Abstract
Topoisomerase II is an essential enzyme that is required for virtually every process that requires movement of DNA within the nucleus or the opening of the double helix. This enzyme helps to regulate DNA under- and overwinding and removes knots and tangles from the genetic material. In order to carry out its critical physiological functions, topoisomerase II generates transient double-stranded breaks in DNA. Consequently, while necessary for cell survival, the enzyme also has the capacity to fragment the genome. The DNA cleavage/ligation reaction of topoisomerase II is the target for some of the most successful anticancer drugs currently in clinical use. However, this same reaction also is believed to trigger chromosomal translocations that are associated with specific types of leukemia. This article will familiarize the reader with the DNA cleavage/ligation reaction of topoisomerase II and other aspects of its catalytic cycle. In addition, it will discuss the interaction of the enzyme with anticancer drugs and the mechanisms by which these agents increase levels of topoisomerase II-generated DNA strand breaks. Finally, it will describe dietary and environmental agents that enhance DNA cleavage mediated by the enzyme.
INTRODUCTION
A number of enzymes that catalyze essential physiological processes also have the capacity to damage the genome during the course of their normal activities. For example, while the cell requires DNA polymerases to copy the genetic material, these enzymes insert an incorrect base approximately every 107 nt (1). Consequently, in the absence of mismatch repair pathways, human DNA polymerases would generate several hundred mutations every round of cell division. Furthermore, while DNA glycosylases initiate base-excision repair pathways, these enzymes can convert innocuous lesions to abasic sites with far greater mutagenic potential (2). Finally, while cytochrome P450 enzymes play critical roles in detoxification pathways, they sometimes convert inert xenobiotic chemicals to compounds with mutagenic properties (3).
Of all the enzymes required to sustain cellular growth, topoisomerase II is one of the most dangerous (4–8). As discussed below, this enzyme unwinds, unknots and untangles the genetic material by generating transient double-stranded breaks in DNA (8–12). Although the cell cannot survive without topoisomerase II, the strand breaks that the enzyme generates have the potential to trigger cell death pathways or chromosomal translocations (8,13).
This article focuses on the mechanism by which topoisomerase II cleaves the genetic material, the ability to exploit this reaction for the chemotherapeutic treatment of human cancers and the role of this reaction in triggering specific types of leukemia.
DNA TOPOLOGY
The existence of topoisomerases is necessitated by the structure of the double helix. Each human cell contains ∼2 m of DNA that are compacted into a nucleus that is ∼10 μm in diameter (14,15). Because the genetic material is anchored to the chromosome scaffold and the two strands of the double helix are plectonemically coiled, accessing the genome is a complex topological challenge (11,12,16–18).
Topological properties of DNA are those that can only be changed when the double helix is broken (12). Two aspects of DNA topology significantly affect nuclear processes. The first deals with topological relationships between the two strands of the double helix. In all living systems, from bacteria to humans, DNA is globally underwound (i.e. negatively supercoiled) by ∼6% (12,19–21). This is important because duplex DNA is merely the storage form for the genetic information. In order to replicate or express this information, the two strands of DNA must be separated. Since global underwinding of the genome imparts increased single-stranded character to the double helix, negative supercoiling greatly facilitates strand separation (12,16–18).
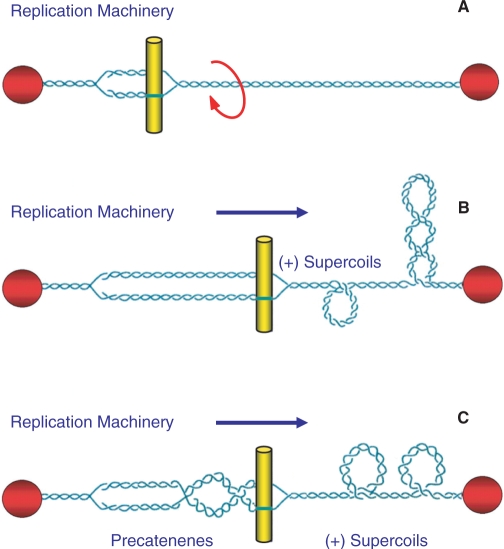
While negative supercoiling promotes many nucleic acid processes, DNA overwinding (i.e. positive supercoiling) inhibits them. The linear movement of tracking enzymes, such as helicases and polymerases, compresses the turns of the double helix into a shorter region (Figure 1) (12,19–21). Consequently, the double helix becomes increasingly overwound ahead of tracking systems. The positive supercoiling that results makes it more difficult to open the two strands of the double helix and ultimately blocks essential nucleic acid processes (10, 12, 16–18).
Figure 1.
Nuclear processes induce changes in DNA topology. DNA replication is used as an example. Although chromosomal DNA is globally underwound in all cells, the movement of DNA tracking systems generates positive supercoils. As shown in (A) chromosomal DNA ends are tethered to membranes or the chromosome scaffold (represented by the red spheres) and are unable to rotate. Therefore, the linear movement of tracking systems (such as the replication machinery represented by the yellow bars) through the immobilized double helix compresses the turns into a shorter segment of the genetic material and induces acute overwinding (i.e. positive supercoiling) ahead of the fork (B). In addition, the compensatory underwinding (i.e. negative supercoiling) behind the replication machinery allows some of the torsional stress that accumulates in the prereplicated DNA to be translated to the newly replicated daughter molecules in the form of precatenanes (C). If these precatenanes are not resolved, they ultimately lead to the formation of intertwined (i.e. tangled) duplex daughter chromosomes. Adapted from ref. 10.
The second aspect of DNA topology deals with relationships between separate DNA segments. Intramolecular knots (formed within the same DNA molecule) are generated during recombination, and intermolecular tangles (formed between daughter DNA molecules) are produced during replication (Figure 1) (8,10,12,17). DNA knots block essential nucleic acid processes because they make it impossible to separate the two strands of the double helix. Moreover, tangled DNA molecules cannot be segregated during mitosis or meiosis (8,10,12,17). Consequently, DNA knots and tangles can be lethal to cells if they are not resolved.
DNA TOPOISOMERASES
The topological state of the genetic material is regulated by enzymes known as topoisomerases (8,10,11,22,23). Topoisomerases are required for the survival of all organisms and alter DNA topology by generating transient breaks in the double helix (8,10,11,22,23). There are two major classes of topoisomerases, type I and type II, that are distinguished by the number of DNA strands that they cleave and the mechanism by which they alter the topological properties of the genetic material (8,10,11,22,23).
Eukaryotic type I topoisomerases are monomeric enzymes that require no high-energy cofactor (11,22,24). Type I enzymes are organized into two subclasses: type IA and type IB. These enzymes alter topology by creating transient single-stranded breaks in the DNA, followed by passage of the opposite intact strand through the break (type IA) or by controlled rotation of the helix around the break (type IB) (11,22,24). Type IA topoisomerases need divalent metal ions for DNA scission and attach covalently to the 5′-terminal phosphate of the DNA (11,22,24). In contrast, type IB enzymes do not require divalent metal ions and covalently link to the 3′-terminal phosphate (11,22,24). As a result of their reaction mechanism, type I topoisomerases can modulate DNA under- and overwinding, but cannot remove knots or tangles from duplex DNA. A number of excellent review articles on type I topoisomerases have appeared recently (22,24,25). Consequently, these enzymes will not be discussed further in this article.
Eukaryotic type II topoisomerases function as homodimers and require divalent metal ions and ATP for complete catalytic activity (5,8,26–28). These enzymes interconvert different topological forms of DNA by a ‘double-stranded DNA passage reaction’ that can be separated into a number of discrete steps (5,8,26–28). Briefly, type II topoisomerases (i) bind two separate segments of DNA, (ii) create a double-stranded break in one of the segments, (iii) translocate the second DNA segment through the cleaved nucleic acid ‘gate’, (iv) rejoin (i.e. ligate) the cleaved DNA, (v) release the translocated segment through a gate in the protein and (vi) close the protein gate and regain the ability to start a new round of catalysis (5,26–34). Because of their double-stranded DNA passage mechanism, type II topoisomerases can modulate DNA supercoiling and also can remove DNA knots and tangles.
TOPOISOMERASE II
Lower eukaryotes and invertebrates encode only a single type II topoisomerase, topoisomerase II (35–38). In contrast, vertebrate species encode two closely related isoforms of the enzyme, topoisomerase IIα and topoisomerase IIβ. These isoforms differ in their protomer molecular masses (170 versus 180 kDa, respectively) and are encoded by separate genes (8,10,22,28,39–46). Topoisomerase IIα and topoisomerase IIβ display a high degree (∼70%) of amino acid sequence identity and similar enzymological characteristics. One notable difference between the two isoforms is that topoisomerase IIα relaxes (i.e. removes) positive superhelical twists ∼10 times faster than it does negative in vitro, while the β isoform is unable to distinguish the geometry of DNA supercoils during DNA relaxation (47).
Topoisomerase IIα and topoisomerase IIβ have distinct patterns of expression and separate cellular functions. Topoisomerase IIα is essential for the survival of proliferating cells, and protein levels rise dramatically during periods of cell growth (48–51). The enzyme is further regulated over the cell cycle, with protein concentrations peaking in G2/M (50,52,53). Topoisomerase IIα is associated with replication forks and remains tightly bound to chromosomes during mitosis (9,51,54–56). Thus, it is believed to be the isoform that functions in growth-related processes, such as DNA replication and chromosome segregation (10,51).
Topoisomerase IIβ is dispensable at the cellular level but appears to be required for proper neural development (57–59). Expression of topoisomerase IIβ is independent of proliferative status and cell cycle, and the enzyme dissociates from chromosomes during mitosis (54,60,61). Topoisomerase IIβ cannot compensate for the loss of topoisomerase IIα in mammalian cells, suggesting that these two isoforms do not play redundant roles in replicative processes (51,60,62,63). Although the physiological functions of topoisomerase IIβ have yet to be defined, recent evidence indicates involvement in the transcription of hormonally or developmentally regulated genes (63,64).
Much of what we understand regarding the mechanism of action of type II enzymes comes from experiments with topoisomerase II from species that express only a single form of the protein. Consequently, eukaryotic type II topoisomerases will be referred to collectively as topoisomerase II, unless the properties being discussed are specific to either the α or β isoform.
