Abstract
The central role of small RNAs in regulating bacterial gene expression has been elucidated in the last years. Typically, small RNAs act via specific basepairing with target mRNAs, leading to modulation of translation initiation and mRNA stability. Quantitative studies suggest that small RNA regulation is characterized by unique features, which allow it to complement regulation at the transcriptional level. In particular, small RNAs are shown to establish a threshold for the expression of their target, providing safety mechanism against random fluctuations and transient signals. The threshold level is set by the transcription rate of the small RNA, and can thus be modulated dynamically to reflect changing environmental conditions.
Introduction
In recent years, it has been noticed that small non-coding RNAs (referred to here as sRNAs) can exert significant regulatory effect on gene expression in bacteria [1,2]. Early on, sRNAs were mainly recognized for their specialized roles in controlling the transposition of insertion elements [3], in regulating plasmid copy number [4,5], and in mediating plasmid maintenance through the toxin-antidote systems [6]. Only a few cases involving chromosomally encoded sRNA were known [7,8]. The advent of genomic and bioinformatic methods has led to the identification and the subsequent verification of numerous chromosomal sRNAs in a variety of bacterial species during this decade [9–11]. These sRNAs are involved predominantly in stress response pathways [12] and pathogenesis [13], but they are also found in the regulation of metabolism [7,14], transport [15], quorum sensing [16] and more.
Small RNAs bind specifically to the mRNA molecules of their target, in a manner that relies both on sequence complementarity and on secondary structure. Like regulatory proteins, many of these sRNAs act in trans, regulate the expression of multiple target genes, and are themselves regulated by one or more factors. Small RNAs – like proteins – can act as activators or repressors of gene expression by modulating translational efficiency and mRNA stability.
A great deal of progress has been made in the past few years in elucidating the mechanisms of sRNA-mediated regulation of gene expression, especially for the major class of post-transcriptional repressors [2,17–20]. Typically, translational repression results from binding of the sRNA to the translational initiation region of the target mRNA. Importantly, this binding may lead to the rapid degradation of both the sRNA and the target mRNA [21]. In other cases, the RNA complex may be highly stable, trapping both RNAs in an inert state [22,23]. Thus, unlike regulatory proteins, which exert their effect on gene expression in a catalytic fashion, these sRNAs are consumed as they exert their regulatory effects.
At the level of “Boolean logic”, namely when gene expression is only considered to be either “on” or “off”, transcriptional and post-transcriptional regulators may seem redundant. Recent studies suggest that sRNA repressors are in fact distinct from transcriptional repressors at a quantitative level, and that its unique characteristics may be closely related to the physiological roles of sRNA regulators. To establish these results, a simple mathematical model (Box 1) has been formulated [16,24,25], and different aspects of it have been studied theoretically [25–30] and experimentally [25–27]. Here we review and expand upon the current state of knowledge on the quantitative aspects of sRNA regulation.
Small RNAs establish threshold for gene expression
Consider the case of an sRNA with a single target at steady state. If transcription of the sRNA (αs) is more frequent than that of the mRNA (αm < αs), then most of the targets are expected to pair rapidly with the sRNAs, and the target will not be expressed. Conversely, if αm > αs then most of the sRNAs pair with the target while the unpaired mRNAs are free to direct protein translation. In the latter case, the expressed protein level would reflect the difference between the two transcription rates, i.e., be linearly proportional to αm − αs. This qualitative expectation is summarized by the schematic response shown in Figure 1, where the target transcription rate αm is taken to be continuously tunable at a given sRNA transcription rate αs. This behavior is referred to as a threshold-linear response [25]. A key feature of this form is that different values of αs shift the position of the threshold, without changing the slope of the linear rise beyond it. Associated with the threshold at αm ≈ αs is the ultrasensitivity of the response, namely an abrupt switch-on of target expression via a small change in target transcription [16,24]. This type of behavior has been observed experimentally in E. coli using a synthetic reporter target [25].
Figure 1. Threshold linear response.
The expression of a target gene is silenced as long as its transcription rate, αm, is smaller than that of the small RNA, αs. Below this dynamically defined threshold, the small RNAs buffer translational noise and transient changes in environmental conditions. Above the threshold, the effect of the small RNA become negligible.
