Abstract
The α-globin poly(C)-binding proteins (αCPs) comprise an abundant and widely expressed set of K-homolog domain RNA-binding proteins. αCPs regulate the expression of a number of cellular and viral mRNAs at the levels of splicing, stability, and translation. Previous surveys have identified 160 mRNAs that are bound by αCP in the human hematopoietic cell line, K562. To explore the functions of these αCP/mRNA interactions, we identified mRNAs whose levels are altered in K562 cells acutely depleted of the two major αCP proteins, αCP1 and αCP2. Microarray analysis identified 27 mRNAs that are down-regulated and 14 mRNAs that are up-regulated in the αCP1/2-co-depleted cells. This αCP1/2 co-depletion was also noted to inhibit cell proliferation and trigger a G1 cell cycle arrest. Targeted analysis of genes involved in cell cycle control revealed a marked increase in p21WAF mRNA and protein. Analysis of mRNP complexes in K562 cells demonstrates in vivo association of p21WAF mRNA with αCP1 and αCP2. In vitro binding assays indicate that a 127-nucleotide region of the 3′-untranslated region of p21WAF interacts with both αCP1 and αCP2, and co-depletion of αCP1/2 results in a marked increase in p21WAF mRNA half-life. p21WAF induction and G1 arrest in the αCP1/2-co-depleted cells occur in the absence of p53 and are not observed in cells depleted of the individual αCP isoforms. The apparent redundancy in the actions of αCP1 and αCP2 upon p21WAF expression correlates with a parallel redundancy in their effects on cell cycle control. These data reveal a pivotal role for αCP1 and αCP2 in a p53-independent pathway of p21WAF control and cell cycle progression.
αCPs,2 also known as heterogeneous nuclear ribonucleoprotein (hnRNP) E (1) or poly(C)-binding proteins (2–4), comprise a family of highly abundant and widely expressed RNA-binding proteins. There are four αCP loci (1, 5, 6, 7), encoding αCP1–αCP4. Two major products of the αCP2 locus, αCP2 and αCP2KL, arise by alternative splicing (8), and a third abundant paralog, αCP1, is encoded from a retrotransposed copy of a fully processed αCP2 transcript (5). αCPs are highly conserved in evolution; orthologs are encoded in the genomes of Xenopus laevis, Drosophila melanogaster, Caenorhabditis elegans, and Saccharomyces cervisiae (6). The abundant expression, widespread tissue distribution (1, 4, 5), and evolutionary conservation of αCPs suggest that they serve a basic cellular function(s).
Each αCP isoform contains three copies of the hnRNP K homology RNA binding domain (9). αCPs, along with hnRNP K, are uniquely characterized by in their strong binding preference for C-rich motifs. This subset of hnRNP K homology domain proteins has been linked to post-transcriptional controls via binding to elements in 5′- and 3′-untranslated regions (UTRs) of cellular and viral mRNAs (10–19). For example, αCP1 and/or αCP2 regulate the stability of the mRNAs encoding α2-globin, tyrosine hydroxylase, and α1(I) collagen via binding to 3′-UTR motifs and mediate control over the translation of specific mRNAs, including 15-lipoxygenase, CCAAT/enhancer-binding protein α, folate receptor, and phosphatase 2A, by binding to either 5′-or3′-UTR elements. In addition to regulating the expression of several cellular mRNAs, αCP can also regulate a number of distinct steps in viral gene expression (11, 20–29). Taken together, these studies indicate that αCPs constitute key regulators in a wide spectrum of post-transcriptional controls.
To develop an understanding of how αCPs impact on cell function, we have screened for in vivo binding targets. Microarray analysis of immunoenriched αCP2-mRNP complexes isolated from K562 cells (30) revealed 160 αCP2-associated mRNAs. These mRNAs could be clustered according to the function(s) of their encoded proteins, suggesting roles for αCP2 in coordination of post-transcriptional controls. One of the larger functional clusters consisted of mRNAs that affect cell growth and proliferation. A role for αCP2 in cell cycle control was consistent with prior observations that a member of the αCP family, αCP4, can induce cell cycle arrest at G2-M and stimulate apoptosis (31, 32).
The current study was initiated to assign functions to αCP interactions with cellular mRNAs (30). To accomplish this goal, we acutely depleted K562 cells of αCP1 and αCP2, either separately or together, and identified mRNAs that were either induced or repressed in their steady state levels. During the course of these studies, we observed that the αCP1/2 co-depletion decreased cell proliferation and triggered a G1 arrest. The basis of the mitotic arrest was explored by determining the effect of the αCP1/2 co-depletion on the expression of genes that play pivotal roles in cell cycle control. These studies revealed an induction of the cyclin-dependent kinase inhibitor 1A (CDKN1A) mRNA and protein. CDKN1A is also known as wild-type p53 activated fragment (p21WAF), and we will use this designation throughout. The induction of p21WAF mRNA and protein correlated with the G1 arrest. p21WAF mRNA was found to be associated with both αCP1 and αCP2 mRNP complexes in untreated cells, and the induction of p21WAF mRNA subsequent to αCP1/2 co-depletion was mechanistically linked to prolongation of the p21WAF mRNA half-life. These data lead us to conclude that αCP1 and αCP2 play a role in cell cycle control via a p53-independent, post-transcriptional modulation of p21WAF gene expression.
EXPERIMENTAL PROCEDURES
Cell Growth and siRNA Transfection—K562 cells (ATCC number CCL-243) were propagated in RPMI 1640 supplemented with 10% fetal bovine serum (HyClone) and antibiotic/antimycotic (Invitrogen) under standard conditions. Cells were transfected with a total of 20 μg of siRNA using Nucleofector V (Amaxa). The following siRNAs were used: αCP1, AAGGGAGAGTCATGACCATTC (Ambion); αCP2, AAGGAUCUACUGAUAGGCAGG; lamin A/C, AACUGGACUUCCAGAAGAACA (Dharmacon). In experiments in which αCP1 or αCP2 siRNAs were used individually, the αCP siRNA was supplemented with lamin A/C siRNA to bring the total siRNA content to 20 μg.
Western Blot Analysis—Radioimmune precipitation assay buffer (1% IGEPAL CA-630, 0.5% sodium deoxycholate, 0.1% SDS in phosphate-buffered saline) lysates were isolated (Santa Cruz Biotechnology, Inc., Santa Cruz, CA), and protein was quantified using the Bio-Rad D/C kit. Extracts prepared from HCT116 cells that had been treated with 50 μm irinotecan for 24 h were provided (kind gift of N. Finnberg; W. El-Deiry laboratory, University of Pennsylvania). αCP1, αCP2, and rpL7 antibodies were generated by our laboratory; antibodies to p53 and p21WAF were purchased from Santa Cruz Biotechnology; and lamin A/C, CCNH, and RB antibodies were purchased from Cell Signaling Technology. Antibody-bound proteins were visualized by Western analysis using ECL Plus (Amersham Biosciences).
