Abstract
Background
Alternative pre-mRNA splicing (AS) plays a central role in generating complex proteomes and influences development and disease. However, the regulation and etiology of AS in human tumorigenesis is not well understood.
Methodology/Principal Findings
A Basic Local Alignment Search Tool database was constructed for the expressed sequence tags (ESTs) from all available databases of human cancer and normal tissues. An insertion or deletion in the alignment of EST/EST was used to identify alternatively spliced transcripts. Alignment of the ESTs with the genomic sequence was further used to confirm AS. Alternatively spliced transcripts in each tissue were then subtractively cross-screened to obtain tissue-specific variants. We systematically identified and characterized cancer/tissue-specific and alternatively spliced variants in the human genome based on a global view. We identified 15,093 cancer-specific variants of 9,989 genes from 27 types of human cancers and 14,376 normal tissue-specific variants of 7,240 genes from 35 normal tissues, which cover the main types of human tumors and normal tissues. Approximately 70% of these transcripts are novel. These data were integrated into a database HCSAS (http://202.114.72.39/database/human.html, pass:68756253). Moreover, we observed that the cancer-specific AS of both oncogenes and tumor suppressor genes are associated with specific cancer types. Cancer shows a preference in the selection of alternative splice-sites and utilization of alternative splicing types.
Conclusions/Significance
These features of human cancer, together with the discovery of huge numbers of novel splice forms for cancer-associated genes, suggest an important and global role of cancer-specific AS during human tumorigenesis. We advise the use of cancer-specific alternative splicing as a potential source of new diagnostic, prognostic, predictive, and therapeutic tools for human cancer. The global view of cancer-specific AS is not only useful for exploring the complexity of the cancer transcriptome but also widens the eyeshot of clinical research.
Introduction
It remains unknown how both intron removal and exon rearrangement are precisely regulated to produce correct proteomes in a cell type- or developmental stage-specific manner. Alternative splicing, the process by which the exons of primary transcripts can be spliced into different arrangements to produce structurally and functionally distinct mRNA and protein variants, is the most widely used mechanism to enhance the protein diversity of higher eukaryotic organisms. It has been estimated that 35%–94% of all human genes appear to undergo alternative splicing [1]–[7], suggesting that this mechanism has a major role in generating protein diversity. As sequence data continue to be generated from projects at an ever-increasing rate, the need for mining the data and constructing a repository for transcriptome information continues to grow as well.
In many pathological conditions, aberrantly spliced pre-mRNAs are generated because they escape the quality control mechanisms within cells (e.g. the nonsense mediated mRNA decay pathway) and are, therefore, translated into aberrant proteins involved in human diseases, including cancer [8]–[11]. It is estimated that approximately 60% of disease mutations in the human genome are splicing mutations [12], [13]. Currently, the analysis of cancer-specific alternative splicing is a promising step forward and potential source of new clinical diagnostic, prognostic, and therapeutic strategies. Evidence is accumulating that supports a connection between tumorigenesis and alternative splicing [14]–[18]. Using bioinformatic approaches, Xu and Lee discovered cancer-specific splice variants in 316 genes [19]. We previously identified testis-/testis cancer-specific splice variants using bioinformatic and experimental approaches [20].
Despite the growing interest in the impact of alternative splicing in various aspects of the biological processes, our understanding of alternative splicing is still scattered, and its general regulatory mechanisms, especially in tumorigenesis, are not well known [21], [22]. However, it is believed that cancer-specific splice variants could be involved in the etiopathogeny of many diseases and some might serve as diagnostic or prognostic markers. Moreover, the direct targeting of protein is probably an advantageous way of correcting cancer-associated splicing alterations. For example, the cancer-restricted splice variant protein could be used as the target for specific antibodies conjugated to tumor cell toxins for cancer treatments. The etiopathogeny concerning the cancer-specific AS and all related applications need to be explored further.
In order to advance our understanding of the biological significance of alternative splicing in human cancers, it is essential to systematically identify cancer-specific splicing events at the transcriptome level. In the present study, we performed a genome-wide analysis of alternative splicing in human cancer and normal tissues using an intersection/subtractive model consisting of the following steps: 1) identifying insertions or deletions in the alignments of expressed sequence tags (ESTs) to identify alternative splicing transcripts based on a previously described method [2], 2) the alignment of EST/genome to confirm the transcripts, and 3) obtaining the tissue-specific and alternatively spliced variants by subtractively cross-screening the alternatively spliced transcripts in each tissue. Our results distinguish distinctive patterns of cancer-specific alternative splicing and identify a large number of cancer- and tissue-specific splicing isoforms, which provides a global view of human cancer-specific alternative splicing in a large-scale approach and a potential source of new clinical diagnostic, prognostic, and therapeutic strategies for human cancer.
Materials and Methods
Data sources and filtration
Human EST data for both cancerous and normal tissues were drawn from the Cancer Genome Anatomy Project (CGAP) (http://cgap.nci.nih.gov/Tissues/LibraryFinder). The CGAP collects EST libraries from all over the world and provides good tissue information. All available EST libraries for both human cancer and normal tissues were downloaded from the CGAP libraries, Mammalian Gene Collection libraries, and Open Reading Frame EST Sequencing libraries. We sought to avoid mixing multiple tissues. Among these libraries, those signed ‘pooled’ were excluded because these procedures affect tissue classification. For normal tissue, ESTs were classified in accordance with the developmental stage information, and libraries without this information were not used. All EST and library data on different tissues that were used are listed in Tables 1 and 2.
