Abstract
Background
The orthoreoviruses are infectious agents that possess a genome comprised of 10 double-stranded RNA segments encased in two concentric protein capsids. Like virtually all RNA viruses, an RNA-dependent RNA polymerase (RdRp) enzyme is required for viral propagation. RdRp sequences have been determined for the prototype mammalian orthoreoviruses and for several other closely-related reoviruses, including aquareoviruses, but have not yet been reported for any avian orthoreoviruses.
Results
We determined the L2 genome segment nucleotide sequences, which encode the RdRp proteins, of two different avian reoviruses, strains ARV138 and ARV176 in order to define conserved and variable regions within reovirus RdRp proteins and to better delineate structure/function of this important enzyme. The ARV138 L2 genome segment was 3829 base pairs long, whereas the ARV176 L2 segment was 3830 nucleotides long. Both segments were predicted to encode λB RdRp proteins 1259 amino acids in length. Alignments of these newly-determined ARV genome segments, and their corresponding proteins, were performed with all currently available homologous mammalian reovirus (MRV) and aquareovirus (AqRV) genome segment and protein sequences. There was ~55% amino acid identity between ARV λB and MRV λ3 proteins, making the RdRp protein the most highly conserved of currently known orthoreovirus proteins, and there was ~28% identity between ARV λB and homologous MRV and AqRV RdRp proteins. Predictive structure/function mapping of identical and conserved residues within the known MRV λ3 atomic structure indicated most identical amino acids and conservative substitutions were located near and within predicted catalytic domains and lining RdRp channels, whereas non-identical amino acids were generally located on the molecule's surfaces.
Conclusion
The ARV λB and MRV λ3 proteins showed the highest ARV:MRV identity values (~55%) amongst all currently known ARV and MRV proteins. This implies significant evolutionary constraints are placed on dsRNA RdRp molecules, particularly in regions comprising the canonical polymerase motifs and residues thought to interact directly with template and nascent mRNA. This may point the way to improved design of anti-viral agents specifically targeting this enzyme.
Background
The avian reoviruses (ARVs) are members of the family Reoviridae, the only group of dsRNA viruses (out of seven dsRNA virus families) that infect mammals [1,2]. The ARVs are the prototypic members of syncytia-inducing, non-enveloped viruses within the Orthoreovirus genus. This genus is divided into 3 subgroups: non-syncytia-inducing mammalian reovirus (MRV; subgroup 1; the prototype of the whole genus), avian reovirus and Nelson Bay virus (subgroup 2), and baboon reovirus (subgroup 3) [3]. In contrast to the MRV, which are rarely associated with human pathology [2,4-6], the ARV are significant pathogens of poultry, and cause a variety of diseases, including infectious enteritis in turkeys [7], viral arthritis/tenosynovitis [8], "pale bird" and runting-stunting syndromes [9], and gastroenteritis, hepatitis, myocarditis, and respiratory illness in chickens [2,8,10].
Like MRV, ARV is a non-enveloped virus with 10 linear double-stranded RNA gene segments surrounded by a double concentric icosahedral capsid shell (inner shell [also called core] and outer shell) of 70–80 nm diameter [11,12]. The ARV genomic segments can be resolved into three size classes based on their electrophoretic mobilities, designated L (large), M (medium), and S (small) [11,12]. In total, the genomic composition includes 3 large segments (Ll, L2, L3), 3 medium sized segments (Ml, M2, M3), and 4 small segments (S1, S2, S3, S4). Nine of the segments are monocistronic and encode a single different protein [11-13] while S1 is tricistronic with partially overlapping open reading frames (ORFs) that encode for three proteins [14,15]. Although ARVs share many features with the prototypic MRVs, several notable differences exist including host range, pathogenicity, hemagglutination properties, and syncytium formation [11,12,16-21].
Genomic coding differences also exist between MRV and ARV. For example, although the ARV and MRV S1 genome segments encode homologous receptor-binding proteins [19,22,23], the ARV S1 genome segment encodes two additional ARV-specific gene products, one of which is responsible for ARV's unusual cell-cell fusion ability [14,15,24], whereas the MRV S1 segment encodes only one additional protein [25]. In addition, available data [12,26] suggest each of the homologous orthoreovirus λ-class proteins are encoded by different ARV and MRV L-class genome segments. Differences in the functional properties of homologous ARV and MRV proteins have also been reported. For example, two non-homologous dsRNA-binding proteins (the ARV σA core protein and the MRV σ3 major outer capsid protein) are predicted to regulate PKR activation [27,28] while the ARV σA core protein displays nucleoside triphosphate phosphohydrolase (NTPase) activity [29], ascribed to the non-homologous MRV μ2 [30] and λ1 [31] core proteins. Based on these early comparative studies, it seems likely that additional analysis of ARV will continue to broaden our understanding of the Reoviridae family, possibly leading to the identification of novel features that impact on the distinct biological and pathogenic properties of ARV.
Recent advances have allowed sequence determinations of a growing number of virus isolates. Many ARV and MRV genome segment sequences have been reported. In addition, the complete genomic sequences of three prototype strains of MRV have been completed [32-34]. In contrast, sequence information from ARV isolates is more limited. While the entire complement of S-class genome segments (for example, [14,15,35-39]) and M-class genome segments (for example, [40,41]) have been determined for some ARV clones, and sequence information is available for some ARV L1 and L3 genome segments [42,43], there is, at present, no sequence information for the ARV L2 genome segment. This segment is presumed to encode for the viral RNA-dependent RNA polymerase (RdRp) protein, an essential enzyme for RNA virus replication. Thus, we determined the genomic sequences of the ARV L2 genome segments from two different strains of ARV (ARV138 and ARV176) in order to expand the available ARV sequence database, determine sequences of the ARV RdRp protein, and to delineate conserved structure/function features of this key viral-encoded enzyme.
