Abstract
We have previously shown that Fhit tumor suppressor protein interacts with Hsp60 chaperone machinery and ferredoxin reductase (Fdxr) protein. Fhit-effector interactions are associated with a Fhit-dependent increase in Fdxr stability, followed by generation of reactive oxygen species and apoptosis induction under conditions of oxidative stress. To define Fhit structural features that affect interactions, downstream signaling, and biological outcomes, we used cancer cells expressing Fhit mutants with amino acid substitutions that alter enzymatic activity, enzyme substrate binding, or phosphorylation at tyrosine 114. Gastric cancer cell clones stably expressing mutants that do not bind substrate or cannot be phosphorylated showed decreased binding to Hsp60 and Fdxr and reduced mitochondrial localization. Expression of Fhit or mutants that bind interactor proteins results in oxidative damage and accumulation of cells in G2/M or sub-G1 fractions after peroxide treatment; noninteracting mutants are defective in these biological effects. Gastric cancer clones expressing noncomplexing Fhit mutants show reduction of Fhit tumor suppressor activity, confirming that substrate binding, interaction with heat shock proteins, mitochondrial localization, and interaction with Fdxr are important for Fhit tumor suppressor function.
Fhit protein is a powerful tumor suppressor that is frequently lost or reduced in cancer cells because of rearrangement of the exquisitely DNA damage-sensitive fragile FHIT gene. Restoration of Fhit expression suppresses tumorigenicity of cancer cells of various types, and the ability to induce apoptosis in cancer cells in vitro is reduced by specific Fhit mutations (1, 2).
Through studies of signal pathways affected by Fhit expression, by searches for Fhit protein effectors, and by in vitro analyses of Fhit activity, we and others have defined Fhit enzymatic activity in vitro (3), apoptotic activity in cells and tumors (4–6), and most recently identification of a Fhit protein complex that affects Fhit stability, mitochondrial localization, and interaction with ferredoxin reductase (Fdxr)5 (7). The complex includes Hsp60 and Hsp10 that mediate Fhit stability and may affect import into mitochondria, where Fhit interacts with Fdxr, which is responsible for transferring electrons from NADPH to cytochrome P450 via ferredoxin. Virally mediated Fhit restoration in Fhit-deficient cancer cells increases production of intracellular reactive oxygen species (ROS), followed by increased apoptosis of cancer cells under oxidative stress conditions; conversely, Fhit-negative cells escape apoptosis, likely carrying oxidative DNA damage that contributes to accumulation of mutations.
The Fhit protein sequence, showing high homology to the histidine triad (HIT) family of proteins, suggested that the protein product would hydrolyze diadenosine tetraphosphate or diadenosine triphosphate (Ap3A) (8), and in vitro studies showed that Ap3A was cleaved into ADP and AMP by Fhit. The catalytic histidine triad within Fhit was essential for catalytic activity (3), and a Fhit mutant that substituted Asn for His at the central histidine (H96N mutant) was catalytically inactive, although it bound substrate well (3). Early tumor suppression studies showed that cancer cells stably transfected with wild type (WT) or H96N mutant Fhit were suppressed for tumor growth in nude mice. This suggested the hypothesis that the Fhit-substrate complex sends the tumor suppression signal (9, 10). To test this hypothesis, a series of FHIT alleles was designed to reduce substrate-binding and/or hydrolytic rates and was characterized by quantitative cell-death assays on cancer cells virally infected with each allele. The allele series covered defects as great as 100,000-fold in kcat and increases as large as 30-fold in Km. Mutants with 2–7-fold increases in Km had significantly reduced apoptotic indices and the mutant with a 30-fold increase in Km retained little apoptotic function. Thus, the proapoptotic function of Fhit, which is likely associated with tumor suppressor function, is limited by substrate binding and is unrelated to substrate hydrolysis (11).
Fhit, a homodimeric protein of 147 amino acids, is a target of tyrosine phosphorylation by the Src family protein kinases, which can phosphorylate Tyr-114 of Fhit in vitro and in vivo (12). After co-expression of Fhit with the Elk tyrosine kinase in Escherichia coli to generate phosphorylated forms of Fhit, unphosphorylated, mono-, and diphosphorylated Fhit were purified, and enzyme kinetics studies showed that monophosphorylated Fhit exhibited monophasic kinetics with Km and kcat values ∼2- and ∼7-fold lower, respectively, than for unphosphorylated Fhit. Diphosphorylated Fhit exhibited biphasic kinetics; one site had Km and kcat values ∼2- and ∼140-fold lower, respectively, than for unphosphorylated Fhit; the second site had a Km ∼60-fold higher and a kcat ∼6-fold lower than for unphosphorylated Fhit (13). Thus, it was possible that the alterations in Km and kcat values for phosphorylated forms of Fhit might favor formation and lifetime of the Fhit-Ap3A complex and enhance tumor suppressor activity (see Table 1 for characteristics of specific Fhit forms).
TABLE 1.
Characteristics of Fhit and Fhit mutants An increase in Km means a decrease in substrate binding; a decrease in kcat means a decrease in substrate hydrolysis. Sub-G1 fractions were determined in different cancer cells using different methods of exogenous Fhit expression, and a representative result is included in the table. The apoptosis column represents results obtained after treatment of stable MKN74 transfectants with H2O2; NT, not tested; NA, not applicable because we cannot study phosho-Fhit forms in vivo due to instability; –/+ or weak, +/– some activity; Cyt & mito, cytosolic fraction and mitochondrial fraction.
|
Fhit forms
|
Kinetic parameters
|
% Sub-G1
|
Direct binding
|
Subcellular location
|
Co-IP in vivo
|
8-OHdG
|
Apoptosis
|
Tumor suppressor
|
||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Km (mm) | kcat (s–1) | A549 | MKN74 | Hsp60 | Fdxr | Hsp60 | Fdxr | |||||
| Fhit WT | 1.6 +/– 0.19 | 2.7 +/– 0.95 | 43 | 24 | Yes | Yes | Cyt & mito | Yes | Yes | Yes | Yes | Yes |
| Catalyt mutants | ||||||||||||
| H96D | Up 2-fold | Down >2 × 104 | 29 | NT | NT | NT | Cyt & mito | Yes | Yes | NT | Yes | NT |
| H96N | Up 2-fold | Down >5 × 105 | 31 | 14.4 | NT | NT | Cyt & mito | Yes | Yes | Yes | Yes | Yes |
| Loop mutants | ||||||||||||
| Y114A | Up 23-fold | Down 2-fold | 3.7 | NT | NT | NT | Cyt | +/– | +/– | +/– | No | No |
| Y114D | NT | NT | 2.9 | 6 | NT | NT | Cyt | +/– | +/– | – | No | –/+ |
| Y114E | NT | NT | NT | NT | NT | NT | Cyt & mito | –/+ | –/+ | – | No | NT |
| Y114F | Up 5-fold | Up 1.1-fold | 11.5 | 3 | NT | NT | Cyt & mito | –/+ | –/+ | – | No | No |
| Y114W | Up 5-fold | Up 1.4-fold | NT | NT | NT | NT | Cyt & mito | –/+ | – | – | NT | NT |
| del113–117 | Up 10-fold | Down 38-fold | 5 | NT | NT | NT | NT | NT | NT | – | No | NT |
| Other mutants | ||||||||||||
| L25W | Up 7-fold | Down 4-fold | 15 | NT | NT | NT | Cyt | – | – | – | NT | –/+ |
| I10W,L25W | Up 32-fold | Down 6-fold | 11 | NT | NT | NT | NT | NT | NT | NT | NT | NT |
| F5W | Up 3.3 fold | NT | NT | 5 | NT | NT | NT | NT | NT | +/– | No | NT |
| Purified pFhit | ||||||||||||
| pFhit | Down 0.4-fold | Down 7-fold | NA | NA | –/+ | Yes | NA | NA | NA | NA | NA | NA |
| ppFhit | Down 0.4-fold | Down > 100-fold | NA | NA | –/+ | Yes | NA | NA | NA | NA | NA | NA |
| Up 60-fold | Down 6-fold | |||||||||||
To explore the in vivo importance of the Tyr-114 phosphorylation site and define Fhit-mediated signaling events, Semba et al. (14) compared the differential biological effects of Ad-FHIT WT and Ad-FHIT Tyr-114 mutant expression in human lung cancer cells. Caspase-dependent apoptosis was effectively induced only by WT Fhit protein. However, the biological significance of phosphorylation at Tyr-114 has been difficult to study because the endogenous phosphorylated forms have very short half-lives; activation of epidermal growth facto receptor family members induces Fhit phosphorylation by Src and proteasome degradation of phosphorylated Fhit (15).
