Abstract
The differentiation of naive CD4 T cells into Th2 cells requires the T cell receptor-mediated activation of the ERK MAPK cascade. Little is known, however, in regard to how the ERK MAPK cascade regulates Th2 cell differentiation. We herein identified Gfi1 (growth factor independent-1) as a downstream target of the ERK MAPK cascade for Th2 cell differentiation. In the absence of Gfi1, interleukin-5 production and the change of histone modification at the interleukin-5 gene locus were severely impaired. Furthermore, the interferon γ gene showed a striking activation in the Gfi1–/– Th2 cells. An enhanced ubiquitin/proteasome-dependent degradation of GATA3 protein was observed in Gfi1–/– Th2 cells, and the overexpression of GATA3 eliminated the defect of Th2 cell function in Gfi1-deficient Th2 cells. These data suggest that the T cell receptor-mediated induction of Gfi1 controls Th2 cell differentiation through the regulation of GATA3 protein stability.
It has been established that effector T helper (Th)2 cells can be classified into at least four subsets, namely Th1, Th2, Th17, and T regulatory cells. Th1 cells produce a large amount of IFNγ and control cell-mediated immunity against intracellular pathogens, whereas Th2 cells produce IL-4, IL-5, and IL-13, and they are also involved in humoral immunity and allergic reactions (1–3). A recently identified new subset, Th17 cells, is thought to be involved in various inflammatory diseases. T regulatory cells are known to suppress various immune responses including those of autoimmune diseases (4). The direction of T helper cell differentiation depends on the types of cytokine in the environmental milieu (5). IL-12 and IFNγ induce Th1 cell differentiation, whereas IL-4 and IL-2 induce Th2 cell differentiation (1, 6, 7). The combination of IL-6 and transforming growth factor-β is required for murine Th17 cell differentiation (8–11), and the transforming growth factor-β-dependent generation of T regulatory cells has also been reported (12, 13).
In addition to the cytokines mentioned above, the activation of TCR-mediated signaling is also indispensable for Th cell differentiation. We previously reported that Th2 cell differentiation is highly dependent on the extent of TCR-mediated activation of the p56lck, calcineurin, and the Ras-ERK MAPK signaling cascade (14–16). In particular, the inhibition of the activation of the Ras-ERK MAPK cascade caused a shift from Th2 to Th1 cell differentiation, thus suggesting that the direction of Th1/Th2 cell differentiation is controlled by the TCR-mediated activation of the Ras-ERK MAPK cascade (15, 17). On the other hand, Th1 cell development appeared to be regulated by another MAPK, c-Jun N-terminal kinase (JNK) (18, 19).
Several transcription factors that govern Th2 cell differentiation have been reported. Among them, GATA3 appears to be a key factor for Th2 cell differentiation (20, 21). The up-regulation of the GATA3 mRNA expression is selectively induced in developing Th2 cells by IL-4-mediated STAT6 activation (22, 23). In addition to the transcriptional regulation, the expression of GATA3 is also regulated by a post-transcriptional mechanism. We recently reported that the Ras-ERK MAPK cascade controls GATA3 stability through the ubiquitin-proteasome-dependent pathway (24).
To identify the candidate genes that play an important role in Th2 cell differentiation and are induced by the activation of the Ras-ERK MAPK cascade, a microarray analysis was performed. We found that the expression of Gfi1 (growth factor independent-1) is dramatically up-regulated by TCR stimulation in an ERK MAPK/calcineurin activation-dependent manner. Originally, Gfi1 has been reported to play an important role in promoting cell proliferation and in preventing apoptosis (25–27). In addition, Gfi1 is involved in the self-renewal of hematopoietic stem cells (28, 29). Recently, the IL-4/STAT6-mediated induction of the Gfi1 expression in developing Th2 cells has been reported (30). Our results in this paper demonstrate that Gfi1 plays an important role in the stable expression of GATA3 protein and the subsequent differentiation of Th2 cells.
EXPERIMENTAL PROCEDURES
Mice—C57BL/6 mice and BALB/c mice were purchased from Clea. The Gfi1-deficient mice were generously provided by Dr. Stuart Orkin (Harvard Medical School, Boston, MA) (31). Gfi1-deficient mice were backcrossed more than 10 times with C57BL/6. All of the mice were maintained under specific pathogen-free conditions and then were used at 4–8 weeks of age. All of the experiments using mice received approval from the Chiba University Administrative Panel for Animal Care. All of the animal care was conducted in accordance with the guidelines of Chiba University (Chiba, Japan).
CD4 T Cell Culture and Inhibitors—Splenic CD4 T cells were prepared using a magnetic cell sorter (AutoMACS; Miltenyi Biotec) yielding a purity of >98%. Where indicated, CD4 T cells with a naive phenotype (CD44low) were purified on a FACSAria cell sorter (Becton Dickinson), yielding a purity of >98% as described previously (32). Purified CD4 T cells (1.5 × 106) were stimulated for 2 days with immobilized anti-TCR mAb (H57–597; 3 μg/ml) in the presence of IL-2 (25 units/ml), IL-12 (10 units/ml), and anti-IL-4 mAb (11B11; 25% culture supernatant) for Th1 conditions, or in the presence of IL-2 (25 units/ml), IL-4 (100 units/ml), and anti-IFNγ mAb (R4.6A2; 25% culture supernatant) for Th2 conditions. The cells were cultured for an additional 3 days in the presence of cytokines. The Th1/Th2 cell differentiation was determined by intracellular staining with anti-IL-4 and anti-IFNγ (14, 32). The production of cytokines was also determined by enzyme-linked immunosorbent assay and quantitative RT-PCR as described (33). For inhibition of TCR-mediated signals, U0126 (Promega), JNK inhibitor (Merck), SB203580 (Merck), FK506 (Sigma), cycloheximide (Merck), and MG132 (SIGMA) were used.
Expression Plasmids and Gene Transfer—The expression plasmids were transfected into 293Tcells using FuGENE reagent (Roche Applied Science) according to the manufacturer's protocol. Retrovirus vector, pMXs-IRES-hNGFR (human nerve growth factor receptor p75) was generated from the pMXs-IRES-GFP plasmid by replacing the EGFP with the cytoplasmic region-deleted hNGFR cDNA. The method for the generation of virus supernatant and the infection into developing Th2 cells was described previously (33, 34). Infected cells were detected by staining with anti-human NGFR mAb (C40-1457; BD Bioscience).
Chromatin Immunoprecipitation Assay—A ChIP assay was performed as previously described (35). Anti-acetylhistone H3-K9/14 antibody was purchased from Upstate Biotechnology, and anti-trimethylhistone H3-K4 antibody (ab8580) was from Abcam Co. (Cambridge, UK). The specific primers and TaqMan probes used in this experiments are described in the supplemental material or described previously (25, 35, 36). In the case of the semi-quantitative PCR analysis, the PCR products were resolved in an agarose gel and visualized by ethidium bromide. The images were recorded and quantified using an ATTO L&S analyzer (ATTO, Tokyo, Japan).
