Abstract
The protein kinase mammalian Sterile 20-like kinase 1 (MST1) plays a critical role in the regulation of cell death. Recent studies suggest that MST1 mediates oxidative stress-induced neuronal cell death by phosphorylating the transcription factor FOXO3 at serine 207, a site that is conserved in other FOXO family members. Here, we show that MST1-induced phosphorylation of FOXO1 at serine 212, corresponding to serine 207 in FOXO3, disrupts the association of FOXO1 with 14-3-3 proteins. Accordingly, MST1 mediates the nuclear translocation of FOXO1 in primary rat cerebellar granule neurons that are deprived of neuronal activity. We also find a requirement for MST1 in cell death of granule neurons upon withdrawal of growth factors and neuronal activity, and MST1 induces cell death in a FOXO1-dependent manner. Finally, we show that the MST1-regulatory, scaffold protein Nore1 is required for survival factor deprivation induced neuronal death. Collectively, these findings define MST1-FOXO1 signaling as an important link survival factor deprivation-induced neuronal cell death with implications for our understanding of brain development and neurological diseases.
During normal development, neurons die by the process of apoptosis to ensure the proper wiring of the nervous system (1, 2). In the mature nervous system, aberrant neuronal cell death contributes to the pathogenesis of a large number of diseases (3–5). Therefore, elucidation of the molecular underpinnings of neuronal cell death is essential for our understanding of brain development and also offers the possibility of identification of targets for the development of drugs that prevent neuronal degeneration in brain diseases.
Granule neurons of the rat cerebellum provide a robust system for the study of the mechanisms that govern neuronal cell death (6, 7). The survival of cerebellar granule neurons during normal development is promoted by growth factors and neuronal activity (8–12). Likewise, the survival of primary rat cerebellar granule neurons is supported by polypeptide growth factors, provided by serum and neuronal activity, and mimicked by the activation of voltage-sensitive calcium channels induced by membrane depolarization (13–15). Granule neurons have also proved useful in the study of mechanisms of neuronal cell death in response to pathologically relevant stimuli such as oxidative stress (16).
The protein kinase mammalian Sterile 20-like kinase 1 (MST1)5 has been implicated in the control of neuronal cell death (16, 17). Exposure of rat cerebellar granule neurons to hydrogen peroxide leads to MST1-dependent cell death (16). Oxidative stress-induced MST1 triggers neuronal cell death via the transcription factor FOXO3. MST1 catalyzes the phosphorylation of FOXO3 at serine 207, a site that lies within its forkhead domain, and thereby promotes the dissociation of FOXO3 from 14-3-3 proteins. The 14-3-3 proteins sequester FOXO3 in the cytoplasm and thus inhibit FOXO3-dependent transcription and cell death. The consequence of the MST1-induced phosphorylation of FOXO3 in neurons treated with hydrogen peroxide is the accumulation of FOXO3 in the nucleus and activation of neuronal cell death (16).
A major question raised by the identification of the MST1-FOXO3 link is whether MST1 also signals to the nucleus via other members of the FOXO family of transcription factors in neurons. The FOXO proteins have overlapping but also distinct functions and regulatory mechanisms (18–21). Thus, the identification of a kinase phosphorylating and regulating a FOXO family member does not automatically mean that another family member will be similarly regulated. Among the FOXO3-related proteins, FOXO1 and FOXO6 are expressed in neurons (22). Although nucleocytoplasmic shuttling does not appear to play a major role in the regulation of FOXO6 function (23), shuttling between the cytoplasm and nucleus is critical to FOXO1 regulation (24).
Recent studies have defined a Cdk1-FOXO1 signaling link in postmitotic neurons whereby neuronal activity deprivation triggers the Cdk1-induced phosphorylation of FOXO1 at a critical regulatory site and releases FOXO1 from 14-3-3 proteins (25, 26). In a similar manner to MST1-FOXO3 signaling, the Cdk1-induced phosphorylation of FOXO1 leads to the accumulation of FOXO1 in the nucleus and the induction of cell death (16, 25). These observations raise the question of whether MST1 might phosphorylate FOXO1 and thus contribute to neuronal cell death upon the withdrawal of survival stimuli.
In this study, we identify FOXO1 as a critical substrate of MST1. MST1 phosphorylates FOXO1 within its forkhead domain at serine 212 in vitro and in vivo. The MST1-induced phosphorylation of FOXO1 at serine 212 triggers the disruption of FOXO1 from 14-3-3 proteins. We also find a requirement for MST1 in the ability of activity deprivation to induce the nuclear accumulation of FOXO1 in granule neurons. Consistent with these results, we identify a function for MST1 in granule neuron cell death induced by the deprivation of growth factors and neuronal activity. Finally, we uncover a novel dual function for the MST1 regulatory scaffold protein Nore1 in granule neuron cell death. Remarkably, whereas Nore1 dampens the ability of oxidative stress to induce neuronal death, we find a requirement for Nore1 in survival factor deprivation-induced neuronal death. Taken together, our findings define a crucial function for MST1 in survival factor deprivation-induced neuronal death. We have also characterized upstream and downstream mechanisms of MST1 function in survival factor deprivation-induced neuronal death. Identification of FOXO1 as a physiologically relevant substrate in primary neurons provides a compelling link between MST1 and the FOXO family of transcription factors with important implications for brain development and disease.