Although there are possible connections among the various pathways known to be altered in Fhit-deficient cells, apoptosis, DNA damage-response checkpoint activation, ROS production, and related biological effects of Fhit loss or overexpression, details of the pathway(s) leading from Fhit overexpression to cell death and tumor suppression have not been delineated. Now that a Fhit signaling complex has been identified, we set out to examine which structural features of Fhit protein might participate in individual steps of the pathway leading from Fhit overexpression through complex formation, subcellular localization, interaction with mitochondrial Fdxr, DNA damage induction, cell cycle changes, apoptosis, and ultimately tumor suppression. The underlying hypotheses were as follows: substrate-binding mutants would behave similarly to WT; nonsubstrate-binding mutants would be defective in some step of the pathway, perhaps complexing with heat shock proteins or Fdxr or perhaps induction of DNA damage; and Tyr-114 mutants, which also affect formation or stability of the enzyme-substrate complex, would also be defective in executing some step of the Fhit overexpression pathway to cell death. One goal was to identify specific mutants that exhibited deficiency in specific steps of the pathway, so that such mutants could be used to dissect each step in more detail. Using in vitro Fhit and Fhit-effector protein interactions, we aimed to determine the following: 1) which proteins of the complex interact directly with Fhit, and 2) the biological role of these interactions in vivo. Using cancer cells expressing exogenous WT and mutant Fhit proteins, we were able to examine the structural features of Fhit that affect the direct interaction with its effectors, participate in ROS production, and are necessary for tumor suppression activity.
EXPERIMENTAL PROCEDURES
Cells, Vectors, and Antisera—H1299, MKN74, and HCT116, which are lung, gastric, and colon cancer cells, respectively, were maintained in RPMI 1640 medium plus 10% fetal bovine serum and penicillin/streptomycin (Sigma). H1299 lung cancer cells are both Fhit and p53 negative, and clonal derivatives stably, inducibly, or transiently expressing Fhit and Fhit mutants were prepared as described (16). MKN74 gastric cancer cell clones stably expressing Fhit and Fhit mutants have been described (9). HCT116 colon cancer cells were obtained from the B. Vogelstein laboratory and carry either three (+/+/+) or one (+/-/-) FDXR allele (17). Cells were transfected using Lipofectamine™ (Invitrogen) following the manufacturer's directions. H1299 cells were transiently infected with AdFHIT and AdFHIT mutant viruses as described previously and lysed for analysis of protein interactions at 48 h after infection (7, 11).
Western Blot Analysis—Immunoblot analyses were performed as described previously (7, 14) using rabbit polyclonal anti-Fhit (Zymed Laboratories Inc.), rabbit polyclonal antisera against Hsp60 and Hsp10 (Santa Cruz Biotechnology), and rabbit polyclonal anti-Fdxr (Abcam). Protein levels were normalized relative to β-actin and/or glyceraldehyde-3-phosphate dehydrogenase level, detected with appropriate antisera (Santa Cruz Biotechnology).
Protein-Protein Interaction—The glutathione S-transferase (GST)/in vitro transcription/translation (IVTT) protein-protein interaction assay was performed essentially as described (18). Briefly, HSP60, HSP10, FHIT, and FDXR cDNAs were subcloned into pGEX-4T-2 (Amersham Biosciences), which allows high expression of C-terminal GST fusion proteins, and were transformed into E. coli XL1-blue. 10-ml overnight cultures were diluted with LB medium to 800 ml and grown at 37 °C until the A600 values were 0.5; isopropyl-1-thio-β-d-galactopyranoside addition to a final concentration of 1 mm and incubation for 2 h at 30 °C induced protein expression. Cultures were harvested by centrifugation at 2000 × g for 15 min at 4 °C. Bacterial pellets were resuspended in phosphate-buffered saline (PBS) containing 1 mm phenylmethylsulfonyl fluoride, 2 μg/ml leupeptin, and 2 μg/ml pepstatin. Cells were lysed by two freeze-thaw cycles in the presence of 100 μg/ml lysozyme, Triton X-100 (0.2%), and 5 mm dithiothreitol. DNase I was added to final concentration of 20 μg/ml. Lysates were incubated on ice for 30 min, and cellular debris was removed by centrifugation. Cleared supernatants were incubated with glutathione-agarose beads (Sigma) for 1 h at 4 °C with gentle continuous agitation. Under these conditions, ∼20–50 ng of protein was bound to 25 μl of beads. Samples were pelleted for 1 min at 300 × g, and pellets were washed three times with 500 μl of binding buffer (20 mm Tris, pH 7.5, 10% glycerol, 150 mm NaCl, 5 mm EDTA, 1 mm dithiothreitol, 0.1% Tween 20, 0.75 mg/ml bovine serum albumin, 0.5 mm phenylmethylsulfonyl fluoride, 1 μg/ml leupeptin, and 1 μg/ml pepstatin). Lysates containing unmodified pGEX vector were treated similarly and used as negative control. GST and GST fusion protein binding to beads was verified by denaturing gel electrophoresis (SDS-PAGE) and quantified using bovine serum albumin as a standard, as described previously (19). To produce 35S-labeled proteins, HSP60, HSP10, FHIT and FDXR cDNAs were subcloned into pET24d (Novagen), and 1 μg of these constructs was used in IVTT (TnT-coupled reticulocyte lysate system; Promega). Labeled proteins were quantified as described (19). Equivalent amounts of protein from each IVTT mix were added to the suspension of protein-bound glutathione beads in 500 μl of binding buffer. Mixtures were incubated at 4 °C for 1 h with gentle agitation. The beads were washed three times with binding buffer and resuspended in SDS-PAGE loading buffer. Proteins were resolved on 4–20% gradient SDS-polyacrylamide gels, visualized, and quantified using a PhosphorImager system (GE Healthcare).
Flow Cytometry—Fhit or Fhit mutant-expressing cells were treated with 0.25 and 0.5 mm H2O2 and incubated for varying times, as indicated in the text and figure. Cells were collected, washed with PBS, and resuspended in cold 70% ethanol. For analysis, cells were spun down, washed in PBS, and suspended in 0.1 mg/ml propidium iodide/Triton X-100 staining solution (0.1% Triton X-100, 0.2 μg/ml DNase-free RNase A) for 30 min at room temperature and analyzed by flow cytometry.
