Abstract
Non-subspecies I salmonellae are commensals of cold-blooded vertebrates and cause sporadic disease in mammals. The reasons why non-subspecies I salmonellae do not circulate in populations of warm-blooded vertebrates, but instead only cause occasional disease in this niche, are unknown. We examined the ability of Salmonella enterica subsp. IIIa (subsp. arizonae) and subsp. IIIb (subsp. diarizonae) isolates to grow competitively with subspecies I (serovar Typhimurium) ATCC 14028 in vitro, to colonize Salmonella-sensitive BALB/c mice, and to persist in the intestine of Salmonella-resistant CBA/J mice in competitive infections. Subspecies IIIa had severely reduced intestinal colonization, intestinal persistence, and systemic spread in mice. Subspecies IIIa is nonmotile on swarming agar and thus may also have reduced motility under viscous conditions in vivo. Surprisingly, subspecies IIIb colonizes the intestinal tract of BALB/c mice normally yet does not spread systemically. Subspecies IIIb colonization of the intestine of CBA/J mice is reduced late in infection. In order to understand why these isolates do not colonize systemic sites, we determined that subspecies IIIa and IIIb are not internalized well and do not replicate in J774-A.1 murine macrophages, despite normal adherence to these cells. We further show that selected effectors of both type III secretion systems 1 and 2 are secreted by subspecies IIIa and IIIb in vitro but that each of these isolates secretes a different combination of effectors. We outline the phenotypic differences between these subspecies and subspecies I and provide a possible explanation for the inability of these strains to spread systemically in murine models.
The species Salmonella enterica contains six subspecies (10). Subspecies I serovar Enterica is responsible for the overwhelming majority of salmonellosis in mammals and birds, while non-subspecies I isolates cause only sporadic disease in mammalian (including humans) and avian species and are primarily described as nonpathogenic commensals of cold-blooded vertebrates (56, 58). The reasons for the overwhelming epidemiologic dominance of subspecies I salmonellae in mammalian and avian salmonellosis are unknown but may be related to an increased ability of this subspecies to be transmitted and persist within the warm-blooded vertebrate population.
An estimated 93,000 cases of human salmonellosis are attributed to amphibian and reptile contact each year; this constitutes ∼7% of the total annual cases of salmonellosis in the United States (14). Subspecies IIIa and IIIb are considered reptile associated, and they colonize the reptilian intestinal tract asymptomatically and are excreted in feces (6, 11, 49, 58). Both subspecies IIIa and IIIb can cause disease in various warm-blooded vertebrates, including humans, domestic poultry, sheep, wild birds, and cats (1, 2, 18, 50). Subspecies IIIa and IIIb are observed as the cause of disease in humans, especially in the young and in immunocompromised individuals. Many human cases of subspecies IIIa and IIIb salmonellosis can be linked to contact with a reptile, although other sources are possible (49).
Subspecies IIIa and IIIb can colonize the human intestine and can be isolated upon fecal culture from infected individuals (26, 49). Furthermore, subspecies IIIa and IIIb infections are invasive in young children and immunocompromised individuals, resulting in serious systemic disease, including sepsis and meningitis (14). Despite the fact that subspecies IIIa and IIIb can colonize the intestinal tract of warm-blooded vertebrates and cause disease in these hosts, the reasons why these subspecies do not circulate more widely in populations of warm-blooded vertebrates are unknown.
We hypothesized that subspecies IIIa and IIIb are not as epidemiologically successful as subspecies I in warm-blooded vertebrates because they may be unable to colonize and persist in the intestinal tracts of these hosts. In this study we determined the ability of subspecies IIIa and IIIb to colonize the mammalian intestine, persist there for prolonged periods, and spread systemically by using competitive infections with subspecies I (serovar Typhimurium) in murine models commonly employed to study salmonellosis.
MATERIALS AND METHODS
Bacterial strains and media.
The bacterial strains used in this study are described in Table 1. The subspecies I isolates used in these experiments are derivatives of S. enterica serovar Typhimurium ATCC 14028. Mutants for the type III secretion system 1 (TTSS-1) effector SipB (STM2885), TTSS-1 machinery (ΔinvA; STM2896), and TTSS-2 effectors SseG (STM1404) and SifA (STM1224) in the subspecies I ATCC 14028 background were generated by Lambda Red swap essentially as previously described (20). The S. enterica subsp. IIIa (subsp. arizonae) isolate used in this study was SGSC4693 (62:z4,z23:-; also known as SARC5 RSK2980, CDC346-86, or ATCC BAA-731), and the complete genome sequence is publicly available for this isolate (GenBank Accession number CP000880.1). The S. enterica subsp. IIIb (diarizonae) isolate used in this study was SGSC4692 [61:1,v:1,5,(7), also known as CDC 01-0005], and this isolate is currently being sequenced (54, 55). Spontaneous nalidixic acid-resistant isolates of these strains were generated in our laboratory by selecting resistant colonies on Luria-Bertani (LB) plates containing 50 μg/ml nalidixic acid, and these strains are listed in Table 1. All strains were routinely grown in LB broth or on LB plates containing 50 μg/ml nalidixic acid or 50 μg/ml kanamycin when appropriate.
TABLE 1.
Strains used in this study
| Strain name | Genotype | Reference or source |
|---|---|---|
| HA348 | ATCC 14028 Nalr | This work |
| HA350 | SGSC4693 Nalr (subsp. IIIa) | This work |
| HA378 | SGSC4692 Nalr (subsp. IIIb) | This work |
| HA458 | HA348 ΔinvA::Kan | This work |
| HA816 | ATCC 14028 ΔsseG::Kan | This work |
| HA817 | ATCC 14028 ΔsifA::Cm | This work |
| HA818 | ATCC 14028 ΔsipB::Kan | This work |
| AJB715 | ATCC 14028 phoN Nalr | 36 |
| SGSC4693 | IIIa (62:z4,z23:-) (SARC5 RSK2980, CDC346-86, and ATCC BAA-731) | 7 |
| SGSC4692 | IIIb [61:1,v:1,5,(7)] (CDC 01-0005 and ATCC BAA-639) | ATCC |
For determination of different subspecies CFU counts from competitive infections we used a simple colorimetric assay for PhoN expression. Mutations in phoN do not affect either virulence or intestinal persistence in subspecies I (36). IIIa (2980) lacks phoN activity and forms white colonies on medium containing 5 bromo-4-chloro-3-indolyl-β-d-phosphate (XP; 20 mg/liter), while IIIb used in these experiments is phoN positive and forms blue colonies on XP-containing medium. IIIa and IIIb can thus be differentiated from subsp. I isolates with either an intact (in the case of IIIa) or deleted phoN locus.