Table 1. Numbers of libraries and ESTs in cancers.
| Cancer types (27 types) | Libraries | ESTs |
| adrenal_cancer | 3 | 10431 |
| bone_marrow_leukemia | 21 | 43943 |
| brain_glioma | 43 | 75546 |
| brain_meningioma | 7 | 860 |
| brain_cancer | 213 | 39845 |
| breast_cancer | 792 | 143423 |
| cervical_cancer | 32 | 48606 |
| chondrosarcoma | 15 | 49638 |
| colorectal_cancer | 803 | 185632 |
| esophageal_cancer | 17 | 15039 |
| germ_cell_cancer | 30 | 161682 |
| head_and_neck_cancer | 291 | 93978 |
| kidney_cancer | 87 | 74290 |
| liver_cancer | 62 | 117276 |
| lung_cancer | 220 | 126775 |
| lymphoma | 6 | 12458 |
| muscle_tissue_cancer | 26 | 83411 |
| ovarian_cancer | 162 | 98087 |
| pancreas_insulinoma | 1 | 33046 |
| pancreatic_cancer | 22 | 82994 |
| primitive_neuroectodermal_cancer_of_CNS | 23 | 72677 |
| prostate_cancer | 169 | 168151 |
| prostatic_intraepithelial_neoplasia | 5 | 9569 |
| retinoblastoma | 3 | 51568 |
| skin_cancer | 36 | 137803 |
| stomach_cancer | 244 | 94950 |
| uterus_cancer | 110 | 46624 |
| Total | 3,443 | 2,078,302 |
Table 2. Numbers of libraries and ESTs in normal tissues.
| Tissue type (35 tissues) | Libraries | ESTs |
| bone-adult | 2 | 1965 |
| brain-adult | 368 | 130619 |
| brain-fetus | 73 | 174889 |
| brain-infant | 11 | 73726 |
| colon-adult | 133 | 25757 |
| colon-fetus | 1 | 5 |
| eye-adult | 33 | 100173 |
| eye-fetus | 7 | 19659 |
| heart-adult | 10 | 3979 |
| heart-fetus | 13 | 62681 |
| kidney-fetus | 16 | 13947 |
| liver-adult | 11 | 29818 |
| liver-fetus | 22 | 142750 |
| lung-adult | 92 | 44457 |
| lung-fetus | 18 | 35607 |
| mammary-gland-adult | 331 | 68603 |
| muscle-adult | 10 | 70211 |
| muscle-fetus | 4 | 2235 |
| ovary-adult | 6 | 8126 |
| pancreas-adult | 10 | 24759 |
| pancreas-fetus | 2 | 5545 |
| peripheral-nerve-adult | 1 | 6571 |
| peripheral-nerve-juvenile | 1 | 9482 |
| pituitary-gland-adult | 6 | 8520 |
| placenta-adult | 358 | 268277 |
| prostate-adult | 127 | 53220 |
| skin-infant | 3 | 10319 |
| spleen-adult | 3 | 1841 |
| spleen-fetus | 1 | 1332 |
| stomach-adult | 67 | 10417 |
| stomach-fetus | 2 | 9 |
| testis-adult | 157 | 30549 |
| thyroid-adult | 78 | 13314 |
| uterus-adult | 10 | 35026 |
| vascular-adult | 5 | 8451 |
| Total | 1,992 | 1,496,839 |
All collection data were then dealt with in three procedures: repeat sequence masking to remove simple repeats in the dataset (program, repeatmasker; repeat database, repbase; girnst server:www.girinst.org), vector and contamination masking to clean the vector sequences (program, crossmatch; vector database, UniVec_Core; National Center for Biotechnology Information ftp server: ftp://ftp.ncbi.nih.gov/), and a final cleaning of short and rubbish sequences (program, seqclean from egassembler server: http://egassembler.hgc.jp). Any Alu repeats were included in, and the filtered ESTs were available for the following analysis.
Computational procedures to identify cancer/tissue-specific alternative splicing
A basic local alignment search tool (BLAST) database was constructed for the ESTs of each tissue. Alternative splicing was analyzed based on a previous method [2]. Transcripts specific to tissue T were identified based on an intersection/subtractive model:
Where TS is the alternatively spliced transcripts specific to tissue T, T is all transcripts in tissue T, and O is all transcripts in the other tissues (∩, intersection).
Briefly, the three steps were as follows:
Tissue T's EST dataset was BLASTed against itself. The e-value was set to 1e-30. Gaps (insertion or deletion) in the ESTs were identified after alignment. Parameters to identify alternative splicing: the gap length, 10 bp; nucleotide identity, 95%.
Tissue T's ESTs were BLASTed against the ESTs of the other tissues. Parameters were the same as step 1.
Subtractive ESTs were identified as tissue T-specific ESTs by insertion/deletion comparisons after BLAST. Computer programs were written using the Perl language.
EST/genomic sequence alignments, chromosome mapping, and splice site analysis
To decrease errors in EST alignments and determine the chromosomal loci of each gene, we localized ESTs to genomic sequences using BLAST-like alignment tools (http://genome.ucsc.edu). We used the default parameters and selected the best score results. The exon position on the chromosome was recorded for each transcript and used to determine splice sites and gene structure. Splice sites for both 5′ and 3′ exon/intron boundaries were aligned online via http://weblogo.berkeley.edu/logo.cgi. We allowed an error of 10 bp in the exon/intron boundary. Based on comparisons of EST/genomic alignments, two possible errors can be checked: (i) if the candidate EST in the same gene was not on the same chromosome and (ii) is the candidate EST in the same gene was not in the same locus on the chromosome. The reasons for these errors mainly included EST sequencing errors, pseudogenes, and multiple copy genes. The two cases were excluded as false positives in the final database.
Function classification of alternative splicing
Each alternatively spliced EST was BLASTed to the RefSeq mRNA database (expectations 1e-30) to identify the corresponding genes. Using PANTHER (http://www.pantherdb.org/tools/genexAnalysis.jsp), these genes were clustered by the gene ontology (GO) process. We also searched the Entrez Gene Database to correct our results.
Alternative splicing database construction
We input all prediction results into the local alternative splicing database. This database was constructed with MySql and programmed by Perl and CGI. All information such as gene ID, gene structure, EST accession, mRNA accession, gene information, and exon location on the chromosome were collected in the database.
