Abstract
The adhesion of bacteria to host tissues is often mediated by interactions with extracellular matrices. Herein, we report on the interactions of the group A streptococcus, Streptococcus pyogenes, with the extracellular matrix protein fibulin-1. S. pyogenes bound purified fibulin-1 in a dose-dependent manner. Genetic ablation of serum opacity factor (SOF), a virulence determinant of S. pyogenes, reduced binding by ∼50%, and a recombinant peptide of SOF inhibited binding of fibulin-1 to streptococci by ∼45%. Fibulin-1 bound to purified SOF2 in a dose-dependent manner with high affinity (Kd = 1.6 nm). The fibulin-1-binding domain was localized to amino acid residues 457–806 of SOF2, whereas the fibronectin-binding domain is contained within residues 807–931 of SOF2, indicating that these two domains are separate and distinct. Fibulin-1 bound to recombinant SOF from M types 2, 4, 28, and 75 of S. pyogenes, indicating that the fibulin-1-binding domain is likely conserved among SOF from different serotypes. Mixed binding experiments suggested that gelatin, fibronectin, fibulin-1, and SOF form a quaternary molecular complex that enhanced the binding of fibulin-1. These data indicate that S. pyogenes can interact with fibulin-1 and that SOF is a major streptococcal receptor for fibulin-1 but not the only receptor. Such interactions with fibulin-1 may be involved in the adhesion of S. pyogenes to extracellular matrices of the host.
Adhesion of bacteria to host surfaces is the first stage in establishing bacterial infections in the human host, and a variety of molecular mechanisms are utilized to initiate adhesion. A common mechanism for adhesion involves interactions between bacterial adhesins and components of the extracellular matrices of the host. The identification and characterization of microbial surface components recognizing adhesive matrix molecules (MSCRAMM) has led to important advances in vaccines and immunotherapies for preventing and treating bacterial infections (1).
The group A streptococcus, Streptococcus pyogenes, is a major human pathogen causing diseases ranging from relative minor infections such as pharyngitis and cellulitis to severe infections with high levels of morbidity and mortality such as necrotizing fasciitis and toxic shock syndrome (2). This pathogen expresses adhesins that interact with various components of the extracellular matrix including laminin, elastin, fibronectin, fibrinogen, and collagen (3–7). The interactions between fibronectin and S. pyogenes have been intensely studied, and these investigations have revealed at least 10 different streptococcal proteins that bind fibronectin (4).
Serum opacity factor (SOF)2 is a major fibronectin-binding protein that is involved in adhesion to host cells (8–11). SOF is a virulence determinant that is expressed by approximately half of the clinical isolates of S. pyogenes (8). SOF opacifies serum by binding and displacing apoA-I in high density lipoproteins (8, 12–15). SOF is covalently linked to the streptococcal cell wall via an LPSTG sortase recognition site and is also released in a soluble form. SOF has two functionally distinct domains, an N-terminal domain that opacifies serum and a C-terminal domain that binds fibronectin. The role of SOF in adhesion involves both its C-terminal fibronectin-binding domain and an N-terminal region (see Fig. 1 for a schematic of structure) (9, 11). However, the nature of the interactions between the N-terminal region of SOF and host components is not well characterized.
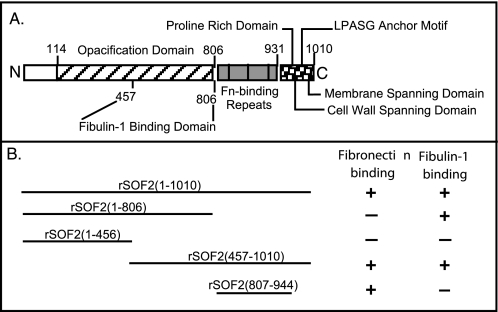
FIGURE 1.
A, a schematic of the structure of SOF and its functional domains is shown. The assignment of functional domains are based on the findings of Rakonjac et al. (33), Kreikemeyer et al. (34), Courtney et al. (8, 13), and results presented in this work. Fn, fibronectin. B, the data for the binding of SOF peptides to fibronectin are from previous publications (8, 13), and the data for fibulin-1 are from the present work.