Methods
Cells and viruses
Avian reovirus strain 138 (ARV138) and strain 176 (ARV176) are laboratory stocks. Virus clones were amplified in the continuous quail cell line QM5 in Medium 199 (Gibco) supplemented to contain 7.5% fetal calf serum (Hyclone), 2 mM glutamine, 100 U/ml penicillin, 100 μg/ml streptomycin, and 1 μg/ml amphotericin B, essentially as previously described [44].
Sequencing the L2 genome segment
Genomic dsRNA was extracted from amplified virus P2 stocks with phenol/chloroform [45]. The extracted dsRNA were resolved in 10% SDS-PAGE and resolved L1, L2, and L3 segments separately excised. Individual segment gel bands were collected into microcentrifuge tubes, macerated, and incubated in 1–2 volumes of diffusion buffer (0.5 M ammonium acetate; 10 mM magnesium acetate; 1 mM EDTA, pH 8.0; 0.1% SDS) at 50°C for 30 minutes. The macerated gel pieces were pelleted by centrifugation at 10,000 × g for 1 min, supernatants were collected and dsRNA precipitated by ethanol. Each pellet was dried and resuspended in ddH2O for 3' ligation-based RT-PCR. All primers used for ligation, RT-PCR, and sequencing were synthesized by Invitrogen. An anchor primer, P-5' CTTATTTATTTGCGAGATGGTTATCATTTTAATTATCTCCATG 3'-Bio (5'-end phosphorylated and 3'-end biotin-blocked) was ligated to the 3' end of each genome segment, using T4 RNA ligase according to the manufacturer's instructions (Promega Inc., Madison, USA). Ligated products were precipitated by mixing with 1/2 volume of (30% PEG 8000 in 30 mM MgCl2), and centrifuged immediately at 10,000 × g for 30 minutes. The supernatants were removed and pellets were dried and dissolved in ddH2O for cDNA synthesis. Full-length cDNA copies of each L2 genome segment were synthesized using a primer (24-mer) complementary to the anchor primer by SuperScript™ II reverse transcriptase according to the manufacturer's instructions (Invitrogen). PCR amplification was performed using cDNA, a forward primer (i.e. primer used for RT), and a reverse primer, 5' ACCGAGGAGAGGgatgaataa 3', designed against highly conserved 3'-end nucleotide sequences of currently known consensus ARV L1 and L3 segment plus strands (shown in lower case) by Expand Long Template PCR System (Roche). PCR products used for DNA sequencing were gel purified using QIAquick® gel extraction kit according to the manufacturer's instructions (Qiagen).
DNA sequencing was performed in both directions by use of an ABI Prism BigDye Terminator v3.1 Cycle Sequencing Ready Reaction Kit (Applied Biosystems) and an Applied Biosystems Genetic Analyzer DNA Model 3100. The first two sequencing reactions were performed with the primers used for PCR amplification. Primers for subsequent reactions were designed from newly obtained sequences to completely sequence each full-length PCR product in both directions. Sequences nearer the ends of each segment were determined from PCR products that were amplified with a primer complementary to the anchor primer and an internal gene-specific primer. Sequences obtained from both directions were assembled and checked for accuracy with SeqMan® (Lasergene®, Version 7.1.0; DNASTAR, Inc.).
Sequence analyses
Sequences were compiled and analyzed using the Lasergene® software suite (Version 7.1.0; DNASTAR, Inc.) Pairwise sequence alignments were performed using the Wilbur-Lipman method [46] for highly divergent nucleotide sequences, the Martinez-NW method [47] for closely related nucleotide sequences, and the Lipman-Pearson method [48] for protein alignments in MegAlign® (Lasergene®). Multiple sequence alignments were performed using Clustal-W [49] and T-Coffee [50], and alignment adjustments were manually performed as needed in MegAlign®. Amino acid alignment images were adjusted in Adobe Photoshop 7.0 (Adobe®). Nucleotide compositions and protein molecular weights were calculated by DNA statistics and protein statistics, respectively, in EditSeq® (Lasergene®). Phylogenetic trees were constructed using Neighbor-Joining and tested with 1000 bootstrap replicates in MEGA version 4 [51].
3-D structural analyses
Molecular graphics coordinates of the mammalian reovirus (MRV) λ3 crystal structure (PDB # 1MUK; [52]), were manipulated with the UCSF Chimera package from the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco ([53]; supported by NIH P41 RR-01081). Resulting images were imported into Adobe Photoshop and assembled with Adobe Illustrator (Adobe).
Results
The sequences of genes that encode the RdRp protein have been determined for a number of members of the Reoviridae family of viruses (Table 1). However, this information was lacking for members of the avian orthoreovirus subgroup. We determined the sequences of two different strains' ARV L2 genome segments. The L2 genome segments of ARV138 and ARV176 were determined to be 3829 (GeneBank accession no. EU707935) and 3830 (GeneBank accession no. EU707936) nucleotides long, respectively (Table 2). The one-nucleotide length difference is attributed to the 5'-end of the non-translated region of the plus-strand, where ARV138 L2 contains a one-base deletion relative to ARV176 L2. No additional deletions or insertions were found elsewhere in the alignment. The nucleotide identity between ARV138 and ARV176 L2 genome segments is 85% (Table 3). BLAST searches indicated the ARV L2 genome segments were most similar to the mammalian reovirus (MRV) and aquareovirus (AqRV) L1 genome segments, which encode the RNA-dependent RNA polymerase [54,55]. Pairwise sequence comparisons between both of these newly-determined ARV genome segments and all currently available homologous MRV and AqRV L1 genome segments (see Table 1) showed a range of nucleotide and protein identity values. Preliminary comparative studies of all currently available AqRV L-class genome segments indicated that the grass carp reovirus (GCRV) and chum salmon reovirus (CSRV) L genes were the most distantly related amongst the AqRV (data not shown). Thus, although all currently available ARV, MRV, and AqRV L-class genome segments were aligned and compared in subsequent analyses, we limited presentation in subsequent tables and figures to these few most-distant clones for clarity. In addition, preliminary attempts to align the ARV138 and ARV176 L2 genome segments with homologous genes in other Reoviridae genera (ie. the Fijivirus Nilaparvata lugens, the Dinovernavirus Aedes pseudoscutellaris, the Coltivirus Eyach virus, the Orbivirus St. Croix River virus, the Seadornavirus Kadipiro virus, the Mimoreovirus Micromonas pusilla reovirus, and the currently unclassified virus Operophtera brumata reovirus) resulted in much lower identity values and significant gaps (data not shown); thus, these other more-distant genera were not included in subsequent analyses. Pairwise nucleotide sequence comparisons between ARV L2 and homologous MRV genome segments showed identities of ~55%, and pairwise nucleotide sequence comparisons of ARV L2 with AqRV homologues revealed ~48% identity (Table 3).
