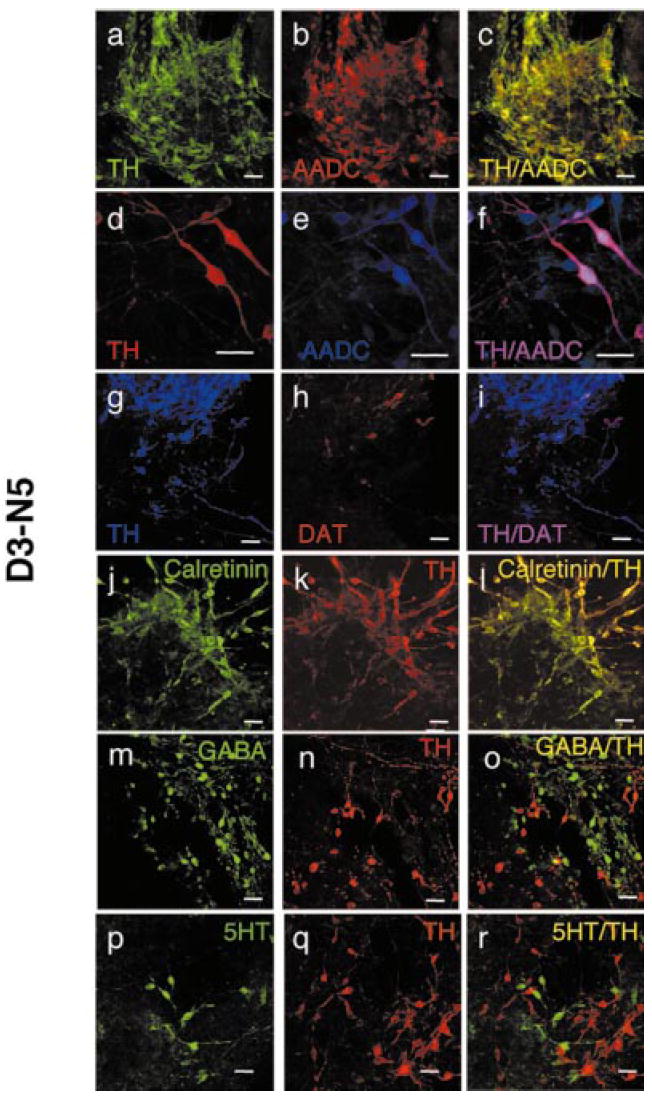
Exogenous expression of Nurr1 correlates with expression of other markers specific for midbrain DA neurons. Detection of TH and AADC (a–f), DAT (g–i), calretinin (j–l), GABA (m–o) and 5HT (p–r) on D3-N5 ES cells after in vitro differentiation. Sheep anti-TH was used for all staining except the double staining with AADC, where rabbit anti-TH was used. All other primary antibodies are as described in materials and methods. Cy2-labelling of secondary antibodies was shown as green, Rhodamine Red-X as red, and Cy5 as blue. Samples were analysed with either 40 × or 80 × magnification. Scale bar, 25 μm.
Semiquantitative RT-PCR analysis was then done in order to determine the expression of midbrain DA neuron-specific molecules at the different stages of in vitro differentiation of D3, D3-E5, D3-N2 and D3-N5 (Fig. 4 and data not shown). It is worth noting that Nurr1 expression was persistently maintained and up-regulated in Nurr1-transduced clones during all three stages of in vitro differentiation, as compared to D3 or D3-E5 (Fig. 4 and data not shown). This contrasts with other studies in which Nurr1 expression was later shut down after the initial screening of Nurr1-expressing clones (Wagner et al., 1999). These dissimilar patterns may be due to the different promoters used in each study to drive Nurr1 expression. The EF-1α promoter used in this study was shown to have active promoter activity during different stages of in vitro ES cell differentiation (Chung et al., 2002). Consistent with the immunocytochemistry data, mRNA expression from the TH, AADC and DAT genes were up-regulated in both D3-N2 and D3-N5 cells after in vitro neuronal differentiation (ND). Interestingly, it was not possible to detect clear induction of TH and DAT gene expression at the initial (ES) or neuronal precursor (NP) stages of differentiation (Fig. 4). In contrast, there was an increase in AADC transcription detected in undifferentiated Nurr1-expressing clones. In addition, the altered expression of other midbrain DA markers was examined including aldehyde dehydrogenase 2 (AHD2), Ptx3 and calbindin. Compared to the parental D3 cells, expression of all three genes was markedly up-regulated in both D3-N2 and D3-N5 clones (Fig. 4 and data not shown). In contrast to these midbrain DA markers, there was no DBH mRNA expression in any of the D3, D3-E5, D3-N2 or D3-N5 cells and no difference in the expression levels of the neuron-specific glutamate transporter mRNA (Tanaka, 1993). Given that these are markers for noradrenergic and glutamatergic neurons, respectively, this data indicates a specific effect of Nurr1 on the expression of midbrain DA neuronal markers (see below).
Fig. 4.