Herein, we report on the interactions between a truncated form of SOF in which its fibronectin-binding domain has been deleted and the extracellular matrix protein fibulin-1. Fibulin-1 is a member of the fibulin family that currently consists of seven glycoproteins. All fibulins contain epidermal growth factor-like repeats and a unique fibulin-type module at its C terminus that define this family (16, 17). Fibulin-1 is found within the extracellular matrices and in human plasma at 30–50 μg/ml (18). It interacts with many of the components of the extracellular matrix including fibronectin, laminin, fibrinogen, nidogen-1, endostatin, aggrecan, and versican (16, 19). Due to its intimate relationship with the extracellular matrix, it is not surprising that the defects in fibulin-1 have a wide-ranging impact. Genetic evidence suggests that fibulin-1 is involved in tissue organization, the maturation and maintenance of blood vessels, and multiple embryonic pathways (16, 20–22).
Although it has been established that many of the other components of the extracellular matrix can interact with bacteria, there has been no previous report on the binding of fibulins to bacteria. Our findings indicate that fibulin-1 does bind to streptococci and that SOF is a major streptococcal receptor for fibulin-1.
EXPERIMENTAL PROCEDURES
Proteins—The construction and purification of recombinant peptides of SOF have been described (8, 9, 13). Essentially, chromosomal DNA from the indicated serotypes of S. pyogenes were used as templates; the desired encoding regions of sof were amplified by PCR, ligated into pTrcHis, and expressed in Escherichia coli; and recombinant SOF was purified by metal affinity chromatography. Purified recombinant SOF from M type 75 S. pyogenes (rSOF) was generously provided by Dr. Mark Walker at the University of Wollongong. Fibronectin was purified by gelatin affinity chromatography from fresh human serum as described by Engvall and Ruoslahti (23). Fibulin-1 was purified from extracts of human placenta by immunoaffinity chromatography using mouse monoclonal 3A11 anti-fibulin-1 IgG-Sepharose (18, 24) and labeled with biotin as described previously (5).
Organisms and Growth Conditions—Streptococci (i.e. T2MR and YL3) were grown overnight at 37 °C in Todd Hewitt broth supplemented with 1.5% yeast extract. T2MR is a clinical isolate of M type 2 S. pyogenes that expresses SOF. YL3 is an isogenic mutant of T2MR in which sof was insertionally inactivated using the Ω-element as described (8). The Ω-element contains translational and transcriptional terminators as well as a kanamycin resistance marker that is expressed in both Gram-positive and Gram-negative organisms (25). Lack of expression of SOF was verified by enzyme-linked immunosorbent assay (ELISA) of whole bacteria, Western blots of streptococcal extracts, and functional analyses (8).
ELISA—Microtiter wells were coated with 3 μg/ml rSOF-(1–806) or bovine serum albumin (BSA) for 4 h at 37 °C and then blocked with 5% nonfat milk in Tris-buffered saline (TBS) for 2 h at room temperature. The wells were then washed and incubated overnight with serial dilutions of fibulin-1 in TBS containing 3% nonfat milk and 0.1% Tween 20 (dilution buffer). The wells were rinsed and incubated with murine monoclonal antibody 3A11 against fibulin-1 (1 μg/ml in dilution buffer) for 1 h. Afterward, the wells were rinsed and incubated at ambient temperature for 1 h with a 1:6,000 dilution of peroxidase-conjugated goat anti-mouse IgG (GE Healthcare). The wells were then rinsed, and the SureBlue tetramethylbenzidine (TMB) substrate (KPL, Gaithersburg, MD) was added, and the absorbance at 650 nm was recorded after color development. The binding curve was generated using SigmaPlot software, and the Kd was calculated by determining the concentration of the ligand required for half-maximal binding.
For ELISA assays measuring the binding of fibulin-1 to SOF peptides, various truncated peptides of SOF or BSA were coated onto microtiter wells at 10 μg/ml in sodium bicarbonate, pH 9.5, for 1 h at 37 °C and then blocked with BSA (1 mg/ml). Wells were rinsed and fibulin-1 (4 μg/ml in TBS with 1 mg/ml BSA) was added to the wells and incubated for 60 min at 37 °C. The wells were then washed, and a 1:1,000 dilution of rabbit anti-fibulin-1 IgG or normal rabbit serum was added to the wells and incubated for 30 min at 37 °C. Afterward, the wells were washed, and a 1:2,000 dilution of peroxidase-conjugated goat anti-rabbit IgG was added. After incubating at 37 °C for 30 min, the wells were washed, and the TMB substrate was added. The absorbance at 650 nm was recorded after color development.