RNA Isolation and Microarray Analysis—Analysis of a G4112A hybridization microarray (Agilent) representing 25,584 human genes was performed by Mogene using a 2-μg aliquot of total RNA isolated from cells 2 days post-siRNA transfection (RNAeasy; Qiagen).
qRT-PCR Analysis—cDNA was synthesized from total RNA (1 μg) (Reaction Ready First Strand cDNA Synthesis; SuperArray), and the cDNA product, diluted 13-fold with H2O, was used as a template for quantitative PCR (RT2 Sybr Green/Rox; SuperArray or Taqman reagents; ABI). The following sets of primers were purchased from ABI: FADS1 (fatty acid desaturase 1) (Hs00203685_m1), RASSF5 (Ras association domain family protein 5) (Hs00739100_m1), and RIG-I (Hs00204833_m1). The following sets of primers were purchased from SuperArray: CCNH (PPH00969A), HIF1α (PPH01361A), PHGDH (PPH07199A), and UBCH2 (PPH18206A). Reactions were run in triplicate on an ABI Prism 7700, and the data were analyzed using Sequence Detection Software version 1.9.1. The Ct values obtained are an average of the triplicates.
Analysis of the Cell Cycle and Viability—Fluorescence-activated cell sorting analysis was performed on cells 4 and 5 days post-siRNA transfection (Easycyte Mini; Guava). Cell cycle reagent (containing propidium iodide) was used to analyze the cell cycle, and a minimum of 2000 cells were analyzed according to the manufacturer's protocol (Guava). The Viacount Dye Exclusion Assay was used to measure viability (Guava).
RNA Co-purification—mRNA content in the immunoenriched RNP preparations was determined by RT-PCR as described (30). p21WAF mRNA (33) and γ-globin mRNA (30) were amplified by RT-PCR as described.
mRNA Half-life Determination—Actinomycin D (Sigma) was added to the media (5 μg/ml) at 2 days post-siRNA transfection and total RNA was collected at subsequent 0, 2, and 4 h time points. 4-μg samples were analyzed by Northern blotting (34) using a p21WAF cDNA probe (Origene) labeled with 32P (RadPrime DNA Labeling Kit; Invitrogen). Band intensities were quantified on a Storm PhosphorImager (Amersham Biosciences).
Cross-linking Immunoprecipitation Analysis—Plasmids containing a T7 promoter upstream of various regions corresponding to the p21WAF 3′-UTR were a kind gift of P. Leedman (University of Western Australia) (33) and are depicted in Fig. 7A. These plasmids were linearized with HindIII prior to transcription. The p21WAF 3′-UTR regions designated WAF1-A to WAF1-E (Fig. 7A) were amplified by PCR. The primers are indicated in Table 4. Note that each forward primer contained a T7 promoter (TAATACGACTCACTATAGG) at its 5′-end, which is not included in the primer sequences listed in Table 4. The p21WAF cDNA (Origene) was used as a template for the PCR. PCR was performed using the Platinum Pfx DNA polymerase (Invitrogen) according to the manufacturer's instructions except that we used 2× Pfx amplification buffer, 0.2 μg of template, 0.4 μl of Pfx DNA polymerase, and 1× PCRx enhancer per reaction. The conditions were 94 °C for 5 min; 30 cycles of 94 °C for 15 s, 55 °C for 30 s, 68 °C for 84 s; and 68 °C for 7 min. Fragments were gel-isolated using the QIAquick Gel Extraction Kit (Qiagen). Linearized plasmids or PCR products were used as templates for transcription of radiolabeled thiolated RNAs as described (30). These RNAs contained thiolated uridines, which allow for cross-linking of the thiol group to a binding protein located within a few Å of the thiol moiety. The RNAs were incubated with cytoplasmic extract from K562 cells and irradiated at 312 nm to activate the protein/RNA cross-link. Following irradiation, the samples were treated with RNase A to remove the unprotected RNA. The samples were then immunoprecipitated with antibodies specific to αCP1 or αCP2 (both generated by our laboratory) or c-Myc antibodies (Santa Cruz Biotechnology) and analyzed by SDS-PAGE as described (30).
FIGURE 7.
αCP1 and αCP2 bind to a 127-nucleotide fragment of the p21WAF 3′-UTR. A, fragments of the p21WAF 3′-UTR used in the initial cross-linking assay (33) and sequences corresponding to subfragments of WAF 1–879 used in higher resolution mapping. B, UV cross-linking assay of fragments of the p21WAF 3′-UTR. Cytoplasmic extracts from K562 cells were incubated with thiolated, 32P-labeled RNA sequences representing different regions of the p21WAF 3′-UTR. The mixture was cross-linked, digested with RNase, and immunoprecipitated (IP) with antibodies directed against c-Myc (Control), αCP1, or αCP2. The resulting complexes were analyzed by SDS-PAGE. Molecular weight markers are shown on the right. The top panel shows the cross-linking results with the RNAs described in the first set of fragments shown in A. An actin antisense RNA (Actin AS) was utilized as a negative control, and the α-globin 3′-UTR (α-globin) was utilized as a positive control. The bottom panel shows the cross-linking results with the RNAs shown in the second set of fragments in A. The WAF 1–879 was included as a positive control in this second study. C, nucleotide sequence of WAF1-A. Three CU-rich patches are underlined.
TABLE 4.
PCR primers used to amplify regions of the WAF 1–879 sequence
| Name | mRNA length | Primer seta |
|---|---|---|
| nucleotides | ||
| WAF1-A | 127 | CTTCATGCCAGCTACTTCCTCCTCCCC (forward) |
| CAGGTCTGAGTGTCCAGGAAAGGGGG (reverse) | ||
| WAF1-B | 127 | AATTCTTTTTCATTTGAGAAGTAAACAGATGGC (forward) |
| GCTCACTTCAGGGTCACCCTGCCCAACC (reverse) | ||
| WAF1-C | 127 | ACAGCCTAGGGCTGAGCTGGGGACC (forward) |
| TCAGAGGGGCCATGAGGGCAGGCGGGG (reverse) | ||
| WAF1-D | 150 | CCTGCACTGGGGAGCCCGTCTCAGTGTTGAGCC (forward) |
| TACTGAAGGGAAAGACAAGGGGGAGGGACAGC (reverse) | ||
| WAF1-E | 98 | CCCTCTCATGCTCCAGGTGGCTCTGAGG (forward) |
| ACTAGGGTGCCCTTCTTCTTGTGTGTCCC (reverse) |
Each forward primer has a T7 promoter linked to its 5′-end (sequence not listed in Table 4). Each primer sequence is shown 5′-3′.
RESULTS
siRNA-mediated Depletion of αCP1 and αCP2 in K562 Cells—αCP1 and αCP2 siRNAs were transfected either individually or in combination. Western blot analyses revealed that the αCP1 and αCP2 siRNAs selectively depleted their targeted proteins and that both αCPs isoforms were depleted when the two siRNAs were used in combination (Fig. 1A). Two controls were included to document specificity of siRNA actions: a “mock” transfection lacking only siRNAs and a transfection with an unrelated siRNA directed against lamin A/C. Expression of lamin A/C protein was unaffected by either of the αCP siRNAs but was effectively cleared by the lamin A/C siRNA.
FIGURE 1.
siRNA-mediated depletion of αCP1 and αCP2. A, cells were mock-transfected (M) or transfected with siRNAs directed against αCP1, αCP2, αCP1 and αCP2 (αCP1 + 2), or lamin A/C (Lam). Protein lysates were analyzed by Western blotting using antibodies directed against the corresponding proteins. The two bands detected with the αCP2-specific antibodies represent αCP2 full-length protein (upper band) and αCP2KL, its major splice variant. The two bands in the lamin A/C Western blot panel represent the two lamins, A and C. B, the simultaneous depletion of αCP1 and αCP2 results in decreased cyclin H protein expression. Cells were mock-transfected or transfected with a mixture of siRNAs directed againstαCP1 andαCP2 (αCP1 + 2) or directed against lamin A/C (Lam). Protein lysates were analyzed by Western blotting using antibodies directed against cyclin H or an antibody that recognizes RB (as a loading control).