Many bacterial small regulatory RNAs are involved in regulating stress response pathways. It is often the case in stress responses that an external signal triggers the execution of a cellular response that may be expensive (in terms of energy and nutrients) or even toxic [39,40]. In such cases, small RNAs may define a threshold for activation of the response. This threshold may reflect the level of tolerance of the cell to that particular stress. For example, the transcription rate of the small RNA RyhB reflects the concentration of free Fe2+ ions in the cell, and thus reflects the level of tolerance to superoxidative stress [32]. Indeed, RyhB sets a threshold for the expression of the superoxide dismutase SodB [25], which responds to that stress.
Small RNAs suppress fluctuations
Gene expression is a stochastic process, involving synthesis of individual molecules through a series of orchestrated yet random events [41,42]. Repression of gene expression may result in a small number of proteins per cell, raising the possibility of significant temporal and cell-to-cell fluctuations.
Consider first a gene whose transcription is tightly repressed by a transcription factor. In this case, transcription events are rather rare (can occur for example only a few times every hour [43,44]). Every transcription event is than amplified by recurring translation events, resulting in a burst of proteins; see figure 2a. The size of this burst is controlled by the rate of translation of this particular mRNA, multiplied by the lifetime of the RNA molecule. Thus, expression of a gene that is repressed transcriptionally is “bursty”, with long periods of silence punctuated with (sometimes large) bursts of proteins [45].
Figure 2. small RNAs suppress fluctuations in gene expression.

Computer simulations were performed for a gene that is repressed by (a) a transcription factor or (b) a small RNA. Upper panels show the cellular protein levels as a function of time, and lower panels show the individual transcription events. In the case of transcriptional repression (a), rare transcription events are accompanied by large protein bursts. In the case of post-transcriptional repression (b), transcription events are frequent, but the small RNA pool reduces the bursts. In these simulations, parameters for the first gene were chosen based on the known parameters of the lac operon in E. coli. Required parameters for the second gene were used to ensure that the mean protein numbers are equal in both cases.
Suppressing the expression of a gene by a small RNA – rather than at the level of transcription – immediately relieves the genetic system from the effect of rare transcription events. The continuous transcription of both the target and the sRNA makes the kinetics much more homogeneous. The two effects of a small RNA – to reduce the rate of translation and to decrease the lifetime of the mRNA molecules – serve to decrease the protein burst size [25,27]. Together, these effects can keep the expression of a gene repressed by a small RNA at a steady low level; compare figures 2a and 2b. Small RNAs may also repress protein bursts that stem from possible bursts of transcriptional initiation events [30,46].
This difference in the noise properties may be very important in genetic circuits where a large burst of proteins will switch a cell from one stable state to another; see Box 2 for an illustrative example. In cases such as stress responses where unintentional entry into the alternative state may be harmful and spontaneous switching is to be avoided, sRNA-mediated regulation might possess a distinct advantage.
Small RNAs can provide rapid response while filtering transient signals
The threshold effect described above has a distinct temporal signature as well. Suppose that a cell is changed from an environment where the small RNA highly suppresses the target (so that αs > αm) to one where its expression is no longer required (αs = 0). The response in target gene expression will occur via two steps [26]. First, the sRNA pool present in the cell at the time of the transition is depleted. Only then can target mRNA and proteins can accumulate. Under the assumptions mentioned above, the transition time (tx) between these two steps can be shown to be approximately by tx = βs−1 log(αs/αm). This scenario is depicted in figure 3a (red), where, for comparison, the de-repressing kinetics for an analogous protein- regulated gene is also shown (blue).
Figure 3. the kinetic of de-repression.
(a) Consider a case where at t=0 the cells are transferred into an environment which suppressed expression of the repressor, either a protein transcription factor (blue) or a sRNA (red). Dynamics in the sRNA case is predicted to be more abrupt, with a delay period tX that can be tuned by αs. (b) Transient signals that suppress sRNA expression are not expected to have significant effects, if their duration is smaller than tX (≈30 min in this case).
Let us focus on two salient features, which make the de-repression kinetics in the two cases very different. First the response time tx can be reduced across the physiologically important range of time scales, from over an hour down to less than 10 min, simply by increasing the target expression αm. This is very different from the case of protein regulators, for which the waiting time is determined primarily by the protein turnover rate, which enforces a “speed limit” for protein-mediated response [47]. Small RNAs do not submit to this limit, since their targets can actively degrade them. Thus, sRNA-mediated regulation may be one of the viable regulatory strategies in situations where a rapid temporal response is physiologically important for the cell [28].