TOPOISOMERASE II-MEDIATED DNA CLEAVAGE AND LIGATION
The ability of topoisomerase II to cleave and ligate DNA is central to all of its catalytic functions (5,8,11,27). All topoisomerases utilize active site tyrosyl residues to mediate DNA cleavage and ligation. Since type II enzymes cut both strands of the double helix, each protomer subunit contains one of these residues (Tyr805 and Tyr821 in human topoisomerase IIα and topoisomerase IIβ, respectively).
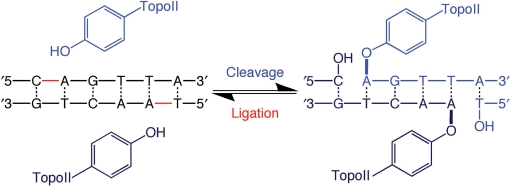
Topoisomerase II initiates DNA cleavage by the nucleophilic attack of the active site tyrosine on the phosphate of the nucleic acid backbone (Figure 2) (11,23,26,27). The resulting transesterification reaction results in the formation of a covalent phosphotyrosyl bond that links the protein to the newly generated 5′-terminus of the DNA chain. It also generates a 3′-hydroxyl moiety on the opposite terminus of the cleaved strand. The scissile bonds on the two strands of the double helix are staggered and are located across the major groove from one another. Thus, topoisomerase II generates cleaved DNA molecules with four-base 5′-single-stranded cohesive ends, each of which is covalently linked to a separate protomer subunit of the enzyme (65–67).
Figure 2.
Double-stranded DNA cleavage mediated by topoisomerase II. Scissile bonds are located four bases apart on opposite strands of the double helix. During cleavage, the active site tyrosine residue of each topoisomerase II protomer subunit becomes covalently linked to the newly generated 5′-terminal phosphate moiety on each strand. This covalent linkage preserves the energy of the sugar-phosphate DNA backbone. The newly generated 3′-hydroxyl group interacts with topoisomerase II in a noncovalent fashion. Ligation represents the reverse of this process and leaves the DNA product chemically unchanged from the initial substrate.
The covalent enzyme–DNA linkage plays two important roles in the topoisomerase II reaction mechanism. First, it conserves the bond energy of the sugar-phosphate DNA backbone. Second, because it does not allow the cleaved DNA chain to dissociate from the enzyme, the protein–DNA linkage maintains the integrity of the genetic material during the cleavage event (11,23,26,27). The covalent topoisomerase II-cleaved DNA reaction intermediate is referred to as the ‘cleavage complex’ and is critical for the pharmacological activities of the enzyme, which are discussed later in this article.
Although topoisomerase II acts globally, it cleaves DNA at preferred sites (68). The consensus sequence for cleavage is weak, and many sites of action do not conform to it (68). Ultimately, the mechanism by which topoisomerase II selects DNA sites is not apparent, and it is nearly impossible to predict de novo whether a given DNA sequence will support scission. Most likely, the specificity of topoisomerase II-mediated cleavage is determined by the local structure, flexibility, or malleability of the DNA that accompanies the sequence, as opposed to a direct recognition of the bases that comprise that sequence (69).
Beyond the nucleophilic attack of the active site tyrosine on the DNA backbone, the details of topoisomerase II-mediated DNA cleavage are not well defined. However, information regarding the roles of specific amino acid residues comes from structural studies on the catalytic core of yeast topoisomerase II generated in the absence of DNA or in a noncovalent complex with its nucleic acid substrate (27,70). For consistency, all amino acid assignments discussed below are for the homologous positions in human topoisomerase IIα.
While not all of the amino acids involved in catalysis have been elucidated, it is believed that the enzyme utilizes a general acid-base mechanism for DNA cleavage (Figure 3). Cleavage is initiated when a general base deprotonates the active site tyrosine hydroxyl, allowing the oxyanion to attack the scissile phosphate. The base has not been identified but is believed to be a conserved histidine residue.
Figure 3.
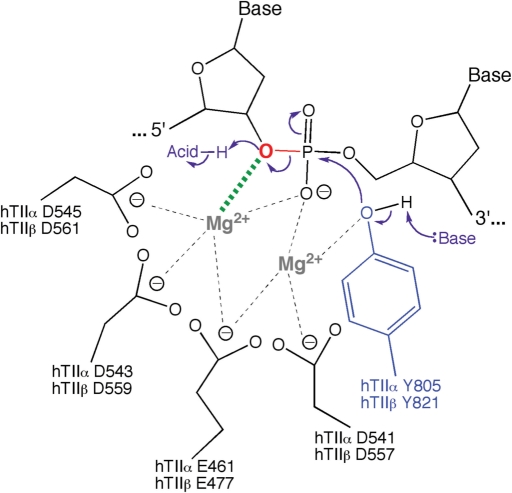
Mechanism of DNA cleavage and ligation mediated by topoisomerase II. The type II enzyme utilizes a two-metal ion mechanism similar to that utilized by primases and polymerases (34,70,71,74–77,155,156). Amino acids that are postulated to interact with the metal ions in the active site of topoisomerase IIα and topoisomerase IIβ are indicated. One of the metal ions (shown at left) makes a critical interaction with the 3′-bridging atom of the scissile phosphate (bond shown in red), which most likely is needed to stabilize the leaving 3′-oxygen (shown in red). A second metal ion (shown at right) is required for DNA scission and may stabilize the DNA transition state and/or help deprotonate the active site tyrosine (Y805 in topoisomerase IIα and Y821 in topoisomerase IIβ). Cleavage is initiated when a general base deprotonates the active site tyrosine hydroxyl, allowing the oxyanion to attack the scissile phosphate. The base has not been identified but is believed to be a conserved histidine residue. Ligation is initiated when a general acid extracts the hydrogen from the 3′-terminal hydroxyl group. The acid may be a water molecule or an unidentified amino acid in the active site of topoisomerase II. Figure adapted from Noble and Maxwell (73).
The DNA cleavage reaction requires a divalent metal ion (5,30,34,71,72). Mg2+ appears to fulfill this function in vivo (5). Recent evidence indicates that human topoisomerase IIα utilizes a ‘two-metal-ion’ mechanism, in which one of the metal ions interacts with the bridging 5′-oxygen of the scissile bond (34). This interaction greatly accelerates rates of enzyme-mediated DNA cleavage and most likely is needed to stabilize the leaving 3′-oxygen. The role of the second metal ion is not known. However, it is believed to make critical contacts with the active site tyrosine and may stabilize the DNA transition state and/or help deprotonate the active site tyrosine (34,73). It has been postulated that the divalent metal ions are coordinated by Glu461, Asp541, Asp543 and Asp545 in human topoisomerase IIα and corresponding residues in the β isoform (Figure 3) (34,70,74,75). A two-metal-ion mechanism for DNA cleavage mediated by the bacterial type II topoisomerase, DNA gyrase, also has been proposed (73).
Topoisomerase II–DNA cleavage complexes normally are short-lived and readily reversible (5,8,76), and the DNA cleavage/ligation equilibrium of the enzyme greatly favors ligation (5,8,11,23,26,27,77,78). Under equilibrium conditions, ∼0.5–1% of topoisomerase II in a DNA scission reaction mixture exists as a cleavage complex (65–67,78,79). Furthermore, when Mg2+ is utilized as the divalent metal ion, ∼1/2–3/4 of the complexes contain double-stranded breaks with the remainder containing single-stranded DNA breaks. The fact that a significant proportion of cleavage complexes contain single-stranded breaks was initially taken as an indication for poor coordination between the two protomer subunits of topoisomerase II (80). However, with hindsight, if the protomers cut the two strands of the double helix in a completely noncoordinated fashion, virtually no double-stranded DNA breaks would be generated (1% × 1% ≈ 0.01% cleavage complexes). These findings suggest that there must be a relatively high degree of coordination between the two protomer active sites of the enzyme, even if they do not act in complete concert with one another (78). To this point, once topoisomerase II cleaves the first strand, it is estimated that the enzyme cuts the second strand ∼20-fold faster (77,78).
Following strand passage, DNA ligation is initiated when a general acid extracts the hydrogen from the 3′-terminal hydroxyl group. The acid may be a water molecule or an unidentified amino acid in the active site of topoisomerase II. The conversion of the terminal hydroxyl moiety to an oxyanion induces a nucleophilic attack on the phosphotyrosyl bond (11,23,26,27). This action represents the reverse of the cleavage event and regenerates an intact DNA chain as well as the active site of topoisomerase II.
It should be noted that the chemical structure of the ligated DNA is identical to that of the original substrate. Only the topological properties of the double helix are altered by the actions of topoisomerase II.
TOPOISOMERASE II AS A CELLULAR TOXIN
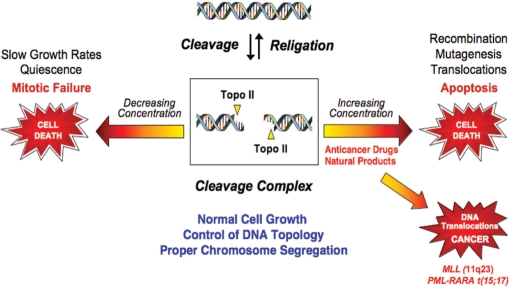
Proliferating cells cannot exist without type II topoisomerases (5,8,10). However, since these enzymes generate obligatory double-stranded DNA breaks as part of their reaction mechanism, they are intrinsically dangerous proteins. Thus, topoisomerase II assumes a Dr. Jekyll/Mr. Hyde character; while essential to cell viability, the enzyme also has the capacity to fragment the genome (Figure 4). Because of this dual persona, levels of cleavage complexes are maintained in a critical balance (5,8). When levels drop below threshold concentrations, daughter chromosomes remain entangled following replication. As a result, chromosomes cannot segregate properly during mitosis and cells die as a result of catastrophic mitotic failure (Figure 4).
Figure 4.