Alterations of mRNA Steady State Levels in αCP1/2-co-depleted Cells—To identify cellular mRNAs whose expression is modulated by αCP1 and αCP2, RNA isolated from control and αCP1/2-co-depleted cells were compared by microarray analysis. The study included six sets of microarray hybridizations, beginning each time with an independent siRNA transfection; three of these studies compared combined treatment with αCP1 and αCP2 siRNAs (“αCP1/2 co-depletion”) with mock-transfected cells, and the other three compared the co-depletion with lamin A/C siRNA treatment. 41 mRNAs were altered 1.7-fold or more in both of these comparisons. Table 1 identifies each mRNA by GenBank™ accession number, average -fold changes, and putative function(s) of the encoded proteins, as identified by GO (gene ontology) terms, OMIM (online Mendelian inheritance in man), and/or manual literature searches. The mRNAs are ranked according to the average -fold change of all six experiments. There were 27 down-regulated mRNAs and 14 up-regulated mRNAs. The mRNAs encoding αCP1 and αCP2 ranked highest on this list of down-regulated mRNAs, with αCP2 (or its variants) ranked at number 1, 2, and 6 with -fold changes of –7, –7, and –3 compared with mock and –5, –4, and –3 compared with lamin A/C. Likewise, αCP1 ranked at number 3 with a -fold change of –4 compared with mock or lamin A/C. These results validate the effective siRNA targeting of αCP1 and αCP2 mRNAs. The fourth highest ranking candidate is the mRNA encoding the cell cycle regulator, CCNH (cyclin H). This mRNA was decreased by 6-fold when αCP1/2 co-depletion was compared with mock depletion and by 3-fold when compared with lamin A/C. The mRNA encoding HIF1α (hypoxia-inducible transcription factor 1α) ranked at number 10. In the list of mRNAs that were up-regulated by the αCP1/2 co-depletion, an anonymous mRNA ranked the highest, with a 6-fold increase in its steady state level when compared with mock and an 11-fold increase when compared with lamin A/C.
TABLE 1.
41 mRNAs altered in steady state level in cells depleted of αCP1 and αCP2
| mRNA name | Accession no. | AFCaversus mocka | AFC versus lamin A/C | Function |
|---|---|---|---|---|
| Down-regulated | ||||
| αCP2, transcript variant 1 | NM_005016 | -7 | -4.5 | mRNA metabolism and expression |
| Strong similarity to αCP2 | I_1903441 | -6.6 | -4.1 | mRNA metabolism and expression |
| αCP1 | NM_006196 | -3.5 | -3.5 | mRNA metabolism and expression |
| Cyclin H (CCNH) | NM_001239 | -5.5 | -3.4 | Regulation of cell cycle |
| Unknown | I_1931775 | -4 | -3 | Unknown |
| αCP2 | I_932189 | -3.2 | -3.2 | mRNA metabolism and expression |
| Chromosome ORF 30 | NM_014145 | -4 | -3 | Unknown |
| Unknown | I_3585116 | -3 | -2.8 | Unknown |
| Fatty acid desaturase 1 (FADS1) | NM_013402 | -3.2 | -2.4 | Fatty acid unsaturation |
| Hypoxia-inducible factor 1α (Hif1α) | I_958733 | -2.1 | -2.9 | Hypoxia |
| Unknown | ENST00000 | -2.9 | -3.1 | Unknown |
| Unknown | THC152941 | -2.4 | -2.5 | Unknown |
| Hypothetical protein FLJ37478 | NM_178557 | -2.3 | -2.5 | Unknown |
| cDNA FLJ10004 clone HEMBA1000076 | AK000866 | -2.5 | -2.2 | Unknown |
| cDNA FLJ90838 clone Y79AA1002129 | AK075319 | -2.5 | -2.1 | Unknown |
| Hypothetical protein FLJ25006 | NM_144610 | -2.3 | -2.2 | Unknown |
| Ras association domain 5 (RASSF5) | NM_031437 | -2.4 | -2.1 | Ras effector |
| Unknown | THC157597 | -2.4 | -2.1 | Unknown |
| Aldo-ketoreductase 7, A2 (AKR7A2) | NM_003689 | -2.5 | -2 | Aldehyde and ketone detoxification |
| cDNA DKFZp547F1714 | AL831830 | -2.1 | -2.2 | Unknown |
| WD repeat SOCS box 2 (WSB2) | NM_018639 | -2.4 | -2 | Unknown |
| Single-stranded DNA-binding protein 4 (SSBP4) | NM_032627 | -2.2 | -2 | Unknown |
| N-Acetylgalactosaminyltransferase (GALNT11) | NM_022087 | -1.9 | -1.7 | Glycosylation of mucins |
| Repressor of estrogen receptor activity (REA) | NM_007273 | -2.1 | -2.1 | Unknown |
| Ribosome-binding protein 1 | I_961859 | -2.2 | -1.9 | Translation and cardiac development |
| FLJ00069 | AK024476 | -2.1 | -2 | Unknown |
| Novel chromosome 22 gene | AL365511 | -2 | -1.9 | Unknown |
| Up-regulated | ||||
| Unknown | THC1570157 | 5.9 | 10.6 | Unknown |
| cDNA FLJ12961 clone NT2RP2005645 | AK023023 | 2.6 | 2.6 | Unknown |
| Angiomotin-like 1 (AMOTL1) | NM_130847 | 2.7 | 2.5 | Control of angiogenesis |
| Phosphoglycerate dehydrogenase (PHGDH) | NM_006623 | 2.2 | 3 | Serine biosynthesis |
| Ubiquitin-conjugating enzyme (UbcH2) | Z29328 | 2.3 | 3.3 | Ubiquitination of cellular substrates |
| mRNA adjacent to integrated HPV16 (INT423) | AJ431620 | 2.5 | 4.4 | Unknown |
| KIAA1541 protein | AB040974 | 3 | 3.8 | Unknown |
| Cytoskeletal tropomysoin isoform (3 kb) | M12127 | 2 | 2.3 | Actin-myosin interaction |
| Phosphotyrosine and phosphoinositides adaptor | NM_014395 | 2.2 | 2.2 | Signal transduction |
| Unknown | XM_209628 | 2.1 | 2.5 | Unknown |
| Unknown | THC1141659 | 2.4 | 2.5 | Unknown |
| cDNA FLJ10656 clone NT2RP2006038 | AK001518 | 2 | 3 | Unknown |
| Hypothetical protein FLJ10656 (P15RS) | NM_018170 | 2.3 | 2.5 | Unknown |
| RNA helicase (RIG-1) | AF038963 | 2.3 | 2.8 | Antiviral signaling |
AFC, average -fold change.