Swimming and swarming motilities of both subspecies IIIa and IIIb were tested and compared to the motility of subsp. I serovar Typhimurium ATCC 14028. Swimming motility was assayed on LB plates containing 0.3% Bacto agar (Difco), while swarming motility was assayed on plates containing 0.6% Bacto agar and 0.5% glucose as previously described (51). Three μl of stationary-phase culture in LB broth was spotted onto motility plates and incubated for 5 h at 37°C, and photographs were taken to capture the movement of all isolates spotted on the same plate. The diameter of each spotted sample was measured at this point and compared to that of subsp. I serovar Typhimurium and was also evaluated after 24 h of incubation on individual swarming motility plates. All assays were performed in triplicate and each experiment was repeated on three separate occasions.
Organ colonization in Salmonella-susceptible BALB/c mice.
We examined the competitive growth of subsp. IIIa (SGSC4693) and IIIb (SGSC4692) isolates with a commonly studied subsp. I isolate, serovar Typhimurium ATCC 14028 (HA348 ATCC 14028 Nalr and AJB715 ATCC 14028 Nalr phoN) in 8- to 10-week-old female Salmonella-sensitive BALB/c mice (Jackson Laboratories) in mixed infections using the following protocol: HA348 (wild type) versus HA350 (IIIa) and AJB715(wild type) versus HA378 (IIIb). Salmonella strains used as inocula were grown to stationary phase at 37°C with aeration and mixed in a 1:1 ratio of subsp. I to non-subsp. I. Inocula were serially diluted and titers were determined for bacterial CFU to determine the exact ratios for these strains.
Groups of six mice were inoculated intragastrically by gavage with approximately 2 × 107 bacteria in 200 μl of LB. Infected mice were observed daily for signs of illness and were euthanized after the development of signs, at 4 to 5 days postinfection (inactivity/reluctance to move, ruffled fur, or crouching together). Immediately after euthanasia Peyer's patches, ceca, mesenteric lymph nodes, livers, and spleens of infected mice were excised and homogenized in 5 ml ice-cold phosphate-buffered saline (PBS). Organ homogenates were serially diluted and plated to determine the ratio of subsp. I to non-subsp. I from the tissues of infected animals. Data are expressed as the ratio of subsp. I CFU versus non-subsp. I CFU, were normalized to the input ratio, and were converted logarithmically. Statistical significance was determined using Student's t test and a P value of <0.05.
Persistence in Salmonella-resistant CBA/J mice.
Subspecies IIIa and subsp. IIIb were tested for the ability to persist in the intestine of Salmonella-resistant CBA/J mice (Jackson Laboratories) in competitive infections with virulent subsp. I ATCC 14028 derivatives. Strains were grown to stationary phase at 37°C with aeration and were mixed 1:1 prior to inoculation. Groups of four to six 8- to 10-week-old CBA/J mice were infected intragastrically by gavage with an equal mixture of subsp. I and non-subsp. I isolates, approximately 2 × 109 total CFU in 100 μl LB (3). Approximately 100 mg of feces was collected at various time intervals (1, 3, 6, 9 12, 15, 21, 30, and 40 days postinfection), serially diluted, and plated for enumeration of CFU of subsp. I strain versus non-subsp. I strain. Data are expressed as the ratio of subsp. I versus the non-subsp. I CFU, were normalized to the input ratio, and were converted logarithmically. Statistical significance was determined using Student's t test and a P value of <0.05.
Cell association, invasion, and intracellular replication.
The ability of subsp. IIIa and IIIb to associate with, be internalized by, and replicate inside J774-A.1 macrophages was tested. J774-A.1 cells were propagated in Dulbecco's modified Eagle's medium (Cellgro) supplemented with 10% fetal bovine serum (PAA Laboratories), and plated at a density of 3.5 × 105 cells per well in 24-well dishes for all infections. Bacteria used for infecting J774-A.1 macrophages were grown to stationary phase without aeration in LB broth supplemented with 0.3 M NaCl, conditions that induce TTSS-1 expression (4, 25). J774-A.1 cells were infected with Salmonella at a multiplicity of infection of 50:1 (Salmonella:J774) and incubated for 1 h at 37°C with 5% CO2 in a humidified tissue culture incubator. The actual titer of the inoculum in each experiment was determined by serial dilution and plating on appropriate bacteriologic media. J774-A.1 monolayers were washed three times with 1 ml of sterile PBS prior to lysis, to enumerate cell-associated bacteria, or treated with 100 μg/ml gentamicin sulfate for 1.5 h at 37°C with 5% CO2 in a humidified tissue culture incubator. For enumeration of intracellular bacteria within J774-A.1 cells, gentamicin was removed and monolayers were washed with sterile PBS three times. Infected monolayers were lysed in 1% Triton X-100, and intracellular CFU were enumerated by serial dilution and plating. For assessment of intracellular growth, infected, gentamicin-treated monolayers were washed with sterile PBS and fresh Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum and 10 μg/ml gentamicin was added. Infected monolayers were incubated for 24 h, washed three times with sterile PBS, and lysed, and intracellular CFU were enumerated. At each stage when infected cells were lysed, the number of viable J774-A.1 cells in duplicate monolayers infected with each strain was assessed by 0.4% trypan blue (Cellgro) exclusion and counting viable cells. Each experiment was performed on three separate occasions and by evaluating samples in triplicate.
Analysis of SPI-1 and SPI-2 effector expression and secretion.
In order to determine whether TTSS-1 and TTSS-2 are functional in subsp. IIIa and IIIb, we grew these strains under previously published conditions for TTSS-1 and TTSS-2 expression and determined whether several effectors of this system were being produced and secreted (17, 47). Cultures were normalized by optical density at 600 nm, and bacterial pellets were collected by centrifugation and resuspended in sodium dodecyl sulfate (SDS) sample buffer to generate whole-cell lysate. Proteins in the culture supernatants were precipitated with trichloroacetic acid (TCA) precipitated (6% TCA final concentration) overnight at 4°C. Both proteins from whole-cell lysates and TCA-precipitated proteins from bacteria grown under TTSS-1- and TTSS-2-inducing conditions were separated by SDS-polyacrylamide gel electrophoresis. These proteins were analyzed by Western blotting using specific antisera against the effector proteins SipB (1:10,000 dilution), SipC (1:10,000 dilution), SopB (1:2,000 dilution), SseG (1:2,000 dilution), and SifA (1:2,000 dilution) (antisera were kind gifts of Daoguo Zhou and John Brumell). For detection of primary antibodies we used the WesternDot 625 Western blot kit according to the manufacturer's instructions (Invitrogen).
RESULTS
Subspecies I colonizes murine models better than non-subsp. I salmonellae.
