Abstract
17β-Estradiol (E2) binds estrogen receptor α (ESR1) in MCF-7 cells and increases cell proliferation and survival through induction or repression of multiple genes. ESR1 interactions with DNA-bound specificity protein (SP) transcription factors is a nonclassical genomic estrogenic pathway and the role of SP transcription factors in mediating hormone-dependent activation or repression of genes in MCF-7 cells was investigated by microarrays and RNA interference. MCF-7 cells were transfected with a nonspecific oligonucleotide or a cocktail of small inhibitory RNAs (iSP), which knockdown SP1, SP3, and SP4 proteins, and treated with dimethylsulfoxide or 10 nM E2 for 6 h. E2 induced 62 and repressed 134 genes and the induction or repression was reversed in ∼62% of the genes in cells transfected with iSP (ESR1/SP dependent), whereas hormonal activation or repression of the remaining genes was unaffected by iSP (SP independent). Analysis of the ESR1/SP-dependent and SP-independent genes showed minimal overlap with respect to the GO terms (functional processes) in genes induced or repressed, suggesting that the different genomic pathways may contribute independently to the hormone-induced phenotype in MCF-7 cells.
Introduction
The estrogen receptor (ER) and other steroid hormone receptors are members of the nuclear receptor (NR) superfamily of transcription factors that are characterized by their common modular structure and similar ligand-dependent mechanisms of action. NRs express an N-terminal A/B domain that contains a ligand-independent activation function-1, a DNA-binding domain C, a flexible hinge region (D), and a C-terminal E/F domain that contains a ligand-dependent AF-2 and a region that binds ligands (Katzenellenbogen et al. 1996, Hall et al. 2001, Nilsson & Gustafsson 2002, Evans 2005, O’Malley 2005).
ESR1 and ESR2 are the two ER subtypes that exhibit overlapping and different patterns of expression in various tissues, and there is extensive evidence from in vitro cell culture, gene ablation, and in vivo studies that these receptors primarily have different functions (Matthews & Gustafsson 2003). ESR1 was the first ER subtype identified and its role is associated with many of the familiar estrogen-mediated functions including female reproductive tract development, bone growth, and overexpression in early stage mammary tumors in women and in ER-positive (ER+) breast cancer cells such as the widely used MCF-7 cell line (Levenson & Jordan 1997). MCF-7 and other ER+ breast cancer cell lines are highly responsive to the mitogenic activity of 17β-estradiol (E2) and have been extensively used to understand the complex molecular biology of estrogen action. The classical mechanism of E2-induced gene expression involves ligand-dependent formation of a nuclear ER homodimer bound to an estrogen-responsive element (ERE) that in turn recruits multiple elements of the transcriptional machinery to activate gene expression (Katzenellenbogen et al. 1996, Hall et al. 2001, Nilsson & Gustafsson 2002, Matthews & Gustafsson 2003, Evans 2005, O’Malley 2005). Subsequent studies have identified multiple pathways of hormone-dependent activation of ER and these include interactions of the receptor with ERE half-sites alone and in combination with other DNA-bound transcription factors such as GATA1, NFκB, and specificity protein-1 (SP1). In addition, ER activates genes through protein-protein interactions with other DNA-bound transcription factors such as the activator protein-1 (JUN) complex and SP proteins (Blobel & Orkin 1996, Paech et al. 1997, Webb et al. 1999, Inadera et al. 2000, Pelzer et al. 2001, Safe 2001, Pratt et al. 2003, Safe & Kim 2004, Chadwick et al. 2005, Ghisletti et al. 2005, Kalaitzidis & Gilmore 2005).
Several E2-responsive genes contain E2-responsive GC-rich promoters (Safe 2001, Safe & Kim 2004), and RNA interference studies using small inhibitory RNAs for SP1 (iSP1), SP3 (iSP3), SP4 (iSP4) or their combination (iSP) show that all three SP proteins play a role in hormonal activation of three E2-responsive genes, namely E2f1, Cad, and Rara (Khan et al. 2007). Transfection with iSP was the most effective inhibitor of hormone activation of these genes.
E2 and various selective ER modulators induce or repress a broad spectrum of genes in breast cancer cells (Soulez & Parker 2001, Inoue et al. 2002, Levenson et al. 2002, Lobenhofer et al. 2002, Coser et al. 2003, Cunliffe et al. 2003, Frasor et al. 2003, 2004, Scafoglio et al. 2006) and, in some microarray studies, E2 decreased expression of more genes than it induced. In this study, we investigated the role of SP proteins in hormonal modulation of gene expression in MCF-7 cells by treating cells for 6 h with dimethylsulfoxide (DMSO; solvent control) or 10 nM E2 and transfecting cells with either a nonspecific oligonucleotide (iNS) or iSP cocktail containing iSP1, iSP3, and iSP4. Using this approach, we showed that E2 induced 67 and repressed 134 genes and 62% of these genes were SP dependent. Genes regulated by ESR1/SP could be further subdivided into three subclasses based on the effects of iSP on basal activity of these genes in which basal expression was unaffected (B0), enhanced (B+), or decreased (B-). Interestingly, the B0 and B+ subcategories were predominant for genes induced and the B- subcategory predominated for genes repressed by E2-mediated activation of ESR1/SP. Our results show that the genomic ESR1/SP pathway is important for hormonal regulation of genes in MCF-7 cells.
Materials and methods
Cell culture
MCF-7 cells were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA) and were maintained in DMEM/nutrient mixture Ham’s F12 (DMEM/F12; Sigma-Aldrich) supplemented with 2·2 g/l sodium bicarbonate, 5% fetal bovine serum (FBS, JRH Biosciences, Lenexa, KS, USA), and 5 ml/liter antibiotic antimycotic solution (AAS, Sigma-Aldrich). Cells were cultured and grown in a 37 °C incubator with humidified 5% CO2:95% air.
Reverse transcription PCR for determination of mRNA levels
MCF-7 cells were cultured in serum-free DMEM/F12 for 1 day before treatment with 20 nM E2 or 20 nM E2+2 μM ICI 182780. DMSO as a solvent control for 24 h or with 20 nM E2 for 0, 2, 4, 6, 9, and 24 h. For the RNA interference studies, cells were cultured in phenol red-free DMEM/F12 supplemented with 2·5% charcoal-stripped FBS in six-well plates until 50-70% confluent. Cells were washed once with serum-free, antibiotic-free, phenol red-free DMEM/F12. The amount of siRNA to give a maximal decrease of each target protein was determined experimentally (50 nM final concentration in the well). Lipofectamine 2000 reagent (Invitrogen) was used to transfect MCF-7 cells with siRNA (iSP1, 3, 4, or iNS) according to the manufacturer’s protocol and as indicated above under RNA interference and microarray assay. The next day, cells were treated with 20 nM E2 or DMSO as a solvent control in serum-free, antibiotic-free, phenol red-free DMEM/F12. Cells were harvested 2, 4, 6, 9, or 24 h after treatment or as indicated in the different experiments. The siRNA oligonucleotides for nonspecific small inhibitory RNA (iNS), SP1, SP3, and SP4 were obtained from Dharmacon, Lafayette, CO, USA as described above. Total RNA from the studies was isolated using the RNeasy Protect Mini Kit (Qiagen) according to the manufacturer’s protocol. RNA concentration was measured by u.v. 260:280 nm absorption ratio; 1 μg RNA was reverse transcribed using Superscript II reverse transcriptase (Invitrogen) according to the manufacturer’s protocol. The genes of interest were amplified from the generated cDNA by PCR cycles using Taq polymerase (Invitrogen). The PCR conditions were as follows: initial denaturation at 94 °C (2 min) followed by 29 cycles (E2F1 and KLK6), 30 cycles (VEGFA), 28 cycles (HMGB1), or 27 cycles (GAPDH) of denaturation for 30 s at 94 °C; annealing for 30 s at 54 °C; extension at 72 °C for 1 min; and a final extension step at 72 °C for 10 min. The mRNA levels were normalized using Gapdh as an internal housekeeping gene. PCR products were electrophoresed on 1·5% agarose gels containing ethidium bromide and visualized under u.v. transillumination, then quantitated using Image-J software. Primers obtained from IDT (Coralville, IA, USA) and used for amplification were E2F1 (sense, 5′-CGC ATC TAT GAC ATC ACC AAC G-3′; antisense, 5′-GAA AGT TCT CCG AAG AGT CCA CG-3′); VEGFA (sense, 5′-CCA TGA ACT TTC TGC TGT CTT-3′; antisense, 5′-ATC GCA TCA GGG GCA CAC AG-3′); HMGB1 (sense, 5′-AAC ATG GGC AAA GGA GAT CC-3′); antisense, 5′-TAC CAG GCA AGG TTA GTG GC-3′); KLK6 (sense, 5′-TAC CAA GCT GCC CTC TAC AC-3′; antisense, 5′-ACA AGG CCT CGG AGG TGG TC-3′); GAPDH (sense, 5′-AAT CCC ATC ACC ATC TTC CA-3′; antisense, 5′-GTC ATC ATA TTT GGC AGG TT-3′).
RNA interference and microarray assay
MCF-7 cells (5×104) were cultured in phenol red-free DMEM/F12 supplemented with 2·5% charcoal-stripped FBS without AAS in 12-well plates overnight. Cells were transfected with nonspecific siRNA (iNS) or the combination of iSP1/iSP3/iSP4 (iSP) by Lipofectamine 2000 reagent (Invitrogen), according to the manufacturer’s protocol. After 6 h, the transfection medium was changed with fresh DMEM/F12 and 2·5% serum without phenol red. Forty-eight hours after transfection, cells were treated with (DMSO, solvent) or 10 nM E2 (98%, Sigma-Aldrich) for 6 h. Three replicate determinations were obtained for each treatment group. Total RNA was isolated using the RNeasy Protection Mini Kit (Qiagen) according to the manufacturer’s protocol.
The siRNA duplexes used in this study are indicated as follows. Silencer Negative Control #1 siRNA purchased from Ambion (Austin, TX, USA) was used as iNS. The siRNA oligonucleotides for SP1, SP3, and SP4 were obtained from Dharmacon as follows: SP1, 5′-AUC ACU CCA UGG AUG AAA UGA dTdT-3′; SP3, 5′-GCG GCA GGU GGA GCC UUC ACU dTdT-3′; and SP4, 5′-GCA GUG ACA CAU UAG UGA GCdT dT-3′.