In assays to examine the effect of potential complexes between the constituents on the binding of fibulin-1, wells were coated with rSOF, fibronectin, gelatin, or BSA (10 μg/ml) for 1 h at 37 °C. The wells were washed and blocked with BSA (1 mg/ml in PBS) for 30 min at 37 °C. Biotinylated fibulin-1 (6 μg/ml) that was premixed with control buffer (1 mg/ml BSA in TBS with 1 mm CaCl2) or 10 μg/ml fibronectin with or without 10 μg/ml rSOF-(1–1010) was then added to the wells and incubated for 1 h at 37 °C. The wells were washed, and a 1:2,000 dilution of Neutralite avidin-peroxidase (Molecular Probes, Eugene, OR) was added to wells and incubated for 30 min at 37 °C. Afterward, the wells were washed, and the TMB substrate was added. The absorbance at 650 nm was recorded after color development. Wells coated with BSA served as negative controls.
Assays for Binding of Fibulin-1 to S. pyogenes—Microtiter wells were coated with 100 μl of wild type strain T2MR of S. pyogenes or its SOF-negative mutant YL3 (A530 = 0.4) in phosphate-buffered saline (PBS) for 30 min at 37 °C. Negative control wells were coated with BSA (10 μg/m PBS). The wells were then blocked with BSA (1 mg/ml in PBS) for 30 min at 37 °C. Serial 2-fold dilutions of biotinylated fibulin-1 were added to wells and incubated for 30 min at 37 °C. The wells were washed, and a 1:2,000 dilution of Neutralite avidin-peroxidase was added. After a 30-min incubation at 37 °C, the wells were washed, and the TMB substrate was added. The absorbance at 650 nm was recorded after color development.
For inhibition experiments with exogenous SOF, microtiter wells were coated with S. pyogenes strain T2MR and blocked with BSA as described above. Wells coated with BSA served as negative controls. Biotinylated fibulin-1 (4 μg/ml) was mixed with serial dilutions of rSOF2-(1–806) in TBS with 1 mg/ml BSA and incubated at 37 °C for 30 min. The wells were then washed, and a 1:2,000 dilution of Neutralite avidin-peroxidase was added and incubated for 30 min at 37 °C. After 30 min, the wells were washed, the TMB substrate was added, and the absorbance at 650 nm was recorded after color development.
SOF Affinity Chromatography of Human Serum—Purified recombinant SOF2 in which the fibronectin-binding domain was deleted (rSOF2-(1–806)) was coupled to Actigel ALD according to the manufacturer's procedures (Sterogene Bioseparations, Inc., Carlsbad, CA). Human serum was added to the column, and then the column was washed with TBS, and the bound material was eluted with 4 m urea in TBS. The bound fraction was dialyzed against TBS, concentrated by ultrafiltration, and subjected to SDS-PAGE under nonreducing conditions. The five major proteins bands were electroeluted from acrylamide gels. The purified proteins were subjected to SDS-PAGE under reducing and nonreducing conditions to determine whether any of the proteins contained more than one polypeptide. The proteins were electrophoretically transferred to polyvinylidene difluoride membrane, and bands were excised and subjected to N-terminal sequencing.
RESULTS
Identification of Fibulin-1 as a Serum Protein That Binds to SOF—The ability of SOF to bind fibulin-1 was discovered during a search for serum proteins that interact with SOF. This search was undertaken because many of the surface proteins of group A streptococci have multiple binding domains for serum proteins, and the binding of these proteins have been linked to increased survival of group A streptococci in blood and to adhesion to host cells (4, 7, 26). To investigate the ability of SOF to bind serum proteins, rSOF2-(1–806) was covalently linked to agarose and used as an affinity column. This construct was selected because the fibronectin-binding domain was deleted (Fig. 1), and we wanted to focus on the potential of SOF to bind serum proteins other than fibronectin. Human serum was applied to the column, and then the column was washed, and bound proteins were eluted with urea. The eluted fraction was subjected to SDS-PAGE under nonreducing conditions (Fig. 2, lane 1).