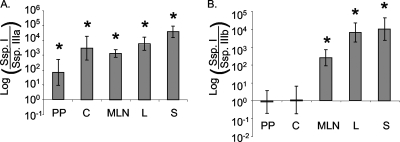
We compared the abilities of subsp. I ATCC 14028 and subsp. IIIa or subsp. IIIb to colonize the organs of orally infected Salmonella-susceptible BALB/c mice 5 days postinfection. The subsp. IIIa isolate we tested, SGSC4693 (HA350), colonized the intestinal tract very poorly compared to subsp. I (Fig. 1A). In the ceca and Peyer's patches of infected mice, subsp. IIIa colonized 100- to 1,000-fold more poorly at this early time after infection than the subsp. I isolate that was concurrently inoculated. Subspecies IIIa also failed to spread systemically, colonizing the liver and spleen in far lower numbers than the subsp. I isolate (concurrent infection). In contrast, subsp. IIIb (SGSC4692) was equally capable of intestinal colonization in Salmonella-susceptible BALB/c mice at 5 days postinfection as subsp. I was (Fig. 1B). Subspecies IIIb colonized the Peyer's patches and cecum in BALB/c mice in numbers equivalent to subsp. I. Despite good colonization of the intestinal tract in BALB/c mice, however, the subsp. IIIb isolate tested here was nearly completely unable to spread systemically and colonized systemic organs very poorly.
FIG. 1.
Non-subspecies I salmonellae colonize BALB/c mice differently than subsp. I ATCC 14028. Mice were infected with an equal mixture of subsp. I (Ssp. I) and non-subsp. I strains and were euthanized 4 days postinfection. Peyer's patches (PP), mesenteric lymph nodes (MLN), spleen (S), liver (L), and cecum (C) were collected, and bacteria were recovered and enumerated. A. Colonization of tissues in BALB/c mice by HA348 (subsp. I) versus HA350 (subsp. IIIa). B. AJB715 (subsp. I) versus HA378 (subsp. IIIb). Data were plotted as logarithmically converted normalized means of output ratios ± standard errors. Statistical significance was determined using a Student's two-tailed t test, and asterisks indicate that normalized output ratios were significantly statistically different from the equivalent ratio in the inoculum with a P value of <0.05.
We assessed intestinal colonization and long-term fecal shedding in Salmonella-resistant mice using intragastric inoculation of groups of six 8-week-old CBA/J mice. We monitored fecal shedding on day 1 and at 3-day intervals for 30 days postinfection. Our results are shown in Fig. 2. Consistent with our experiments in Salmonella-susceptible BALB/c mice (Fig. 1), subsp. IIIa also had a defect in initial intestinal colonization of Salmonella-resistant CBA/J mice compared to subsp. I (Fig. 2A). This colonization defect was followed by severely reduced fecal shedding of subsp. IIIa from Salmonella-resistant mice for 40 days of infection compared to subsp. I ATCC 14028. In contrast, subsp. IIIb colonized the intestine of Salmonella-resistant CBA/J indistinguishably from subsp. I and was shed in the feces at levels similar to subsp. I for over 20 days postinfection (Fig. 2B). After 27 days of infection, subsp. IIIb exhibited a statistically significant defect in fecal shedding compared to ATCC 14028.
FIG. 2.
Non-subsp. I salmonellae differ from subsp. I ATCC 14028 in their ability to persist in the intestine of Salmonella-resistant CBA/J mice. (A) S. enterica subsp. IIIa (HA350) does not persist in the intestine relative to S. enterica subsp. I over 40 days of infection. (B) S. enterica subsp. IIIb (HA378) has an intestinal persistence defect relative to subsp. I. Data are represented as the log (mean of the subsp. I CFU/non-subsp. I CFU ratio) recovered from feces at the indicated time points postinfection. Data were converted logarithmically and displayed graphically, and statistical significance was determined using a Student's two-tailed t test and a P value of <0.05. Error bars denote standard errors, and a * denotes a result statistically significantly different from inoculum.
Cell association, invasion, and intracellular growth in macrophages.
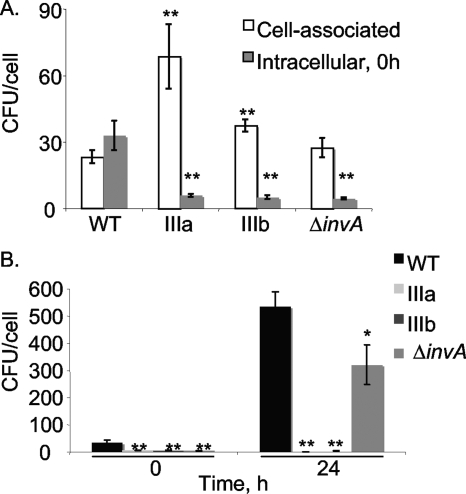
The ability to adhere to, be internalized by, and replicate in eukaryotic cells, notably macrophages, is intimately associated with the pathogenesis of subsp. I (24) and is important for virulence and systemic colonization by this organism (12, 23). Because subsp. IIIa and IIIb were unable to spread systemically in BALB/c mice and colonize systemic organs, we postulated that defects in cell association, internalization, and intracellular growth in macrophages may be responsible for these defects in intestinal colonization and systemic spread. Thus, we tested the ability of subsp. IIIa and subsp. IIIb isolates to associate with, become intracellular in, and replicate in cultured murine macrophages of the J774A.1 lineage. Both subsp. IIIa and IIIb associated with cultured J774 macrophages in higher numbers than subsp. I in our assays (Fig. 3A). Despite this finding, both subsp. IIIa and subsp. IIIb were internalized poorly and at levels similar to our noninvasive serovar Typhimurium ΔinvA mutant (Fig. 3A). In addition, both the subsp. IIIa and IIIb isolates that we studied replicated poorly inside macrophages in our assays, while both the subsp. I isolate used here and the ΔinvA mutant replicated intracellularly (Fig. 3B). Additional experiments showed that the growth of subsp. IIIa and subsp. IIIb during competitive growth in vitro in rich media is indistinguishable from subsp. I ATCC 14028 (data not shown).
FIG. 3.
Subspecies IIIa and IIIb isolates have reduced invasiveness and do not replicate intracellularly in J774 murine macrophages. (A) Attachment and invasion of J774.A1 murine macrophages of virulent wild-type HA348 (WT; subsp. I ATCC 14028 derivative), HA350 (IIIa), HA378 (IIIb), and HA458 (HA348 ΔinvA::Kan) at a multiplicity of infection of 50. (B) Replication of bacteria inside J774-A.1 murine macrophages. Intracellular growth was quantified after a standard gentamicin protection assay as described in Materials and Methods. Data are shown as the means of three experiments, with each assay performed in triplicate, and standard errors. Statistical significance: *, P < 0.05; **, P < 0.001.
Analysis of SPI-1 and SPI-2 secreted effectors.
We used published conditions for the expression of the subsp. I TTSS-1 and TTSS-2 and the secretion of effectors from these systems to determine whether our subspecies IIIa and IIIb isolates were secreting effectors of these systems (Table 2) (45). Bacteria were grown under SPI-1- or SPI-2-expressing conditions (17, 47) and collected by centrifugation, and the secreted proteins in the supernatant were precipitated with TCA. Whole-cell lysates and TCA precipitates were examined by SDS-polyacrylamide gel electrophoresis and Western analysis with antibodies specific to particular effectors of these TTSS.
TABLE 2.
