Abstract
In Salmonella enterica serovar Typhimurium, gmk encodes guanylate kinase, an essential enzyme involved in the synthesis and salvage of guanine nucleotides. Here we report the isolation of a mutation in gmk that results in a nutritional requirement for adenine at low temperature. Comparisons of kinetic parameters from the wild-type and mutant Gmk enzymes revealed that the mutant enzyme had a more than 20-fold-higher Km for ATP than the wild-type enzyme. The growth dependence of the mutant on temperature and/or adenine could not be explained as a direct result of this kinetic difference. We propose a model in which previously described regulatory effects of GMP are responsible for these phenotypes.
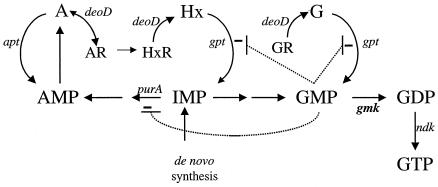
The biochemistry of de novo purine nucleotide biosynthesis and salvage is a broadly conserved and well-understood component of cellular metabolism. The first nucleotide formed in the purine biosynthetic pathway of Salmonella enterica is IMP. The synthesis of GTP from IMP involves four steps, with the intermediate formation of GMP and GDP. The enzyme guanylate kinase (Gmk) is a component of this pathway, catalyzing the transfer of the terminal phosphoryl group of ATP to the acceptor molecule GMP (or dGMP) (Fig. 1).
FIG. 1.
Simplified schematic representation of purine biosynthesis and salvage. The enzymes relevant to the processes discussed herein are identified by gene symbols below the reactions they catalyze; guanylate kinase (gmk) is shown in bold. Sites of allosteric inhibition by GMP that are significant for the model proposed are indicated. Abbreviations: A, adenine; G, guanine; Hx, hypoxanthine; HxR, inosine; AR, adenosine; GR, guanosine; apt, adenine phosphoribosyltransferase; gpt, guanine phosphoribosyltransferase; deoD, purine nucleoside phosphorylase; ndk, nucleoside diphosphate kinase. (While the DeoD-catalyzed reaction is reversible, the phosphorolysis of adenosine has been shown to be of minor quantitative importance.)
As an essential enzyme in the synthesis and salvage of guanine nucleotides, guanylate kinase has been the subject of numerous investigations. In addition, the biomedical role of human Gmk in the activation of antiviral agents has attracted significant attention (2, 13, 14). As a result, a large amount of information has amassed about the biochemistry, genetics, and regulation of these enzymes for a variety of organisms (2, 5, 6, 8, 12, 13, 15, 17, 20, 23) and a high-resolution structure for the Saccharomyces cerevisiae enzyme (GUK1) is available (21). Most guanylate kinases have significant sequence similarity, and the conserved motifs allow the prediction of substrate binding sites. Near the N terminus of the proteins is a canonical nucleoside triphosphate binding motif, GXXXXGK, reflecting the phosphate binding loop (P-loop) that is present in many ATP/GTP binding proteins. This site has been assigned to ATP binding, and the GMP binding site was shown to contain additional conserved residues just C terminal to the P-loop (21).
Since the activity of Gmk is essential for growth (e.g., null mutations are lethal), few studies have probed the consequences of altered guanylate kinase activity for cellular metabolism. Mutations in the GUK1 gene of S. cerevisiae were identified for their ability to bypass the purine biosynthetic gene repression normally caused by exogenous adenine (14). In this study, multiple metabolic phenotypes were described for strains that had a detectable, but >10-fold-decreased, level of GUK activity. The authors attributed the phenotypes of these mutants to the accumulation of the substrate GMP that resulted from the decrease in GUK activity (14). The molecular nature of the causative mutations was not explored in the yeast study, and potential kinetic differences in the mutant enzymes were not addressed.