In Vivo Experiments—Mouse studies were conducted in compliance with institutional guidelines using 6–8-week-old female Nu/nu mice (Charles River Laboratories Inc., Wilmington, MA) that had been injected with 2 × 106 HCT116 FDXR+/+/+ or FDXR+/-/- cells subcutaneously. Mice were randomly assigned to one of two treatment groups for the two cell types injected: (a) HCT116 FDXR+/+/+/FHIT-siRNAs, (b) HCT116 FDXR+/+/+/siNSR (nonspecific sequence), (c) FDXR+/-/-/FHIT-siRNAs, and (d) HCT116 FDXR+/-/-/siNSR; at days 7 and 14 after subcutaneous injection, tumors were injected with 16 nmol of lipo-nanoparticles complexed with 1 nmol of siRNA. Tumor volume was assessed at regular intervals by measuring the two diameters of the tumors (V (mm3) = a × b2/2). One week after the last treatment (day 21), mice were sacrificed, and tumors were excised and measured, and intratumoral protein expression was assessed by immunohistochemical and immunoblot analyses. MKN74 cell clones stably expressing WT Fhit (A116), or Y114A, Y114D, Y114F, H96N, or L25W mutant Fhit were injected (5 × 106 cells/mouse) into the right flanks of nude mice, four mice per clone, and tumor growth was compared with growth of MKN74/pRcCMV (empty vector) clone. At days 7 and 10, the tumors were measured, and on day 14, mice were sacrificed, and tumors were measured. Statistical analyses for tumor size were done by Student's t test, and p < 0.05 was considered as significant.
Immunohistochemistry—Immunohistochemical studies were performed using the streptavidin-peroxidase method. Immunohistochemical detection was done using Fhit and Fdxr rabbit polyclonal antisera, diluted 1:3000 and 1:100, respectively. Endogenous peroxidase activity was quenched by methanol containing 3% hydrogen peroxide (Sigma). Nonspecific binding was blocked by applying normal goat serum in a humidified chamber at a dilution of 1:10 for 30 min. Primary antisera were applied overnight at 4 °C. Secondary antiserum (goat anti-mouse or anti-rabbit immunoglobulin, DAKO, Denmark) was applied for 1 h at room temperature. Peroxidase-anti-peroxidase (PAP rabbit, DAKO) conjugate diluted at 1:100 in PBS was applied for 45 min at room temperature. Sections were stained with diaminobenzidine tetrahydrochloride (Sigma) and lightly counterstained with hematoxylin. Negative control slides were prepared by omitting primary antiserum. As positive controls for Fhit and Fdxr protein, paraffin sections of normal human tissues were used. Frequency of apoptosis was assessed in mouse xenograft tumor tissues by staining sections using the In Situ Ligation (ISOL) apoptosis detection kit (Chemicon), as directed by the manufacturer.
Detection of 8-OHdG—MKN74 cells stably expressing Fhit or Fhit mutants or H1299 cells expressing induced Fhit or Fhit mutants (10,000 cells/well, LAB-TEC 8 well) were incubated for 48 h and treated with or without 0.5 mm H2O2 for 5 h. Cells were fixed for 15 min with 3.7% buffered formaldehyde and permeabilized with 0.5% Triton X-100 for 10 min. The slides were then incubated with goat anti-8-hydroxyguanosine (8-OHdG) polyclonal antiserum (Chemicon) overnight at 4 °C (1:200) after blocking with 5% milk for 30 min. Negative controls were performed by replacing primary antiserum with PBS. Texas Red-conjugated anti-goat serum (Santa Cruz Biotechnology) was used as secondary antiserum (1:500). Cells were counterstained with 4′,6-diamidino-2-phenylindole. Images were captured with Zeiss Axio Cam and Axio Visionsystem (×400).
Assessment of Fdxr Protein Stability—1 × 106 H1299 cancer cell clones expressing Fhit or Fhit mutants were seeded in 6-well plates, and after 24 h, a cycloheximide (10 μg/ml) chase for 0, 6, 12, and 24 h was performed, and the protein lysates were analyzed by immunoblot detection of Fdxr expression level. H1299 cells were chosen for this experiment because they are completely negative for p53, which is a transcriptional inducer of FDXR expression.
RESULTS
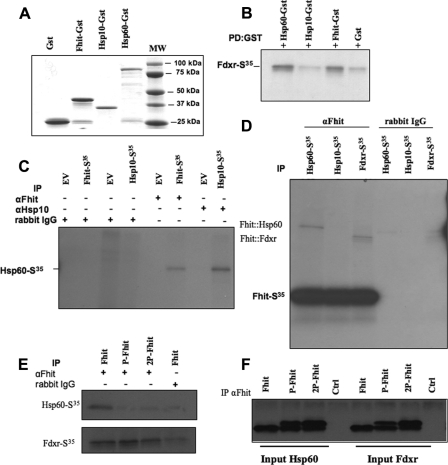
Interaction of Purified Fhit Complex Proteins in Vitro—We previously demonstrated the physical interaction of Fhit with Hsp60/Hsp10 and with Fdxr using protein-protein cross-linking and immunoprecipitation (IP) with Fhit antiserum (7). To further explore interactions in the Fhit protein complex, we did GST/IVTT bait-prey interaction experiments (19) to determine whether Fhit is able to interact directly with these proteins or might be part of a larger complex driving indirect interactions with the candidate proteins. We used an assay (19) that relies on use of a GST fusion protein expressed in E. coli as “bait” and IVTT protein as “prey.” The degree of purity of all GST fusion proteins and normalization among proteins is shown in Fig. 1A; this assay proved effective for Hsp60-GST-IVTT-Fdxr and Fhit-GST-IVTT-Fdxr complex interactions but did not detect interaction of Hsp10-GST-IVTT-Fdxr. We detected Hsp10-GST-IVTT-Hsp60 interaction (data not shown) but did not detect direct interaction between Fhit-GST and IVTT-Hsp60 or IVTT-Hsp10 (data not shown). We are confident that the interaction for each of these IVTT proteins was specific, because we observed nearly undetectable background nonspecific binding, as demonstrated by incubation and centrifugal precipitation of IVTT-Fdxr with purified GST protein previously normalized as control (Fig. 1B). To determine whether the C-terminal GST tag might influence correct Fhit interactions, we examined protein interactions using only IVTT proteins with anti-Fhit IP as the pulldown method. As shown in Fig. 1C, this strategy was effective with all candidate proteins. We detected the IVTT-Hsp10-IVTT-Hsp60 (Fig. 1C) and IVTT-Fhit-IVTT-Fdxr (Fig. 1D) interactions and were able also to detect the IVTT-Fhit-IVTT-Hsp60 interaction (Fig. 1C) but not an IVTT-Fhit-IVTT-Hsp10 interaction (Fig. 1D), confirming that the GST tag on the C terminus of Fhit-GST protein likely interfered with direct interaction with Hsp60, but not Hsp10, suggesting that Fhit does not interact directly with Hsp10. To determine whether mono- or diphospho-Fhit interacts with Hsp60 or Fdxr, purified Fhit, phospho-Fhit, or diphospho-Fhit forms (shown on denaturing gel in Fig. 1F) were individually combined with purified, 35S-labeled Hsp60 or Fdxr and IP performed using anti-Fhit. As shown in Fig. 1E, lower panel, the two phospho-Fhit forms did not bind Hsp60 but bound Fdxr; however, without Hsp60 binding and chaperoning to the mitochondria, these phospho forms may not encounter much Fdxr within cells. The recombinant Fhit and phospho-Fhit proteins were normalized to each other; to show that the unstable phospho-Fhit was still present after IP, we assessed the presence of the slower migrating phospho-Fhit band by Western blot of the immunoprecipitated protein (Fig. 1F).
FIGURE 1.