Table 1.
Nucleotide sequences used in this study
| Strain | GenBank Accession Number |
| ARVa | |
| 138 | EU707935 |
| 176 | EU707936 |
| MRVb | |
| T1L | NC_004271 |
| T2J | NC_004272 |
| T3D | EF494435 |
| T4N | AF368033 |
| BYD1 | DQ664184 |
| SC-A | DQ997719 |
| AqRVc | |
| GCRV | AF260512 |
| GCHV | AF284502 |
| GSRV | NC_005167 |
| AGCRV | NC_010585 |
| CSRV | NC_007583 |
| ASRV | EF434978 |
a ARV, avian reovirus.
b MRV, mammalian reovirus. T1L, type 1Lang; T2J, type 2 Jones; T3D, type 3 Dearing; T4N, type 4 Ndelle.
c AqRV, Aquareovirus. GCRV, Grass carp reovirus; GCHV, Grass carp hemorrhagic virus; GSRV, Golden shiner reovirus; AGCRV, American grass carp reovirus; CSRV, Chum salmon reovirus; ASRV, Atlantic salmon reovirus.
Table 2.
Genome-segment lengths, non-translated regions, and encoded proteins of ARV138 and ARV176
| Genome segment | Base pairsa | 5' NTRb | 3' NTR | ORFc | Codonsd | Protein | Molecular weight (kDa)e | |
| (no. of bases) | (no. of bases) | ARV138 | ARV176 | |||||
| L1f | 3958 | 20 | 56 | 21–3899 | 1293 | λA | 142.3 | 142.2 |
| L2 | 3829g | 13h | 36 | 14–3790i | 1259 | λB | 139.7 | 139.8 |
| L3f | 3907 | 12 | 37 | 13–3867 | 1285 | λC | 141.9 | 142.2 |
| M1 | 2283 | 12 | 72 | 13–2208 | 732 | μA | 82.0 | 82.2 |
| M2 | 2158 | 29 | 98 | 30–2057 | 676 | μB | 73.1 | 73.3 |
| M3 | 1996 | 24 | 64 | 25–1929 | 635 | μC | 70.9 | 70.8 |
| S1 | 1643 | 24 | 33 | 25–318 | 98 | p10 | 10.3 | 10.3 |
| 293–730 | 146 | p17 | 16.9 | 16.9 | ||||
| 630–1607 | 326 | σC | 34.9 | 34.8 | ||||
| S2 | 1324 | 15 | 58 | 16–1263 | 416 | σA | 46.1 | 46.1 |
| S3 | 1202 | 30 | 68 | 31–1131 | 367 | σB | 40.9 | 40.9 |
| S4 | 1192 | 23 | 65 | 24–1124 | 367 | σNS | 40.5 | 40.6 |
| Total | 23492j | |||||||
a Total nucleotides on each strand.
bNTR, non-translated region.
cNucleotide positions indicated for starting and ending codons.
dTotal number of amino acids in deduced protein.
eMolecular weight calculated from deduced protein and rounded to closest 0.1 kDa.
fUnpublished.
g3830 for ARV176.
h14 for ARV176.
i15–3791 for ARV176.
j 23,493 for ARV176.
Table 3.
Percent identities of the ARV L2 genome segments and homologous encoded proteins of MRV and Aquareovirusesa
| Strain | ARV138 | ARV176 | T1L | T2J | T3D | T4N | GCRV | CSRV |
| ARV138 | 98 | 55 | 55 | 55 | 55 | 42 | 41 | |
| ARV176 | 85 | 55 | 55 | 55 | 55 | 42 | 41 | |
| T1L | 55 | 55 | 92 | 99 | 97 | 42 | 41 | |
| T2J | 55 | 55 | 75 | 92 | 91 | 42 | 40 | |
| T3D | 55 | 55 | 96 | 76 | 98 | 42 | 41 | |
| T4N | 56 | 56 | 89 | 75 | 90 | 42 | 41 | |
| GCRV | 49 | 49 | 48 | 47 | 48 | 47 | 58 | |
| CSRV | 47 | 47 | 47 | 46 | 47 | 47 | 59 | |
a Percent amino acid identities indicated in upper triangle; percent nucleotide identities are in lower triangle, in bold.