We report the identification of a mutation in the Salmonella gene that encodes guanylate kinase (gmk). This mutation causes a cold-sensitive growth defect that can be corrected by the addition of exogenous adenine. The mutant protein displayed a significantly increased apparent Km for ATP, yet this difference could not fully account for the growth behavior with changing temperature. On the basis of the results reported herein and the known regulatory properties of enzymes involved in the biosynthesis and salvage of purines, a model to explain the role of temperature and adenine is proposed.
Initial observations.
In the course of screening mutants generated by localized hydroxylamine mutagenesis (11) around the pyrE locus, a cold-sensitive mutant was identified. Phenotypic characterization showed that this mutant strain (DM922) was able to grow at 37°C on minimal medium but required exogenous adenine for growth at 30°C.
A requirement for adenine at low temperature is caused by a mutation in gmk.
A plasmid clone able to complement the growth defect of strain DM922 was isolated from a plasmid library of Sau3A partially digested chromosomal DNA from S. enterica. The complementing plasmid contained a 3.5-kb insert. Subsequent cloning of a 1.2-kb fragment into the mid-copy vector pSU19 (16) provided a plasmid (pMK1) that was able to complement the growth defect in DM922. Sequence analysis determined that this fragment contained the entire gmk locus and no other complete open reading frames (data not shown). In Salmonella and Escherichia coli, the gmk locus is the promoter proximal gene in the spoT operon and encodes the essential protein guanylate kinase (8). Since gmk is <10 kb from the pyrE locus, growth allowed by plasmid pMK1 was considered to be due to functional complementation rather than to multicopy suppression by an unlinked gene.
The gmk gene was amplified by PCR from the wild-type strain LT2 and mutant strain DM922. Comparison of the nucleotide sequences revealed that the gmk gene from DM922 contained a single base substitution (G→A) consistent with the hydroxylamine mutagenesis. This base substitution at nucleotide 56 of the coding sequence results in a serine-to-asparagine (S19N) change in the protein at a location immediately adjacent to the ATP binding site of the protein (13).
A pair of isogenic strains differing at the gmk locus were constructed and subjected to liquid growth analyses at different temperatures. The data from representative experiments are shown in Table 1. The gmk101 strain had a significant growth defect at 30°C in minimal medium. Growth was restored by the addition of adenine (>0.1 mM) or by elevated temperature. The addition of other purines, including adenosine, guanosine, and hypoxanthine, had no effect on the growth of the gmk101 mutant at 30°C (data not shown).
TABLE 1.
Cold-sensitive adenine requirement of gmk101 mutant
| Strain | Genotype | Specific growth rate in indicated medium ata:
|
|||
|---|---|---|---|---|---|
| 30°
|
37°C
|
||||
| Min | Ade | Min | Ade | ||
| DM7062 | WTb | 0.31 | 0.28 | 0.36 | 0.33 |
| DM7061 | gmk101 | 0.11 | 0.34 | 0.25 | 0.31 |
Thiamine (100 nM) was present in minimal glucose medium (Min) and the same medium with adenine (Ade). Overnight cultures of the respective strains were grown in nutrient broth, pelleted, and resuspended in NaCl prior to inoculation (1:25) into the designated medium. Growth was monitored as absorbance at 650 nm. Specific growth rate (μ) was determined according to the equation ln(X/X0)/T, where X is A650, X0 is A650 at time zero, and T is time (in hours).
WT, wild type.
The S19N variant of Gmk has an increased Km for ATP.