Topoisomerase II is an essential but genotoxic enzyme. The formation of topoisomerase II–DNA cleavage complexes is required for the enzyme to perform its critical cellular functions. If the level of topoisomerase II–DNA cleavage complexes falls too low (left arrow), cells are not able to untangle daughter chromosomes and ultimately die of mitotic failure. If the level of cleavage complexes becomes too high (right arrow) the actions of DNA tracking systems can convert these transient complexes to permanent double-stranded breaks. The resulting DNA breaks, as well as the inhibition of essential DNA processes, initiate recombination/repair pathways and generate chromosome translocations and other DNA aberrations. If the strand breaks overwhelm the cell, they can trigger apoptosis. This is the basis for the actions of several widely prescribed anticancer drugs. If the concentration of topoisomerase-mediated DNA strand breaks is too low to overwhelm the cell, mutations or chromosomal aberrations may be present in surviving populations. In some cases, exposure to topoisomerase II poisons has been associated with the formation of specific types of leukemia that involve the MLL (mixed lineage leukemia) gene at chromosome band 11q23 or the chromosome 15;17 translocation that joins the PML (promyelocytic leukemia) and RARA (retinoic acid receptor α) genes (lower right arrow).
When levels of cleavage complexes rise too high, cells also die, but for different reasons (Figure 4). Accumulated topoisomerase II–DNA cleavage intermediates are converted to permanent strand breaks when replication forks, transcription complexes or DNA tracking enzymes such as helicases attempt to traverse the covalently bound protein ‘roadblock’ in the genetic material (5,8,76,81). The resulting collision disrupts cleavage complexes and ultimately converts transient topoisomerase II-associated DNA breaks to permanent double-stranded breaks that are no longer tethered by proteinaceous bridges (5,8,76,81). The resulting damage and induction of recombination/repair pathways can trigger mutations, chromosomal translocations and other aberrations. When these permanent DNA breaks are present in sufficient numbers, they can overwhelm the cell and initiate death pathways in eukaryotes (4,5,7,8,13,76,81).
TOPOISOMERASE II POISONS
Compounds that impact the catalytic activity of topoisomerase II can be separated into two categories. Chemicals in the first category decrease the overall activity of the enzyme and are known as catalytic inhibitors (5,8,76). Chemicals in the second category increase levels of topoisomerase II–DNA cleavage complexes. These latter compounds are said to ‘poison’ the type II enzyme and convert it to a cellular toxin that initiates the mutagenic and lethal consequences described above (5,8,76). Because of their actions, these compounds are referred to as ‘topoisomerase II poisons’ to distinguish them from inhibitors that do not affect enzyme-mediated DNA cleavage/ligation (5,8,76). Although some topoisomerase II poisons also inhibit overall activity, the ‘gain of function’ induced by these compounds in the cell (i.e. increased levels of cleavage complexes) is a dominant phenotype (8,76).
Topoisomerase II poisons increase the concentration of cleavage complexes by two nonmutually exclusive pathways. Some compounds, such as the anticancer drug etoposide (see below), act by inhibiting the ability of the enzyme to ligate cleaved DNA molecules (5,8,76,78). Other poisons, such as abasic sites and other forms of DNA damage (see below) work primarily by enhancing the forward rate of scission (8,76,78). Because of the manner in which they act, abasic sites poison topoisomerase II without inhibiting overall catalytic activity.
Beyond their effects on DNA scission versus ligation, topoisomerase II poisons (with the exception of DNA lesions) act by two distinct mechanisms. Compounds in the first group are referred to as traditional, noncovalent, interfacial or redox-independent topoisomerase II poisons (8,46,76,82). These chemicals form noncovalent interactions with topoisomerase II at the protein–DNA interface in the vicinity of the active site tyrosine (8,46,84–87). Because these compounds also interact with DNA within the ternary enzyme–DNA–poison complex, they generally alter the DNA cleavage site specificity of the enzyme (88). Finally, their actions against topoisomerase II are not affected by reducing agents, such as dithiothreitol, and these compounds induce similar levels of enzyme-mediated DNA cleavage whether they are added to the binary topoisomerase II–DNA complex or are incubated with the enzyme prior to the addition of nucleic acid substrates (8,82,83,89).
Unlike the traditional poisons, compounds that use the second mechanism require redox activity to facilitate their actions against topoisomerase II. The redox-dependent poisons covalently adduct to the enzyme at amino acid residues outside of the active site (8,76,89–96) and generally enhance DNA cleavage at sites that are intrinsically cut by the enzyme (89,94). Moreover, because these compounds require redox chemistry for activation, their ability to poison topoisomerase II is abrogated by reducing agents (89,91,94,97–99). Finally, compounds within this group enhance DNA cleavage when added to the protein–DNA complex, but display the distinguishing feature of inhibiting topoisomerase II activity when incubated with the enzyme prior to the addition of DNA (89,91,94,97–99).
TOPOISOMERASE II AS A TARGET FOR ANTICANCER DRUGS
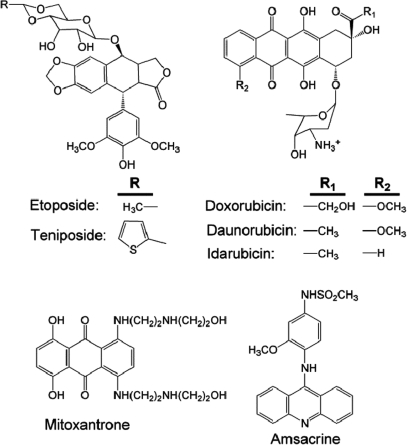
Topoisomerase II poisons represent some of the most important and widely prescribed anticancer drugs currently in clinical use (Figure 5). These drugs encompass a diverse group of natural and synthetic compounds that are commonly used to treat a variety of human malignancies (5,8,76,82,100,101). At the present time, six topoisomerase II-targeted anticancer agents are approved for use in the United States, and additional drugs are prescribed elsewhere in the world (101). These agents all act as traditional topoisomerase II poisons and function primarily by inhibiting enzyme-mediated DNA ligation.
Figure 5.
Topoisomerase II anticancer drugs. Structures of selected topoisomerase II-targeted anticancer drugs are shown.
One of the first topoisomerase II-targeted agents to be discovered was etoposide, which is derived from podophyllotoxin (100,102). This natural product is found in Podophyllum peltatum, more commonly known as the mayapple or mandrake plant (100,102). Podophyllotoxin has been used as a folk remedy for over a 1000 years (100,102). The clinical use of this compound as an antineoplastic agent was prevented by high toxicity, but two synthetic analogs, etoposide and teniposide, displayed increased antineoplastic activity and decreased toxicity (100,102). Etoposide was approved for clinical use against cancer in the mid-1980s and for several years was the most widely prescribed anticancer drug in the world (100,102).
Etoposide and other drugs such as doxorubicin (and its derivatives) are front-line therapy for a variety of systemic cancers and solid tumors, including leukemias, lymphomas, sarcomas and breast, lung and germline cancers (100–102). Mitoxantrone is used to treat breast cancer, and both it and amsacrine are used to treat relapsed acute myeloid leukemia (103,104). Every form of cancer that is considered to be curable by chemotherapy utilizes treatment regimens that include topoisomerase II-targeted drugs (100–102,105). In addition to the use of mitoxantrone in anticancer regimens, it is used as a treatment for autoimmune diseases, such as multiple sclerosis (106).
Although topoisomerase II is the cytotoxic target of the drugs shown in Figure 5, the relative contributions of topoisomerase IIα and topoisomerase IIβ to the chemotherapeutic effects of these agents has yet to be resolved. Some drugs appear to favor one isoform or the other; however, no truly ‘isoform-specific’ agents have been identified. The issue of isoform specificity has potential clinical ramifications. For example, since topoisomerase IIα is not expressed appreciably in quiescent cells, the actions of topoisomerase II-targeted agents against the β isoform in differentiated tissues, such as cardiac cells, most likely are responsible for much of the off-target toxicity of these drugs (107–109). Alternatively, since topoisomerase IIα and topoisomerase IIβ are involved in different cellular processes, it may be that cleavage complexes formed with one or the other isoform are more likely to be converted to permanent DNA strand breaks.
DIETARY TOPOISOMERASE II POISONS
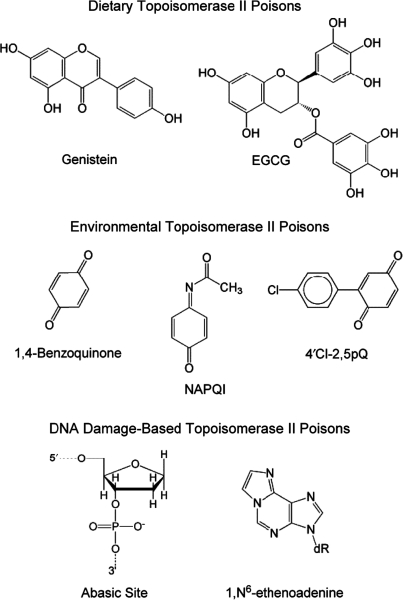
Many foods consumed in the human diet contain naturally occurring topoisomerase II poisons (Figure 6). The most prominent natural products with activity against the mammalian type II enzymes are the bioflavonoids (i.e. phytoestrogens) (110–113). Bioflavonoids represent a broad group of polyphenolic compounds (including flavones, flavonols, isoflavones and catechins) that are components of many fruits, vegetables and plant leaves (114–117). These compounds affect human cells through a variety of pathways; they are strong antioxidants and efficient inhibitors of growth factor receptor tyrosine kinases (114–117). In addition, many bioflavonoids, especially genistein, are potent topoisomerase II poisons (90,110–113,118–122).
Figure 6.
Dietary, environmental and DNA-based topoisomerase II poisons. Abbreviations used are epigallocatechin gallate (EGCG), N-acetyl-p-benzoquinone imine (NAPQI) and 2-(4-chloro-phenyl)-[1,4]benzoquinone (4′Cl-2,5pQ).
Genistein, which is prominent in soy, is believed to be a chemopreventative agent in adults that contributes to the low incidence of breast and colorectal cancers in the Pacific Rim (Figure 6) (116,117). However, as discussed below, there also is evidence associating genistein consumption during pregnancy with the development of infant leukemias (111,123–126). In addition, (–)-epigallocatechin gallate (EGCG), the most abundant and biologically active polyphenol in green tea, is a topoisomerase II poison (Figure 6) (90,116,121). Many of the therapeutic benefits of green tea have been attributed to this compound.