EXPERIMENTAL PROCEDURES
Plasmids and Antibodies—The plasmids encoding GST-FOXO1-forkhead domain (GST-FOXO1(FD)) were generated by PCR cloning in pGEX4T1 at the EcoRI and XhoI restriction sites. The FOXO1(FD) S212A, 3A, and 4A plasmids (where 3A refers to the 212A, S234A, S235A FOXO1(FD) mutant, and 4A refers to the S212A, S218A, S234A, and S235A FOXO1(FD) mutant) were generated by site-directed mutagenesis. All mutations were verified by sequencing. The Nore1 short hairpin RNAs (shRNA) targeting sequence is conserved in mice and rats and corresponds to: GAAGATTGACAGCTATAACAGC. Antibodies to MST1 (Zymed Laboratories Inc.); GFP (Molecular Probes); GST, 14-3-3β (Santa Cruz Biotechnology); FLAG-M2 (Sigma); and β-galactosidase (Promega) were purchased. The rabbit antibody to Nore1 was generated by injecting New Zealand rabbits with full-length His-Nore1 and purified over a protein A column (Pierce).
Neuronal Cultures—Cultures of cerebellar granule neurons were prepared from postnatal day 6 rat (P6) pups as described (27). For RNAi experiments, cultures of P6 granule neurons were transfected by a calcium phosphate method after 2 days in vitro (P6 + 2DIV) with the MST1 RNAi or control U6 plasmid together with a plasmid encoding β-galactosidase. Following incubation with conditioned medium containing calf serum (HyClone) and membrane-depolarizing concentrations of KCl (30 mm) for 3 days, cultures were washed and returned to conditioned medium or placed in medium deprived of serum- and membrane-depolarizing concentrations of KCl (5 mm) for 24 h. Cultures were fixed with paraformaldehyde and subjected to indirect immunofluorescence as described (28). Cell survival was assessed in β-galactosidase-expressing neurons based on the integrity of neurites and nuclear morphology visualized using the DNA dye bisbenzimide (Hoechst 33258; Sigma). Although healthy neurons harbor robust processes and intact nuclei, dying neurons have disintegrated and degenerating neuronal processes and pyknotic or fragmented nuclei (16, 25, 28, 29). Cell counts were carried out in a blinded manner and analyzed for statistical significance by ANOVA and Fisher's protected least significance difference (PLSD) post hoc test. Approximately 150 cells were counted per experiment.
In analyses of FOXO1 localization, neurons were transfected after 2 days in vitro (P6 + 2DIV) with the MST1 RNAi or control U6 plasmid together with the GFP-FOXO1 expression plasmid. After 24 h, neurons were placed in full medium (serum + 30 mm KCl) or in medium in which membrane depolarization was inhibited (serum + 5 mm KCl). Cells were fixed 30 h later and analyzed by fluorescence microscopy.
Immunoprecipitation and Immunoblotting—Immunoprecipitation and immunoblotting were carried out as described (16, 25). Cells were lysed in a buffer containing 50 mm HEPES (pH 7.5), 150 mm NaCl, 10% (v/v) glycerol, 1% Triton X-100, 1.5 mm MgCl2, 1 mm EGTA, 100 mm NaF, 1 mm dithiothreitol, 1 mm sodium vanadate, 2 mm phenylmethylsulfonyl fluoride, and 2 mg/ml aprotinin and leupeptin. Lysates were next centrifuged at 14,000 × g for 15 min at 4 °C prior to immunoprecipitation or immunoblotting. In immunoprecipitation analyses, lysates were precleared with protein A beads at 4 °C for 20 min, and supernatants were incubated with appropriate antibodies in the presence of 10 ml of protein A-protein G (2:1) agarose beads for at least 2 h at 4 °C. Beads were washed with lysis buffer, and the immunoprecipitates were subjected to immunoblotting.
In Vitro Kinase Assays—In vitro kinase assays were carried out as described (30). Immunoprecipitated MST1 or recombinant active MST1 was incubated in the following conditions: 20 mm Tris, pH 7.5, 2 mm MgCl2, 1 mm dithiothreitol, and 1 μg of full-length GST-FOXO1 or GST-FOXO1-FD as the substrate. Kinase reactions were separated by SDS-PAGE gel electrophoresis and analyzed by autoradiography or immunoblotting using the phospho-FOXO antibody.
RESULTS AND DISCUSSION
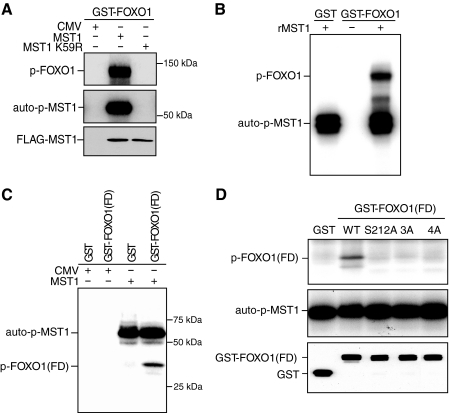
MST1 Phosphorylates FOXO1 at Serine 212 in Vitro and in Vivo—We investigated whether the transcription factor FOXO1 might play a role in propagating the MST1 signal to the nucleus in neurons. First, we asked whether MST1 phosphorylates full-length FOXO1 in vitro. We expressed MST1 tagged with the FLAG epitope or a kinase-dead FLAG-tagged MST1 in which the ATP binding site was mutated (MST1 K59R) in 293T cells. MST1 or MST1 K59R were immunoprecipitated from lysates of transfected cells using the FLAG antibody and subjected to an in vitro kinase assay in which recombinant full-length FOXO1 fused to GST (GST-FOXO1) and 32P-labeled ATP were added. MST1, but not the kinase-dead MST1, robustly phosphorylated FOXO1 (Fig. 1A). In other experiments, we found that purified recombinant MST1 also phosphorylated FOXO1 in vitro (Fig. 1B).
FIGURE 1.