Verification of the microarray data was carried out on selected mRNAs by targeted qRT-PCR (Table 2). The level of each mRNA was determined as the -fold change versus the mock transfection control and the -fold change versus the Lamin A/C siRNA control. The mRNA encoding CCNH was 3- and 2-fold lower in the αCP1/2 co-depleted cells compared with mock or lamin A/C knockdowns, respectively. Likewise, the mRNAs encoding HIF1α, FADS1 (fatty acid desaturase 1), and RASSF5 (Ras association domain family protein 5) were decreased in the αCP1/2-co-depleted cells, all in agreement with the microarray analysis. The increase in the levels of the mRNAs encoding PHGDH (phosphoglycerate dehydrogenase), RIG-1 (retinoic acid-inducible gene 1), and UbcH2 (ubiquitin-conjugating enzyme E2H), as determined by the microarray analysis of αCP1/2 co-depleted cells, were confirmed by the qRT-PCR analysis, as was the lowest ranked up-regulated mRNA (rank number 14) in the microarray analysis (RIG-I; Table 1). These qRT-PCR studies support the reliability of the microarray data set.
TABLE 2.
Verification of mRNAs that are altered in steady state level in the absence of αCP1 and αCP2
| mRNA name | AFCaversus mock | AFC versus lamin A/C |
|---|---|---|
| Down-regulated | ||
| CCNH | -2.8 | -2.2 |
| HIF1α | -3 | -3.6 |
| FADS1 | -11.7 | -5.7 |
| RASSF5 | -3 | -3.4 |
| Up-regulated | ||
| PHGDH | 4 | 5.4 |
| RIG-I | 10.6 | 4 |
| UbcH2 | 2.4 | 1.8 |
AFC, average -fold change among two replicates of double knockdown.
Expression of Cyclin H Is Decreased in αCP1/2-co-depleted Cells—Since CCNH mRNA was the most strongly down-regulated mRNA in the αCP1/2 co-depleted cells (excluding αCPs), we assessed the corresponding impact on CCNH protein. Western blot analysis was consistent with the mRNA analysis, revealing that CCNH protein was reduced by ∼50% in the αCP1/2-co-depleted cells (Fig. 1B). The levels of CCNH protein in cells individually depleted of either αCP1 or αCP2 were reduced by ∼20–30% in each case (data not shown). These data suggest that αCP1 or αCP2 can each regulate CCNH protein expression, but together the effect is additive. Taken together, the data support the conclusion that CCNH mRNA and protein are markedly and coordinately reduced in cells depleted for αCP1 and/or αCP2.
Co-depletion of αCP1 and αCP2 Results in a G1 Cell Cycle Arrest—The observed reduction of CCNH protein in αCP-depleted cells suggested that αCP1 and αCP2 levels might impact on cell cycle kinetics. To test this possibility, αCP1 and αCP2 were depleted from K562 cells both individually and in combination, and cell replication parameters were evaluated in comparison with mock transfection and lamin A/C siRNA transfection controls. This analysis revealed a 53% reduction in cell number subsequent to co-depletion of αCP1 and αCP2 compared with mock-treated cells, and a 41% reduction in cell number when the αCP1/2 co-depletion was compared with lamin A/C siRNA-transfected cells at 4 days post-transfection of siRNAs. In contrast, there was no significant decrease in the density of cells individually depleted of αCP1 or αCP2 or in the controls (data not shown). Fluorescence-activated cell sorting analysis of cells transfected with siRNAs revealed that the αCP1/2 co-depletion resulted in accumulation of cells in the G1 phase that was not apparent in the mock-transfected or lamin A/C-transfected controls (Fig. 2). The G1 arrest in the αCP1/2 knockdown was observed in three independent knockdown studies. When αCP1/2-co-depleted cells were compared with control cells, the differences in the accumulation of G1 phase cells were highly significant (p = 0.0009 when compared with mock-transfected and p = 0.0023 when compared with lamin A/C siRNA-transfected cells). This G1 arrest was accompanied by a reciprocal decrease of cells in S phase (Fig. 2) that was significant when the double αCP1/2 knockdown was compared with mock-transfected (p = 0.0043) or with lamin A/C-transfected (p = 0.0068) cells. These alterations in the cell cycle were not observed in cells individually depleted for αCP1 or αCP2 (Fig. 2). There was no substantial impact on viability among the various treatments when assessed by dye exclusion (data not shown). Taken together, the data reveal that co-depletion of αCP1 and αCP2 reduced cellular proliferation and resulted in a G1 arrest.
FIGURE 2.
K562 cells that are acutely co-depleted of αCP1 and αCP2 accumulate in the G1 phase of the cell cycle. A, cells were mock-transfected or transfected with siRNAs directed against αCP1, αCP2, αCP1 and αCP2 (αCP1 + 2), or lamin A/C (Lam) and were subject to cell cycle analysis at 4 or 5 days after siRNA treatment. The x axis shows DNA content as determined by propidium iodide (PI) fluorescence, and the y axis shows number of cells. The regions representing G1, S, and G2 are indicated. B, analysis and quantitation of the percentage of cells found in each phase of the cell cycle from replicate experiments. The mean and S.D. are shown.
Phosphorylation of Serine 795 on the Retinoblastoma (RB) Protein Is Increased in the αCP1/2-co-depleted K562 Cells—The RB protein is a pivotal factor in the G1 to S transition of the cell cycle, and phosphorylation of specific residues in RB has been implicated in this activity (35). The observation that cells co-depleted of αCP1 and αCP2 accumulate in G1 led us to monitor for changes in the phosphorylation status of RB. Western blot analysis revealed that RB phosphorylation at Ser795 was increased in the αCP1/2-co-depleted cells relative to controls (Fig. 3A, top). There was also a slight but consistent increase of Ser795 phosphorylation in cells treated with lamin A/C siRNA relative to the mock treatment. Analysis of cells individually depleted for αCP1 revealed a slight increase in Ser795 phosphorylation, whereas the modification of this residue in the cells individually depleted of αCP2 remained unchanged (data not shown). The Western analysis of Ser780 phosphorylation (Fig. 3A, middle) failed to reveal any changes in any of the conditions tested. The overall levels of RB protein were also found to be unaltered in any depleted cells (Fig. 3, A and B, bottom panels). Phosphorylation of RB at Ser807/811 was marginally increased by the αCP1/2 co-depletion relative to the mock control, but a marginal increase was also observed in the cells treated with the lamin A/C siRNA (Fig. 3B). Taken together, these data reveal a selective increase in phosphorylation of RB at Ser795 in cells co-depleted of αCP1 and αCP2. However, since previous reports (36) have shown that phosphorylation of RB at Ser795 correlates with entry into S phase, the linkage of this change to the observed G1 arrest appeared unlikely. For this reason, we decided to search for additional targets of αCP that might be causative in the observed G1 arrest in the αCP1/2-co-depleted cells.
FIGURE 3.
Co-depletion of αCP1 and αCP2 increases the phosphorylation of RB at serine 795. Cells were mock-transfected (M) or transfected with siRNAs directed against αCP1 and αCP2 (αCP1 + 2) or lamin A/C (Lam). Protein lysates were analyzed by Western blotting using antibodies directed against the following phosphorylated forms of RB: Ser795 and Ser780 (A) and Ser807/811 (B). An antibody that recognizes RB was utilized as a load control.