The ring structure of genistein is remarkably similar to that of quinolones (122). These latter compounds, which target the prokaryotic type II topoisomerases DNA gyrase and topoisomerase IV, represent the most active and broad-spectrum antibacterials currently in clinical use (127–129). Like the quinolones and the anticancer drugs discussed above, genistein increases levels of topoisomerase II–DNA cleavage complexes as a traditional topoisomerase II poison (122). In contrast, since many bioflavonoids undergo redox chemistry, several members of the class act as redox-dependent topoisomerase II points. EGCG and some related catechins poison topoisomerase II by this latter mechanism (90,121).
ENVIRONMENTAL TOPOISOMERASE II POISONS
The toxic metabolites of some drugs and industrial chemicals are topoisomerase II poisons (Figure 6) (92,93,95,130,131). In all cases described to date, these chemicals include quinones (aromatic rings that feature ketone groups) as part of their structures (132–135). Quinones commonly are produced in the body as a result of detoxification or metabolism pathways (132–135). These compounds are highly reactive and often damage cells by generating oxidative radicals and by covalently modifying proteins and (to a lesser extent) nucleic acids (132–135).
Although acetaminophen is the most widely utilized analgesic in the world, the second most prevalent cause of toxic drug admissions to emergency departments in the United States is overdosage (both accidental and intentional) of this drug (136). The toxic metabolite of acetaminophen, N-acetyl p-benzoquinone imine (NAPQI), is a potent topoisomerase II poison that produces liver failure (96). Benzene is an industrial solvent that is associated with the development of human leukemias (130,132–135). One of the most prevalent metabolites of benzene, 1,4-benzoquinone, is a highly reactive topoisomerase II poison (92,93,95,130,131). Finally, polychlorinated biphenyls (PCBs), which have multiple industrial uses, have been linked to a variety of human health issues (94). The quinone metabolites of these compounds display activity against human type II topoisomerases. Consistent with their highly active redox chemistry, all quinone-based topoisomerase II poisons act in a redox-dependent manner that involves covalent attachment to the enzyme (91–93,97).
DNA DAMAGE AS TOPOISOMERASE II POISONS
Several forms of nucleic acid damage enhance topoisomerase II-mediated DNA cleavage (Figure 6) (69,78,137–144). The type II enzymes are particularly sensitive to abasic sites, alkylated bases that contain exocyclic rings, and other lesions that distort the double helix.
DNA damage increases cleavage at naturally occurring sites of topoisomerase II action (69,78,137–144). In order to enhance cleavage, lesions must be located within the four-base stagger that separates the two scissile bonds (69,78,137–144). Unlike the traditional and redox-dependent topoisomerase II poisons discussed above, DNA damage has no obvious effect on rates of topoisomerase II-mediated ligation and appears to act primarily by enhancing the forward rate of scission (69,78,137–144).
The physiological benefits of DNA lesions as topoisomerase II poisons, if any, are unclear. However, human topoisomerase IIα and topoisomerase IIβ both appear to play roles in fragmenting genomic DNA and releasing chromosomal loops during apoptosis (145,146). It has been suggested that the apoptotic activities of topoisomerase II are enhanced (or perhaps triggered) by DNA lesions that are generated following the release of oxidative radicals from permeable mitochondria in apoptotic cells (145,146).
TOPOISOMERASE II AND LEUKEMIA
Despite the importance of topoisomerase II in cancer chemotherapy, evidence suggests that DNA cleavage mediated by the enzyme can trigger chromosomal translocations that lead to specific types of leukemia (Figure 4) (5–8,76,147,148). To this point, 2–3% of patients who receive regimens that include etoposide or other topoisomerase II-targeted drugs eventually develop acute myeloid leukemias (AMLs). Most of these leukemias are accompanied by translocations with breakpoints in the MLL (mixed lineage leukemia) gene at chromosomal band 11q23 (6,7,76). The MLL protein is a histone methyltransferase that regulates the Hox genes, which control proliferation in hematopoietic cells (6,7,76). Several breakpoints in MLL have been identified and are located in close proximity to topoisomerase II–DNA cleavage sites (6,7,76).
Recently, a link between topoisomerase II-targeted drugs and the development of acute promyelocytic leukemias (APLs) has been observed. Patients with these leukemias display translocations between the PML (promyelocytic leukemia) gene on chromosome 15 and the RARA (retinoic acid receptor α) gene on chromosome 17 (149,150).
In addition to treatment-related leukmias, ∼80% of infants with AML or acute lymphoblastic leukemia (ALL) display translocations that involve the MLL gene (6,7,111,123–125,147,148,151). The chromosomal translocations associated with these cancers have been observed in utero, indicating that infant leukemias are initiated during pregnancy. Epidemiological studies indicate that the risk of developing these infant leukemias is increased >3-fold by the maternal consumption of foods that are high in naturally occurring topoisomerase II poisons such as genistein or other bioflavonoids (111,123–126).
The ability of topoisomerase II poisons to ‘cause’ rather than ‘cure’ cancer may be related to cellular levels of topoisomerase II-mediated DNA cleavage complexes. If the concentration of enzyme-associated DNA breaks is sufficient, DNA recombination/repair pathways can be overwhelmed and cells will die (5,8,76). However, if the levels of breaks are not adequate to induce death, pathways that promote cell survival can lead to the formation of stable chromosomal translocations that ultimately lead to cancerous growth (Figure 4) (5,8,76).
Finally, the specific contributions of topoisomerase IIα and topoisomerase IIβ to cancer therapy versus leukemogenesis are unclear. However, evidence suggests that (with at least some drugs) topoisomerase IIα plays a more important role in cytotoxicity (107,152), while topoisomerase IIβ may play a greater role in triggering drug-induced cancers (107). Although no isoform-specific topoisomerase II-targeted drugs are available at the present time, it may be possible to preferentially target topoisomerase IIα by scheduling. In this regard, topoisomerase IIα–DNA cleavage complexes induced by etoposide persist approximately three to four times longer than those formed with topoisomerase IIβ (153,154). Therefore, it has been suggested that the use of pulsed chemotherapeutic regimens, in which patients receive cycles of drug treatment followed by recovery, may maintain higher levels of topoisomerase IIα as compared to topoisomerase IIβ cleavage complexes over the course of therapy (154).
FUNDING
The National Institutes of Health (GM33944 and GM53960); National Institutes of Health (T32 CA09592 to J.E.D.). Funding for open access charge: National Institutes of Health (GM33944 and GM53960).
Conflict of interest statement. None declared.
ACKNOWLEDGEMENTS
We are grateful to Amanda C. Gentry and Adam C. Ketron for critical reading of the manuscript.
REFERENCES
- 1.Loeb LA, Monnat R.J., Jr. DNA polymerases and human disease. Nat. Rev. Genet. 2008;9:594–604. doi: 10.1038/nrg2345. [DOI] [PubMed] [Google Scholar]
- 2.Hedge ML, Hazra TK, Mitra S. Early steps in the DNA base excision/single-strand interruption repair pathway in mammalian cells. Cell Res. 2008;18:27–47. doi: 10.1038/cr.2008.8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Guengerich FP. Cytochrome P450s and other enzymes in drug metabolism and toxicity. AAPS J. 2006;8:E101–E111. doi: 10.1208/aapsj080112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Baguley BC, Ferguson LR. Mutagenic properties of topoisomerase-targeted drugs. Biochim. Biophys. Acta. 1998;1400:213–222. doi: 10.1016/s0167-4781(98)00137-7. [DOI] [PubMed] [Google Scholar]
- 5.Fortune JM, Osheroff N. Topoisomerase II as a target for anticancer drugs: when enzymes stop being nice. Prog. Nucleic Acid Res. Mol. Biol. 2000;64:221–253. doi: 10.1016/s0079-6603(00)64006-0. [DOI] [PubMed] [Google Scholar]
- 6.Felix CA. Leukemias related to treatment with DNA topoisomerase II inhibitors. Med. Pediatr. Oncol. 2001;36:525–535. doi: 10.1002/mpo.1125. [DOI] [PubMed] [Google Scholar]
- 7.Felix CA, Kolaris CP, Osheroff N. Topoisomerase II and the etiology of chromosomal translocations. DNA Repair (Amst.) 2006;5:1093–1108. doi: 10.1016/j.dnarep.2006.05.031. [DOI] [PubMed] [Google Scholar]