TTSS-1 and TTSS-2 effector homologs in S. enterica subsp. IIIa examined in this study
| TTSS and effector(s) | SARI homolog | Length (aa)a in subspecies:
|
% Identity | % Similarity | |
|---|---|---|---|---|---|
| I (LT2) | IIIa | ||||
| TTSS-1 | |||||
| SipB, SspB | SARI00088 | 593 | 594 | 92 | 94 |
| SipC, SspC | SARI00089 | 409 | 409 | 91 | 94 |
| SopB, SigD | SARI01910 | 561 | 561 | 90 | 94 |
| TTSS-2 | |||||
| SseG | SARI01576 | 228 | 196 | 83 | 91 |
| SifA | SARI01766 | 336 | 336 | 66 | 79 |
aa, amino acids.
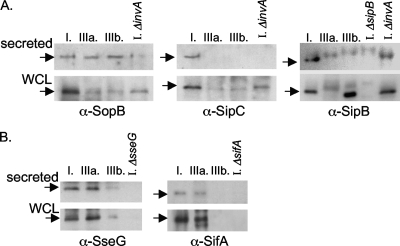
We determined that the TTSS-1 effectors SopB and SipC (see Table 2 for S. enterica subsp. arizonae homologs SARDI01910 and SARDI00089) are present in the whole-cell lysates of subsp. I, IIIa, and IIIb grown under TTSS-1-inducing conditions (Fig. 4A). Of these effectors, we found that SopB and its homologs (SARI01910 in IIIa; IIIb was not sequenced) are secreted into the medium from subsp. I, IIIa, and IIIb under TTSS-1-inducing in vitro conditions. Thus, the TTSS-1 secretion system in all of the subspecies isolates examined here is likely functional and the effector SopB is produced and secreted. In contrast, proteins recognized by anti-SipC antiserum were produced by all the subspecies we examined but they were only secreted by subsp. I. Therefore, although the TTSS-1 appears to be present and functional as shown by the secretion of SopB in all subspecies examined here, a different complement of effectors is being secreted from subsp. I than from the IIIa and IIIb isolates that we studied.
FIG. 4.
TTSS-1 and TTSS-2 effector production and secretion from non-subsp. I isolates. We studied the production and secretion of effectors of TTSS-1 and TTSS-2 by using published conditions for the expression of these systems combined with Western analysis with specific antisera. (A) TTSS-1 effector production and secretion were examined in whole-cell lysates (WCL) and from TCA precipitates of secreted proteins using specific antisera against SopB, SipC, and SipB. (B) TTSS-2 effector protein production and secretion were examined in whole-cell lysates and TCA precipitates of secreted proteins using specific antisera against SseG and SifA.
Both IIIa and IIIb encode a homolog of the sipB gene (Table 2) (41). In our experiments, however, only subsp. I and subsp. IIIb produced a protein that was strongly reactive with anti-SipB antiserum in the whole-cell lysate under TTSS-1-inducing conditions (Fig. 4A). However, this strongly anti-SipB-reactive protein was not secreted from subsp. IIIb into the medium in our experiments. Although subsp. IIIa encodes a homolog with 92% identity and 94% similarity at the protein level to SipB (SARDI000880 (Table 2), subsp. IIIa did not produce a protein that reacted strongly with anti-SipB antiserum in our experiments. Further mutational analysis will determine whether the weakly reactive anti-SipB bands of similar size produced by subsp. IIIa are SipB homologs. To summarize our findings for the TTSS-1 effectors we examined, there was a variable pattern of production and secretion of these effectors from subsp. IIIa and IIIb relative to subsp. I under the in vitro conditions used here.
We also examined the production and secretion of effectors of TTSS-2 in subsp. I, IIIa, and IIIb, using published conditions for TTSS-2 expression and effector secretion. Subspecies IIIa and IIIb encode homologs of the sseG gene, while only subsp. I and subsp. IIIa encode the TTSS-2 effector SifA (45). In our experiments, proteins that reacted with anti-SseG antiserum were produced and secreted from subsp. I, IIIa, and IIIb (Fig. 4B). These data suggest that TTSS-2 in subsp. IIIa and IIIb is functional and that the effector SseG is secreted. Subspecies IIIa produces and secretes a protein that is recognized by anti-SifA antiserum, while subsp. IIIb does not (Fig. 4B). These data are in agreement with the fact that subsp. IIIb does not encode a sifA homolog (45). To summarize, the TTSS-2 of subsp. IIIa and IIIb is likely functional and the effector SseG is secreted by both of these non-subspecies I serovars, while the effector SifA is produced and secreted only from subsp. IIIa.
Motility.
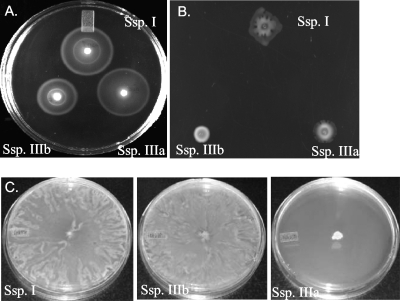
We tested our subsp. IIIa and IIIb isolates for both swimming and swarming motilities by using previously established assay methods. Subspecies IIIb is diphasic and was motile in both our swimming (Fig. 5A) and swarming (Fig. 5B) assays and swarmed to the same diameter at 24 h compared to subsp. I (Fig. 5C). Subspecies IIIa, in contrast, is known to be monophasic and was comparable to subsp. I in swimming motility (Fig. 5A) but was completely unable to swarm (Fig. 5B and C). After 24 h of incubation on motility plates, subsp. I isolates had a mean swarm diameter of 85 mm (standard deviation, 0), while subsp. IIIa had a mean swarm diameter of 6 mm (standard deviation, 0.81 mm) (Student's two-tailed t test, P < 1.5 × 10−9) (Fig. 5C). Using video microscopy we noted that this subsp. IIIa isolate is completely nonmotile on swarming agar (data not shown). Subspecies IIIb, in contrast, is diphasic and was motile in both our swimming and swarming assays and swarmed to the same diameter at 24 h as subsp. I (Fig. 5C).
FIG. 5.
Subspecies (Ssp.) IIIa and IIIb have altered swimming and swarming motility compared to subsp. I. Three microliters of bacterial culture was spotted onto swimming assay plates (0.3% Bacto Agar) (A) or swarming assay plates (0.6% Bacto agar with 0.5% glucose) (B) and incubated for 5 h at 37°C. (C) Three μl of each isolate was spotted on individual swarming plates and incubated for 24 h at 37°C. The diameter of each area of spotted bacteria was measured at both 5 h and 24 h postinoculation.
DISCUSSION
Subspecies I salmonellae cause the vast majority of salmonellosis in mammals and birds, although non-subspecies I serovars are capable of causing sporadic disease. The factors that allow subspecies I salmonellae to be so epidemiologically dominant in mammals and birds are unknown. In this study we determined that the non-subspecies I isolates that we examined, IIIa SGSC4693 and IIIb SGSC4692, poorly colonize murine models that are commonly used to study the pathogenesis of subsp. I salmonellae compared to subsp. I ATCC 14028.