Targeted Analysis of mRNAs Encoding Cell Cycle Control Proteins Reveals a Subset of mRNAs Whose Expression Is Altered by the αCP1/2 Co-depletion—To further explore the mechanism by which αCPs impact on cell cycle control(s), we screened a set of 84 human genes that play key roles in cell cycle regulation for alterations in mRNA levels subsequent to the αCP1/2 co-depletion (RT2 Profiler PCR array). Eleven mRNAs that were found to be either up- or down-regulated in the αCP1/2-co-depleted cells when compared with the mock and the Lamin A/C depleted cells are shown in Table 3. Included in this set were the mRNAs encoding p53 and p21WAF. When compared with mock- or lamin A/C-depleted cells, the αCP1/2 co-depleted cells had levels of p53 mRNA and p21WAF mRNA that were increased by 3.1- and 2.6-fold and 3- and 4.1-fold, respectively. It should be noted that the p21WAF gene was not detected on the microarray platform used in our initial study (Table 1). This may be due to differences in the detection limits of qRT-PCR and microarray analysis. The increases in p53 and p21WAF mRNA levels seen in the αCP1/2-co-depleted cells were of particular interest, since either could contribute to the G1 arrest.
TABLE 3.
RT2 Profiler PCR array human cell cycle analysis
| mRNA name | FCaversus mock | FC versus lamin A/C | Function |
|---|---|---|---|
| Down-regulated | |||
| RAD9A | -2.4 | -3 | DNA repair and DNA damage sensor |
| B cell CLL/Lymphoma 1 (BCL1) or cyclin D1 (CCND1) | -3.7 | -3 | Proto-oncogene |
| Cyclin G1 (CCNG1) | -3.5 | -2.6 | G2 phase and G2/M transition control |
| Hypoxanthine guanine phosphoribosyltransferase 1 (HPRT1) | -2.1 | -2 | Enzyme in purine salvage pathway |
| Minichromosome maintenenance 4 (MCM4) | -2 | -2 | S phase and DNA replication |
| Up-regulated | |||
| Distinct subgroup of the Ras family, member 3 (DIRAS3) | 3 | 4.2 | Tumor suppressor |
| Cyclin-dependent kinase inhibitor 1A (CDKN1A) or p21WAF | 3 | 4.1 | Inhibitor of cyclin dependent kinases |
| B cell CLL/lymphoma 2 (BCL2) | 2.8 | 3.9 | Proto-oncogene |
| Proliferating Cell Nuclear Antigen (PCNA) | 3.1 | 3 | DNA polymerase δ auxilliary protein |
| Tumor protein p53 (TP53) | 3.1 | 2.6 | Tumor suppressor |
| G2 and S phase-expressed gene 1 (GTSE1) | 2.1 | 2 | Overexpression induces G2 to M block |
FC, -fold change of double knockdown.
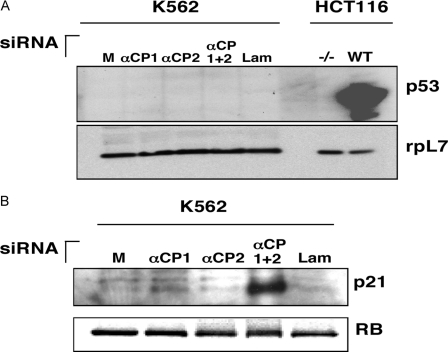
p21WAF Protein Is Up-regulated in the Cells Co-depleted of αCP1 and αCP2 via a p53-independent Mechanism—The observation that the levels of p53 and p21WAF mRNAs were both enhanced in cells co-depleted of αCP1 and αCP2 was followed by determining whether these changes were reflected at the protein level. Western blot analysis of extracts derived from cells individually transfected with αCP1 siRNA, αCP2 siRNA, or a combination of αCP1 and αCP2 siRNAs was compared with mock-transfected and lamin A/C siRNA-transfected controls (Fig. 4A). The human colon cancer cell line (HCT116) was used as a control for p53 detection. HCT116 cells containing the p53 gene (WT) and derivative HCT116 cells lacking the p53 gene (–/–) were treated with irinotecan to induce p53 expression. p53 was robustly and selectively induced by irinotecan in the wild type HCT116 cells. Parallel analysis of the K562 cells revealed a complete absence of p53 protein. This lack of p53 was consistent with previous reports (37) showing that the p53 gene is inactivated in K562 cells. The mutation consists of an insertion of a cytosine between codons 135 and 136. This insertion creates a frameshift, leading to a truncated protein of 147 amino acids. In contrast to the lack of p53 expression, p21WAF was strongly induced in the cells co-depleted for αCP1 and αCP2 (Fig. 4B). Interestingly, the individual αCP1 and αCP2 depletions had no apparent effect on p21WAF protein levels. In summary, the data confirm that p21WAF protein is strongly induced by co-depletion of αCP1 and αCP2 and that this effect is p53-independent.
FIGURE 4.
Co-depletion ofαCP1 andαCP2 results in p53-independent induction of p21WAF expression. A, K562 cells were mock-transfected (M) or transfected with siRNAs directed against αCP1, αCP2, αCP1 and αCP2 (αCP1 + 2), or lamin A/C (Lam). HCT116 cells that lack (–/–) or contain the wild type (WT) p53 gene were treated with irinotecan to induce p53 expression and were used as controls. Protein lysates were analyzed by Western blotting using antibodies directed against p53. The loading control is the ribosomal protein L7 (rpL7). B, K562 cells were transfected with siRNAs as above and analyzed by Western blotting using antibodies directed against p21WAF. An antibody directed against RB was utilized as a loading control.
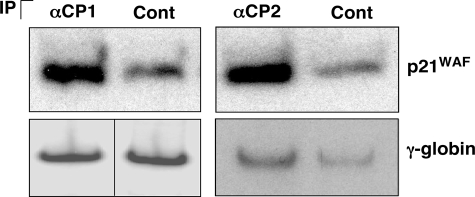
The mRNA encoding p21WAF Interacts with both αCP1 and αCP2 in Vivo—Since p21WAF mRNA and protein were both found to be up-regulated by the αCP1/2 co-depletion and since αCPs are known to modulate gene expression via targeted binding to mRNA, we asked whether αCP1 and αCP2 interacted with p21WAF mRNA in vivo. αCP1- and αCP2-containing mRNPs were individually enriched from K562 cytosolic extracts by immunoprecipitation with isoform-specific antibodies. mRNAs isolated from both sets of mRNP immunoprecipitations were assessed for enrichment of p21WAF mRNA by a semiquantitative RT-PCR analysis (Fig. 5). The analysis revealed that the p21WAF mRNA was enriched by ∼3- and 3.4-fold in the αCP1 and αCP2 mRNP isolates, respectively. In contrast, levels of γ-globin mRNA levels were not significantly different in the αCP and control immunoprecipitates. These data lead us to conclude that both αCP1 and αCP2 bind to the p21WAF mRNA in vivo.
FIGURE 5.
The mRNA encoding p21WAF is associated with αCP1 and αCP2 in vivo. K562 cell extracts were immunoprecipitated (IP) with antibodies to αCP1 or αCP2 or a c-Myc control (Cont) antibody. Following immunoprecipitation, RNA was isolated from the RNP complexes and subjected to RT-PCR analysis to detect p21WAF or γ-globin mRNAs. The γ-globin image for the αCP1 and control immunoprecipitate was from different areas of the same gel.
