Abstract
Major histocompatibility complex (MHC) molecules bind short peptides resulting from intracellular processing of foreign and self proteins, and present them on the cell surface for recognition by T-cell receptors. We propose a new robust approach to quantitatively model the binding affinities of MHC molecules by quantitative structure-activity relationships (QSAR) that use the physical-chemical amino acid descriptors E1–E5. These QSAR models are robust, sequence-based, and can be used as a fast and reliable filter to predict the MHC binding affinity for large protein databases.
Keywords: Major histocompatibility complex, peptide binding affinity, quantitative structure-activity relationships, amino acid descriptors
INTRODUCTION
The ability to distinguish between foreign and self proteins is one of the most important characteristics of the immune system. Short peptides resulting from intracellular processing of foreign and self proteins form a complex with the major histocompatibility complex (MHC) molecules, and T cells recognize the MHC-bound peptides. There are two classes of MHC molecules: (i) MHC class I, which binds and presents to the T cells peptides derived from endogenously expressed proteins that are degraded by cytosolic proteases, usually by the proteasome; and (ii) MHC class II, which presents to the T cells peptides derived mainly from exogenous or transmembrane proteins, but also from cytosolic proteins that are degraded by various proteases which originate from the lysosomal compartment. Peptides that bind to MHC class I have a usual length of 8–12 amino acids, and are transported into the endoplasmic reticulum by the transporter associated with antigen processing (TAP), where they bind to MHC class I molecules. The MHC I molecule with bound peptide on the surface of infected cells and tumor cells can be recognized by a complementary-shaped T cell receptor (TCR) of CD8+ cytotoxic T cells (CTL), initiating the destruction of the cell containing the endogenous antigen.
The structural descriptors used in quantitative structure-activity relationships (QSAR) have a substantial contribution to a successful model. Because the present paper advocates a novel group of physical-chemical descriptors [1] for the modeling of the MHC binding, we briefly review the most important descriptors used in MHC QSAR. Lin et al. found that the isotropic surface area, ISA, and the electronic charge index, ECI, are effective descriptors for the QSAR modeling of peptide IC50 for class I MHC HLA-A*0201 [2]. CoMSIA molecular field descriptors were successful in 3D-QSAR for peptide IC50 binding to class I MHC HLA-A*0201 [3; 4], HLA-A2 superfamily [5], HLA-A3 superfamily (A*1101, A*0301, A*3101, and A*6801) [5–7], and mouse class I MHC alleles H2-Db, H2-Kb, and H2-Kk [8]. In a similar approach, CoMFA molecular field descriptors gave good predictions for the binding to HLA-A*0201 [3]. A series of sequence-based descriptors were evaluated by Guan et al. in QSAR predictions of binding affinities for HLA-A*0201 [9]. The first group of models used 93 amino acid properties from the AAindex database [10], whereas the second approach tested the principal component amino acid indices z1–z5 [11]. The study found that the QSAR models obtained with z1–z5 have the highest predictivity. Doytchinova et al. compared three classes of descriptors for their ability to predict peptide BL50 for HLA-A*0201 [12], namely a 20-element binary encoding for each amino acid, the z1–z5 scale, and a collection of molecular descriptors that included molecular connectivity indices, shape indices, electrotopological state indices, hydrophobicity, polarizability, surface area, and volume. The predictions tests showed that the best results are obtained with the simple 20-element binary encoding, followed by the z1–z5 scale, whereas the molecular descriptors did not provide meaningful predictions. The 20-element binary encoding approach [13; 14] was extended to include also cross-interaction terms [5; 15–17], with good results for HLA-A*0201 [18], the HLA-A3 superfamily [19], and MHC class II alleles [20].
The heuristic molecular lipophilicity potential (HMLP) [21] was used by Chou and co-workers to quantify the lipophilicity and hydrophilicity of amino acid side chains [22]. HMLP descriptors were also used to model the binding affinity of class I MHC peptides [23] and for QSAR studies of neuraminidase inhibitors [24]. A support vector machines regression approach in the quantitative modeling of the peptide binding affinity for MHC molecules [25] was applied by Liu et al. to model the peptide IC50 for mouse class I MHC alleles H2-Db, H2-Kb, and H2-Kk [26]. Each amino acid in a peptide was encoded with 17 physico-chemical properties, such as polarity, isoelectric point, volume, and hydrophobicity. Genetic function approximation and genetic partial least squares were used by Davies et al. to obtain QSAR models for the binding level BL50 of 118 peptides with class I MHC HLA-A*0201 [27]. The QSAR descriptors for BL50 modeling represent peptide-MHC interaction energies computed with AMBER. The average relative binding (ARB) matrix [28] is a quantitative tool that was calibrated for the prediction of binding affinities [29]. Using experimental data from the Immune Epitope Database and Analysis Resource (IEDB) [30], Bui et al. computed 85 matrices for class I MHC alleles and 13 matrices for class II MHC alleles. The ARB matrices were used to model the MHC class I pathway for several alleles [31].
High quality predictions for protein and peptide properties may be obtained only with a proper encoding of the amino acids sequence into a series of structural descriptors. As example of successful amino acids descriptors we mention here the pseudo-amino acid composition indices [32], which were used to model a wide range of bioinformatics problems, such as classification of membrane proteins [33], protease classification [34], protein-protein interactions [35], and protein subcellular location [32]. From the large list of machine learning procedures used in bioinformatics, we mention here k-nearest neighbors classifier [36], partial least squares [8; 13; 18; 27], artificial neural networks (ANN) [37–39], support vector machines (SVM) [33; 40; 41], and LogitBoost [42]. The combination of several machine learning predictions with ensemble, jury or fusion methods proved to be a very efficient strategy in order to obtain robust and predictive models [36; 43–45].
Our approach for the prediction of MHC binding affinity is based on five physical-chemical descriptors E1–E5 [1], where each amino acid position in a peptide is encoded by the five descriptors of the amino acid type at that position. The descriptors capture the physical-chemical similarity of amino acids, and are able to encode the peptide sequence more efficiently than previously used representations. Especially our method can still be applied when for certain peptide positions not all 20 amino acids are present in the training set. Most current QSAR models for predicting binding affinities of MHC-binding peptides aim for a high goodness-of-fit criteria in developing the mathematical models. Typically they use the 20 amino acids in a binary code that leads to a large number of variables, 20×n, to encode a peptide of length n. In many practical applications this number exceeds the available data points used as the training set for generating the computational model. High-quality predictions of the MHC-peptide binding affinity depends both on a reliable set of amino acid descriptors and on the use of predictive machine learning and pattern recognition models.
A large variety of methods have been proposed for the prediction of peptide binding affinity to MHC molecules: position specific scoring matrices [46; 47], ANN [37–39], hidden Makov models [48; 49], SVM [40; 41]. In this paper we explore new avenues to find a minimal number of variables in the mathematical modeling of binding affinities of peptides to MHC molecules. We previously demonstrated that our descriptors are a suitable mathematical framework to find sequence similarities between cross-reactive IgE epitopes using a property distance measure (PD value) [50–55]. We show here that the E1–E5 descriptors and three multivariate quantitative structure-activity relationships (QSAR) methods (multilinear regression (MLR), partial least squares (PLS), and multi-layer feed-forward artificial neural networks (ANN), can be efficiently used to find robust quantitative models of MHC binding, and to explore the physical-chemical properties that determine the binding interaction between the MHC molecule and ligand peptides. Our new method is illustrated for the binding affinities (IC50) for the class I MHC HLA-A*0201 of a set of 152 nona-peptides [3].
MATERIALS AND METHODS
Data Set
The sequences and binding affinities for class I MHC HLA-A*0201 of 152 nona-peptides (Table 5) were taken from a recent study in which the IC50 of these peptides was modeled with the CoMFA and CoMSIA techniques [3]. We have selected the same set of nona-peptides in order to compare the QSAR value of the E1–E5 descriptors [1] with that of well-established QSAR models CoMFA and CoMSIA.
Table 5.