Microarray hybridization and data analysis
Microarray studies were carried out with the three samples obtained for each treatment group using the CodeLink Whole Genome Bioarrays (#300026) with over 50 000 probe targets per slide and, after treatment with 10 nM E2 for 6 h, RNA was isolated for reverse transcription PCR. The microarray data for each sample were imported into GeneSpring BX7 software (Silicon Genetics, Redwood City, CA, USA) for data analysis. The data were normalized in two steps. First, for each array, the expression value of each gene was divided by the median of the expression values in that array. Second, for each gene, the expression value in each array was divided by the median expression value of that gene across all of the arrays. Genes that were flagged ‘L’ by the CodeLink preprocessing software, i.e. with low signal to background noise ratio, were excluded from the subsequent statistical analysis. One-way parametric ANOVA test was performed to detect significant changes of gene expression. Benjamini-Hochberg false discovery rate as multiple testing correction and Tukey post hoc test were applied. The false discovery rate cut-off was set to 0·05 due to the large number of genes on the chip and the small number of microarrays used in the experiment.
Western blot analysis
MCF-7 cells were seeded into six-well plates in DMEM/F12 supplemented with 2·5% charcoal-stripped FBS. The next day, cells were transfected with siRNA as described above and high-salt extracts were obtained by harvesting cells in a high-salt lysis buffer (50 mM HEPES (pH 7·5), 500 mM NaCl, 10% (vol/vol) glycerol, 1% Triton X-100, 1·5 mM MgCl2, 1 mM EGTA, protease inhibitor cocktail (Sigma-Aldrich)) on ice for 45-60 min with frequent vortex and centrifugation at 20 000 g for 10 min at 4 °C. Protein concentrations were determined using a BioRad protein assay reagent. Protein (60 μg) was diluted with Laemmli’s loading buffer, boiled, and loaded onto 7·5% SDS-PAGE. Samples were resolved using electrophoresis at 150 V for 3-4 h and transferred (transfer buffer, 48 mM Tris-HCl, 29 mM glycine, and 0·025% sodium dodecyl sulfate) to a polyvinylidene difluoride membrane (BioRad) by electrophoresis at 0·2 Å for ∼12-16 h. Membranes were blocked in 5% TBS-Tween 20-Blotto (10 mmol/l Tris-HCl, 150 mmol/l NaCl (pH 8·0), 0·05% Triton X-100, 5% nonfat dry milk) with gentle shaking for 30 min and incubated in fresh 5% TBS-Tween 20-Blotto with 1:1000 (for SP1 and SP3,), 1:500 (for SP4), and 1:5000 (for β-actin) primary antibody overnight with gentle shaking at 4 °C. The primary antibodies for SP1 (PEP2), SP3 (D-20), and SP4 (V-20) were purchased from Santa Cruz Biotechnology Inc. (Santa Cruz, CA, USA) and antibody for β-actin was purchased from Sigma-Aldrich. Membranes were probed with a HRP-conjugated secondary antibody (1:5000) for 3-6 h at 4 °C. Blots were visualized using the chemiluminescence substrate (Perkin-Elmer Life Sciences Warwick, RI, USA) and exposure on Image Tek-H X-ray film (American X-ray Supply). Band quantitation was performed by ImageJ (National Institutes of Health).
Results
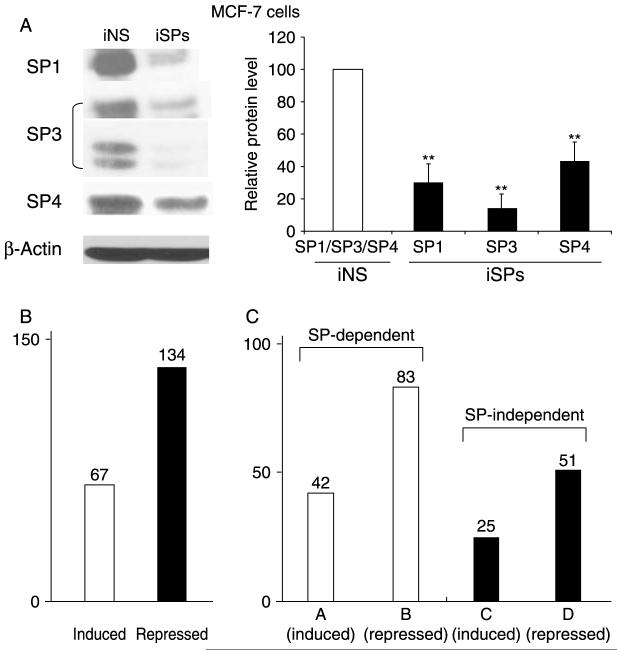
Previous microarray studies have investigated hormonal modulation of genes in breast cancer cells under various conditions and there is evidence that genes, with different and overlapping functions, are induced and repressed (Soulez & Parker 2001, Inoue et al. 2002, Levenson et al. 2002, Lobenhofer et al. 2002, Coser et al. 2003, Cunliffe et al. 2003, Frasor et al. 2003, 2004, Scafoglio et al. 2006, Jonsson et al. 2007). Several reports show that E2 induces and represses genes that are activated through a nonclassical ESR1/SP mechanism in which ESR1 binds SP proteins but not promoter DNA (Safe 2001, Safe & Kim 2004); however, the contributions of this pathway to hormonal activation of genes is not well defined. In this study, MCF-7 cells were treated with DMSO (solvent) or 10 nM E2 for 6 h and transfected with a nonspecific oligonucleotide (iNS) or a combined cocktail (iSP) of small inhibitory RNAs for SP1 (iSP1), SP3 (iSP3), and SP4 (iSP4), which has previously been used to simultaneously knockdown all three SP proteins (Khan et al. 2007). Western blot analysis of whole-cell lysates from cells transfected with INS or iSP confirms efficient knockdown of all three SP proteins (Fig. 1A). Results in Fig. 1B show that in cells transfected with iNS and treated with DMSO or E2, there were 67 and 134 genes significantly (P<0·01) induced and repressed respectively, and this was consistent with other studies in MCF-7 cells showing that E2 primarily decreases gene expression (Frasor et al. 2003, 2004). It is important to point out that our selection of significantly expressed genes is based on a stringent condition (P<0·01) that reflects the large number of probes on the slides and the small number of samples. Selection of these stringent conditions is consistent with a report by Hu and coworkers (Hu et al. 2005), which uses the false discovery rate to estimate the number of required samples per condition for achieving a desired level of significance. This has to be taken into account when comparing the results of our statistical analysis with the already published similar studies. For example, Frasor and coworkers (Frasor et al. 2003) identified over 400 genes with ‘robust level of regulation,’ but the microarrays used in their experiment contained ∼12 000 genes and the decision to place a gene in the reported list was based on a different scoring procedure.
Figure 1.
Role of SP transcription factors in E2-dependent gene expression. (A) Effect of iSP on SP1, SP3, and SP4. MCF-7 cells were transfected with the iSP cocktail or a nonspecific oligonucleotide (iNS), and SP proteins were analyzed in triplicate by western blotting as described in the Materials and methods. Significantly (P<0·05) decreased expression is indicated (**). (B) Number of genes up- or downregulated by 10 nM E2. (C) SP-dependent and -independent gene categories. Categories A and B represent SP-dependent (ESR1/SP) genes induced or repressed by E2. Categories C and D represent SP-independent (ESR1) genes induced or repressed by E2. Category F genes are induced or repressed by E2 only after SP knockdown.
After transfection of cells with iSP, hormone-regulated genes in MCF-7 cells can be subdivided into four major categories (Fig. 1C). Categories A and B represent SP-dependent genes induced or repressed by E2 (Tables 1 and 2) and categories C and D represent SP-independent genes induced or repressed by E2 respectively (Tables 3 and 4). A fifth category, F, represents a substantial number of genes (85) that are not affected by treatment with E2; however, after SP protein knockdown (Fig. 1A), category F genes are either induced or repressed by E2 (Table 5). SP proteins are overexpressed in breast tumor compared with nontumor cells (Mertens-Talcott et al. 2007), and it is possible that as SP proteins increase during mammary carcinogenesis, category F genes become hormone insensitive. The decision to assign a gene to one of the five categories was based on two cut-off points; genes with fold change of 0·8 or less were considered repressed while the genes with fold change of 1·3 or greater were considered induced. The selection of these thresholds was based on the observation that varying them by as much as 0·02 produced small changes in the categories; only one to five genes would change their category membership.
Table 1.