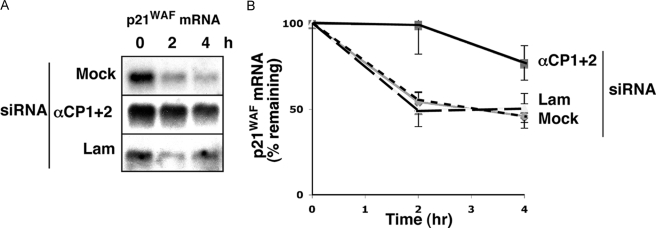
Co-depletion of αCP1 and αCP2 Stabilizes Endogenous p21WAF mRNA in K562 Cells—The induction of p21WAF mRNA levels in the αCP1/2-co-depleted cells and the observation that both αCP proteins interacted with the p21WAF mRNA in vivo suggested that control over p21WAF mRNA levels might be mediated by an effect of αCP on p21WAF mRNA stability. This model was tested. At 2 days post-siRNA transfection, the cells were treated with the transcriptional inhibitor actinomycin D, and RNA harvested at subsequent time points was quantified for p21WAF mRNA by Northern analysis. This analysis revealed that the rates of p21WAF mRNA decay in mock- or lamin A/C siRNA-transfected cells were similar, with a half-life of ∼3 h (Fig. 6). In contrast, the αCP1/2 co-depletion resulted in prolongation of the half-life to ∼13 h. This alteration in p21WAF mRNA stability in the αCP1/2-co-depleted cells is consistent with the observed increase in levels of p21WAF mRNA and protein.
FIGURE 6.
The p21WAF mRNA is stabilized in cells co-depleted of αCP1 and αCP2. A, K562 cells were mock-transfected (M) or transfected with siRNAs directed against αCP1 and αCP2 (αCP1 + 2) or against lamin A/C (Lam). At 2 days post-transfection, actinomycin D was added to inhibit transcription, and total RNA was collected at 0, 2, and 4 h. Northern blot analysis was used to monitor p21WAF mRNA levels. Representative blots are shown. B, the experiment shown in A was performed three times, and the p21WAF mRNA levels at each time point were quantified on a PhosphorImager. For each set of siRNA transfections, the band intensity at the 0 h time point was set to 100%, and the percentage of mRNA remaining was plotted over time. The error bars show the S.E. among the three experiments.
αCP1 and αCP2 Bind to a 127-nucleotide Fragment of the p21WAF 3′-UTR—Since we found that the p21WAF mRNA interacted with αCP1 and αCP2 in vivo (Fig. 5) and since depletion of both proteins stabilized the p21WAF mRNA (Fig. 6), we decided to test whether regions of the p21WAF 3′-UTR interacted with αCP. Our rationale for this experiment was that several examples exist in the literature where αCP binding sites occur in the 3′-UTR of specific mRNAs and regulate mRNA stability (14–16). We obtained a series of plasmids each containing a T7 promoter that drives the synthesis of different regions of the p21WAF 3′-UTR (33). The map of these regions is shown in Fig. 7A. We synthesized thiolated RNA corresponding to these regions and used them in cross-linking assays. After UV cross-linking and RNase A digestion, the resulting mRNP complexes were immunoprecipitated using antibodies directed against αCP1, αCP2, or a c-Myc control. The results are shown in Fig. 7B (top). Actin antisense RNA and the α-globin 3′-UTR were used as negative and positive controls, respectively. We did not detect any cross-linked immunoprecipitated product using any of the antibodies when the actin antisense RNA was used in the assay. In contrast, when the α-globin 3′-UTR was used, we observed a cross-linked product when the immunoprecipitation was carried out using αCP2- or αCP1-specific antibodies. Both of these products were of the appropriate size for αCP2 or αCP1. The only fragment of the p21WAF 3′-UTR that interacted with both αCP1 and αCP2 was WAF 1–879, which corresponds to nucleotides 879–1512 of the p21WAF 3′-UTR. Next, we further mapped the αCP binding site on the 3′-UTR by generating smaller fragments of WAF 1–879 (WAF1-A to WAF1-E) to be used in the cross-linking assay (Fig. 7A and Table 4). The cross-linking results are shown in Fig. 7B (bottom). WAF 1–879 was used as a positive control, and we observed the expected immunoprecipitation of αCP1 and αCP2. The only subfragment of WAF 1–879 that showed significant binding to αCP was WAF1-A. WAF1-D had very faint cross-linked products, but it was not reproducible. Therefore, we conclude that the major binding determinant of both αCP1 and αCP2 resides in the WAF1-A fragment. The sequence of WAF1-A is indicated in Fig. 7C. The triplication of C-rich regions (underlined) bears striking resemblance to the triple C-rich motifs previously identified as the αCP binding site in human α-globin mRNA (3, 16).
DISCUSSION
Our current observations lead us to conclude that αCP1 and αCP2 play a significant role in the control of p21WAF expression. This control appears to reflect a direct in vivo association of these proteins with the p21WAF mRNA with consequent mRNA stabilization and increase in p21WAF protein expression. Co-depletion of αCP1 and αCP2 results in a decrease in cell proliferation and a G1 cell cycle arrest (Fig. 2). These functions appear to be mechanistically linked to the increase in p21WAF protein levels (Figs. 4, 5, 6). This direct, post-transcriptional control of p21WAF expression by the αCP proteins is consistent with the observation that this control is independent of p53, the major transcriptional modulator of p21 expression. Of note, the alteration in cell cycle kinetics and increase in p21WAF expression in the αCP1/2 co-depleted cells were not apparent in cells individually depleted of αCP1 or αCP2 (Figs. 2 and 4). These data suggest that the αCP1 and αCP2 isoforms have overlapping and/or redundant functions that are required for control of p21WAF expression and normal progression through the cell cycle.
CCNH Expression in the αCP1/2-co-depleted Cells—The mechanism of the G1 arrest in the cells co-depleted for αCP1 and αCP2 was investigated by defining alterations in the expression of mRNAs that encode proteins involved in cell cycle control. We found that the CCNH mRNA was down-regulated by the αCP1/2 co-depletion (Table 1) with a 50% decrease in protein expression (Fig. 1B). The results of the individual αCP1 or αCP2 knockdowns suggested that these isoforms can individually and additively regulate CCNH expression. CCNH is a regulatory subunit for a Cdk (cyclin-dependent kinase)-activating kinase involved in multiple cell cycle transitions (38). Selective inhibition of Cdk7 (a Cdk-activating kinase subunit) delays entry into S phase (39). Therefore, it is possible that a decrease in CCNH expression in cells depleted of αCP1 and αCP2 could disrupt Cdk-activating kinase function(s) and contribute to the observed G1 arrest. However, the observation that individual αCP1 and αCP2 depletions are sufficient for repression of CCNH expression and yet fail to trigger the G1 arrest leads us to conclude that the additive effect of the combined knockdowns on CCNH may be contributory to but are not the defining determinants of the G1 arrest seen in the combined αCP1/2 depletion.
RB Phosphorylation in the αCP1 and αCP2 Co-depletion—Phosphorylation of RB plays a major role in RB-mediated cell cycle controls. We observed an increase in the phosphorylation of RB at Ser795 in cells co-depleted for αCP1 and αCP2 (Fig. 3). However, it seems unlikely that this alteration in RB phosphorylation is the cause of the G1 arrest subsequent to αCP1/2 co-depletion. Cells treated with the control lamin A/C siRNA showed a reproducible, albeit moderate, increase in Ser795 phosphorylation without a corresponding alteration in the cell cycle. In addition, previous reports in osteogenic sarcoma (SAOS-2) cells show that phosphorylation of RB at Ser795 correlates with entry into S phase rather than G1 arrest (36). Although it is possible that the impact of the RB phosphorylation in K562 cells may differ from that in other cells, this linkage remains untested.