Immunoprecipitation and Immunoblotting—Nuclear extracts and cytoplasmic extracts were prepared using NE-PER nuclear and cytoplasmic extraction reagent (78833; Pierce). For immunoblotting, anti-GATA3 (HG3–31; Santa Cruz Biotechnology), anti-c-Maf antisera (M153; Santa Cruz Biotechnology), anti-NFAT1 mAb(4G6-G5; Santa Cruz Biotechnology), anti-tubulin-α mAb(DM1A; Lab Vision Corporation), anti-JunB mAb (C-11; Santa Cruz Biotechnology), anti-T-bet mAb (39D; Santa Cruz Biotechnology), anti-Gfi1 antisera (N20; Santa Cruz Biotechnology), anti-FLAG mAb (M2; Sigma-Aldrich), and anti-Myc tag mAb (PL14; MBL, Japan) were used. The detection of the ubiquitinated form of GATA3 was performed as described previously (24). In brief, the cells were treated with proteasome inhibitor MG132 (20 μm) for 2 h, and then the cells were lysed with radioimmunoprecipitation assay buffer (1% Nonidet P-40, 0.25% sodium deoxychorate, 150 mm NaCl, 1 mm EDTA, 1 mm phenylmethylsulfonyl fluoride, 1 μg/ml aprotinin, leupeptin, and pepstatin, 1 mm Na3VO4, 1 mm NaF, and 50 mm Tris-HCl, pH 7.4). GATA3 protein in lysates was immunoprecipitated with anti-GATA3 mAb (D16; Santa Cruz Biotechnology) and was subjected to immunoblotting with anti-multiubiquitin mAb (FK2; MBL).
Quantitative RT-PCR—Total RNA was isolated using TRIzol reagent (Invitrogen). Reverse transcription was done using Superscript II (Invitrogen). For quantitative PCR, a TaqMan universal PCR Master Mix was used for all reactions (Applied Biosystems), and the ABI Prism 7000 sequence detection system was used (Applied Biosystems). The primers and TaqMan probes for the detection of mouse GATA3, Gfi1, IL-4, IL-5, IL-13, IFNγ, hypoxanthine phosphoribosyltransferase (HPRT), and human 18 S rRNA were purchased from Applied Biosystems. The expression of mRNA was normalized using the HPRT or human 18 S rRNA signal.
Pulse-Chase Experiment—293T cells cotransfected with GATA3 and Gfi1 were washed, preincubated for 30 min in methionine/cysteine-free medium, and pulsed for 30 min with 200 μCi/ml [35S]methionine/cysteine (ICN). Then the cells were washed twice with Dulbecco's modified Eagle's medium containing nonradioactive 5 mm l-methionine, 3 mm l-cysteine, and 0.25% fetal calf serum.
RESULTS
TCR-mediated Activation of the ERK MAPK and Calcineurin Pathways Induces Gfi1 mRNA Expression—We previously reported that the TCR-mediated activation of the Ras-ERK MAPK cascade is required for Th2 cell differentiation (15, 17). To identify candidate genes, which are induced as downstream targets of the Ras-ERK MAPK and regulate Th2 cell differentiation, a DNA microarray analysis was performed. Naïve CD4 T cells were stimulated with immobilized anti-TCR mAb under Th2 conditions (IL-2, IL-4, and anti-IFNγ) for 48 h in the absence or presence of a specific inhibitor of MEK1/2, U0126. Next, total RNA was prepared and subjected to a DNA microarray analysis. We focused on the nuclear factors, and the potentially interesting 27 genes were selected (Table 1). Among the candidate genes, Gfi1 mRNA was rapidly induced after TCR stimulation in CD4 T cells, cultured under both Th1 and Th2 conditions (supplemental Fig. S1A). The STAT6-dependent induction of Gfi1 mRNA has been previously reported (30). As shown in supplemental Fig. S1B, the induction (2 h of stimulation) of Gfi1 mRNA was not dependent on STAT6; however, the reduction was more severe in STAT6-deficient cells cultured under Th2 conditions for 5 days. In contrast, the induction of GATA3 mRNA expression was completely dependent on the STAT6 activation (supplemental Fig. S1B, right panel). We confirmed the ERK MAPK-dependent induction of Gfi1 mRNA in developing Th2 cells (Fig. 1A). Splenic CD4 T cells were stimulated under Th2 conditions in the absence or presence of the indicated inhibitors for 2 h, and the expression level of Gfi1 mRNA was determined by quantitative RT-PCR. As expected, the induction of Gfi1 mRNA expression was completely blocked by the treatment with MEK inhibitor (U0126). A calcineurin inhibitor, FK506, also exhibited an inhibitory effect on Gfi1 induction. In contrast, a specific inhibitor for the p38 MAPK (SB203580) and for the JNK had no effect on the expression of Gfi1 mRNA. These results indicated that the induction of Gfi1 mRNA at the early phase of T cell activation was dependent on the ERK MAPK and the calcineurin signaling pathways.
TABLE 1.
| Gene symbol | Name of gene | U0126 treatment/no treatment (-fold increase) |
|---|---|---|
| Mybl2 | Myeloblastosis oncogene-like 2 | -7.5 |
| Gfi1 | Growth factor-independent 1 | -6.5 |
| Sap30 | sin3-associated polypeptide | -5.7 |
| Bhlhb2 (DEC1, STRA13) | Basic helix-loop-helix domain-containing, class B2 | -5.7 |
| IRF4 | Interferon regulatory factor 4 | -14.9 |
| Egr2 | Early growth response 2 | -6.5 |
| PBX1 | Pre-B cell leukemia transcription factor 1 | -6.5 |
| Id3 | Inhibitor of DNA binding 3 | 6.5 |
| IRF7 | Interferon regulatory factor 7 | 4.3 |
| Klf2 | Kruppel-like factor 2 | 39.4 |
| Klf3 | Kruppel-like factor 3 | 13.9 |
| Klf7 | Kruppel-like factor 7 | 4.3 |
| Foxp1 | Forkhead box P1 | 4.0 |
| Foxp3 | Forkhead box P3 | 4.0 |
| TCF7 | Transcription factor 7 | 5.3 |
| PIAS1 | Protein inhibitor of activated STAT1 | 4.3 |
| — | KRAB box-containing protein | 3.2 |
| Spi-B | Ets transcription factor Spi-B | 5.3 |
| DPZF | BTBPOZ zinc finger protein DPZF | 9.8 |
| Bcl11b | B cell lymphomaleukaemia 11B | 2.8 |
| Gilz | Glucocorticoid-induced leucine zipper | 22.6 |
| Asb2 | Ankyrin repeat- and SOCS box-containing protein 2 | 2.3 |
| Asb13 | Ankyrin repeat domain-containing SOCS box protein 13 | 4.3 |
| Aes | N-terminal enhancer of split | 3.0 |
| makorin | Makorin, ring finger protein, 1 | 3.5 |
| Tox | Thymus high mobility group box protein TOX | 4.0 |
| Mta3 | Metastasis-associated 3 | 4.0 |
FIGURE 1.
ERK MAPK-dependent induction of Gfi1 was required for Th2 cell differentiation. A, freshly prepared splenic CD4 T cells were stimulated under Th2 conditions in the presence of U0126 (10 μm), or JNK inhibitor (10 μm), or SB203580 (10 μm), or FK506 (100 nm) for 2 h. The expression of Gfi1 mRNA was determined by a quantitative RT-PCR analysis. The relative intensity (/HPRT) (mean of three samples) is shown with standard deviations. B, naïve (CD44low) CD4 T cells from Gfi1–/– mice were stimulated with immobilized anti-TCR mAb in the presence of IL-2 (10 units/ml), anti-IFNγ mAb, and the indicated concentrations of IL-4 for 5 days. Next, the cells were restimulated with immobilized anti-TCR mAb for 6 h and subjected to intracellular staining. The staining profiles of anti-IFNγ mAb and anti-IL-4 mAb are shown with the percentages of cells in each area. Three independent experiments were performed with similar results. C, Gfi1–/– naïve CD4 T cells were cultured with IL-4 (100 units/ml) and were restimulated with immobilized anti-TCR mAb for 24 h. The amount of cytokine (IL-4, IL-5, IL-13, and IFNγ) in the culture supernatants was determined by enzyme-linked immunosorbent assay. D, 2 h after restimulation with immobilized anti-TCR mAb, total RNA was prepared, and the expression levels IL-4, IL-5, IL-13, and IFNγ mRNA were determined by quantitative RT-PCR. The relative intensity (/HPRT) with standard deviations is shown. E, the levels of histone H3-K4 methylation at the IL-4, IL-5, IL-13, and IFNγ promoter in Gfi1–/– Th2 cells were determined by ChIP assay with quantitative PCR. The relative intensity (/Input) is shown with standard deviations.