Results
HCSAS: A database for cancer-specific alternative splicing
For analyzing cancer-specific alternative splicing, we carefully classified all available EST libraries into 35 distinct normal tissue classes and 27 types of cancer to avoid mixing multiple tissues. Our final classification consisted of 1,992 libraries with 1,496,839 ESTs for normal human samples and 3,443 libraries with 2,078,302 ESTs for cancer samples (Tables 1 and 2). Through computationally subtractive analysis, we detected 15,093 cancer-specific transcripts in 9,989 genes from the 27 types of cancer, and 14,376 normal tissue-specific transcripts in 7,240 genes from the 35 tissues (Tables 3 and 4), which cover the main types of human tumors and tissues. Cancer-specific transcript numbers per gene detected were 1 to 1.69 with an average of 1.51, whereas there were 1 to 6 normal tissue-specific transcripts with an average of 1.99 (Tables 3 and 4), indicating fewer alternative splicing events (cancer-specific) in cancer compared to normal tissues.
Table 3. Numbers of cancer-specific AS transcripts and their genes.
| Cancer types (27 types) | Genes | Transcripts | Transcripts/Gene |
| adrenal_cancer | 61 | 80 | 1.31 |
| bone_marrow_leukemia | 237 | 356 | 1.50 |
| brain_glioma | 485 | 720 | 1.48 |
| brain_meningioma | 1 | 1 | 1.00 |
| brain_cancer | 22 | 30 | 1.36 |
| breast_cancer | 550 | 757 | 1.38 |
| cervical_cancer | 316 | 428 | 1.35 |
| chondrosarcoma | 239 | 352 | 1.47 |
| colorectal_cancer | 397 | 578 | 1.46 |
| esophageal_cancer | 175 | 226 | 1.29 |
| germ_cell_cancer | 1307 | 2167 | 1.66 |
| head_and_neck_cancer | 135 | 182 | 1.35 |
| kidney_cancer | 467 | 716 | 1.53 |
| liver_cancer | 950 | 1410 | 1.48 |
| lung_cancer | 605 | 880 | 1.45 |
| lymphoma | 16 | 21 | 1.31 |
| muscle_tissue_cancer | 692 | 1044 | 1.51 |
| ovarian_cancer | 351 | 512 | 1.46 |
| pancreas_insulinoma | 45 | 76 | 1.69 |
| pancreatic_cancer | 426 | 643 | 1.51 |
| primitive_neuroectodermal_cancer_of_CNS | 659 | 1018 | 1.54 |
| prostate_cancer | 92 | 111 | 1.21 |
| prostatic_intraepithelial_neoplasia | 11 | 18 | 1.64 |
| retinoblastoma | 434 | 705 | 1.62 |
| skin_cancer | 939 | 1557 | 1.66 |
| stomach_cancer | 249 | 326 | 1.31 |
| uterus_cancer | 128 | 179 | 1.40 |
| Total | 9,989 | 15,093 | 1.51 |
Table 4. Numbers of normal tissue-specific AS transcripts and their genes.
| Tissue types (35 tissues) | Genes | Transcripts | Transcripts/Gene |
| bone-adult | 1 | 2 | 2 |
| brain-adult | 924 | 1613 | 1.75 |
| brain-fetus | 2231 | 5545 | 2.49 |
| brain-infant | 53 | 72 | 1.36 |
| colon-adult | 17 | 22 | 1.29 |
| colon-fetus | 1 | 2 | 2.00 |
| eye-adult | 582 | 907 | 1.56 |
| eye-fetus | 58 | 76 | 1.31 |
| heart-adult | 6 | 9 | 1.50 |
| heart-fetus | 100 | 146 | 1.46 |
| kidney-fetus | 110 | 142 | 1.29 |
| liver-adult | 335 | 961 | 2.87 |
| liver-fetus | 292 | 506 | 1.73 |
| lung-adult | 78 | 160 | 2.05 |
| lung-fetus | 110 | 152 | 1.38 |
| mammary-adult | 41 | 57 | 1.39 |
| muscle-adult | 250 | 384 | 1.54 |
| muscle-fetus | 18 | 27 | 1.50 |
| ovary-adult | 19 | 26 | 1.37 |
| pancreas-adult | 28 | 39 | 1.39 |
| pancreas-fetus | 23 | 33 | 1.43 |
| peripheral-nerve-adult | 18 | 23 | 1.28 |
| peripheral-nerve-juvenile | 18 | 23 | 1.28 |
| pituitary-gland-adult | 28 | 84 | 3.00 |
| placenta-adult | 1567 | 2902 | 1.85 |
| prostate-adult | 102 | 162 | 1.59 |
| skin-infant | 99 | 121 | 1.22 |
| spleen-adult | 2 | 2 | 1.00 |
| spleen-fetus | 1 | 1 | 1.00 |
| stomach-adult | 3 | 5 | 1.67 |
| stomach-fetus | 1 | 6 | 6.00 |
| testis-adult | 47 | 73 | 1.55 |
| thyroid-adult | 16 | 20 | 1.25 |
| uterus-adult | 51 | 63 | 1.2 |
| vascular-adult | 10 | 10 | 1.00 |
| Total | 7240 | 14,376 | 1.99 |
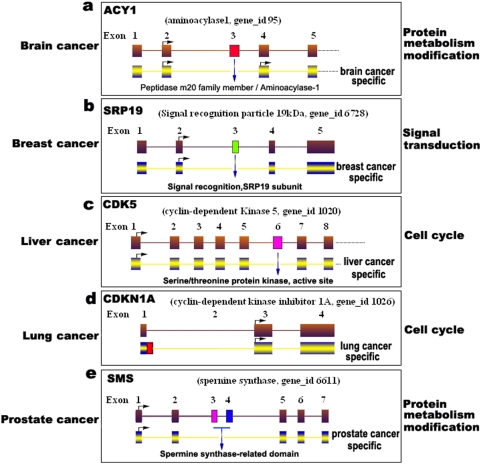
To facilitate future studies and referencing of alternatively spliced genes, for both human cancer and normal tissues, we constructed a human cancer- and normal tissue-specific alternative splicing database (HCSAS) based on our analysis, which was divided into two parts: cancer-specific (15,093 transcripts) and normal tissue-specific (14,376) alternative splicing. Of these cancer- or tissue-specific AS, approximately 70% are novel isoforms. For example, in brain cancer, because of the alternative splicing and deletion of domain of the peptidase m20 family member, the aminoacylase-1 gene (ACY1) was spliced to produce a brain cancer-specific transcript (Figure 1a), and alternative splicing occurs in the SRP19 gene to produce a breast cancer-specific transcript by an alternative deletion of exon 3 (Figure 1b). Similarly, in liver cancer, lung cancer, and prostate cancer, cancer-specific isoforms were detected in our subtractive screening (Figure 1c–e).