Although the genome sequence of the subsp. IIIa isolate used in these studies has recently become publicly available, the remaining isolates used here are currently being sequenced, making a direct comparison of their genomic content and comprehensive analysis of our observations possible in the near future. Both small-scale gene-by-gene studies and more comprehensive genomic comparisons between subsp. I salmonellae and non-subspecies I salmonellae have been undertaken (8, 15, 19, 27, 34, 36, 40, 41, 45, 46, 52, 53). These studies identified several areas of genomic variability, including fimbrial operons and other adhesins (36, 37, 45), flagellins, and TTSS apparatus and effectors that, along with other loci, may be relevant to our observations.
We have shown that subsp. IIIa SGSC4693 is highly defective in intestinal colonization and persistence compared to subsp. I. One major area of genetic difference between subsp. I and subsp. IIIa is in the complement of fimbriae and other adhesins that are encoded by these subspecies (15, 45, 52). In subsp. I salmonellae, fimbriae are important factors for colonization of mucosal surfaces in the intestine of mammalian hosts (57). Subspecies I LT2 possesses 12 fimbrial operons of the chaperone-usher assembly class, while subspecies IIIa possesses only two of these, fim and bcf, but may have additional still-undefined fimbrial operons (45). Subspecies I mutants with the fim operon deleted have a reduced ability to attach to HeLa cell monolayers (5), but these mutants colonize and persist in the intestine of Salmonella-resistant CBA/J mice to levels very similar to the isogenic wild type (57). Similarly, in subsp. I the bcf operon is unnecessary for colonization of the murine intestine until 20 days postinfection (57), but other fimbrial operons such as std and stb are critical both for colonization and persistence. These studies are consistent with our data that subsp. IIIa is heavily disadvantaged both in initial colonization of the murine intestine and in persistence in this niche, and the two fimbrial operons IIIa is known to possess (fim and bcf) are unimportant for these processes during infection of murine models with subsp. I.
We have also shown that subsp. IIIa is motile individually in liquid media but is completely unable to swarm across the surface of more solid media. Subspecies IIIa is the most divergent subspecies among S. enterica and is monophasic (45). The ability of subsp. IIIa to colonize the intestinal tract may also be limited by its lack of motility in environments of high viscosity. Perhaps this isolate is unable to penetrate viscous environment of the mucous layer covering the intestinal epithelium, although this remains to be shown.
In contrast to subsp. IIIa, subsp. IIIb SGSC4692 colonizes the intestine well in the murine models used in our study but has a persistence defect in this niche. Subspecies IIIb is known to possess four fimbrial operons, fim, bcf, std, and stb, but may also have additional, still-unidentified fimbrial operons (45). Clinical isolates of subsp. IIIb have also been reported to have agfA (43). In subsp. I deletions of the std or stb fimbrial operons result in strains that are defective for initial colonization of the intestine and for persistence in this niche (57). Our data, and those of others showing that subsp. IIIb possesses the std and stb operons and colonizes the murine intestinal tract well, are consistent with the hypothesis that fimbria encoded by stb and std are very important for intestinal colonization. Subspecies IIIb also lacks the lpf operon, which is important for intestinal persistence after 20 days of infection in murine models and for biofilm formation on Hep-2 cells and explanted chicken intestinal tissue (39, 57). The apparent lack of lpf in subsp. IIIb may contribute to the mild intestinal persistence defect of this isolate.
Other adhesins known to be important for intestinal persistence in subsp. I include proteins encoded at the CS54 island by shdA (STM2513), the neighboring gene ratB (STM2514), and a third protein chromosomally encoded elsewhere by misL (STM3757). ShdA and MisL are autotransporter proteins that bind extracellular matrix components in vitro (21, 35). Mutants for misL and shdA in subsp. I show reduced colonization of the cecum and/or Peyer's patches in BALB/c mice at 5 days postinfection (21, 36). These mutants also have very late defects (21 to 25 days postinfection) in intestinal persistence in Salmonella-resistant CBA/J mice (21, 36). ratB mutants in subsp. I colonize the intestine well but are highly defective for intestinal persistence beginning at 5 days postinfection (36). shdA, ratB, and misL are absent in the subsp. IIIa and subsp. IIIb isolates that have been examined, including the subsp. IIIa isolate used in our studies (36, 37, 45). These data are consistent with our observations that subsp. IIIa is defective for colonization and persistence in the intestinal tract of mice. However, despite the apparent lack of these factors in subsp. IIIb, the subsp. IIIb isolate tested here was not defective for colonization of the Peyer's patches and cecum in BALB/c mice (21, 36, 37) and had a milder persistence defect in identical animal models than either misL, shdA, or ratB mutants of subsp. I. Our subsp. IIIb isolate may possess alternate adhesins that partially complement the functions of MisL, ShdA, and RatB, and we may be able to determine this when the complete genome sequence for this isolate is completed.
The ability to replicate in macrophages is important for subsp. I to colonize systemic sites in the mouse (12, 23). Both the subsp. IIIa and subsp. IIIb isolates we tested failed to spread beyond the intestine and colonize systemic organs in murine models, and we hypothesized that subsp. IIIa and IIIb may be unable to replicate in macrophages. We showed that these isolates adhere to J774-A.1 murine macrophages in culture, yet they are not internalized and do not replicate once intracellular. The SPI-1-encoded TTSS-1 and the SPI-2-encoded TTSS-2 are essential for cellular invasion and for intracellular replication, abilities that are important for salmonellae to invade host cells and spread systemically. Both subsp. IIIa and subsp. IIIb possess SPI-1 and SPI-2, but the genetic islands encoding these TTSS, the fidelity of these secretion systems, and the distribution, production, and secretion of the effectors of these systems, and the spv genes, have not been extensively studied outside of subsp. I salmonellae (15, 24, 29, 30, 45).
Recent reports have indicated that subsp. IIIa either lacks or has several divergent components of TTSS-1 encoded in SPI-1, including invA and invH (15). avrA, an effector of the TTSS-1 that inhibits interleukin-8 production by inhibiting NF-κΒ signaling as well as preventing the ubiquitination of β-catein, is also absent from subsp. IIIa (16, 22, 33, 41). We showed that the TTSS-1 effector SopB, an inositol phosphatase that activates CDC42, Rac1, RhoG, AktA, and chloride secretion and disrupts tight junctions (9, 38, 42, 44, 59), is produced and secreted from subsp. IIIa in vitro, indicating that the TTSS-1 is likely functional in subsp. IIIa in our experiments. Despite its presence in whole-cell lysate, we could not detect the secretion of SipC, a second TTSS-1 effector that nucleates and bundles actin (13, 28, 48) under in vitro conditions that allow the secretion of this protein from subsp. I. Finally, the TTSS-1 effector SipB, which binds to and activates caspase-1 to induce autophagy and apoptosis in macrophages (31, 32), does not appear to be produced or secreted from subsp. IIIa in our experiments. Thus, the TTSS-1 of subsp. IIIa is functional but secretes a different complement of effectors than the TTSS-1 of subsp. I under similar conditions. A more complete analysis of the presence or absence of TTSS-1 effector homologs and their production and secretion in vivo from subsp. IIIa may provide additional insight into the inability of subsp. IIIa to colonize the murine intestine.