The other feature is that changes in the target transcription level do not propagate to changes in protein levels at times shorter than tx. Transient changes in the signals that control αm that persist for shorter periods are filtered by the small RNA regulator (figure 3b) [26]. This, for example, may be an important feature in SOS response, where the stress is transient in nature (persisting between the occurrence of DNA damage and its repair), and of regulation of iron homeostasis that must overcome iron pulses used by some hosts to fight pathogens [48].
Coordinated response of sRNA targets
Many of the known small RNAs have more than one mRNA target [49–51]. These targets are expected to have different affinities for association with the small RNA, resulting from different lengths of the base-pairing regions and different secondary structure. Different interaction strengths between sRNA and its targets should have little effect on the threshold when the target is expressed alone; however, when expressed simultaneously, the different mRNA species will compete for association with the same pool of sRNA molecules. The outcome of this competition necessarily depends on both the affinities and the abundances of the different mRNA species involved.
Generalizing the model (Box 1) to the case of multiple targets is straightforward [25,29]. One predicted consequence of this hierarchy is an ordering (or prioritization) of the targets. For example, consider a case of two targets, with k1≫k2. In this case, depending on the value of αs, the steady state may be such that either both targets are expressed (low αs), only target 2 is expressed (intermediate αs), or none. This prioritization is similar to the ordered transcriptional response of different genes controlled by a common transcription factor, where prioritization is governed by the equilibrium binding constants to various operator sites [52].
However, targets of a small RNA are not only affected by their regulator, they also deplete it. Thus, an increased expression of one target enhances the expression of another target, by reducing the level of its inhibitor [25]. Consider for example the case where the transcription rate of a sRNA regulator is larger than that of either targets (α1 and α2) but smaller than their sum (α1+α2) Due to the threshold-linear nature of the response (Figure 1), either target is expected to be silenced when transcribed alone, but can be expressed if both are transcribed together.
A more quantitative analysis of this case reveals a hierarchy between the expression of different targets. Expression of a strongly bound target is predicted to have a strong effect on the expression of weakly bound targets, while the expression of a weakly-bound target is predicted to have a much weaker effect on the former. This behavior has been observed for different targets of the small RNA RyhB in E. coli [25] The existence of this mechanism offers a possibility for sRNA regulatory systems to set up a complex hierarchy of responses based on the expression of the targets as well as the regulator.
Conclusions
Quantitative studies of gene regulation by small RNA in bacteria suggest that small RNAs establish a threshold for gene expression. This threshold can be dynamically tuned as a response to changes in environmental conditions. Consequently, fluctuations that stem from the stochastic nature of gene expression, as well as transient changes in environmental conditions, can be strongly suppressed.
Although a number of the novel features involving sRNA regulation have been demonstrated for specific systems, it remains to be shown how such features may be utilized in physiological responses. Regardless, these features and possibilities should be taken into account when analyzing the responses of systems involving sRNA regulation.
Box 1: Quantitative model for gene regulation by small RNA.
A quantitative study of gene regulation by small RNA starts with a simple mathematical model. In constructing such model, the most prominent feature is the possibility that the small RNA itself is consumed via the interaction between sRNAs and their mRNA targets. Two additional assumptions, namely that the association and dissociation rates occur rapidly, and that the RNA complex cannot be translated, lead to a minimal model which only accounts for the free small RNA and mRNA molecules. This model takes the form [16,25,31]
| (1) |
where αs, αm are the transcription rates of the sRNA, mRNA, βs, βm are the turnover rates for the RNA species alone, and k is the rate of coupled degradation of sRNA with its target. The parameter p accounts for the possibility that the degradation of the small RNA molecule due to binding may not be the same as that of the mRNA; it is set to 1 throughout this review as it does not make a qualitative difference to the results [25].