Th2 Cell Differentiation Is Impaired in Gfi1–/– CD4 T Cells—In Gfi1–/– mice, although a reduction of T cell numbers in the lymphoid organs was seen (supplemental Fig. S2A), the cell surface expression of TCRβ, CD3ε, CD69, CD25, CD62L, IL-7Rα, IL-2Rβ, IL-4Rα, and common γ-chain (Cγ) on naïve (CD44low) Gfi1–/– splenic CD4 T cells was normal (supplemental Fig. S2B). Splenic naive CD4 T cells were purified by sorting and then were subjected to in vitro Th1/Th2 cell differentiation cultures. Under the Th2 conditions, almost equivalent numbers of IL-4-producing cells (the sum of the upper left and upper right areas) were generated both from Gfi1+/+ and Gfi1–/– naïve CD4 T cells (IL-4 0 units/ml; 3.2% versus 2.5%, 10 units/ml; 12.1% versus 16.5%, 100 units/ml; 23.4% versus 20.5%) (Fig. 1B). Interestingly, the generation of IFNγ-producing cells (the sum of the lower right and upper right areas) was markedly evaluated at all doses of IL-4 in Gfi1–/– CD4 T cell cultures (IL-4, 0 units/ml; 13.5% versus 58.5%, 10 units/ml; 9.4% versus 50.2%, 100 units/ml; 15.3% versus 42.2%) (Fig. 1B). IL-12-dependent IFNγ-producing cell (the sum of the lower right and upper right areas) generation was not enhanced in Gfi1–/– CD4 T cell under Th1 conditions (IL-12, 1 unit/ml; 69.2% versus 69.0%, 10 units/ml; 91.4% versus 90.5%), whereas the IL-12-independent generation of IFNγ-producing cells (the sum of the lower right and upper right areas) was enhanced (IL-12, 0 units/ml; 29.7% versus 48.5%) (supplemental Fig. S3A). The cytokine production from the effector Th cells was also determined by enzyme-linked immunosorbent assay (supplemental Fig. S3B). Surprisingly, the IL-5 production from Gfi1–/– Th2 cells dramatically decreased, whereas the production of IL-4 and IL-13 was equivalent in comparison with Gfi1+/+ Th2 cells (Fig. 1C). The increase in IFNγ production from Gfi1–/– Th2 cells was confirmed (Fig. 1C). A decrease in IL-5 and an increase in IFNγ mRNA expression in Gfi1–/– Th2 cells were confirmed by quantitative RT-PCR (Fig. 1D). The levels of IL-4 and IL-13 mRNA showed a modest increase.
We next assessed the histone modification status of the Th2 cytokine gene loci and the IFNγ gene locus in Gfi1–/– developing Th2 cells by ChIP assay. As expected, the trimethylation level of H3-K4 at the IL-5 promoter locus was severely impaired in Gfi1–/– Th2 cells, whereas the methylation at the IFNγ promoter was enhanced (Fig. 1E). The methylation status of histone H3-K4 at the IL-4 promoter, IL-13 promoter, and RAD50 promoter in Gfi1–/– Th2 cells was equivalent to those in the Gfi1+/+ Th2 cells (Fig. 1E). These results suggest that Gfi1 plays an important role in the acquisition of IL-5 production ability and for the suppression of the IFNγ gene in developing Th2 cells.
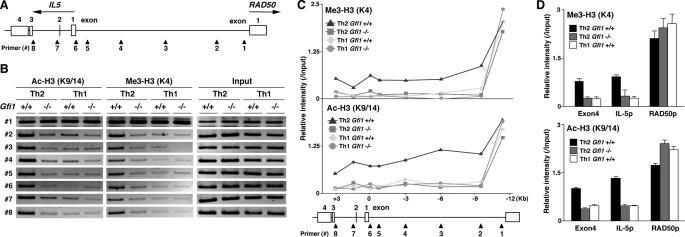
Gfi1 Is Required for the Establishment of an Active Chromatin Status at the IL-5 Gene Locus—We previously reported the induction of Th2 cell-specific hyperacetylation at the intergenic region between the IL-5 and the RAD50 gene during Th2 cell differentiation, as well as the IL-5 gene locus (37). Therefore, we analyzed the histone modification status around the IL-5 gene locus in Gfi1–/– Th2 cells more precisely. A schematic representation of the IL-5 gene locus and the location of the designed primer pairs is indicated in Fig. 2A. The actual band patterns of each ChIP assay (Fig. 2B) and the relative band intensities (Me3 H3-K4/Input DNA or Ac H3-K9/14/Input DNA) of the eight primer pairs (Fig. 2C) were depicted. As previously reported, a long range Th2-specific histone modification was observed from the upstream region of the RAD50 promoter (corresponding to primer 1) through the end of the IL-5 intron 2 (primer 8) in Gfi1+/+ Th2 cells. Nevertheless, the H3-K4 methylation and H3-K9/14 acetylation status from the primer 2 to the primer 8 regions were lower in Gfi1–/– Th2 cells, and the levels were similar to those in Th1 condition cells (Fig. 2C). The reduction of histone modifications of the IL-5 gene locus was confirmed by a ChIP assay with quantitative PCR (Fig. 2D). Neither of the modifications of the RAD50 promoter was affected (Fig. 2D). These results suggest that Gfi1 is required for the formation of active chromatin at the IL-5 gene locus.
FIGURE 2.
Histone modifications at the IL-5 gene locus in Gfi1–/– Th2 cells. A, a schematic representation of the intergenic region of the IL-5 and RAD50 locus is shown with the location of the specific primers used in B and C. B and C, histone H3-K4 trimethylation and H3-K9/14 acetylation from the promoter region of RAD50 through the end of IL-5 exon 4 in Gfi1–/– Th2 and Th1 cells was determined by ChIP assay. The bands of PCR products in B and the relative intensity for each of the primer pairs in C are indicated. Three independent experiments were performed, and similar results were obtained. D, the levels of trimethylation of H3-K4 and acetylation of H3-K9/14 at the indicated regions in Gfi1–/– Th2 cells were determined by a ChIP assay with a quantitative PCR analysis. Three independent experiments were performed with similar results.
Gfi1 Is Required for the Suppression of the IFNγ Gene Locus Activation in Developing Th2 Cells—To assess the status of histone modifications at the IFNγ locus in Gfi1–/– Th2 cells more precisely, we designed a series of primers of the IFNγ locus, including the promoter, regulatory elements, and DNase I-hypersensitive sites (Fig. 3A) and performed a ChIP assay. The actual band patterns of each ChIP assay (Fig. 3B) and the relative band intensities (Me3 H3-K4/Input DNA or Ac H3-K9/14/Input DNA) of the 14 selected primer pairs (Fig. 3C) are shown. Th1-specific hypermethylation of H3-K4 at the IFNγ locus was similarly observed both in Gfi1+/+ and in Gfi1–/– Th1 cells (Fig. 3C, upper panel). H3-K9/14 acetylation in the Gfi1–/– Th1 cells was moderately increased (Fig. 3C, lower panel). In Gfi1+/+ Th2 cells, the histone methylation and the acetylation at the IFNγ gene locus was suppressed, and those levels were significantly lower in comparison with Th1 condition cells (Fig. 3, B and C). However, in Gfi1–/– Th2 cells, high level H3-K4 methylation and H3-K9/14 acetylation were observed at the IFNγ gene locus (Fig. 3, B and C). A substantial increase in the H3-K4 methylation and H3-K9/14 acetylation at the IFNγ gene locus in Gfi1–/– Th2 cells was confirmed by a ChIP assay with quantitative PCR (Fig. 3D). These results suggest that Gfi1 plays a functional role in the IFNγ gene suppression of developing Th2 cells.