Sequences of the 152 Nona-Peptides, their Experimental pIC50 Values, Calibration and Prediction Values Obtained with the MLR QSAR Model 7 (Table 1), and the Binding Affinities Predicted with BIMAS, SYFPEITHI, and Rankpep Score
| No | Peptide | pIC50 Exp | MLR Cal | MLR Pre | BIMAS | SYFPEITHI | Rankpep score |
|---|---|---|---|---|---|---|---|
| 1 | VALVGLFVL | 5.146 | 5.838 | 6.445 | 10.207 | 23 | 33.0 |
| 2 | VCMTVDSLV | 5.146 | 5.611 | 5.656 | 2.856 | 12 | 44.0 |
| 3 | HLESLFTAV | 5.301 | 6.398 | 6.659 | 0.288 | 21 | 85.0 |
| 4 | LLGCAANWI | 5.301 | 5.681 | 5.664 | 97.547 | 21 | 51.0 |
| 5 | GTLVALVGL | 5.342 | 5.553 | 5.491 | 2.525 | 24 | 35.0 |
| 6 | SAANDPIFV | 5.342 | 6.069 | 6.453 | 5.313 | 18 | 34.0 |
| 7 | TTAEEAAGI | 5.380 | 5.835 | 5.749 | 0.594 | 19 | 52.0 |
| 8 | LLSCLGCKI | 5.447 | 6.277 | 5.958 | 17.736 | 23 | 52.0 |
| 9 | LQTTIHDII | 5.501 | 5.615 | 5.840 | 0.361 | 9 | 22.0 |
| 10 | TLLVVMGTL | 5.580 | 6.282 | 6.482 | 23.633 | 24 | 76.0 |
| 11 | LTVILGVLL | 5.580 | 5.574 | 5.588 | 0.504 | 19 | 25.0 |
| 12 | AMFQDPQER | 5.740 | 5.881 | 5.903 | 0.040 | 13 | 18.0 |
| 13 | HLLVGSSGL | 5.792 | 6.287 | 6.374 | 2.687 | 25 | 63.0 |
| 14 | SLHVGTQCA | 5.842 | 5.672 | 5.656 | 4.968 | 20 | 47.0 |
| 15 | ALPYWNFAT | 5.869 | 6.181 | 6.221 | 43.222 | 16 | 29.0 |
| 16 | LLVVMGTLV | 5.869 | 6.415 | 6.180 | 118.238 | 22 | 70.0 |
| 17 | SLNFMGYVI | 5.881 | 5.898 | 5.869 | 4.277 | 21 | 57.0 |
| 18 | NLQSLTNLL | 6.000 | 6.319 | 6.443 | 21.362 | 24 | 82.0 |
| 19 | GIGILTVIL | 6.000 | 5.665 | 5.807 | 1.204 | 24 | 52.0 |
| 20 | FVTWHRYHL | 6.025 | 5.835 | 6.037 | 8.598 | 15 | 23.0 |
| 21 | TVILGVLLL | 6.072 | 6.791 | 6.610 | 4.299 | 23 | 55.0 |
| 22 | WLEPGPVTA | 6.082 | 7.184 | 7.216 | 1.463 | 21 | 36.0 |
| 23 | WTDQVPFSV | 6.145 | 6.706 | 7.071 | 10.309 | 16 | 27.0 |
| 24 | QVMSLHNLV | 6.170 | 6.124 | 6.089 | 22.517 | 17 | 63.0 |
| 25 | DPKVKQWPL | 6.176 | 5.632 | 5.593 | 0.003 | 9 | −19.0 |
| 26 | AIAKAAAAV | 6.176 | 6.689 | 6.733 | 9.563 | 25 | 93.0 |
| 27 | ITSQVPFSV | 6.196 | 6.744 | 6.811 | 9.525 | 18 | 48.0 |
| 28 | ALAKAAAAI | 6.211 | 6.601 | 6.267 | 10.433 | 25 | 102.0 |
| 29 | GLGQVPLIV | 6.301 | 7.026 | 6.824 | 28.516 | 22 | 65.0 |
| 30 | MLDLQPETT | 6.335 | 6.218 | 6.511 | 2.483 | 16 | 38.0 |
| 31 | LLSSNLSWL | 6.342 | 6.365 | 6.240 | 459.398 | 26 | 80.0 |
| 32 | GLACHQLCA | 6.380 | 5.749 | 5.758 | 4.968 | 18 | 30.0 |
| 33 | VLHSFTDAI | 6.380 | 7.086 | 7.623 | 33.024 | 23 | 82.0 |
| 34 | AAAKAAAAV | 6.398 | 5.930 | 5.909 | 0.966 | 21 | 69.0 |
| 35 | LIGNESFAL | 6.415 | 6.519 | 6.365 | 28.962 | 21 | 61.0 |
| 36 | ALAKAAAAV | 6.419 | 6.945 | 6.866 | 69.552 | 27 | 103.0 |
| 37 | ILTVILGVL | 6.419 | 6.439 | 6.458 | 4.452 | 25 | 68.0 |
| 38 | LLAVGATKV | 6.477 | 5.956 | 5.904 | 118.238 | 28 | 80.0 |
| 39 | MLLAVLYCL | 6.477 | 7.087 | 6.799 | 309.050 | 27 | 77.0 |
| 40 | KLPQLCTEL | 6.484 | 6.495 | 6.212 | 74.768 | 24 | 69.0 |
| 41 | AVAKAAAAV | 6.495 | 6.159 | 6.354 | 6.086 | 21 | 87.0 |
| 42 | ALAKAAAAL | 6.511 | 6.293 | 6.247 | 21.362 | 27 | 99.0 |
| 43 | WILRGTSFV | 6.556 | 7.094 | 7.004 | 895.230 | 24 | 58.0 |
| 44 | IISCTCPTV | 6.580 | 6.640 | 6.651 | 16.258 | 23 | 57.0 |
| 45 | AAGIGILTV | 6.581 | 6.051 | 5.969 | 2.222 | 25 | 40.0 |
| 46 | FLGGTPVCL | 6.623 | 6.713 | 6.673 | 98.267 | 25 | 54.0 |
| 47 | ALIHHNTHL | 6.623 | 5.952 | 5.846 | 21.362 | 24 | 66.0 |
| 48 | ILDEAYVMA | 6.623 | 6.499 | 6.153 | 20.776 | 20 | 62.0 |
| 49 | NLSWLSLDV | 6.639 | 6.528 | 6.975 | 69.552 | 22 | 62.0 |
| 50 | YMIMVKCWM | 6.663 | 6.650 | 7.122 | 90.776 | 16 | 32.0 |
| 51 | YLEPGPVTA | 6.668 | 7.061 | 7.252 | 1.463 | 23 | 47.0 |
| 52 | VLQAGFFLL | 6.682 | 6.730 | 6.739 | 400.203 | 23 | 67.0 |
| 53 | LLWFHISCL | 6.682 | 6.532 | 6.733 | 693.274 | 26 | 55.0 |
| 54 | GTLGIVCPI | 6.714 | 6.766 | 6.697 | 1.233 | 21 | 32.0 |
| 55 | TLHEYMLDL | 6.726 | 7.283 | 7.390 | 201.447 | 24 | 81.0 |
| 56 | VILGVLLLI | 6.785 | 7.756 | 7.660 | 20.753 | 26 | 75.0 |
| 57 | VTWHRYHLL | 6.793 | 6.000 | 5.844 | 6.280 | 17 | 22.0 |
| 58 | TLDSQVMSL | 6.793 | 6.940 | 6.874 | 19.653 | 26 | 93.0 |
| 59 | PLLPIFFCL | 6.796 | 7.761 | 8.028 | 19.163 | 21 | 48.0 |
| 60 | TLGIVCPIC | 6.815 | 6.651 | 6.910 | 2.037 | 11 | 32.0 |
| 61 | CLTSTVQLV | 6.832 | 7.181 | 7.358 | 159.970 | 23 | 88.0 |
| 62 | HLYQGCQVV | 6.832 | 6.928 | 6.944 | 3.103 | 23 | 71.0 |
| 63 | ILLLCLIFL | 6.845 | 6.949 | 6.158 | 1699.774 | 28 | 58.0 |
| 64 | FLCKQYLNL | 6.875 | 8.128 | 8.196 | 147.401 | 22 | 65.0 |
| 65 | FAFRDLCIV | 6.886 | 6.869 | 6.756 | 15.504 | 20 | 28.0 |
| 66 | QLFHLCLII | 6.886 | 7.015 | 7.111 | 15.827 | 21 | 67.0 |
| 67 | FLEPGPVTA | 6.898 | 6.993 | 6.971 | 1.463 | 22 | 44.0 |
| 68 | ALAKAAAAA | 6.947 | 6.816 | 6.557 | 4.968 | 21 | 83.0 |
| 69 | ITDQVPFSV | 6.947 | 6.910 | 6.556 | 3.810 | 18 | 37.0 |
| 70 | LMAVVLASL | 6.954 | 6.761 | 6.803 | 60.325 | 30 | 79.0 |
| 71 | YVITTQHWL | 6.983 | 6.871 | 7.238 | 47.291 | 19 | 42.0 |
| 72 | LLCLIFLLV | 6.996 | 7.324 | 7.262 | 224.653 | 23 | 63.0 |
| 73 | HLAVIGALL | 7.000 | 6.462 | 6.529 | 0.726 | 24 | 65.0 |
| 74 | ALCRWGLLL | 7.000 | 6.948 | 6.955 | 21.362 | 23 | 55.0 |
| 75 | ITAQVPFSV | 7.020 | 7.091 | 7.299 | 9.525 | 20 | 46.0 |
| 76 | YLEPGPVTL | 7.058 | 7.033 | 6.703 | 6.289 | 29 | 63.0 |
| 77 | YTDQVPFSV | 7.066 | 6.849 | 6.871 | 10.309 | 18 | 38.0 |
| 78 | NLYVSLLLL | 7.114 | 6.967 | 6.670 | 157.227 | 26 | 77.0 |
| 79 | NLGNLNVSI | 7.119 | 6.528 | 7.061 | 10.433 | 23 | 74.0 |
| 80 | ILHNGAYSL | 7.127 | 6.962 | 6.621 | 36.316 | 27 | 73.0 |
| 81 | HLYSHPIIL | 7.131 | 7.064 | 7.299 | 0.953 | 21 | 70.0 |
| 82 | SIISAVVGI | 7.159 | 7.100 | 7.124 | 3.299 | 27 | 83.0 |
| 83 | VVMGTLVAL | 7.174 | 6.858 | 7.162 | 27.042 | 25 | 74.0 |