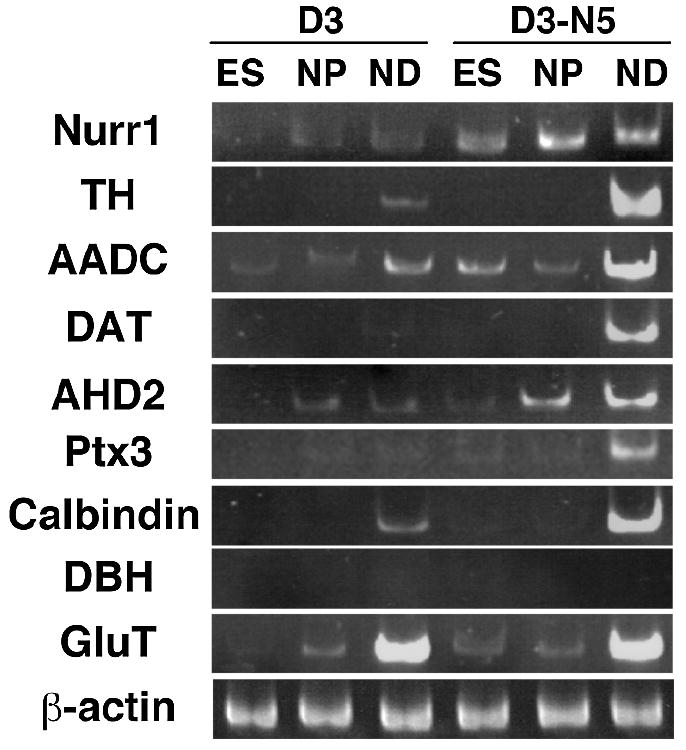
Semi-quantitative RT-PCR analysis of midbrain DA markers during in vitro differentiation of D3 vs. D3-N5 cells. Samples from undifferentiated ES cell stage (ES) in the presence of LIF, nestin+ neuronal precursors (NP) and fully differentiated neurons after 15 days of neuronal differentiation (ND) were analysed for specific marker gene expression. To avoid saturation effect, the numbers of PCR cycles were experimentally determined for each cDNA, such that amplification of specific transcript is in a linear range. RT-PCR analyses were repeated twice using RNA samples prepared from two independent in vitro differentiation experiments, resulting in similar results. In addition, a same pattern was obtained when D3-E5 and D3-N2 substituted for D3 and D3-N5, respectively.
Effect of exogenous Nurr1 expression on the differentiation of neural phenotypes other than DA neurons
The effect of Nurr1 on the differentiation of ES cells into other neural phenotypes was next investigated, using various neuronal and non-neuronal molecules. Both D3 and D3-N5 cells generated GABA (Fig. 5a and b), choline acetyltransferase (ChAT, Fig. 5c and d), and glutamate (Glu, Fig. 5e and f) positive neurons in similar amounts, indicating that Nurr1 does not affect the differentiation into these neuronal subtypes. In case of the glutamatergic neuronal phenotype, the immunohistochemical results were confirmed by RT-PCR showing that the neuronal glutamate transporter gene was equally expressed in D3 and D3-N5 clones (Fig. 4). Based on previous studies showing that ES cells can differentiate to a serotonergic phenotype in vivo (Deacon et al., 1998; Björklund et al., 2002) or in vitro (Lee et al., 2000), ES cells were also examined using antibodies against 5HT following in vitro differentiation. The numbers of 5HT+ neurons in the D3-N2 and D3-N5 clones were 1.8- and 3.4-fold that seen in the D3-E5 clone, respectively (data not shown), suggesting the possibility that exogenous expression of Nurr1 may modestly increase the proportion of serotonergic neurons. One possible explanation for this observation is that increased expression of AADC may contribute to the higher level of serotonergic neurons as well as DA neurons. Finally, the glial cell population was analysed using specific antibodies for astrocytes (GFAP) and oligodendrocytes (GalC). Both glial cell populations could be detected in similar amounts in D3 and D3-N5 clones (Fig. 5g–j; other data not shown). Taken together, these data suggest that Nurr1 may have a relatively specific effect on ES cell differentiation into DA neurons.
Fig. 5.
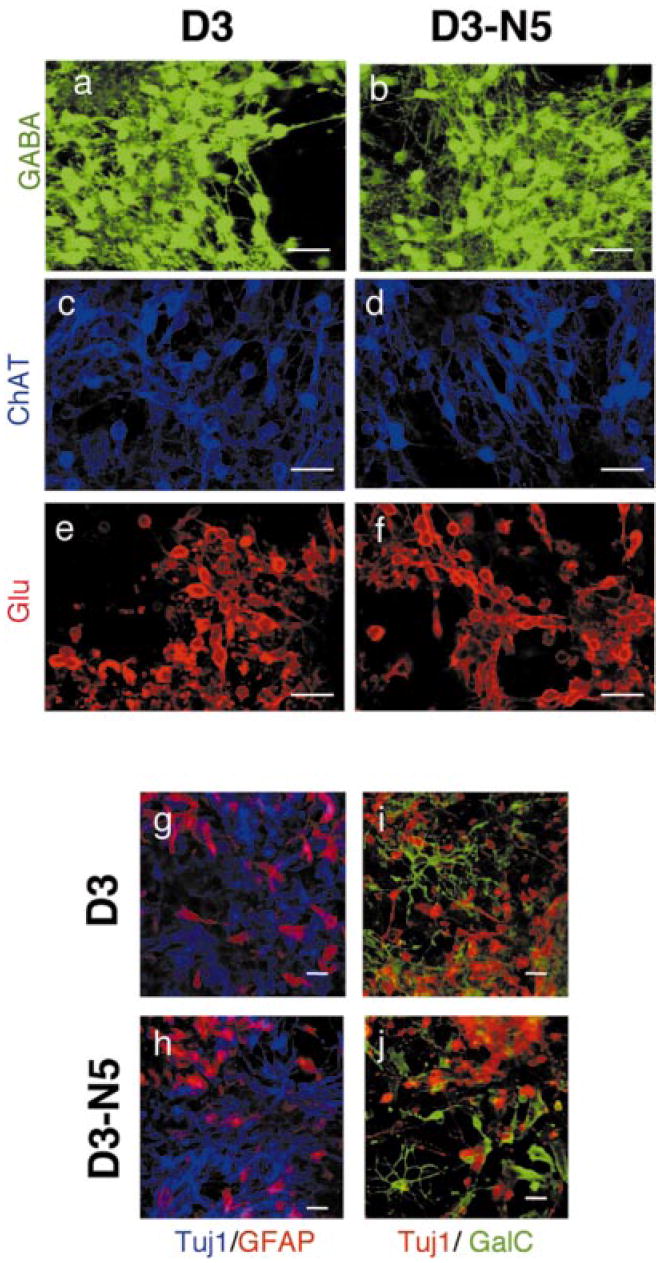
Nurr1 overexpression in ES cells does not influence the differentiation into neural phenotypes other than DA and serotonergic neurons. Detection of GABA (a and b), choline acetyltransferase ChAT; (c and d) and glutamate Glu; (e and f) positive neurons as well as β-tubulin and glial fibrillary acidic protein GFAP (g and h) or galactocerebrosidase C GalC (i and j) on in vitro differentiated D3 and D3-N5 cells. For double labelling with GFAP, mouse Tuj1 was used and for double labelling with GalC, rabbit antiβ-tubulin was used. All other primary antibodies are as described in materials and methods. For a-f, Rhodamine Red-X-labelled secondary antibodies are used, and for g-j, Cy2-labelling of secondary antibodies was shown as green, Rhodamine Red-X as red, and Cy5 as blue. Confocal microscopy at 40 × magnification. Scale bar, 25 μm.