Relationship of αCPs to p21WAF Expression and Cell Cycle Controls—Our data are most consistent with a pathway in which co-depletion of αCP1 and αCP2 lead to an induction of p21WAF protein expression (Fig. 4B) via stabilization of p21WAF mRNA (Fig. 6). The increase in p21WAF protein correlates with the G1 arrest; both occur only in the αCP1/2-co-depleted cells and not in cells where the mRNAs encoding these two proteins are individually targeted. The impact of the increased p21WAF expression on cell growth is fully concordant with the known ability of p21WAF to mediate a G1 block of the cell cycle (40). Therefore, our data suggest that activation of p21WAF protein expression, triggered by depletion of αCP1 and αCP2, mediates G1 arrest in the K562 cells.
p21WAF is a direct mediator of cell cycle arrest at the G1 phase (41). p21WAF can inhibit specific Cdks, resulting in inhibition of RB phosphorylation (40). Unphosphorylated RB protein binds to several proteins involved in the regulation of the G1 to S transition, including the E2F family of transcription factors (42). The RB-E2F complex acts as a transcriptional repressor whose targets include several genes required for S phase, contributing to the mechanism of G1 arrest (43). p21WAF also appears to be required for maintaining the G2 checkpoint in human cells (44). The pathways by which p21WAF levels in the cell are controlled and modulated appear to be complex and remain to be fully defined.
Since p53 has been shown to transcriptionally up-regulate p21WAF (45, 46), an important parameter of the p21WAF-induced G1 arrest in the K562 cells is that this effect occurs in the absence of p53. The p53 gene in K562 cells contains a single base insertion that leads to a translational frameshift and a truncated protein (37). Sequencing of the p53 locus in K562 cells reveals only the mutant sequence, indicating that the wild type allele has been either lost or converted to the mutant allele (37). Consistent with this mutation, the p53 mRNA could be detected (Table 3), whereas the protein was not detected by Western blotting (Fig. 4A).
Post-transcriptional Regulation of p21WAF mRNA Expression—Although p21WAF expression is under p53-mediated transcriptional control, an extensive body of literature documents that p21WAF expression is also subject to post-transcriptional modulation. For example, the RNA-binding proteins hnRNP K (47) or Msi-1 (48) can block translation of the p21WAF mRNA, and p21WAF mRNA stability can be altered by a number of mRNA-binding proteins. The half-life of the p21WAF mRNA can be increased by the binding of HuR in response to UV light (49) or prostaglandin A2 treatment (50) or by the binding of RNPC1a (51). In the last situation, the stabilization of p21WAF mRNA is accompanied by G1 arrest (51). Treatment of cells with hydroxyurea has been shown to stabilize p21WAF mRNA (52), although the mechanism remains undefined. The 3′-UTR of p21WAF mRNA has also been shown to be bound by a number of RNA-binding proteins, including αCP1, although the functional impact of αCP1 binding in that study was not explored (33). In that case, recombinant αCP1 was used, and binding was detected using the WAF1-1/6 fragment (referred to as WAF1–571 in Fig. 7A). In contrast, our study indicates that αCP1 and αCP2 both bind to WAF 1–879, and we did not detect binding to WAF1-1/6 (Fig. 7D). It is possible that differential RNA binding can occur with recombinant αCP versus cellular extracts containing αCP. Taken together, these reports indicate that expression of p21WAF is subject to multiple layers of post-transcriptional control.
What is the mechanism of the increased expression of p21WAF mRNA in the current study? Since previous work has linked αCP to the regulation of a number of mRNA targets, we monitored the impact of the siRNA treatments on p21WAF mRNA stability (Fig. 6). These studies revealed that the p21WAF mRNA half-life was increased in cells co-depleted of αCP1 and αCP2. Interestingly, αCP1 and αCP2 each bind to the p21WAF mRNA in vivo (Fig. 5). We mapped the αCP1 and αCP2 binding site on the 3′-UTR of p21WAF to a 127-nucleotide sequence. This sequence contains three CU-rich patches, reminiscent of the αCP binding site on the α-globin 3′-UTR (3, 16). The finding that co-depletion of αCP1 and αCP2 stabilizes the p21WAF mRNA suggests that under normal conditions, the p21WAF mRNA is destabilized by these two proteins. This finding is of particular interest, since αCP binding has been previously linked to mRNA stabilization rather than destabilization. Thus, the present study points to a novel activity of these hnRNP K homology domain proteins. However, our data suggest that this control via mRNA destabilization may not be unique; Table 1 lists 14 mRNAs whose steady state levels are increased in cells co-depleted of αCP1 and αCP2. The questions of whether these mRNAs are coordinately stabilized in the co-depleted cells in some manner, whether they are all direct binding targets of αCP1 and/or αCP2, and whether the alteration in any of these additional mRNAs contributes to the cell cycle arrest in the co-depleted cells can now be addressed.
Acknowledgments
We greatly appreciate the gift of HCT116 cell extracts from Dr. Wafik El-Deiry. We thank Dr. Christian Sell and Jeff Thomas for technical assistance. We thank Dr. Donna George for experimental advice and critical reading of the manuscript.
This work was supported, in whole or in part, by National Institutes of Health Grants PO1-CA72765, MERIT HL 65449, and K01 DK-071137-04.
Footnotes
The abbreviations used are: αCP, α-globin poly(C)-binding protein; hnRNP, heterogeneous nuclear ribonucleoprotein; UTR, untranslated region; siRNA, small interfering RNA; RT, reverse transcription; qRT-PCR, quantitative reverse transcription-PCR.