Direct interactions of purified Fhit forms with Hsp60/10 and Fdxr. A, immunoblot of preparations of purified GST-tagged proteins for in vitro interaction studies. Examples of the purified proteins made in bacteria are shown. B–D, interactions between purified Fhit and interactor proteins. GST-tagged or 35S-labeled proteins made by in vitro transcription-translation were prepared and combined in pairs as indicated, followed by radiographic imaging of pulled down or immunoprecipitated partner proteins. E, interaction of purified Fhit and phospho-Fhit forms with effectors. Purified Fhit and phospho-Fhit forms were combined with labeled interactors; IP was with anti-Fhit. The upper panel shows the radiographic image of co-IP of Hsp60 and the lower panel co-IP of Fdxr with the Fhit forms. F, anti-Fhit immunoprecipitates phospho-Fhit monomers. This panel shows the immunoblotted Fhit proteins precipitated for each of the IP experiments shown in E; the anti-Fhit serum precipitates and detects phosphorylated and unphosphorylated forms of Fhit.
In multiple experiments we have attempted to examine the effect of including Fhit substrate, Ap3A, in the protein binding experiments, but we find that MnCl2, included in Fhit-Ap3A binding buffer, causes nonspecific sticking of proteins to the agarose beads.
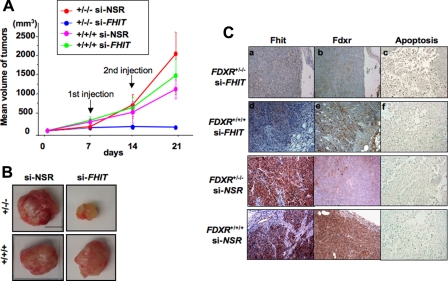
Effect of Abrogation of the Fhit-Fdxr Interaction on HCT116 Tumorigenicity—To confirm that interaction of endogenous Fhit and Fdxr is biologically relevant in vivo, nude mice were inoculated with HCT116 FDXR+/+/+ and FDXR+/-/- cells subcutaneously, 2 × 106 cells/mouse. Fhit is expressed at normal levels (7) in the parental HCT116 cells (with three copies of FDXR) and in the mutant (with two copies of FDXR knocked out; complete knock-out of FDXR is incompatible with viability (17)), illustrating that Fhit is not acting as a tumor suppressor in this cancer. We have taken advantage of the lethality of FDXR loss to show a biological effect of the Fhit-Fdxr interaction, in which Fhit is necessary for stabilizing the Fdxr protein, especially when this protein is limiting, as in the case of the HCT116+/-/- cells (7). We considered a model in which Fhit acts as a suppressor so that silencing Fdxr would abrogate the Fhit suppressor activity, but the lethal effect of Fdxr turnoff makes this model untenable. At 7 days post-injection, when the average tumor size was 8.4 ± 2 mm in diameter, mice were randomly divided into two treatment groups, and Lipofectamine nanoparticles were used to deliver FHIT-siRNAs or nonspecific siRNA (siNSR). Tumors were injected at days 7 and 14 with 1 nmol of siRNA mixed with 16 nmol of lipoparticles in PBS; tumors were removed at day 21 when the experiment was terminated. Dramatic inhibition of HCT116 FDXR+/-/- tumor growth was observed with siFHIT treatment (p < 0.0001) compared with other experimental groups, siNSR-lipoparticles in HCT116 FDXR+/-/- cells, siFHIT, or siNSR-lipoparticles in HCT116 FDXR+/+/+ cells (Fig. 2, A and B). HCT116 FDXR+/+/+/siFHIT and HCT116 FDXR+/+/+/siNSR tumors showed similar growth profiles. Fig. 2A shows the time course of tumor growth for each group. The effect of siRNAs on protein expression in the tumors was assessed by immunohistochemical analysis of fixed tumors (Fig. 2C) after in vivo treatment; a summary of the results is given in Table 2. The effect of in vivo FHIT knockdown on HCT116 tumor growth is consistent with the role of Fhit in stabilization of Fdxr, i.e. the HCT116 FDXR+/-/- cells require Fhit interaction to keep Fdxr at a level compatible with viability, as illustrated by the high level of apoptosis in the shrinking siFHIT-treated HCT116 FDXR+/-/- tumors (Fig. 2C). To understand this experiment, it is important to note that Fhit is not acting as a tumor suppressor in this experiment. Thus, the in vivo experiment confirms a physiologic role of the Fhit-Fdxr interaction.
FIGURE 2.
In vivo effect of abrogation of the Fhit-Fdxr interaction. Human colon cancer cells, HCT116 FDXR+/+/+, and FDXR+/-/- (1 × 106) cells were injected into the flanks of nude mice subcutaneously, and siFHIT or nonspecific sequence (siNSR) oligonucleotides with Lipofectamine were injected into the tumors on days 7 and 14. Mice were sacrificed on day 21 and tumor volumes calculated. A and B, FDXR+/-/- tumors treated with siFHIT were significantly smaller than tumors of other groups (scale bar, 10 mm). A, time course of tumor growth in all groups. B, tumors were excised on day 21, and portions of each tumor were taken for fixation, processing, and immunohistochemical analyses. C, tissue sections of siFHIT-treated tumors were stained for Fhit and Fdxr expression; representative results are shown; staining for apoptotic cells (by ISOL method) is also shown on the right. Expression of Fhit was low in both si-FHIT-treated tissues (panels a and d); expression of Fdxr was low in the tissue from FDXR+/-/- tumor (panel b) and high in tissue from FDXR+/+/+ tumor (panel e). Many apoptotic cells with black-brown nuclei were observed in tissue from FDXR+/-/- tumor (panel c) but not in tissue from FDXR+/+/+ tumor (panel f) panels a, b, d, and e, ×200; panels c and f, ×400. Similarly stained tissue sections of si-NSR treated tumors are shown below the si-FHIT-treated sections.
TABLE 2.
Fhit and Fdxr expression and apoptosis index in siFHIT treated tumors Fhit and Fdxr expression is as follows: +++, 100–90%; ++, 90–50%; +, 50–10%. Apoptotic index is as follows: ISOL positive cells/1000 cancer cells. *, p < 0.001.
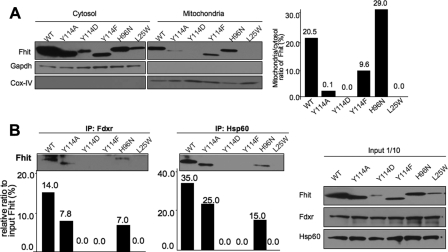
Fhit Mutants, Subcellular Localization, and Complex Formation—Fhit expression is reduced or absent in many cancer-derived cell lines, and stable or transient expression of exogenous Fhit in cancer cells has various biological effects. We have isolated and characterized stable and inducible Fhit and Fhit mutant-expressing cell lines that have now been used to examine subcellular localization of Fhit and Fhit mutants and then to examine the composition of the Fhit mutant-effector protein complexes in the transfected cells. Some of the mutants and their enzyme kinetics have been examined previously (3, 11, 14); characteristics of specific Fhit mutants and phospho forms are summarized in Table 1. Stable FHIT and FHITH96N mutant transfectants of MKN74 gastric cancer cells were shown to be suppressed for tumorigenicity in nude mice (9). The Fhit H96N protein with mutation of the central and critical His of the histidine triad is catalytically inactive with a kcat reduced by >10,000-fold, but the Km is only increased ∼2-fold (see Table 1), allowing continued substrate binding and prompting the hypothesis that substrate binding, but not cleavage, was important for suppressor activity (9, 10). Other mutations that affect substrate binding or enzyme cleavage differentially (11) and mutations at the Tyr-114 site that affect phosphorylation (14) have been isolated and some characteristics studied (see Table 1). MKN74/FHIT and MKN74/FHIT mutant stable transfectant cells were fractionated into cytosolic and mitochondrial fractions, and subcellular locations of the WT and mutant proteins were assessed (see Fig. 3A). In this experiment only WT and H96N protein showed substantial localization in the mitochondrial fractions (20 and 29%, respectively), with Y114F protein showing some mitochondrial localization (9.6%). The Y114A, Y114D, and L25W proteins remained mainly cytosolic in MKN74 stably expressing cells.