Category A genes that are induced by E2 in MCF-7 cells and reversed by SP protein knockdown (SP dependent)
| Gene name |
Fold inductiona (iNSE/iNSD) |
GO biol process |
|
|---|---|---|---|
| Gene accession numbers | |||
| Category A (B0) | |||
| BM459591 | 1·366667 | No info about GO biol process | |
| NM_178501 | 2·447368 | No info about GO biol process | |
| NM_005225 | E2f1 | 2·338661 | Apoptosis; + multiple GO terms |
| BF056555 | 2·294118 | No info about GO biol process | |
| AW002058 | 2·189189 | No info about GO biol process | |
| AI654853 | 2·179487 | No info about GO biol process | |
| H23439 | 2·095238 | No info about GO biol process | |
| H93315 | 2·022222 | No info about GO biol process | |
| AW977008 | 1·97561 | No info about GO biol process | |
| NM_003125 | Sprr1b | 1·826185 | Epidermis development |
| NM_130830 | Lrrc15 | 1·626459 | No info about GO biol process |
| BQ931737 | 1·549451 | No info about GO biol process | |
| NM_012429 | Sec14l2 | 1·477273 | Transport; + multiple GO terms |
| NM_005551 | Klk2 | 1·423529 | Proteolysis and peptidolysis |
| NM_004973 | Jarid2 | 1·32288 | Central nervous system development |
| BG029979 | 1·465116 | No info about GO biol process | |
| Category A (B+) | |||
| BF512856 | 1·84127 | No info about GO biol process | |
| NM_001051 | Sstr3 | 1·65625 | Cell-cell signaling; + multiple GO terms |
| NM_003647 | Dgke | 1·326923 | Intracellular signaling cascade; + multiple GO terms |
| BM714396 | 1·6 | No info about GO biol process | |
| CF147798 | P2ry10 | 4·5 | No info about GO biol process |
| AW028308 | 4·333333 | No info about GO biol process | |
| U91328 | 4·384615 | No info about GO biol process | |
| BC038785 | 2·914286 | No info about GO biol process | |
| AI444574 | 2·83871 | No info about GO biol process | |
| NM_000171 | Glra1 | 2·433333 | Ion transport; + multiple GO terms |
| BG541228 | 2·382979 | No info about GO biol process | |
| BC012108 | Ca14 | 1·806452 | No info about GO biol process |
| AI202342 | 1·745763 | No info about GO biol process | |
| AF130075 | 1·597015 | No info about GO biol process | |
| BC033385 | 1·604938 | No info about GO biol process | |
| NM_144715 | Efhb | 1·475728 | No info about GO biol process |
| NM_016594 | Fkbp11 | 1·347765 | Protein folding |
| NM_033315 | Rasl10b | 1·309091 | Small GTPase-mediated signal transduction |
| AW274516 | Itpkb | 3·607143 | GO:0007165 |
| BG498282 | 3·071429 | No info about GO biol process | |
| BU616743 | 2·761905 | No info about GO biol process | |
| Category A (B-) | |||
| BG211832 | 1·842105 | No info about GO biol process | |
| BE927766 | 1·755556 | No info about GO biol process | |
| NM_001165 | Birc3 | 1·620818 | Anti-apoptosis; + multiple GO terms |
| NM_005416 | Sprr3 | 1·391586 | GO:8544 (epidermis development);+ multiple GO terms |
| BX095305 | 1·319797 | No info about GO biol process | |
MCF-7 cells transfected with iNS (nonspecific oligonucleotide) and treated with either 10 nM E2 (iNSE) or DMSO (iNSD).
Table 2.
Category B genes that are repressed by E2 in MCF-7 cells and reversed by SP protein knockdown (SP dependent)
| Gene name |
Fold inductiona (iNSE/iNSD) |
GO biol process |
|
|---|---|---|---|
| Gene accession numbers | |||
| Category B (B0) | |||
| BI520070 | 0·75475 | No info about GO biol process | |
| NM_005919 | Mef2b | 0·755932 | Muscle development; + multiple GO terms |
| BG194714 | 0·751412 | No info about GO biol process | |
| AI748967 | 0·694444 | No info about GO biol process | |
| BI791886 | 0·685832 | No info about GO biol process | |
| BI559738 | 0·664634 | No info about GO biol process | |
| NM_020909 | Epb41l5 | 0·661818 | No info about GO biol process |
| NM_152429 | Fgfbp3 | 0·657258 | No info about GO biol process |
| AI459284 | 0·609756 | No info about GO biol process | |
| NM_018374 | Tmem106b | 0·582677 | No info about GO biol process |
| BX093288 | 0·555556 | No info about GO biol process | |
| AK091299 | 0·556604 | No info about GO biol process | |
| XM_061614 | 0·527273 | No info about GO biol process | |
| AI348944 | 0·451923 | No info about GO biol process | |
| NM_173580 | 0·736842 | No info about GO biol process | |
| AI632459 | 0·573964 | No info about GO biol process | |
| AK023938 | Lrrfip1 | 0·724638 | GO:0016481;+ multiple GO terms |
| U66581 | Gpr22 | 0·75 | G-protein coupled receptor protein signaling pathway |
| NM_020340 | Kiaa1244 | 0·795064 | GO:32012 (regulation of ARF protein signal transduction) |
| BX103217 | 0·731898 | No info about GO biol process | |
| W69651 | 0·56 | No info about GO biol process | |
| NM_173580 | 0·736842 | No info about GO biol process | |
| Category B (B+) | |||
| NM_004799 | Zfyve9 | 0·765625 | Endocytosis; + multiple GO terms |
| Category B(B-) | |||
| AA323614 | 0·799242 | No info about GO biol process | |
| NM_024056 | Tmem106c | 0·796373 | No info about GO biol process |
| NM_018186 | C1orf112 | 0·790563 | No info about GO biol process |
| NM_014793 | Lcmt2 | 0·786538 | No info about GO biol process |
| NM_014920 | Ick | 0·768975 | Protein amino acid phosphorylation |
| NM_001310 | Crebl2 | 0·767041 | Signal transduction; transcription |
| NM_022746 | Mosc1 | 0·765499 | No info about GO biol process |
| BM723321 | 0·762322 | No info about GO biol process | |
| NM_152524 | Sgol2 | 0·758025 | GO:7049 (cell cycle);+ multiple GO terms |
| NM_025004 | Ccdc15 | 0·754331 | No info about GO biol process |
| NM_002774 | Klk6 | 0·752992 | Regulation of cell differentiation; + multiple GO terms |
| NM_016076 | Fam152a | 0·741884 | No info about GO biol process |
| BG740877 | 0·741463 | No info about GO biol process | |
| NM_024430 | Pstpip2 | 0·736111 | No info about GO biol process |
| AA046178 | 0·731949 | No info about GO biol process | |
| AA809349 | 0·732218 | No info about GO biol process | |
| AK026856 | 0·729258 | No info about GO biol process | |
| CB962212 | 0·728643 | No info about GO biol process | |
| NM_003400 | Xpo1 | 0·720289 | GO:6406 (mRNA export from nucleus);+ multiple GO terms |
| NM_014240 | Limd1 | 0·719298 | Development; + multiple GO terms |
| NM_016551 | Tm7sf3 | 0·718053 | No info about GO biol process |
| NM_006479 | Rad51ap1 | 0·71855 | GO:6281 (DNA repair);+ multiple GO terms |
| NM_022374 | Atl2 | 0·712202 | Immune response |
| NM_012287 | Acap2 | 0·703775 | GO:43087 (regulation of GTPase activity) |
| NM_005226 | S1pr3 | 0·703278 | Morphogenesis; + multiple GO terms |
| AI979035 | 0·698925 | No info about GO biol process | |
| NM_016203 | Prkag2 | 0·696429 | Cholesterol biosynthesis; + multiple GO terms |
| AA261924 | 0·695868 | No info about GO biol process | |
| NM_001278 | Chuk | 0·687229 | Immune response; + multiple GO terms |
| AA653936 | 0·686189 | No info about GO biol process | |
| NM_020132 | Agpat3 | 0·682759 | Metabolism; + multiple GO terms |
| NM_007027 | Topbp1 | 0·677027 | DNA metabolism; + multiple GO terms |
| BQ013447 | 0·675 | No info about GO biol process | |
| BI754506 | 0·670455 | No info about GO biol process | |
| NM_032876 | 0·668553 | No info about GO biol process | |
| NM_153826 | Capg | 0·6639 | Complement activation, classical pathway |
| NM_016824 | Add3 | 0·664298 | Apoptosis; + multiple GO terms |
| NM_016359 | Nusap1 | 0·663537 | G0:0000281;+ multiple GO terms |
| BG285682 | 0·65847 | No info about GO biol process | |
| NM_172240 | Wdr51b | 0·652372 | No info about GO biol process |
| N40495 | 0·647321 | No info about GO biol process | |
| NM_003415 | Znf268 | 0·646 | Regulation of transcription, DNA dependent |
| NM_139279 |
Mcfd2; Lman1; Mcfd2 |
0·637809 | GO:6888 (ER to Golgi vesicle-mediated transport);+ multiple GO terms |
| NM_032525 | Tubb6 | 0·633021 | Microtubule-based movement |
| AK091460 | Znf813 | 0·626354 | No info about GO biol process |
| NM_017709 | Fam46c | 0·624658 | No info about GO biol process |
| NM_138635 | H2afv | 0·621495 | GO:6334 (nucleosome assembly);+ multiple GO terms |
| AK056236 | 0·621074 | No info about GO biol process | |
| NM_173556 | Ccdc83 | 0·605682 | No info about GO biol process |
| NM_002128 | Hmgb1 | 0·59 | DNA unwinding; + multiple GO terms |
| NM_020927 | Vat1l | 0·582734 | No info about GO biol process |
| NM_016516 | Vps54 | 0·564955 | GO:15031 (protein transport);+ multiple GO terms |
| NM_018836 | Ajap1 | 0·533835 | No info about GO biol process |
| NM_019087 | Arl15 | 0·515924 | No info about GO biol process |
| AI218135 | 0·647059 | No info about GO biol process | |
| AI742318 | 0·314286 | No info about GO biol process | |
| BU069371 | 0·224138 | No info about GO biol process | |
| NM_001701 | Baat | 0·772727 | Bile acid metabolism; + multiple GO terms |
| NM_007023 | Rapgef4 | 0·675676 | Exocytosis; + multiple GO terms |
| AI480013 | 0·52381 | No info about GO biol process | |
MCF-7 cells transfected with iNS (nonspecific oligonucleotide) and treated with either 10 nM E2 (iNSE) or DMSO (iNSD).
Table 3.