FIGURE 3.
Histone modifications at the IFNγ gene locus in Gfi1–/– Th2 cells. A, a schematic representation of the IFNγ locus is shown with the location of the specific primers used in B and C. B and C, the levels of trimethylation of histone H3-K4 and acetylation of histone H3-K9/14 at the IFNγ gene locus in Gfi1–/– Th1/Th2 cells were determined by ChIP assay. The bands of PCR product in B and the relative intensity for each of the primer pairs in C are indicated. Three independent experiments with different Th1/Th2 cell preparations were performed and obtained similar results. D, the levels of trimethylation of H3-K4 and acetylation of H3-K9/14 at the several region around the IFNγ locus in Gfi1–/– Th2 were determined by a ChIP assay with quantitative PCR as described for Fig. 2D. Three independent experiments were performed with similar results.
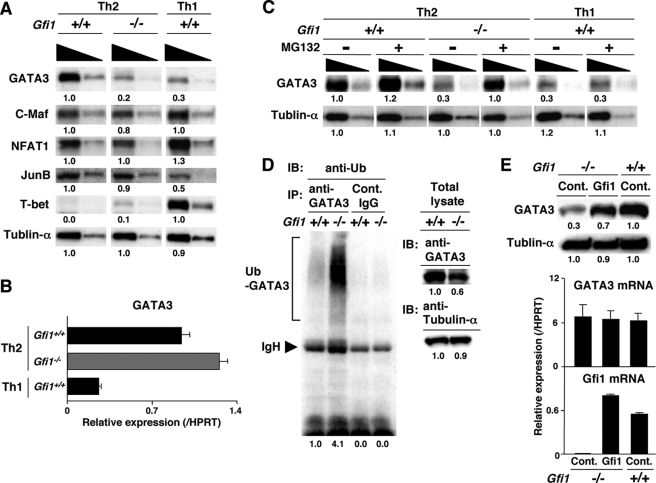
Enhanced Ubiquitin/Proteasome-dependent Degradation of GATA3 Protein in Gfi1–/– Developing Th2 Cells—GATA3 has been reported to be a critical transcriptional factor for the suppression of IFNγ production (38–40) and the activation of IL-5 transcription (41–43). Therefore, we checked the expression of GATA3 in the Gfi1–/– Th2 cells. As shown in Fig. 4A, the protein expression level of GATA3 in Gfi1–/– Th2 cells was substantially decreased, and its level was similar to that in developing Th1 cells. The expression of c-Maf, NFAT1, JunB, and T-bet proteins in Gfi1–/– Th2 cells was equivalent to that of Gfi1+/+ Th2 cells (Fig. 4A). Interestingly, the quantitative RT-PCR revealed that GATA3 mRNA was not decreased in the Gfi1–/– Th2 cells (Fig. 4B). We previously reported that GATA3 is rapidly degraded through the 26 S proteasome-dependent pathway (24). Consequently, we wanted to determine whether the 26 S proteasome pathway is involved in the reduction of GATA3 protein in Gfi1–/– Th2 cells. The Gfi1–/– Th2 cells were treated with a 26 S proteasome inhibitor, MG132, for 2 h and were subjected to Western blotting. The expression of GATA3 protein in the Gfi1–/– Th2 cells was strikingly increased by the treatment with MG132, whereas GATA3 protein in Gfi1+/+ Th2 cells showed a moderate increase (Fig. 4C). Enhanced protein expression for GATA3 was not elicited in developing Th1 cells by MG132 treatment (Fig. 4C, right panel). Collectively, these results imply that GATA3 protein rapidly degraded in Gfi1–/– Th2 cells via the 26 S proteasome-dependent pathway. Accordingly, we assessed the levels of the ubiquitinated-form of GATA3 protein in Gfi1–/– Th2 cells in the presence of MG132. The ubiquitination of GATA3 protein was dramatically increased in Gfi1–/– Th2 cells, whereas the total protein amount of GATA3 slightly decreased (Fig. 4D).
FIGURE 4.
Increased ubiquitin/proteasome-dependent degradation of GATA3 in Gfi1–/– Th2 cells. A, Gfi1+/+ and Gfi1–/– naïve CD4 T cells were cultured under Th1 or Th2 conditions for 5 days, and nuclear lysates were prepared. The nuclear lysates with a 3-fold serial dilution (1.0 × 106 and 0.3 × 106/lane) were subjected to immunoblotting with the antibodies against indicated proteins. In case of immunoblotting with anti-tubulin-α, the cytoplasmic lysates with a 3-fold serial dilution (1.0 × 105 and 0.3 × 105/lane) were used. Three experiments were performed with similar results. B, Th2 and Th1 cells were prepared as described for A. The expression levels of GATA3 mRNA were determined by a quantitative RT-PCR analysis. Three independent experiments were performed with similar results. C, Gfi1+/+ and Gfi1–/– developing Th2 cells were treated with MG132 (20 μm) for 2 h. Then the nuclear and cytoplasmic lysates were prepared, and the amount of GATA3 protein was assessed by immunoblotting. D, Gfi1+/+ and Gfi1–/– developing Th2 cells were treated with MG132 (20 μm) for 2 h, and then the ubiquitination of GATA3 was assessed. GATA3 were immunoprecipitated (IP) with anti-GATA3 mAb, and the level of ubiquitination was assessed by immunoblotting (IB) with anti-ubiquitin (Ub) mAb (left panel). The positions of migration of ubiquitinated GATA3 (Ub-GATA3) and IgH are indicated. The levels of expression of GATA3 and tubulin-α were also assessed by immunoblotting (right panel). E, naive CD4 T cells from Gfi1–/– mice were stimulated under the Th2 conditions for 2 days, and then the cells were infected with a retrovirus vector containing a Gfi1 gene (pMXs-Gfi1-IRES-hNGFR). hNGFR-positive infected cells were enriched by magnetic cell sorting, and the levels of GATA3 protein (upper panel) and mRNA (lower panel) were assessed. Two experiments were performed with similar results.
To confirm the effect of Gfi1 on the stabilization of GATA3 protein, Gfi1 was introduced into Gfi1–/– developing Th2 cells. Naive CD4 T cells from Gfi1–/– mice were stimulated with immobilized anti-TCR mAb under the Th2 conditions for 2 days, and then the cells were infected with a retrovirus vector containing Gfi1 cDNA. Three days after infection, the protein and mRNA expression levels of GATA3 were assessed. A significant increase in the level of GATA3 protein was detected in the Gfi1-introducing Gfi1–/– Th2 cells in comparison with the control mock-infected Gfi1–/– cells (Fig. 4E, upper panel). The GATA3 mRNA expression was not affected by the introduction of Gfi1 (Fig. 4E, lower panel). The enhancement of IL-5 production and the reduction of IFNγ production from Gfi1-introducing Gfi1–/– Th2 cells were confirmed by intracellular staining (supplemental Fig. S4A) and quantitative RT-PCR (supplemental Fig. S4B). These results suggest that Gfi1 stabilizes GATA3 protein through the inhibition of the ubiquitin/proteasomemediated degradation of GATA3 protein.