The biochemical characterization of the mutant and wild-type enzymes was performed. The wild-type and mutant gmk genes were cloned into pET-20b (Novagen, Madison, Wis.) to generate a 6-His tag fusion to the carboxy terminus of the protein. Gmk and GmkS19N proteins were overexpressed in E. coli BL21(DE3). The resulting 24-kDa proteins were purified using standard nickel column affinity column chromatography and determined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis to be >95% pure. Guanylate kinase activity was measured in an assay coupled to lactate dehydrogenase activity, as has been previously described (8, 17). Initial velocity measurements were made using 5.1 nmol of either mutant or wild-type enzyme per reaction. The results of representative experiments are shown in Table 2 and Fig. 2. Data were analyzed and fitted to the Michaelis-Menten equation using the enzyme kinetics template of a Prism program (GraphPad, San Diego, Calif.). Pseudo first-order kinetics were used to determine the apparent Km and Vmax values presented. The kinetic constants found for the wild-type enzyme were consistent with those reported for the E. coli enzyme (17). As shown in Table 2, the apparent GMP Km values of the wild-type and mutant enzymes (∼10−4 M) were not significantly different. Kinetic analyses to determine the Km for ATP were carried out at 30 and 37°C. At both temperatures, the two proteins had comparable Vmax values. In contrast, there was a significant difference between the mutant and wild-type enzymes in their apparent Km values for ATP. The data showed that a more than 20-fold difference in Km values for ATP existed between the two enzymes. This finding was consistent with a model in which the larger asparagine residue of the mutant protein obstructed the ATP binding site and the serine residue of the wild-type protein did not. The experimental data used to determine the apparent Km value for ATP (plotted as initial velocity versus ATP concentration) are presented in Fig. 2.
TABLE 2.
Kinetic constants for Gmk and GmkS19Nalegend
| Constant | Temp (°C) | Gmk | GmkS19N |
|---|---|---|---|
| Km (GMP)b | 30 | 0.13 ± 0.24 | 0.11 ± 0.02 |
| Km (ATP)b | 30 | 0.18 ± 0.04 | 3.72 ± 0.61 |
| 37 | 0.15 ± 0.02 | 4.32 ± 0.66 | |
| Vmaxc | 30 | 0.08 ± 0.06 | 0.11 ± 0.005 |
| Vmax | 37 | 0.11 ± 0.007 | 0.16 ± 0.004 |
aGuanylate kinase was assayed spectrophotometrically by coupling the GMP to GDP conversion with the lactic dehydrogenase-catalyzed conversion of pyruvate and NADH to lactate and NAD (8, 17). Initial velocities were determined using 5.1 nmol of enzyme in each reaction. Kinetic constants were calculated from the data obtained using the enzyme kinetics template of the Prism program.
The Km values for GMP were determined with 5 mM ATP and GMP concentrations ranging from 0.0625 to 3 mM. The Km values for ATP were determined with saturating GMP (3 mM) and ATP concentrations ranging from 0.15 to 10 mM. Km values are reported as millimolar.
Vmax values were determined from the data shown in Fig. 2 and are reported as micromoles per minute.
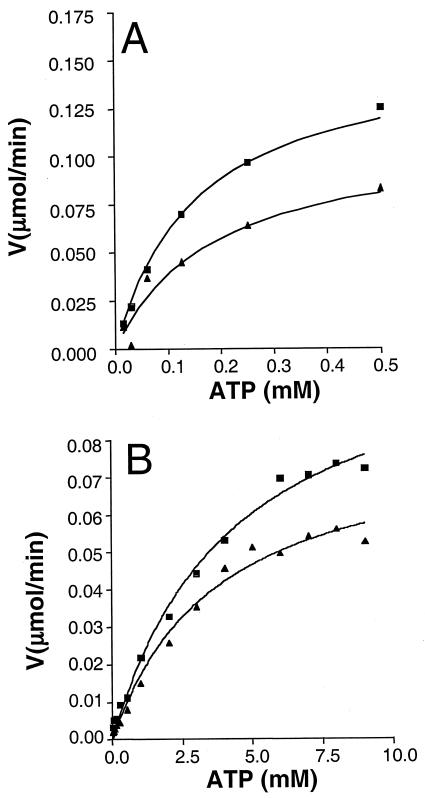
FIG. 2.