- 8.McClendon AK, Osheroff N. DNA topoisomerase II, genotoxicity, and cancer. Mutat. Res. 2007;623:83–97. doi: 10.1016/j.mrfmmm.2007.06.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Nitiss JL. Investigating the biological functions of DNA topoisomerases in eukaryotic cells. Biochim. Biophys. Acta. 1998;1400:63–81. doi: 10.1016/s0167-4781(98)00128-6. [DOI] [PubMed] [Google Scholar]
- 10.Wang JC. Cellular roles of DNA topoisomerases: a molecular perspective. Nat. Rev. Mol. Cell Biol. 2002;3:430–440. doi: 10.1038/nrm831. [DOI] [PubMed] [Google Scholar]
- 11.Wang JC. DNA topoisomerases. Annu. Rev. Biochem. 1996;65:635–692. doi: 10.1146/annurev.bi.65.070196.003223. [DOI] [PubMed] [Google Scholar]
- 12.Bates AD, Maxwell A. DNA Topology. New York: Oxford University Press; 2005. [Google Scholar]
- 13.Kaufmann SH. Cell death induced by topoisomerase-targeted drugs: more questions than answers. Biochim. Biophys. Acta. 1998;1400:195–211. doi: 10.1016/s0167-4781(98)00136-5. [DOI] [PubMed] [Google Scholar]
- 14.Kornberg A, Baker TA. DNA Replication. 2nd. New York: W. H. Freeman; 1992. [Google Scholar]
- 15.Voet D, Voet JG, Pratt CW. Fundamentals of Biochemistry. 2nd. Hoboken, N.J: John Wiley & Sons; 2006. [Google Scholar]
- 16.Espeli O, Marians KJ. Untangling intracellular DNA topology. Mol. Microbiol. 2004;52:925–931. doi: 10.1111/j.1365-2958.2004.04047.x. [DOI] [PubMed] [Google Scholar]
- 17.Falaschi A, Abdurashidova G, Sandoval O, Radulescu S, Biamonti G, Riva S. Molecular and structural transactions at human DNA replication origins. Cell Cycle. 2007;6:1705–1712. doi: 10.4161/cc.6.14.4495. [DOI] [PubMed] [Google Scholar]
- 18.Travers A, Muskhelishvili G. A common topology for bacterial and eukaryotic transcription initiation? EMBO Rep. 2007;8:147–151. doi: 10.1038/sj.embor.7400898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bauer WR, Crick FH, White JH. Supercoiled DNA. Sci. Am. 1980;243:100–113. [PubMed] [Google Scholar]
- 20.Vologodskii AV, Cozzarelli NR. Conformational and thermodynamic properties of supercoiled DNA. Annu. Rev. Biophys. Biomol. Struct. 1994;23:609–643. doi: 10.1146/annurev.bb.23.060194.003141. [DOI] [PubMed] [Google Scholar]
- 21.White JH, Cozzarelli NR. A simple topological method for describing stereoisomers of DNA catenanes and knots. Proc. Natl Acad. Sci. USA. 1984;81:3322–3326. doi: 10.1073/pnas.81.11.3322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Champoux JJ. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 2001;70:369–413. doi: 10.1146/annurev.biochem.70.1.369. [DOI] [PubMed] [Google Scholar]
- 23.Schoeffler AJ, Berger JM. Recent advances in understanding structure-function relationships in the type II topoisomerase mechanism. Biochem. Soc. Trans. 2005;33:1465–1470. doi: 10.1042/BST0331465. [DOI] [PubMed] [Google Scholar]
- 24.Leppard JB, Champoux JJ. Human DNA topoisomerase I: relaxation, roles, and damage control. Chromosoma. 2005;114:75–85. doi: 10.1007/s00412-005-0345-5. [DOI] [PubMed] [Google Scholar]
- 25.Pommier Y. Topoisomerase I inhibitors: camptothecins and beyond. Nat. Rev. Cancer. 2006;6:789–802. doi: 10.1038/nrc1977. [DOI] [PubMed] [Google Scholar]
- 26.Wang JC. Moving one DNA double helix through another by a type II DNA topoisomerase: the story of a simple molecular machine. Quart. Rev. Biophys. 1998;31:107–144. doi: 10.1017/s0033583598003424. [DOI] [PubMed] [Google Scholar]
- 27.Berger JM, Gamblin SJ, Harrison SC, Wang JC. Structure and mechanism of DNA topoisomerase II. Nature. 1996;379:225–232. doi: 10.1038/379225a0. [DOI] [PubMed] [Google Scholar]
- 28.Velez-Cruz R, Osheroff N. Encyclopedia of Biological Chemistry. San Diego, CA: Elsevier Inc.; 2004. pp. 806–811. [Google Scholar]
- 29.Osheroff N. Eukaryotic topoisomerase II. Characterization of enzyme turnover. J. Biol. Chem. 1986;261:9944–9950. [PubMed] [Google Scholar]
- 30.Osheroff N. Role of the divalent cation in topoisomerase II mediated reactions. Biochemistry. 1987;26:6402–6406. doi: 10.1021/bi00394a015. [DOI] [PubMed] [Google Scholar]
- 31.Lindsley JE, Wang JC. On the coupling between ATP usage and DNA transport by yeast DNA topoisomerase II. J. Biol. Chem. 1993;268:8096–8104. [PubMed] [Google Scholar]
- 32.Roca J, Wang JC. The capture of a DNA double helix by an ATP-dependent protein clamp: a key step in DNA transport by type II DNA topoisomerases. Cell. 1992;71:833–840. doi: 10.1016/0092-8674(92)90558-t. [DOI] [PubMed] [Google Scholar]
- 33.Wilstermann AM, Osheroff N. Positioning the 3′-DNA terminus for topoisomerase II-mediated religation. J. Biol. Chem. 2001;276:17727–17731. doi: 10.1074/jbc.M100197200. [DOI] [PubMed] [Google Scholar]
- 34.Deweese JE, Burgin AB, Osheroff N. Human topoisomerase IIα uses a two-metal-ion mechanism for DNA cleavage. Nucleic Acids Res. 2008;36:4883–4893. doi: 10.1093/nar/gkn466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Goto T, Wang JC. Yeast DNA topoisomerase II. An ATP-dependent type II topoisomerase that catalyzes the catenation, decatenation, unknotting, and relaxation of double-stranded DNA rings. J. Biol. Chem. 1982;257:5866–5872. [PubMed] [Google Scholar]
- 36.Goto T, Laipis P, Wang JC. The purification and characterization of DNA topoisomerases I and II of the yeast Saccharomyces cerevisiae. J. Biol. Chem. 1984;259:10422–10429. [PubMed] [Google Scholar]
- 37.Goto T, Wang JC. Yeast DNA topoisomerase II is encoded by a single-copy, essential gene. Cell. 1984;36:1073–1080. doi: 10.1016/0092-8674(84)90057-6. [DOI] [PubMed] [Google Scholar]
- 38.Nolan JM, Lee MP, Wyckoff E, Hsieh TS. Isolation and characterization of the gene encoding Drosophila DNA topoisomerase II. Proc. Natl Acad. Sci. USA. 1986;83:3664–3668. doi: 10.1073/pnas.83.11.3664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Drake FH, Zimmerman JP, McCabe FL, Bartus HF, Per SR, Sullivan DM, Ross WE, Mattern MR, Johnson RK, Crooke ST. Purification of topoisomerase II from amsacrine-resistant P388 leukemia cells. Evidence for two forms of the enzyme. J. Biol. Chem. 1987;262:16739–16747. [PubMed] [Google Scholar]
- 40.Tsai-Pflugfelder M, Liu LF, Liu AA, Tewey KM, Whang-Peng J, Knutsen T, Huebner K, Croce CM, Wang JC. Cloning and sequencing of cDNA encoding human DNA topoisomerase II and localization of the gene to chromosome region 17q21-22. Proc. Natl Acad. Sci. USA. 1988;85:7177–7181. doi: 10.1073/pnas.85.19.7177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Drake FH, Hofmann GA, Bartus HF, Mattern MR, Crooke ST, Mirabelli CK. Biochemical and pharmacological properties of p170 and p180 forms of topoisomerase II. Biochemistry. 1989;28:8154–8160. doi: 10.1021/bi00446a029. [DOI] [PubMed] [Google Scholar]
- 42.Chung TD, Drake FH, Tan KB, Per SR, Crooke ST, Mirabelli CK. Characterization and immunological identification of cDNA clones encoding two human DNA topoisomerase II isozymes. Proc. Natl Acad. Sci. USA. 1989;86:9431–9435. doi: 10.1073/pnas.86.23.9431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Austin CA, Fisher LM. Isolation and characterization of a human cDNA clone encoding a novel DNA topoisomerase II homologue from HeLa cells. FEBS Lett. 1990;266:115–117. doi: 10.1016/0014-5793(90)81520-x. [DOI] [PubMed] [Google Scholar]
- 44.Jenkins JR, Ayton P, Jones T, Davies SL, Simmons DL, Harris AL, Sheer D, Hickson ID. Isolation of cDNA clones encoding the beta isozyme of human DNA topoisomerase II and localisation of the gene to chromosome 3p24. Nucleic Acids Res. 1992;20:5587–5592. doi: 10.1093/nar/20.21.5587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tan KB, Dorman TE, Falls KM, Chung TD, Mirabelli CK, Crooke ST, Mao J. Topoisomerase II alpha and topoisomerase II beta genes: characterization and mapping to human chromosomes 17 and 3, respectively. Cancer Res. 1992;52:231–234. [PubMed] [Google Scholar]
- 46.Wilstermann AM, Osheroff N. Stabilization of eukaryotic topoisomerase II-DNA cleavage complexes. Curr. Top. Med. Chem. 2003;3:321–338. doi: 10.2174/1568026033452519. [DOI] [PubMed] [Google Scholar]
- 47.McClendon AK, Rodriguez AC, Osheroff N. Human topoisomerase IIα rapidly relaxes positively supercoiled DNA: implications for enzyme action ahead of replication forks. J. Biol. Chem. 2005;280:39337–39345. doi: 10.1074/jbc.M503320200. [DOI] [PubMed] [Google Scholar]
- 48.Heck MM, Earnshaw WC. Topoisomerase II: A specific marker for cell proliferation. J. Cell Biol. 1986;103:2569–2581. doi: 10.1083/jcb.103.6.2569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Hsiang YH, Wu HY, Liu LF. Proliferation-dependent regulation of DNA topoisomerase II in cultured human cells. Cancer Res. 1988;48:3230–3235. [PubMed] [Google Scholar]