Genetic variability in SPI-2, encoding the TTSS-2, in subsp. IIIa has also been reported recently (15). We showed that the TTSS-2 effector SseG, a participant in Salmonella-induced filament (Sif) formation and microtubule bundling during the intracellular growth of subsp. I, is produced and secreted from subsp. IIIa. A second effector of the TTSS-2, SifA, involved in inducing Sif formation and maintenance of the Salmonella-containing vacuoule, is also produced and secreted from subsp. IIIa. These data strongly suggest that the TTSS-2 machinery is functional in our in vitro experiments in subsp. IIIa. A more complete comparison of the presence and primary sequence of the 20 or so known effector protein homologs of the TTSS-2 in subsp. IIIa may yield additional clues as to why subsp. IIIa is unable to replicate intracellularly in macrophages and spread systemically in murine models of infection.
SPI-1 and SPI-2 regions encoding the TTSS-1 and TTSS-2 secretion systems themselves have not been extensively studied in subsp. IIIb beyond microarray analysis for genomic content (45). Subspecies IIIb is known to encode homologs for the following TTSS-1 effectors: SipA, SipB, SipC, AvrA, SopB, and SopD (41), but the presence of the remaining known effectors of TTSS-1 has been studied only by microarray analysis. The gene for the TTSS-1 effector SopA appears to be missing or divergent in subsp. IIIb in these microarray studies (45). We showed that subsp. IIIb produces and secretes SopB, suggesting that the TTSS-1 is functional in our assays. However, although subsp. IIIb produced two additional effector proteins, SipB and SipC, these proteins were not detectable in the secreted fraction. This finding indicates that despite a functional TTSS-1, subsp. IIIb secretes a different complement of effectors than subsp. I. Further work will be necessary to determine whether a unique complement of TTSS-1 effectors is secreted from subsp. IIIb during infection and how this may affect intestinal colonization and persistence.
Finally, subsp. IIIb produced and secreted the TTSS-2 effector SseG, indicating that the TTSS-2 apparatus is also functional in our subsp. IIIb isolate. An additional effector called SifA does not have a homolog in subsp. IIIb (45). In agreement with this published finding, SifA was not detected in whole-cell lysates in our experiments. Thus, the TTSS-2 in subsp. IIIb also secretes a different complement of effectors, lacking at least one effector known to be important for intracellular growth of subsp. I. If variability in effector production and secretion also occurs in vivo, it may contribute to the inability of IIIb to grow intracellularly and to its failure to colonize systemic sites in Salmonella-susceptible BALB/c mice. Completion of the genome sequence and subsequent complete analysis of the TTSS-2 effector homologs present in subsp. IIIb should provide additional insight into the differences between subsp. I and subsp. IIIb TTSS-2 effector complements present in both organisms.
The non-subsp. I Salmonella isolates we studied in this work colonize the intestine differently than subsp. I, and they are defective for fecal shedding and intestinal persistence. This altered colonization of the murine intestinal tract is likely the result of multiple factors, including different complement of fimbrial operons and adhesins and potentially differential production and secretion of effectors of the TTSS in non-subsp. I isolates relative to subsp. I. The inabilities of subsp. IIIa and IIIb to colonize and/or persist in the intestinal tract are likely to be partially responsible for the failure of non-subsp. I isolates to circulate in populations of warm-blooded vertebrates relative to subsp. I salmonellae, as these factors are critical to transmission of salmonellae from host to host. Furthermore, we showed that subsp. IIIa and IIIb are strongly defective for systemic colonization. The inability of the subsp. IIIa and IIIb isolates tested here to access systemic sites may be partially the result of different complements of effectors of the type three secretion systems in these isolates relative to subsp. I and may reduce the ability of these isolates to persist long term in an infected host, thus reducing transmission of these subspecies in warm-blooded vertebrates.
Acknowledgments
We thank Michael McClelland for the kind gift of subsp. IIIb isolate SGSC4692. We thank Daoguo Zhou and John Brumell for the kind gift of antibodies to TTSS-1 and TTSS-2 effectors. We thank M. Megan Reynolds, Andrew Bugbee, and Christine Shields for technical assistance.
This work was supported by the Texas A&M University System start-up funding awarded to H.A.-P.
Footnotes
Published ahead of print on 13 February 2009.
REFERENCES
- 1.Alvseike, O., and E. Skjerve. 2002. Prevalence of a Salmonella subspecies diarizonae in Norwegian sheep herds. Prev. Vet. Med. 52277-285. [DOI] [PubMed] [Google Scholar]
- 2.Alvseike, O., T. Vardund, B. Lindstedt, E. Heir, E. Eriksson, and G. Kapperud. 2004. Molecular epidemiology and population genetics of Salmonella subspecies diarizonae in sheep in Norway and Sweden. Epidemiol. Infect. 132253-261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Andrews-Polymenis, H. L., W. Rabsch, S. Porwollik, M. McClelland, C. Rosetti, L. G. Adams, and A. J. Baumler. 2004. Host restriction of Salmonella enterica serotype Typhimurium pigeon isolates does not correlate with loss of discrete genes. J. Bacteriol. 1862619-2628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bajaj, V., R. L. Lucas, C. Hwang, and C. A. Lee. 1996. Co-odinate regulation of Salmonella typhimurium invasion genes by environmental factors is mediated by control of hilA expression. Mol. Microbiol. 22703-714. [DOI] [PubMed] [Google Scholar]
- 5.Bäumler, A. J., R. M. Tsolis, and F. Heffron. 1996. Contribution of fimbrial operons to attachment to and invasion of epithelial cell lines by Salmonella typhimurium. Infect. Immun. 641862-1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bemis, D. A., L. M. Grupka, S. Liamthong, D. W. Folland, J. M. Sykes, and E. C. Ramsay. 2007. Clonal relatedness of Salmonella isolates associated with invasive infections in captive and wild-caught rattlesnakes. Vet. Microbiol. 120300-307. [DOI] [PubMed] [Google Scholar]
- 7.Boyd, E. F., F.-S. Wang, T. S. Whittam, and R. K. Selander. 1996. Molecular genetic relationship of the salmonellae. Appl. Environ. Microbiol. 62804-808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Boyd, F. E., and D. L. Hartl. 1998. Salmonella virulence plasmid: modular acquisition of the spv virulence region by an F-plasmid in Salmonella enterica subspecies I and insertion into the chromosome in subspecies II, IIIa, IV, and VII isolates. Genetics 1491183-1190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Boyle, E. C., N. F. Brown, and B. B. Finlay. 2006. Salmonella enterica serovar Typhimurium effectors SopB, SopE, SopE2 and SipA disrupt tight junction structure and function. Cell. Microbiol. 81946-1957. [DOI] [PubMed] [Google Scholar]