Signalling molecules synergistically increase the differentiation of Nurr1-expressing ES cells to DA neurons
Several signalling molecules such as Sonic hedgehog (Shh) and FGF8 have been well characterized to facilitate DA differentiation (Ye et al., 1998; Lee et al., 2000). One interesting possibility is that Nurr1 acts downstream of one or more of these signalling pathways. Alternatively, Nurr1 may act independently of these pathways and, if so, may co-operatively facilitate induction of DA phenotypes. To address these possibilities, we sought to apply the recently optimized treatment of Shh, FGF8, and ascorbic acid (AA) (Lee et al., 2000) to our Nurr1-expressing clone. When these molecules were added to the wild type D3 cells, formation of TH+ neurons increased from 3.6 ± 1.3% to 11.1 ± 1.6% of the β-tubulin+ neurons (Fig. 6a). Therefore, while the proportion of TH+ neurons was lower than that reported by Lee et al., (2000), the amount of increase was similar (approximately 3-fold). The same treatment was next applied to the D3-N2 clone during in vitro differentiation. In this case, almost all cell clusters contained large numbers of TH+ neurons and the counting analysis showed that approximately 61.9 ± 3.9% of all β-tubulin+ neurons were TH+ (Fig. 6a). Thus, combined treatment of Shh, FGF8, and AA increased the proportion of D3-N2-derived TH+ neurons 4- to 5-fold, indicating that Nurr1 may facilitate DA neuron differentiation independently of these signal molecules.
Fig. 6.
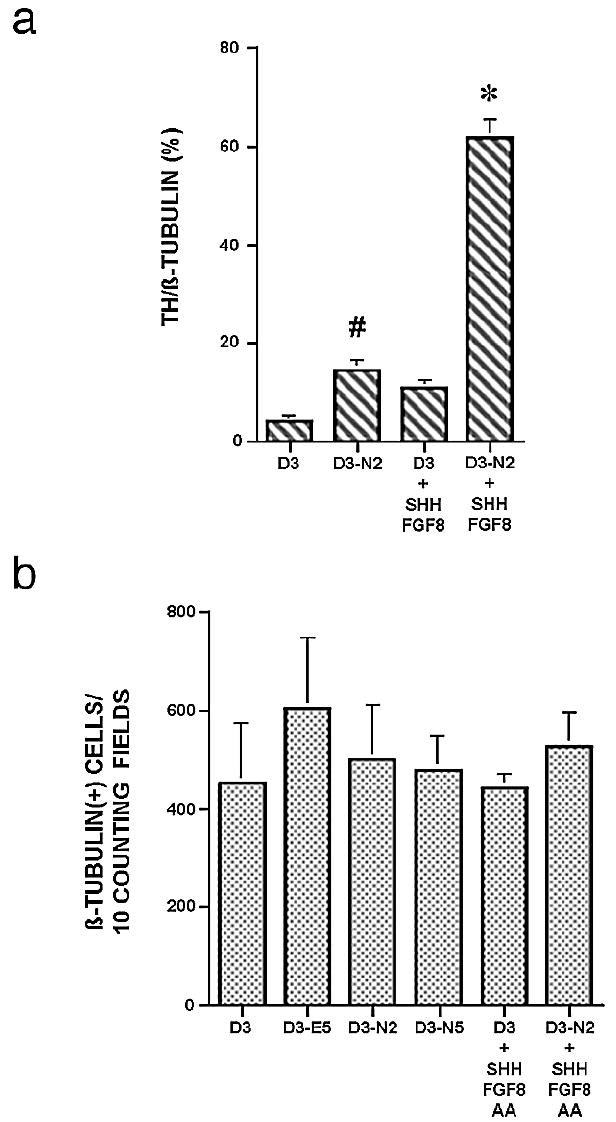
(a) Induction of TH+ neurons by Nurr1 in the presence of signalling molecules. D3 or D3-N2 cells were differentiated in vitro with or without signalling molecules (Shh, FGF8, and AA). TH+ neurons were quantified as described in Materials and methods and in Fig. 2g. Each group represents an average of 3–5 samples from 2 to 3 independent experiments. anova revealed F = 68.5, P < 0.0001. Fisher’s PLSD posthoc analysis was performed with a significance level of 0.05. #P < 0.05 compared with D3 cells. *P < 0.05 compared with all other groups. b. Quantification of β-tubulin+ cells. Shown are the numbers of total β-tubulin+ cells from 10 random fields per each sample (n = 4–5). anova revealed F = 0.296, P = 0.909, showing that there is no significant difference among group means.
To address whether Nurr1 and/or these signalling molecules may increase the proportion of TH+ neurons by facilitating neurogenesis, the numbers of β-tubulin+ neurons for 10 fields of each sample (n = 4–5) of different ES cell clones were counted, with or without the addition of signalling molecules. As shown in Fig. 6b, Nurr1 overexpression, signalling molecules, or their combination did not obviously increase the formation of β-tubulin+ neurons.
Nurr1-expressing ES cell-derived neurons efficiently produce and release DA in response to membrane depolarization
An important physiological aspect of authentic DA neuron phenotypes is the ability to synthesize DA and release it in response to membrane depolarization. When ES cells were harvested at time points earlier than the ND stage (day 5) and assayed by HPLC, DA was not detectable in the cell pellets (data not shown) or in the media after KCl depolarization (Fig. 7), 0.00 ± 0.00, 0.00 ± 0.00, 0.01 ± 0.01 pg/μg cellular proteins, for D3, D3-E5, and D3-N2, respectively. In contrast, when fully in vitro differentiated ES cell clones (day 11 of the ND stage) were assayed, total DA contents within the cells were 0.54 ± 0.24, 0.37 ± 0.15, and 1.11 ± 0.15 pg/μg cellular proteins, for D3, D3-E5, and D3-N2, respectively (n = 3, P < 0.001). Following these results it was tested whether DA can be released following membrane depolarisation, after treatment of fully differentiated ES cell clones (day 11 of ND stage) with 50 mm KCl, the released DA in the media was 0.31 ± 0.09, 0.21 ± 0.11, and 0.68 ± 0.10 pg/μg cellular proteins for D3, D3-E5, and D3-N2, respectively (Fig. 7). These data demonstrate that neurons derived from Nurr1-overexpressing ES clones have the enhanced ability to produce and release DA as compared to naïve ES cells. Taken together, one may conclude that exogenous Nurr1 expression in ES cells increases the efficiency of generating phenotypically and functionally mature midbrain DA neurons in vitro.