Category C genes that are induced by E2 in MCF-7 cells and are unaffected by SP protein knockdown (SP independent)
| Gene name |
Fold inductiona (iNSE/iNSD) |
GO biol process |
|
|---|---|---|---|
| Gene accession numbers | |||
| Category C (B0) | |||
| AK098294 | 1·7 | No info about GO biol process | |
| BX107797 | 1·395349 | No info about GO biol process | |
| BC041884 | 3·5 | No info about GO biol process | |
| AW513673 | 1·895833 | No info about GO biol process | |
| NM_001321 | Csrp2 | 1·529201 | Cell growth; + multiple GO terms |
| Category C (B+) | |||
| AA234276 | 3·173387 | No info about GO biol process | |
| CA310410 | 2·073394 | No info about GO biol process | |
| NM_022467 | Chst8 | 1·350649 | Hormone biosynthesis; + multiple GO terms |
| NM_173505 | Ankrd29 | 2·151515 | No info about GO biol process |
| NM_000817 | Gad1 | 1·39375 | Amino acid metabolism; + multiple GO terms |
| NM_000360 | Th | 1·447205 | Morphogenesis; + multiple GO terms |
| NM_024522 | Nkain1 | 1·460317 | No info about GO biol process |
| NM_030781 | 4·384615 | No info about GO biol process | |
| Category C (B-) | |||
| BU677122 | 2·575758 | No info about GO biol process | |
| BF594491 | 1·455682 | No info about GO biol process | |
| BM995921 | 1·756098 | No info about GO biol process | |
| AA470089 | 1·41206 | No info about GO biol process | |
| BG188765 | 2·434783 | No info about GO biol process | |
| AK126800 | 2·34375 | No info about GO biol process | |
| BX091274 | 2·189189 | No info about GO biol process | |
| BQ182928 | 2·095238 | No info about GO biol process | |
| AW291057 | 1·967742 | No info about GO biol process | |
| AI356266 | 1·630137 | No info about GO biol process | |
| NM_001657 | Areg | 1·587224 | Cell proliferation; cell-cell signaling |
| BG025371 | 1·465385 | No info about GO biol process | |
MCF-7 cells transfected with iNS (nonspecific oligonucleotide) and treated with either 10 nM E2 (iNSE) or DMSO (iNSD).
Table 4.
Category D genes that are repressed by E2 in MCF-7 cells and are unaffected by SP protein knockdown (SP independent)
| Gene name |
Fold inductiona (iNSE/iNSD) |
GO biol process |
|
|---|---|---|---|
| Gene accession numbers | |||
| Category D (B0) | |||
| NM_004982 | Kcnj8 | 0·54631 | Ion transport; + multiple GO terms |
| AK022053 | 0·430769 | No info about GO biol process | |
| CB321977 | 0·56338 | No info about GO biol process | |
| NM_012106 | Arl2bp | 0·600355 | Signal transduction |
| NM_001056 | Sult1c2 | 0·492958 | Amine metabolism |
| AI492256 | 0·560606 | No info about GO biol process | |
| NM_015033 | Fnbp1 | 0·718391 | GO:6412 (protein biosynthesis) |
| BX114748 | 0·554545 | No info about GO biol process | |
| BQ024005 | 0·630137 | No info about GO biol process | |
| BM990273 | 0·730769 | No info about GO biol process | |
| AB067496 | Plekhg4b | 0·653333 | GO:35023 (regulation of Rho protein signal transduction) |
| BC042089 | 0·772727 | No info about GO biol process | |
| NM_004260 | Recql4 | 0·730203 | DNA repair; + multiple GO terms |
| AW298780 | 0·5 | No info about GO biol process | |
| Category D (B+) | |||
| NM_000304 | Pmp22 | 0·737374 | Mechanosensory behavior; + multiple GO terms |
| NM_004089 | Tsc22d3 | 0·794749 | Regulation of transcription, DNA dependent |
| AV652591 | 0·788945 | No info about GO biol process | |
| AK126952 | 0·674419 | No info about GO biol process | |
| Category D (B-) | |||
| NM_018393 | Tcp11l1 | 0·756677 | No info about GO biol process |
| NM_003025 | Sh3gl1 | 0·74812 | Signal transduction; + multiple GO terms |
| NM_018492 | Pbk | 0·771031 | Protein amino acid phosphorylation |
| NM_133638 | Adamts19 | 0·759615 | Proteolysis and peptidolysis |
| BI602086 | Abat | 0·756691 | Aminobutyrate metabolism; + multiple GO terms |
| NM_017634 | Kctd9 | 0·743668 | GO:6813 (potassium ion transport) |
| NM_001668 | Arnt | 0·741218 | Signal transduction; + multiple GO terms |
| W31852 | 0·73236 | No info about GO biol process | |
| NM_001798 | Cdk2 | 0·726695 | Cytokinesis; + multiple GO terms |
| NM_006751 | Ssfa2 | 0·720841 | No info about GO biol process |
| NM_015368 | Panx1 | 0·707317 | GO:0006812;+ multiple GO terms |
| NM_014900 | Cobll1 | 0·701372 | No info about GO biol process |
| NM_004523 | Kif11 | 0·698872 | Mitotic spindle assembly |
| NM_001067 | Top2a | 0·697188 | GO:6266 (DNA ligation);+ multiple GO terms |
| NM_021211 | 0·693424 | No info about GO biol process | |
| BQ350534 | 0·671694 | No info about GO biol process | |
| NM_012381 | Orc3l | 0·667582 | DNA replication |
| NM_005573 | Lmnb1 | 0·66129 | No info about GO biol process |
| NM_022748 | Tns3 | 0·655586 | Intracellular signaling cascade; + multiple GO terms |
| NM_004298 | Nup155 | 0·649412 | Nucleocytoplasmic transport |
| NM_016121 | Kctd3 | 0·649156 | GO:6813 (potassium ion transport) |
| NM_022918 | Tmem135 | 0·637079 | No info about GO biol process |
| NM_004856 | Kif23 | 0·637931 | GO:7049 (cell cycle);+ multiple GO terms |
| NM_144717 | Il20rb | 0·628205 | No info about GO biol process |
| NM_005139 | Anxa3 | 0·624911 | GO:7165 (signal transduction) |
| NM_006219 | Pik3cb | 0·623397 | Activation of MAPK; + multiple GO terms |
| W38778 | 0·591923 | No info about GO biol process | |
| NM_007145 | Znf146 | 0·587372 | Regulation of transcription, DNA dependent |
| NM_004087 | Dlg1 | 0·58216 | GO:31575 (G1/S transition checkpoint);+ multiple GO terms |
| NM_005180 | Commd3 | 0·576584 | Cell growth and/or maintenance; + multiple GO terms |
| BM145609 | 0·56044 | No info about GO biol process | |
| AW573130 | 0·506667 | No info about GO biol process | |
| AW237319 | Kiaa1614 | 0·477273 | No info about GO biol process |
MCF-7 cells transfected with iNS (nonspecific oligonucleotide) and treated with either 10 nM E2 (iNSE) or DMSO (iNSD).
Table 5.
Category F genes that are only induced or repressed by E2 after SP protein knockdown
| Gene name |
Fold inductiona (iNSE/iNSD) |
GO biol process |
|
|---|---|---|---|
| Gene accession numbers | |||
| Category F (B0) | |||
| AI693142 | 0·880041365 | No info about GO biol process | |
| NM_013314 | Blnk | 0·937625755 | B-cell differentiation; + multiple GO terms |
| BM724201 | 0·90070922 | No info about GO biol process | |
| NM_006849 | Pdia2 | 1·201550388 | Electron transport; + multiple GO terms |
| NM_001343 | Dab2 | 0·82967033 | Cell proliferation |
| AL080280 | 1·157894737 | No info about GO biol process | |
| AW294582 | 1·278106509 | No info about GO biol process | |
| NM_004998 | Myo1e | 1·052631579 | Actin filament-based movement |
| NM_014930 | Znf510 | 0·919254658 | No info about GO biol process |
| NM_153020 | Rbm24 | 0·883024251 | No info about GO biol process |
| BE350156 | 0·843283582 | No info about GO biol process | |
| NM_006731 | Fktn | 1·070850202 | Muscle development; + multiple GO terms |
| NM_004948 | Dsc3 | 0·905405405 | Cell adhesion; + multiple GO terms |
| NM_144726 | Rnf145 | 0·898989899 | GO:6512 (ubiquitin cycle) |
| NM_145646 | 0·954356846 | No info about GO biol process | |
| NM_001408 | Celsr2 | 0·935185185 | Development; + multiple GO terms |
| NM_016424 | LUC7A | 0·936090226 | RNA splicing |
| NM_033083 | Eaf1 | 0·844650206 | GO:6350 (transcription);+ multiple GO terms |
| AW869316 | 0·956521739 | No info about GO biol process | |
| NM_016014 | Fam108b1 | 1·03 | No info about GO biol process |
| NM_014650 | Znf432 | 0·861751152 | GO:6350 (transcription);+ multiple GO terms |
| NM_173683 | Xkr6 | 0·851851852 | No info about GO biol process |
| AW007727 | 0·80952381 | No info about GO biol process | |
| N52436 | 0·827922078 | No info about GO biol process | |
| CD104312 | 0·830508475 | No info about GO biol process | |
| NM_012193 | Fzd4 | 0·830065359 | Development; + multiple GO terms |
| NM_003839 | Tnfrsf11a | 0·993197279 | Cell-cell signaling; + multiple GO terms |
| BF963928 | 0·903100775 | No info about GO biol process | |
| NM_020246 | Trip6 | 1·103030303 | Amino acid transport |
| NM_006651 | Cplx1 | 0·873563218 | Exocytosis; + multiple GO terms |
| NM_002750 | Mapk8 | 0·91503268 | JNK cascade; + multiple GO terms |
| Category F (B+) | |||
| NM_013301 | Ccdc106 | 0·923547401 | No info about GO biol process |
| NM_012324 | Mapk8ip2 | 1·079207921 | Positive regulation of anti-apoptosis; + multiple GO terms |
| NM_032431 | SVN1 | 0·857777778 | GO:0030433 |
| NM_020820 | Prex1 | 0·897926635 | Actin filament polymerization; + multiple GO terms |
| NM_052880 | Pik3ip1 | 0·934362934 | No info about GO biol process |
| CA337205 | 0·947174447 | No info about GO biol process | |
| BX101252 | 1·01509434 | No info about GO biol process | |
| BX089473 | 1·120930233 | No info about GO biol process | |
| NM_031464 | Rps6kl1 | 1·095 | GO:6468 (protein amino acid phosphorylation) |
| AB065764 | 0·952380952 | No info about GO biol process | |
| BX470892 | 0·873015873 | No info about GO biol process | |
| NM_181535 | 1·056603774 | No info about GO biol process | |
| AL049274 | 0·945273632 | No info about GO biol process | |
| NM_022818 | Map1lc3b | 0·948138298 | GO:6914 (autophagy);+ multiple GO terms |
| AK097239 | 0·892592593 | No info about GO biol process | |
| BQ276710 | 0·949275362 | No info about GO biol process | |
| BM712945 | 1·021276596 | No info about GO biol process | |