| 84 | ITFQVPFSV | 7.179 | 7.422 | 7.435 | 35.242 | 19 | 42.0 |
| 85 | YLEPGPVTI | 7.187 | 6.852 | 6.739 | 3.071 | 27 | 66.0 |
| 86 | FTDQVPFSV | 7.212 | 6.857 | 6.586 | 10.309 | 17 | 35.0 |
| 87 | GLSRYVARL | 7.248 | 7.148 | 7.169 | 49.134 | 28 | 87.0 |
| 88 | LLAQFTSAI | 7.301 | 6.915 | 6.675 | 67.396 | 26 | 96.0 |
| 89 | YMLDLQPET | 7.310 | 7.673 | 8.002 | 375.567 | 21 | 41.0 |
| 90 | VLLDYQGML | 7.328 | 7.645 | 7.368 | 71.619 | 25 | 70.0 |
| 91 | YLEPGPVTV | 7.342 | 7.103 | 7.369 | 20.476 | 29 | 67.0 |
| 92 | RLMKQDFSV | 7.342 | 8.027 | 8.032 | 1492.586 | 23 | 74.0 |
| 93 | ILSPFMPLL | 7.347 | 7.337 | 7.394 | 317.403 | 26 | 77.0 |
| 94 | KLHLYSHPI | 7.352 | 7.483 | 7.503 | 36.515 | 21 | 61.0 |
| 95 | YLSPGPVTA | 7.383 | 7.405 | 7.372 | 22.853 | 24 | 55.0 |
| 96 | ALAKAAAAM | 7.398 | 6.547 | 6.697 | 4.968 | 21 | 93.0 |
| 97 | IIDQVPFSV | 7.398 | 7.841 | 7.928 | 37.718 | 22 | 60.0 |
| 98 | ITMQVPFSV | 7.398 | 7.035 | 6.966 | 35.242 | 19 | 47.0 |
| 99 | YMNGTMSQV | 7.398 | 7.234 | 7.381 | 531.455 | 23 | 80.0 |
| 100 | SVYDFFVWL | 7.444 | 6.939 | 6.853 | 973.849 | 21 | 51.0 |
| 101 | ITWQVPFSV | 7.463 | 7.538 | 7.773 | 79.056 | 19 | 37.0 |
| 102 | KIFGSLAFL | 7.478 | 7.766 | 7.865 | 481.186 | 28 | 69.0 |
| 103 | ITYQVPFSV | 7.480 | 7.246 | 7.568 | 30.479 | 19 | 44.0 |
| 104 | GLYSSTVPV | 7.481 | 7.791 | 7.816 | 222.566 | 26 | 84.0 |
| 105 | VMGTLVALV | 7.553 | 6.937 | 6.787 | 196.407 | 26 | 79.0 |
| 106 | LLLCLIFLL | 7.585 | 7.279 | 7.033 | 1792.489 | 29 | 65.0 |
| 107 | SLDDYNHLV | 7.585 | 7.186 | 7.033 | 114.065 | 25 | 70.0 |
| 108 | ALVGLFVLL | 7.585 | 7.070 | 7.550 | 40.589 | 27 | 78.0 |
| 109 | YLSPGPVTV | 7.642 | 7.560 | 7.349 | 319.939 | 30 | 75.0 |
| 110 | VLIQRNPQL | 7.644 | 7.705 | 7.609 | 36.316 | 24 | 67.0 |
| 111 | SLYADSPSV | 7.658 | 7.823 | 7.588 | 222.566 | 26 | 82.0 |
| 112 | RLLQETELV | 7.682 | 7.300 | 7.266 | 126.098 | 24 | 83.0 |
| 113 | ILSQVPFSV | 7.699 | 8.070 | 8.106 | 685.783 | 24 | 81.0 |
| 114 | GLYSSTVPV | 7.699 | 7.791 | 7.816 | 222.566 | 26 | 84.0 |
| 115 | IMDQVPFSV | 7.719 | 7.770 | 7.705 | 198.115 | 22 | 60.0 |
| 116 | QLFEDNYAL | 7.764 | 6.963 | 6.884 | 324.068 | 25 | 64.0 |
| 117 | ALMDKSLHV | 7.770 | 7.882 | 7.897 | 1055.104 | 26 | 85.0 |
| 118 | YLYPGPVTA | 7.772 | 8.000 | 7.982 | 73.129 | 25 | 51.0 |
| 119 | YAIDLPVSV | 7.796 | 7.835 | 7.965 | 18.219 | 24 | 44.0 |
| 120 | YLAPGPVTV | 7.818 | 7.583 | 7.642 | 319.939 | 32 | 73.0 |
| 121 | FVWLHYYSV | 7.824 | 7.419 | 7.563 | 348.534 | 19 | 42.0 |
| 122 | MLGTHTMEV | 7.845 | 6.966 | 6.867 | 118.238 | 24 | 87.0 |
| 123 | VVLGVVFGI | 7.845 | 8.072 | 7.974 | 76.598 | 22 | 59.0 |
| 124 | LLFGYPVYV | 7.886 | 8.162 | 8.223 | 2406.151 | 28 | 71.0 |
| 125 | ILKEPVHGV | 7.921 | 7.836 | 7.810 | 39.025 | 30 | 66.0 |
| 126 | MMWYWGPSL | 7.921 | 7.789 | 7.662 | 217.695 | 21 | 34.0 |
| 127 | YLMPGPVTV | 7.932 | 7.932 | 7.926 | 1183.775 | 31 | 74.0 |
| 128 | WLDQVPFSV | 7.939 | 8.139 | 7.978 | 742.259 | 22 | 60.0 |
| 129 | ILAQVPFSV | 7.939 | 8.077 | 7.996 | 685.783 | 26 | 79.0 |
| 130 | KTWGQYWQV | 7.955 | 7.829 | 7.416 | 315.701 | 18 | 25.0 |
| 131 | ALMPLYACI | 8.000 | 7.325 | 7.765 | 57.902 | 26 | 77.0 |
| 132 | YLAPGPVTA | 8.032 | 7.442 | 7.459 | 22.853 | 26 | 53.0 |
| 133 | WLSLLVPFV | 8.048 | 7.543 | 7.852 | 4047.231 | 26 | 63.0 |
| 134 | YLYPGPVTV | 8.051 | 8.058 | 8.114 | 1023.805 | 31 | 71.0 |
| 135 | FLLSLGIHL | 8.053 | 7.372 | 7.427 | 363.588 | 24 | 65.0 |
| 136 | LLMGTLGIV | 8.097 | 7.694 | 7.983 | 53.631 | 29 | 78.0 |
| 137 | YLWPGPVTV | 8.125 | 8.133 | 8.103 | 2655.495 | 31 | 64.0 |
| 138 | ILMQVPFSV | 8.125 | 8.187 | 8.061 | 2537.396 | 25 | 80.0 |
| 139 | FLLTRILTI | 8.149 | 8.180 | 8.134 | 408.402 | 28 | 85.0 |
| 140 | GLLGWSPQA | 8.237 | 7.921 | 8.215 | 18.382 | 20 | 46.0 |
| 141 | YLFPGPVTV | 8.237 | 8.103 | 8.184 | 1183.775 | 31 | 69.0 |
| 142 | ILYQVPFSV | 8.310 | 8.226 | 8.178 | 2194.505 | 25 | 77.0 |
| 143 | GILTVILGV | 8.347 | 7.879 | 7.641 | 81.385 | 27 | 67.0 |
| 144 | YLMPGPVTA | 8.367 | 7.846 | 7.873 | 84.555 | 25 | 54.0 |
| 145 | NMVPFFPPV | 8.398 | 8.168 | 8.204 | 362.675 | 22 | 50.0 |
| 146 | ILDQVPFSV | 8.481 | 7.968 | 7.782 | 274.313 | 24 | 70.0 |
| 147 | YLFPGPVTA | 8.495 | 8.055 | 8.035 | 84.555 | 25 | 49.0 |
| 148 | YLWPGPVTA | 8.495 | 8.093 | 8.048 | 189.678 | 25 | 44.0 |
| 149 | YLDQVPFSV | 8.638 | 8.086 | 7.955 | 742.259 | 24 | 71.0 |
| 150 | FLDQVPFSV | 8.658 | 8.043 | 7.737 | 742.259 | 23 | 68.0 |
| 151 | ILFQVPFSV | 8.699 | 8.240 | 8.351 | 2537.396 | 25 | 75.0 |
| 152 | ILWQVPFSV | 8.770 | 8.249 | 8.190 | 5691.997 | 25 | 70.0 |
E1–E5 Physical-Chemical Descriptors
Numerous amino acids properties were developed for predicting a wide range of protein proteins, such as secondary structure, fold type, or subcellular localization. However, these scales numerically encode similar or related amino acids property, such as hydrophobicity or polarity. In order to cluster together related numerical scales, the principal components-like amino acids descriptors E1–E5 were developed [1]. Starting from 237 physical-chemical properties for all 20 naturally occurring amino acids, multidimensional scaling was used to condense the structural information into five descriptors. The mathematical procedure used in deriving these five descriptors ensures that the main variations of all 237 properties for the 20 amino acids are reflected by E1–E5. Every position in a nona-peptide was characterized by five E1–E5 descriptors of the corresponding amino acid, giving a vector with 45 components for each peptide. Similarly with the computation of IgE epitopes sequence similarity index PD [50; 52], each E component was weighted with the square root of the corresponding eigenvalue.