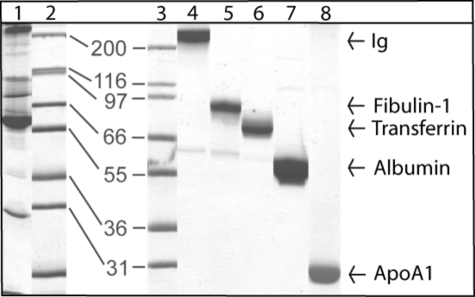
FIGURE 2.
Identification of serum proteins that bind to SOF. Human serum was applied to a SOF affinity column, and bound proteins were eluted with urea and subjected to SDS-PAGE under nonreducing conditions. Lane 1, peak fraction eluted from the column; lanes 2 and 3, molecular mass standards shown in kilodaltons; lanes 4–8, five major SOF-binding proteins purified from the gel by electroelution. The protein in lane 4 was identified as an immunoglobulin (Ig) by immunoblot analysis. Proteins in lanes 5–8 were identified by N-terminal amino acid sequencing.
Five major protein bands were present in the eluate from the SOF affinity column. The bands were purified from the acrylamide gels by electroelution (Fig. 2, lanes 4–8). SDS-PAGE under reducing conditions suggested that the protein in lane 4 of Fig. 2 could be an immunoglobulin. This was confirmed by immunoblot analysis using antibodies specific for human immunoglobulins (data not shown). It is not surprising that immunoglobulins were found to bind to the SOF affinity column, as antibodies to SOF are commonly found in human sera in response to infections with SOF-positive S. pyogenes (27).
N-terminal sequencing of the other proteins indicated that the other major bands corresponded to fibulin-1, transferrin, albumin, and apoA-I (Fig. 2, lanes 5–8). The finding that apoA-I bound to SOF was anticipated, as SOF was reported previously to bind to apoA-I with high affinity (13). Characterizations of the binding of albumin and transferrin to SOF will be the subject of future studies.
Characterization of Fibulin-1 Binding to SOF—To further investigate the interactions between fibulin-1 and SOF, fibulin-1 was purified from human placentas and incubated with microtiter wells coated with purified rSOF2-(1–806). Fibulin-1 bound to SOF in a dose-dependent fashion but did not bind to BSA (Fig. 3). Fibulin-1 bound to SOF with high affinity (Kd ∼ 1.6 nm), as determined by calculating the concentration required for half-maximal binding. To further localize the fibulin-1-binding domain, various recombinant peptides of SOF were constructed and tested for their ability to bind fibulin-1 (Fig. 4). In addition to binding to rSOF2-(1–806), fibulin-1 also bound to wells coated with rSOF2-(457–1010). In contrast, no significant levels of fibulin-1 binding were detected to wells coated with rSOF2-(1–456) and rSOF2-(807–944). As rSOF2-(1–806) does not contain the fibronectin-binding domain, and rSOF2-(807–944) does contain this domain, these data clearly indicate that the fibronectin-binding domain of SOF is not required for the binding of fibulin-1. These data further suggest that a fibulin-1-binding domain resides between amino acid residues 457–806 of SOF2.
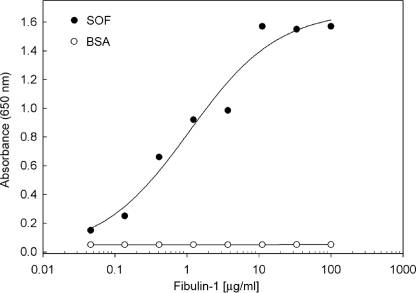
FIGURE 3.
Binding of fibulin-1 to SOF. Varying concentrations of fibulin-1 were incubated with microtiter wells coated with BSA or with rSOF2-(1–806). Bound fibulin-1 was quantified by ELISA using a monoclonal fibulin-1 antibody. The data shown are averages of duplicate wells. The experiment was repeated, and the results confirmed the presented data.
FIGURE 4.
Localization of the fibulin-1-binding domain of SOF. Microtiter wells were coated with the indicated recombinant peptides of SOF2 and then incubated with a constant amount of fibulin-1 (4 μg/ml). Bound fibulin-1 was quantified by ELISA as described under “Experimental Procedures.” Experiments were done in triplicate, and the mean ± S.D. is shown.