FIGURE 3.
Localization and co-immunoprecipitation of mutant Fhit proteins in stable expressors. A, mitochondrial and cytosol fractions of MKN74 transfectants were isolated using mitochondria/cytosol fractionation kit (BioVision), and fractionated lysates were separated on a polyacrylamide gel and probed with anti-Fhit serum (left panel). Western blot images were analyzed with GS800 calibrated densitometer (Bio-Rad) and Quantity One software system to determine the ratio of Fhit or mutant Fhit protein in mitochondria versus cytosol (right panel). Note that the Y114D residue adds a negative charge to the protein, effecting slower migration on the gel, whereas the Y114F and Y114A mutants migrate faster than WT Fhit. Note also that the Y114D mutant was made to mimic a phospho-Fhit and is apparently much less stable than WT, in accord with the report that phospho-Fhit is targeted for degradation (15). B, MKN74/FHIT and FHIT mutant transfectants (WT, Y114A, Y114D, Y114F, H96N, and L25W) were cultured, and 106 cells of each were collected. Cells were lysed; DSP cross-linker was added, and immunoprecipitation with Fdxr or Hsp60 antiserum was carried out; proteins were separated on polyacrylamide gel and probed with anti-Fhit serum. Western blot images were analyzed with GS800 calibrated densitometer (Bio-Rad) and Quantity One software system.
In co-immunoprecipitation (co-IP) experiments using antiserum against Fdxr or Hsp60, WT and H96N proteins were co-precipitated with both Fdxr and Hsp60 (Fig. 3B). In this experiment we observed some co-precipitation of Y114A protein with Fdxr and Hsp60, although the sub-localization experiment in Fig. 3A suggested that Y114A protein did not localize to mitochondria, where most Fdxr protein is found.
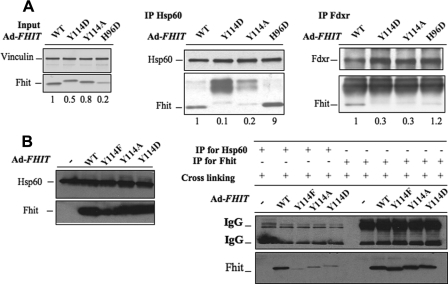
Similar experiments were performed to examine complex formation of Fhit mutants in H1299 lung cancer cells after transient transfection with FHIT or FHIT mutant plasmids, infection with AdFHIT and AdFHIT mutant viruses, or in H1299 cells inducibly expressing Fhit or Fhit mutant proteins. Examples of results in H1299 AdFHIT WT and mutant infected cells are shown in Fig. 4. In Fig. 4A, co-IP of Fhit with Hsp60 (middle panel) illustrates that both WT and Fhit H96D, which bind Fhit substrate well, interact well with Hsp60, whereas the mutants Y114D and Y114A interact poorly with Hsp60; similarly, these two mutants, which bind Fhit substrate poorly, also bind Fdxr poorly. A second experiment is shown in Fig. 4B. Fig. 4B, left panel, shows the level of Fhit and Hsp60 in lysates of H1299 cells infected with AdFHIT WT or AdFHIT Y114F, Y114A, or Y114D mutants. As shown in Fig. 4B, right co-IP panel, the three Tyr-114 mutant proteins interact poorly with Hsp60.
FIGURE 4.
FhitY114 mutants bind poorly to Fhit effector proteins. A, H1299 cells were infected with AdFHIT or AdFHIT mutants at a multiplicity of infection of 5. At 48 h after infection cells were lysed for immunoprecipitation experiments. In the left panel, the immunoblot shows the level of Fhit or mutant Fhit protein expressed in infected cells, normalized to the level of Vinculin. The lysates, in amounts that would give equivalent amounts of Fhit protein, were then used in IP experiments, as shown in the middle and right panels; upper nonspecific bands in these panels include rabbit IgG. Numbers below each lane indicate the relative amounts of Fhit or Fhit mutant protein co-precipitated with Hsp60 or Fdxr, respectively. B, similar experiment was performed using Y114F, Y114A, and Y114D mutants; the left panel shows the input level of Hsp60 and Fhit after infection of H1299 cells. The right panel shows the IP with Hsp60 and Fhit antisera. Western blot images were analyzed with GS800 calibrated densitometer (Bio-Rad) and Quantity One software system.
The cumulative results of these co-localization and co-IP experiments show that not only are nonsubstrate-binding proteins not good apoptosis inducers as shown previously (11), but in general they do not participate in interaction with the Fhit effector proteins, a function that precedes induction of apoptosis by Fhit. Although the Fhit viral protein expression was not perfectly homogeneous among Fhit WT and mutants (Fig. 4A), the IP for Hsp60 and Fdxr was normalized for Fhit protein expression (input), previously normalized for two housekeeping proteins Vinculin (Fig. 4A) and glyceraldehyde-3-phosphate dehydrogenase (data not shown).
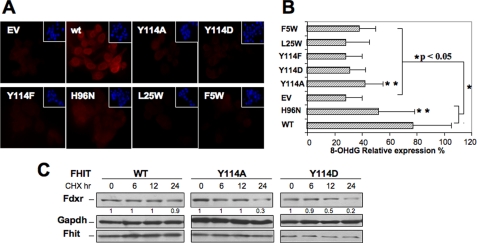
Fhit-related Biological Functions of Cancer Cells Expressing Fhit Mutants—To examine the responses of cancer cells expressing exogenous WT and mutant Fhit proteins, the stable MKN74 Fhit expressors and the induced H1299 expressors were treated with 0.5 mm H2O2, a concentration known to differentially effect ROS and apoptosis in Fhit-negative and -positive cancer cells (7), and the occurrence of DNA damage in the nuclear genomes was assessed by immunofluorescent detection of 8-OHdG residues in the DNA of the respective cell lines (Fig. 5). Fig. 5A illustrates results for the MKN74 stable transfectants, in which the red nuclear stain detects 8-OHdG and the blue stain is 4′,6-diamidino-2-phenylindole nuclear DNA staining (insets). In the MKN74 stable transfectants only the WT and H96N-expressing cells exhibit strong 8-OHdG; Y114A and F5W expressing cells show weak 8-OHdG staining, whereas empty vector, Y114D, Y114F, and L25W-expressing cells are negative for 8-OHdG staining. These results are an indication of more extensive oxidative DNA damage in WT and H96N mutant expressing cells, as shown in the bar graph in Fig. 5B.
FIGURE 5.
H2O2 induction of 8-OHdG in DNA of Fhit and Fhit mutant stable and induced expressors. A, stable MKN74 Fhit and mutant expressors were treated with 0.5 mm H2O2, and DNA damage in the nuclear genomes was assessed by immunofluorescent detection of 8-OHdG residues in the DNA of the respective cell lines. The red nuclear stain detects 8-OHdG and the blue stain is 4′,6-diamidino-2-phenylindole nuclear DNA staining (insets). Negative controls for all cells showed no staining (data not shown). B, bar graph ± S.D. shows the relative frequency of 8-OHdG detection in MKN74 WT and mutant expressors. The double asterisk on Y114A and H96D denotes a p value >0.05. C, immunoblot analysis of Fdxr, Fhit, and glyceraldehyde-3-phosphate dehydrogenase in H1299 cells expressing Fhit WT, Y114A, and Y114D proteins, showing the Fdxr level after cycloheximide (CHX) chase (30 μg/ml) for 6–24 h. Densitometry based on glyceraldehyde-3-phosphate dehydrogenase levels shows enhanced stability of Fdxr in presence of WT Fhit.