Kinetic analysis of Gmk and GmkS19N enzymes. Kinetic analyses were performed with wild-type (A) and mutant (B) Gmk enzymes. Initial velocity measurements were made using 5.1 nmol of enzyme in each reaction. The phosphoryl transfer of ATP to GMP was coupled to the pyruvate kinase-lactate dehydrogenase enzymes such that the absorbance change at 340 nm, detecting the decrease in NADH, revealed the guanylate kinase activity (8). In all cases the concentration of GMP was held constant at 3 mM and the concentration of ATP ranged from 0.015 mM to 10 mM. Kinetic analyses were carried out at both 37°C (▪) and 30°C (▴). The kinetic constants derived from these data are given in Table 2.
The temperature profile of Gmk activity does not explain the in vivo temperature-dependent phenotype.
A temperature profile of the mutant and wild-type enzyme activity was performed to identify a parameter that would correlate with the temperature-dependent phenotype in vivo. Initial velocity was measured from 25 to 52°C. Over this temperature range the change in initial velocity was linear, with slopes of 0.0016 and 0.0037 for the mutant and wild-type protein, respectively (data not shown). These slopes indicted that the disparity in initial velocity between the wild-type and mutant protein increased with temperature. In contrast, the phenotypic difference between the two strains decreased with temperature, as indicated by the prototrophic growth of the mutant strain at 37 but not at 30°C. This result indicated the temperature-dependent growth of the mutant was not a direct consequence of the altered kinetics of GmkS19N.
Accumulation of GMP contributes to two phenotypes associated with gmk101.
Two simple explanations for the temperature-dependent phenotype were considered and eliminated. First, similar ATP pools (∼0.5 mM) were detected with an Enliten ATP assay system (Promega, Madison, Wis.) at both temperatures, eliminating the possibility that increased ATP levels at 30°C could overcome the difference in Km value of GmkS19N. Although the presence of exogenous adenine has been reported to increase ATP pool size by ∼40%, other purine sources result in a similar elevation of pool size and yet fail to allow growth of the gmk101 mutant (1). Second, regulation of gmk expression was considered. Polyclonal antibodies were obtained from M. Cashel (National Institutes of Health) and used in immunoblot analyses to qualitatively assess accumulation of Gmk in mutant and wild-type strains at the two relevant temperatures. No significant difference in Gmk accumulation levels was noted.
Considering the data presented above, a general model to explain the purine requirement at 30°C and the specificity for adenine caused by the gmk101 mutation was suggested. Reduced activity of Gmk results in the accumulation of GMP in the cell (14). Since GMP is an allosteric inhibitor of adenylsuccinate synthetase (PurA) (7, 22), a partial starvation for AMP occurred in the gmk101 mutant strain. The starvation was exacerbated at 30°C, resulting in the adenine requirement; we suggest two possible means for this. First, the enzymatic activity of PurA may be decreased at 30°C. There is a precedent for a temperature dependence of biosynthetic enzymes (e.g., that of PurH and MetA) (4, 19). To the best of our knowledge, a temperature profile for PurA has not been reported. Second, a small increase in ppGpp levels would prevent AMP formation, since ppGpp is the most potent inhibitor of PurA, occupying a site distinct from that of GMP (7, 22). The metabolism of ppGpp is complex; thus, there are multiple factors that could generate the postulated increase. For instance, in one report a mutant defective in ndk (nucleoside diphosphate kinase) (Fig. 1) was found have a cold-sensitive growth defect (9, 18). At the low temperature the ppGpp levels in this mutant were elevated with respect to those seen at the permissive temperature. While the mechanism of this effect was not clear, the similarity to the result described herein suggests that the proposed scenario is feasible.
