Abstract
Elongation by RNA polymerase II (RNAPII) is a finely regulated process in which many elongation factors contribute to gene regulation. Among these factors are the polymerase-associated factor (PAF) complex, which associates with RNAPII, and several cyclin-dependent kinases, including positive transcription elongation factor b (P-TEFb) in humans and BUR kinase (Bur1–Bur2) and C-terminal domain (CTD) kinase 1 (CTDK1) in Saccharomyces cerevisiae. An important target of P-TEFb and CTDK1, but not BUR kinase, is the CTD of the Rpb1 subunit of RNAPII. Although the essential BUR kinase phosphorylates Rad6, which is required for histone H2B ubiquitination on K123, Rad6 is not essential, leaving a critical substrate(s) of BUR kinase unidentified. Here we show that BUR kinase is important for the phosphorylation in vivo of Spt5, a subunit of the essential yeast RNAPII elongation factor Spt4/Spt5, whose human orthologue is DRB sensitivity-inducing factor. BUR kinase can also phosphorylate the C-terminal region (CTR) of Spt5 in vitro. Like BUR kinase, the Spt5 CTR is important for promoting elongation by RNAPII and recruiting the PAF complex to transcribed regions. Also like BUR kinase and the PAF complex, the Spt5 CTR is important for histone H2B K123 monoubiquitination and histone H3 K4 and K36 trimethylation during transcription elongation. Our results suggest that the Spt5 CTR, which contains 15 repeats of a hexapeptide whose consensus sequence is S[T/A]WGG[A/Q], is a substrate of BUR kinase and a platform for the association of proteins that promote both transcription elongation and histone modification in transcribed regions.
Keywords: chromatin, PAF, Rtf1, RNAPII, DSIF
Elongation by RNA polymerase II (RNAPII) involves a number of factors that interact directly or indirectly with the enzyme (1). Among these RNAPII-interacting proteins in the budding yeast Saccharomyces cerevisiae are the RNA chain cleavage factor TFIIS, the chromatin assembly factor Spt16/Pob3 (known as FACT in higher eukaryotes), the polymerase-associated factor (PAF) complex, and Spt4/Spt5, which is orthologous to DRB sensitivity-inducing factor (DSIF) in humans. These factors can associate with the transcription complex from the promoter-proximal region to the cleavage and polyadenylation signal or the transcription termination site. In higher eukaryotes, DSIF and the negative elongation factor (NELF), which appears not to exist in S. cerevisiae, restrain elongation by RNAPII soon after initiation (2). As a consequence, many inactive genes in higher eukaryotes have “paused” or “poised” RNAPII molecules downstream of their promoters (3).
An important event during elongation by RNAPII is phosphorylation of the heptapeptide repeats (consensus sequence YSPTSPS) in the C-terminal domain (CTD) of Rpb1, the largest subunit of RNAPII (4). Phosphorylation on S5 of the CTD repeats is catalyzed by the Kin28 subunit (CDK7 in humans) of the general initiation factor TFIIH. S5 phosphorylation tends to decline across yeast transcription units (5, 6) and is reversed by the Ssu72 subunit (7) of the cleavage and polyadenylation factor, which first associates with the transcription complex at or near promoters (8). S2 phosphorylation is catalyzed mainly or entirely by Ctk1, the catalytic subunit of CTD kinase 1 (CTDK1) in yeast (9, 10), and is reversed by the CTD phosphatase Fcp1 (11). Although there is little evidence that S2 phosphorylation in yeast affects elongation by RNAPII, S2 phosphorylation does promote recruitment of the cleavage and polyadenylation factors (12, 13).
S2 phosphorylation in higher eukaryotes is catalyzed by the CDK9 subunit of positive transcription elongation factor b (P-TEFb) (14). Although there is some evidence that S2 phosphorylation can promote elongation by human RNAPII (15), relief of the negative effects of DSIF and NELF also involves their phosphorylation by P-TEFb (2). P-TEFb phosphorylates the C-terminal region (CTR) of the SPT5 subunit of human DSIF on the threonine residues of pentapeptide repeats whose consensus sequence is GS[R/Q]TP (16). In S. cerevisiae, the 2 cyclin-dependent kinases Ctk1 and Bur1 (also known as Sgv1) are most closely related to CDK9. Bur1 and its cyclin, Bur2, promote elongation by RNAPII (10) and can phosphorylate RNAPII in vitro (17), but there is little evidence that Bur1 phosphorylates the CTD of Rpb1 in vivo (10). Thus, the mechanism by which BUR kinase promotes elongation by RNAPII has been unclear.