Strategies to Find a Robust MLR Model
To find a robust MLR QSAR model of the binding affinities, standard multiple linear regression methods were then applied by progressively reducing the number of parameters from 45 to 9 in blocks of 9. The model with all 45 components was used as a reference model. Two different strategies were tested in this reduction process: (1) a position independent strategy where in each round 9 descriptors with the lowest absolute correlation coefficient to the IC50 values were discarded and (2) a position dependent strategy where we eliminated in each round the descriptors with the lowest absolute correlation coefficient IC50 of each position in the nona-peptide.
Software
The PLS models were computed with Unscrambler (CAMO Inc., Corvalis, OR, http://www.camo.com), while MLR and ANN models were obtained with in-house developed C programs.
Artificial Neural Networks
The present study uses multilayer feed-forward neural networks provided with a single hidden layer. The number of neurons in the hidden layer was selected on the basis of systematic empirical trials in which ANNs with increasing number of hidden neurons were trained to predict the experimental pIC50 binding affinities. Each network was provided with a bias neuron connected to all neurons in the hidden and output layers, and with one output neuron providing the calculated pIC50 value.
Activation Functions
The most commonly used activation function in biological applications of neural networks has a sigmoidal shape and takes values between 0 and 1. For large negative arguments its value is close to 0, and practice demonstrated that ANN training is difficult in such conditions. To overcome this deficiency of the sigmoid function, the hyperbolic tangent (tanh) which takes values between −1 and 1 was used in the present study for the hidden and output layers.
Preprocessing of the Data
Each component of the input (PLS factors) and output (pIC50 binding affinities) patterns was linearly scaled between −0.9 and 0.9. For the tanh output activation function the scaling is required by the range of values of the function, while for the linear function experiments showed that a linear scaling improves the learning process.
Learning Method
The training of the ANNs was performed with the Polack-Ribiere method, for 2000 epochs. One epoch corresponds to the presentation of one complete set of examples. The patterns were presented randomly to the network, and the weights were updated after the presentation of each pattern. Random values between −0.1 and 0.1 were used as initial weights.
Performance Indicators
The performances of the neural networks were evaluated both for the model calibration and prediction. The quality of model calibration is estimated by comparing the calculated pIC50 during the training phase (pIC50calc) with the target values (pIC50exp), while the predictive quality was estimated by a cross-validation method by comparing the predicted (pIC50pr) and experimental values. In order to compare the performance of the ANN models with the statistical results of the MLR models, we have used the correlation coefficient r and the standard deviation s of the linear correlation between experimental and calibration or prediction pIC50: pIC50exp = A + B·pIC50calc/pr. In order to evaluate the effect of random initial weights, each ANN was trained 10 times, and in each case we obtained the same values for the QSAR statistical indices, at the precision reported in this paper.
Cross-Validation
Because the scope of our study is to develop quantitative models that can offer reliable predictions for the pIC50 of nona-peptides not used in the calibration of the QSAR model, we have estimated the prediction power of the MLR and ANN models with the leave-one-out (LOO) and leave-10%-out (L10%O) cross-validation methods. The LOO (or the “jackknife test”) test is a rigorous and objective method to evaluate the accuracy of a statistical prediction method, as shown in a comprehensive review [56] and in numerous bioinformatics applications [36; 43–45; 57]. In the L10%O algorithm 10% of the patterns are selected from the complete data set and form the prediction set. In the following step the QSAR model is calibrated with a learning set consisting of the remaining 90% of the data and the neural model obtained in the calibration phase is used to predict the pIC50 values for the patterns in the prediction set. This procedure is repeated ten times, until all patterns are selected for prediction once and only once. In order to compare the QSAR models obtained with MLR, PLS and ANN, we will give the correlation coefficient and standard deviation between experimental and calculated pIC50 for calibration/training/fitting (rcal and scal) and for the cross-validation prediction (rpre and spre). Some QSAR studies, including the CoMFA and CoMSIA models for the class I MHC HLA-A*0201 binding of the 152 nona-peptides [3], use as prediction parameter the leave-one-out q2 statistics:
In order to compare our E1–E5 MLR and ANN models with the CoMFA and CoMSIA results, we will report also the q2 statistics for LOO and L10%O cross-validation experiments.
RESULTS
Linear Regression Model
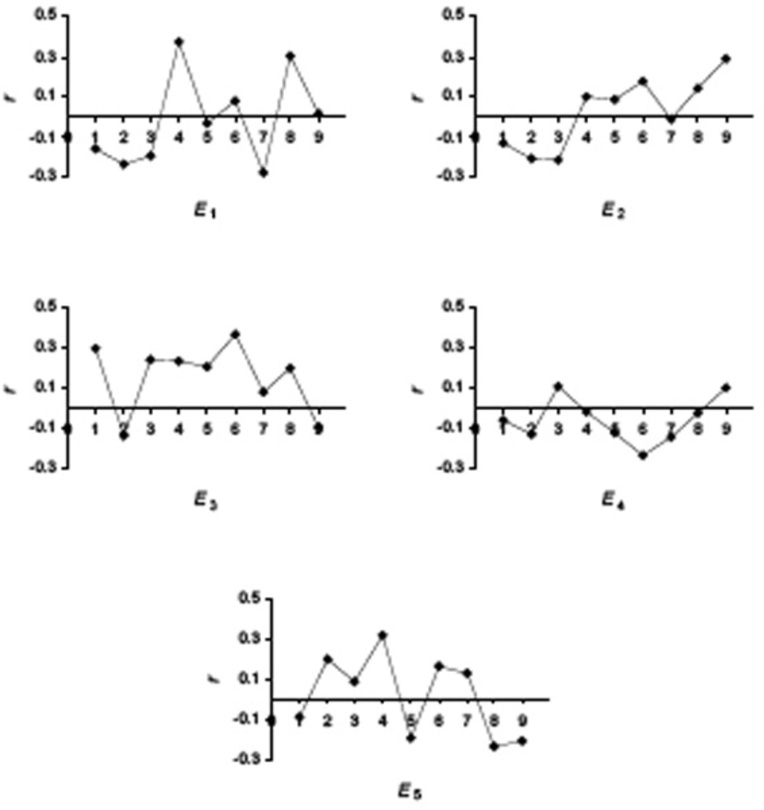
In the first set of experiments we have used linear regression to evaluate the contribution of the quantitative descriptors and individual positions in a nona-peptide to the binding affinity. In Fig. (1) we present the plots of the correlation coefficient r for each E descriptor across the nona-peptides length. From these plots it is apparent that the structure-activity correlation is distributed among the five E descriptors and across the nine positions in the ligands. Each position has a small but significant contribution to the overall binding, with different physical-chemical properties determining the affinity. The most important descriptors for each position will be discussed below, when we present our main QSAR model.
Figure 1.
Correlation coefficients between E1–E5 and pIC50 for each position of the 152 nona-peptides.