- 50.Woessner RD, Mattern MR, Mirabelli CK, Johnson RK, Drake FH. Proliferation- and cell cycle-dependent differences in expression of the 170 kilodalton and 180 kilodalton forms of topoisomerase II in NIH-3T3 cells. Cell Growth Differ. 1991;2:209–214. [PubMed] [Google Scholar]
- 51.Grue P, Grasser A, Sehested M, Jensen PB, Uhse A, Straub T, Ness W, Boege F. Essential mitotic functions of DNA topoisomerase IIα are not adopted by topoisomerase IIβ in human H69 cells. J. Biol. Chem. 1998;273:33660–33666. doi: 10.1074/jbc.273.50.33660. [DOI] [PubMed] [Google Scholar]
- 52.Heck MM, Hittelman WN, Earnshaw WC. Differential expression of DNA topoisomerases I and II during the eukaryotic cell cycle. Proc. Natl Acad. Sci. USA. 1988;85:1086–1090. doi: 10.1073/pnas.85.4.1086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kimura K, Saijo M, Ui M, Enomoto T. Growth state- and cell cycle-dependent fluctuation in the expression of two forms of DNA topoisomerase II and possible specific modification of the higher molecular weight form in the M phase. J. Biol. Chem. 1994;269:1173–1176. [PubMed] [Google Scholar]
- 54.Linka RM, Porter AC, Volkov A, Mielke C, Boege F, Christensen MO. C-terminal regions of topoisomerase IIα and IIβ determine isoform-specific functioning of the enzymes in vivo. Nucleic Acids Res. 2007;35:3810–3822. doi: 10.1093/nar/gkm102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Bauman ME, Holden JA, Brown KA, Harker WG, Perkins SL. Differential immunohistochemical staining for DNA topoisomerase IIα and β in human tissues and for DNA topoisomerase IIβ in non-Hodgkin's lymphomas. Mod. Pathol. 1997;10:168–175. [PubMed] [Google Scholar]
- 56.Christensen MO, Larsen MK, Barthelmes HU, Hock R, Andersen CL, Kjeldsen E, Knudsen BR, Westergaard O, Boege F, Mielke C. Dynamics of human DNA topoisomerases IIα and IIβ in living cells. J. Cell Biol. 2002;157:31–44. doi: 10.1083/jcb.200112023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Chen M, Beck WT. DNA topoisomerase II expression, stability, and phosphorylation in two VM-26-resistant human leukemic CEM sublines. Oncol. Res. 1995;7:103–111. [PubMed] [Google Scholar]
- 58.Dereuddre S, Delaporte C, Jacquemin-Sablon A. Role of topoisomerase II beta in the resistance of 9-OH-ellipticine-resistant Chinese hamster fibroblasts to topoisomerase II inhibitors. Cancer Res. 1997;57:4301–4308. [PubMed] [Google Scholar]
- 59.Yang X, Li W, Prescott ED, Burden SJ, Wang JC. DNA topoisomerase IIβ and neural development. Science. 2000;287:131–134. doi: 10.1126/science.287.5450.131. [DOI] [PubMed] [Google Scholar]
- 60.Austin CA, Marsh KL. Eukaryotic DNA topoisomerase IIβ. BioEssays. 1998;20:215–226. doi: 10.1002/(SICI)1521-1878(199803)20:3<215::AID-BIES5>3.0.CO;2-Q. [DOI] [PubMed] [Google Scholar]
- 61.Isaacs RJ, Davies SL, Sandri MI, Redwood C, Wells NJ, Hickson ID. Physiological regulation of eukaryotic topoisomerase II. Biochim. Biophys. Acta. 1998;1400:121–137. doi: 10.1016/s0167-4781(98)00131-6. [DOI] [PubMed] [Google Scholar]
- 62.Sakaguchi A, Kikuchi A. Functional compatibility between isoform alpha and beta of type II DNA topoisomerase. J. Cell Sci. 2004;117:1047–1054. doi: 10.1242/jcs.00977. [DOI] [PubMed] [Google Scholar]
- 63.Ju BG, Lunyak VV, Perissi V, Garcia-Bassets I, Rose DW, Glass CK, Rosenfeld MG. A topoisomerase IIβ-mediated dsDNA break required for regulated transcription. Science. 2006;312:1798–1802. doi: 10.1126/science.1127196. [DOI] [PubMed] [Google Scholar]
- 64.Haince JF, Rouleau M, Poirier GG. Transcription. Gene expression needs a break to unwind before carrying on. Science. 2006;312:1752–1753. doi: 10.1126/science.1129808. [DOI] [PubMed] [Google Scholar]
- 65.Zechiedrich EL, Christiansen K, Andersen AH, Westergaard O, Osheroff N. Double-stranded DNA cleavage/religation reaction of eukaryotic topoisomerase II: evidence for a nicked DNA intermediate. Biochemistry. 1989;28:6229–6236. doi: 10.1021/bi00441a014. [DOI] [PubMed] [Google Scholar]
- 66.Liu LF, Rowe TC, Yang L, Tewey KM, Chen GL. Cleavage of DNA by mammalian DNA topoisomerase II. J. Biol. Chem. 1983;258:15365–15370. [PubMed] [Google Scholar]
- 67.Sander M, Hsieh T. Double strand DNA cleavage by type II DNA topoisomerase from Drosophila melanogaster. J. Biol. Chem. 1983;258:8421–8428. [PubMed] [Google Scholar]
- 68.Capranico G, Binaschi M. DNA sequence selectivity of topoisomerases and topoisomerase poisons. Biochim. Biophys. Acta. 1998;1400:185–194. doi: 10.1016/s0167-4781(98)00135-3. [DOI] [PubMed] [Google Scholar]
- 69.Velez-Cruz R, Riggins JN, Daniels JS, Cai H, Guengerich FP, Marnett LJ, Osheroff N. Exocyclic DNA lesions stimulate DNA cleavage mediated by human topoisomerase IIα in vitro and in cultured cells. Biochemistry. 2005;44:3972–3981. doi: 10.1021/bi0478289. [DOI] [PubMed] [Google Scholar]
- 70.Dong KC, Berger JM. Structural basis for gate-DNA recognition and bending by type IIA topoisomerases. Nature. 2007;450:1201–1205. doi: 10.1038/nature06396. [DOI] [PubMed] [Google Scholar]
- 71.Aravind L, Leipe DD, Koonin EV. Toprim–a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 1998;26:4205–4213. doi: 10.1093/nar/26.18.4205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Berger JM, Fass D, Wang JC, Harrison SC. Structural similarities between topoisomerases that cleave one or both DNA strands. Proc. Natl Acad. Sci. USA. 1998;95:7876–7881. doi: 10.1073/pnas.95.14.7876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Noble CG, Maxwell A. The role of GyrB in the DNA cleavage-religation reaction of DNA gyrase: a proposed two metal-ion mechanism. J. Mol. Biol. 2002;318:361–371. doi: 10.1016/S0022-2836(02)00049-9. [DOI] [PubMed] [Google Scholar]
- 74.West KL, Meczes EL, Thorn R, Turnbull RM, Marshall R, Austin CA. Mutagenesis of E477 or K505 in the B' domain of human topoisomerase II beta increases the requirement for magnesium ions during strand passage. Biochemistry. 2000;39:1223–1233. doi: 10.1021/bi991328b. [DOI] [PubMed] [Google Scholar]
- 75.Leontiou C, Lakey JH, Lightowlers R, Turnbull RM, Austin CA. Mutation P732L in human DNA topoisomerase IIbeta abolishes DNA cleavage in the presence of calcium and confers drug resistance. Mol. Pharmacol. 2006;69:130–139. doi: 10.1124/mol.105.015933. [DOI] [PubMed] [Google Scholar]
- 76.Bender RP, Osheroff N. Dai W. Checkpoint Responses in Cancer Therapy. Totowa, NJ: Humana Press; 2008. DNA topoisomerases as targets for the chemotherapeutic treatment of cancer. In; pp. 57–91. [Google Scholar]
- 77.Mueller-Planitz F, Herschlag D. Coupling between ATP binding and DNA cleavage by DNA topoisomerase II: A unifying kinetic and structural mechanism. J. Biol. Chem. 2008;283:17463–17476. doi: 10.1074/jbc.M710014200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Deweese JE, Burgin AB, Osheroff N. Using 3′-bridging phosphorothiolates to isolate the forward DNA cleavage reaction of human topoisomerase IIα. Biochemistry. 2008;47:4129–4140. doi: 10.1021/bi702194x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Andersen AH, Christiansen K, Zechiedrich EL, Jensen PS, Osheroff N, Westergaard O. Strand specificity of the topoisomerase II mediated double-stranded DNA cleavage reaction. Biochemistry. 1989;28:6237–6244. doi: 10.1021/bi00441a015. [DOI] [PubMed] [Google Scholar]
- 80.Bromberg KD, Burgin AB, Osheroff N. A two-drug model for etoposide action against human topoisomerase IIα. J. Biol. Chem. 2003;278:7406–7412. doi: 10.1074/jbc.M212056200. [DOI] [PubMed] [Google Scholar]
- 81.D'Arpa P, Beardmore C, Liu LF. Involvement of nucleic acid synthesis in cell killing mechanisms of topoisomerase poisons. Cancer Res. 1990;50:6919–6924. [PubMed] [Google Scholar]
- 82.Pommier Y, Marchand C. Interfacial inhibitors of protein-nucleic acid interactions. Curr. Med. Chem. Anticancer Agents. 2005;5:421–429. doi: 10.2174/1568011054222337. [DOI] [PubMed] [Google Scholar]
- 83.Burden DA, Kingma PS, Froelich-Ammon SJ, Bjornsti M.-A, Patchan MW, Thompson RB, Osheroff N. Topoisomerase II•etoposide interactions direct the formation of drug-induced enzyme-DNA cleavage complexes. J. Biol. Chem. 1996;271:29238–29244. doi: 10.1074/jbc.271.46.29238. [DOI] [PubMed] [Google Scholar]
- 84.Kingma PS, Burden DA, Osheroff N. Binding of etoposide to topoisomerase II in the absence of DNA: decreased affinity as a mechanism of drug resistance. Biochemistry. 1999;38:3457–3461. doi: 10.1021/bi982855i. [DOI] [PubMed] [Google Scholar]
- 85.Leroy D, Kajava AV, Frei C, Gasser SM. Analysis of etoposide binding to subdomains of human DNA topoisomerase IIα in the absence of DNA. Biochemistry. 2001;40:1624–1634. doi: 10.1021/bi0019141. [DOI] [PubMed] [Google Scholar]