Transcription by RNAPII is closely connected to histone modification in transcribed regions. Methylation of histone H3 on K36 by Set2 depends on the PAF complex that associates with elongating RNAPII, as well as association of Set2 with RNAPII that has been phosphorylated on S2 by Ctk1 (18–20). H3 K36 methylation causes histone deacetylation by the Rpd3S complex, which prevents aberrant initiation by RNAPII at cryptic TATA boxes in coding regions (21–23). Ubiquitination of histone H2B on K123 involves the E2 ubiquitin conjugating enzyme Rad6 and the E3 ubiquitin ligase Bre1 (24–26) and depends on the PAF complex (27, 28). Both ubiquitination of H2B and its deubiquitination by the Ubp8 subunit of SAGA assist elongation by RNAPII (29, 30). Ubiquitination also depends on BUR kinase, partly because BUR is needed for the efficient recruitment of the PAF complex (28) and also because BUR phosphorylates Rad6 on S120 (31). Trimethylation of histone H3 on K4 by the Set1 subunit of COMPASS depends on phosphorylation of the CTD repeats on S5 by Kin28 (32). Because it also depends on ubiquitination of H2B K123 (33, 34) and the PAF complex, which mediates the association of COMPASS with RNAPII (35), trimethylation of H3 K4 depends on the BUR kinase as well (28, 31, 36). Thus, the identification of BUR kinase substrates should provide considerable insight into histone modification in transcribed regions.
We show here that BUR kinase is important for phosphorylation of yeast Spt5 in vivo and is capable of phosphorylating the Spt5 CTR in vitro. In turn, the Spt5 CTR is needed for both efficient elongation by RNAPII and histone H2B ubiquitination and H3 methylation in transcribed regions. Thus, the Spt5 CTR mediates, at least in part, the effects of BUR kinase on both transcription elongation and histone modification.
Results
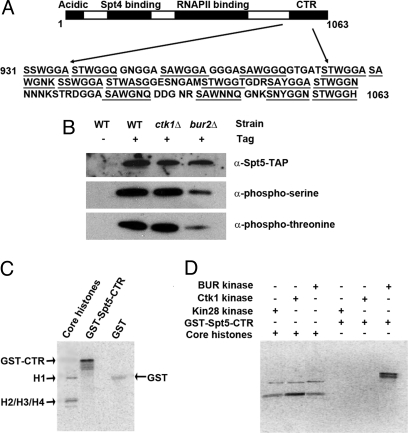
The C-terminal 133 aa of yeast Spt5 contain 15 repeats of a hexapeptide whose consensus sequence is S[T/A]WGG[Q/A] (Fig. 1A). Because the yeast cyclin-dependent kinases Ctk1 and Bur1 are related, to a similar extent, to CDK9 in higher eukaryotes and the CDK9 subunit of P-TEFb phosphorylates threonine residues in repeats whose consensus sequence is G-S-R/Q-T-P in the SPT5 subunit of DSIF (16), we decided to test whether mutation of Ctk1 or the BUR kinase affects the phosphorylation of Spt5 in S. cerevisiae. Because BUR1 is an essential gene, we made use of a deletion of BUR2, which encodes the cyclin subunit of BUR kinase. Spt5 was derivatized with a C-terminal tandem affinity purification (TAP) tag (37) in a wild-type strain or strains with a deletion of CTK1 or BUR2, then extracts from these cells were precipitated with IgG-Sepharose, which binds the protein A component of the TAP tag, and the precipitates were subjected to Western blotting with antibodies that recognize phosphoserine or phosphothreonine and IgG recognizing the TAP tag.
Fig. 1.
Yeast Spt5 is a substrate of the BUR kinase. (A) Sequence of the Spt5 C-terminal region. The region contains 15 repeats (underlined) of a hexapeptide whose consensus sequence is S[T/A]WGG[Q/A]. (B) BUR kinase is needed for phosphorylation of Spt5 in vivo. Spt5-TAP was immunoprecipitated from extracts of the indicated strains with IgG Sepharose and the precipitates were Western-blotted with the indicated antibodies. (C) BUR kinase can directly phosphorylate the Spt5 CTR in vitro. (D) Bur2-TAP, Ctk1-TAP, and Kin28-TAP were purified from TAP-tagged yeast strains and used, as indicated, to phosphorylate chicken core histones and GST-Spt5 CTR fusion protein in vitro.