MST1 phosphorylates FOXO1 in vitro. A, lysates of 293T cells transfected with an expression plasmid encoding FLAG-MST1, kinase-dead FLAG-MST1 K59R, or the control plasmid were immunoprecipitated with the FLAG antibody and subjected to an in vitro kinase assay using full-length GST-FOXO1 as substrate in the presence of 32P-labeled ATP. Reactions were analyzed by SDS-PAGE followed by autoradiography. Auto-p-MST1 denotes autophosphorylated MST1; p-FOXO1 denotes phosphorylated FOXO1. The lower panels show the expression of FLAG-MST1 and FLAG-MST1 K59R by immunoblotting with the FLAG antibody. CMV, cytomegalovirus. B, in vitro kinase assays were carried out by incubating recombinant MST1 (rMST1) (Upstate Biotechnology) together with recombinant GST or GST-FOXO1 as substrate in the presence of 32P-labeled ATP. Reactions were analyzed by SDS-PAGE followed by autoradiography. Recombinant MST1 phosphorylated FOXO1 in vitro. C, immunoprecipitated MST1 was subjected to an in vitro kinase assay as in A using the forkhead domain of FOXO1 fused to GST (GST-FOXO1(FD)) as substrate. D, MST1 was immunoprecipitated from 293T cells and subjected to in vitro kinase assays as in A using wild type or serine 212 mutants of GST-FOXO1(FD). The designation 3A refers to the S212A, S234A, S235A FOXO1(FD) mutant; the designation 4A refers to the S212A, S218A, S234A, and S235A FOXO1(FD) mutant. Reaction products were analyzed by SDS-PAGE followed by autoradiography. Immunoblotting with the GST antibody is shown in the lower panel. Mutation of serine 212 dramatically reduced MST1-induced phosphorylation of FOXO1(FD).
MST1 phosphorylates FOXO3 at serine 207, a conserved site within the forkhead domain of the FOXO family of transcription factors (16). Therefore, we next determined whether MST1 phosphorylates FOXO1 at serine 212, the site that corresponds to serine 207 in FOXO3. In in vitro kinase assays, MST1 phosphorylated the forkhead domain of FOXO1 fused to GST (GST-FOXO1(FD)) (Fig. 1C) (16). Upon tandem mass spectrometry analysis followed by data-dependent MS/MS/MS (MS3), we identified four serine residues in FOXO1 (serines 212, 218, 234, and 245) that were phosphorylated by MST1 (data not shown) (16). We next determined the ability of MST1 to catalyze the phosphorylation of wild-type FOXO1(FD) or FOXO1(FD) mutants in which serine 212 was mutated. Although MST1 robustly phosphorylated FOXO1(FD), MST1 failed to phosphorylate FOXO1(FD) mutants in which serine 212 was replaced with alanine (Fig. 1D) (16). These results indicate that MST1 phosphorylates FOXO1 at serine 212 in vitro.
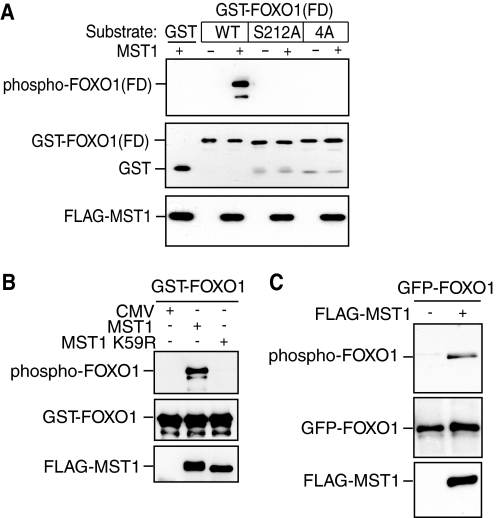
To determine whether MST1 phosphorylates FOXO1 at serine 212 in vivo, we used a phospho-FOXO antibody raised to specifically recognize FOXO3 or FOXO1 when phosphorylated at serine 207 or serine 212, respectively (16). The phospho-FOXO antibody recognized recombinant FOXO1(FD) that was phosphorylated by MST1 in vitro but failed to recognize unphosphorylated FOXO1(FD) (Fig. 2A). Importantly, the phospho-FOXO antibody failed to recognize serine 212 mutants of GST-FOXO1(FD) that were incubated with MST1 in vitro (Fig. 2A). We also found that the phospho-FOXO antibody recognized recombinant full-length FOXO1 when phosphorylated by MST1 in vitro but failed to recognize unphosphorylated full-length FOXO1 or FOXO1 that was incubated with the kinase-dead MST1 (MST1 K59R) (Fig. 2B). These results establish that the phospho-FOXO antibody recognizes serine 212-phosphorylated FOXO1 and that MST1 phosphorylates FOXO1 at serine 212 in vitro. We next determined the capacity of MST1 to phosphorylate FOXO1 in cells. We expressed FOXO1 fused to GFP (GFP-FOXO1) together with FLAG-tagged MST1 in 293T cells and subjected lysates of transfected cells to immunoblotting with the phospho-FOXO, GFP, or FLAG antibody. We found that expression of MST1 triggered the phosphorylation of FOXO1 at serine 212 (Fig. 2C). These results show that MST1 phosphorylates FOXO1 at serine 212 in vivo.
FIGURE 2.