Fig. 7.
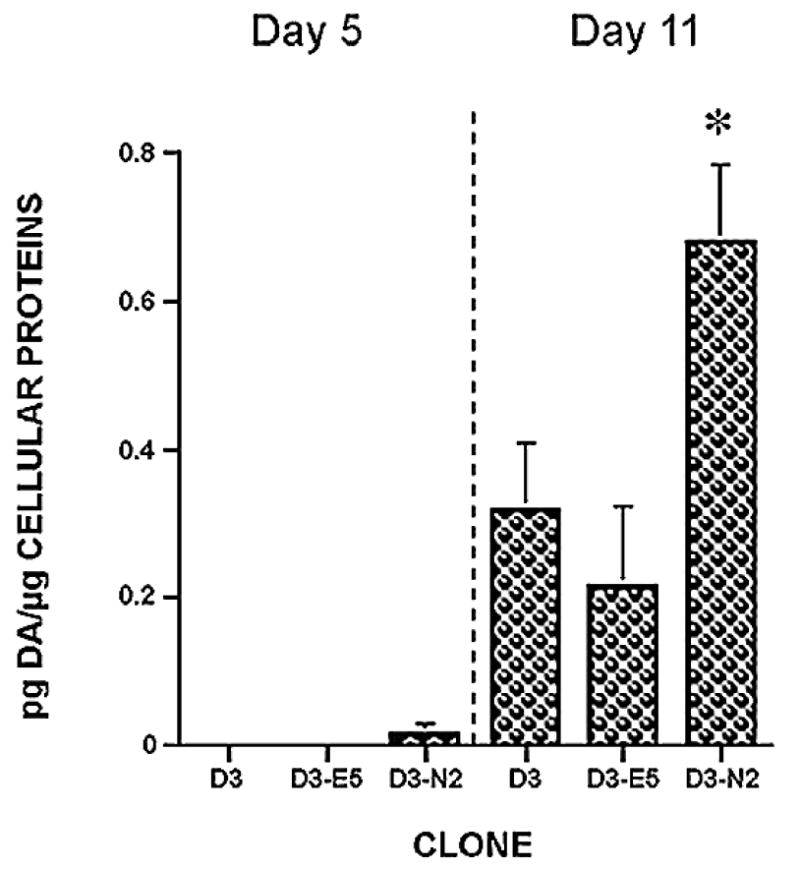
Analysis of dopamine release. In vitro differentiated ES cell clones were challenged with 50 mm KCl, and the media assayed for DA levels by reverse phase HPLC. DA release from ES cell clone-derived DA neurons was measured at day 5 and day 11 of ND stage. Each group represents an average of 3–4 samples from independent experiments. anova revealed F = 14.19, P < 0.0001. Fisher’s PLSD posthoc analysis was performed with a significance level of 0.05. *P < 0.05 compared with all other samples.
Discussion
The current study analysed the effect of Nurr1 on uncommitted pluripotent mouse ES cells and showed that overexpression of this transcription factor enhances the development of these cells into midbrain DA neurons. The effect of Nurr1 expression in ES cells on differential gene expression during cell development was also examined. It was found that there was selective up-regulation of the midbrain DA-specific markers TH, AADC, DAT, Ptx3, calbindin, and AHD2 in Nurr1-expressing ES cell clones (Figs 3 and 4). While TH and AADC mediate the first and last step of DA biosynthesis, respectively, DAT is responsible for DA reuptake into the presynaptic terminal. AADC is also expressed in serotonergic neurons, mediating the last step of serotonin biosynthesis. Ptx3 is one of the most selective midbrain DA markers (Smidt et al., 1997) and is a putative transcription factor whose function is unknown. AHD2 has been defined as the earliest marker identifiable in developing DA cells (Wallen et al., 1999). Some of these markers were either absent or significantly down-regulated in midbrain DA neurons of Nurr1-null mice (Zetterstrom et al., 1997; Castillo et al., 1998; Saucedo-Cardenas et al., 1998; Wallen et al., 1999; Witta et al., 2000). However, it was not clear whether this reduction was due to the direct action of Nurr1 or defective neurogenesis. The gene expression analysis, using a gain of function approach, demonstrated the importance of Nurr1 for the expression of these genes in developing ES cells. A clear up-regulation of AADC, Ptx3 and AHD2 gene expression was observed in the Nurr1-transgenic ES clones, compared to the D3 or D3-E5 clones during in vitro differentiation (Fig. 4 and other data not shown). Interestingly, the markers for the mature DA phenotype such as TH and DAT were expressed only at the fully differentiated stage. Collectively, this data suggests that Nurr1 is involved in the induction and/or maintenance of expression of various midbrain DA neuronal markers.
The observation that TH gene induction occurs after attainment of a mature neuronal phenotype indicates that Nurr1 alone is not sufficient for activating TH gene expression, but rather requires specific cellular (or neuronal) environments or cofactors. Such cofactors or conditions may be present in Nurr1-overexpressing adult hippocampal precursors (Sakurada et al., 1999), or provided by mesencephalic astrocytes to allow TH induction in C17.2 neuronal precursors (Wagner et al., 1999). In addition, we and others have recently demonstrated that Nurr1 can directly activate the TH promoter in a cell line-specific manner (Sakurada et al., 1999; Chung et al., 2000; Iwawaki et al., 2000; Jensen & O’Malley, 2001), supporting the view that certain transcriptional conditions are required for TH gene induction by Nurr1.