Multilinear Regression Model
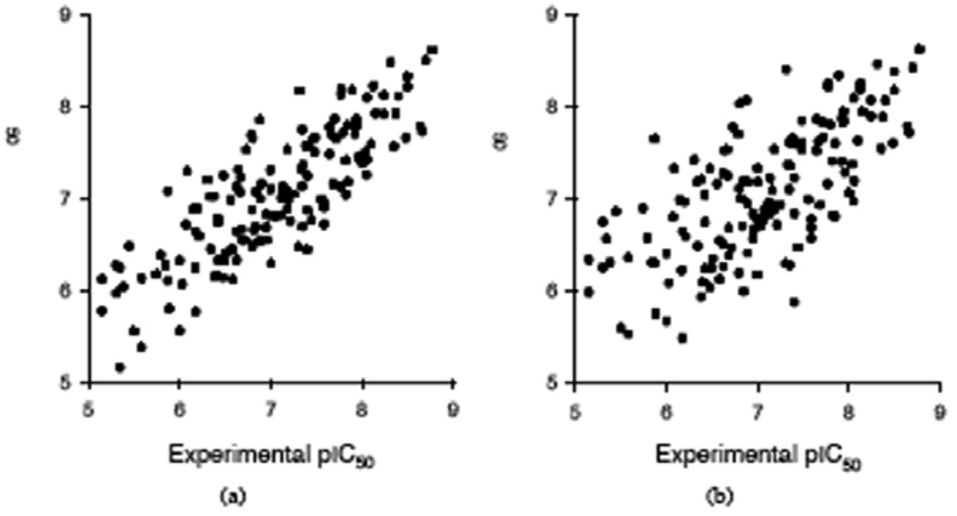
In order to identify a robust MLR model with good prediction statistics, we have pursued two strategies, a position independent and a position dependent strategy, as described in the Methods part. As a reference we used the MLR model with all 45 E1–E5 descriptors (Table 1, model 1), with the following statistics: calibration rcal = 0.866 and scal = 0.510, LOO rpre = 0.685 and spre = 0.625, L10%O rpre = 0.676 and spre = 0.633. In the first position independent strategy (Table 1, models 2–5) we eliminated in each round 9 descriptors that have the lowest absolute correlation coefficient with pIC50. The statistical indices for these for MLR models show that their prediction ability is lower than in the case when all 45 E1–E5 descriptors are used in an MLR model. However, in the second position dependent strategy, where we kept for each position the same number of best descriptors, the predictive capability determined by rpre, increases to a maximum for model 7 with 27 position best descriptors (rcal = 0.825 and scal = 0.534, LOO rpre = 0.709 and spre= 0.605, L10%O rpre= 0.700 and spre= 0.613). This MLR model with 27 E1–E5 has better prediction statistics than the reference model 1 with all 45 E1–E5 descriptors, showing that the eliminated descriptors are not important in modeling the binding affinity. The plot of experimental vs. calculated pIC50, and experimental vs. L10%O predicted pIC50 obtained with the best MLR model (Table 1, model 7) do not show any particular clustering of data or regions with abnormal high prediction errors, as can be seen from Fig. (2).
Table 1.
Calibration and Prediction (Leave-10%-out and Leave-one-out Cross-Validation) Statistics for the MLR QSAR Models. Model 1 was Obtained with all 45 E1–E5 Descriptors, Models 2–5 were Obtained by Eliminating in Each Step 9 Descriptors with the Lowest Correlation Coefficient, and Models 6–9 were Obtained by Eliminating in Each Step and for each Position in a Nona-Peptide the Descriptor with Minimum Correlation Coefficient
| Model | Descriptors | L10%O | LOO | ||||||
|---|---|---|---|---|---|---|---|---|---|
| rcal | scal | rpre | spre | q2 | rpre | spre | q2 | ||
| 1 | all 45 E1–E5 | 0.866 | 0.510 | 0.676 | 0.633 | 0.377 | 0.685 | 0.625 | 0.403 |
| 2 | overall best 36 E1–E5 | 0.839 | 0.534 | 0.665 | 0.641 | 0.396 | 0.664 | 0.642 | 0.385 |
| 3 | overall best 27 E1–E5 | 0.786 | 0.583 | 0.644 | 0.657 | 0.380 | 0.634 | 0.664 | 0.363 |
| 4 | overall best 18 E1–E5 | 0.716 | 0.636 | 0.590 | 0.693 | 0.324 | 0.591 | 0.692 | 0.330 |
| 5 | overall best 9 E1–E5 | 0.659 | 0.664 | 0.599 | 0.687 | 0.355 | 0.595 | 0.690 | 0.350 |
| 6 | position best 36 X | 0.838 | 0.535 | 0.666 | 0.641 | 0.396 | 0.668 | 0.639 | 0.395 |
| 7 | position best 27 X | 0.825 | 0.534 | 0.700 | 0.613 | 0.467 | 0.709 | 0.605 | 0.484 |
| 8 | position best 18 X | 0.717 | 0.636 | 0.596 | 0.689 | 0.335 | 0.591 | 0.693 | 0.328 |
| 9 | position best 9 X | 0.664 | 0.680 | 0.602 | 0.685 | 0.359 | 0.599 | 0.687 | 0.354 |
Figure 2.
MLR calibration and prediction (leave-10%-out cross-validation) results obtained with 27 E1–E5 descriptors (the best three E1–E5 descriptors for each position in a nona-peptide, model 7 in Table 2). (a) Calibrated versus experimental pIC50 values; (b) Predicted versus experimental pIC50 values.
Neural Network Model
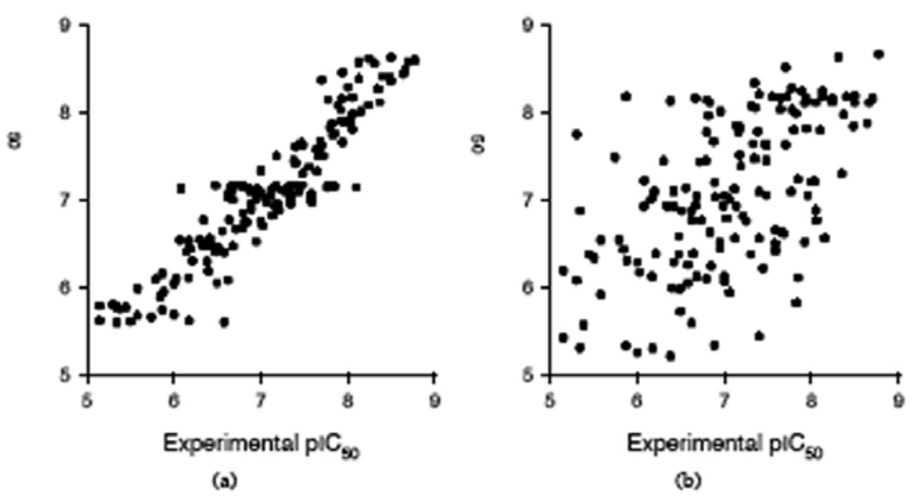
Artificial neural networks form a class of computational models that can explore non-linear relationships between structural descriptors and biological activities, with many applications in drug design. The ANN hidden neurons, that have non-linear transfer functions, can model both the interactions between the input descriptors and the non-linear relationships between input and output variables. We have used ANN models to investigate the non-linear relationships between the E1–E5 descriptors and pIC50, and the cross-interaction between the input descriptors. In preliminary tests we have determined that the optimum number of hidden neurons is two, because a larger number of hidden neurons gives lower prediction statistics. In Table 2 we give the statistics of the ANN models obtained with the same sets of descriptors as the MLR models in Table 1. While the fitting (calibration) statistics are better than those of the corresponding MLR models, the LOO and L10%O prediction tests show a drastic decrease, indicating that the neural networks are over-fitted and give less reliable prediction, compared with those obtained with MLR models. One explanation for the poor prediction performance of the ANN models is the larger number of parameters for optimization. When all 45 E1–E5 descriptors are used as input for the ANN, each nona-peptide is described by a 45 elements vector and 45 input neurons are required. For an ANN with 2 hidden neurons and one output neuron, the number of network connections (parameters to optimize) is (45 +1)×2 + (2 + 1)×1 = 95. In this case, the ratio between the number of experimental data and the number of network connections is too small (ρ = 152/95 = 1.6), compared to the MLR model (ρ = 152/46 = 3.3). Another explanation comes from the nature of the pIC50 data, which are affected by experimental errors. When these errors are large, the non-linear mapping of the ANN model gives erroneous predictions. The plot of experimental vs. calculated pIC50, and experimental vs. L10%O predicted pIC50 obtained with the best ANN model (Table 2, model 7), shown in Fig. (3), indicate that the neural model is over-fitted, compared with the corresponding MLR QSAR.
Table 2.
Calibration and Prediction (Leave-10%-Out and Leave-one-out Cross-Validation) Statistics for the ANN QSAR Models. See Table 1 Caption for a Description of the Experiments
| Model | Descriptors | L10%O | LOO | ||||||
|---|---|---|---|---|---|---|---|---|---|
| rcal | scal | rpre | spre | q2 | rpre | spre | q2 | ||
| 1 | all 45 E1–E5 | 0.968 | 0.215 | 0.489 | 0.749 | −0.045 | 0.540 | 0.722 | 0.014 |
| 2 | overall best 36 E1–E5 | 0.960 | 0.239 | 0.514 | 0.736 | −0.113 | 0.573 | 0.703 | 0.081 |
| 3 | overall best 27 E1–E5 | 0.904 | 0.367 | 0.536 | 0.725 | 0.023 | 0.623 | 0.671 | 0.249 |
| 4 | overall best 18 E1–E5 | 0.838 | 0.469 | 0.426 | 0.776 | 0.013 | 0.435 | 0.773 | −0.043 |
| 5 | overall best 9 E1–E5 | 0.767 | 0.551 | 0.476 | 0.755 | 0.093 | 0.444 | 0.769 | 0.068 |
| 6 | position best 36 X | 0.958 | 0.246 | 0.509 | 0.739 | −0.218 | 0.532 | 0.727 | 0.024 |
| 7 | position best 27 X | 0.929 | 0.317 | 0.561 | 0.710 | 0.112 | 0.446 | 0.768 | −0.091 |
| 8 | position best 18 X | 0.865 | 0.431 | 0.439 | 0.771 | −0.074 | 0.599 | 0.688 | 0.259 |
| 9 | position best 9 X | 0.727 | 0.589 | 0.496 | 0.745 | 0.148 | 0.498 | 0.744 | 0.168 |
Figure 3.