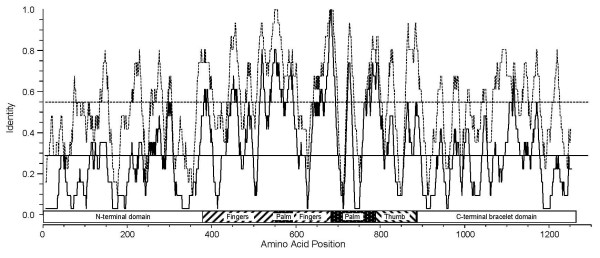
The predicted open reading frames for both ARV L2 segments were determined to be nucleotides 14–3790 for ARV138 L2 and 15–3791 for ARV176 L2, resulting in deduced λB proteins of 1259 residues (Table 2). The calculated molecular weights for ARV138 λB and ARV176 λB are ~140 kDa each (Table 2). The amino acid identity between the two ARV λB proteins is 97.5%, with no insertions or deletions relative to one another. ARV protein λB is the only ARV protein whose sequence has not been reported previously. Thus, completion of the L2 sequence in this study has allowed us to assign its function at the sequence level. Amino acid alignments of ARV λB, MRV λ3, and AqRV VP2 proteins revealed several regions of high amino acid identity (Fig. 1), many of which correspond to previously identified polymerase domains [56]. A large number of amino acids were completely conserved across all 14 currently known ARV, MRV, and AqRV RdRp protein sequences (Fig. 1, closed circles). Amino acid identities between ARV λB and homologous MRV λ3 or AqRV VP2 are ~55% and ~42%, respectively (Table 3), suggesting the ARV and MRV are more closely related to each other than either are to AqRV (also seen in phylogenetic analysis – Fig. 2), reflecting that ARV and MRV belong to different species in the Orthoreovirus genus [36] whereas AqRV are members of the different Aquareovirus genus in the Reoviridae family. Window-averaged analysis of ARV λB and MRV λ3 protein identities (Fig. 3, dashed lines) revealed several regions of high amino acid identity. The highest identity scores, with window-averaged identity values > 90%, were located within canonical polymerase regions, including "fingers" domains (MRV residues 452 – 467 and 514 – 530) "fingers"/"palm" interface domains (MRV residues 542 – 571 and 673 – 699), "palm" domains (MRV residues 725 – 738, which includes the GDD motif, which is common to all viral RNA-dependent RNA polymerases [57-59]), "thumb" domains (MRV residues 864 – 878), and an "undefined" domain (MRV residues 881 – 896). Addition of the AqRV VP2 protein to the above analyses provided additional information about potentially important conserved domains. Clustal-W (Fig. 1) and T-Coffee (data not shown) alignments identified 359 amino acid residues that were identical in the 6 aligned sequences (overall average identity = 28.3% Fig. 3, horizontal solid line]). There were numerous window-averaged regions of very low conservation, with most attributed to AqRV regions that were poorly conserved compared to corresponding ARV/MRV regions, a feature also noted in MRV:AqRV comparisons [60]. Three regions showed higher-than-average conservation in the ARV:MRV:AqRV alignments, with window-averaged identity values > 75%, suggesting these polymerase regions (ARV residues G516LRNQVQRRPRTIMP530, H542TLS/CADYINYHMNLSTTSGSAV563, and T677TTFPSGSTATSTEHTANNSTM698, that correspond to MRV residues G516LRNQVQRRPRSIMP530, H542TLTADYINYHMNLSTTSGSAV563, and T677TTFPSGSTATSTEHTANNSTM698, respectively) contain important structural/functional domains. The GDD motif was located within a region of slightly lower window-averaged scores (~60%), but in a sequence (in ARV) I724QxxYVCQGDDG735 that, apart from the residues at positions 726 and 727, were completely conserved in all 14 currently-available ARV, MRV, and AqRV RdRp sequences. In addition to the 359 identical residues found in all 6 sequences discussed above, blossum50 weighting alignments indicated that an additional 206 positions contained either identical amino acid residues or conservative substitutions in at least 4 of the 6 aligned sequences.
Figure 1.
Alignment of the deduced ARV138 and ARV176 λB amino acid sequences. All 14 currently available homologous ARV λB, MRV λ3, and AqRV VP2 proteins (determined for each clone shown in Table 1) were aligned, both by T-Coffee [50] (data not shown) and by Clustal-W [49], with only minor differences in the alignments created by different gap penalties (data not shown). Only the two most-distant ARV, MRV, and AqRV sequences (see text for details) are shown for clarity. Clones are: MRV – T1L (GenBank No. NC_004271) and T2J (GenBank No. NC_004272); ARV – ARV138 (GenBank No. EU707935) and ARV176 (GenBank No. EU707936); AqRV – Grass Carp reovirus (GCRV) (GenBank No. AF260512) and Chum Salmon reovirus (CSRV) (GenBank No. NC_007583). Amino acid residues that are identical in at least four of the sequences are indicated by black background shading. The single letter amino acid code is used. Previously identified polymerase domains (labeled I – III) [56] are indicated with solid horizontal lines above the sequences. Amino acid residues that are completely conserved in all 14 sequences are indicated by closed circles, and the GDD motif found in all polymerases is indicated by open circles, shown above the sequences.
Figure 2.
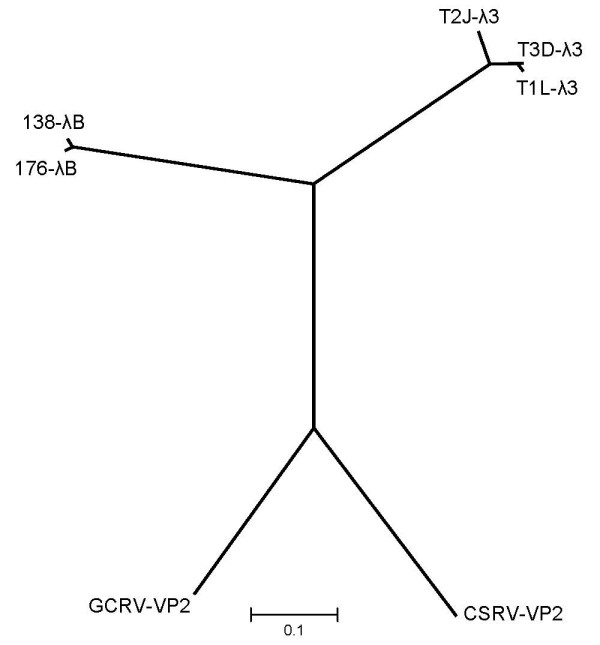
Phylogenetic tree analyses of the prototype ARV L2 genome segments and homologous genes in other reoviruses. Abbreviations are as defined in the legend to Fig. 1. Lines are proportional in length to nucleotide substitution. Alignments were performed by Neighbor-Joining and tested with 1000 bootstrap replicates in MEGA version 4 [51].