The specificity for adenine for satisfying the mutant growth requirement can be explained by a regulatory role for GMP. GMP allosterically inhibits guanine phosphoribosyl transferase (Gpt), the primary route for the conversion of hypoxanthine and guanine to their respective nucleotides (3, 10) (Fig. 1). While a single purine source can be converted to all others in a wild-type strain, a block in Gpt would slow, or prevent, the generation of adenine derivatives from the others (22), since ultimately this conversion must go through IMP via adenylsuccinate synthetase (PurA). Somewhat surprisingly, the preferred route for conversion of adenosine to adenine is via a route through IMP that also depends on Gpt and PurA. The direct phosphorolysis of adenosine to adenine is of minor quantitative importance (emphasized by the fact that an adenine-requiring mutant [e.g., purA or purB] grows poorly on adenosine) (22). Thus, specificity for adenine is consistent with the known routes of salvage in cases in which PurA is allosterically inhibited.
Interestingly, when the de novo purine pathway is blocked via mutation (purG) in a gmk101 mutant, the specificity for adenine is lost (data not shown). This result is consistent with the above model and suggests that the distribution of flux from IMP can be different when purines are utilized solely via the salvage pathways.
Conclusions.
The work described here identified a lesion in an essential gene and probed the metabolic consequences of partially disrupting the relevant metabolic activity. Two clear phenotypes were noted, and a model to explain them is presented. Many of the purine biosynthetic and salvage enzymes are allosterically regulated positively and negatively by various purine nucleotides and ppGpp. As shown here, significant metabolic disruptions can result from the sum of these multiple effects.
The nucleotide sequence for the gmk gene was deposited in GenBank under accession number AF140283.
Acknowledgments
We thank M. Cashel (NIH) and R. Gourse for helpful discussion and M. Cashel for providing antibodies to Gmk. We thank Katrina Forest for discussion of possible structural implications of the mutant gmk allele.
This work was supported by competitive grant GM47296 from the NIH. Funds were also provided from a 21st Century Scientist Scholars Award from the J. S. McDonnell Foundation.
REFERENCES
- 1.Bagnara, A. S., and L. R. Finch. 1974. The effects of bases and nucleosides on the intracellular contents of nucleotides and 5-phosphoribosyl 1-pyrophosphate in Escherichia coli. Eur. J. Biochem. 41:421-430. [DOI] [PubMed] [Google Scholar]
- 2.Brady, W. A., M. S. Kokoris, M. Fitzgibbon, and M. E. Black. 1996. Cloning, characterization, and modeling of mouse and human guanylate kinases. J. Biol. Chem. 271:16734-16740. [DOI] [PubMed] [Google Scholar]
- 3.Deo, S. S., W. C. Tseng, R. Saini, R. S. Coles, and R. S. Athwal. 1985. Purification and characterization of Escherichia coli xanthine-guanine phosphoribosyltransferase produced by plasmid pSV2gpt. Biochim. Biophys. Acta 839:233-239. [DOI] [PubMed] [Google Scholar]
- 4.Downs, D. M. 1987. Purine metabolism and cryptic prophages in Salmonella typhimurium. Ph.D. thesis. University of Utah, Salt Lake City.