MST1 phosphorylates FOXO1 at serine 212 in vitro and in vivo. A, immunoprecipitated MST1 subjected to kinase assays using wild type or serine 212 mutants of GST-FOXO1(FD). Proteins were analyzed by immunoblotting using the phospho-FOXO (top panel), GST (middle panel), or FLAG antibody (lower panel). MST1 induced phosphorylation of FOXO1 at serine 212. The designation 4A refers to the S212A, S218A, S234A, and S235A FOXO1(FD) mutant. B, lysates of 293T cells transfected with an expression plasmid encoding FLAG-MST1, kinase-dead FLAG-MST1 K59R, or the control plasmid were immunoprecipitated with the FLAG antibody and subjected to an in vitro cold kinase assay using full-length GST-FOXO1 as substrate. Reactions were analyzed by SDS-PAGE followed by immunoblotting with the phospho-FOXO antibody. GST-FOXO1 protein levels were detected with the GST antibody. The lower panels show the expression of FLAG-MST1 and FLAG-MST1 K59R by immunoblotting with the FLAG antibody. CMV, cytomegalovirus. C, lysates of 293T cells transfected with the GFP-FOXO1 expression plasmid together with the FLAG-MST1 expression plasmid or its control vector were immunoblotted with the phospho-FOXO (top panel), GFP (middle panel), or FLAG antibody (lower panel). MST1 expression induced the phosphorylation of FOXO1 at serine 212 in cells.
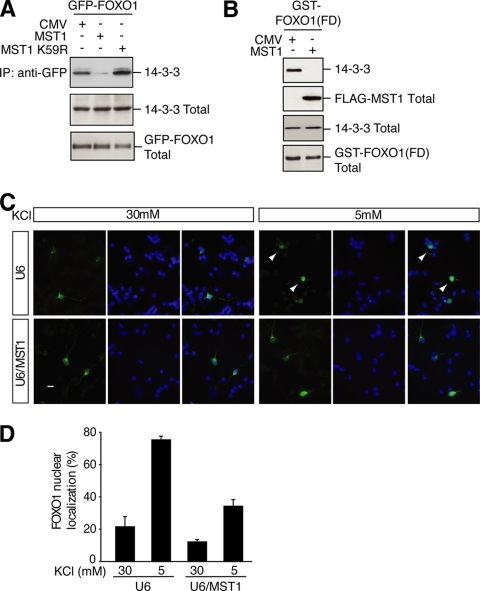
The MST1-induced FOXO1 Phosphorylation Disrupts FOXO1's Interaction with 14-3-3 Proteins and Promotes the Nuclear Accumulation of FOXO1—The ability of MST1 to phosphorylate FOXO1 at serine 212 led us next to determine the functional consequences of the FOXO1 phosphorylation. Because MST1-induced phosphorylation of FOXO3 at serine 207 disrupts FOXO3's interaction with 14-3-3 proteins (16), we assessed the effect of the MST1-induced phosphorylation of FOXO1 at serine 212 on FOXO1's association with 14-3-3 proteins. We expressed MST1 or kinase-dead MST (MST1 K59R) together with GFP-FOXO1 in cells and immunoblotted GFP immunoprecipitates of transfected cells with 14-3-3 antibodies. In control-transfected cells, FOXO1 interacted robustly with 14-3-3 proteins (Fig. 3A). By contrast, expression of MST1 dramatically reduced the ability of FOXO1 to interact with 14-3-3 proteins (Fig. 3A). Expression of MST1 K59R failed to disrupt FOXO1's interaction with 14-3-3 proteins, suggesting that the ability of MST1 to disrupt the FOXO1/14-3-3 association required the kinase activity of MST1 (Fig. 3A). In other experiments, MST1 inhibited the ability of the forkhead domain of FOXO1 to associate with 14-3-3 proteins in cells (Fig. 3B). Collectively, these results support the conclusion that MST1-induced phosphorylation of FOXO1 disrupts FOXO1's association with 14-3-3 proteins.
FIGURE 3.
MST1-induced phosphorylation of FOXO1 disrupts FOXO1's interaction with 14-3-3 proteins and promotes the nuclear accumulation of FOXO1. A, lysates of 293T cells transfected with the GFP-FOXO1 expression plasmid together with an expression vector encoding FLAG-MST1, FLAG-MST1 K59R, or the control vector were immunoprecipitated (IP) with the GFP antibody followed by immunoblotting with a 14-3-3 antibody (top panel). The total 14-3-3 and GFP-FOXO is shown in the middle and lower panels. MST1, but not the kinase-dead MST1 K59R, disrupted FOXO1's interaction with 14-3-3 proteins. CMV, cytomegalovirus. B, lysates of 293T cells transfected with a mammalian expression vector encoding GST-FOXO1(FD) together with the MST1 expression plasmid or its control vector were subjected to GST pull-down assays followed by immunoblotting with the 14-3-3 antibody (top panel). The total MST1, 14-3-3, and FOXO1(FD) is shown in the lower panels. C, granule neurons transfected with GFP-FOXO1 together with the MST1 RNAi or control U6 plasmid maintained in membrane-depolarizing medium (30 mm KCl) or deprived of membrane depolarization (5 mm KCl) were analyzed by fluorescence microscopy. Representative images are shown (left, GFP-FOXO1; center, Hoechst; right, merged). Scale bar, 10 μm. Arrowheads indicate neurons displaying nuclear localization of FOXO1. D, quantification of results shown in Fig. 3C. MST1 knockdown significantly decreased GFP-FOXO1 nuclear accumulation in activity-deprived neurons. For each experiment, at least 100 neurons were counted (n = 3; p < 0.0001, ANOVA followed by Fisher's PLSD post hoc test).