ANN calibration and prediction (leave-10%-out cross-validation) results obtained with 27 E1–E5 descriptors (the best three E1–E5 descriptors for each position in a nona-peptide, model 7 in Table 3). (a) Calibrated versus experimental pIC50 values; (b) Predicted versus experimental pIC50 values.
Partial Least Squares Model
Because the physical-chemical information from the 45 E1–E5 descriptors is correlated, we have used PLS to extract principal components and develop a regression model (Table 3). The first group of experiments tried to estimate the importance of each individual E descriptor (Table 3, models 1–5), while in the second group of experiments the descriptors were added step-wise (Table 3, models 6–9). Based on L10%O cross-validation results, a PLS QSAR with three principal components gives best predictions, with rcal= 0.795, scal= 0.517, rpre= 0.696, and spre= 0.616. Both calibration and prediction statistics are slightly lower than those from the best MLR QSAR (Table 1, model 7), showing that the data compression into PLS components does not improve the predictive power of the QSAR model.
Table 3.
Calibration and Prediction Statistics for PLS QSAR Models Obtained with Various Combinations of E1-E5 Descriptors
| Model | E1 | E2 | E3 | E4 | E5 | rcal | scal | rpre | spre | PC |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 1 | 0 | 0 | 0 | 0 | 0.631 | 0.662 | 0.569 | 0.704 | 3 |
| 2 | 0 | 1 | 0 | 0 | 0 | 0.480 | 0.748 | 0.338 | 0.814 | 6 |
| 3 | 0 | 0 | 1 | 0 | 0 | 0.436 | 0.768 | 0.396 | 0.784 | 1 |
| 4 | 0 | 0 | 0 | 1 | 0 | 0.370 | 0.792 | 0.206 | 0.845 | 1 |
| 5 | 0 | 0 | 0 | 0 | 1 | 0.534 | 0.721 | 0.458 | 0.761 | 2 |
| 6 | 1 | 1 | 0 | 0 | 0 | 0.645 | 0.652 | 0.560 | 0.709 | 2 |
| 7 | 1 | 1 | 1 | 0 | 0 | 0.735 | 0.578 | 0.644 | 0.657 | 3 |
| 8 | 1 | 1 | 1 | 1 | 0 | 0.752 | 0.562 | 0.657 | 0.656 | 3 |
| 9 | 1 | 1 | 1 | 1 | 1 | 0.795 | 0.517 | 0.696 | 0.616 | 3 |
DISCUSSION
After comparing a wide range of QSAR models (MLR, PLS, and ANN) we found that the MLR model with 27 E1–E5 descriptors (the best three E1–E5 descriptors for each position in a nona-peptide, model 7 in Table 1) has the best prediction statistics and the lowest difference between calibration and prediction statistics. This shows that the MLR model is more robust and more predictive than the PLS or ANN QSAR models. It is also of interest to compare the results from our sequence-based MLR prediction of MHC peptide binding affinities with the 3D-QSAR models for the same dataset [3] but which are more difficult to compute and cannot be done automatically. The CoMSIA model has calibration r2 = 0.870 and LOO q2 = 0.542. Our best MLR model, with LOO q2 = 0.484, is slightly lower in predictive power. However, the CoMSIA model requires a 3D modeling of all peptides, a computational intensive step that also requires human assistance, whereas our QSAR model is fast, sequence-based, and can be easily applied to an automatic screening of large protein databases.
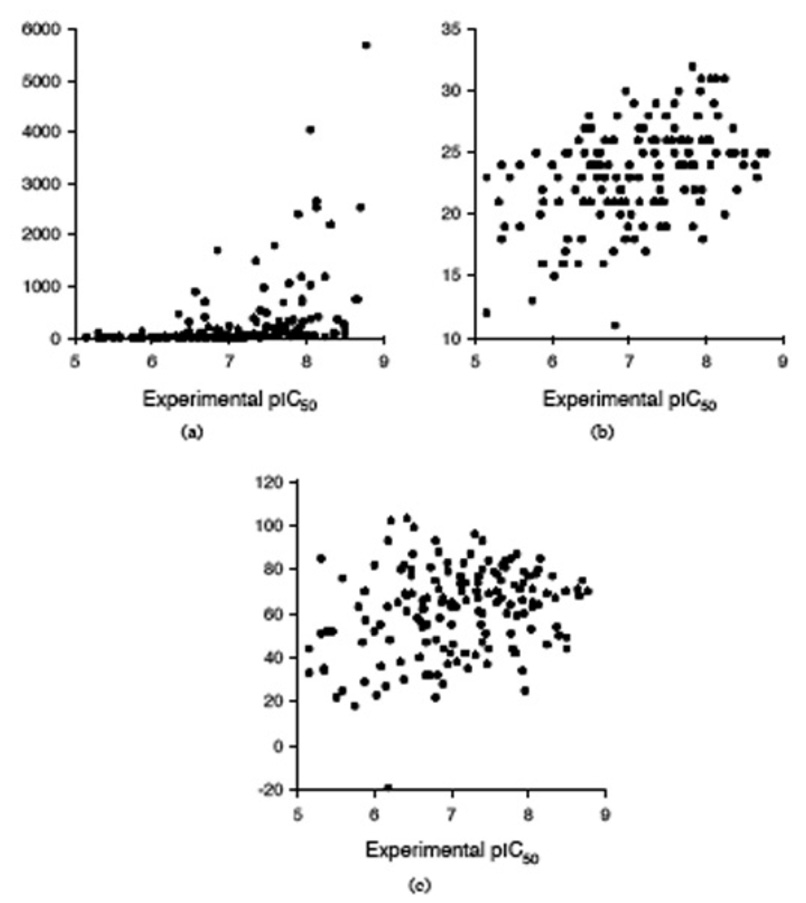
The peptides affinity for the MHC class I HLA-A*0201 may be also predicted with several servers, such as BIMAS (http://bimas.dcrt.nih.gov/molbio/hla_bind/) [58], SYFPEITHI (http://www.syfpeithi.de/) [59], and Rankpep (http://bio.dfci.harvard.edu/Tools/rankpep.html) [47; 60]. All 152 nonapeptides from the dataset were submitted to these three servers, and the predictions are collected in Table 5. Our aim is to compare the predictions obtained with our MLR QSAR model with those computed with BIMAS, SYFPEITHI, and Rankpep, and we use the predictions of the MLR model 7 (Table 1) as a benchmark. The predictions obtained with BIMAS, SYFPEITHI, and Rankpep are compared with the experimental pIC50 in Fig. (4). To compare all these methods, a linear regression was performed between the predicted binding affinity and experimental pIC50 values, and the correlation coefficient is used to evaluate the predictions. We selected this test because the predictions are not on the same scale with the pIC50 values used to develop the QSAR models. The correlation coefficients for all four predictions are as follows: r = 0.823 for MLR QSAR, r = 0.428 for BIMAS, r = 0.445 for SYFPEITHI, and r = 0.258 for Rankpep. It is obvious from this comparison that the MLR QSAR model outperforms other algorithms that predict the peptides affinity for the MHC class I HLA-A*0201.
Figure 4.
Comparison between experimental pIC50 and predictions with three servers: (a) BIMAS, (b) SYFPEITHI, and (c) Rankpep.
The MLR QSAR model based on physical chemical descriptors can also provide guidance in the design of high-affinity peptides, similar as 3D-QSAR models do. The MLR QSAR for the modeling of the MHC binding affinity (Table 1, model 7) is presented in Table 4. An inspection of the partial correlation coefficients r and MLR coefficients for individual E descriptors can offer an insight for the preferences for a particular amino acid type at each of the nine positions in a nona-peptide.
Table 4.