Figure 1. A schematic representation of cancer-specific alternative gene splicing.
(a) Brain cancer (gene ACYl), (b) breast cancer (SRP19), (c) liver cancer (CDK5), (d) lung cancer (CDKN1A), and (e) prostate cancer (SMS). Cancer-specific isoforms are showed on the bottom in each panel. The biological processes of these transcripts (GO process) are indicated on the right. Deleted domains are shown with blue arrows. Arrows with a right angle indicate the start codon, ATG.
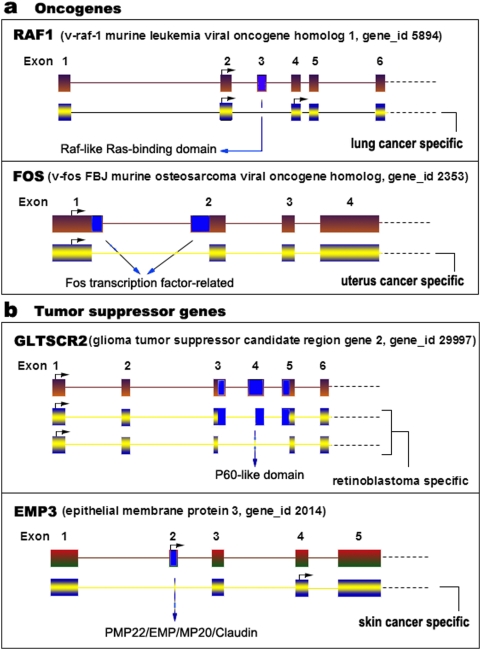
Furthermore, we systematically identified cancer-specific transcripts in both oncogenes and tumor suppressors. Thirty-nine oncogene isoforms and 38 tumor suppressor gene isoforms with cancer-specific AS events were detected (Table 5). For example, we identified a lung cancer-specific transcript in the oncogene RAF1 with a deletion of the Raf-like Ras-binding domain, an uterus cancer-specific transcript in oncogene FOS (Figure 2a), and a retinoblastoma-specific transcript in the tumor suppressor GLTSCR2, and a skin-cancer-specific transcript in the tumor suppressor EMP3 (Figure 2b).
Table 5. Oncogenes and tumor suppressors with cancer-specific AS events.
| Gene ID | Symbol | Gene Description | AS |
| Oncogenes | |||
| 25 | ABL1 | v-abl Abelson murine leukemia viral oncogene homolog 1 | 1 |
| 3726 | JUNB | jun B proto-oncogene | 2 |
| 7409 | VAV1 | vav 1 oncogene | 1 |
| 6757 | SSX2 | synovial sarcoma, X breakpoint 2 | 3 |
| 2130 | EWSR1 | Ewing sarcoma breakpoint region 1 | 3 |
| 2241 | FER | fer (fps/fes related) tyrosine kinase (phosphoprotein NCP94) | 2 |
| 369 | ARAF | v-raf murine sarcoma 3611 viral oncogene homolog | 1 |
| 4613 | MYCN | v-myc myelocytomatosis viral related oncogene | 2 |
| 2534 | FYN | FYN oncogene related to SRC, FGR, YES | 2 |
| 727735 | unassigned | similar to TBC1 domain family member 3 | 1 |
| 51513 | ETV7 | ets variant gene 7 (TEL2 oncogene) | 2 |
| 5894 | RAF1 | v-raf-1 murine leukemia viral oncogene homolog 1 | 2 |
| 4193 | MDM2 | Mdm2, transformed 3T3 cell double minute 2, p53 binding protein | 1 |
| 4609 | MYC | v-myc myelocytomatosis viral oncogene homolog | 1 |
| 2353 | FOS | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 |
| 7410 | VAV2 | vav 2 oncogene | 1 |
| 4194 | MDM4 | Mdm4, transformed 3T3 cell double minute 4, p53 binding protein | 1 |
| 2118 | ETV4 | ets variant gene 4 (E1A enhancer binding protein, E1AF) | 3 |
| 598 | BCL2L1 | BCL2-like 1 | 3 |
| 55885 | LMO3 | LIM domain only 3 (rhombotin-like 2) | 1 |
| 3265 | HRAS | v-Ha-ras Harvey rat sarcoma viral oncogene homolog | 2 |
| 4893 | NRAS | neuroblastoma RAS viral (v-ras) oncogene homolog | 1 |
| Total | 39 | ||
| Tumor suppressors | |||
| 5934 | RBL2 | retinoblastoma-like 2 (p130) | 1 |
| 3482 | IGF2R | insulin-like growth factor 2 receptor | 1 |
| 5925 | RB1 | retinoblastoma 1 (including osteosarcoma) | 1 |
| 54984 | unassigned | PIN2-interacting protein 1 | 1 |
| 4017 | LOXL2 | lysyl oxidase-like 2 | 3 |
| 29997 | GLTSCR2 | glioma tumor suppressor candidate region gene 2 | 3 |
| 2014 | EMP3 | epithelial membrane protein 3 | 1 |
| 672 | BRCA1 | breast cancer 1, early onset | 2 |
| 54879 | ST7L | suppression of cancerigenicity 7 like | 2 |
| 51147 | ING4 | inhibitor of growth family, member 4 | 1 |
| 7982 | ST7 | suppression of cancerigenicity 7 | 1 |
| 51566 | ARMCX3 | armadillo repeat containing, X-linked 3 | 4 |
| 84695 | LOXL3 | lysyl oxidase-like 3 | 2 |
| 79961 | DENND2D | DENN/MADD domain containing 2D | 2 |
| 7157 | TP53 | cancer protein p53 (Li-Fraumeni syndrome) | 1 |
| 7248 | TSC1 | tuberous sclerosis 1 | 1 |
| 54768 | HYDIN | hydrocephalus inducing homolog | 3 |
| 581 | BAX | BCL2-associated X protein | 1 |
| 1026 | CDKN1A | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 5 |
| 1029 | CDKN2A | cyclin-dependent kinase inhibitor 2A | 1 |
| 10263 | CDK2AP2 | CDK2-associated protein 2 | 1 |
| Total | 38 | ||
Figure 2. A schematic representation of cancer-specific alternative gene splicing.