References
- 1.Makeyev, A. V., and Liebhaber, S. A. (2000) Genomics 67 301–316 [DOI] [PubMed] [Google Scholar]
- 2.Aasheim, H. C., Loukianova, T., Deggerdal, A., and Smeland, E. B. (1994) Nucleic Acids Res. 22 959–964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kiledjian, M., Wang, X., and Liebhaber, S. A. (1995) EMBO J. 14 4357–4364 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Leffers, H., Dejgaard, K., and Celis, J. E. (1995) Eur. J. Biochem. 230 447–453 [PubMed] [Google Scholar]
- 5.Makeyev, A. V., Chkheidze, A. N., and Liebhaber, S. A. (1999) J. Biol. Chem. 35 24849–24857 [DOI] [PubMed] [Google Scholar]
- 6.Makeyev, A. V., and Liebhaber, S. A. (2002) RNA 8 265–278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Tommerup, N., and Leffers, H. (1996) Genomics 32 297–298 [DOI] [PubMed] [Google Scholar]
- 8.Funke, B., Zuleger, R., Benavente, R., Schuster, T., Goller, M., Stevenin, J., and Horak, I. (1996) Nucleic Acids Res. 24 3821–3828 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gibson, T. J., Thompson, J. D., and Heringa, J. (1993) FEBS Lett. 324 361–366 [DOI] [PubMed] [Google Scholar]
- 10.Andino, R., Rieckhof, G. Achacose, P. L, and Baltimore, D. (1993) EMBO J. 12 3587–3598 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gamarnik, A. V., and Andino, R. (2000) J. Virol. 74 2219–2226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ostareck, D. H., Ostareck-Lederer, A., Wilm, M., Thiele, B. J., Mann, M., and Hentze, M. W. (1997) Cell 89 597–606 [DOI] [PubMed] [Google Scholar]
- 13.Parsley, T. B., Towner, J. S., Blyn, L. B., Ehrenfeld, E., and Semler, B. L. (1997) RNA 3 1124–1134 [PMC free article] [PubMed] [Google Scholar]
- 14.Paulding, W. R., and Czyzyk-Krzeska, M. F. (1999) J. Biol. Chem. 274 2532–2538 [DOI] [PubMed] [Google Scholar]
- 15.Stefanovic, B., Hellerbrand, C., Holcik, M., Briendl, M., Liebhaber, S. A., and Brenner, D. A. (1997) Mol. Cell. Biol. 17 5201–5209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang, X., Kiledjian, M., Weiss, I. M., and Liebhaber, S. A. (1995) Mol. Cell. Biol. 15 1769–1777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Weiss, I. M., and Liebhaber, S. A. (1995) Mol. Cell. Biol. 15 2457–2465 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Perrotti, D., Cesi, V., Trotta, R., Guerzoni, C., Santilli, G., Campbell, K., Iervolino, A., Condorelli, F., Gambacorti-Passerini, C., Caligiuri, M. A., and Calabretta, B. (2002) Nat. Gen. 30 48–58 [DOI] [PubMed] [Google Scholar]
- 19.Xiao, X., Tang, Y. S., Mackins, J. Y., Sun, X. L., Jayaram, H. N., Hansen, D. K., and Antony, A. C. (2001) J. Biol. Chem. 276 41510–41517 [DOI] [PubMed] [Google Scholar]
- 20.Blyn, L. B., Towner, J. S., Semler, B. L., and Ehrenfeld, E. (1997) J. Virol. 8 6243–6246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Blyn, L. B., Swiderek, K. M., Richards, O., Stahl, D. C., Semler, B. L., and Ehrenfeld, E. (1996) Proc. Natl. Acad. Sci. U. S. A. 93 11115–11120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Crichton, R. R., Wilmet, S., Legssyer, R., and Ward, R. J. (2002) J. Inorg. Biochem. 91 9–18 [DOI] [PubMed] [Google Scholar]
- 23.Gamarnik, A. V., and Andino, R. (1997) RNA 3 882–892 [PMC free article] [PubMed] [Google Scholar]
- 24.Graff, J., Cha, J., Blyn, L. B., and Ehrenfeld, E. (1998) J. Virol. 72 9668–9675 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gutierrez-Escolano, A. L., Brito, Z. U., delAngel, R. M., and Jiang, X. (2000) J. Virol. 74 8558–8562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Murray, K. E., Roberts, A. W., and Barton, D. J. (2001) RNA 7 1126–1141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Paillard, L., Maniey, D., Lachaume, P., Legagneux, V. H., and Osborne, B. (2000) Mech. Dev. 93 117–125 [DOI] [PubMed] [Google Scholar]
- 28.Spangberg, K., and Schwartz, S. (1999) J. Gen. Virol. 80 1371–1376 [DOI] [PubMed] [Google Scholar]
- 29.Walter, B. L., Nguyen, J. H. C., Ehrenfeld, E., and Semler, B. (1999) RNA 5 1570–1585 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Waggoner, S. A., and Liebhaber, S. A. (2003) Mol. Cell Biol. 23 7055–7067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Castano, Z., Vergara-Irigaray, N., Pajares, M. J., Montuenga, L. M., and Pio, R. (2008) Int. J. Cancer 122 1512–1520 [DOI] [PubMed] [Google Scholar]
- 32.Zhu, J., and Chen, X. (2000) Mol. Cell Biol. 20 5602–5618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Giles, K. M., Daly, J. M., Beveridge, D. J., Thomson, A. M., Voon, D. C., Furneaux, H. M., Jazayeri, J. A., and Leedman, P. J. (2003) J. Biol. Chem. 278 2937–2946 [DOI] [PubMed] [Google Scholar]
- 34.Johannes, G., and Sarnow, P. (1998) RNA 4 1500–1513 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hatakeyama, M., and Weinberg, R. A. (1995) Prog. Cell Cycle Res. 1 9–19 [DOI] [PubMed] [Google Scholar]
- 36.Connell-Crowley, L., Harper, J. W., and Goodrich, D. W. (1997) Mol. Biol. Cell 8 287–301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Law, J. C., Ritke, M. K., Yalowich, J. C., Leder, G. H., and Ferrell, R. E. (1993) Leuk. Res. 17 1045–1050 [DOI] [PubMed] [Google Scholar]
- 38.Fisher, R. P., and Morgon, D. O. (1994) Cell 26 713–724 [DOI] [PubMed] [Google Scholar]
- 39.Larochelle, S., Merrick, K.A., Terret, M. E., Wohlbold, L., Barboza, N. M., Zhang, C., Shokat, K. M., Jallepalli, P. V., and Fisher, R. P. (2007) Mol. Cell 25 839–850 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K., and Elledge, S. J. (1993) Cell 75 805–816 [DOI] [PubMed] [Google Scholar]
- 41.Hengst, L., Dulic, V., Slingerland, J. M., Lees, E., and Reed, S. I. (1994) Proc. Natl. Acad. Sci. U. S. A. 91 5291–5295 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Nevins, J. R. (1992) Science 258 424–429 [DOI] [PubMed] [Google Scholar]
- 43.Zhang, H. S., Postigo, A. A., and Dean, D. C. (1999) Cell 97 53–61 [DOI] [PubMed] [Google Scholar]
- 44.Bunz, F., Dutriaux, A., Lengauer, C., Waldman, T., Zhou, S., Brown, J. P., Sedivy, J. M., Kinzler, K. W., and Vogelstein, B. (1998) Science 282 1497–1501 [DOI] [PubMed] [Google Scholar]
- 45.Del Sal, G., Murphy, M., Ruaro, E., Lazarevic, D., Levine, A. J., and Schneider, C. (1996) Oncogene 12 177–185 [PubMed] [Google Scholar]
- 46.El-Deiry, W. S., Tokino, T., Velculescu, V. E., Levy, D. B., Parsons, R., Trent, J. M., Lin, D., Mercer, W. E., Kinzler, K. W., and Vogelstein, B. (1993) Cell 75 817–825 [DOI] [PubMed] [Google Scholar]
- 47.Yano, M., J. Hirotaka, and H. Okano. (2005) J. Biol. Chem. 280 12690–12699 [DOI] [PubMed] [Google Scholar]
- 48.Battelli, C., Nikopoulos, G. N., Mitchell, J. G., and Verdi, J. M. (2006) Mol. Cell Neurosci. 31 85–96 [DOI] [PubMed] [Google Scholar]
- 49.Wengong, W., Furneaux, H., Cheng, H., Caldwell, M. C., Hutter, D., Liu, Y., Holbrook, N., and Gorospe, M. (2000) Mol. Cell. Biol. 20 760–769 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Yang, X., Wang, W., Fan, J., Lal, A., Yang, D., Cheng, H., and Gorospe, M. (2004) J. Biol. Chem. 279 49298–49306 [DOI] [PubMed] [Google Scholar]
- 51.Shu, L., Yan, W., and Chen, X. (2006) Gen. Dev. 20 2961–2972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kim, H. S., Yeo, E. J., Park, S. H., Park, J. I., Park, S. C., Shin, J. Y., Kim, M. J., Oh, S. J., Won, M. H., Kang, T. C., Park, J. B., Kim, J., Kim, J. I., Lee, H. Y., and Lee, J. Y. (2005) Mech. Ageing Dev. 126 1255–1261 [DOI] [PubMed] [Google Scholar]