| AK092442 | PP14571 | 0·854166667 | No info about GO biol process |
| NM_014751 | Mtss1 | 0·9360519 | Cell adhesion; + multiple GO terms |
| AK097032 | MGC17403 | 0·950657895 | Transcription |
| NM_032412 | C5orf32 | 0·988957055 | Visual perception; + multiple GO terms |
| BC042976 | Jmjd2c | 1·114035088 | GO:0008150 |
| AA324557 | 1·056818182 | No info about GO biol process | |
| NM_014428 | Tjp3 | 0·847682119 | No info about GO biol process |
| BG009563 | 0·873846154 | No info about GO biol process | |
| NM_000076 | Cdkn1c | 1·008213552 | Cell cycle arrest; + multiple GO terms |
| AW299364 | 1 | No info about GO biol process | |
| BG032839 | PCOTH | 1·118715084 | No info about GO biol process |
| BM743413 | 1·162576687 | No info about GO biol process | |
| NM_022912 | Reep1 | 0·972665148 | GO:0051205 |
| NM_014509 | Serhl2 | 0·991071429 | No info about GO biol process |
| NM_033467 | Mmel1 | 1·016348774 | Proteolysis and peptidolysis |
| BM991890 | 0·819787986 | No info about GO biol process | |
| AK130231 | 1·009433962 | No info about GO biol process | |
| NM_016162 | Ing4 | 0·878472222 | Regulation of transcription, DNA dependent |
| BF062651 | 0·875 | No info about GO biol process | |
| Category F (B-) | |||
| NM_006461 | Spag5 | 0·835327234 | GO:7049 (cell cycle);+ multiple GO terms |
| NM_199050 | C2cd2 | 0·826548673 | No info about GO biol process |
| NM_015341 | Ncaph | 0·814073227 | Cell cycle; + multiple GO terms |
| NM_181353 | Id1 | 0·807936508 | Development; + multiple GO terms |
| NM_005547 | Ivl | 0·885714286 | Keratinocyte differentiation |
| NM_153750 | 0·913043478 | No info about GO biol process | |
| BX451454 | 1·083333333 | No info about GO biol process | |
| CB052574 | 1·153153153 | No info about GO biol process | |
| NM_138573 | Nrg4 | 1·015384615 | No info about GO biol process |
| AI349515 | 0·824561404 | No info about GO biol process | |
| NM_016823 | Crk | 0·869067103 | Cell motility; + multiple GO terms |
| NM_002766 | Prpsap1 | 0·814229249 | Nucleoside metabolism; + multiple GO terms |
| BG164253 | 1·119565217 | No info about GO biol process | |
| NM_004494 | Hdgf | 0·875396825 | Cell proliferation; + multiple GO terms |
| NM_002755 | Map2k1 | 0·806122449 | GO:6928 (cell motility);+ multiple GO terms |
| NM_014553 | Tfcp2l1 | 0·877777778 | Steroid biosynthesis; + multiple GO terms |
| NM_001262 | Cdkn2c | 0·826974268 | Cell cycle; + multiple GO terms |
| NM_024590 | Arsj | 0·87037037 | GO:8152 (metabolism) |
MCF-7 cells transfected with iNS (nonspecific oligonucleotide) and treated with either 10 nM E2 (iNSE) or DMSO (iNSD).
Category A genes were induced by E2 and induction of these genes was inhibited after cotransfection with small inhibitory RNAs for SP1, SP3, and SP4 (combined; iSP). Table 1 summarizes one subset of genes induced by E2 (1·3- to 2·4-fold) in which transfection with iSP significantly decreased fold induction, whereas basal activity was minimally affected by iSP since the ratio of the DMSO values in the iSP and iNS treated groups (iSPD/iNSD) varied only from 0·81 to 1·77 and were considered to be unchanged. These were classified as +ESR1/SP (B0)-inducible genes. A second subset of genes was also induced by E2 (1·30- to 4·38-fold), but the iSPD/iNSD ratios varied from 1·35 to 3·9, suggesting that basal activities of these hormone-inducible genes were repressed by SP proteins. These genes were classified as +ESR1/SP (B+). The third subset of +ESR1/SP-regulated genes were induced 1·32- to 1·84-fold by E2 and after cotransfection with iSP, the fold induction responses were significantly decreased to 0·83- to 1·17-fold. This was accompanied by iSPD/iNSD ratios of 0·54-0·79. This subset of genes induced by E2 (category A) was designated at +ESR1/SP (B-), signifying that their basal activity was also SP dependent. There were 42 genes induced by E2 that were classified as +ESR1/SP (B0), +ESR1/SP (B+), or +ESR1/SP (B-).
The classification of genes downregulated by E2 and reversed after cotransfection with iSP (category B) was also subdivided into -ESR1/SP (B0), -ESR1/SP (B+), and -ESR1/SP (B-) subsets using the same criteria as indicated for genes induced by E2 (category A). Table 2 summarizes the 21 genes decreased (0·45 to 0·80) after treatment with E2 (+ iNS) and this response was reversed after transfection with iSP (category B). The iSPD/iNSD ratios (0·80-1·10) were relatively unchanged and these genes were designated as -ESR1/SP (B0). We detected only a single gene classified as -ESR1/SP (B+) where loss of SP protein increased basal activity. Sixty genes were classified as ESR1/SP (B-); E2 decreased expression (0·80-0·52) of these genes and transfection with iSP reversed this response (0·80-1·20) for 54 of these genes. Noticeably, the remaining six genes showed similar yet much more pronounced fold induction (1·35-4·55) after transfection with iSP. Thus, E2 decreased the expression of 75 genes that were SP dependent (category B) compared with the induction of 42 SP-dependent genes (category A).
Categories C and D contain 25 and 51 genes induced (≥1·3-fold) or repressed (≤ 0·8) by E2 respectively, and after transfection with iSP, the fold induction or repression was either unaffected or only partially modulated by SP protein knockdown. Despite the fact that E2-mediated modulation of these genes was SP independent, the categories C (+ER) and D (-ER) genes (Tables 3 and 4 respectively) were subdivided into B0, B+, and B- subcategories where iSPD/iNSD ratios were unchanged (B0), increased (B+), or decreased (B-). There were 5, 8, and 12 genes in the +ER (B0), +ER (B+), and +ER (B-) subcategories respectively, and 14, 4, and 33 genes in the -ER (B0), -ER (B+), and -ER (B-) subcategories respectively, indicating that the major subcategory for each set involved hormone-responsive genes whose basal expression was, in part, SP dependent (B-). Statistical analysis of the data was also carried out with relaxed conditions, which included a new cut-off P-value of 0·05, adjustment of the thresholds for inducibility (1·1 instead of 1·3) and repression (0·9 instead of 0·8), and no cross-array normalization. Our new analysis used the software package GeneSifter and the parameters used were selected as close as possible to those in the original analysis. The number of genes in the four different categories were 57 (A), 244 (B), 93 (C), and 350 (D) compared with 43 (A), 83 (B), 25 (C), and 51 (D) genes reported in Tables 1-4 respectively. In the less stringent analysis, only 40% of the E2-modulated genes were SP dependent and this change was primarily due to the large increase in the number of E2-repressed genes in category D.
GeneSpring BX7 software was used to investigate the statistical significance of genes induced or repressed by E2, and the adjusted P-values were obtained using Bonferroni correction. During the processing, GO terms on biological processes were assigned to each one of the significantly expressed genes in categories A-D. Tables 1-4 summarize the individual genes induced or repressed by E2 and also includes the GO terms associated with these genes. In categories A, B, C, and D, the hormone-induced or -repressed genes were associated with 37, 68, 19, and 78 different biological processes, and most processes contained only one gene with a maximum number of four genes per individual process. However, the largest number of genes in categories A, B, C, and D were genes that were not assigned to any specific biological process and these included 30, 56, 20, and 24 genes respectively. A comparison of the biological processes induced by E2 by ESR1/SP (category A) and SP-independent (category C) pathways exhibited minimal overlap with only ‘synaptic transmission’, ‘cell-cell signaling’, and ‘central nervous system’ genes activated in common. A comparison of the 68 and 74 biological processes downregulated by E2 in categories B (SP dependent) and D (SP independent) indicated that only nine processes were in common. These results suggest that the SP-dependent (A, B) and SP-independent (C and D) pathways for hormone-mediated induction or inhibition of gene expression primarily target different biological processes (GO terms). At the same time, it is important to emphasize that this minimal functional overlap in the GO categories could be attributed to incomplete or inaccurate annotation in the existing databases and could change if one accepts a less conservative gene selection criteria in terms of P-values or FDR or if multiple cell lines with multiple time points for data collection were used. Thus, further analysis is warranted.
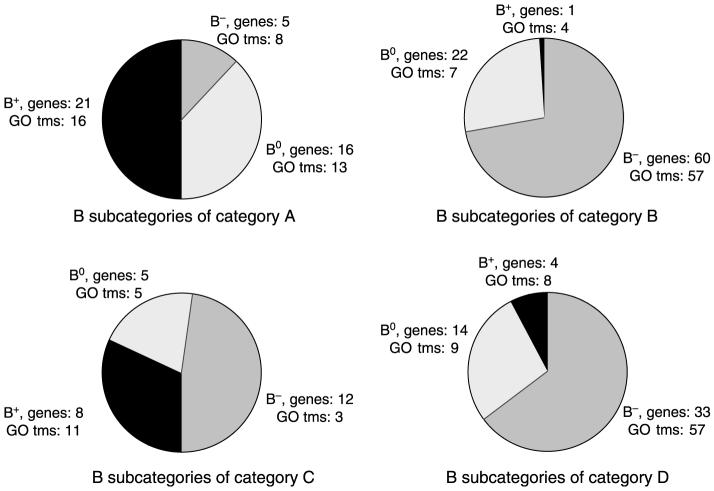
Distribution of genes in categories A-D and their subcategories B0, B+, and B- is illustrated in Fig. 2 along with the number of biological processes associated with the genes in each category. Not surprisingly, the results are complex with minimal overlap between categories.
Figure 2.
Subclassification of genes in categories A-D. The effects of iSP on basal activities (B0, B+, and B-) and the number of GO terms within each subclassification of categories A-D genes that are induced or repressed by E2 in MCF-7 cells are indicated (also see Tables 1-4).