Up-regulation of DAT gene expression was observed in the fully differentiated Nurr1-transgenic cells. Whereas few DAT+ cells could be detected after neuronal differentiation of naïve D3 cells (data not shown), many DAT+ neurons were detected among the TH+ cell population of differentiated D3-Nurr1 cells (Fig. 3). This was consistent with the RT-PCR analysis demonstrating that expression of DAT mRNA was significantly up-regulated in D3-Nurr1 cells after differentiation. Nurr1 has been shown to enhance the transcriptional activity of the human DAT gene (Sacchetti et al., 1999), and in addition, DAT up-regulation in vivo depends on interaction between presynaptic DA fibres and target striatal neurons (Perrone-Capano et al., 1994; Perrone-Capano et al., 1996). While only some of the TH+ cells expressed DAT in these experiments, it might be possible to increase the proportion of DAT expressing cells by providing target cells. In support of this, it has previously been shown that even in naive D3 cells, there is abundant DAT expression after in vivo transplantation into striatum of rodent PD model animals (Björklund et al., 2002).
Various signalling molecules have been extensively tested for their potential to facilitate DA lineage differentiation using ES and progenitor cells (Ye et al., 1998; Kawasaki et al., 2000; Lee et al., 2000; Rolletschek et al., 2001). Among these factors, we tested if the combined treatment of Shh, FGF8, and AA (Lee et al., 2000) can increase formation of TH+ neurons in the Nurr1-transduced ES cell clone. This treatment increased formation of TH+ neurons in D3 ES cells approximately 3-fold (from 3.6 ± 1.3% to 11.1 ± 1.6%; Fig. 6a). When the same treatment was applied to the D3-N2 clone during in vitro differentiation, formation of TH+ neurons was robustly increased. In this condition, most of neuronal clusters contained large numbers of TH+ neurons and our counting analysis showed that approximately 61.9 ± 3.9% of all β-tubulin+ neurons were TH+ (Fig. 6a). This observation therefore does not support the idea that Nurr1 may act downstream of one of these signalling molecules, but rather, suggests that Nurr1 and these molecules may act independently or synergistically to induce the DA lineage differentiation.
Different mechanisms may underlie the increase of TH+ neurons by Nurr1 overexpression and/or signalling molecules. First, they may facilitate neurogenesis at the level of formation of neuroprogenitor cells or neurons, the results described here show that the numbers of β-tubulin+ neurons were not changed in any of ES cells and therefore do not support this possibility (Fig. 6b). Consistent with this, analyses of Nurr1-deficient mice also indicated that Nurr1 acts at the later stage of DA differentiation (Zetterstrom et al., 1997; Castillo et al., 1998; Saucedo-Cardenas et al., 1998; Wallen et al., 1999; Witta et al., 2000). Another possible mechanism by which Nurr1 can increase DA differentiation is by influencing DA cell populations at the cell survival and/or death level. In Nurr1-null mice, Saucedo-Cardenas et al., (1998) observed increased apoptosis and decreased cell numbers in the midbrain of Nurr1-null neonates, indicating that Nurr1 is important in the survival of midbrain DA neurons. This possibility is consistent with the observation that Nurr1 critically regulates expression of Ret, an important receptor for glial cell line-derived and related neurotrophic factors (Wallen et al., 2001). On the other hand, Witta et al., (2000) did not observe such an effect in their Nurr1-null mice. Thus, at this point in time, the effect of Nurr1 on DA cell survival is not clear. The data reported here does not address these possibilities, as cell survival or death was not measured directly in this culture system. It is thus an open possibility that Nurr1 may have pleiotropic effects on DA gene induction and cell survival, thereby enhancing the fraction of DA neurons in our in vitro system.
It appears that exogenous expression of Nurr1 has a relatively specific effect on DA lineage differentiation. When other neurotransmitter subtypes such as GABA, cholinergic, noradrenergic, and glutamatergic marker proteins were analysed, formation of these neuronal subtypes was apparently unaltered in Nurr1-transduced ES cells. In support of this, semiquantitative RT-PCR analysis confirmed that expression of the neuronal glutamate transporter and DBH genes, markers of glutamatergic and noradrenergic neurons, respectively, was not changed in the D3 and D3-N5 clones (Fig. 4). In addition, Nurr1 had no detectable effect on the formation of glial cell populations such as astrocytes and oligodendrocytes, suggesting that Nurr1 may specifically influence the DA neuron lineage. One possible exception may be the number of serotonergic neurons, which seem to have been modestly increased by Nurr1 overexpression. A possible explanation for this observation is that AADC, the final step for both DA and serotonin synthesis, was up-regulated in Nurr1-overexpressing clones in our study (Fig. 4), possibly contributing to the appearance of 5HT+ neurons. However, as only one Nurr1-expressing clone (out of two) significantly elevated 5HT+ neuronal numbers, this observation should be considered preliminary and awaits further investigation.
Several lines of evidence suggest that the TH+ neurons observed in this study have the midbrain DA neuronal phenotype. First, double-immunostaining analysis showed that most of TH+ neurons coexpress other midbrain DA neuron markers such as AADC and calretinin. Additionally, some of these cells also coexpressed DAT, another essential marker protein of mature DA neurons, and none of them expressed DBH, a noradrenergic marker. Notably, most of these TH+/calretinin+ cells did not express GABA (Fig. 3m–o), suggesting that these neurons have the midbrain, but not olfactory, DA phenotype. Furthermore, these TH+ cells did not coexpress other neurotransmitters such as 5HT (Fig. 3p–r). Finally, a critical criterion of authentic DA neuron phenotypes is the ability to synthesize DA and release it in response to membrane depolarization. The HPLC analysis of in vitro differentiated ES cells showed that significant amounts of DA were produced and, then, released in response to membrane depolarization by 50 mm KCl (Fig. 7), strongly suggesting that Nurr1 overexpression facilitates the differentiation of ES cells to the authentic midbrain DA neuron lineage.