- 86.Li TK, Liu LF. Tumor cell death induced by topoisomerase-targeting drugs. Annu. Rev. Pharmacol. Toxicol. 2001;41:53–77. doi: 10.1146/annurev.pharmtox.41.1.53. [DOI] [PubMed] [Google Scholar]
- 87.Walker JV, Nitiss JL. DNA topoisomerase II as a target for cancer chemotherapy. Cancer Invest. 2002;20:570–589. doi: 10.1081/cnv-120002156. [DOI] [PubMed] [Google Scholar]
- 88.Capranico G, Guano F, Moro S, Zagotto G, Sissi C, Gatto B, Zunino F, Menta E, Palumbo M. Mapping drug interactions at the covalent topoisomerase II-DNA complex by bisantrene/amsacrine congeners. J. Biol. Chem. 1998;273:12732–12739. doi: 10.1074/jbc.273.21.12732. [DOI] [PubMed] [Google Scholar]
- 89.Lindsey R.H, Jr, Bromberg KD, Felix CA, Osheroff N. 1,4-Benzoquinone is a topoisomerase II poison. Biochemistry. 2004;43:7563–7574. doi: 10.1021/bi049756r. [DOI] [PubMed] [Google Scholar]
- 90.Bandele OJ, Osheroff N. (-)-Epigallocatechin gallate, a major constituent of green tea, poisons human type II topoisomerases. Chem. Res. Toxicol. 2008;21:936–943. doi: 10.1021/tx700434v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Wang H, Mao Y, Chen AY, Zhou N, LaVoie EJ, Liu LF. Stimulation of topoisomerase II-mediated DNA damage via a mechanism involving protein thiolation. Biochemistry. 2001;40:3316–3323. doi: 10.1021/bi002786j. [DOI] [PubMed] [Google Scholar]
- 92.Bender RP, Osheroff N. Mutation of cysteine residue 455 to alanine in human topoisomerase IIα confers hypersensitivity to quinones: enhancing DNA scission by closing the N-terminal protein gate. Chem. Res. Toxicol. 2007;20:975–981. doi: 10.1021/tx700062t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Bender RP, Ham AJ, Osheroff N. Quinone-induced enhancement of DNA cleavage by human topoisomerase IIα: adduction of cysteine residues 392 and 405. Biochemistry. 2007;46:2856–2864. doi: 10.1021/bi062017l. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Bender RP, Lehmler HJ, Robertson LW, Ludewig G, Osheroff N. Polychlorinated biphenyl quinone metabolites poison human topoisomerase IIα: altering enzyme function by blocking the N-terminal protein gate. Biochemistry. 2006;45:10140–10152. doi: 10.1021/bi0524666. [DOI] [PubMed] [Google Scholar]
- 95.Lindsey R.H., Jr, Bender RP, Osheroff N. Effects of benzene metabolites on DNA cleavage mediated by human topoisomerase IIα: 1,4-hydroquinone is a topoisomerase II poison. Chem. Res. Toxicol. 2005;18:761–770. doi: 10.1021/tx049659z. [DOI] [PubMed] [Google Scholar]
- 96.Bender RP, Lindsey R.H., Jr, Burden DA, Osheroff N. N-acetyl-p-benzoquinone imine, the toxic metabolite of acetaminophen, is a topoisomerase II poison. Biochemistry. 2004;43:3731–3739. doi: 10.1021/bi036107r. [DOI] [PubMed] [Google Scholar]
- 97.Frydman B, Marton LJ, Sun JS, Neder K, Witiak DT, Liu AA, Wang HM, Mao Y, Wu HY, Sanders MM, et al. Induction of DNA topoisomerase II-mediated DNA cleavage by beta-lapachone and related naphthoquinones. Cancer Res. 1997;57:620–627. [PubMed] [Google Scholar]
- 98.Frantz CE, Chen H, Eastmond DA. Inhibition of human topoisomerase II in vitro by bioactive benzene metabolites. Environ. Health Perspect. 1996;104(Suppl. 6):1319–1323. doi: 10.1289/ehp.961041319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Baker RK, Kurz EU, Pyatt DW, Irons RD, Kroll DJ. Benzene metabolites antagonize etoposide-stabilized cleavable complexes of DNA topoisomerase IIα. Blood. 2001;98:830–833. doi: 10.1182/blood.v98.3.830. [DOI] [PubMed] [Google Scholar]
- 100.Baldwin EL, Osheroff N. Etoposide, topoisomerase II and cancer. Curr. Med. Chem. Anticancer Agents. 2005;5:363–372. doi: 10.2174/1568011054222364. [DOI] [PubMed] [Google Scholar]
- 101.Martincic D, Hande KR. Topoisomerase II inhibitors. Cancer Chemother. Biol. Response Modif. 2005;22:101–121. doi: 10.1016/s0921-4410(04)22005-1. [DOI] [PubMed] [Google Scholar]
- 102.Hande KR. Etoposide: four decades of development of a topoisomerase II inhibitor. Eur. J. Cancer. 1998;34:1514–1521. doi: 10.1016/s0959-8049(98)00228-7. [DOI] [PubMed] [Google Scholar]
- 103.Kell J. Treatment of relapsed acute myeloid leukaemia. Rev. Recent Clin. Trials. 2006;1:103–111. doi: 10.2174/157488706776876445. [DOI] [PubMed] [Google Scholar]
- 104.Coley HM. Mechanisms and strategies to overcome chemotherapy resistance in metastatic breast cancer. Cancer Treat. Rev. 2008;34:378–390. doi: 10.1016/j.ctrv.2008.01.007. [DOI] [PubMed] [Google Scholar]
- 105.Burden DA, Osheroff N. Mechanism of action of eukaryotic topoisomerase II and drugs targeted to the enzyme. Biochim. Biophys. Acta. 1998;1400:139–154. doi: 10.1016/s0167-4781(98)00132-8. [DOI] [PubMed] [Google Scholar]
- 106.Gold R. Combination therapies in multiple sclerosis. J. Neurol. 2008;255(Suppl. 1):51–60. doi: 10.1007/s00415-008-1008-2. [DOI] [PubMed] [Google Scholar]
- 107.Azarova AM, Lyu YL, Lin CP, Tsai YC, Lau JY, Wang JC, Liu LF. Roles of DNA topoisomerase II isozymes in chemotherapy and secondary malignancies. Proc. Natl Acad. Sci. USA. 2007;104:11014–11019. doi: 10.1073/pnas.0704002104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Lyu YL, Lin CP, Azarova AM, Cai L, Wang JC, Liu LF. Role of topoisomerase IIβ in the expression of developmentally regulated genes. Mol. Cell Biol. 2006;26:7929–7941. doi: 10.1128/MCB.00617-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Sehested M, Jensen PB, Sorensen BS, Holm B, Friche E, Demant EJ. Antagonistic effect of the cardioprotector (+)-1,2-bis(3,5-dioxopiperazinyl-1-yl)propane (ICRF-187) on DNA breaks and cytotoxicity induced by the topoisomerase II directed drugs daunorubicin and etoposide (VP-16) Biochem. Pharmacol. 1993;46:389–393. doi: 10.1016/0006-2952(93)90514-w. [DOI] [PubMed] [Google Scholar]
- 110.Constantinou A, Mehta R, Runyan C, Rao K, Vaughan A, Moon R. Flavonoids as DNA topoisomerase antagonists and poisons: structure-activity relationships. J. Nat. Prod. 1995;58:217–225. doi: 10.1021/np50116a009. [DOI] [PubMed] [Google Scholar]
- 111.Strick R, Strissel PL, Borgers S, Smith SL, Rowley JD. Dietary bioflavonoids induce cleavage in the MLL gene and may contribute to infant leukemia. Proc. Natl Acad. Sci. USA. 2000;97:4790–4795. doi: 10.1073/pnas.070061297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Martin-Cordero C, Lopez-Lazaro M, Pinero J, Ortiz T, Cortes F, Ayuso MJ. Glucosylated isoflavones as DNA topoisomerase II poisons. J. Enzyme Inhib. 2000;15:455–460. doi: 10.3109/14756360009040701. [DOI] [PubMed] [Google Scholar]
- 113.Galati G, O’Brien PJ. Potential toxicity of flavonoids and other dietary phenolics: significance for their chemopreventive and anticancer properties. Free Radic. Biol. Med. 2004;37:287–303. doi: 10.1016/j.freeradbiomed.2004.04.034. [DOI] [PubMed] [Google Scholar]
- 114.Adlercreutz H, Markkanen H, Watanabe S. Plasma concentrations of phyto-oestrogens in Japanese men. Lancet. 1993;342:1209–1210. doi: 10.1016/0140-6736(93)92188-y. [DOI] [PubMed] [Google Scholar]
- 115.Lamartiniere CA. Protection against breast cancer with genistein: a component of soy. Am. J. Clin. Nutr. 2000;71:1705S–1707S. doi: 10.1093/ajcn/71.6.1705S. discussion 1708S–1709S. [DOI] [PubMed] [Google Scholar]
- 116.Siddiqui IA, Adhami VM, Saleem M, Mukhtar H. Beneficial effects of tea and its polyphenols against prostate cancer. Mol. Nutr. Food Res. 2006;50:130–143. doi: 10.1002/mnfr.200500113. [DOI] [PubMed] [Google Scholar]
- 117.Sarkar FH, Adsule S, Padhye S, Kulkarni S, Li Y. The role of genistein and synthetic derivatives of isoflavone in cancer prevention and therapy. Mini Rev. Med. Chem. 2006;6:401–407. doi: 10.2174/138955706776361439. [DOI] [PubMed] [Google Scholar]
- 118.Markovits J, Linassier C, Fosse P, Couprie J, Pierre J, Jacquemin-Sablon A, Saucier JM, Le Pecq JB, Larsen AK. Inhibitory effects of the tyrosine kinase inhibitor genistein on mammalian DNA topoisomerase II. Cancer Res. 1989;49:5111–5117. [PubMed] [Google Scholar]
- 119.Austin CA, Patel S, Ono K, Nakane H, Fisher LM. Site-specific DNA cleavage by mammalian DNA topoisomerase II induced by novel flavone and catechin derivatives. Biochem. J. 1992;282:883–889. doi: 10.1042/bj2820883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Markovits J, Junqua S, Goldwasser F, Venuat AM, Luccioni C, Beaumatin J, Saucier JM, Bernheim A, Jacquemin-Sablon A. Genistein resistance in human leukaemic CCRF-CEM cells: selection of a diploid cell line with reduced DNA topoisomerase II beta isoform. Biochem. Pharmacol. 1995;50:177–186. doi: 10.1016/0006-2952(95)00131-i. [DOI] [PubMed] [Google Scholar]