The transcription rates of the two RNAs can readily be changed dynamically by regulating the activity at their promoters. More hard-wired are the other parameters, namely the turnover rates of the two RNAs and their coupled degradation. Small RNAs that are involved in gene regulation are likely to be stable in the absence of their targets, as demonstrated for both cis and trans acting sRNAs [32–34]. Thus, the value of βs is dictated by the growth rate of the cell. Messenger RNAs in E. coli are typically unstable, with degradation rate of a few minutes. It is possible that mRNAs that are targeted by sRNAs are on the more stable side [23,32–35], say βm ≤ (10 min)−1. The binding rate k can depend both on sequence complementarity and secondary structure of the RNA molecules. The value of k has been estimated for several sRNA/target pairs to be around 1/50 (nM min)−1 [21,25,35,36]. Some authors have estimated this value to be larger; but since this value is already close to the diffusion-limited association rate for typical small proteins [37,38], we do not expect it can be much higher in vivo. In any case, the results presented here remain valid as long as k ≫ βmβs/αthreshold ~ 1/500 (nM/min)−1.
Box 2: small RNAs stabilize multi-stable genetic systems.
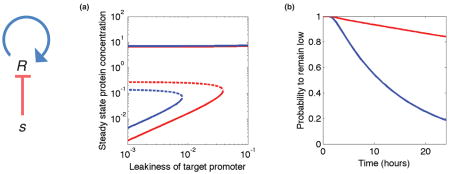
We consider as an example an auto-activated gene (R), that may be (slightly) repressed by a small RNA (s), as depicted on the left. In some cases, this system may sustain two stable states, of high or low expression. Leakiness in the expression of the target is the main reason why a low-expression state is typically unstable. (a) A bifurcation diagram for this system, with (red) and without (blue) the small RNA. Solid and dashed lines represent protein concentrations that are stable and unstable, resp. Note that in the presence of the small RNA, bi-stability is achieved over a wider range of leakiness. Conversely, the presence of the small RNA has almost no effect on the high state. (b) Even when the system is bi-stable, random bursts of protein may take the system from the low-expression state to the high-expression one. Thus, the probability to remain in the low state decreases with time. However, since small RNA serve to suppress these bursts, it decreases much slower in the presence of sRNA (red) than in its absence (blue).
Acknowledgments
We thank Mathew Scott, Ned Wingreen, and Johan Paulsson for discussions, and the experimental members of the Hwa lab, in particular Zhongge Zhang and Tom Kuhlman, for initiating the experimental studies. Our sRNA work is supported by NIH (Grant RO1GM77298), and by the NSF (Grant PHY-0822283) through the PFC-sponsored Center for Theoretical Biological Physics.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and recommended reading
- 1•.Storz G, Gottesman S. Versatile roles of small RNA regulators in bacteria. In: Gesteland RF, Cech TR, Atkins JF, editors. The RNA World. 3. Cold Spring Harbor Laboratory Press; 2006. pp. 567–594. A recent review of bacterial small RNA regulation, covering sRNAs encoded on plasmids, transposons, and the chromosome. [Google Scholar]
- 2.Marzi S, Fechter P, Chevalier C, Romby P, Geissmann T. RNA switches regulate initiation of translation in bacteria (*) Biol Chem. 2008 doi: 10.1515/bc.2008.055. [DOI] [PubMed] [Google Scholar]
- 3.Kleckner N. Regulation of transposition in bacteria. Annu Rev Cell Biol. 1990;6:297–327. doi: 10.1146/annurev.cb.06.110190.001501. [DOI] [PubMed] [Google Scholar]
- 4.Wagner EG, Simons RW. Antisense RNA control in bacteria, phages, and plasmids. Annu Rev Microbiol. 1994;48:713–742. doi: 10.1146/annurev.mi.48.100194.003433. [DOI] [PubMed] [Google Scholar]