In vitro differentiation of the genetically modified ES cells to the DA lineage can be further increased, when combined with the induction procedure using several signalling molecules (Ye et al., 1998; Hynes & Rosenthal, 1999, 2000; Kawasaki et al., 2000; Lee et al., 2000; Caldwell et al., 2001; Rolletschek et al., 2001). Two groups have recently shown that human ES cells can differentiate into neuronal cell types in vitro (Reubinoff et al., 2001; Studer, 2001; Zhang et al., 2001). However, generation of DA neurons was very limited in both studies. Therefore, it is of great interest to determine whether the genetic and inductive manipulations described here could be applied to improve the generation of DA neurons in human ES cells. Previous work has also demonstrated that transplantation of naïve ES cells in a rat model of PD resulted in spontaneous differentiation into functional DA neurons partially restoring behavioural deficits (Deacon et al., 1998; Björklund et al., 2002). Given the increased yield of DA neurons in Nurr1-transgenic ES cells, it may be to evaluate the function of these genetically modified cells in rodent and primate models of PD. In parallel with the above findings, McKay and colleagues similarly report that DA neurons can be efficiently generated from Nurr1-overexpressing embryonic stem cells and showed that grafting of these in vitro differentiated DA neurons resulted in functional recovery in a rodent model of PD (Kim et al., 2002).
Taken together, these results demonstrate that ES cells can be genetically manipulated by key cell fate-determining transcription factor(s) to facilitate their differentiation into specific cell lineages.
Acknowledgments
We thank Orla M. Conneely at Baylor College of Medicine for the Nurr1-expression plasmid pSV40Nurr1 and Devanand Kowlessur for his expert technical help with HPLC analysis. This work was supported by NIH grants (P50)NS39793 (to OI and KSK), MH48866 (to KSK), NS32080 (UJK), and DAMD-17-01-1-0763 (KSK), DAMD-17-01-1-0762 (OI), and Kinetics Foundation and Parkinson Foundation National Capital Area (OI). Most results of this study were previously presented at the Annual Conference of the Society for Neuroscience, 2001, 245.1, http://sfn.scholarone.com/itin2001/.
Abbreviations
- AA
ascorbic acid
- AADC
aromatic l-amino acid decarboxylase
- AHD2
aldehyde dehydrogenase 2
- anova
analysis of variance
- ChAT
choline acetyltransferase
- DA
dopaminergic
- DAT
dopamine transporter
- DBH
dopamine β-hydroxylase
- EB
embryoid body
- EF
elongation factor-1α
- ES cells
embryonic stem cells
- FGF
fibroblast growth factor
- GABA
gamma aminobutyric acid
- GalC
galactocerebrosidase C
- GFAP
glial fibrillary acidic protein
- Glu
glutamate
- HPLC
high performance liquid chromatography
- 5HT
serotonin
- LIF
leukaemia inhibitory factor
- ND
neuronal differentiation
- NDS
normal donkey serum
- NP
neuronal precursor
- PCA
perchloric acid
- PD
Parkinson’s disease
- SEM
standard error of mean
- Shh
sonic hedgehog
- TH
tyrosine hydroxylase
References
- Backman C, Perlmann T, Wallen A, Hoffer BJ, Morales M. A selective group of dopaminergic neurons express Nurr1 in the adult mouse brain. Brain Res. 1999;851:125–132. doi: 10.1016/s0006-8993(99)02149-6. [DOI] [PubMed] [Google Scholar]
- Björklund L, Sánchez-Pernaute R, Chung S, Andersson T, Chen I, McNaught K, Brownell A, Jenkins B, Wahlestedt C, Kim K-S, Isacson O. Embryonic stem cells develop into functional dopaminergic neurons after transplantation in a Parkinson rat model. Proc Natl Acad Sci USA. 2002;99:2344–2349. doi: 10.1073/pnas.022438099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford MM. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Brustle O, Jones K, Learish R, Karram K, Choudhary K, Wiestler O, Duncan I, McKay R. Embryonic stem cell-derived glial precursors: a source of myelinating transplants. Science. 1999;285:754–756. doi: 10.1126/science.285.5428.754. [DOI] [PubMed] [Google Scholar]
- Caldwell MA, He X, Wilkie N, Pollack S, Marshall G, Wafford KA, Svendsen CN. Growth factors regulate the survival and fate of cells derived from human neurospheres. Nature Biotechnol. 2001;19:475–479. doi: 10.1038/88158. [DOI] [PubMed] [Google Scholar]
- Castillo SO, Baffi JS, Palkovits M, Goldstein DS, Kopin IJ, Witta J, Magnuson MA, Nikodem VM. dopamine biosynthesis is selectively abolished in substantia nigra/ventral tegmental area but not in hpothalamic neurons in mice with targeted disruption of the Nurr1 gene. Mol Cell Neurosci. 1998;11:36–46. doi: 10.1006/mcne.1998.0673. [DOI] [PubMed] [Google Scholar]
- Chung S, Andersson T, Sonntag K-C, Björklund L, Isacson O, Kim KS. Analysis of different promoter systems for efficient transgene expression in mouse embryonic stem cell lines. Stem Cells. 2002;20:139–145. doi: 10.1634/stemcells.20-2-139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung S, Kim CH, Andersson T, Isacson O, Kim KS. The nuclear orphan receptor Nurr1 binds and transactivates the promoter of the tyrosine hydroxylase but not that of the dopamine β-hydroxylase gene. Soc Neurosci Abstr. 2000;26:19.1. [Google Scholar]
- Deacon T, Dinsmore J, Costantini LC, Ratliff J, Isacson O. Blastula-derived stem cells can differentiate into dopaminergic and serotonergic neurons after transplantation. Exp Neurol. 1998;149:28–41. doi: 10.1006/exnr.1997.6674. [DOI] [PubMed] [Google Scholar]