The sequence of SOF varies among different serotypes of S. pyogenes with homology ranging from 56 to 65% (8). Therefore, it was of interest to determine whether SOF expressed by other serotypes could also bind fibulin-1. The sof genes from M serotypes 4, 28, and 75 were cloned into E. coli and expressed, and the recombinant forms of SOF were purified. Microtiter wells were coated with these purified products as well as with rSOF2-(1–806) and then reacted with fibulin-1. Fibulin-1 bound to rSOF2, rSOF4, rSOF28, and rSOF75 but not to control wells coated with BSA (Fig. 5). These findings indicate that fibulin-1 can bind to SOF from multiple serotypes and suggest that the fibulin-1-binding domain is probably conserved within SOF from these different serotypes.
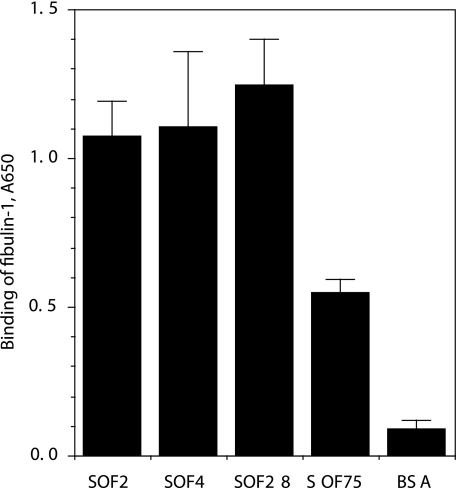
FIGURE 5.
Fibulin-1 binds to SOF from multiple serotypes of S. pyogenes. Purified, recombinant forms of SOF from the indicated serotypes were immobilized on microtiter wells and incubated with fibulin-1. Bound fibulin-1 was quantified by ELISA as described under “Experimental Procedures.” The experiments were done in triplicate, and the mean ± S.D. is shown.
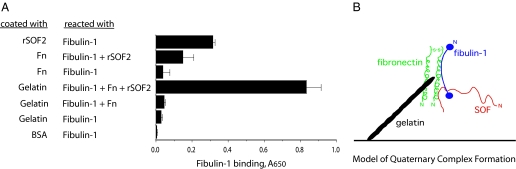
Quaternary Complex Formation between Gelatin, Fibronectin, Fibulin-1, and SOF—Fibronectin has multiple ligand-binding domains including ones for SOF, fibulin-1, and gelatin. Given our findings that fibulin-1 can also bind to SOF, there is the potential for these four proteins to form a complex, and the formation of such a complex could influence the degree to which fibulin-1 is bound. To determine whether this may be the case, microtiter wells were first coated with gelatin, fibronectin, SOF, or BSA and then incubated with solutions of fibulin-1 ± rSOF2-(1–1010) or fibronectin (Fig. 6A). Note that this form of SOF2 contains the fibronectin-binding domain. The highest degree of fibulin-1 binding occurred in wells that were first coated with gelatin and then incubated with a mixture of fibulin-1, fibronectin, and SOF. These findings suggest that these proteins may interact to form a quaternary complex (Fig. 6B).
FIGURE 6.
Interactions between gelatin, fibronectin, fibulin-1, and SOF. A, the left column indicates proteins that were coated on microtiter wells. The next column indicates the protein or mixture of proteins that were then added to these coated wells. Note that the full-length rSOF2-(1–1010) fragment containing the fibronectin-binding domain was used in these experiments. Biotinylated fibulin-1 was used in these experiments, and the degree of binding was determined using avidin-peroxidase as described under “Experimental Procedures.” The experiments were done in quadruplicate, and the mean ± S.D. is shown. Fn, fibronectin. B, shown is a schematic of quaternary complex that may form between gelatin coated on wells and fibulin-1, SOF, and fibronectin in solution.