Figure 3.
Window-averaged scores for sequence identity among the ARV λB, AqRV VP2, and MRV λ3 RNA-dependent RNA polymerase proteins. To provide consistent weighting to the averaged scores, only the two most-distant clones from each of the three groups (ARV: ARV138 and ARV176; AqRV: GCRV and CSRV; MRV: T1L and T2J – see text for details) were used. Identity scores averaged over running windows of 15 amino acids and centered at consecutive amino acid positions are shown for ARV:MRV comparisons (dashed lines) and ARV:MRV:AqRV comparisons (solid line). The global identity scores for each of the compared sequence sets are indicated by the horizontal lines. Previously-identified enzymatic motifs are indicated with boxes below the plots.
Discussion
The atomic structure of few ARV proteins have been reported [61], and such high-resolution structures are not known for any λ-class ARV proteins. By contrast, atomic structures are known for most MRV proteins, including the RdRp [52]. Comparative sequence analyses described in this report have indicated that ARV and MRV RdRp proteins share ~55% amino acid identity, ARV and AqRV RdRp proteins share ~42% identity, and that only 359 (~28%) amino acids are completely conserved (identical) when ARV138, ARV176, MRV T1L, MRV T2J, AqRV CSRV, and AqRV GCRV are aligned (Fig. 1). Thus, to gain structure/function information about this key viral-encoded enzyme, ARV, MRV, and AqRV identical amino acids, conservative substitutions, and non-conservative substitutions were modeled in the MRV λ3 crystal structure (Fig. 4). This comparative analysis indicated that most non-conserved amino acids were located on the surfaces of the protein exposed to the core interior and in the N-terminal and C-terminal bracelet domains, whereas most identical amino acids and conservative substitutions were located within canonical fingers, palm, and thumb polymerase motifs, particularly those lining channels used by template and nascent RNA during transcription (Fig. 4). Similar observations had been reported from MRV:AqRV comparisons [60] and our results support and extend these earlier observations. As was previously reported from MRV:AqRV comparisons [60], conserved residues surround the GDD motif and additional residues shown to be important for a variety of polymerase functions are also conserved, including Arg522, Arg523, Arg525, Ala587 (which are needed to properly position the incoming NTP triphosphate), Ile527 and Pro529 (needed to help position template nucleosides), Thr557, Ser558, Gly559, Ser560, and Val562 (portions of a loop that maintains priming NTP), and Asp589, Ser681, and Gln731 (specifies ribonucleotides). Each of these residues is located one amino acid more N-terminal in the MRV sequence (ie. ARV Arg522 = MRV Arg523 and all (as well as numerous others) are completely conserved in all 14 currently available ARV, MRV, and AqRV RdRp sequences (Fig. 1, indicated by closed circles). In addition, our comparative analyses indicated many identical amino acids and conservative substitutions were located on the surface of the protein that is predicted to interact with the core shell [62]. This might imply that conserved domains are needed to help tether the RdRp to the underside of the core shell. This hypothesis could be tested by extending such ARV:MRV:AqRV sequence comparisons to the other core proteins.
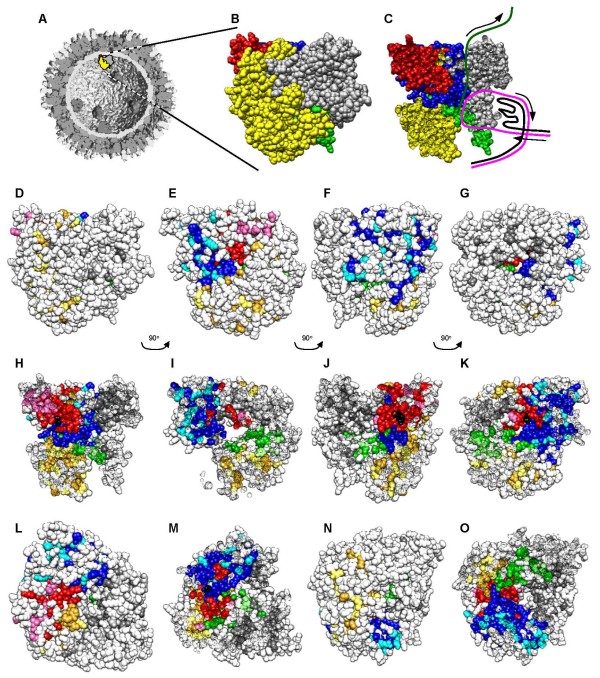
Figure 4.