- 5••.Paulsson J, Ehrenberg M. Noise in a minimal regulatory network: plasmid copy number control. Q Rev Biophys. 2001;34:1–59. doi: 10.1017/s0033583501003663. This is a comprehensive quantitative study of the role played by cis-acting small RNAs in controlling plasmid copy number. While the mechanism - and hence the details of the model - are somewhat different from those discussed here, small RNA also provide threshold behavior and noise suppression in this system. [DOI] [PubMed] [Google Scholar]
- 6.Gerdes K, Gultyaev AP, Franch T, Pedersen K, Mikkelsen ND. Antisense RNA-regulated programmed cell death. Annu Rev Genet. 1997;31:1–31. doi: 10.1146/annurev.genet.31.1.1. [DOI] [PubMed] [Google Scholar]
- 7.Sahagan BG, Dahlberg JE. A small, unstable RNA molecule of Escherichia coli: spot 42 RNA. I Nucleotide sequence analysis. J Mol Biol. 1979;131:573–592. doi: 10.1016/0022-2836(79)90008-1. [DOI] [PubMed] [Google Scholar]
- 8•.Aiba H, Matsuyama S, Mizuno T, Mizushima S. Function of micF as an antisense RNA in osmoregulatory expression of the ompF gene in Escherichia coli. J Bacteriol. 1987;169:3007–3012. doi: 10.1128/jb.169.7.3007-3012.1987. In this review, Aiba presents a detailed picture of the mechanism by which the best-studied class of small RNAs suppress the expression of their targets. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rivas E, Klein RJ, Jones TA, Eddy SR. Computational identification of noncoding RNAs in E. coli by comparative genomics. Curr Biol. 2001;11:1369–1373. doi: 10.1016/s0960-9822(01)00401-8. [DOI] [PubMed] [Google Scholar]
- 10.Tjaden B, Saxena RM, Stolyar S, Haynor DR, Kolker E, Rosenow C. Transcriptome analysis of Escherichia coli using high-density oligonucleotide probe arrays. Nucleic Acids Res. 2002;30:3732–3738. doi: 10.1093/nar/gkf505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wassarman KM, Repoila F, Rosenow C, Storz G, Gottesman S. Identification of novel small RNAs using comparative genomics and microarrays. Genes Dev. 2001;15:1637–1651. doi: 10.1101/gad.901001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gottesman S, McCullen CA, Guillier M, Vanderpool CK, Majdalani N, Benhammou J, Thompson KM, FitzGerald PC, Sowa NA, FitzGerald DJ. Small RNA regulators and the bacterial response to stress. Cold Spring Harb Symp Quant Biol. 2006;71:1–11. doi: 10.1101/sqb.2006.71.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Toledo-Arana A, Repoila F, Cossart P. Small noncoding RNAs controlling pathogenesis. Curr Opin Microbiol. 2007;10:182–188. doi: 10.1016/j.mib.2007.03.004. [DOI] [PubMed] [Google Scholar]
- 14.Prevost K, Salvail H, Desnoyers G, Jacques JF, Phaneuf E, Masse E. The small RNA RyhB activates the translation of shiA mRNA encoding a permease of shikimate, a compound involved in siderophore synthesis. Mol Microbiol. 2007;64:1260–1273. doi: 10.1111/j.1365-2958.2007.05733.x. [DOI] [PubMed] [Google Scholar]
- 15.Vanderpool CK, Gottesman S. Involvement of a novel transcriptional activator and small RNA in post-transcriptional regulation of the glucose phosphoenolpyruvate phosphotransferase system. Mol Microbiol. 2004;54:1076–1089. doi: 10.1111/j.1365-2958.2004.04348.x. [DOI] [PubMed] [Google Scholar]
- 16.Lenz DH, Mok KC, Lilley BN, Kulkarni RV, Wingreen NS, Bassler BL. The small RNA chaperone Hfq and multiple small RNAs control quorum sensing in Vibrio harveyi and Vibrio cholerae. Cell. 2004;118:69–82. doi: 10.1016/j.cell.2004.06.009. [DOI] [PubMed] [Google Scholar]
- 17.Geissmann TA, Touati D. Hfq, a new chaperoning role: binding to messenger RNA determines access for small RNA regulator. EMBO J. 2004;23:396–405. doi: 10.1038/sj.emboj.7600058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kawamoto H, Koide Y, Morita T, Aiba H. Base-pairing requirement for RNA silencing by a bacterial small RNA and acceleration of duplex formation by Hfq. Mol Microbiol. 2006;61:1013–1022. doi: 10.1111/j.1365-2958.2006.05288.x. [DOI] [PubMed] [Google Scholar]
- 19.Aiba H. Mechanism of RNA silencing by Hfq-binding small RNAs. Curr Opin Microbiol. 2007;10:134–139. doi: 10.1016/j.mib.2007.03.010. [DOI] [PubMed] [Google Scholar]