Statistical indices for the MLR Model 7 from Table 1 (E Descriptors, Individual Statistics r, s and F, and MLR Model Coefficients)
| No | r | s | F | MLR Coef | |
|---|---|---|---|---|---|
| 0 | 6.251572 | ||||
| 1 | E1,1 | −0.1618 | 0.847 | 4.0 | −0.007284 |
| 2 | E2,1 | −0.1340 | 0.851 | 2.7 | −0.008691 |
| 3 | E3,1 | 0.2955 | 0.820 | 14.3 | 0.014139 |
| 4 | E1,2 | −0.2395 | 0.833 | 9.1 | −0.038484 |
| 5 | E2,2 | −0.2062 | 0.840 | 6.7 | −0.025657 |
| 6 | E5,2 | 0.1948 | 0.842 | 5.9 | 0.042089 |
| 7 | E1,3 | −0.1935 | 0.842 | 5.8 | −0.028070 |
| 8 | E2,3 | 0.2157 | 0.838 | 7.3 | 0.009419 |
| 9 | E3,3 | 0.2410 | 0.833 | 9.2 | 0.026485 |
| 10 | E1,4 | 0.3719 | 0.797 | 24.1 | 0.025164 |
| 11 | E3,4 | −0.2343 | 0.834 | 8.7 | −0.000594 |
| 12 | E5,4 | 0.3194 | 0.813 | 17.0 | 0.036336 |
| 13 | E3,5 | 0.2034 | 0.840 | 6.5 | 0.022777 |
| 14 | E4,5 | −0.1260 | 0.851 | 2.4 | −0.049741 |
| 15 | E5,5 | −0.1951 | 0.842 | 5.9 | −0.012579 |
| 16 | E2,6 | 0.1753 | 0.845 | 4.8 | 0.013265 |
| 17 | E3,6 | −0.3681 | 0.798 | 23.5 | −0.000668 |
| 18 | E4,6 | −0.2343 | 0.834 | 8.7 | −0.049334 |
| 19 | E1,7 | −0.2816 | 0.824 | 12.9 | −0.012053 |
| 20 | E4,7 | 0.1490 | 0.849 | 3.4 | 0.008458 |
| 21 | E5,7 | 0.1258 | 0.852 | 2.4 | 0.062446 |
| 22 | E1,8 | 0.3035 | 0.818 | 15.2 | 0.014795 |
| 23 | E3,8 | 0.2017 | 0.841 | 6.4 | 0.010158 |
| 24 | E5,8 | 0.2358 | 0.834 | 8.8 | 0.008948 |
| 25 | E2,9 | 0.2920 | 0.821 | 14.0 | 0.080997 |
| 26 | E4,9 | 0.0993 | 0.854 | 1.5 | 0.039239 |
| 27 | E5,9 | −0.2091 | 0.839 | 6.9 | −0.000990 |
Position 1
The negative correlation with E1,1 (mainly correlated with hydrophobicity) favors Ile, Phe and Val for position 1. Similarly, the negative correlation with E2,1 (mainly correlated with side chain size) indicates that Arg, Lys, and Glu are favored. The positive correlation with E3,1 indicates a preference for α-helix breaking residues, such as Tyr, Pro, and Trp.
Position 2
Amino acids with large hydrophobic side chains are preferred for position 2 (E1,2 and E2,2 have negative correlations with pIC50). The positive correlation with E5,2 points out that β-strand breakers (Pro, Glu, Trp) are also favored here.
Position 3
The statistical results suggest that for this position the binding affinity of the peptides is enhanced by hydrophobic (negative correlation with E1,3), small side-chain (positive correlation with E2,3; Gly, Pro, or Ser are favored), or α-helix breaking residues (positive correlation with E3,3).
Position 4
The binding affinity is increased by polar and charged residues (positive correlation with E1,4; Asp, Asn, or Lys are favored), β-strand breakers (positive correlation with E5,4), and by residues with high α-helix forming propensity (negative correlation with E3,4; Ala, Glu, or Leu are favored).
Position 5
The MLR model indicates that the binding affinity is enhanced by the presence in position 5 of α-helix breaking residues (positive correlation with E3,5), most abundant residues (negative correlation with E4,5; Lys, Leu, or Arg are favored), or β-strand forming residues (negative correlation with E5,5; Cys, Arg, or Val are favored).
Position 6
IC50 for a peptide increases when position 6 is occupied by small side-chain residues (positive correlation with E2,6), most abundant residues (negative correlation with E4,5), or α-helix forming residues (negative correlation with E3,6).
Position 7
Binding affinity increases when position 7 corresponds to by hydrophobic residues (negative correlation with E1,7), less abundant residues (positive correlation with E4,7; Cys, Met, or His are favored), or β-strand breakers (positive correlation with E5,7).
Position 8
The statistical correlation suggests that peptide binding increases when position 8 is occupied by polar and charged residues (positive correlation with E1,8), α-helix breaking residues (positive correlation with E3,8), or β-strand breakers (positive correlation with E5,8).
Position 9
The C-terminal residue increases the peptide binding affinity when it corresponds to a small side-chain residue (positive correlation with E2,9), less abundant residue (positive correlation with E4,9), or to β-strand forming residues (negative correlation with E5,9).
CONCLUSIONS
In this paper we presented a successful QSAR application of the amino acids E1–E5 descriptors [1] for the modeling of the binding affinities of 152 nona-peptides for class I MHC HLA-A*0201. After a comparative modeling with multiple linear regression, partial least squares, and artificial neural networks, we found that the MLR model has the highest predictive power. The predictions of the MLR model are as good as those obtained with CoMSIA [3], and much better than the predictions of BIMAS, SYFPEITHI, and Rankpep. The MLR QSAR based on the E1–E5 descriptors has several advantages over the CoMSIA method, because CoMSIA requires a 3D modeling of all peptides, a computational intensive step that also requires human assistance. In contrast, the E1–E5 MLR model is sequence-based, fast, and can be easily applied to an automatic screening of large protein databases.
ACKNOWLEDGEMENTS
This work was supported by National Institutes of Health Grant R01 AI 064913, and a grant from the U.S. Environmental Protection Agency under a STAR Research Assistance Agreement (No. RD 833137). The article has not been formally reviewed by the EPA, and the views expressed in this document are solely those of the authors.
REFERENCES
- 1.Venkatarajan MS, Braun W. J. Mol. Model. 2001;7:445–453. [Google Scholar]
- 2.Lin Z, Wu Y, Zhu B, Ni B, Wang L. J. Comput. Biol. 2004;11:683–694. doi: 10.1089/cmb.2004.11.683. [DOI] [PubMed] [Google Scholar]
- 3.Doytchinova IA, Flower DR. J. Med. Chem. 2001;44:3572–3581. doi: 10.1021/jm010021j. [DOI] [PubMed] [Google Scholar]
- 4.Doytchinova IA, Flower DR. Proteins. 2002;48:505–518. doi: 10.1002/prot.10154. [DOI] [PubMed] [Google Scholar]
- 5.Doytchinova IA, Guan PP, Flower DR. Methods. 2004;34:444–453. doi: 10.1016/j.ymeth.2004.06.007. [DOI] [PubMed] [Google Scholar]
- 6.Guan P, Doytchinova IA, Flower DR. Bioorg. Med. Chem. 2003;11:2307–2311. doi: 10.1016/s0968-0896(03)00109-3. [DOI] [PubMed] [Google Scholar]
- 7.Doytchinova IA, Flower DR. J. Med. Chem. 2006;49:2193–2199. doi: 10.1021/jm050876m. [DOI] [PubMed] [Google Scholar]
- 8.Hattotuwagama CK, Doytchinova IA, Flower DR. J. Chem. Inf. Model. 2005;45:1415–1423. doi: 10.1021/ci049667l. [DOI] [PubMed] [Google Scholar]
- 9.Guan P, Doytchinova IA, Walshe VA, Borrow P, Flower DR. J. Med. Chem. 2005;48:7418–7425. doi: 10.1021/jm0505258. [DOI] [PubMed] [Google Scholar]
- 10.Kawashima S, Kanehisa M. Nucl. Acids Res. 2000;28:374. doi: 10.1093/nar/28.1.374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sandberg M, Eriksson L, Jonsson J, Sjöström M, Wold S. J. Med. Chem. 1998;41:2481–2491. doi: 10.1021/jm9700575. [DOI] [PubMed] [Google Scholar]
- 12.Doytchinova IA, Walshe V, Borrow P, Flower DR. J. Comput.-Aided Mol. Des. 2005;19:203–212. doi: 10.1007/s10822-005-3993-x. [DOI] [PubMed] [Google Scholar]
- 13.Doytchinova IA, Flower DR. Bioinformatics. 2003;19:2263–2270. doi: 10.1093/bioinformatics/btg312. [DOI] [PubMed] [Google Scholar]