Localization of conserved, non-conserved, and identical amino acids in ARV, MRV, and AqRV RdRp proteins. The MRV λ3 crystal structure (PDB # 1MUK [52]) was manipulated with Chimera [53]. A, Low-resolution, cutaway model of the reovirus core structure (modified from [26] with permission). B, Blow-up of indicated λ3 molecule in 'A', and C, cut-away of "B" with presumptive paths of genomic (+) RNA (black line), template genomic (-) RNA (magenta line) and nascent mRNA (dark green line) shown (adapted from and as described in [62]); Specific motifs in 'B' – 'O' are color-coded, with N-terminal region in yellow, C-terminal "bracelet" in grey, and canonical polymerase "fingers", "palm", and "thumb" depicted in blue, red, and green, respectively. D, Same as 'B', but in "D" – "O", amino acids that are identical in all 6 ARV, MRV, and AqRV sequences (see Fig. 1) are shown in darker versions of each motif color (goldenrod, dim grey, blue, red, and green, respectively), amino acids that represent conservative substitutions (as determined by Blossum50 matrix) are shown in lighter versions of each motif color (yellow, medium grey, cyan, hotpink, and light green, respectively), and non-conserved amino acids are shown in white. The canonical GDD motif is depicted in black. D - G, represent successive 90° rotations counter-clockwise around vertical axis, of entire RdRp protein, to correspond to front (as depicted in "A"), left side, back, and right side. H - K, represent same views as "D - G", respectively, but with the front approximate half of each view removed. L and N, represent top and bottom view, respectively, of RdRp molecule. M, represents top view, after upper approximately 40% of view removed, and O, represents bottom view, after lower approximately half of view removed. The top surface depicted in "L" is believed to interact with the λ-class core shell protein.
In conclusion, we report the first sequence analysis of the avian reovirus RdRp gene and protein. The ARV λB and MRV λ3 proteins showed the highest ARV:MRV identity values (~55%) amongst currently known ARV and MRV proteins, suggesting significant evolutionary constraints are placed on dsRNA RdRp molecules, particularly in regions comprising the canonical polymerase motifs and residues thought to interact directly with template and nascent mRNA.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
WX performed the experiments and analyses and WX and KC wrote the manuscript.
Acknowledgments
Acknowledgements
We thank members of our laboratory for critical reviews of this manuscript and Kolawole Opanubi for expert technical assistance. This research was supported by grant FRN-11630 from the Canadian Institutes of Health Research.
Contributor Information
Wanhong Xu, Email: wanhongxu@hotmail.com.
Kevin M Coombs, Email: kcoombs@ms.umanitoba.ca.
References
- Mertens P. The dsRNA viruses. Virus Research. 2004;101:3–13. doi: 10.1016/j.virusres.2003.12.002. [DOI] [PubMed] [Google Scholar]
- Schiff LA, Nibert ML, Tyler KL. Orthoreoviruses and their replication. In: Knipe DM, Howley PM, editor. Fields Virology. Philadelphia: Lippincott Williams & Wilkins; 2007. pp. 1853–1915. [Google Scholar]
- Chappell JD, Duncan R, Mertens PPC, Dermody TS. "Genus Orthoreovirus". In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA, editor. Virus Taxonomy Eighth Report of the International Committee on Taxonomy of Viruses. San Diego, CA: Elsevier Inc; 2005. [Google Scholar]
- Johansson PJ, Sveger T, Ahlfors K, Ekstrand J, Svensson L. Reovirus type 1 associated with meningitis. Scand J Infect Dis. 1996;28:117–120. doi: 10.3109/00365549609049060. [DOI] [PubMed] [Google Scholar]
- Hermann L, Embree J, Hazelton P, Wells B, Coombs K. Reovirus type 2 isolated from cerebrospinal fluid. Ped Infect Dis J. 2004;23:373–375. doi: 10.1097/00006454-200404000-00026. [DOI] [PubMed] [Google Scholar]
- Tyler KL, Barton ES, Ibach ML, Robinson C, Campbell JA, O'Donnell SM, et al. Isolation and molecular characterization of a novel type 3 reovirus from a child with meningitis. J Infect Dis. 2004;189:1664–1675. doi: 10.1086/383129. [DOI] [PubMed] [Google Scholar]
- Gershowitz A, Wooley RW. Characterization of two reoviruses isolated from turkeys with infectious enteritis. Avian Dis. 1973;17:406–414. doi: 10.2307/1589225. [DOI] [PubMed] [Google Scholar]
- Olson NO. "Reovirus infections". In: Hofstad MS, editor. Diseases of poultry. Ames, Iowa: Iowa State University Press; 1978. [Google Scholar]
- Kouwenhoven B, Vertommen M, Eck JHv. Runting and leg weakness in broilers: involvement of infectious factors. Vet Sci Commun. 1978;2:253–259. doi: 10.1007/BF02291456. [DOI] [Google Scholar]
- Rosenberger JK, Olson NO. "Reovirus infections". In: Calnek BW, Burnes MJ, Beard CW, Reid WM, Yoder HW, editor. Diseases of poultry. Ames, Iowa: Iowa State University Press; 1991. [Google Scholar]
- Spandidos DA, Graham AF. Physical and chemical characterization of an avian reovirus. J Virol. 1976;19:968–976. doi: 10.1128/jvi.19.3.968-976.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benavente J, Martinez-Costas J. Avian reovirus: Structure and biology. Virus Research. 2007;123:105–119. doi: 10.1016/j.virusres.2006.09.005. [DOI] [PubMed] [Google Scholar]
- Gouvea VS, Schnitzer TJ. Polymorphism of the genomic RNAs among the avian reoviruses. J Gen Virol. 1982;61:87–91. doi: 10.1099/0022-1317-61-1-87. [DOI] [PubMed] [Google Scholar]