- 10.Brenner, F. W., R. G. Villar, F. J. Angulo, R. Tauxe, and B. Swaminathan. 2000. Salmonella nomenclature. J. Clin. Microbiol. 382465-2467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Briones, V., S. Tellez, J. Goyache, C. Ballesteros, M. del Pilar Lanzarot, L. Dominguez, and J. F. Fernandez-Garayzabal. 2004. Salmonella diversity associated with wild reptiles and amphibians in Spain. Environ. Microbiol. 6868-871. [DOI] [PubMed] [Google Scholar]
- 12.Buchmeier, N. A., and F. Heffron. 1989. Intracellular survival of wild-type Salmonella typhimurium and macrophage-sensitive mutants in diverse populations of macrophages. Infect. Immun. 571-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Carlson, S. A., M. B. Omary, and B. D. Jones. 2002. Identification of cytokeratins as accessory mediators of Salmonella entry into eukaryotic cells. Life Sci. 701415-1426. [DOI] [PubMed] [Google Scholar]
- 14.Centers for Disease Control and Prevention. 1999. Reptile-associated salmonellosis—selected states, 1996-1998. Morbid. Mortal. Wkly. Rep. 481009-1013. [PubMed] [Google Scholar]
- 15.Chan, K., S. Baker, C. C. Kim, C. S. Detweiler, G. Dougan, and S. Falkow. 2003. Genomic comparison of Salmonella enterica serovars and Salmonella bongori by use of an S. enterica serovar Typhimurium DNA microarray. J. Bacteriol. 185553-563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Collier-Hyams, L. S., H. Zeng, J. Sun, A. D. Tomlinson, Z. Q. Bao, H. Chen, J. L. Madara, K. Orth, and A. S. Neish. 2002. Cutting edge: Salmonella AvrA effector inhibits the key proinflammatory, anti-apoptotic NF-kappa B pathway. J. Immunol. 1692846-2850. [DOI] [PubMed] [Google Scholar]
- 17.Coombes, B. K., N. F. Brown, Y. Valdez, J. H. Brumell, and B. Finlay. 2004. Expression and secretion of Salmonella pathogenicity island-2 virulence genes in response to acidification exhibit differential requirements of a functional type III secretion apparatus and SsaL. J. Biol. Chem. 27949804-49815. [DOI] [PubMed] [Google Scholar]
- 18.Crespo, R., J. S. Jeffrey, R. P. Chin, G. Senties-Cue, and H. L. Shivaprasad. 2004. Phenotypic and genotypic characterization of Salmonella arizonae from an integrated turkey operation. Avian Dis. 48344-350. [DOI] [PubMed] [Google Scholar]
- 19.Crosa, J. H., D. J. Brenner, W. H. Ewing, and S. Falkow. 1973. Molecular relationships among the salmonelleae. J. Bacteriol. 115307-315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Datsenko, K. A., and B. L. Wanner. 2000. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 976640-6645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dorsey, C. W., M. C. Laarakker, A. D. Humphries, E. H. Weening, and A. J. Baumler. 2005. Salmonella enterica serotype Typhimurium MisL is an intestinal colonization factor that binds fibronectin. Mol. Microbiol. 57196-211. [DOI] [PubMed] [Google Scholar]
- 22.Ehrbar, K., S. Mirold, A. Friebel, S. Stender, and W. D. Hardt. 2002. Characterization of effector proteins translocated via the SPI1 type III secretion system of Salmonella typhimurium. Int. J. Med. Microbiol. 291479-485. [DOI] [PubMed] [Google Scholar]
- 23.Fields, P. I., R. V. Swanson, C. G. Haidaris, and F. Heffron. 1986. Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Natl. Acad. Sci. USA 835189-5193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Galán, J. E., and R. Curtiss III. 1989. Cloning and molecular characterization of genes whose products allow Salmonella typhimurium to penetrate tissue culture cells. Proc. Natl. Acad. Sci. USA 866383-6387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Galan, J. E., and R. Curtiss III. 1990. Expression of Salmonella typhimurium genes required for invasion is regulated by changes in DNA supercoiling. Infect. Immun. 581879-1885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hall, M. L., and B. Rowe. 1992. Salmonella arizonae in the United Kingdom from 1966 to 1990. Epidemiol. Infect. 10859-65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hansen-Wester, I., and M. Hensel. 2002. Genome-based identification of chromosomal regions specific for Salmonella spp. Infect. Immun. 702351-2360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hayward, R. D., and V. Koronakis. 1999. Direct nucleation and bundling of actin by the SipC protein of invasive Salmonella. EMBO J. 184926-4934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hensel, M., J. E. Shea, C. Gleeson, M. D. Jones, E. Dalton, and D. W. Holden. 1995. Simultaneous identification of bacterial virulence genes by negative selection. Science 269400-403. [DOI] [PubMed] [Google Scholar]
- 30.Hensel, M., J. E. Shea, S. R. Waterman, R. Mundy, T. Nikolaus, G. Banks, A. Vazquez-Torres, C. Gleeson, F. C. Fang, and D. W. Holden. 1998. Genes encoding putative effector proteins of the type III secretion system of Salmonella pathogenicity island 2 are required for bacterial virulence and proliferation in macrophages. Mol. Microbiol. 30163-174. [DOI] [PubMed] [Google Scholar]
- 31.Hernandez, L. D., M. Pypaert, R. A. Flavell, and J. E. Galan. 2003. A Salmonella protein causes macrophage cell death by inducing autophagy. J. Cell Biol. 1631123-1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hersh, D., D. M. Monack, M. R. Smith, N. Ghori, S. Falkow, and A. Zychlinsky. 1999. The salmonella invasin SipB induces macrophage apoptosis by binding to caspase-1. Proc. Natl. Acad. Sci. USA 962396-2401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Jones, R. M., H. Wu, C. Wentworth, L. Luo, L. Collier-Hyams, and A. S. Neish. 2008. Salmonella AvrA coordinates suppression of host immune and apoptotic defenses via JNK pathway blockade. Cell Host Microbe 3233-244. [DOI] [PubMed] [Google Scholar]