Enforced Expression of Gfi1 Stabilizes GATA3 Protein in 293 T Cells—To elucidate the direct stabilization of GATA3 protein by Gfi1, Myc-tagged human GATA3 (hGATA3) and FLAG-tagged Gfi1 or FLAG-tagged control vector was cotransfected into 293T cells, and then the protein expression level of GATA3 was determined. As shown in Fig. 5A, a Gfi1 dose-dependent expression of GATA3 protein (Fig. 5A, upper panel) was detected without affecting the GATA3 mRNA level (Fig. 5A, lower panel). The stabilization of GATA3 protein by Gfi1 was confirmed using hGATA3-IRES-EGFP vector, which encodes GATA3 and EGFP on the same mRNA. As expected, GATA3 protein substantially increased by the expression of Gfi1 without affecting of the green fluorescent protein level (Fig. 5B). Consequently, we studied the effect of Gfi1 expression on the degradation of GATA3 protein by a plus-chase analysis. 293T cells were transfected with GATA3 plus Gfi1 or control vector, and the amount of GATA3 protein was assessed at the indicated time points. The degradation of 35S-labeled nascent GATA3 protein in Gfi1-expressed 293T cells was slower in comparison with mock vector-introduced control cells (Fig. 5C). After a 2-h chase, 70% of the labeled GATA3 remained in the Gfi1-expressing cells, whereas the level decreased at 30% in the control cells. We next examined whether the enforced expression of Gfi1 can suppress the ubiquitination of GATA3. 293T cells were introduced with Myc-tagged hGATA3 plus FLAG-tagged Gfi1 or control FLAG vector, and the ubiquitination of GATA3 was assessed by immunoblotting with anti-ubiquitin mAb following immunoprecipitation with anti-GATA3 mAb. In the Gfi1 introduced 293T cells, the ubiquitination of GATA3 dramatically reduced (Fig. 5D, left panel), whereas the expression levels of GATA3 protein were increased (Fig. 5D, right panel). These results suggest that Gfi1 controls the protein expression of GATA3 protein through regulation of the ubiquitin/proteasome-dependent GATA3 protein.
FIGURE 5.
Gfi1 stabilizes GATA3 protein in 293 T cells. A, 293T cells were transfected with the expression plasmids for Myc-tagged human GATA3 and the indicated amounts (μg) of FLAG-tagged Gfi1. Two days after transfection, the nuclear extracts were prepared, and the levels of Myc-tagged GATA3 and FLAG-tagged Gfi1 were determined by immunoblotting with anti-Myc, and anti-FLAG mAb, respectively (upper panel). The expression levels of introduced human GATA3 mRNA were assessed by a quantitative RT-PCR analysis. The relative intensity (/HPRT) with standard deviations is shown. (lower panel). Two experiments were performed with similar results. B, 293T cells were transfected with GATA3-IRES-EGFP and FLAG-tagged Gfi1. Two days after transfection, the expression of GATA3, EGFP and Gfi1 proteins was assessed by immunoblotting. Two experiments were performed with similar results. C, the degradation of GATA3 was assessed by a pulse-chase analysis. GATA3 expression plasmid was introduced into 293T cells with control or Gfi1 expression plasmid. Two days after transfection, the cells were labeled with [35S]methionine and [35S]cysteine and chased for the indicated periods. Labeled GATA3 protein was immunoprecipitated and visualized by autoradiography. Two experiments were performed with similar results. D. The ubiquitination of GATA3 was determined by immunoblotting. 293T cells were transfected with GATA3 expression plasmid plus control or Gfi1 expression plasmid. 48 h after transfection, the cells were treated with MG132 (50 μm) for 2 h. GATA3 protein was immunoprecipitated (IP) with anti-GATA3 mAb, and the level of ubiquitination was assessed by immunoblotting (IB) with anti-Ub mAb (left panel). The positions of migration of ubiquitinated GATA3 (Ub-GATA3) and IgH are indicated. Three independent experiments were performed with similar results.
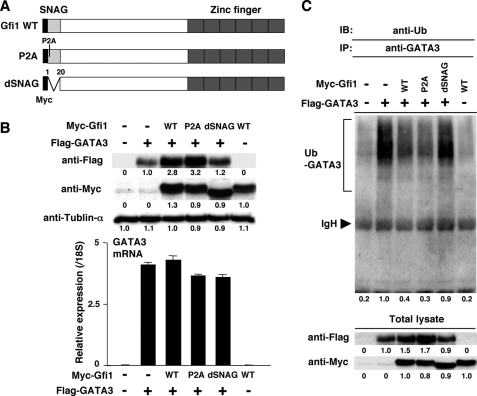
The SNAG Domain of Gfi1 Is Required for the Stabilization of GATA3 Protein—In addition to the C2H2 type zinc finger domain, Gfi1 contains a unique region called the Snail/Gfi1 (SNAG) domain. The importance of the SNAG domain in the Gfi1-mediated transcriptional repression has been reported (26). Gfi1 can form a repressor complex, which contains the CoREST corepressor, the histone demethylase LSD1, and histone deacetylases via the SNAG domain (44). The proline residue at position 2 in the SNAG domain is crucial for the Gfi1-mediated formation of the repressor complex. To assess the role of the SNAG domain for the Gfi1-mediated stabilization of GATA3 protein, a SNAG domain deletion mutant (dSNAG) and a proline substitution mutant (P2A) were generated (Fig. 6A). FLAG-tagged human GATA3 (hGATA3) and Myc-tagged WT or mutant Gfi1 was cotransfected into 293T cells, and then the protein expression level of GATA3 was determined. As shown in Fig. 6B, the dSNAG mutant failed to stabilize GATA3 protein (upper panel, fifth lane), whereas the P2A mutant succeeded to stabilize (upper panel, fourth lane) without affecting the GATA3 mRNA level (lower panel). WT and mutants Gfi1 proteins expressed at a similar level. Next, we studied the effect of Gfi1 mutants on GATA3 ubiquitination. As expected, the inhibitory function for GATA3 ubiquitination was dramatically reduced in the dSNAG mutant (lane 5), whereas the P2A mutant suppressed GATA3 ubiquitination equivalently (lane 4) in comparison with WT Gfi1 (lane 3) (Fig. 6C, upper panel). These results suggest that Gfi1 controls GATA3 protein stability through its SNAG domain. However, the repressor activity of Gfi1 is not essential for GATA3 stabilization.
FIGURE 6.
The SNAG domain of Gfi1 is required for the stabilization of GATA3 protein. A, a schematic representation of Myc-tagged Gfi1 mutants. WT Gfi1 and two mutants (P2A and dSNAG) are shown with location of Myc tag. B, 293T cells were transfected with the expression plasmids for FLAG-tagged human GATA3 plus Myc-tagged WT or mutant Gfi1. Two days after transfection, the nuclear extracts were prepared, and the levels of FLAG-tagged GATA3 and Myc-tagged Gfi1 were determined by immunoblotting with anti-FLAG, and anti-Myc mAb, respectively (upper panel). The expression levels of introduced human GATA3 mRNA were assessed by a quantitative RT-PCR analysis (lower panel). The relative intensity with standard deviations is shown. Three experiments were performed with similar results. C, the ubiquitination of GATA3 was determined by immunoblotting with anti-Ub mAb. 293T cells were transfected with FLAG-tagged GATA3 expression plasmid plus Myc-tagged WT or mutant Gfi1 expression plasmid. Two days after transfection, the cells were treated with MG132 (50 μm) for 2 h. GATA3 protein was immunoprecipitated (IP) with anti-GATA3 mAb, and the level of ubiquitination was assessed by immunoblotting (IB) with anti-Ub mAb (upper panel). The positions of migration of ubiquitinated GATA3 (Ub-GATA3) and IgH are indicated. Three independent experiments were performed with similar results.