Because 14-3-3 proteins reside predominantly in the cytoplasm (31), the finding that the MST1-induced phosphorylation of FOXO1 disrupts its interaction with 14-3-3 proteins raised the possibility that the MST1-induced phosphorylation of FOXO1 might promote the nuclear translocation of FOXO1. We therefore determined the role of MST1 in the control of the subcellular localization of FOXO1 in primary cerebellar granule neurons. In recent studies, we found that FOXO1 translocates to the nucleus upon activity deprivation (25). We confirmed that the withdrawal of neuronal activity triggers the nuclear accumulation of FOXO1 in primary granule neurons (Fig. 3C). To determine the role of MST1 in activity deprivation-induced nuclear accumulation of FOXO1, we induced the knockdown of MST1 in neurons. We used a plasmid-based method of RNAi encoding shRNAs targeting MST1 (16, 32). MST1 knockdown reduced significantly the level of FOXO1 in the nucleus in activity-deprived granule neurons (Fig. 3, C and D). These results suggest that MST1 plays a critical role in the control of FOXO1 nucleocytoplasmic shuttling and support the interpretation that MST1-FOXO1 signaling promotes the nuclear accumulation of FOXO1 in the nucleus in neurons.
MST1 Mediates Growth Factor and Activity Deprivation-induced Neuronal Cell Death—A requirement for MST1 in the nuclear accumulation of FOXO1 in activity-deprived neurons led us to test the role of MST1 in neuronal cell death. The deprivation of neuronal activity and growth factors, mimicked by the withdrawal of serum and membrane depolarization, potently induces cell death in cerebellar granule neurons (13–15, 28, 29). We therefore tested the effect of MST1 knockdown on the ability of growth factor and activity deprivation to induce neuronal cell death. Primary granule neurons were transfected with the MST1 RNAi or control U6 plasmid together with an expression plasmid encoding β-galactosidase. Transfected neurons were left in full survival medium containing serum- and membrane-depolarizing concentrations of KCl (30 mm) or deprived of serum and membrane depolarization (5 mm KCl) and then subjected to immunocytochemical analyses using an antibody to β-galactosidase and the DNA dye bisbenzimide (Hoechst 33258). Transfected β-galactosidase-positive neurons were analyzed for cell survival and death based on the integrity of neuronal processes and nuclear morphology.
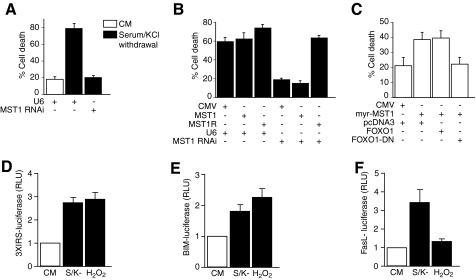
As expected, the deprivation of membrane depolarization and growth factors induced cell death in granule neurons (Fig. 4A). However, the knockdown of MST1 significantly reduced the level of cell death in activity and growth factor-deprived neurons (Fig. 4A). MST1 knockdown neurons appeared healthy, displaying robust nuclei and processes (data not shown). These results suggest that MST1 plays a critical role in inducing cell death in neurons upon the deprivation of survival stimuli. To demonstrate the specificity of the MST1 RNAi-induced survival phenotype, we performed a rescue experiment. We expressed MST1 encoded by cDNA designed to be resistant to MST1 shRNAs (MST1R) (16). We found that expression of MST1R, but not MST1 encoded by wild-type cDNA, reversed the ability of MST1 RNAi to protect neurons against activity and growth factor deprivation-induced apoptosis (Fig. 4B). These results suggest that the MST1 RNAi-induced protection of neurons from cell death is the result of specific knockdown of MST1 rather than off-target effects of RNAi.
FIGURE 4.
MST1-FOXO1 signaling promotes cell death in neurons upon survival factor deprivation. A, granule neurons were transfected with the MST1 RNAi or control U6 plasmid together with the β-galactosidase expression vector. After 72 h, cultures were left in full survival (conditioned medium (CM)) or deprived of serum and membrane depolarization for 24 h. Transfected granule neurons were analyzed for cell survival and death based on the integrity of neuronal processes and nuclear morphology (see “Experimental Procedures”). The percentage of cell death in transfected β-galactosidase-positive-staining neurons is represented as mean ± S.E. Cell death was significantly reduced in MST1 knockdown neurons as compared with U6 control neurons upon serum and activity deprivation (n = 4; p < 0.05, ANOVA followed by Fisher's PLSD post hoc test). B, neurons transfected with FLAG-MST1 or FLAG-MST1R plasmid and the β-galactosidase expression vector were treated and analyzed as in A. MST1 encoded by the RNAi-resistant cDNA (MST1R), but not MST1 encoded by wild-type cDNA (MST1), blocked the ability of MST1 RNAi to protect neurons from serum- and activity deprivation-induced cell death (n = 3; p < 0.0001; ANOVA followed by Fisher's PLSD post hoc test). CMV, cytomegalovirus. C, neurons transfected with myr-MST1 or its control vector, together with the FOXO1, FOXO1-DN, or control expression plasmid, were analyzed as in A. FOXO1-DN, but not FOXO1, blocked the ability of myr-MST1 to promote neuronal death (n = 3; p < 0.05; ANOVA followed by Fisher's PLSD post hoc test). D–F, granule neurons were transfected with the 3×IRS (IGFBP1)-luciferase reporter gene (D), BIM-luciferase reporter gene (E), or FasL-luciferase reporter gene (F) together with the tk-Renilla reporter gene, which served as an internal control for transfection efficiency. After 24 h, cultures were left in full conditioned medium, deprived of serum and membrane depolarization (S/K-), or treated with H2O2 for 24 h. Neurons were subjected to luciferase assays. The mean ± S.E. of normalized firefly/Renilla luciferase values (RLU) relative to neurons maintained in conditioned medium is shown in the graphs (n = 3). Withdrawal of serum and membrane depolarization increased the level of 3×IRS, BIM, and FasL reporter genes (p < 0.05; ANOVA followed by Fisher's PLSD post hoc test). Hydrogen peroxide significantly stimulated the 3×IRS and BIM reporter genes (p < 0.05; ANOVA followed by Fisher's PLSD post hoc test) but failed to induce the FasL reporter gene.