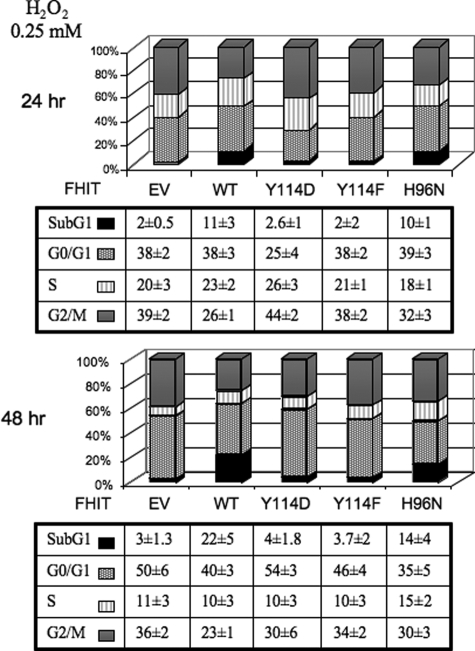
Similarly, H1299D1 cells expressing induced WT Fhit showed strong nuclear 8-OHdG, whereas Y114E, Y114F, Y114W, and del113–117-expressing cells showed lack of evidence of 8-OHdG in peroxide-treated cells (not shown). Negative controls for all cells showed no staining (data not shown). To determine whether the variable 8-OHdG staining in WT and Fhit mutant-expressing cells could be a result of varying Fdxr stability in the cells, a cycloheximide chase experiment was performed. We found that in accord with previous results (7), Fdxr was relatively stable when WT Fhit was present (Fig. 5C). Instead, in the presence of Y114D mutant, we observed a decrease in Fdxr stability at 12 (0.5) and at 24 h (0.2). Interestingly, in the presence of the Y114A mutant protein, which exhibited only partial mitochondrial localization, immunoprecipitated with Hsp60 and Fdxr (Fig. 3), but did not function as tumor suppressor in vivo (see below), Fdxr was more stable at 12 h (1), compared with Y114D (0.5), but was less stable at 24 h (0.3), compared with the WT (Fig. 5C). This result strongly suggests that Tyr-114 is not only important for binding but directly affects Fdxr stability when interaction occurs. In previous experiments (7) we have shown that after H2O2 treatment, Fhit-negative MKN74 and H1299 cancer cells escape apoptosis, whereas the same cells expressing WT Fhit are more susceptible to apoptosis induced by H2O2, as well as cisplatin and paclitaxel. Thus, we also tested mutant Fhit-expressing cells for response to H2O2 treatment by flow cytometric measurement of cell cycle kinetics and DNA content. Results are shown in Fig. 6 for 0.25 mm H2O2 at 24 and 48 h after addition. By 48 h, WT Fhit-expressing MKN74 cells showed 22% sub-G1 cells, and H96N-expressing cells showed 14% sub-G1 cells, compared with 4% or less sub-G1 cells for Y114D, Y114F, and empty vector expressing cells, confirming that the Y114D and Y114F mutants are deficient in apoptosis induction. Furthermore, although at 24 h we did not find a significant sub-G1 population in Tyr-114 mutant-expressing cells, compared with WT and H96N expressors, there was a consistent increase in G2/M population consistent with the fact that stressed p53 minus or mutant cells undergo G2/M arrest; at 48 h the Tyr-114 mutant expressors overcame arrest and re-entered the cell cycle. Consistent with previous observation, Fhit-deficient cells avoid G2/M arrest and apoptosis, likely carrying unrepaired DNA damage.
FIGURE 6.
Cell cycle kinetics of H2O2-treated MKN74/Fhit and Fhit mutant expressors. MKN74 stable Fhit or mutant-expressing cells were treated with 0.25 mm H2O2 and incubated for 24 (top panels) or 48 h (bottom panels). Cells were collected, washed with PBS, and resuspended in cold 70% ethanol. For analysis, cells were spun down, washed in PBS, and suspended in 0.1 mg/ml propidium iodide/Triton X-100 staining solution (0.1% Triton X-100, 0.2 μg/ml DNase-free RNase A) for 30 min at room temperature and analyzed by flow cytometry.
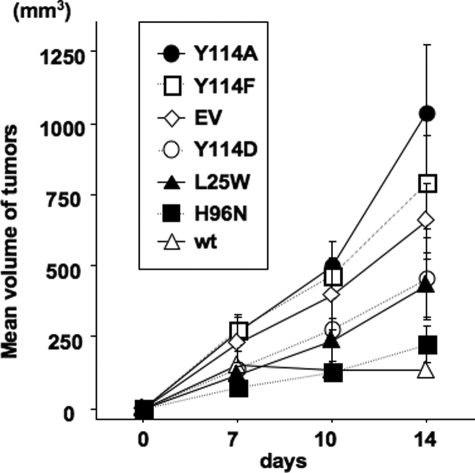
Finally, Fhit WT and Fhit mutant-expressing MKN74 cells were injected into nude mice, and growth of tumors was assessed to determine the tumor suppressor function of Fhit mutants relative to WT Fhit and Fhit negative (empty vector) MKN74 cells. Results of the experiment are shown in Fig. 7, which illustrates that WT and H96N-expressing cells are almost completely suppressed for tumor growth, whereas Y114A- and Y114F-expressing cells formed large tumors by day 14; Y114D- and L25W-expressing cells showed intermediate growth, suggesting that these mutants retain some tumor suppressor function.
FIGURE 7.
Tumorigenicity of MKN74/Fhit and Fhit mutant expressors in nude mice. 5 × 106 WT or mutant (Y114A, Y114D, Y114F, H96N, and L25W) stable expressing MKN74 cells were injected subcutaneously into the right flanks of four nude mice for each cell line. Tumor sizes were measured over a 2-week period and volumes of tumors calculated and graphed with standard deviations as shown. Significant differences in pairwise comparisons of tumor sizes of different MKN74/Fhit and mutant expressors were determined by Student's t test. The tumors from MKN74/Fhit WT cells were significantly smaller than tumors of all mutant expressors (values ranging from p = 0.008 for Y114A to p = 0.04 for Y114D and L25W) except H96N (p = 0.2). The MKN74/H96N tumors were significantly smaller than Y114A (p = 0.01) and Y114F (p = 0.02) tumors. There were no other significant differences.
DISCUSSION
The earlier finding that Fhit interacts with the Hsp60/10 complex and is imported into mitochondria where it interacts with and stabilizes Fdxr is of interest for a number of reasons. 1) Under stress conditions Fdxr accumulates in mitochondria to a higher level in Fhit-positive cells (data not shown). 2) Fdxr accumulation under stress conditions increases the electron transport process, depleting the reduced NADPH pool (20), and in the absence of NADPH to detoxify ROS, ROS-induced apoptosis is amplified (7). 3) In the absence of Fhit, a condition occurring in most cancers, there is less Fdxr in the mitochondria, allowing better detoxification of ROS and less apoptosis. 4) This can allow survival of Fhit-deficient cells carrying a low level of oxidative damage that can contribute to neoplastic progression. Fhit-negative pre-neoplastic lesions would escape apoptosis, carrying some oxidative damage and resulting mutations, setting pre-neoplasias on the path to progression. Antioxidants may contribute to cancer prevention by blocking this route to neoplastic progression.