Introduction of GATA3 Compensates Gfi1–/– Th2 Cell Phenotypes—Finally, we examined whether the introduction of GATA3 can compensate the phenotypes of Gfi1–/– Th2 cells. The GATA3 was introduced into Gfi1-deficient developing Th2 cells, and the IFNγ/IL-5 production profile was determined by intracellular staining. The number of IL-5-producing cells (the sum of the upper left and upper right areas) in Gfi1–/– Th2 cells was substantially recovered by the introduction of GATA3 (2.9% versus 10.7%) (Fig. 7A). Moreover, the generation of IFNγ-producing cells (the sum of the lower right and upper right areas) was inhibited (25.4% versus 8.9%) (Fig. 7A). The compensatory effects of GATA3 on the expression of IL-5 and IFNγ in Gfi1–/– Th2 cells were confirmed by quantitative RT-PCR (Fig. 7B). Finally, we assessed the histone modification at the IL-5 and IFNγ gene locus in GATA3-introduced Gfi1–/– Th2 cells. The trimethylation of H3-K4 at the IL-5 promoter substantially increased in Gfi1–/– Th2 cells by the introduction of GATA3 (Fig. 7C, left panel), whereas the level at the IFNγ promoter was suppressed (Fig. 7C, middle panel). The methylation at the IL-4 promoter was not affected by the introduction of GATA3 (Fig. 7C, right panel).
FIGURE 7.
Introduction of GATA3 into developing Gfi1–/– Th2 cells rendered Th2 cell function. A, naive CD4 T cells from Gfi1–/– mice were stimulated under the Th2 conditions for 2 days, and then the cells were infected with a retrovirus vector containing a GATA3 (pMXs-GATA3-IRES-hNGFR). Three days after infection, the IFNγ/IL-5 staining profile of GATA3-infected cells was determined by intracellular staining. The percentages of cells in each quadrant are indicated. Three independent experiments were performed with similar results. B, hNGFR-positive infected cells were enriched as described in Fig. 4E and restimulated by anti-TCR mAb for 2 h. The expression of each cytokine mRNA was assessed by quantitative RT-PCR. C, the levels of trimethylation of H3-K4 were determined by a ChIP assay with a quantitative PCR as described in Fig. 1E. Three independent experiments were performed with similar results.
DISCUSSION
In this report, we demonstrate that Gfi1 plays an important role in the regulation of IL-5 and IFNγ production in Th2 cells. Gfi1 appears to control these Th2 functions through the regulation of GATA3 protein stability via the repression of ubiquitin/proteasome-dependent GATA3 degradation. Therefore, the cooperation of Gfi1 with GATA3 is required for the proper Th2 cell differentiation.
We previously reported that the activation of the Ras-ERK MAPK cascade inhibits the ubiquitin/proteasome-dependent degradation of GATA3 (24). In the present study, we found that Gfi1 is rapidly induced after TCR-mediated activation in an ERK MAPK/calcineurin-dependent manner (Fig. 1A) and suppressed the degradation of GATA3 protein (Figs. 4, 5, 6). These results raise the possibility that ERK MAPK regulates GATA3 stability through the induction of Gfi1 expression. Gfi1 has been reported to be associated with PIAS3, a small ubiquitin-like modifier (SUMO) ligase, and inhibits the binding with STAT3 (45). STAT3 could escape from PIAS-mediated sumoylation and increase transcriptional activity. We previously reported that Mdm2 binds to GATA3 and acts as an ubiquitin ligase for GATA3 (24). As in the case of PIAS3, it is possible that Gfi1 binds to Mdm2 and sequesters its E3 ligase activity for GATA3. In fact, the association of Gfi1 with Mdm2 was detected in the transfected 293T cells and Th2 cells.3 Furthermore, the ubiquitination of Gfi1 protein has also been reported (46, 47). Although further studies are required to elucidate the precise mechanism of Gfi1-mediated inhibition in the GATA3 ubiquitination, it is likely that Gfi1 regulates GATA3 protein stability through the repression of ubiquitination.
Gfi1 acts as a transcriptional repressor through a unique N-terminal SNAG domain (26). The proline residue at position 2 was crucial for repressor activity of Gfi1, because the substitution of proline to alanine (Gfi1 P2A mutant) abolished the transcriptional repressor ability. Although the SNAG domain of Gfi1 was required for the stabilization of GATA3 protein, Gfi1 P2A mutant could suppress GATA3 ubiquitination and stabilize GATA3 protein (Fig. 6, B and C). These results demonstrated that the repressor activity of Gfi1 was not essential for GATA3 stabilization and that the SNAG domain might has a distinct function from transcriptional repression.
In this report, we mostly analyzed the Gfi1 effects on the GATA3 stabilization using 293 T cells. Because Gfi1-mediated stabilization of GATA3 was observed in 293T cells as well as Th2 cells (Figs. 4, 5, 6), it is likely that Gfi1 stabilized GATA3 protein in Th2 cells through the similar machinery that would operate in 293T cells. However, GATA3 and Gfi1 were not expressed endogenously in 293T cells. The identification of a Gfi1 target ubiquitin E3 ligase may help us to better understand the molecular mechanism controlling GATA3 degradation.
It was previously reported that Gfi1 is required for IL-2-dependent expansion of GATA3high Th2 cells (48). In our experiment, the mRNA expression level of GATA3 was not changed in expanded Gfi1–/– developing Th2 cells (Fig. 4B). Retrovirus-mediated introduction of Gfi1 into Gfi1–/– developing Th2 cells was sufficient to induce GATA3 protein expression (Fig. 4E). Furthermore, the IL-2-dependent expression of GATA3 protein was impaired in Gfi1–/– effector Th2 cells.3 Therefore, the reduced GATA3high cell number in Gfi1–/– Th2 cells might be also due to the enhanced degradation of GATA3 protein.
In Gfi1–/– developing Th2 cells, the production of IL-5 was severely decreased, whereas IL-4 and IL-13 production was not affected (Fig. 1, C and D). The methylation status of histone H3-K4 and the acetylation status of histone H3-K9/14 at the IL-5 gene locus were significantly decreased in developing Gfi1–/– Th2 cells (Figs. 1E and 2). Again, histone modifications at the IL-13/IL-4 gene loci were not affected (Fig. 1E). Although the induction of H3-K4 methylation and H3-K9/14 acetylation at the IL-4, IL-5, and IL-13 gene loci was dependent on GATA3 expression (34, 35, 37, 49), the higher expression level of GATA3 is required for the induction of IL-5 gene activation in comparison with those of the IL-4/IL-13 gene (37). A different dependence on GATA3 expression level may explain the difference of histone modifications (H3-K4 methylation and H3-K9/14 acetylation) in Gfi1–/– Th2 cells between the IL-4/IL-13 gene loci and the IL-5 gene locus.