- 121.Bandele OJ, Clawson SJ, Osheroff N. Dietary polyphenols as topoisomerase II poisons: B ring and C ring substituents determine the mechanism of enzyme-mediated DNA cleavage enhancement. Chem. Res. Toxicol. 2008;21:1253–1260. doi: 10.1021/tx8000785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Bandele OJ, Osheroff N. Bioflavonoids as poisons of human topoisomerase IIα and IIβ. Biochemistry. 2007;46:6097–6108. doi: 10.1021/bi7000664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Ross JA, Potter JD, Robison LL. Infant leukemia, topoisomerase II inhibitors, and the MLL gene. J. Natl. Cancer Inst. 1994;86:1678–1680. doi: 10.1093/jnci/86.22.1678. [DOI] [PubMed] [Google Scholar]
- 124.Ross JA, Potter JD, Reaman GH, Pendergrass TW, Robison LL. Maternal exposure to potential inhibitors of DNA topoisomerase II and infant leukemia (United States): a report from the Children's Cancer Group. Cancer Causes Control. 1996;7:581–590. doi: 10.1007/BF00051700. [DOI] [PubMed] [Google Scholar]
- 125.Ross JA. Dietary flavonoids and the MLL gene: a pathway to infant leukemia? Proc. Natl Acad. Sci. USA. 2000;97:4411–4413. doi: 10.1073/pnas.97.9.4411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Spector LG, Xie Y, Robison LL, Heerema NA, Hilden JM, Lange B, Felix CA, Davies SM, Slavin J, Potter JD, et al. Maternal diet and infant leukemia: the DNA topoisomerase II inhibitor hypothesis: a report from the children's oncology group. Cancer Epidemiol. Biomarkers Prev. 2005;14:651–655. doi: 10.1158/1055-9965.EPI-04-0602. [DOI] [PubMed] [Google Scholar]
- 127.Anderson VE, Osheroff N. Type II topoisomerases as targets for quinolone antibacterials: turning Dr. Jekyll into Mr. Hyde. Curr. Pharm. Des. 2001;7:337–353. doi: 10.2174/1381612013398013. [DOI] [PubMed] [Google Scholar]
- 128.Drlica K, Malik M. Fluoroquinolones: action and resistance. Curr. Top. Med. Chem. 2003;3:249–282. doi: 10.2174/1568026033452537. [DOI] [PubMed] [Google Scholar]
- 129.Levine C, Hiasa H, Marians KJ. DNA gyrase and topoisomerase IV: biochemical activities, physiological roles during chromosome replication, and drug sensitivities. Biochim. Biophys. Acta. 1998;1400:29–43. doi: 10.1016/s0167-4781(98)00126-2. [DOI] [PubMed] [Google Scholar]
- 130.Ross D. The role of metabolism and specific metabolites in benzene-induced toxicity: evidence and issues. J. Toxicol. Environ. Health A. 2000;61:357–372. doi: 10.1080/00984100050166361. [DOI] [PubMed] [Google Scholar]
- 131.Lindsey RH, Bender RP, Osheroff N. Stimulation of topoisomerase II-mediated DNA cleavage by benzene metabolites. Chem. Biol. Interact. 2005;153-154:197–205. doi: 10.1016/j.cbi.2005.03.035. [DOI] [PubMed] [Google Scholar]
- 132.Shen Y, Shen HM, Shi CY, Ong CN. Benzene metabolites enhance reactive oxygen species generation in HL60 human leukemia cells. Hum. Exp. Toxicol. 1996;15:422–427. doi: 10.1177/096032719601500510. [DOI] [PubMed] [Google Scholar]
- 133.Kuo ML, Shiah SG, Wang CJ, Chuang SE. Suppression of apoptosis by Bcl-2 to enhance benzene metabolites-induced oxidative DNA damage and mutagenesis: a possible mechanism of carcinogenesis. Mol. Pharmacol. 1999;55:894–901. [PubMed] [Google Scholar]
- 134.Lovern MR, Cole CE, Schlosser PM. A review of quantitative studies of benzene metabolism. Crit. Rev. Toxicol. 2001;31:285–311. doi: 10.1080/20014091111703. [DOI] [PubMed] [Google Scholar]
- 135.Rappaport SM, Waidyanatha S, Qu Q, Shore R, Jin X, Cohen B, Chen LC, Melikian AA, Li G, Yin S, et al. Albumin adducts of benzene oxide and 1,4-benzoquinone as measures of human benzene metabolism. Cancer Res. 2002;62:1330–1337. [PubMed] [Google Scholar]
- 136.Prescott LF. Paracetamol: past, present, and future. Am. J. Ther. 2000;7:143–147. [PubMed] [Google Scholar]
- 137.Kingma PS, Greider CA, Osheroff N. Spontaneous DNA lesions poison human topoisomerase IIα and stimulate cleavage proximal to leukemic 11q23 chromosomal breakpoints. Biochemistry. 1997;36:5934–5939. doi: 10.1021/bi970507v. [DOI] [PubMed] [Google Scholar]
- 138.Kingma PS, Osheroff N. Apurinic sites are position-specific topoisomerase II-poisons. J. Biol. Chem. 1997;272:1148–1155. doi: 10.1074/jbc.272.2.1148. [DOI] [PubMed] [Google Scholar]
- 139.Kingma PS, Osheroff N. Spontaneous DNA damage stimulates topoisomerase II-mediated DNA cleavage. J. Biol. Chem. 1997;272:7488–7493. doi: 10.1074/jbc.272.11.7488. [DOI] [PubMed] [Google Scholar]
- 140.Kingma PS, Osheroff N. The response of eukaryotic topoisomerases to DNA damage. Biochim. Biophys. Acta. 1998;1400:223–232. doi: 10.1016/s0167-4781(98)00138-9. [DOI] [PubMed] [Google Scholar]
- 141.Cline SD, Jones WR, Stone MP, Osheroff N. DNA abasic lesions in a different light: solution structure of an endogenous topoisomerase II poison. Biochemistry. 1999;38:15500–15507. doi: 10.1021/bi991750s. [DOI] [PubMed] [Google Scholar]
- 142.Cline SD, Osheroff N. Cytosine arabinoside lesions are position-specific topoisomerase II poisons and stimulate DNA cleavage mediated by human type II enzymes. J. Biol. Chem. 1999;274:29740–29743. doi: 10.1074/jbc.274.42.29740. [DOI] [PubMed] [Google Scholar]
- 143.Sabourin M, Osheroff N. Sensitivity of human type II topoisomerases to DNA damage: stimulation of enzyme-mediated DNA cleavage by abasic, oxidized and alkylated lesions. Nucleic Acids Res. 2000;28:1947–1954. doi: 10.1093/nar/28.9.1947. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Khan QA, Kohlhagen G, Marshall R, Austin CA, Kalena GP, Kroth H, Sayer JM, Jerina DM, Pommier Y. Position-specific trapping of topoisomerase II by benzo[a]pyrene diol epoxide adducts: implications for interactions with intercalating anticancer agents. Proc. Natl Acad. Sci. USA. 2003;100:12498–12503. doi: 10.1073/pnas.2032456100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Solovyan VT, Bezvenyuk ZA, Salminen A, Austin CA, Courtney MJ. The role of topoisomerase II in the excision of DNA loop domains during apoptosis. J. Biol. Chem. 2002;277:21458–21467. doi: 10.1074/jbc.M110621200. [DOI] [PubMed] [Google Scholar]
- 146.Belyaev IY. DNA loop organization and DNA fragmentation during radiation-induced apoptosis in human lymphocytes. Radiats. Biol. Radioecol. 2005;45:541–548. [PubMed] [Google Scholar]
- 147.Felix CA, Hosler MR, Winick NJ, Masterson M, Wilson AE, Lange BJ. ALL-1 gene rearrangements in DNA topoisomerase II inhibitor-related leukemia in children. Blood. 1995;85:3250–3256. [PubMed] [Google Scholar]
- 148.Felix CA. Secondary leukemias induced by topoisomerase-targeted drugs. Biochim. Biophys. Acta. 1998;1400:233–255. doi: 10.1016/s0167-4781(98)00139-0. [DOI] [PubMed] [Google Scholar]
- 149.Hasan SK, Mays AN, Ottone T, Ledda A, La Nasa G, Cattaneo C, Borlenghi E, Melillo L, Montefusco E, Cervera J, et al. Molecular analysis of t(15;17) genomic breakpoints in secondary acute promyelocytic leukemia arising after treatment of multiple sclerosis. Blood. 2008;112:3383–3390. doi: 10.1182/blood-2007-10-115600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Mistry AR, Felix CA, Whitmarsh RJ, Mason A, Reiter A, Cassinat B, Parry A, Walz C, Wiemels JL, Segal MR, et al. DNA topoisomerase II in therapy-related acute promyelocytic leukemia. N. Engl. J. Med. 2005;352:1529–1538. doi: 10.1056/NEJMoa042715. [DOI] [PubMed] [Google Scholar]
- 151.Gilliland DG, Jordan CT, Felix CA. The molecular basis of leukemia. Hematol. Am. Soc. Hematol. Educ. Program. 2004:80–97. doi: 10.1182/asheducation-2004.1.80. [DOI] [PubMed] [Google Scholar]
- 152.Errington F, Willmore E, Tilby MJ, Li L, Li G, Li W, Baguley BC, Austin CA. Murine transgenic cells lacking DNA topoisomerase IIbeta are resistant to acridines and mitoxantrone: analysis of cytotoxicity and cleavable complex formation. Mol. Pharmacol. 1999;56:1309–1316. doi: 10.1124/mol.56.6.1309. [DOI] [PubMed] [Google Scholar]
- 153.Willmore E, Frank AJ, Padget K, Tilby MJ, Austin CA. Etoposide targets topoisomerase IIα and IIβ in leukemic cells: isoform-specific cleavable complexes visualized and quantified in situ by a novel immunofluorescence technique. Mol. Pharm. 1998;54:78–85. doi: 10.1124/mol.54.1.78. [DOI] [PubMed] [Google Scholar]
- 154.Bandele OJ, Osheroff N. The efficacy of topoisomerase II-targeted anticancer agents reflects the persistence of drug-induced cleavage complexes in cells. Biochemistry. 2008;47:11900–11908. doi: 10.1021/bi800981j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Beese LS, Friedman JM, Steitz TA. Crystal structures of the Klenow fragment of DNA polymerase I complexed with deoxynucleoside triphosphate and pyrophosphate. Biochemistry. 1993;32:14095–14101. doi: 10.1021/bi00214a004. [DOI] [PubMed] [Google Scholar]
- 156.Curley JF, Joyce CM, Piccirilli JA. Functional evidence that the 3′-5′ exonuclease domain of Escherichia coli DNA polymerase I employs a divalent metal ion in leaving group stabilization. J. Am. Chem. Soc. 1997;119:12691–12692. [Google Scholar]