(a) Oncogene, (b) tumor suppressor gene. The alternative splicing of RAF1 generates a lung cancer-specific transcript, whereas the alternative splicing of FOS produces an uterus cancer-specific transcript. Tumor suppressor GLTSCR2 is alternatively spliced to produce two retinoblastoma-specific transcripts and EMP3 to generate a skin cancer-specific transcript. Deleted domains are shown with blue arrows. Arrows with a right angle indicate the start codon, ATG.
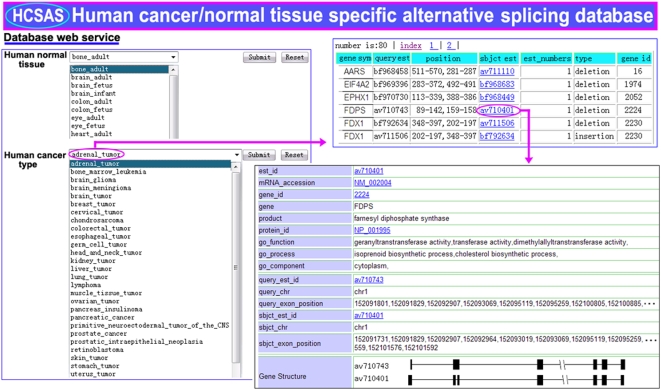
The HCSAS database presents a global overview of cancer-specific alternative splicing in humans and is essential for understanding tumorigenesis at a systematic level. The main information in this database includes the specific alternative splicing in both cancer and normal tissues, gene ID, gene structure, splicing sites, chromosome localization, DNA and protein sequences linked with the NCBI website, and GO process, function, and subcellular localization. An example page set shows the details of an adrenal cancer gene, FDPS (Figure 3). The HCSAS database can be accessed at http://202.114.72.39/database/human.html.
Figure 3. A database of cancer- and normal tissue-specific alternative splicing.
An example page set from the database shows the details of an adrenal cancer gene, FDPS. The information includes the specific alternative splicing of both cancer and normal tissues, gene ID, gene structure, splicing sites, chromosome localization, DNA and protein sequences linked with the NCBI website, and GO process, function, and subcellular localization.
Biased utilization of alternative splicing types in cancer
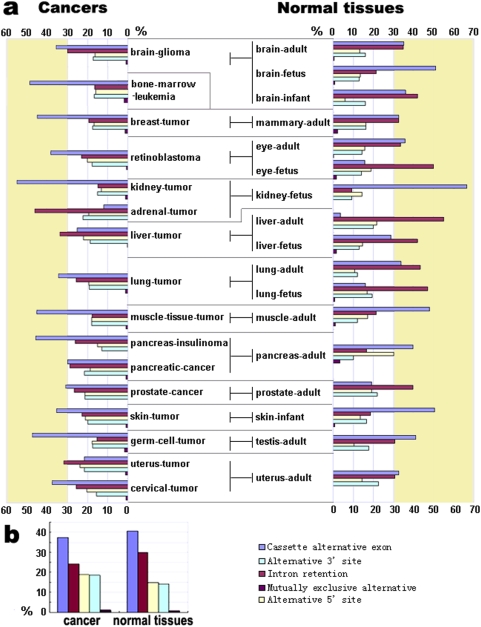
An examination of cancer-specific alternative splicing revealed a biased distribution of alternative splicing types in cancer. Both the alternative 3′ splice site and 5′ splice site were used more often in cancer; however, a lower proportion of intron retention and cassette alternative exon occurred in cancer tissues compared to normal tissues (Figure 4b). Moreover, alternative splicing types differ between different kinds of cancer (Figure 4a). For example, in liver cancer, breast cancer, and prostate cancer, intron retention decreased and cassette alternative exons increased significantly, whereas in uterus cancer and skin cancer, cassette alternative exons markedly decreased.
Figure 4. The frequencies (percentages) of the five types of cancer- and normal tissue-specific alternative splicing.
(a) 16 types of human cancer and 17 normal tissues, (b) the average values between tumors and normal tissues. The five colors indicate the five types of tissue-specific alternative splicing: cassette alternative exon, alternative 5′ splice site, alternative 3′ splice site, intron retention, and mutually exclusive alternative exons. Yellowish regions indicate over 30% of the frequencies.