- Bodelon G, Labrada L, Martinez-Costas J, Benavente J. The avian reovirus genome segment S1 is a functionally tricistronic gene that expresses one structural and two nonstructural proteins in infected cells. Virology. 2001;290:181–191. doi: 10.1006/viro.2001.1159. [DOI] [PubMed] [Google Scholar]
- Shmulevitz M, Yameen Z, Dawe S, Shou J, O'Hara D, Holmes I, et al. Sequential partially overlapping gene arrangement in the tricistronic S1 genome segments of avian reovirus and Nelson Bay reovirus: implications for translation initiation. J Virol. 2002;76:609–618. doi: 10.1128/JVI.76.2.609-618.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnitzer TJ. Protein coding assignment of the S genes of the avian reovirus S1133. Virology. 1985;141:167–170. doi: 10.1016/0042-6822(85)90194-1. [DOI] [PubMed] [Google Scholar]
- Ni Y, Ramig RF. Characterization of avian reovirus-induced cell fusion: the role of viral structural proteins. Virology. 1993;194:705–714. doi: 10.1006/viro.1993.1311. [DOI] [PubMed] [Google Scholar]
- Theophilos MB, Huang JA, Holmes IH. Avian reovirus sigma C protein contains a putative fusion sequence and induces fusion when expressed in mammalian cells. Virology. 1995;208:678–684. doi: 10.1006/viro.1995.1199. [DOI] [PubMed] [Google Scholar]
- Martinez-Costas J, Grande A, Varela R, Garcia-Martinez C, Benavente J. Protein architecture of avian reovirus S1133 and identification of the cell attachment protein. J Virol. 1997;71:59–64. doi: 10.1128/jvi.71.1.59-64.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones RC. Avian reovirus infections. Rev Sci Tech. 2000;19:614–625. doi: 10.20506/rst.19.2.1237. [DOI] [PubMed] [Google Scholar]
- Zhang X, Tang J, Walker SB, O'Hara D, Nibert ML, Duncan R, et al. Structure of avian orthoreovirus virion by electron cryomicroscopy and image reconstruction. Virology. 2005;343:25–35. doi: 10.1016/j.virol.2005.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner HL, Ault KA, Fields BN. Interaction of reovirus with cell surface receptors. I. Murine and human lymphocytes have a receptor for the hemagglutinin of reovirus type 3. J Immunol. 1980;124:2143–2148. [PubMed] [Google Scholar]
- Grande A, Costas C, Benavente J. Subunit composition and conformational stability of the oligomeric form of the avian reovirus cell-attachment protein sigmaC. J Gen Virol. 2002;83:131–139. doi: 10.1099/0022-1317-83-1-131. [DOI] [PubMed] [Google Scholar]
- Shmulevitz M, Duncan R. A new class of fusion-associated small transmembrane (FAST) proteins encoded by the non-enveloped fusogenic reoviruses. EMBO J. 2000;19:902–912. doi: 10.1093/emboj/19.5.902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munemitsu SM, Atwater JA, Samuel CE. Biosynthesis of reovirus-specified polypeptides. Molecular cDNA cloning and nucleotide sequence of the reovirus serotype 1 Lang strain bicistronic s1 mRNA which encodes the minor capsid polypeptide sigma 1a and the nonstructural polypeptide sigma 1bNS. Biochem Biophys Res Commun. 1986;140:508–514. doi: 10.1016/0006-291X(86)90761-8. [DOI] [PubMed] [Google Scholar]
- Dryden KA, Coombs KM, Yeager M. The structure of orthoreoviruses. In: Patton JT, editor. Segmented Double-stranded RNA Viruses: Structure and Molecular Biology. Horizon Press; 2008. pp. 3–25. [Google Scholar]
- Schiff LA, Nibert ML, Co MS, Brown EG, Fields BN. Distinct binding sites for zinc and double-stranded RNA in the reovirus outer capsid protein sigma 3. Mol Cell Biol. 1988;8:273–283. doi: 10.1128/mcb.8.1.273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez-Lopez C, Martinez-Costas J, Esteban M, Benavente J. Evidence that avian reovirus sigmaA protein is an inhibitor of the double-stranded RNA-dependent protein kinase. J Gen Virol. 2003;84:1629–1639. doi: 10.1099/vir.0.19004-0. [DOI] [PubMed] [Google Scholar]
- Yin HS, Su YP, Lee LH. Evidence of nucleotidyl phosphatase activity associated with core protein sigma A of avian reovirus S1133. Virology. 2002;293:379–385. doi: 10.1006/viro.2001.1292. [DOI] [PubMed] [Google Scholar]
- Noble S, Nibert ML. Core protein mu2 is a second determinant of nucleoside triphosphatase activities by reovirus cores. J Virol. 1997;71:7728–7735. doi: 10.1128/jvi.71.10.7728-7735.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bisaillon M, Bergeron J, Lemay G. Characterization of the nucleoside triphosphate phosphohydrolase and helicase activities of the reovirus lambda1 protein. J Biol Chem. 1997;272:18298–18303. doi: 10.1074/jbc.272.29.18298. [DOI] [PubMed] [Google Scholar]
- Wiener JR, Bartlett JA, Joklik WK. The sequences of reovirus serotype 3 genome segments M1 and M3 encoding the minor protein mu 2 and the major nonstructural protein mu NS, respectively. Virology. 1989;169:293–304. doi: 10.1016/0042-6822(89)90154-2. [DOI] [PubMed] [Google Scholar]
- Breun LA, Broering TJ, McCutcheon AM, Harrison SJ, Luongo CL, Nibert ML. Mammalian reovirus L2 gene and lambda2 core spike protein sequences and whole-genome comparisons of reoviruses type 1 Lang, type 2 Jones, and type 3 Dearing. Virology. 2001;287:333–348. doi: 10.1006/viro.2001.1052. [DOI] [PubMed] [Google Scholar]
- Yin P, Keirstead ND, Broering TJ, Arnold MM, Parker JSL, Nibert ML, et al. Comparisons of the M1 genome segments and encoded μ2 proteins of different reovirus isolates. Virol J. 1 doi: 10.1186/1743-422X-1-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiu CJ, Lee LH. Cloning and nucleotide sequencing of the S4 genome segment of avian reovirus S1133. Arch Virol. 1997;142:2515–2520. doi: 10.1007/s007050050258. [DOI] [PubMed] [Google Scholar]