The role of SP proteins in mediating hormonal regulation of select genes was also confirmed by real-time PCR, and the four genes selected include E2f1, Vegfa, Hmg, and Klk6. Although VEGFA was not detected in the microarray studies due to the stringent conditions, we included VEGFA since ongoing studies show consistent downregulation of this gene by E2 in MCF-7 cells (data not shown). Figure 3 illustrates the time course induction or repression of these genes after treatment with 20 nM E2 for 2, 4, 6, 9, and 24 h. Significant effects were observed within 2 h and persisted for up to 24 h. We used a higher concentration of E2 (20 nM) for the RT-PCR experiments and this was based on preliminary studies showing a maximal response observed using 20 nM E2. The role of iSP in mediating hormone-dependent activation of these genes was determined in MCF-7 cells (Fig. 4) and the results were similar to those observed in the microarray experiments. Moreover, we also observed that the antiestrogen ICI 182780 (fulvestrant) also significantly inhibited the E2-induced or -repressed genes. These results demonstrate that E2 induces and represses a diverse spectrum of genes in MCF-7 cells as previously reported (Soulez & Parker 2001, Inoue et al. 2002, Levenson et al. 2002, Lobenhofer et al. 2002, Coser et al. 2003, Cunliffe et al. 2003, Frasor et al. 2003, 2004, Scafoglio et al. 2006, Jonsson et al. 2007), and this study shows that >60% of these genes are dependent on SP proteins.
Figure 3.
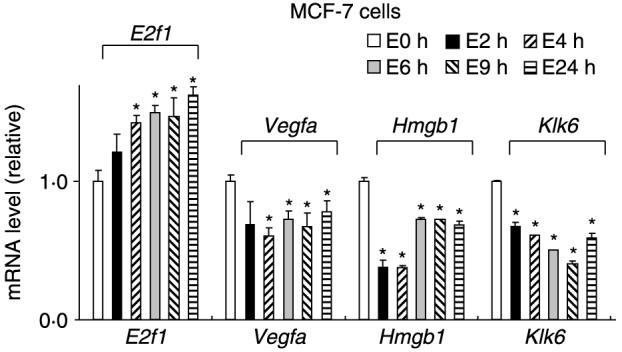
Time course effects of E2. MCF-7 cells were treated with 20 nM E2 for various times and expression of E2f1, Vegfa, Hmgb1, and Klk6 was determined (in triplicate) by real-time PCR (normalized to TBP) as outlined in the Materials and methods. Significant (P<0·05) induction or inhibition is indicated by *.
Figure 4.
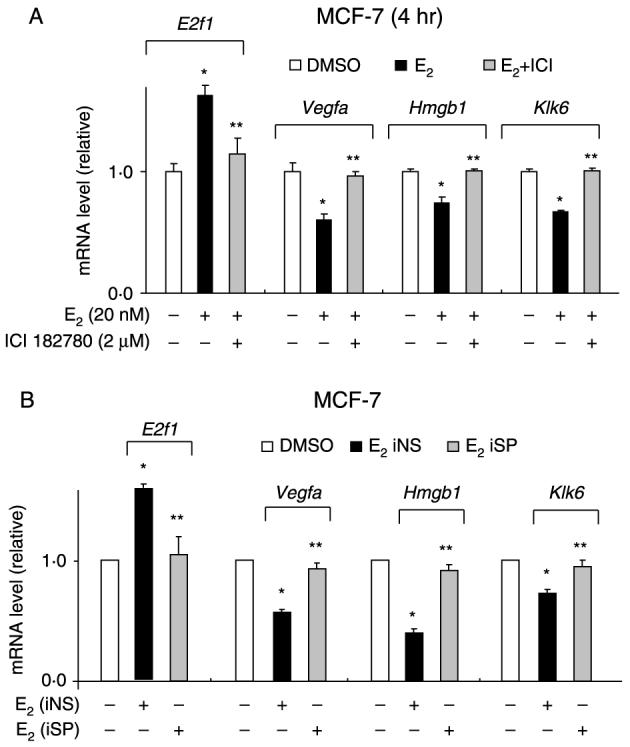
Effects of (A) RNA interference or (B) ICI 182780 on E2-dependent induction or repression of genes. MCF-7 cells were transfected with iNS or iSP and treated with DMSO or 20 nM E2 or treated with DMSO, E2 alone, or E2 plus 2 μM ICI 182780, and gene expression was determined by real-time PCR as described in the Materials and methods. Results are means±s.e.m. for three replicate determinations and significant (P<0·05) induction or repression by E2 (*) and reversal of these effects by iSP or ICI 182780 (**) is indicated.
Discussion
Gene expression profiling has been extensively used to understand global changes in gene expression in various endogenous biochemical processes and during development of various diseases including breast cancer (Sorlie et al. 2003, Andersson et al. 2005). The patterns of gene expression in different tumor types have been extensively investigated and are used for disease prognosis and for predicting responses to various chemotherapies. For example, gene expression profiling has been used for diagnosis, prognosis, and treatment of breast cancer metastasis and for recommending adjuvant chemotherapy for breast cancer (Driouch et al. 2007, Henry & Hayes 2007). Gene expression profiles in breast cancer cells have been extensively investigated in several different ER+ and ER- breast cancer cell lines, and the effects of time, concentration, and structure of ER agonists and antagonists and other growth regulatory factors on modulation of gene expression have been investigated (Soulez & Parker 2001, Inoue et al. 2002, Levenson et al. 2002, Lobenhofer et al. 2002, Coser et al. 2003, Cunliffe et al. 2003, Frasor et al. 2003, 2004, Scafoglio et al. 2006, Jonsson et al. 2007). Results of these studies demonstrate that hormone-activated genes and gene networks are highly complex and variable and demonstrate the broad effects of E2 on gene expression. For example, Frasor et al. (2003) showed that E2 regulated over 400 genes in 12 different functional categories in MCF-7 cells and the majority (76%) of these genes were downregulated by E2. Similar results were observed in this study (Fig. 1B and C). Frasor et al. also demonstrated distinct temporal differences in the expression of some E2-induced and repressed genes (Frasor et al. 2003, 2004).
Several studies show that SP proteins play a pivotal role in hormone-dependent induction of genes (Safe 2001, Safe & Kim 2004), and recent reports have shown that these transcription factors are also involved in gene repression (Higgins et al. 2006, Stossi et al. 2006). Recent studies on genome-wide interactions of ESR1 with DNA demonstrate that many ESR1-binding sites for specific genes are distal from their corresponding transcription start sites (Carroll & Brown 2006, Carroll et al. 2006, Lin et al. 2007), whereas most promoter studies have focused on proximal E2-responsive promoter sites. Thus, the E2-responsive proximal GC-sites identified in previous studies (Safe & Kim 2005) may also contain functional E2-responsive distal sites and further analysis of the potentially ESR1/SP-dependent genes listed in Tables 1 and 2 will have to examine both proximal and distal cis-elements. For example, based on proximal promoter analysis, we previously identified retinoic acid receptor α1 (RARA) as an E2-responsive gene regulated by ESR1/SP (Sun et al. 1998); however, Laganiere et al. (2005) used a functional genomics approach to identify a functional ERE 3·7 kb downstream from the RARA transcription start site. Another study demonstrated the potential involvement of GC-rich binding sites in the global regulation of some ESR1-dependent genes using a whole-genome cartography approach for identifying ESR1-binding sites (Lin et al. 2007).
Most of the previous research have focused on SP1 protein; however, studies in this laboratory have now shown that SP1, SP3, and SP4 proteins are overexpressed in breast cancer cells (Mertens-Talcott et al. 2007) and simultaneous knockdown of one or all three SP proteins block E2-dependent expression of the Rara, E2f, and Cad genes (Khan et al. 2007). These genes are primarily activated through ESR1/SP binding to GC-rich promoter sites (ER:DNA independent). In this study, we have selected a single concentration (10 nM) of E2 and one time point (6 hr) to investigate the role of SP proteins in mediating hormonal activation of genes in MCF-7 cells. This approach was not designed to identify all hormonally regulated genes but to investigate the importance of SP proteins in this induction response. Results of this study demonstrate that in MCF-7 cells treated with E2, 67 genes are induced and 134 genes are repressed (Fig. 1B), and this ratio of induced/repressed hormone-responsive genes is similar to the results of previous studies (Frasor et al. 2003). However, the use of RNA interference shows that out of these 201 genes, over 60% of the genes affected by E2 are also dependent on SP proteins (categories A and B). We also observed that within the four categories of genes (Tables 1-4 and Fig. 2), SP protein also markedly affected basal gene expression. By grouping genes into subcategories of B0, B+, and B-, we observed that among the 201 genes induced or repressed by E2, the basal activity of 57 genes was unaffected after transfection with iSP, whereas the basal activity of 144 genes was either decreased (110 genes; B-) or increased (34 genes; B+) (Fig. 3). Since SP proteins are important for constitutive expression of multiple mammalian and viral genes, it is not surprising that loss of SP1, SP3, and SP4 resulted in decreased basal expression of 110 out of 144 of those genes affected by iSP. However, SP protein knockdown also enhanced basal activity of 34 genes and this may be due to the SP3 protein that can act as both an enhancer and a suppressor of gene expression (Black et al. 2001, Suske et al. 2005). It should also be noted that knockdown of SP1, SP3, and SP4 proteins will not only affect ESR1/SP-regulated genes but may also result in altered expression of ESR1, coactivators, and other nuclear cofactors required for gene induction or expression. We are currently investigating the molecular mechanisms associated with the E2-dependent induction and repression of individual SP-dependent genes listed in Tables 1 and 2. These studies will determine the role of SP proteins and identify both proximal and distal GC-rich sites that are functional cis-elements in ESR1/SP action.