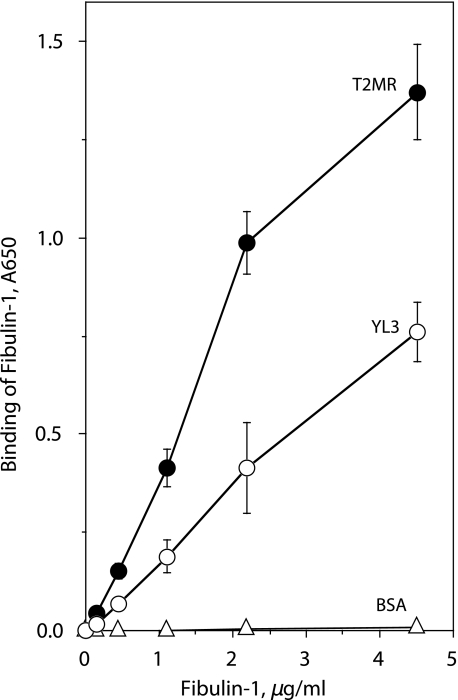
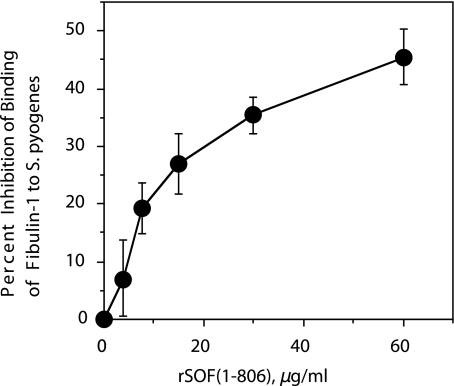
Binding of Fibulin-1 to S. pyogenes—The above findings provide ample evidence that SOF interacts with fibulin-1. Next, we wanted to determine whether SOF on the surface of streptococci could serve as a receptor for fibulin-1. Microtiter wells were coated with wild type strain T2MR of S. pyogenes and its SOF-negative mutant, YL3, and reacted with various concentrations of biotinylated fibulin-1. Fibulin-1 was found to bind to T2MR in a dose-related fashion (Fig. 7). However, the level of fibulin-1 binding to the SOF-negative mutant was 50% less than that bound to T2MR. Next, we evaluated the ability of purified rSOF2-(1–806) to block the binding of fibulin-1 to wild type S. pyogenes. As shown in Fig. 8, exogenously added SOF blocked the binding of fibulin-1 in a dose-dependent fashion, reaching a level of 45% inhibition at the highest concentration tested.
FIGURE 7.
SOF-deficient mutant displays reduced binding of fibulin-1. Microtiter wells were coated with wild type S. pyogenes T2MR or its isogenic mutant defective in expressing SOF, YL3. Wells coated with BSA served as negative controls. Various concentrations of biotinylated fibulin-1 were added to the wells, and the degree of binding was determined with avidin-peroxidase as described under “Experimental Procedures.” The experiments were done in triplicate, and the mean ± S.D. is shown.
FIGURE 8.
SOF blocks binding of fibulin-1 to S. pyogenes. Biotinylated fibulin-1 and varying concentrations of rSOF2-(1–806) were incubated with microtiter wells coated with wild type S. pyogenes T2MR. Bound fibulin-1 was detected with avidin-peroxidase as described under “Experimental Procedures.” The experiments were done in triplicate, and the mean ± S.D. is shown.
DISCUSSION
Interactions between bacteria and sites within the extracellular matrix that have been exposed by trauma can lead to initial bacterial adhesion and the nidus of an infection. In this study, we described our investigations on the interactions between SOF, a surface-expressed virulence factor of S. pyogenes, and fibulin-1, a protein found in the extracellular matrix and in the blood of humans. The following lines of evidence indicate that SOF is a major streptococcal receptor for fibulin-1. (i) Fibulin-1 in human serum bound to an SOF affinity column; (ii) purified fibulin-1 bound with high affinity to SOF in solid phase assays; (iii) fibulin-1 bound to wild type S. pyogenes; (iv) inactivation of the sof gene resulted in a 50% decrease in the binding of fibulin-1 to S. pyogenes; and (v) purified SOF blocked the binding of fibulin-1 to S. pyogenes.
However, the finding that inactivation of sof2 and excess rSOF2 resulted in diminished binding of fibulin-1 no greater than 50% suggests that an additional streptococcal receptor(s) for fibulin-1 may exist. The nature of the other streptococcal receptor(s) remains to be determined. That exogenous SOF only blocked binding of fibulin-1 by 45% also suggested that SOF and the other receptor(s) bind to different sites in fibulin-1 because the inhibition of binding should be greater if they bound to the same site in fibulin-1. Thus, it appears the group A streptococci express at least two different receptors that bind to different domains within fibulin-1.