Similarly, p53 is activated and induces apoptosis in response to various cellular stresses in part through ROS, and simultaneously p53 increases transcription of the FDXR gene, which in turn enhances p53 function by increasing ROS-induced apoptosis (17, 20). Sablina et al. (21) showed that through the level of p53 expression, FDXR expression and thus ROS generation are fine-tuned, such that with normal p53 expression levels and mild oxidative damage, p53 induces antioxidant genes, and ROS is lowered and DNA repaired; with extensive stress and irreparable damage, p53 induces expression of pro-oxidant genes, including FDXR, and ROS is generated and damaged cells die by apoptosis.
Direct Protein-Protein Interactions in the Fhit Complex—In vitro studies of direct interactions among Fhit and effector proteins showed that Fhit interacts directly with Hsp60 and Fdxr and Hsp60 interacts directly with Hsp10, suggesting that Fhit indirectly interacts with Hsp10 through binding to Hsp60.
As for interaction of phospho-Fhit forms, it was hypothesized (13) that monophospho-Fhit should bind Ap3A well because of its decreased Km value, whereas it is difficult to predict expectations for diphospho-Fhit because of its biphasic kinetics (13). It is possible that the diphosphorylated form of Fhit does not occur in cells because phosphorylation of one Fhit monomer leads to rapid degradation (15). Results shown in Fig. 1E suggest binding of neither mono- nor diphospho-Fhit to Hsp60 but are consistent with some binding of monophospho-Fhit to Fdxr in vitro and little binding of diphospho-Fhit, especially considering that Fig. 1F results suggest the presence of some monophospho-Fhit in the diphospho-Fhit preparation because of dephosphorylation during storage (see lower band in the lanes labeled 2P-Fhit, indicating a nonphosphorylated Fhit monomer on this denaturing gel). In any case, Fdxr probably does not encounter phospho-Fhit in the mitochondria because of lack of Hsp60 interaction and the probability that phospho-Fhit is rapidly degraded. Thus, phospho-Fhit proteins likely do not participate in the tumor suppressor function of Fhit.
In Vivo Effect of Fhit-Fdxr Interaction—To demonstrate that the interaction of Fhit and Fdxr had biological consequences, we needed a cell system in which Fhit expression has a biological effect, such as an effect on tumor growth or apoptosis. Fhit is expressed in few tumorigenic cancer-derived cell lines, and in those with Fhit expression, Fhit is not suppressing growth; thus, the ideal cell model, a cancer cell that is suppressed for tumor growth because of endogenous Fhit expression so that we could test effects of Fdxr knockdown on tumorigenicity, is an oxymoron. Therefore, we chose to test the effect of Fhit knockdown in tumors with varying levels of Fdxr. The fact that Fhit knockdown in HCT116 FDXR+/-/- tumors, but not HCT116 FDXR+/+/+ tumors, causes highly significant eradication of tumors, as illustrated in Fig. 2 and Table 2, may seem antithetical to the role of Fhit as a tumor suppressor, but we interpret this experiment not only as proof of the biological necessity of the Fhit-Fdxr interaction but as evidence that further reduction of the level of Fdxr in HCT116 FDXR+/-/- cells, because of instability in the absence of Fhit, leads to loss of viability.
Biochemical and Biological Functions of Fhit Mutants—Specific Fhit amino acids were previously targeted for mutation based upon predicted effects on substrate binding, catalytic activity, or the ability to be phosphorylated (3, 9, 11, 14), and it was determined that mutants with reduced substrate binding showed reduced ability to induce apoptosis, whereas mutants within the catalytic domain still showed apoptotic and suppressor function (9, 11). The highly conserved (from Schizosaccharomyces pombe to Homo sapiens) tyrosine at position 114 of human Fhit is within an unstructured 20-amino acid loop that has been invisible in all reported Fhit structures (10). Preliminary x-ray crystallographic studies show that the loop is less flexible (i.e. more visible) when Tyr-114 is phosphorylated,6 and we hypothesize that the loop in phosphorylated forms may hover over the substrate surface, simultaneously allowing lowered Km values but possibly obscuring the Fhit-substrate signaling surface. Tyr-114 mutant Fhit proteins had higher Km values and were less capable of inducing caspase-dependent apoptosis in human lung cancers (14).
To determine which Fhit biological activities are retained by specific Fhit mutants that were previously studied for the ability to induce apoptosis (see Table 1, left 5 columns) in cancer cells (9, 11), we have used two cell models with stable or transient expression of exogenous Fhit and Fhit mutants; the gastric cancer-derived cells, MKN74 with homozygous deletion of FHIT exon 5 (9) and mutant p53, are highly tumorigenic in nude mice, and WT Fhit suppresses tumorigenicity (9); and H1299 lung cancer cells express no endogenous Fhit or p53. The results obtained with the different cellular models were similar but not identical, possibly because of cancer cell differences or methods of producing Fhit expression, or both. The results shown in the right 6 columns of Table 1 are a summary of the consensus results from the three cell models, including some data that were not shown. The Fhit H96N and H96D catalytic mutants with near zero catalytic activity but only 2-fold reduced substrate binding ability are very similar to WT Fhit in each measured biological function, including sub-localization to cytosol and mitochondria, binding to Hsp60 and Fdxr, production of oxidative DNA damage, apoptosis, and tumor suppression. Y114D mutant produced little protein, which did not bind Hsp60 or Fdxr and did not localize to mitochondria, perhaps because Fhit Y114D mimics phospho-Fhit and is rapidly degraded (see Fig. 3B, right panel). The Y114F and L25W mutants did not bind Fhit substrate well, did not bind Hsp60 or Fdxr, nor induce production of 8-OHdG or apoptosis after H2O2 treatment; however, these mutants when expressed in MKN74 cells produced tumors that, although significantly larger than tumors produced by WT Fhit-expressing cells (p = 0.04), were not much larger than tumors produced by H96N expressors (p = 0.2). Does this mean that these mutants retain some tumor suppressor activity? It seems unlikely, and we believe that we would see a significant difference in sizes relative to H96N tumors with another few days growth.
Perhaps the most interesting mutant, Fhit Y114A in MKN74 cells, did bind Hsp60 and Fdxr (Fig. 3B), apparently as well as H96N, and showed some H2O2-induced 8-OHdG (Fig. 5B) but completely lacked tumor suppressor activity. Fhit Y114A stands out from the other loop mutants in having the highest Km value, i.e. it binds substrate most poorly of all the mutants, consistent with its lack of tumor suppressor activity. The deletion mutant, del113–117, has the next highest Km value, consistent with the idea that the Y114A and deletion mutants have the most drastic effect on the biochemical or structural function of the loop region, perhaps eliminating the ability of the loop to support binding of the substrate, or perhaps obscuring the binding site altogether, preventing substrate binding and suppressor signaling. Another possible conclusion from the Y114A result is that substrate binding is not essential for interaction with Hsp60 or Fdxr, although it may influence mitochondrial localization. More detailed analysis of biochemical and biological properties of the Y114A, del113–117, and Y114D mutants will be useful. For example, it would be interesting to determine the crystal structure of the Y114A protein to observe the effect on the loop structure. Because Tyr-114 mutants are not in the catalytic site but affect substrate binding, Tyr-114 clearly has other functions in addition to signaling degradation, because Tyr-114 mutants lack several biological and biochemical functions. The results of sub-localization and in vivo effector binding studies raise several questions. Is substrate binding necessary for interaction of Fhit with Hsp60, for interaction with Fdxr, for mitochondrial localization, or for all three, and is substrate binding the critical feature for all of these functions? In H1299 cells transiently expressing these mutant proteins (Fig. 4), Hsp60 interaction seems equally poor for the three Tyr-114 mutants. Interestingly, mitochondrial localization of rat dinucleoside triphosphate hydrolase, almost certainly the rat Fhit homolog, and the Ap3A substrate, in the matrix space of rat liver mitochondria was reported as early as 1991 (22). The results of flow cytometric detection of apoptotic fractions, 8-OhdG, and xenograft suppression in MKN74 stable transfectants, as well as H1299-inducible transfectants, showed that Fhit mutants that lack ability to bind Fhit substrate also showed reduced ability to participate in Fhit biological activities.