Analysis of the effects of SP knockdown on hormone-responsive gene expression revealed a fifth category (F) of genes (Fig. 1C) that were not induced or repressed by E2 in MCF-7 cells transfected with iNS. There were 85 E2-nonresponsive genes in category F; however, after transfection of iSP, treatment with E2 induced or repressed expression of these genes that were further subdivided into B0, B+, and B- subcategories. Category F genes were further subdivided into multiple (112) biological processes and only 30 of these were in common with biological processes associated with genes in categories A-D. It has been reported that SP proteins are overexpressed in multiple tumor types (Zannetti et al. 2000, Shi et al. 2001, Chiefari et al. 2002, Wang et al. 2003, Hosoi et al. 2004, Yao et al. 2004, Mertens-Talcott et al. 2007), and our recent studies have shown that SP proteins are highly expressed in breast cancer cells but barely detectable in non-transformed mammary cells (Mertens-Talcott et al. 2007). Lou and coworkers also showed that malignant transformation of human fibroblast cells results in an 8- to 18-fold increase in the expression of SP1 in the tumor cells compared with the parental cells (Lou et al. 2005). These studies suggest that over-expression of SP proteins contributes to tumor formation; thus, genes in category F, which regain hormone responsiveness in MCF-7 cells transfected with iSP, may play an endogenous role in normal or precancerous mammary cells where SP protein levels are relatively low. Their hormone responsiveness is subsequently silenced with increased expression of SP proteins found in MCF-7 and other breast cancer cell lines. Current studies are examining a variety of tumorigenic and non-tumorigenic mammary cells to further investigate the functions and changes in expression of category F genes and the role of SP proteins in mediating gene silencing.
In summary, results of these microarray studies confirm that in MCF-7 cells, E2 predominantly inhibits expression of genes. By combining the microarray data with simultaneous knockdown of SP1, SP3, and SP4 proteins, we have demonstrated that these transcription factors are critical regulators of hormone-dependent gene induction and repression. Moreover, the latter response has recently been confirmed in two studies on SP-dependent downregulation of KDR and cyclin G2 in MCF-7 cells treated with E2 (Stossi et al. 2006, Higgins et al. 2008). Previous studies show that VEGFA and KDR are induced by E2 in ZR-75 cells (Stoner et al. 2004, Higgins et al. 2006), whereas in MCF-7 cells, E2 decreases expression of both genes (Higgins et al. 2008; Figs 3 and 4). This unusual cell context-dependent difference in hormonal regulation of VEGFA/KDR in two ER-positive breast cancer cells lines may be due to altered expression of critical cofactors and this is currently being investigated. It should also be noted that ESR1 is constitutively associated with proximal E2-responsive GC-rich sequences and these interactions are not enhanced by E2 in a ChIP assay (Higgins et al. 2006, 2008, Khan et al. 2007). We are now examining coactivators as potential markers of ESR1/SP-mediated transactivation. Thus, in MCF-7 cells treated with E2 for 6 h, most hormonally regulated genes were SP dependent and, for the 76 genes in categories C and D where iSP did not affect hormone inducibility, loss of SP affected basal expression of 53 out of these 76 genes. Thus, among the 201 genes in categories A-D, loss of SP proteins affected hormone-induced or basal expression of over 88% (178/201) of all genes. It was also apparent that hormone-mediated responses such as increased cell proliferation involves mobilization genes that regulate a large number of functional processes (Tables 1-4), and current studies are focused on interconnections between pathways that lead to E2-induced proliferation and survival pathways in MCF-7 cells.
Acknowledgments
Funding
This work was supported by the National Institutes of Health (ES04917, CA104116, and ES09106) and the Texas Agricultural Experiment Station.
Footnotes
Declaration of interest
The authors declare that there is no conflict of interest that could be perceived as prejudicing the impartiality of the research reported.
References
- Andersson A, Eden P, Lindgren D, Nilsson J, Lassen C, Heldrup J, Fontes M, Borg A, Mitelman F, Johansson B, et al. Gene expression profiling of leukemic cell lines reveals conserved molecular signatures among subtypes with specific genetic aberrations. Leukemia. 2005;19:1042–1050. doi: 10.1038/sj.leu.2403749. [DOI] [PubMed] [Google Scholar]
- Black AR, Black JD, Azizkhan-Clifford J. Sp1 and Krüppel-like factor family of transcription factors in cell growth regulation and cancer. Journal of Cellular Physiology. 2001;188:143–160. doi: 10.1002/jcp.1111. [DOI] [PubMed] [Google Scholar]
- Blobel GA, Orkin SH. Estrogen-induced apoptosis by inhibition of the erythroid transcription factor GATA-1. Molecular and Cellular Biology. 1996;16:1687–1694. doi: 10.1128/mcb.16.4.1687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carroll JS, Brown M. Estrogen receptor target gene: an evolving concept. Molecular Endocrinology. 2006;20:1707–1714. doi: 10.1210/me.2005-0334. [DOI] [PubMed] [Google Scholar]
- Carroll JS, Meyer CA, Song J, Li W, Geistlinger TR, Eeckhoute J, Brodsky AS, Keeton EK, Fertuck KC, Hall GF, et al. Genome-wide analysis of estrogen receptor binding sites. Nature Genetics. 2006;38:1289–1297. doi: 10.1038/ng1901. [DOI] [PubMed] [Google Scholar]
- Chadwick CC, Chippari S, Matelan E, Borges-Marcucci L, Eckert AM, Keith JC, Jr, Albert LM, Leathurby Y, Harris HA, Bhat RA, et al. Identification of pathway-selective estrogen receptor ligands that inhibit NF-κB transcriptional activity. PNAS. 2005;102:2543–2548. doi: 10.1073/pnas.0405841102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiefari E, Brunetti A, Arturi F, Bidart JM, Russo D, Schlumberger M, Filetti S. Increased expression of AP2 and Sp1 transcription factors in human thyroid tumors: a role in NIS expression regulation? BMC Cancer. 2002;2:35. doi: 10.1186/1471-2407-2-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coser KR, Chesnes J, Hur J, Ray S, Isselbacher KJ, Shioda T. Global analysis of ligand sensitivity of estrogen inducible and suppressible genes in MCF7/BUS breast cancer cells by DNA microarray. PNAS. 2003;100:13994–13999. doi: 10.1073/pnas.2235866100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cunliffe HE, Ringner M, Bilke S, Walker RL, Cheung JM, Chen Y, Meltzer PS. The gene expression response of breast cancer to growth regulators: patterns and correlation with tumor expression profiles. Cancer Research. 2003;63:7158–7166. [PubMed] [Google Scholar]
- Driouch K, Landemaine T, Sin S, Wang S, Lidereau R. Gene arrays for diagnosis, prognosis and treatment of breast cancer metastasis. Clinical and Experimental Metastasis. 2007;24:575–585. doi: 10.1007/s10585-007-9110-x. [DOI] [PubMed] [Google Scholar]
- Evans RM. The nuclear receptor superfamily: a rosetta stone for physiology. Molecular Endocrinology. 2005;19:1429–1438. doi: 10.1210/me.2005-0046. [DOI] [PubMed] [Google Scholar]
- Frasor J, Danes JM, Komm B, Chang KC, Lyttle CR, Katzenellenbogen BS. Profiling of estrogen up- and down-regulated gene expression in human breast cancer cells: insights into gene networks and pathways underlying estrogenic control of proliferation and cell phenotype. Endocrinology. 2003;144:4562–4574. doi: 10.1210/en.2003-0567. [DOI] [PubMed] [Google Scholar]
- Frasor J, Stossi F, Danes JM, Komm B, Lyttle CR, Katzenellenbogen BS. Selective estrogen receptor modulators: discrimination of agonistic versus antagonistic activities by gene expression profiling in breast cancer cells. Cancer Research. 2004;64:1522–1533. doi: 10.1158/0008-5472.can-03-3326. [DOI] [PubMed] [Google Scholar]
- Ghisletti S, Meda C, Maggi A, Vegeto E. 17β-Estradiol inhibits inflammatory gene expression by controlling NF-κB intracellular localization. Molecular and Cellular Biology. 2005;25:2957–2968. doi: 10.1128/MCB.25.8.2957-2968.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall JM, Couse JF, Korach KS. The multifaceted mechanisms of estradiol and estrogen receptor signaling. Journal of Biological Chemistry. 2001;276:36869–36872. doi: 10.1074/jbc.R100029200. [DOI] [PubMed] [Google Scholar]
- Henry NL, Hayes DF. Use of gene-expression profiling to recommend adjuvant chemotherapy for breast cancer. Oncology. 2007;21:1301–1309. [PubMed] [Google Scholar]
- Higgins KJ, Liu S, Abdelrahim M, Yoon K, Vanderlaag K, Porter W, Metz RP, Safe S. Vascular endothelial growth factor receptor-2 expression is induced by 17β-estradiol in ZR-75 breast cancer cells by estrogen receptor α/Sp proteins. Endocrinology. 2006;147:3285–3295. doi: 10.1210/en.2006-0081. [DOI] [PubMed] [Google Scholar]
- Higgins KJ, Liu S, Abdelrahim M, Vanderlaag K, Liu X, Porter W, Metz R, Safe S. Vascular endothelial growth factor receptor-2 expression is downregulated by 17β-estradiol in MCF-7 breast cancer cells by estrogen receptor α/Sp proteins. Molecular Endocrinology. 2008;22:388–402. doi: 10.1210/me.2007-0319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hosoi Y, Watanabe T, Nakagawa K, Matsumoto Y, Enomoto A, Morita A, Nagawa H, Suzuki N. Up-regulation of DNA-dependent protein kinase activity and Sp1 in colorectal cancer. International Journal of Oncology. 2004;25:461–468. [PubMed] [Google Scholar]