- Desbaillets I, Ziegler U, Groscurth P, Gassmann M. Embryoid bodies: an in vitro model of mouse embryogenesis. Exp Physiol. 2000;85:645–651. [PubMed] [Google Scholar]
- Dinsmore J, Ratliff J, Deacon T, Pakzaban P, Jacoby D, Galpern W, Isacson O. Embryonic stem cells differentiated in vitro as a novel source of cells for transplantation. Cell Transplantation. 1996;5:131–143. doi: 10.1177/096368979600500205. [DOI] [PubMed] [Google Scholar]
- Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292:154–156. doi: 10.1038/292154a0. [DOI] [PubMed] [Google Scholar]
- Goridis C, Brunet J-F. Transcriptional control of neurotransmitter phenotype. Curr Opin Neurobiol. 1999;9:47–53. doi: 10.1016/s0959-4388(99)80006-3. [DOI] [PubMed] [Google Scholar]
- Hooper M, Hardy K, Handyside A, Hunter S, Monk M. HPRT-deficient (Lesch-Nyhan) mouse embryos derived from germline colonization by cultured cells. Nature. 1987;326:292–295. doi: 10.1038/326292a0. [DOI] [PubMed] [Google Scholar]
- Hynes M, Rosenthal A. Specification of dopaminergic and serotonergic neurons in the vertebrate CNS. Curr Opin Neurobiol. 1999;9:26–36. doi: 10.1016/s0959-4388(99)80004-x. [DOI] [PubMed] [Google Scholar]
- Hynes M, Rosenthal A. Embryonic stem cells go dopaminergic. Neuron. 2000;28:11–14. doi: 10.1016/s0896-6273(00)00079-9. [DOI] [PubMed] [Google Scholar]
- Iwawaki T, Kohno K, Kobayashi K. Identification of a potential nurr1 response element that activates the tyrosine hydroxylase gene promoter in cultured cells. Biochem Biophys Res Commun. 2000;274:590–595. doi: 10.1006/bbrc.2000.3204. [DOI] [PubMed] [Google Scholar]
- Jensen PJ, O’Malley K. Transactivation of the tyrosine hydrosylase gene by Nurr1 is dependent upon specific cellular factors. Soc Neurosci Abstr. 2001;27:588.17. [Google Scholar]
- Johe KK, Hazel TG, Muller T, Dugich-Djordjevic MM, McKay RD, Vicario-Abejon C, Collazo D. Single factors direct the differentiation of stem cells from the fetal and adult central nervous system. Functions of basic fibroblast growth factor and neurotrophins in the differentiation of hippocampal neurons. Genes Dev. 1996;10:3129–3140. doi: 10.1101/gad.10.24.3129. [DOI] [PubMed] [Google Scholar]
- Kawasaki H, Mizuseki K, Nishikawa S, Kaneko S, Kuwana Y, Nakanishi S, Nishikawa SI, Sasai Y. Induction of midbrain dopaminergic neurons from ES cells by stromal cell-derived inducing activity. Neuron. 2000;28:31–40. doi: 10.1016/s0896-6273(00)00083-0. [DOI] [PubMed] [Google Scholar]
- Kim JH, Auerbach JM, Rodriguez-Gomez JA, Velasco I, Gavin D, Lumelsky N, Lee SH, Nguyen J, Sanchez-Pernaute R, Bankiewicz K, McKay R. Dopamine neurons derived from embryonic stem cells function in an animal model of Parkinson’s disease. Nature. 2002;418:50–56. doi: 10.1038/nature00900. [DOI] [PubMed] [Google Scholar]
- Law SW, Conneely OM, DeMayo FJ, O’Malley BW. Identification of a new brain-specific transcription factor, Nurr1. Mol Endocrinol. 1992;6:2129–2135. doi: 10.1210/mend.6.12.1491694. [DOI] [PubMed] [Google Scholar]
- Lee SH, Lumelsky N, Studer L, Auerbach JM, McKay RD. Efficient generation of midbrain and hindbrain neurons from mouse embryonic stem cells. Nature Biotechnol. 2000;18:675–679. doi: 10.1038/76536. [DOI] [PubMed] [Google Scholar]
- Lumelsky N, Blondel O, Laeng P, Velasco I, Ravin R, McKay R. Differentiation of embryonic stem cells to insulin-secreting structures similar to pancreatic islets. Science. 2001;292:1389–1394. doi: 10.1126/science.1058866. [DOI] [PubMed] [Google Scholar]
- Martin GR. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci USA. 1981;78:7634–7638. doi: 10.1073/pnas.78.12.7634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murphy EP, Dobson AD, Keller C, Conneely OM. Differential regulation of transcription by the NURR1/NUR77 subfamily of nuclear transcription factors. Gene Expr. 1996;5:169–179. [PMC free article] [PubMed] [Google Scholar]
- Nagy A, Gozca E, Diaz EM, Prideaux VR, Ivanyi E, Markkula M, Rossant J. Embryonic stem cells alone are able to support fetal development in the mouse. Development. 1990;110:815–821. doi: 10.1242/dev.110.3.815. [DOI] [PubMed] [Google Scholar]
- Nagy A, Rossant J, Nagy R, Abramow-Newerly W, Roder JC. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc Natl Acad Sci USA. 1993;90:8424–8428. doi: 10.1073/pnas.90.18.8424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nemoto C, Hida T, Arai R. Calretinin and calbindin-D28k in dopaminergic neurons of the rat midbrain: a triple-labeling immunohistochemical study. Brain Res. 1999;846:129–136. doi: 10.1016/s0006-8993(99)01950-2. [DOI] [PubMed] [Google Scholar]
- Okabe S, Forsberg-Nilsson K, Spiro AC, Segal M, McKay RD. Development of neuronal precursor cells and functional postmitotic neurons from embryonic stem cells in vitro. Mech Dev. 1996;59:89–102. doi: 10.1016/0925-4773(96)00572-2. [DOI] [PubMed] [Google Scholar]
- Perrone-Capano C, Tino A, Amadoro G, Pernas-Alonso R, di Porzio U. Dopamine transporter gene expression in rat mesencephalic dopaminergic neurons is increased by direct interaction with target striatal cells in vitro. Brain Res Mol Brain Res. 1996;39:160–166. doi: 10.1016/0169-328x(96)00022-8. [DOI] [PubMed] [Google Scholar]