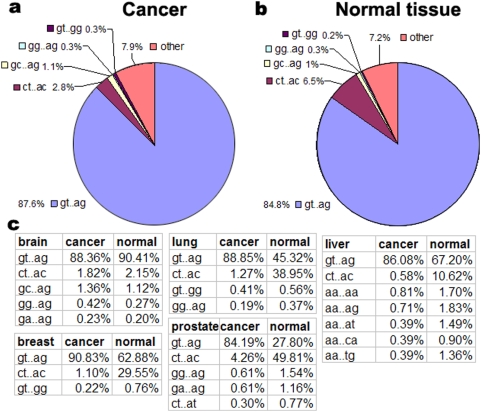
Preference in the selection of alternative splice sites in cancer
To explore the preference/diversification of alternative splice sites in cancer, we analyzed all splice sites in the 27 types of cancer and 35 normal tissues by comparing each EST with its genomic sequence and mapping it onto the chromosome. We detected five basic donor-acceptor splice sites: GT-AG, CT-AC, GC-AG, GG-AG, and GT-GG, of which GT-AG are the most dominant sites. The others were classified into rare splice sites. We found that cancer uses rare splice sites and GT-AG more frequently, but less CT-AC compared to normal tissues (Figure 5a, b). Moreover, the selection of splice sites differs between different kinds of cancer (Figure 5c). For example, CT-AC sites are seldom used in breast cancer, liver cancer, lung cancer, and prostate cancer; in liver cancer, 5′ sites of rare splicing are almost AA.
Figure 5. Percentages of the types of alternative splice sites.
The splice sites include GT-AG, GC-AG, GG-AG, GT-GG, and the others (a) in human cancer (b) and normal tissues. (c) Percentage distribution of the splice sites in five types of cancer and normal tissues (brain, breast, lung, liver, and prostate).
Association of cancer-specific alternative splicing of both oncogenes and tumor suppressor genes with cancer
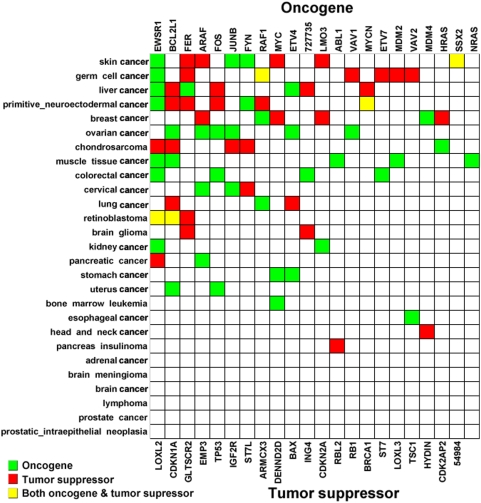
Although both oncogenes and tumor suppressors are thought to be vital factors in tumorigenesis, we sought to identify cancer-specific variants and their possible involvement in cancer. We observed that oncogenes with cancer-specific AS are more often present in ovary cancer (6 oncogenes) and muscle cancer (5 oncogenes), whereas tumor suppressor genes with cancer-specific AS are more frequent in germ cell cancer (6), skin cancer (5), and primitive neuroectodermal cancer (5) (Figure 6). Some oncogenes and tumor suppressors with cancer-specific alternative splicing, such as EWSR1, CDKN1A, and GLTSCR2, are present in more types of cancer. Moreover, neither oncogenes nor tumor suppressors with cancer-specific AS were detected in brain cancer, prostate cancer, adrenal cancer, or lymphoma. This distribution bias for cancer-specific AS implies that the cancer-specific alternative splicing of both oncogenes and tumor suppressor genes is associated with specific cancer types.
Figure 6. Distribution of oncogenes and tumor suppressors with cancer-specific alternative splicing in cancer.
Blue squares indicate oncogenes, red squares indicate tumor suppressors, and yellow squares show both oncogenes and tumor suppressors.
Biological relevance of the cancer-specific transcripts in the diversification of protein functions
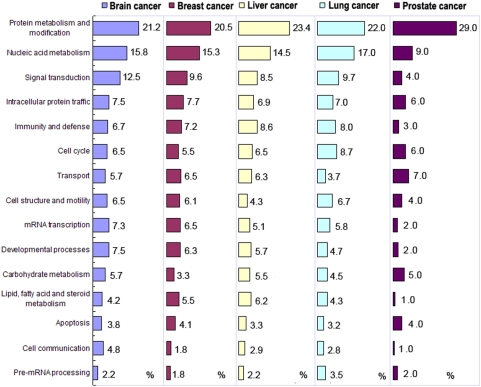
The cancer-specific transcripts were classified based on gene function by searching the RefSeq database and GO. We classified 15,093 cancer-specific transcripts from 9,989 genes into 15 function groups. Protein metabolism and modification, and nucleic acid metabolism are the most prevalent functional processes in cancer. However, the function groups of these cancer-specific transcripts differ in different cancers. For example, the least common process in breast cancer is pre-mRNA processing, whereas the function groups of cell communication and lipid, fatty acid, and steroid metabolism are seldom found in prostate cancer (Figure 7).
Figure 7. Biological processes of alternatively spliced transcripts specific to cancer.
The five cancer types are brain, breast, liver, lung, and prostate cancer. The numbers indicate the percentages for each process in the cancer. The GO process classification is based on the PANTHER (http://www.pantherdb.org/tools/genexAnalysis.jsp).
Discussion
The complexity of the transcriptome has been underestimated. In this paper, we described the transcriptome-wide identification and characterization of cancer-specific and alternatively spliced variants in human cancer based on a global view of cancer-specific alternative splicing developed by subtractive transcriptome-wide analysis. Based on an intersection/subtractive model, we have developed an analysis method for precisely screening cancer-specific alternative splicing. The EST sequences were aligned first, compared with their genomic sequences, and then mapped onto chromosomes. These procedures eliminated many EST errors, pseudogene, and multiple-copy/repeat gene problems when data were from diverse EST databases. Finally, the alternatively spliced transcripts were subject to the subtractive screening of a tissue versus all other tissues, and these analyses finally yielded cancer-specific transcripts. We identified a large number of cancer- / normal tissue-specific transcripts. Beyond all doubt, this is an abundant resource for research and the development of new diagnostic, prognostic, predictive, and therapeutic tools against human cancer. Furthermore, these resources are integrated into an available database. The HCSAS database presents a global overview of cancer-specific alternative splicing in humans and is essential for understanding tumorigenesis at a systematic level.