- Hu J, Zou F, Wright FA. Practical FDR-based sample size calculations in microarray experiments. Bioinformatics. 2005;21:3264–3272. doi: 10.1093/bioinformatics/bti519. [DOI] [PubMed] [Google Scholar]
- Inadera H, Sekiya T, Yoshimura T, Matsushima K. Molecular analysis of the inhibition of monocyte chemoattractant protein-1 gene expression by estrogens and xenoestrogens in MCF-7 cells. Endocrinology. 2000;141:50–59. doi: 10.1210/endo.141.1.7233. [DOI] [PubMed] [Google Scholar]
- Inoue A, Yoshida N, Omoto Y, Oguchi S, Yamori T, Kiyama R, Hayashi S. Development of cDNA microarray for expression profiling of estrogen-responsive genes. Journal of Molecular Endocrinology. 2002;29:175–192. doi: 10.1677/jme.0.0290175. [DOI] [PubMed] [Google Scholar]
- Jonsson G, Staaf J, Olsson E, Heidenblad M, Vallon-Christersson J, Osoegawa K, de Jong P, Oredsson S, Ringner M, Hoglund M, et al. High-resolution genomic profiles of breast cancer cell lines assessed by tiling BAC array comparative genomic hybridization. Genes, Chromosomes and Cancer. 2007;46:543–558. doi: 10.1002/gcc.20438. [DOI] [PubMed] [Google Scholar]
- Kalaitzidis D, Gilmore TD. Transcription factor cross-talk: the estrogen receptor and NF-κB. Trends in Endocrinology and Metabolism. 2005;16:46–52. doi: 10.1016/j.tem.2005.01.004. [DOI] [PubMed] [Google Scholar]
- Katzenellenbogen JA, O’Malley BW, Katzenellenbogen BS. Tripartite steroid hormone receptor pharmacology - interaction with multiple effector sites as a basis for the cell- and promoter-specific action of these hormones. Molecular Endocrinology. 1996;10:119–131. doi: 10.1210/mend.10.2.8825552. [DOI] [PubMed] [Google Scholar]
- Khan S, Wu F, Liu S, Wu Q, Safe S. Role of specificity protein (Sp) transcription factors in estrogen-induced gene expression in MCF-7 breast cancer cells. Journal of Molecular Endocrinology. 2007;39:289–304. doi: 10.1677/JME-07-0043. [DOI] [PubMed] [Google Scholar]
- Laganiere J, Deblois G, Giguere V. Functional genomics identifies a mechanism for estrogen activation of the retinoic acid receptor alpha1 gene in breast cancer cells. Molecular Endocrinology. 2005;19:1584–1592. doi: 10.1210/me.2005-0040. [DOI] [PubMed] [Google Scholar]
- Levenson AS, Jordan VC. MCF-7: the first hormone-responsive breast cancer cell line. Cancer Research. 1997;57:3071–3078. [PubMed] [Google Scholar]
- Levenson AS, Svoboda KM, Pease KM, Kaiser SA, Chen B, Simons LA, Jovanovic BD, Dyck PA, Jordan VC. Gene expression profiles with activation of the estrogen receptor α-selective estrogen receptor modulator complex in breast cancer cells expressing wild-type estrogen receptor. Cancer Research. 2002;62:4419–4426. [PubMed] [Google Scholar]
- Lin CY, Vega VB, Thomsen JS, Zhang T, Kong SL, Xie M, Chiu KP, Lipovich L, Barnett DH, Stossi F, et al. Whole-genome cartography of estrogen receptor α binding sites. PLoS Genetics. 2007;3:e87. doi: 10.1371/journal.pgen.0030087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lobenhofer EK, Bennett L, Cable PL, Li L, Bushel PR, Afshari CA. Regulation of DNA replication fork genes by 17β-estradiol. Molecular Endocrinology. 2002;16:1215–1229. doi: 10.1210/mend.16.6.0858. [DOI] [PubMed] [Google Scholar]
- Lou Z, O’Reilly S, Liang H, Maher VM, Sleight SD, McCormick JJ. Down-regulation of overexpressed Sp1 protein in human fibrosarcoma cell lines inhibits tumor formation. Cancer Research. 2005;65:1007–1017. [PubMed] [Google Scholar]
- Matthews J, Gustafsson JA. Estrogen signaling: a subtle balance between ERα and ERβ. Molecular Interventions. 2003;3:281–292. doi: 10.1124/mi.3.5.281. [DOI] [PubMed] [Google Scholar]
- Mertens-Talcott SU, Chintharlapalli S, Li X, Safe S. The oncogenic microRNA-27a targets genes that regulate specificity protein (Sp) transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells. Cancer Research. 2007;67:11001–11011. doi: 10.1158/0008-5472.CAN-07-2416. [DOI] [PubMed] [Google Scholar]
- Nilsson S, Gustafsson JA. Biological role of estrogen and estrogen receptors. Critical Reviews in Biochemistry and Molecular Biology. 2002;37:1–28. doi: 10.1080/10409230290771438. [DOI] [PubMed] [Google Scholar]
- O’Malley BW. A life-long search for the molecular pathways of steroid hormone action. Molecular Endocrinology. 2005;19:1402–1411. doi: 10.1210/me.2004-0480. [DOI] [PubMed] [Google Scholar]
- Paech K, Webb P, Kuiper GG, Nilsson S, Gustafsson J, Kushner PJ, Scanlan TS. Differential ligand activation of estrogen receptors ERα and ERβ at AP1 sites. Science. 1997;277:1508–1510. doi: 10.1126/science.277.5331.1508. [DOI] [PubMed] [Google Scholar]
- Pelzer T, Neumann M, de Jager T, Jazbutyte V, Neyses L. Estrogen effects in the myocardium: inhibition of NF-κB DNA binding by estrogen receptor-α and -β. Biochemical and Biophysical Research Communications. 2001;286:1153–1157. doi: 10.1006/bbrc.2001.5519. [DOI] [PubMed] [Google Scholar]
- Pratt MA, Bishop TE, White D, Yasvinski G, Menard M, Niu MY, Clarke R. Estrogen withdrawal-induced NF-κB activity and bcl-3 expression in breast cancer cells: roles in growth and hormone independence. Molecular and Cellular Biology. 2003;23:6887–6900. doi: 10.1128/MCB.23.19.6887-6900.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Safe S. Transcriptional activation of genes by 17β-estradiol through estrogen receptor-Sp1 interactions. Vitamins and Hormones. 2001;62:231–252. doi: 10.1016/s0083-6729(01)62006-5. [DOI] [PubMed] [Google Scholar]
- Safe S, Kim K. Nuclear receptor-mediated transactivation through interaction with Sp proteins. Progress in Nucleic Acid Research and Molecular Biology. 2004;77:1–36. doi: 10.1016/S0079-6603(04)77001-4. [DOI] [PubMed] [Google Scholar]
- Scafoglio C, Ambrosino C, Cicatiello L, Altucci L, Ardovino M, Bontempo P, Medici N, Molinari AM, Nebbioso A, Facchiano A, et al. Comparative gene expression profiling reveals partially overlapping but distinct genomic actions of different antiestrogens in human breast cancer cells. Journal of Cellular Biochemistry. 2006;98:1163–1184. doi: 10.1002/jcb.20820. [DOI] [PubMed] [Google Scholar]
- Shi Q, Le X, Abbruzzese JL, Peng Z, Qian CN, Tang H, Xiong Q, Wang B, Li XC, Xie K. Constitutive Sp1 activity is essential for differential constitutive expression of vascular endothelial growth factor in human pancreatic adenocarcinoma. Cancer Research. 2001;61:4143–4154. [PubMed] [Google Scholar]
- Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A, Deng S, Johnsen H, Pesich R, Geisler S, et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. PNAS. 2003;100:8418–8423. doi: 10.1073/pnas.0932692100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soulez M, Parker MG. Identification of novel oestrogen receptor target genes in human ZR75-1 breast cancer cells by expression profiling. Journal of Molecular Endocrinology. 2001;27:259–274. doi: 10.1677/jme.0.0270259. [DOI] [PubMed] [Google Scholar]
- Stoner M, Wormke M, Saville B, Samudio I, Qin C, Abdelrahim M, Safe S. Estrogen regulation of vascular endothelial growth factor gene expression in ZR-75 breast cancer cells through interaction of estrogen receptor α and Sp proteins. Oncogene. 2004;23:1052–1063. doi: 10.1038/sj.onc.1207201. [DOI] [PubMed] [Google Scholar]
- Stossi F, Likhite VS, Katzenellenbogen JA, Katzenellenbogen BS. Estrogen-occupied estrogen receptor represses cyclin G2 gene expression and recruits a repressor complex at the cyclin G2 promoter. Journal of Biological Chemistry. 2006;281:16272–16278. doi: 10.1074/jbc.M513405200. [DOI] [PubMed] [Google Scholar]
- Sun G, Porter W, Safe S. Estrogen-induced retinoic acid receptor α1 gene expression: role of estrogen receptor-Sp1 complex. Molecular Endocrinology. 1998;12:882–890. doi: 10.1210/mend.12.6.0125. [DOI] [PubMed] [Google Scholar]
- Suske G, Bruford E, Philipsen S. Mammalian SP/KLF transcription factors: bring in the family. Genomics. 2005;85:551–556. doi: 10.1016/j.ygeno.2005.01.005. [DOI] [PubMed] [Google Scholar]
- Wang L, Wei D, Huang S, Peng Z, Le X, Wu TT, Yao J, Ajani J, Xie K. Transcription factor Sp1 expression is a significant predictor of survival in human gastric cancer. Clinical Cancer Research. 2003;9:6371–6380. [PubMed] [Google Scholar]
- Webb P, Nguyen P, Valentine C, Lopez GN, Kwok GR, McInerney E, Katzenellenbogen BS, Enmark E, Gustafsson J-Å, Nilsson S, et al. The estrogen receptor enhances AP-1 activity by two distinct mechanisms with different requirements for receptor transactivation functions. Molecular Endocrinology. 1999;13:1672–1685. doi: 10.1210/mend.13.10.0357. [DOI] [PubMed] [Google Scholar]
- Yao JC, Wang L, Wei D, Gong W, Hassan M, Wu TT, Mansfield P, Ajani J, Xie K. Association between expression of transcription factor Sp1 and increased vascular endothelial growth factor expression, advanced stage, and poor survival in patients with resected gastric cancer. Clinical Cancer Research. 2004;10:4109–4117. doi: 10.1158/1078-0432.CCR-03-0628. [DOI] [PubMed] [Google Scholar]
- Zannetti A, Del Vecchio S, Carriero MV, Fonti R, Franco P, Botti G, D’Aiuto G, Stoppelli MP, Salvatore M. Coordinate up-regulation of Sp1 DNA-binding activity and urokinase receptor expression in breast carcinoma. Cancer Research. 2000;60:1546–1551. [PubMed] [Google Scholar]