- 34.Kim, H. J., S. H. Park, and H. Y. Kim. 2006. Comparison of Salmonella enterica serovar Typhimurium LT2 and non-LT2 Salmonella genomic sequences and genotyping of salmonellae by using PCR. Appl. Environ. Microbiol. 726142-6151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kingsley, R. A., D. Abi Ghanem, N. Puebla-Osorio, A. M. Keestra, L. Berghman, and A. J. Baumler. 2004. Fibronectin binding to the Salmonella enterica serotype Typhimurium ShdA autotransporter protein is inhibited by a monoclonal antibody recognizing the A3 repeat. J. Bacteriol. 1864931-4939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kingsley, R. A., A. D. Humphries, E. H. Weening, M. R. De Zoete, S. Winter, A. Papaconstantinopoulou, G. Dougan, and A. J. Baumler. 2003. Molecular and phenotypic analysis of the CS54 island of Salmonella enterica serotype Typhimurium: identification of intestinal colonization and persistence determinants. Infect. Immun. 71629-640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kingsley, R. A., K. van Amsterdam, N. Kramer, and A. J. Baumler. 2000. The shdA gene is restricted to serotypes of Salmonella enterica subspecies I and contributes to efficient and prolonged fecal shedding. Infect. Immun. 682720-2727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Knodler, L. A., B. B. Finlay, and O. Steele-Mortimer. 2005. The Salmonella effector protein SopB protects epithelial cells from apoptosis by sustained activation of Akt. J. Biol. Chem. 2809058-9064. [DOI] [PubMed] [Google Scholar]
- 39.Ledeboer, N. A., J. G. Frye, M. McClelland, and B. D. Jones. 2006. Salmonella enterica serovar Typhimurium requires the Lpf, Pef, and Tafi fimbriae for biofilm formation on HEp-2 tissue culture cells and chicken intestinal epithelium. Infect. Immun. 743156-3169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Libby, S. J., M. Lesnick, P. Hasegawa, M. Kurth, C. Belcher, J. Fierer, and D. G. Guiney. 2002. Characterization of the spv locus in Salmonella enterica serovar Arizona. Infect. Immun. 703290-3294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mirold, S., K. Ehrbar, A. Weissmuller, R. Prager, H. Tschape, H. Russmann, and W. D. Hardt. 2001. Salmonella host cell invasion emerged by acquisition of a mosaic of separate genetic elements, including salmonella pathogenicity island 1 (SPI1), SPI5, and sopE2. J. Bacteriol. 1832348-2358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Norris, F. A., M. P. Wilson, T. S. Wallis, E. E. Galyov, and P. W. Majerus. 1998. SopB, a protein required for virulence of Salmonella dublin, is an inositol phosphate phosphatase. Proc. Natl. Acad. Sci. USA 9514057-14059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Pasmans, F., A. Martel, F. Boyen, D. Vandekerchove, I. Wybo, F. V. Immerseel, M. Heyndrickx, J. M. Collard, R. Ducatelle, and F. Haesebrouck. 2005. Characterization of Salmonella isolates from captive lizards. Vet. Microbiol. 110285-291. [DOI] [PubMed] [Google Scholar]
- 44.Patel, J. C., and J. E. Galan. 2006. Differential activation and function of Rho GTPases during Salmonella-host cell interactions. J. Cell Biol. 175453-463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Porwollik, S., R. M. Wong, and Michael McClelland. 2002. Evolutionary genomics of Salmonella: gene acquisitions revealed by microarray analysis. Proc. Natl. Acad. Sci. 998956-8961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Prager, R., S. Mirold, E. Tietze, U. Strutz, B. Knuppel, W. Rabsch, W. D. Hardt, and H. Tschape. 2000. Prevalence and polymorphism of genes encoding translocated effector proteins among clinical isolates of Salmonella enterica. Int. J. Med. Microbiol. 290605-617. [DOI] [PubMed] [Google Scholar]
- 47.Raffatellu, M., R. P. Wilson, D. Chessa, H. Andrews-Polymenis, Q. T. Tran, S. Lawhon, S. Khare, L. G. Adams, and A. J. Baumler. 2005. SipA, SopA, SopB, SopD, and SopE2 contribute to Salmonella enterica serotype Typhimurium invasion of epithelial cells. Infect. Immun. 73146-154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Scherer, C. A., E. Cooper, and S. I. Miller. 2000. The Salmonella type III secretion translocon protein SspC is inserted into the epithelial cell plasma membrane upon infection. Mol. Microbiol. 371133-1145. [DOI] [PubMed] [Google Scholar]
- 49.Schroter, M., P. Roggentin, J. Hofmann, A. Speicher, R. Laufs, and D. Mack. 2004. Pet snakes as a reservoir for Salmonella enterica subsp. diarizonae (serogroup IIIb): a prospective study. Appl. Environ. Microbiol. 70613-615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Shivaprasad, H. L., P. Cortes, and R. Crespo. 2006. Otitis interna (labyrinthitis) associated with Salmonella enterica arizonae in turkey poults. Avian Dis. 50135-138. [DOI] [PubMed] [Google Scholar]
- 51.Toguchi, A., M. Siano, M. Burkart, and R. M. Harshey. 2000. Genetics of swarming motility in Salmonella enterica serovar Typhimurium: critical role for lipopolysaccharide. J. Bacteriol. 1826308-6321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Townsend, S. M., N. E. Kramer, R. Edwards, S. Baker, N. Hamlin, M. Simmonds, K. Stevens, S. Maloy, J. Parkhill, G. Dougan, and A. J. Bäumler. 2001. Salmonella enterica serotype Typhi possesses a unique repertoire of fimbrial gene sequences. Infect. Immun. 692894-2901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Tsolis, R. M., S. M. Townsend, E. A. Miao, S. I. Miller, T. A. Ficht, L. G. Adams, and A. J. Baumler. 1999. Identification of a putative Salmonella enterica serotype Typhimurium host range factor with homology to IpaH and YopM by signature-tagged mutagenesis. Infect. Immun. 676385-6393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Washington University. 2008. Salmonella enterica subspecies IIIa (Arizonae) serovar 62:z4,z23:-. Washington University Genome Sequencing Center, St. Louis, MO. http://genome.wustl.edu/genome.cgi?GENOME=Salmonella%20enterica%20subspecies%20IIIa%20(Arizonae)%20serovar%2062:z4,z23:-.
- 55.Washington University. 2008. Salmonella enterica subspecies IIIa (Arizonae) serovar 62:z4,z23:-. Washington University Genome Sequencing Center, St. Louis, MO. http://genome.wustl.edu/genome.cgi?GENOME=Salmonella%20enterica%20subspecies%20IIIa%20(Diarizonae)%20serovar%2061:1,v:1,5,(7).
- 56.Waterman, S. H., G. Juarez, S. J. Carr, and L. Kilman. 1990. Salmonella arizona infections in Latinos associated with rattlesnake folk medicine. Am. J. Public Health 80286-289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Weening, E. H., J. D. Barker, M. C. Laarakker, A. D. Humphries, R. M. Tsolis, and A. J. Baumler. 2005. The Salmonella enterica serotype Typhimurium lpf, bcf, stb, stc, std, and sth fimbrial operons are required for intestinal persistence in mice. Infect. Immun. 733358-3366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Weiss, S. H., M. J. Blase, F. P. Paleologo, R. E. Black, A. C. McWorther, M. A. Asbury, G. P. Carter, R. A. Feldman, and D. J. Brenner. 1986. Occurrence and distribution of serotypes of the Arizona subgroup of Salmonella strains in the United States from 1967 to 1976. J. Clin. Microbiol. 231056-1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhou, D., L. M. Chen, L. Hernandez, S. B. Shears, and J. E. Galan. 2001. A Salmonella inositol polyphosphatase acts in conjunction with other bacterial effectors to promote host cell actin cytoskeleton rearrangements and bacterial internalization. Mol. Microbiol. 39248-259. [DOI] [PubMed] [Google Scholar]