There are two main approaches for the global analysis of alternative splicing. First, based on the availability of sequenced genomes and large databases of sequenced transcripts (ESTs and cDNAs), alternative splicing events may be searched through reciprocal transcript alignments and alignments to genomic sequences. Several analyses in this manner have been reported [6], [23]–[29]. Because of its major limitation of EST coverage bias, a microarray-based technology has been developed to search for the alternative splicing events [3], [30]–[36]. Large sets of oligonucleotide probes may be designed specifically for individual exons and/or splice junction sequences, which allow the identification of new AS events. Here we have further developed a systematic method to search for cancer- or tissue-specific AS events in transcriptomes based on the intersection/subtractive screening analyses of transcriptomes, which is especially useful for identifying cancer/tissue-specific variants. Using this method, large numbers of cancer-specific isoforms were identified for the main human cancers. Nevertheless, these transcripts need to be further confirmed for their cancer/tissue specialization. RT-PCR technology and/or microarrays may be useful screening tools for this analysis.
Based on the transcriptome-wide analysis, we did observe special patterns of cancer-specific alternative splicing. 1) Less cancer-specific AS events occur in cancer compared to normal tissues. 2) Cancer possesses distribution bias for alternative splicing types. 3) Cancer uses rare splice sites and GT-AG more frequently, but less CT-AC compared to normal tissues. 4) The selection of splice sites differs between different kinds of cancer. 5) The cancer-specific alternative splicing of both oncogenes and tumor suppressor genes is associated with the specific cancer type. And finally, the functional groups of these cancer-specific transcripts differ in different cancers, indicating that individual cancers prefer combination controls of pathways in preference of using AS in tumorigenesis. These special features of human cancers indicate that 1) the cellular splicing machinery is changed during the transformation from normal to cancerous, 2) alternative splicing plays an important role during tumorigenesis, and 3) individual cancers have unique regulatory combinations at the alternative splicing level, which further support the prediction that approximately 60% of disease mutations in the human genome are splicing mutations [12], [13]. Our data includes the discovery of many novel splice forms of cancer-associated genes and alternative-splicing patterns in cancer, and it suggests a significant new direction for human cancer research. We strongly advise the use of cancer-specific alternative splicing as a potential source of new diagnostic, prognostic, predictive, and therapeutic tools against human cancer. The global view of cancer-specific AS is not only useful for exploring the complexity of the cancer transcriptome, but it also widens the eyeshot of clinical research.
Acknowledgments
The authors thank J.Yuan, X. Fu, Y. Sheng, and H. Qi for their data collections.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: The work was supported by the National Natural Science Foundation of China, the National Key Basic Research project (2006CB102103, 2004CB117401), the Program for New Century Excellent Talents in University, and the 111 project #B06018. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Mironov AA, Fickett JW, Gelfand MS. Frequent alternative splicing of human genes. Genome Res. 1999;9:1288–1293. doi: 10.1101/gr.9.12.1288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brett D, Hanke J, Lehmann G, Haase S, Delbruck S, et al. EST comparison indicates 38% of human mRNAs contain possible alternative splice forms. FEBS Lett. 2000;474:83–86. doi: 10.1016/s0014-5793(00)01581-7. [DOI] [PubMed] [Google Scholar]
- 3.Johnson JM, Castle J, Garrett-Engele P, Kan Z, Loerch PM, et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science. 2003;302:2141–2144. doi: 10.1126/science.1090100. [DOI] [PubMed] [Google Scholar]
- 4.Kan Z, States D, Gish W. Selecting for functional alternative splices in ESTs. Genome Res. 2002;12:1837–1845. doi: 10.1101/gr.764102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim N, Alekseyenko AV, Roy M, Lee C. The ASAP II database: analysis and comparative genomics of alternative splicing in 15 animal species. Nucleic Acids Res. 2007;35:D93–98. doi: 10.1093/nar/gkl884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Modrek B, Lee C. A genomic view of alternative splicing. Nat Genet. 2002;30:13–19. doi: 10.1038/ng0102-13. [DOI] [PubMed] [Google Scholar]
- 7.Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, et al. Alternative isoform regulation in human tissue transcriptomes. Nature. 2008;456:470–476. doi: 10.1038/nature07509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Venables JP. Unbalanced alternative splicing and its significance in cancer. Bioessays. 2006;28:378–386. doi: 10.1002/bies.20390. [DOI] [PubMed] [Google Scholar]
- 9.Garcia-Blanco MA, Baraniak AP, Lasda EL. Alternative splicing in disease and therapy. Nat Biotechnol. 2004;22:535–546. doi: 10.1038/nbt964. [DOI] [PubMed] [Google Scholar]
- 10.Faustino NA, Cooper TA. Pre-mRNA splicing and human disease. Genes Dev. 2003;17:419–437. doi: 10.1101/gad.1048803. [DOI] [PubMed] [Google Scholar]
- 11.Caceres JF, Kornblihtt AR. Alternative splicing: multiple control mechanisms and involvement in human disease. Trends Genet. 2002;18:186–193. doi: 10.1016/s0168-9525(01)02626-9. [DOI] [PubMed] [Google Scholar]
- 12.Lopez-Bigas N, Audit B, Ouzounis C, Parra G, Guigo R. Are splicing mutations the most frequent cause of hereditary disease? FEBS Lett. 2005;579:1900–1903. doi: 10.1016/j.febslet.2005.02.047. [DOI] [PubMed] [Google Scholar]