We next determined whether MST1 induces cell death in a FOXO1-dependent manner. We expressed an activated form of MST1 in which MST1 is fused with a myristoylation signal (myr-MST1) in cerebellar granule neurons together with an expression plasmid encoding a dominant interfering form of FOXO1 in which the transactivation domain is deleted (FOXO1-DN) or its control vector (33, 34). Expression of myr-MST1 induced cell death robustly in granule neurons (Fig. 4C). However, expression of FOXO1-DN reduced myr-MST1-induced cell death in neurons (Fig. 4C). Collectively, our results suggest that MST1-FOXO1 signaling contributes to cell death upon the withdrawal of growth factors and neuronal activity.
In view of the role of MST1-FOXO3 signaling in oxidative stress-induced neuronal death (16), the identification of MST1-FOXO1 signaling in survival factor deprivation-induced neuronal cell death in this study raised the question of how MST1 function is differentially controlled in neurons upon exposure to distinct cell death stimuli. We first characterized the effect of survival factor deprivation and oxidative stress on transcriptional responses of apoptotic gene targets of the FOXO transcription factors in granule neurons. Both hydrogen peroxide and survival factor deprivation triggered the expression of a FOXO-responsive reporter gene in granule neurons (Fig. 4D) (16, 25). Consistent with these results, both hydrogen peroxide and survival factor deprivation similarly induced the expression of a reporter gene controlled by the promoter of the apoptotic FOXO target gene BIM (Fig. 4E) (16, 25). By contrast, the FOXO target gene FasL appeared to respond differentially to hydrogen peroxide and survival factor deprivation. Although survival factor deprivation robustly induced the expression of a reporter gene controlled by the FasL promoter, hydrogen peroxide marginally induced the FasL-luciferase reporter gene in neurons (Fig. 4F). Together, these results suggest that distinct sets of FOXO target genes may be activated in neurons upon exposure to survival factor deprivation and oxidative stress. It will be important to determine in future studies the full complement of genes that are regulated by FOXO1 and FOXO3 that couple the MST1 signal to cell death in neurons.
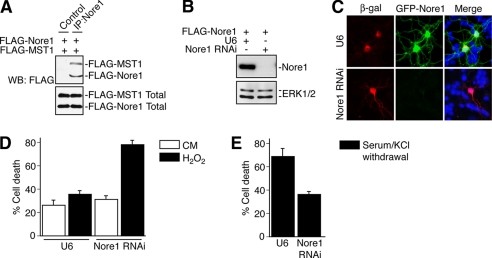
The MST1 Regulatory Scaffold Protein Nore1 Plays Distinct Roles in Neuronal Cell Death—We next investigated the role of MST1 regulatory mechanisms in neuronal responses to oxidative stress and survival factor deprivation. The scaffold protein Nore1 has emerged as a critical upstream regulator of MST1 in non-neural cells (33, 34). We therefore asked whether Nore1 plays a role in the control of neuronal death. Nore1 is expressed in the brain, including granule neurons of the rodent cerebellar cortex (the Allen Mouse Brain Atlas and data not shown). We confirmed that Nore1 forms a physical complex with MST1 in cells (Fig. 5A). To determine Nore1 function in neuronal death, we generated a plasmid encoding Nore1 shRNAs. Nore1 RNAi triggered robust knockdown of Nore1 in 293T cells and neurons (Fig. 5, B and C). We next analyzed the effect of Nore1 RNAi on neuronal death. Remarkably, Nore1 knockdown led to distinct cellular responses in neurons upon exposure to oxidative stress and survival factor deprivation. Nore1 knockdown sensitized granule neurons to hydrogen peroxide-induced cell death (Fig. 5D). By contrast, we found that Nore1 knockdown robustly protected neurons from survival factor deprivation-induced cell death (Fig. 5E). These results suggest that the MST1-regulating scaffold protein Nore1 plays a dual role in neuronal death depending on the stimulus. Although Nore1 dampens the effect of oxidative stress on cell death in neurons, Nore1 is required for survival factor deprivation-induced neuronal cell death.
FIGURE 5.
The MST1 regulatory scaffold protein Nore1 plays distinct roles in neuronal death. A, lysates of 293T cells transfected with the FLAG-Nore1 and FLAG-MST1 expression vectors were immunoprecipitated (IP) using a rabbit polyclonal antibody to Nore1 and immunoblotted (WB) using the FLAG antibody. The total FLAG-Nore1 and FLAG-MST1 is shown in the lower panel. B, lysates of 293T cells transfected with the FLAG-Nore1 expression plasmid together with an RNAi plasmid encoding Nore1 shRNAs or the control U6 plasmid were immunoblotted with the Nore1 or ERK1/2 antibody. Nore1 RNAi triggered robust knockdown of Nore1. C, granule neurons transfected with expression plasmids encoding GFP-Nore1 and β-galactosidase (β-gal) together with the Nore1 RNAi plasmid or control U6 plasmid were subjected to immunocytochemical analysis using a mouse monoclonal antibody to β-galactosidase and a rabbit polyclonal antibody to GFP. Nore1 RNAi induced the knockdown of Nore1 in nearly all cells examined. D, granule neurons were transfected with the Nore1 RNAi plasmid or control U6 plasmid together with a plasmid encoding β-galactosidase. After 72 h, cultures were left untreated or treated with H2O2, and 24 h later cultures were analyzed as described in Fig. 4A. Nore1 knockdown sensitized neurons to H2O2-induced cell death as compared with control (n = 3, p < 0.05, ANOVA followed by Fisher's PLSD post hoc test). CM, conditioned medium. E, granule neurons were transfected with the Nore1 RNAi plasmid or control U6 plasmid together with the β-galactosidase expression plasmid. After 72 h, cultures were deprived of survival factors and left untreated for 24 h and analyzed as in Fig. 4A. Nore1 knockdown protected neurons from survival factor deprivation-induced cell death as compared with control (n = 3, p < 0.05, t test).