- Duncan R. Extensive sequence divergence and phylogenetic relationships between the fusogenic and nonfusogenic orthoreoviruses: a species proposal. Virology. 1999;260:316–328. doi: 10.1006/viro.1999.9832. [DOI] [PubMed] [Google Scholar]
- Liu HJ, Huang PH. Sequence and phylogenetic analysis of the sigma A-encoding gene of avian reovirus. J Virol Meth. 2001;98:99–107. doi: 10.1016/S0166-0934(01)00328-7. [DOI] [PubMed] [Google Scholar]
- Kapczynski DR, Sellers HS, Simmons V, Schultz-Cherry S. Sequence analysis of the S3 gene from a turkey reovirus. Virus Genes. 2002;25:95–100. doi: 10.1023/A:1020130410601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sellers HS, Linnemann EG, Pereira L, Kapczynski DR. Phylogenetic analysis of the sigma 2 protein gene of turkey reoviruses. Avian Dis. 2004;48:651–657. doi: 10.1637/7181-032304R. [DOI] [PubMed] [Google Scholar]
- Touris-Otero F, Cortez-San Martín M, Martinez-Costas J, Benavente J. Avian reovirus morphogenesis occurs within viral factories and begins with the selective recruitment of sigma NS and lambda A to mu NS inclusions. J Mol Biol. 2004;341:361–374. doi: 10.1016/j.jmb.2004.06.026. [DOI] [PubMed] [Google Scholar]
- Noad L, Shou JY, Coombs KM, Duncan R. Sequences of avian reovirus M1, M2 and M3 genes and predicted structure/function of the encoded mu proteins. Virus Research. 2006;116:45–57. doi: 10.1016/j.virusres.2005.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiao J, Martinez-Costas J, Benavente J, Vakharia VN. Cloning, expression, and characterization of avian reovirus guanylyltransferase. Virology. 2002;296:288–299. doi: 10.1006/viro.2002.1427. [DOI] [PubMed] [Google Scholar]
- Shen PC, Chiou YF, Liu HJ, Song CH, Su YP, Lee LH. Genetic variation of the lambda A and lambda C protein encoding genes of avian reoviruses. Res Vet Sci. 2007;83:394–402. doi: 10.1016/j.rvsc.2007.01.002. [DOI] [PubMed] [Google Scholar]
- Patrick M, Duncan R, Coombs KM. Generation and genetic characterization of avian reovirus temperature-sensitive mutants. Virology. 2001;284:113–122. doi: 10.1006/viro.2001.0915. [DOI] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. 2. Cold Spring Harbor: Cold Spring Harbor Laboratory; 1989. [Google Scholar]
- Wilbur WJ, Lipman DJ. Rapid Similarity Searches of Nucleic-Acid and Protein Data Banks. Proc Natl Acad Sci (USA) 1983;80:726–730. doi: 10.1073/pnas.80.3.726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinez HM. An Efficient Method for Finding Repeats in Molecular Sequences. Nucl Acids Res. 1983;11:4629–4634. doi: 10.1093/nar/11.13.4629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lipman DJ, Pearson WR. Rapid and Sensitive Protein Similarity Searches. Science. 1985;227:1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Notredame C, Higgins DG, Heringa J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J Mol Biol. 2000;302:205–217. doi: 10.1006/jmbi.2000.4042. [DOI] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- Tao Y, Farsetta DL, Nibert ML, Harrison SC. RNA synthesis in a cage – Structural studies of reovirus polymerase lambda3. Cell. 2002;111:733–745. doi: 10.1016/S0092-8674(02)01110-8. [DOI] [PubMed] [Google Scholar]
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF chimera – A visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–1612. doi: 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- Starnes MC, Joklik WK. Reovirus protein lambda 3 is a poly(C)-dependent poly(G) polymerase. Virology. 1993;193:356–366. doi: 10.1006/viro.1993.1132. [DOI] [PubMed] [Google Scholar]
- Fang Q, Attoui H, Cantaloube JF, Biagini P, Zhu Z, de Micco P, et al. Sequence of genome segments 1, 2, and 3 of the grass carp reovirus (Genus Aquareovirus, family Reoviridae) Biochem Biophys Res Commun. 2000;274:762–766. doi: 10.1006/bbrc.2000.3215. [DOI] [PubMed] [Google Scholar]
- Bisaillon M, Lemay G. Computational sequence analysis of mammalian reovirus proteins. Virus Genes. 1999;18:13–37. doi: 10.1023/A:1008013117929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morozov SY. A possible relationship of reovirus putative RNA polymerase to polymerases of positive-strand RNA viruses. Nucleic Acids Res. 1989;17:5394. doi: 10.1093/nar/17.13.5394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruenn JA. Relationships Among the Positive Strand and Double-Strand Rna Viruses As Viewed Through Their Rna-Dependent Rna-Polymerases. Nucl Acids Res. 1991;19:217–226. doi: 10.1093/nar/19.2.217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruenn JA. A structural and primary sequence comparison of the viral RNA-dependent RNA polymerases. Nucl Acids Res. 2003;31:1821–1829. doi: 10.1093/nar/gkg277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim J, Tao Y, Reinisch KM, Harrison SC, Nibert ML. Orthoreovirus and Aquareovirus core proteins: conserved enzymatic surfaces, but not protein-protein interfaces. Virus Research. 2004;101:15–28. doi: 10.1016/j.virusres.2003.12.003. [DOI] [PubMed] [Google Scholar]
- Guardado-Calvo P, Vazquez-Iglesias L, Martinez-Costas J, Llamas-Saiz AL, Schoehn G, Fox GC, et al. Crystal structure of the avian reovirus inner capsid protein sigmaA. J Virol. 2008;82:11208–11216. doi: 10.1128/JVI.00733-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Walker SB, Chipman PR, Nibert ML, Baker TS. Reovirus polymerase lambda 3 localized by cryo-electron microscopy of virions at a resolution of 7.6 angstrom. Nature Structural Biology. 2003;10:1011–1018. doi: 10.1038/nsb1009. [DOI] [PMC free article] [PubMed] [Google Scholar]