- 20.Darfeuille F, Unoson C, Vogel J, Wagner EG. An antisense RNA inhibits translation by competing with standby ribosomes. Mol Cell. 2007;26:381–392. doi: 10.1016/j.molcel.2007.04.003. [DOI] [PubMed] [Google Scholar]
- 21.Masse E, Escorcia FE, Gottesman S. Coupled degradation of a small regulatory RNA and its mRNA targets in Escherichia coli. Genes Dev. 2003;17:2374–2383. doi: 10.1101/gad.1127103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Simons RW, Kleckner N. Translational control of IS10 transposition. Cell. 1983;34:683–691. doi: 10.1016/0092-8674(83)90401-4. [DOI] [PubMed] [Google Scholar]
- 23.Hjalt TA, Wagner EG. Bulged-out nucleotides in an antisense RNA are required for rapid target RNA binding in vitro and inhibition in vivo. Nucleic Acids Res. 1995;23:580–587. doi: 10.1093/nar/23.4.580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Elf J, Paulsson J, Berg OG, Ehrenberg M. Near-critical phenomena in intracellular metabolite pools. Biophys J. 2003;84:154–170. doi: 10.1016/S0006-3495(03)74839-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25••.Levine E, Zhang Z, Kuhlman T, Hwa T. Quantitative characteristics of gene regulation by small RNA. PLoS Biol. 2007;5:e229. doi: 10.1371/journal.pbio.0050229. This detailed quantitative study of small RNA regulation in E. coli demonstrates how interplay between theoretical modeling and quantitative experiments may provide a unique perspective on the features of small RNAs. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26•.Legewie S, Dienst D, Wilde A, Herzel H, Axmann I. Small RNAs Establish Delays and Temporal Thresholds in Gene Expression. Biophys J. 2008 doi: 10.1529/biophysj.108.133819. Legewie et al use a simple model to interpret their previous experiments on the cyanobacterial sRNA IsrR, and suggest that sRNAs may filter transient signals. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Levine E, Huang M, Huang Y, Kuhlman T, Zhang Z, Hwa T. On noise and silence in gene regulation by small. RNA. 2008 [Google Scholar]
- 28.Shimoni Y, Friedlander G, Hetzroni G, Niv G, Altuvia S, Biham O, Margalit H. Regulation of gene expression by small non-coding RNAs: a quantitative view. Mol Syst Biol. 2007;3:138. doi: 10.1038/msb4100181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mitarai N, Andersson AM, Krishna S, Semsey S, Sneppen K. Efficient degradation and expression prioritization with small RNAs. Phys Biol. 2007;4:164–171. doi: 10.1088/1478-3975/4/3/003. [DOI] [PubMed] [Google Scholar]
- 30.Goyal S, Mehta P, Wingreen NS. A quantitative comparison of sRNA-based and protein-based gene regulation. Mol Syst Biol. 2008 doi: 10.1038/msb.2008.58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Elf J, Ehrenberg M. Fast evaluation of fluctuations in biochemical networks with the linear noise approximation. Genome Res. 2003;13:2475–2484. doi: 10.1101/gr.1196503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Masse E, Gottesman S. A small RNA regulates the expression of genes involved in iron metabolism in Escherichia coli. Proc Natl Acad Sci USA. 2002;99:4620–4625. doi: 10.1073/pnas.032066599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wagner EG, Altuvia S, Romby P. Antisense RNAs in bacteria and their genetic elements. Adv Genet. 2002;46:361–398. doi: 10.1016/s0065-2660(02)46013-0. [DOI] [PubMed] [Google Scholar]
- 34.Case CC, Roels SM, Jensen PD, Lee J, Kleckner N, Simons RW. The unusual stability of the IS10 anti-sense RNA is critical for its function and is determined by the structure of its stem-domain. Embo J. 1989;8:4297–4305. doi: 10.1002/j.1460-2075.1989.tb08616.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Argaman L, Altuvia S. fhlA repression by OxyS RNA: kissing complex formation at two sites results in a stable antisense-target RNA complex. J Mol Biol. 2000;300:1101–1112. doi: 10.1006/jmbi.2000.3942. [DOI] [PubMed] [Google Scholar]