In this study, we have identified a link between MST1 and FOXO1 and thus uncovered a critical function for MST1 in survival factor deprivation-induced neuronal death. Our major findings are: 1) MST1 phosphorylates FOXO1 at the conserved forkhead binding site of serine 212 in vitro and in vivo, 2) the MST1-induced phosphorylation of FOXO1 promotes the disruption of FOXO1 from 14-3-3 proteins, 3) MST1 triggers the nuclear accumulation of FOXO1 in primary neurons upon activity deprivation, 4) MST1 plays a critical role in cell death of neurons upon growth factor and activity deprivation, and 5) the MST1 regulatory scaffold protein Nore1 plays a critical role in neuronal death upon growth factor and activity deprivation. Collectively, we have identified a novel function for MST1 in survival factor deprivation-induced neuronal death, and we have characterized upstream and downstream mechanisms of MST1 function in neuronal death upon survival factor deprivation.
The identification of MST1-induced FOXO1 phosphorylation and consequent dissociation of FOXO1 from 14-3-3 proteins supports the growing concept that the disruption of 14-3-3 proteins from pro-apoptotic proteins plays a key role in determining the balance of life and death in neurons (16, 25, 26, 29, 35, 36). Intriguingly, a number of apoptotic protein kinases, including Cdk1, JNK1, and MST1, trigger the dissociation of 14-3-3 proteins from apoptotic proteins including the BH3-only protein BAD and the transcription factors FOXO1 and FOXO3 (16, 25, 26, 29, 35). Cdk1 promotes cell death by acting on both BAD and FOXO1 (25, 29, 37). Thus, Cdk1 and MST1 appear to converge upon FOXO1 by phosphorylating two distinct sites, serine 249 and serine 212, respectively, each promoting the disruption of FOXO1 from 14-3-3 proteins. Thus, the combined actions of Cdk1 and MST1 in neurons deprived of survival stimuli may ensure the robust activation of FOXO1 and induction of cell death. In future studies, it will be important to determine whether MST1, like Cdk1 and JNK1, also acts directly at the mitochondria to promote cell death.
Our findings generalize the substrates of MST1 from FOXO3 to other family members of the FOXO family of transcription factors in mammalian cells. Thus, MST1 is likely to influence diverse biological processes beyond cell death in neurons. Consistent with this interpretation, MST-FOXO signaling promotes lifespan and health span in Caenorhabditis elegans (16). The scope of MST1-FOXO1 as well as MST1-FOXO3 signaling in the regulation of additional biological functions in mammalian cells from cell proliferation to differentiation and metabolism remains to be explored.
The finding that the MST1 regulatory protein Nore1 exerts distinct roles in neuronal cell death depending on the nature of the death stimulus raises the important question of how these stimuli in turn regulate Nore1. In agreement with our finding that Nore1 is critical for survival factor deprivation-induced death in primary neurons, Nore1 plays an apoptotic tumor suppressive role in non-neural cells (33). Whether the ability of Nore1 to dampen oxidative stress-induced neuronal cell death extends beyond neurons to cells outside the nervous system remains an interesting question for future studies.
The identification of MST1 signaling to FOXO1 further builds the case that MST1 is a key regulator in neuronal cell death. It will be important in future studies to determine the role of these pathways in brain development and to investigate the role of MST1-FOXO1 signaling in the pathogenesis of neurological diseases.
Acknowledgments
We thank M. Smidt for providing the GFP-FOXO1 plasmid, P. Bois for the dominant negative FOXO1 plasmid, J. Avruch for MST1 and Nore1 plasmids, and members of the Bonni laboratory for critical reading of the manuscript.
This work was supported, in whole or in part, by National Institutes of Health Grants NS047188 and NS41021 (to A. B.). This work was also supported by a pilot grant from the Paul Glenn laboratories (to A. B.) and a grant from the Lefler Foundation (to Z. Y.).
Footnotes
The abbreviations used are: MST1, mammalian Sterile 20-like kinase 1; GST, glutathione S-transferase; RNAi, RNA interference; shRNA, short hairpin RNA; ANOVA, analysis of variance; PLSD, protected least significance difference; GFP, green fluorescent protein; FD, forkhead domain; myr, myristoylation.