Enhanced production of IFNγ was observed in Gfi1–/– cells cultured under the Th2 condition and under the Th1 condition with low concentration of IL-12 (supplemental Fig. S3). Furthermore, the levels of histone H3-K9/14 acetylation and of H3-K4 methylation at the IFNγ gene locus was dramatically increased in the Gfi1–/– Th2 cells (Fig. 3). For the repression of the IFNγ gene activation including the suppression of H3-K4 methylation and H3-K9/14 acetylation, the expression of GATA3 is required (34, 38–40). In addition, the expression of the T-box type transcription factor, T-bet promotes IFNγ production in Th1 cells (50). Although the level of T-bet was increased slightly in the Gfi1–/– Th2 cells, the level was quite low in comparison with that in the Th1 cells (Fig. 4A). Therefore, the de-repression status of the IFNγ gene locus in Gfi1–/– Th2 cells appeared to be due to the decreased level of GATA3 protein.
Gfi1 contains six C2H2 zinc fingers and is able to bind to specific DNA target sequence (51). The consensus recognition sequence, determined by selection for Gfi1 binding from a random DNA sequence library, is TAAA(T/G)CAC(A/T)GCA. We identified a potential Gfi1-binding site within the promoter region of the IL-13 gene, whereas there are no potential binding sites for Gfi1 in other regulatory regions of the Th2 cytokine gene locus (IL-4 promoter, IL-5 promoter, CGRE, CNS1, and VA enhancer) and the IFNγ gene locus (IFNγ promoter, CNS1, and CNS2). We also performed a ChIP assay and confirmed the Gfi1 binding within the Th2 cytokine gene locus and the IFNγ gene locus. As expected, the binding of Gfi1 was only detected at the IL-13 promoter.3 Therefore, it is unlikely that Gfi1 directly regulates IL-5 and IFNγ production in Th2 cells.
Although the expression of Gfi1 mRNA was equivalently induced in CD4 T cells cultured under both Th1 and Th2 conditions, its expression was preferentially maintained in developing Th2 cells.3 STAT6-deficient CD4 T cells cultured under Th2 conditions also failed to maintain Gfi1 mRNA expression (supplemental Fig. S1B). These results are consistent with previous data, in which Gfi1 expression is Th2 cell-specific and STAT6-dependent (30). Th2 cell-specific maintenance of Gfi1 expression may contribute to the Th2 cell-specific stabilization of GATA3 protein. In addition, our preliminary results indicate that the introduction of GATA3 can induce Gfi1 mRNA expression in developing Th1 cells.3 These data raise the possibility that GATA3 and Gfi1 make a positive feedback loop for stable expression of both genes and control Th2 cell differentiation. In summary, the results of this study indicate that Gfi1 is a downstream target of the ERK MAPK cascade, and it plays an important role in the regulation of the GATA3 protein expression and Th2 cell functions.
Supplementary Material
Acknowledgments
We thank Toshihiro Ito, Hikari Asou, Satoko Norikane, and Kaoru Sugaya for excellent technical assistance.
This work was supported by Ministry of Education, Culture, Sports, Science and Technology Grants-in-Aid for Scientific Research on Priority Areas 20060003, Scientific Research (B) 17390139, Scientific Research (C) 20590485, Exploratory Research 19659121, and Young Scientists (Start-up) 19890041; by Special Coordination Funds for Promoting Science and Technology; and by the Ministry of Health, Labor and Welfar, the Kanae Foundation, the Uehara Memorial Foundation, the Astellas Foundation, and the Sagawa Foundation. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact.
The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S4 and supplemental text.
Footnotes
The abbreviations used are: Th, T helper; IL, interleukin; IFN, interferon; ERK, extracellular signal-regulated kinase; MAPK, mitogen-activated protein kinase; JNK, c-Jun N-terminal kinase; STAT, signal transducers and activators of transcription; mAb, monoclonal antibody; RT, reverse transcription; EGFP, enhanced green fluorescent protein; ChIP, chromatin immunoprecipitation; HPRT, hypoxanthine phosphoribosyltransferase; MEK, MAPK/ERK kinase; E3, ubiquitin-protein isopeptide ligase; WT, wild type; NGFR, nerve growth factor receptor; TCR, T cell receptor.
M. Yamashita and R. Shinnakasu, unpublished observation.
References
- 1.Constant, S. L., and Bottomly, K. (1997) Annu. Rev. Immunol. 15 297–322 [DOI] [PubMed] [Google Scholar]
- 2.Seder, R. A., and Paul, W. E. (1994) Annu. Rev. Immunol. 12 635–673 [DOI] [PubMed] [Google Scholar]
- 3.Reiner, S. L., and Locksley, R. M. (1995) Annu. Rev. Immunol. 13 151–177 [DOI] [PubMed] [Google Scholar]
- 4.Stockinger, B., and Veldhoen, M. (2007) Curr. Opin. Immunol. 19 281–286 [DOI] [PubMed] [Google Scholar]
- 5.Bettelli, E., Oukka, M., and Kuchroo, V. K. (2007) Nat. Immun. 8 345–350 [DOI] [PubMed] [Google Scholar]
- 6.Abbas, A. K., Murphy, K. M., and Sher, A. (1996) Nature 383 787–793 [DOI] [PubMed] [Google Scholar]
- 7.O'Garra, A. (1998) Immunity 8 275–283 [DOI] [PubMed] [Google Scholar]
- 8.Ivanov, II, McKenzie, B. S., Zhou, L., Tadokoro, C. E., Lepelley, A., Lafaille, J. J., Cua, D. J., and Littman, D. R. (2006) Cell 126 1121–1133 [DOI] [PubMed] [Google Scholar]
- 9.Bettelli, E., Carrier, Y., Gao, W., Korn, T., Strom, T. B., Oukka, M., Weiner, H. L., and Kuchroo, V. K. (2006) Nature 441 235–238 [DOI] [PubMed] [Google Scholar]
- 10.Mangan, P. R., Harrington, L. E., O'Quinn, D. B., Helms, W. S., Bullard, D. C., Elson, C. O., Hatton, R. D., Wahl, S. M., Schoeb, T. R., and Weaver, C. T. (2006) Nature 441 231–234 [DOI] [PubMed] [Google Scholar]
- 11.Veldhoen, M., Hocking, R. J., Atkins, C. J., Locksley, R. M., and Stockinger, B. (2006) Immunity 24 179–189 [DOI] [PubMed] [Google Scholar]
- 12.Shevach, E. M. (2006) Immunity 25 195–201 [DOI] [PubMed] [Google Scholar]
- 13.Wan, Y. Y., and Flavell, R. A. (2006) Immunol. Rev. 212 114–130 [DOI] [PubMed] [Google Scholar]