- 36.Franch T, Petersen M, Wagner EG, Jacobsen JP, Gerdes K. Antisense RNA regulation in prokaryotes: rapid RNA/RNA interaction facilitated by a general U-turn loop structure. J Mol Biol. 1999;294:1115–1125. doi: 10.1006/jmbi.1999.3306. [DOI] [PubMed] [Google Scholar]
- 37.Koren R, Hammes GG. A kinetic study of protein-protein interactions. Biochemistry. 1976;15:1165–1171. doi: 10.1021/bi00650a032. [DOI] [PubMed] [Google Scholar]
- 38.Northrup SH, Erickson HP. Kinetics of protein-protein association explained by Brownian dynamics computer simulation. Proc Natl Acad Sci U S A. 1992;89:3338–3342. doi: 10.1073/pnas.89.8.3338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kawano M, Aravind L, Storz G. An antisense RNA controls synthesis of an SOS-induced toxin evolved from an antitoxin. Mol Microbiol. 2007;64:738–754. doi: 10.1111/j.1365-2958.2007.05688.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Fozo EM, Kawano M, Fontaine F, Kaya Y, Mendieta KS, Jones KL, Ocampo A, Rudd KE, Storz G. Repression of small toxic protein synthesis by the Sib and OhsC small RNAs. Mol Microbiol. 2008 doi: 10.1111/j.1365-2958.2008.06394.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.McAdams HH, Arkin A. Stochastic mechanisms in gene expression. Proc Natl Acad Sci U S A. 1997;94:814–819. doi: 10.1073/pnas.94.3.814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Maheshri N, O’Shea EK. Living with noisy genes: how cells function reliably with inherent variability in gene expression. Annu Rev Biophys Biomol Struct. 2007;36:413–434. doi: 10.1146/annurev.biophys.36.040306.132705. [DOI] [PubMed] [Google Scholar]
- 43.Cai L, Friedman N, Xie XS. Stochastic protein expression in individual cells at the single molecule level. Nature. 2006;440:358–362. doi: 10.1038/nature04599. [DOI] [PubMed] [Google Scholar]
- 44.Yu J, Xiao J, Ren X, Lao K, Xie XS. Probing gene expression in live cells, one protein molecule at a time. Science. 2006;311:1600–1603. doi: 10.1126/science.1119623. [DOI] [PubMed] [Google Scholar]
- 45.Ozbudak EM, Thattai M, Kurtser I, Grossman AD, van Oudenaarden A. Regulation of noise in the expression of a single gene. Nat Genet. 2002;31:69–73. doi: 10.1038/ng869. [DOI] [PubMed] [Google Scholar]
- 46.Golding I, Paulsson J, Zawilski SM, Cox EC. Real-time kinetics of gene activity in individual bacteria. Cell. 2005;123:1025–1036. doi: 10.1016/j.cell.2005.09.031. [DOI] [PubMed] [Google Scholar]
- 47.Rosenfeld N, Elowitz MB, Alon U. Negative autoregulation speeds the response times of transcription networks. J Mol Biol. 2002;323:785–793. doi: 10.1016/s0022-2836(02)00994-4. [DOI] [PubMed] [Google Scholar]
- 48.Fischbach MA, Lin H, Liu DR, Walsh CT. How pathogenic bacteria evade mammalian sabotage in the battle for iron. Nat Chem Biol. 2006;2:132–138. doi: 10.1038/nchembio771. [DOI] [PubMed] [Google Scholar]
- 49.Gottesman S. The small RNA regulators of Escherichia coli: roles and mechanisms. Annu Rev Microbiol. 2004;58:303–328. doi: 10.1146/annurev.micro.58.030603.123841. [DOI] [PubMed] [Google Scholar]
- 50.Lease RA, Smith D, McDonough K, Belfort M. The small noncoding DsrA RNA is an acid resistance regulator in Escherichia coli. J Bacteriol. 2004;186:6179–6185. doi: 10.1128/JB.186.18.6179-6185.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Masse E, Vanderpool CK, Gottesman S. Effect of RyhB small RNA on global iron use in Escherichia coli. J Bacteriol. 2005;187:6962–6971. doi: 10.1128/JB.187.20.6962-6971.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kalir S, McClure J, Pabbaraju K, Southward C, Ronen M, Leibler S, Surette MG, Alon U. Ordering genes in a flagella pathway by analysis of expression kinetics from living bacteria. Science. 2001;292:2080–2083. doi: 10.1126/science.1058758. [DOI] [PubMed] [Google Scholar]