- 5.Escobar-Henriques, M., and B. Daignan-Fornier. 2001. Transcriptional regulation of the yeast GMP synthesis pathway by its end products. J. Biol. Chem. 276:1523-1530. [DOI] [PubMed] [Google Scholar]
- 6.Fitzgibbon, J., N. Katsanis, D. Wells, J. Delhanty, W. Vallins, and D. M. Hunt. 1996. Human guanylate kinase (GUK1): cDNA sequence, expression and chromosomal localization. FEBS Lett. 385:185-188. [DOI] [PubMed] [Google Scholar]
- 7.Gallant, J., J. Irr, and M. Cashel. 1971. The mechanism of amino acid control of guanylate and adenylate biosynthesis. J. Biol. Chem. 246:5812-5816. [PubMed] [Google Scholar]
- 8.Gentry, D., C. Bengra, K. Ikehara, and M. Cashel. 1993. Guanylate kinase of Escherichia coli K-12. J. Biol. Chem. 268:14316-14321. [PubMed] [Google Scholar]
- 9.Ginther, C. L., and J. L. Ingraham. 1974. Cold-sensitive mutant of Salmonella typhimurium defective in nucleosidediphosphokinase. J. Bacteriol. 118:1020-1026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hochstadt, J. 1978. Hypoxanthine phosphoribosyltransferase and guanine phosphoribosyltransferase from enteric bacteria. Methods Enzymol. 51:549-558. [DOI] [PubMed] [Google Scholar]
- 11.Hong, J. S., and B. N. Ames. 1971. Localized mutagenesis of any specific small region of the bacterial chromosome. Proc. Natl. Acad. Sci. USA 68:3158-3162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Konrad, M. 1992. Cloning and expression of the essential gene for guanylate kinase from yeast. J. Biol. Chem. 267:25652-25655. [PubMed] [Google Scholar]
- 13.Kumar, V., O. Spangenberg, and M. Konrad. 2000. Cloning of the guanylate kinase homologues AGK-1 and AGK-2 from Arabidopsis thaliana and characterization of AGK-1. Eur. J. Biochem. 267:606-615. [DOI] [PubMed] [Google Scholar]
- 14.Lecoq, K., M. Konrad, and B. Daignan-Fornier. 2000. Yeast GMP kinase mutants constitutively express AMP biosynthesis genes by phenocopying a hypoxanthine-guanine phosphoribosyltransferase defect. Genetics 156:953-961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li, Y., Y. Zhang, and H. Yan. 1996. Kinetic and thermodynamic characterizations of yeast guanylate kinase. J. Biol. Chem. 271:28038-28044. [DOI] [PubMed] [Google Scholar]
- 16.Martinez, E., B. Bartolome, and F. de la Cruz. 1988. pACYC184-derived cloning vectors containing the multiple cloning site and lacAa reporter gene of pUC8/9 and pUC18/19 plasmids. Gene 68:159-162. [DOI] [PubMed] [Google Scholar]
- 17.Oeschger, M. P., and M. J. Bessman. 1966. Purification and properties of guanylate kinase from Escherichia coli. J. Biol. Chem. 241:5452-5460. [PubMed] [Google Scholar]
- 18.Rodriguez, S. B., and J. L. Ingraham. 1983. Location on the Salmonella typhimurium chromosome of the gene encoding nucleoside diphosphokinase (ndk). J. Bacteriol. 153:1101-1103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ron, E. Z., and M. Shani. 1971. Growth rate of Escherichia coli at elevated temperatures: reversible inhibition of homoserine trans-succinylase. J. Bacteriol. 107:397-400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Shen, H., B. Y. Yao, and D. M. Mueller. 1994. Primary structural constraints of P-loop of mitochondrial F1-ATPase from yeast. J. Biol. Chem. 269:9424-9428. [PubMed] [Google Scholar]
- 21.Stehle, T., and G. E. Schulz. 1992. Refined structure of the complex between guanylate kinase and its substrate GMP at 2.0 A resolution. J. Mol. Biol. 224:1127-1141. [DOI] [PubMed] [Google Scholar]
- 22.Zalkin, H., and P. Nygaard. 1996. Biosynthesis of purine nucleotides, p. 561-579. In F. C. Neidhardt, R. Curtiss III, J. L. Ingraham, E. C. C. Lin, K. B. Low, B. Magasanik, W. S. Resnikoff, M. Riley, M. Schaechter, and H. E. Umbarger (ed.), Escherichia coli and Salmonella: cellular and molecular biology, vol. 1. ASM Press, Washington, D.C.
- 23.Zhang, Y., Y. Li, Y. Wu, and H. Yan. 1997. Structural and functional roles of tyrosine 78 of yeast guanylate kinase. J. Biol. Chem. 272:19343-19350. [DOI] [PubMed] [Google Scholar]