References
- 1.Oppenheim, R. (1989) Annu. Rev. Neurosci. 14, 453-501 [DOI] [PubMed] [Google Scholar]
- 2.Pettmann, B., and Henderson, C. E. (1998) Neuron 20, 633-647 [DOI] [PubMed] [Google Scholar]
- 3.Yuan, J., and Yankner, B. A. (2000) Nature 407, 802-809 [DOI] [PubMed] [Google Scholar]
- 4.Mattson, M. P. (2000) Nat. Rev. Mol. Cell Biol. 1, 120-129 [DOI] [PubMed] [Google Scholar]
- 5.Choi, D. W. (1996) Curr. Opin. Neurobiol. 6, 667-672 [DOI] [PubMed] [Google Scholar]
- 6.Williams, R. W., and Herrup, K. (1988) Annu. Rev. Neurosci. 11, 423-453 [DOI] [PubMed] [Google Scholar]
- 7.Becker, E. B., and Bonni, A. (2004) Prog. Neurobiol. 72, 1-25 [DOI] [PubMed] [Google Scholar]
- 8.Schwartz, P. M., Borghesani, P. R., Levy, R. L., Pomeroy, S. L., and Segal, R. A. (1997) Neuron 19, 269-281 [DOI] [PubMed] [Google Scholar]
- 9.Bates, B., Rios, M., Trumpp, A., Chen, C., Fan, G., Bishop, J. M., and Jaenisch, R. (1999) Nat. Neurosci. 2, 115-117 [DOI] [PubMed] [Google Scholar]
- 10.Suh, Y. S., Oda, S., Kang, Y. H., Kim, H., and Rhyu, I. J. (2002) Neurosci. Lett. 325, 1-4 [DOI] [PubMed] [Google Scholar]
- 11.Mennerick, S., and Zorumski, C. F. (2000) Mol. Neurobiol. 22, 41-54 [DOI] [PubMed] [Google Scholar]
- 12.Ikonomidou, C., Bosch, F., Miksa, M., Bittigau, P., Vockler, J., Dikranian, K., Tenkova, T. I., Stefovska, V., Turski, L., and Olney, J. W. (1999) Science 283, 70-74 [DOI] [PubMed] [Google Scholar]
- 13.D'Mello, S. R., Galli, C., Ciotti, T., and Calissano, P. (1993) Proc. Natl. Acad. Acad. Sci. U. S. A. 90, 10989-10993 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Padmanabhan, J., Park, D., Greene, L. A., and Shelanski, M. L. (1999) J. Neurosci. 19, 8747-8756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Miller, T. M., and Johnson, E. M. (1996) J. Neurosci. 16, 7487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lehtinen, M. K., Yuan, Z., Boag, P. R., Yang, Y., Villen, J., Becker, E. B., DiBacco, S., de la Iglesia, N., Gygi, S., Blackwell, T. K., and Bonni, A. (2006) Cell 125, 987-1001 [DOI] [PubMed] [Google Scholar]
- 17.Lehtinen, M. K., and Bonni, A. (2008) Curr. Mol. Med. (Hilversum) 8, 313-318 [DOI] [PubMed] [Google Scholar]
- 18.Obsil, T., and Obsilova, V. (2008) Oncogene. 27, 2263-2275 [DOI] [PubMed] [Google Scholar]
- 19.Calnan, D. R., and Brunet, A. (2008) Oncogene 27, 2276-2288 [DOI] [PubMed] [Google Scholar]
- 20.van der Vos, K. E., and Coffer, P. J. (2008) Oncogene 27, 2289-2299 [DOI] [PubMed] [Google Scholar]
- 21.Arden, K. C. (2008) Oncogene 27, 2345-2350 [DOI] [PubMed] [Google Scholar]
- 22.Hoekman, M. F., Jacobs, F. M., Smidt, M. P., and Burbach, J. P. (2006) Gene. Expr. Patterns 6, 134-140 [DOI] [PubMed] [Google Scholar]
- 23.van der Heide, L. P., Jacobs, F. M., Burbach, J. P., Hoekman, M. F., and Smidt, M. P. (2005) Biochem. J. 391, 623-629 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Biggs, W. H., III, Meisenhelder, J., Hunter, T., Cavenee, W. K., and Arden, K. C. (1999) Proc. Natl. Acad. Sci. U. S. A. 96, 7421-7426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yuan, Z., Becker, E. B., Merlo, P., Yamada, T., DiBacco, S., Konishi, Y., Schaefer, E. M., and Bonni, A. (2008) Science 319, 1665-1668 [DOI] [PubMed] [Google Scholar]
- 26.Kim, A. H., and Bonni, A. (2008) Cell Cycle 7, 3819-3822 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Shalizi, A., Lehtinen, M., Gaudilliere, B., Donovan, N., Han, J., Konishi, Y., and Bonni, A. (2003) J. Neurosci. 23, 7326-7336 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Becker, E. B., and Bonni, A. (2006) Neuron 49, 655-662 [DOI] [PubMed] [Google Scholar]
- 29.Konishi, Y., Lehtinen, M., Donovan, N., and Bonni, A. (2002) Mol. Cell 9, 1005-1016 [DOI] [PubMed] [Google Scholar]
- 30.Graves, J. D., Gotoh, Y., Draves, K. E., Ambrose, D., Han, D. K., Wright, M., Chernoff, J., Clark, E. A., and Krebs, E. G. (1998) EMBO J. 17, 2224-2234 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Van Der Heide, L. P., Hoekman, M. F., and Smidt, M. P. (2004) Biochem. J. 380, 297-309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gaudilliere, B., Shi, Y., and Bonni, A. (2002) J. Biol. Chem. 277, 46442-46446 [DOI] [PubMed] [Google Scholar]
- 33.Khokhlatchev, A., Rabizadeh, S., Xavier, R., Nedwidek, M., Chen, T., Zhang, X. F., Seed, B., and Avruch, J. (2002) Curr. Biol. 12, 253-265 [DOI] [PubMed] [Google Scholar]
- 34.Bois, P. R., and Grosveld, G. C. (2003) EMBO J. 22, 1147-1157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Donovan, N., Becker, E. B., Konishi, Y., and Bonni, A. (2002) J. Biol. Chem. 277, 40944-40949 [DOI] [PubMed] [Google Scholar]
- 36.Becker, E. B., and Bonni, A. (2005) Semin. Cell Dev. Biol. 16, 439-448 [DOI] [PubMed] [Google Scholar]
- 37.Konishi, Y., and Bonni, A. (2003) J. Neurosci. 23, 1649-1658 [DOI] [PMC free article] [PubMed] [Google Scholar]