- Perrone-Capano C, Tino A, Di Porzio U. Target cells modulate dopamine transporter gene expression during brain development. Neuroreport. 1994;5:1145–1148. doi: 10.1097/00001756-199405000-00031. [DOI] [PubMed] [Google Scholar]
- Reubinoff BE, Itsykson P, Turetsky T, Pera MF, Reinhartz E, Itzik A, Ben-Hur T. Neural progenitors from human embryonic stem cells. Nature Biotechnol. 2001;19:1134–1140. doi: 10.1038/nbt1201-1134. [DOI] [PubMed] [Google Scholar]
- Rolletschek A, Chang H, Guan K, Czyz J, Meyer M, Wobus AM. Differentiation of embryonic stem cell-derived dopaminergic neurons is enhanced by survival-promoting factors. Mech Dev. 2001;105:93–104. doi: 10.1016/s0925-4773(01)00385-9. [DOI] [PubMed] [Google Scholar]
- Sacchetti P, Brownschidle LA, Granneman JG, Bannon MJ. Characterization of the 5′-flanking region of the human dopamine transporter gene. Brain Res Mol Brain Res. 1999;74:167–174. doi: 10.1016/s0169-328x(99)00275-2. [DOI] [PubMed] [Google Scholar]
- Sakurada K, Ohshima-Sakurada M, Palmer TD, Gage FH. Nurr1, an orphan nuclear receptor, is a transcriptional activator of endogenous tyrosine hydroxylase in neural progenitor cells derived from the adult brain. Development. 1999;126:4017–4026. doi: 10.1242/dev.126.18.4017. [DOI] [PubMed] [Google Scholar]
- Saucedo-Cardenas O, Quintana-Hau JD, Le W-D, Smidt MP, Cox JJ, De Mayo F, Burbach JPH, Conneely OM. Nurr1 is essential for the induction of the dopaminergic phenotype and the survival of ventral mesencephalic late dopaminergic precursor neurons. Proc Natl Acad Sci USA. 1998;95:4013–4018. doi: 10.1073/pnas.95.7.4013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye W, Shimamura K, Rubenstein JL, Hynes MA, Rosenthal A. FGF and Shh signals control dopaminergic and serotonergic cell fate in the anterior neural plate. Cell. 1998;93:755–766. doi: 10.1016/s0092-8674(00)81437-3. [DOI] [PubMed] [Google Scholar]
- Smidt MP, van Schaick HS, Lanctot C, Tremblay JJ, Cox JJ, van der Kleij AA, Wolterink G, Drouin J, Burbach JP. A homeodomain gene Ptx3 has highly restricted brain expression in mesencephalic dopaminergic neurons. Proc Natl Acad Sci USA. 1997;94:13305–13310. doi: 10.1073/pnas.94.24.13305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith AG. Culture and differentiation of embryonic stem cells. J Tiss Cult Meth. 1991;13:89–94. [Google Scholar]
- Studer L. Stem cells with brainpower. Nature Biotechnol. 2001;19:1117–1118. doi: 10.1038/nbt1201-1117. [DOI] [PubMed] [Google Scholar]
- Tanaka K. Cloning and expression of a glutamate transporter from mouse brain. Neurosci Lett. 1993;159:183–186. doi: 10.1016/0304-3940(93)90829-a. [DOI] [PubMed] [Google Scholar]
- Thomas KR, Capecchi MR. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell. 1987;51:503–512. doi: 10.1016/0092-8674(87)90646-5. [DOI] [PubMed] [Google Scholar]
- Wachtel SR, Bencsics C, Kang UJ. Role of aromatic 1-amino acid decarboxylase for dopamine replacement by genetically modified fibroblasts in a rat model of Parkinson’s disease. J Neurochem. 1997;69:2055–2063. doi: 10.1046/j.1471-4159.1997.69052055.x. [DOI] [PubMed] [Google Scholar]
- Wagner J, Akerud P, Castro DS, Holm PC, Snyder EY, Perlmann T, Arenas E. Induction of a midbrain dopaminergic phenotype in Nurr1-overexpressing neural stem cells by tytpe1 astrocytes. Nature Biotechnol. 1999;17:653–659. doi: 10.1038/10862. [DOI] [PubMed] [Google Scholar]
- Wallen AA, Castro DS, Zetterstrom RH, Karlen M, Olson L, Ericson J, Perlmann T. Orphan nuclear receptor Nurr1 is essential for Ret expression in midbrain dopamine neurons and in the brain stem. Mol Cell Neurosci. 2001;18:649–663. doi: 10.1006/mcne.2001.1057. [DOI] [PubMed] [Google Scholar]
- Wallen A, Zetterstrom RH, Solomin L, Arvidsson M, Olson L, Perlmann T. Fate of mesencephalic AHD2-expressing dopamine progenitor cells in NURR1 mutant mice. Exp Cell Res. 1999;253:737–746. doi: 10.1006/excr.1999.4691. [DOI] [PubMed] [Google Scholar]
- Witta J, Baffi JS, Palkovits M, Mezey E, Castillo SO, Nikodem VM. Nigrostriatal innervation is preserved in Nurr1-null mice, although dopaminergic neuron precursors are arrested from terminal differentiation. Brain Res Mol Brain Res. 2000;84:67–78. doi: 10.1016/s0169-328x(00)00211-4. [DOI] [PubMed] [Google Scholar]
- Wutz A, Jaenisch R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol Cell. 2000;5:695–705. doi: 10.1016/s1097-2765(00)80248-8. [DOI] [PubMed] [Google Scholar]
- Zetterstrom RH, Solomin L, Jansson L, Hoffer BJ, Olson L, Perlmann T. Dopamine neuron agenesis in Nurr1-deficient mice. Science. 1997;276:248–250. doi: 10.1126/science.276.5310.248. [DOI] [PubMed] [Google Scholar]
- Zetterstrom RH, Williams R, Perlmann T, Olson L. Cellular expression of the immediate early transcription factors Nurr1 and NGFI-B suggests a gene regulatory role in several brain regions including the nigrostriatal dopamine system. Mol Brain Res. 1996;41:111–120. doi: 10.1016/0169-328x(96)00074-5. [DOI] [PubMed] [Google Scholar]
- Zhang S, Wernig M, Duncan ID, Brüstle O, Thomson JA. In vitro differentiation of transplantable neural precursors from human embryonic stem cells. Nature Biotechnol. 2001;19:1129–1133. doi: 10.1038/nbt1201-1129. [DOI] [PubMed] [Google Scholar]