- 13.Xing Y, Lee C. Alternative splicing and RNA selection pressure–evolutionary consequences for eukaryotic genomes. Nat Rev Genet. 2006;7:499–509. doi: 10.1038/nrg1896. [DOI] [PubMed] [Google Scholar]
- 14.Kim E, Goren A, Ast G. Insights into the connection between cancer and alternative splicing. Trends Genet. 2008;24:7–10. doi: 10.1016/j.tig.2007.10.001. [DOI] [PubMed] [Google Scholar]
- 15.Wang Z, Lo HS, Yang H, Gere S, Hu Y, et al. Computational analysis and experimental validation of tumor-associated alternative RNA splicing in human cancer. Cancer Res. 2003;63:655–657. [PubMed] [Google Scholar]
- 16.Hui L, Zhang X, Wu X, Lin Z, Wang Q, et al. Identification of alternatively spliced mRNA variants related to cancers by genome-wide ESTs alignment. Oncogene. 2004;23:3013–3023. doi: 10.1038/sj.onc.1207362. [DOI] [PubMed] [Google Scholar]
- 17.Liu HX, Cartegni L, Zhang MQ, Krainer AR. A mechanism for exon skipping caused by nonsense or missense mutations in BRCA1 and other genes. Nat Genet. 2001;27:55–58. doi: 10.1038/83762. [DOI] [PubMed] [Google Scholar]
- 18.Roy M, Xu Q, Lee C. Evidence that public database records for many cancer-associated genes reflect a splice form found in tumors and lack normal splice forms. Nucleic Acids Res. 2005;33:5026–5033. doi: 10.1093/nar/gki792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Xu Q, Lee C. Discovery of novel splice forms and functional analysis of cancer-specific alternative splicing in human expressed sequences. Nucleic Acids Res. 2003;31:5635–5643. doi: 10.1093/nar/gkg786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.He C, Zuo Z, Chen H, Zhang L, Zhou F, et al. Genome-wide detection of testis- and testicular cancer-specific alternative splicing. Carcinogenesis. 2007;28:2484–2490. doi: 10.1093/carcin/bgm194. [DOI] [PubMed] [Google Scholar]
- 21.Takeda J, Suzuki Y, Nakao M, Barrero RA, Koyanagi KO, et al. Large-scale identification and characterization of alternative splicing variants of human gene transcripts using 56,419 completely sequenced and manually annotated full-length cDNAs. Nucleic Acids Res. 2006;34:3917–3928. doi: 10.1093/nar/gkl507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ladd AN, Cooper TA. Finding signals that regulate alternative splicing in the post-genomic era. Genome Biol. 2002;3:reviews0008. doi: 10.1186/gb-2002-3-11-reviews0008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lee C, Atanelov L, Modrek B, Xing Y. ASAP: the Alternative Splicing Annotation Project. Nucleic Acids Res. 2003;31:101–105. doi: 10.1093/nar/gkg029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Thanaraj TA, Stamm S, Clark F, Riethoven JJ, Le Texier V, et al. ASD: the Alternative Splicing Database. Nucleic Acids Res. 2004;32:D64–69. doi: 10.1093/nar/gkh030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zheng CL, Kwon YS, Li HR, Zhang K, Coutinho-Mansfield G, et al. MAASE: an alternative splicing database designed for supporting splicing microarray applications. Rna. 2005;11:1767–1776. doi: 10.1261/rna.2650905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gupta S, Zink D, Korn B, Vingron M, Haas SA. Genome wide identification and classification of alternative splicing based on EST data. Bioinformatics. 2004;20:2579–2585. doi: 10.1093/bioinformatics/bth288. [DOI] [PubMed] [Google Scholar]
- 27.Sharov AA, Dudekula DB, Ko MS. Genome-wide assembly and analysis of alternative transcripts in mouse. Genome Res. 2005;15:748–754. doi: 10.1101/gr.3269805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yeo GW, Van Nostrand E, Holste D, Poggio T, Burge CB. Identification and analysis of alternative splicing events conserved in human and mouse. Proc Natl Acad Sci U S A. 2005;102:2850–2855. doi: 10.1073/pnas.0409742102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Brentani H, Caballero OL, Camargo AA, da Silva AM, da Silva WA, Jr, et al. The generation and utilization of a cancer-oriented representation of the human transcriptome by using expressed sequence tags. Proc Natl Acad Sci U S A. 2003;100:13418–13423. doi: 10.1073/pnas.1233632100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Das D, Clark TA, Schweitzer A, Yamamoto M, Marr H, et al. A correlation with exon expression approach to identify cis-regulatory elements for tissue-specific alternative splicing. Nucleic Acids Res. 2007;35:4845–4857. doi: 10.1093/nar/gkm485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kwan T, Benovoy D, Dias C, Gurd S, Serre D, et al. Heritability of alternative splicing in the human genome. Genome Res. 2007;17:1210–1218. doi: 10.1101/gr.6281007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Pan Q, Shai O, Misquitta C, Zhang W, Saltzman AL, et al. Revealing global regulatory features of mammalian alternative splicing using a quantitative microarray platform. Mol Cell. 2004;16:929–941. doi: 10.1016/j.molcel.2004.12.004. [DOI] [PubMed] [Google Scholar]
- 33.Yeakley JM, Fan JB, Doucet D, Luo L, Wickham E, et al. Profiling alternative splicing on fiber-optic arrays. Nat Biotechnol. 2002;20:353–358. doi: 10.1038/nbt0402-353. [DOI] [PubMed] [Google Scholar]
- 34.Li C, Kato M, Shiue L, Shively JE, Ares M, Jr, et al. Cell type and culture condition-dependent alternative splicing in human breast cancer cells revealed by splicing-sensitive microarrays. Cancer Res. 2006;66:1990–1999. doi: 10.1158/0008-5472.CAN-05-2593. [DOI] [PubMed] [Google Scholar]
- 35.Zhang C, Li HR, Fan JB, Wang-Rodriguez J, Downs T, et al. Profiling alternatively spliced mRNA isoforms for prostate cancer classification. BMC Bioinformatics. 2006;7:202. doi: 10.1186/1471-2105-7-202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Clark TA, Schweitzer AC, Chen TX, Staples MK, Lu G, et al. Discovery of tissue-specific exons using comprehensive human exon microarrays. Genome Biol. 2007;8:R64. doi: 10.1186/gb-2007-8-4-r64. [DOI] [PMC free article] [PubMed] [Google Scholar]