The binding of serum proteins to proteins expressed on the surface of group A streptococci has been linked to the ability of these organisms to escape phagocytosis and to multiply in human blood (7, 26, 28, 29). The expression of SOF has also been linked to the increased survival of streptococci in blood (26). Therefore, it was of interest to identify SOF-binding proteins in serum. The finding that fibulin-1 in serum bound to a SOF affinity column is intriguing and indicates that fibulin-1 and SOF can interact with each other, even in the presence of high concentrations of other serum components. The potential relevance of the binding of SOF to soluble fibulin-1 in blood is suggested by the findings that platelets in blood can bind to surfaces coated with fibulin-1 via a fibrinogen bridge (30) and that platelet adhesion can trigger aggregation and formation of a platelet plug. It remains to be determined whether or not fibulin-1 bound to the surface of S. pyogenes can also bind to platelets and the impact such binding might have on platelet aggregation.
Although the fibronectin-binding domain of SOF does not interact with fibulin-1, experiments with full-length SOF suggested that this domain may have a role in forming complexes between SOF, fibulin-1, and fibronectin. The finding that immobilization of fibronectin onto gelatin-coated surfaces enhanced fibulin-1 binding suggested that gelatin, fibronectin, SOF, and fibulin-1 form a quaternary complex as illustrated in Fig. 6B. We propose that the interactions between gelatin and fibronectin may induce a conformational change in fibronectin that enhances or stabilizes interactions with other ligands and that this conformational shift does not occur when fibronectin is directly immobilized on the microtiter wells. Such a concept has precedence, as others have demonstrated that the surface used for immobilization of fibronectin can alter its conformation and binding to ligands (31). Moreover, conformational changes in fibronectin can regulate the assembly of the extracellular matrix (32).
Group A streptococci use a variety of mechanisms for adhesion including interactions with the extracellular matrices of the host. Our findings raise the possibility that interactions between fibulin-1 and SOF on the surface of S. pyogenes may be an additional mechanism used by these streptococci to initiate adhesion to components of the extracellular matrix and to establish an infection.
This work was supported, in whole or in part, by National Institutes of Health Grant HL095067(to W. S. A.). This work was also supported by the Department of Veterans Affairs (to H. S. C.) and American Heart Association Grant 0755346U (to W. S. A.).
Footnotes
The abbreviations used are: SOF, serum opacity factor; ELISA, enzyme-linked immunosorbent assay; BSA, bovine serum albumin; TBS, Tris-buffered saline.
References
- 1.Rivas, J. M., Speziale, P., Patti, J. M., and Hook, M. (2004) Curr. Opin. Drug. Discovery Dev. 7 223-227 [PubMed] [Google Scholar]
- 2.Cunningham, M. W. (2000) Clin. Microbiol. Rev. 13 470-511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Courtney, H. S., Dale, J. B., and Hasty, D. I. (1996) Infect. Immun. 64 2415-2419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Courtney, H. S., Hasty, D. L., and Dale, J. B. (2002) Ann. Med. 34 77-87 [DOI] [PubMed] [Google Scholar]
- 5.Courtney, H. S., Li, Y., Dale, J. B., and Hasty, D. L. (1994) Infect. Immun. 62 3937-3946 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kreikemeyer, B., Klenk, M., and Podbielski, A. (2004) Int. J. Med. Microbiol. 294 177-188 [DOI] [PubMed] [Google Scholar]
- 7.Courtney, H. S., and Podbielski, A. (2004) in Bacterial Invasion of Host Cells (Lamont, R. J., ed) pp. 239-273, Cambridge University Press, Cambridge