- 14.Doytchinova IA, Flower DR. Mol. Immunol. 2006;43:2037–2044. doi: 10.1016/j.molimm.2005.12.013. [DOI] [PubMed] [Google Scholar]
- 15.Doytchinova IA, Flower DR. Immunol. Cell Biol. 2002;80:270–279. doi: 10.1046/j.1440-1711.2002.01076.x. [DOI] [PubMed] [Google Scholar]
- 16.Guan PP, Doytchinova IA, Zygouri C, Flower DR. Nucl. Acids Res. 2003;31:3621–3624. doi: 10.1093/nar/gkg510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hattotuwagama CK, Guan PP, Doytchinova IA, Zygouri C, Flower DR. J. Mol. Graph. Modell. 2004;22:195–207. doi: 10.1016/S1093-3263(03)00160-8. [DOI] [PubMed] [Google Scholar]
- 18.Doytchinova IA, Blythe MJ, Flower DR. J. Proteome Res. 2002;1:263–272. doi: 10.1021/pr015513z. [DOI] [PubMed] [Google Scholar]
- 19.Guan P, Doytchinova IA, Flower DR. Protein Eng. 2003;16:11–18. doi: 10.1093/proeng/gzg005. [DOI] [PubMed] [Google Scholar]
- 20.Hattotuwagama CK, Toseland CP, Guan P, Taylor DJ, Hemsley SL, Doytchinova IA, Flower DR. J. Chem. Inf. Model. 2006;46:1491–1502. doi: 10.1021/ci050380d. [DOI] [PubMed] [Google Scholar]
- 21.Du Q-S, Mezey PG, Chou K-C. J. Comput. Chem. 2005;26:461–470. doi: 10.1002/jcc.20174. [DOI] [PubMed] [Google Scholar]
- 22.Du Q-S, Li D-P, He W-Z, Chou K-C. J. Comput. Chem. 2006;27:685–692. doi: 10.1002/jcc.20369. [DOI] [PubMed] [Google Scholar]
- 23.Du Q-S, Huang R-B, Wei Y-T, Wang C-H, Chou K-C. J. Comput. Chem. 2007;28:000–000. doi: 10.1002/jcc.20732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Du Q-S, Huang R-B, Wei Y-T, Du L-Q, Chou K-C. J. Comput. Chem. 2007;28:000–000. doi: 10.1002/jcc.20732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wan J, Liu W, Xu Q, Ren Y, Flower D, Li T. BMC Bioinformatics. 2006;7:463. doi: 10.1186/1471-2105-7-463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Liu W, Meng XS, Xu Q, Flower D, Li T. BMC Bioinformatics. 2006;7:182. doi: 10.1186/1471-2105-7-182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Davies M, Hattotuwagama C, Moss D, Drew M, Flower D. BMC Struct. Biol. 2006;6:5. doi: 10.1186/1472-6807-6-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Peters B, Sette A. BMC Bioinformatics. 2005;6:132. doi: 10.1186/1471-2105-6-132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bui H-H, Sidney J, Peters B, Sathiamurthy M, Sinichi A, Purton K-A, Mothé BR, Chisari FV, Watkins DI, Sette A. Immunogenetics. 2005;57:304–314. doi: 10.1007/s00251-005-0798-y. [DOI] [PubMed] [Google Scholar]
- 30.Vita R, Vaughan K, Zarebski L, Salimi N, Fleri W, Grey H, Sathiamurthy M, Mokili J, Bui H-H, Bourne P, Ponomarenko J, de Castro R, Chan R, Sidney J, Wilson S, Stewart S, Way S, Peters B, Sette A. BMC Bioinformatics. 2006;7:341. doi: 10.1186/1471-2105-7-341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tenzer S, Peters B, Bulik S, Schoor O, Lemmel C, Schatz MM, Kloetzel P-M, Rammensee H-G, Schild H, Holzhütter H-G. Cell. Mol. Life Sci. 2005;62:1025–1037. doi: 10.1007/s00018-005-4528-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chou K-C. Proteins. 2001;43:246–255. doi: 10.1002/prot.1035. [DOI] [PubMed] [Google Scholar]
- 33.Cai Y-D, Zhou G-P, Chou K-C. Biophys. J. 2003;84:3257–3263. doi: 10.1016/S0006-3495(03)70050-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chou K-C, Cai Y-D. Biochem. Biophys. Res. Commun. 2006;339:1015–1020. doi: 10.1016/j.bbrc.2005.10.196. [DOI] [PubMed] [Google Scholar]
- 35.Chou K-C, Cai Y-D. J. Proteome Res. 2006;5:316–322. doi: 10.1021/pr050331g. [DOI] [PubMed] [Google Scholar]
- 36.Chou K-C, Shen H-B. J. Proteome Res. 2006;5:1888–1897. doi: 10.1021/pr060167c. [DOI] [PubMed] [Google Scholar]
- 37.Brusic V, Rudy G, Honeyman M, Hammer J, Harrison L. 1998;14:121–130. doi: 10.1093/bioinformatics/14.2.121. [DOI] [PubMed] [Google Scholar]
- 38.Milik M, Sauer D, Brunmark AP, Yuan LL, Vitiello A, Jackson MR, Peterson PA, Skolnick J, Glass CA. Nature Biotechnol. 1998;16:753–756. doi: 10.1038/nbt0898-753. [DOI] [PubMed] [Google Scholar]
- 39.Nielsen M, Lundegaard C, Worning P, Lauemoller SL, Lamberth K, Buus S, Brunak S, Lund O. Protein Sci. 2003;12:1007–1017. doi: 10.1110/ps.0239403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Donnes P, Elofsson A. BMC Bioinformatics. 2002;3 doi: 10.1186/1471-2105-3-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhao YD, Pinilla C, Valmori D, Martin R, Simon R. Bioinformatics. 2003;19:1978–1984. doi: 10.1093/bioinformatics/btg255. [DOI] [PubMed] [Google Scholar]
- 42.Cai Y-D, Feng K-Y, Lu W-C, Chou K-C. J. Theor. Biol. 2006;238:172–176. doi: 10.1016/j.jtbi.2005.05.034. [DOI] [PubMed] [Google Scholar]
- 43.Chou K-C, Shen H-B. Biochem. Biophys. Res. Commun. 2006;347:150–157. doi: 10.1016/j.bbrc.2006.06.059. [DOI] [PubMed] [Google Scholar]
- 44.Shen H-B, Chou K-C. Biopolymers. 2007;85:233–240. doi: 10.1002/bip.20640. [DOI] [PubMed] [Google Scholar]
- 45.Shen H-B, Chou K-C. Biochem. Biophys. Res. Commun. 2007;355:1006–1011. doi: 10.1016/j.bbrc.2007.02.071. [DOI] [PubMed] [Google Scholar]
- 46.Lauemoller SL, Holm A, Hilden J, Brunak S, Nissen MH, Stryhn A, Pedersen LO, Buus S. Tissue Antigens. 2001;57:405–414. doi: 10.1034/j.1399-0039.2001.057005405.x. [DOI] [PubMed] [Google Scholar]
- 47.Reche PA, Glutting JP, Reinherz EL. Human Immunol. 2002;63:701–709. doi: 10.1016/s0198-8859(02)00432-9. [DOI] [PubMed] [Google Scholar]
- 48.Kato R, Noguchi H, Honda H, Kobayashi T. Enzyme Microb. Technol. 2003;33:472–481. [Google Scholar]
- 49.Noguchi H, Kato R, Hanai T, Matsubara Y, Honda H, Brusic V, Kobayashi T. J. Biosci. Bioeng. 2002;94:264–270. doi: 10.1263/jbb.94.264. [DOI] [PubMed] [Google Scholar]
- 50.Ivanciuc O, Schein CH, Braun W. 2002;18:1358–1364. doi: 10.1093/bioinformatics/18.10.1358. [DOI] [PubMed] [Google Scholar]
- 51.Ivanciuc O, Mathura V, Midoro-Horiuti T, Braun W, Goldblum RM, Schein CH. J. Agric. Food Chem. 2003;51:4830–4837. doi: 10.1021/jf034218r. [DOI] [PubMed] [Google Scholar]
- 52.Ivanciuc O, Schein CH, Braun W. Nucl. Acids Res. 2003;31:359–362. doi: 10.1093/nar/gkg010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Schein CH, Ivanciuc O, Braun W. J. Agric. Food Chem. 2005;53:8752–8759. doi: 10.1021/jf051148a. [DOI] [PubMed] [Google Scholar]
- 54.Schein CH, Ivanciuc O, Braun W. In: Food Allergy. Maleki SJ, Burks AW, Helm RM, editors. Washington, D.C: ASM Press; 2006. pp. 257–283. [Google Scholar]
- 55.Schein CH, Ivanciuc O, Braun W. Immunol. Allerg. Clin. North Am. 2007;27:1–27. doi: 10.1016/j.iac.2006.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Chou K-C, Zhang CT. Crit. Rev. Biochem. Mol. Biol. 1995;30:275–349. doi: 10.3109/10409239509083488. [DOI] [PubMed] [Google Scholar]
- 57.Chou KC, Shen HB. J. Cell. Biochem. 2007;100:665–678. doi: 10.1002/jcb.21096. [DOI] [PubMed] [Google Scholar]
- 58.Parker KC, Bednarek MA, Coligan JE. J. Immunol. 1994;152:163–175. [PubMed] [Google Scholar]
- 59.Rammensee HG, Bachmann J, Emmerich NPN, Bachor OA, Stevanovic S. Immunogenetics. 1999;50:213–219. doi: 10.1007/s002510050595. [DOI] [PubMed] [Google Scholar]
- 60.Reche PA, Glutting JP, Zhang H, Reinherz EL. Immunogenetics. 2004;56:405–419. doi: 10.1007/s00251-004-0709-7. [DOI] [PubMed] [Google Scholar]