- 14.Yamashita, M., Katsumata, M., Iwashima, M., Kimura, M., Shimizu, C., Kamata, T., Shin, T., Seki, N., Suzuki, S., Taniguchi, M., and Nakayama, T. (2000) J. Exp. Med. 191 1869–1879 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yamashita, M., Kimura, M., Kubo, M., Shimizu, C., Tada, T., Perlmutter, R. M., and Nakayama, T. (1999) Proc. Natl. Acad. Sci. U. S. A. 96 1024–1029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Yamashita, M., Hashimoto, K., Kimura, M., Kubo, M., Tada, T., and Nakayama, T. (1998) Int. Immunol. 10 577–591 [DOI] [PubMed] [Google Scholar]
- 17.Shibata, Y., Kamata, T., Kimura, M., Yamashita, M., Wang, C. R., Murata, K., Miyazaki, M., Taniguchi, M., Watanabe, N., and Nakayama, T. (2002) J. Immunol. 169 2134–2140 [DOI] [PubMed] [Google Scholar]
- 18.Dong, C., Yang, D. D., Wysk, M., Whitmarsh, A. J., Davis, R. J., and Flavell, R. A. (1998) Science 282 2092–2095 [DOI] [PubMed] [Google Scholar]
- 19.Yang, D. D., Conze, D., Whitmarsh, A. J., Barrett, T., Davis, R. J., Rincon, M., and Flavell, R. A. (1998) Immunity 9 575–585 [DOI] [PubMed] [Google Scholar]
- 20.Zhang, D. H., Cohn, L., Ray, P., Bottomly, K., and Ray, A. (1997) J. Biol. Chem. 272 21597–21603 [DOI] [PubMed] [Google Scholar]
- 21.Zheng, W., and Flavell, R. A. (1997) Cell 89 587–596 [DOI] [PubMed] [Google Scholar]
- 22.Murphy, K. M., Ouyang, W., Farrar, J. D., Yang, J., Ranganath, S., Asnagli, H., Afkarian, M., and Murphy, T. L. (2000) Annu. Rev. Immunol. 18 451–494 [DOI] [PubMed] [Google Scholar]
- 23.Nelms, K., Keegan, A. D., Zamorano, J., Ryan, J. J., and Paul, W. E. (1999) Annu. Rev. Immunol. 17 701–738 [DOI] [PubMed] [Google Scholar]
- 24.Yamashita, M., Shinnakasu, R., Asou, H., Kimura, M., Hasegawa, A., Hashimoto, K., Hatano, N., Ogata, M., and Nakayama, T. (2005) J. Biol. Chem. 280 29409–29419 [DOI] [PubMed] [Google Scholar]
- 25.Karsunky, H., Mende, I., Schmidt, T., and Moroy, T. (2002) Oncogene 21 1571–1579 [DOI] [PubMed] [Google Scholar]
- 26.Grimes, H. L., Chan, T. O., Zweidler-McKay, P. A., Tong, B., and Tsichlis, P. N. (1996) Mol. Cell. Biol. 16 6263–6272 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gilks, C. B., Bear, S. E., Grimes, H. L., and Tsichlis, P. N. (1993) Mol. Cell. Biol. 13 1759–1768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hock, H., Hamblen, M. J., Rooke, H. M., Schindler, J. W., Saleque, S., Fujiwara, Y., and Orkin, S. H. (2004) Nature 431 1002–1007 [DOI] [PubMed] [Google Scholar]
- 29.Zeng, H., Yucel, R., Kosan, C., Klein-Hitpass, L., and Moroy, T. (2004) EMBO J. 23 4116–4125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhu, J., Guo, L., Min, B., Watson, C. J., Hu-Li, J., Young, H. A., Tsichlis, P. N., and Paul, W. E. (2002) Immunity 16 733–744 [DOI] [PubMed] [Google Scholar]
- 31.Hock, H., Hamblen, M. J., Rooke, H. M., Traver, D., Bronson, R. T., Cameron, S., and Orkin, S. H. (2003) Immunity 18 109–120 [DOI] [PubMed] [Google Scholar]
- 32.Kimura, M. Y., Hosokawa, H., Yamashita, M., Hasegawa, A., Iwamura, C., Watarai, H., Taniguchi, M., Takagi, T., Ishii, S., and Nakayama, T. (2005) J. Exp. Med. 201 397–408 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kimura, M., Koseki, Y., Yamashita, M., Watanabe, N., Shimizu, C., Katsumoto, T., Kitamura, T., Taniguchi, M., Koseki, H., and Nakayama, T. (2001) Immunity 15 275–287 [DOI] [PubMed] [Google Scholar]
- 34.Shinnakasu, R., Yamashita, M., Shinoda, K., Endo, Y., Hosokawa, H., Hasegawa, A., Ikemizu, S., and Nakayama, T. (2006) J. Immunol. 177 5801–5810 [DOI] [PubMed] [Google Scholar]
- 35.Yamashita, M., Ukai-Tadenuma, M., Kimura, M., Omori, M., Inami, M., Taniguchi, M., and Nakayama, T. (2002) J. Biol. Chem. 277 42399–42408 [DOI] [PubMed] [Google Scholar]
- 36.Yamashita, M., Hirahara, K., Shinnakasu, R., Hosokawa, H., Norikane, S., Kimura, M. Y., Hasegawa, A., and Nakayama, T. (2006) Immunity 24 611–622 [DOI] [PubMed] [Google Scholar]
- 37.Inami, M., Yamashita, M., Tenda, Y., Hasegawa, A., Kimura, M., Hashimoto, K., Seki, N., Taniguchi, M., and Nakayama, T. (2004) J. Biol. Chem. 279 23123–23133 [DOI] [PubMed] [Google Scholar]
- 38.Ouyang, W., Ranganath, S. H., Weindel, K., Bhattacharya, D., Murphy, T. L., Sha, W. C., and Murphy, K. M. (1998) Immunity 9 745–755 [DOI] [PubMed] [Google Scholar]
- 39.Kaminuma, O., Kitamura, F., Kitamura, N., Miyagishi, M., Taira, K., Yamamoto, K., Miura, O., and Miyatake, S. (2004) FEBS Lett. 570 63–68 [DOI] [PubMed] [Google Scholar]
- 40.Usui, T., Nishikomori, R., Kitani, A., and Strober, W. (2003) Immunity 18 415–428 [DOI] [PubMed] [Google Scholar]
- 41.Lee, H. J., O'Garra, A., Arai, K., and Arai, N. (1998) J. Immunol. 160 2343–2352 [PubMed] [Google Scholar]
- 42.Zhang, D. H., Yang, L., and Ray, A. (1998) J. Immunol. 161 3817–3821 [PubMed] [Google Scholar]
- 43.Schwenger, G. T., Fournier, R., Kok, C. C., Mordvinov, V. A., Yeoman, D., and Sanderson, C. J. (2001) J. Biol. Chem. 276 48502–48509 [DOI] [PubMed] [Google Scholar]
- 44.Saleque, S., Kim, J., Rooke, H. M., and Orkin, S. H. (2007) Mol. cell 27 562–572 [DOI] [PubMed] [Google Scholar]
- 45.Rodel, B., Tavassoli, K., Karsunky, H., Schmidt, T., Bachmann, M., Schaper, F., Heinrich, P., Shuai, K., Elsasser, H. P., and Moroy, T. (2000) EMBO J. 19 5845–5855 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Marteijn, J. A., van der Meer, L. T., van Emst, L., van Reijmersdal, S., Wissink, W., de Witte, T., Jansen, J. H., and Van der Reijden, B. A. (2007) Blood 110 3128–3135 [DOI] [PubMed] [Google Scholar]
- 47.Marteijn, J. A., van der Meer, L. T., Van Emst, L., de Witte, T., Jansen, J. H., and van der Reijden, B. A. (2007) Blood 109 100–108 [DOI] [PubMed] [Google Scholar]
- 48.Zhu, J., Jankovic, D., Grinberg, A., Guo, L., and Paul, W. E. (2006) Proc. Natl. Acad. Sci. U. S. A. 103 18214–18219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Omori, M., Yamashita, M., Inami, M., Ukai-Tadenuma, M., Kimura, M., Nigo, Y., Hosokawa, H., Hasegawa, A., Taniguchi, M., and Nakayama, T. (2003) Immunity 19 281–294 [DOI] [PubMed] [Google Scholar]
- 50.Szabo, S. J., Sullivan, B. M., Stemmann, C., Satoskar, A. R., Sleckman, B. P., and Glimcher, L. H. (2002) Science 295 338–342 [DOI] [PubMed] [Google Scholar]
- 51.Zweidler-Mckay, P. A., Grimes, H. L., Flubacher, M. M., and Tsichlis, P. N. (1996) Mol. Cell. Biol. 16 4024–4034 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.