The convergence of the biochemical assay results with the biological outcomes were satisfying yet suggest the following possibilities: 1) there may be additional protein members of the Fhit complex that could differ in specific cancer cell types and could influence mitochondrial localization and effector binding; 2) substrate binding may not be the only function that affects Fhit binding to Hsp60 and Fdxr; and 3) it will be necessary to test more cell types and more protein interactors (7) to clarify differences in biochemical activities of Fhit within cells and how these biochemical functions transmit signals for biological functions.
Acknowledgments
We are grateful to Bert Vogelstein for providing us with HCT116 FDXR+/-/- and parental cells and to Richard Fishel for providing pGEX-4T-2 and for important suggestions for the study of protein-protein interactions.
This work was supported, in whole or in part, by National Institutes of Health Grants CA77738, CA132453, and CA115965 (USPHS). This work was also supported by grants from the Associazione Italiana Ricerca Cancro. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact.
Footnotes
The abbreviations used are: Fdxr, ferredoxin reductase; Ap3A, diadenosine triphosphate; IVTT, in vitro transcription-translation; IP, immunoprecipitation; PBS, phosphate-buffered saline; ROS, reactive oxygen species; 8-OHdG, 8-oxo-deoxyguanosine; GST, glutathione S-transferase; siRNA, short interfering RNA; siNSR, nonspecific siRNA.
H. Pace and L. D. Barnes, unpublished data.
References
- 1.Huebner, K., and Croce, C. M. (2001) Nat. Rev. Cancer 1 214-221 [DOI] [PubMed] [Google Scholar]
- 2.Huebner, K., and Croce, C. M. (2003) Br. J. Cancer 88 1501-1506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Barnes, L. D., Garrison, P. N., Siprashvili, Z., Guranowski, A., Robinson, A. K., Ingram, S. W., Croce, C. M., Ohta, M., and Huebner, K. (1996) Biochemistry 35 11529-11535 [DOI] [PubMed] [Google Scholar]
- 4.Ji, L., Fang, B., Yen, N., Fong, K., Minna, J. D., and Roth, J. A. (1999) Cancer Res. 59 3333-3339 [PubMed] [Google Scholar]
- 5.Ishii, H., Dumon, K. R., Vecchione, A., Trapasso, F., Mimori, K., Alder, H., Mori, M., Sozzi, G., Baffa, R., Huebner, K., and Croce, C. M. (2001) Cancer Res. 61 1578-1584 [PubMed] [Google Scholar]
- 6.Roz, L., Gramegna, M., Ishii, H., Croce, C. M., and Sozzi, G. (2002) Proc. Natl. Acad. Sci. U. S. A. 99 3615-3620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Trapasso, F., Pichiorri, F., Gasparo, M., Palombo, T., Aqeilan, R. I., Gaudio, E., Okumura, H., Iuliano, R., Di Leva, G., Fabbri, M., Birk, D. E., Raso, C., Green-Church, K., Spagnoli, L. G., Venuta, S., Huebner, K., and Croce, C. M. (2008) J. Biol. Chem. 283 13736-13744 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 8.Ohta, M., Inoue, H., Cotticelli, M. G., Kastury, K., Baffa, R., Palazzo, J., Siprashvili, Z., Mori, M., McCue, P., Druck, T., Croce, C. M., and Huebner, K. (1996) Cell 84 587-597 [DOI] [PubMed] [Google Scholar]
- 9.Siprashvili, Z., Sozzi, G., Barnes, L. D., McCue, P., Robinson, A. K., Eryomin, V., Sard, L., Tagliabue, E., Greco, A., Fusetti, L., Schwartz, G., Pierotti, M. A., Croce, C. M., and Huebner, K. (1997) Proc. Natl. Acad. Sci. U. S. A. 94 13771-13776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pace, H. C., Garrison, P. N., Robinson, A. K., Barnes, L. D., Draganescu, A., Rösler, A., Blackburn, G. M., Siprashvili, Z., Croce, C. M., Huebner, K., and Brenner, C. (1998) Proc. Natl. Acad. Sci. U. S. A. 95 5484-5489 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Trapasso, F., Krakowiak, A., Cesari, R., Arkles, J., Yendamuri, S., Ishii, H., Vecchione, A., Kuroki, T., Bieganowski, P., Pace, H. C., Huebner, K., Croce, C. M., and Brenner, C. (2003) Proc. Natl. Acad. Sci. U. S. A. 100 1592-1597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pekarsky, Y., Garrison, P. N., Palamarchuk, A., Zanesi, N., Aqeilan, R. I., Huebner, K., Barnes, L. D., and Croce, C. M. (2004) Proc. Natl. Acad. Sci. U. S. A. 101 3775-3779 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Garrison, P. N., Robinson, A. K., Pekarsky, Y., Croce, C. M., and Barnes, L. D. (2005) Biochemistry 44 6286-6292 [DOI] [PubMed] [Google Scholar]
- 14.Semba, S., Trapasso, F., Fabbri, M., McCorkell, K. A., Volinia, S., Druck, T., Iliopoulos, D., Pekarsky, Y., Ishii, H., Garrison, P. N., Barnes, L. D., Croce, C. M., and Huebner, K. (2006) Oncogene 25 2860-2872 [DOI] [PubMed] [Google Scholar]
- 15.Bianchi, F., Magnifico, A., Olgiati, C., Zanesi, N., Pekarsky, Y., Tagliabue, E., Croce, C. M., Ménard, S., and Campiglio, M. (2006) Proc. Natl. Acad. Sci. U. S. A. 103 18981-18986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cantor, J. P., Iliopoulos, D., Rao, A. S., Druck, T., Semba, S., Han, S. Y., McCorkell, K. A., Lakshman, T. V., Collins, J. E., Wachsberger, P., Friedberg, J. S., and Huebner, K. (2007) Int. J. Cancer 120 24-31 [DOI] [PubMed] [Google Scholar]
- 17.Hwang, P. M., Bunz, F., Yu, J., Rago, C., Chan, T. A., Murphy, M. P., Kelso, G. F., Smith, R. A., Kinzler, K. W., and Vogelstein, B. (2001) Nat. Med. 7 1111-1117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schmutte, C., Marinescu, R. C., Sadoff, M. M., Guerrette, S., Overhauser, J., and Fishel, R. (1998) Cancer Res. 58 4537-4542 [PubMed] [Google Scholar]
- 19.Guerrette, S., Wilson, T., Gradia, S., and Fishel, R. (1998) Mol. Cell. Biol. 18 6616-6623 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu, G., and Chen, X. (2002) Oncogene 21 7195-7204 [DOI] [PubMed] [Google Scholar]
- 21.Sablina, A. A., Budanov, A. V., Ilyinskaya, G. V., Agapova, L. S., Kravchenko, J. E., and Chumakov, P. M. (2005) Nat. Med. 11 1306-1313 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bernet, D., Pinto, R. M., Sillero, A., and Cameselle, J. C. (1991) FEBS Lett. 283 286-288 [DOI] [PubMed] [Google Scholar]