- 8.Courtney, H. S., Hasty, D. L., Li, Y., Chiang, H. C., Thacker, J. L., and Dale, J. B. (1999) Mol. Microbiol. 32 89-98 [DOI] [PubMed] [Google Scholar]
- 9.Gillen, C. M., Courtney, H. S., Schulze, K., Rohde, M., Wilson, M. R., Timmer, A. M., Guzman, C. A., Nizet, V., Chhatwal, G. S., and Walker, M. J. (2008) J. Biol. Chem. 283 6359-6366 [DOI] [PubMed] [Google Scholar]
- 10.Timmer, A. M., Kristian, S. A., Datta, V., Jeng, A., Gillen, C. M., Walker, M. J., Beall, B., and Nizet, V. (2006) Mol. Microbiol. 62 15-25 [DOI] [PubMed] [Google Scholar]
- 11.Oehmcke, S., Podbielski, A., and Kreikemeyer, B. (2004) Infect. Immun. 72 4302-4308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gillard, B. K., Courtney, H. S., Massey, J. B., and Pownall, H. J. (2007) Biochemistry 46 12968-12978 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Courtney, H. S., Zhang, Y. M., Frank, M. W., and Rock, C. O. (2006) J. Biol. Chem. 281 5515-5521 [DOI] [PubMed] [Google Scholar]
- 14.Pownall, H. J., Courtney, H. S., Gillard, B. K., and Massey, J. B. (2008) Chem. Phys. Lipids 156 45-51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Han, M., Gillard, B. K., Courtney, H. S., Ward, K., Rosales, C., Khant, H., Ludtke, S. J., and Pownall, H. J. (2009) Biochemistry 48 1481-1487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Argraves, W. S., Greene, L. M., Cooley, M. A., and Gallagher, W. M. (2003) EMBO Rep. 4 1127-1131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.de Vega, S., Iwamoto, T., and Yamada, Y. (2009) CMLS Cell. Mol. Life Sci., in press [DOI] [PMC free article] [PubMed]
- 18.Argraves, W. S., Tran, H., Burgess, W. H., and Dickerson, K. (1990) J. Cell Biol. 111 3155-3164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Timpl, R., Sasaki, T., Kostka, G., and Chu, M. L. (2003) Nat. Rev. Mol. Cell Biol. 4 479-489 [DOI] [PubMed] [Google Scholar]
- 20.Chu, M. L., and Tsuda, T. (2004) Birth Defects Res. C Embryo Today 72 25-36 [DOI] [PubMed] [Google Scholar]
- 21.Muriel, J. M., Dong, C., Hutter, H., and Vogel, B. E. (2005) Development (Camb.) 132 4223-4234 [DOI] [PubMed] [Google Scholar]
- 22.Cooley, M. A., Kern, C. B., Fresco, V. M., Wessels, A., Thompson, R. P., McQuinn, T. C., Twal, W. O., Mjaatvedt, C. H., Drake, C. J., and Argraves, W. S. (2008) Dev. Biol. 319 336-345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Engvall, E., and Ruoslahti, E. (1977) Int. J. Cancer 20 1-5 [DOI] [PubMed] [Google Scholar]
- 24.Godyna, S., Mann, D. M., and Argraves, W. S. (1995) Matrix Biol. 14 467-477 [DOI] [PubMed] [Google Scholar]
- 25.Prentki, P., and Krisch, H. M. (1984) Gene (Amst.) 29 303-313 [DOI] [PubMed] [Google Scholar]
- 26.Courtney, H. S., Hasty, D. L., and Dale, J. B. (2006) Mol. Microbiol. 59 936-947 [DOI] [PubMed] [Google Scholar]
- 27.Kim, S., and Lee, N. Y. (2001) J. Clin. Microbiol. 39 1316-1318 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Carlsson, F., Sandin, C., and Lindahl, G. (2005) Mol. Microbiol. 56 28-39 [DOI] [PubMed] [Google Scholar]
- 29.Podbielski, A., Schnitzler, N., Beyhs, P., and Boyle, M. D. (1996) Mol. Microbiol. 19 429-441 [DOI] [PubMed] [Google Scholar]
- 30.Tran, H., Tanaka, A., Litvinovich, S. V., Medved, L. V., Haudenschild, C. C., and Argraves, W. S. (1995) J. Biol. Chem. 270 19458-19464 [DOI] [PubMed] [Google Scholar]
- 31.Bergkvist, M., Carlsson, J., and Oscarsson, S. (2003) J. Biomed. Mater. Res. 64 349-356 [DOI] [PubMed] [Google Scholar]
- 32.Karuri, N. W., Lin, Z., Rye, H. S., and Schwarzbauer, J. E. (2009) J. Biol. Chem. 284 3445-3452 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Rakonjac, J. V., Robbins, J. C., and Fischetti, V. A. (1995) Infect. Immun. 63 622-631 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kreikemeyer, B., Martin, D. R., and Chhatwal, G. S. (1999) FEMS Microbiol. Lett. 178 305-311 [DOI] [PubMed] [Google Scholar]