As shown in Fig. 1B, Spt5 is phosphorylated on both serine and threonine in a wild-type strain. Moreover, both types of phosphorylation are essentially unaffected when CTK1 is deleted, but substantially reduced when BUR2 is deleted. Because the levels of Spt5, as reflected by Western blotting of whole-cell extracts with IgG, are only slightly reduced by deletion of CTK1 or BUR2, these results suggested that BUR kinase but not CTDK1 is responsible, directly or indirectly, for the phosphorylation in vivo of yeast Spt5 on both serine and threonine residues.
Because P-TEFb phosphorylates human SPT5 directly, we hypothesized BUR would do the same in yeast. To test this hypothesis, we affinity-purified BUR kinase from a strain containing a TAP tag on the C-terminus of Bur2 and tested its ability to phosphorylate the CTR of yeast Spt5 (amino acids 931-1063), produced in Escherichia coli as a fusion protein with N-terminal GST. As shown in Fig. 1C, BUR kinase phosphorylates the GST-Spt5CTR fusion protein in vitro, although BUR kinase is also able to phosphorylate GST to a small extent when it is included in the reaction at ≈10-fold molar excess over the GST-Spt5CTR. To control for kinase specificity, we also purified 2 other yeast CDKs, TFIIH and CTDK1, both of which also participate in transcription by RNAPII via TAP tags on their kinase catalytic subunits, Kin28 and Ctk1, respectively. As shown in Fig. 1D, only BUR kinase, but not TFIIH or CTDK1, is able to phosphorylate GST-Spt5CTR in vitro even though all 3 enzymes phosphorylate recombinant chicken core histones, presumably nonspecifically, to about the same extent. These results implied that BUR kinase can specifically phosphorylate the Spt5CTR in vitro. Because BUR kinase is important or required for the phosphorylation of Spt5 in vivo and can phosphorylate the Spt5 CTR in vitro, we concluded that it is likely BUR kinase directly phosphorylates the Spt5 CTR in vivo.
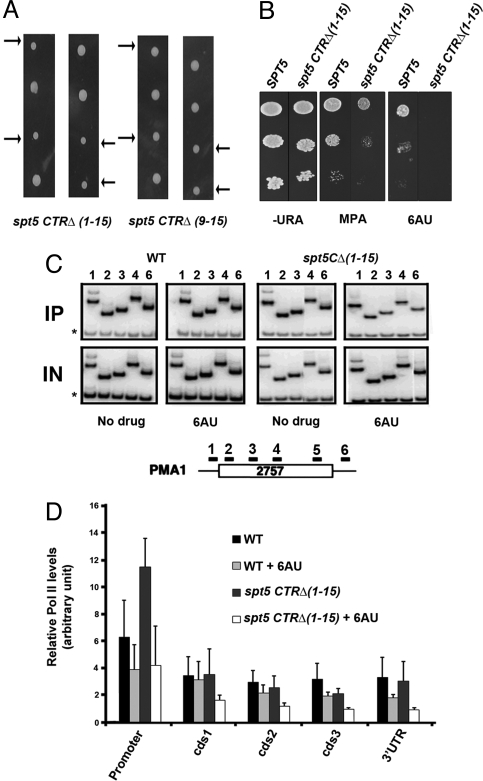
Because the BUR kinase is required for transcriptional elongation (10, 28), we tested whether the Spt5 CTR is also associated with this process. First, to examine the effect on cell growth of deleting the CTR, we created diploid strains in which one of the SPT5 genes was truncated such that 6 of the 15 CTR repeats or all 15 repeats were deleted and replaced with a HA tag, then examined the growth of haploid spores derived from these strains. As shown in Fig. 2A Right, deletion of the 6 C-terminal repeats causes only a very slight growth defect, whereas deletion of all 15 repeats causes a more substantial growth defect (Fig. 2A Left).
Fig. 2.
The Spt5 CTR is important for growth and promotes transcription elongation. (A) A diploid strain was transformed with a 3xHA-His3MX6 cassette replacing either the entire C-terminal region (repeats 1–15) of Spt5 or only part of the CTR (repeats 10–15). After sporulation, tetrads were dissected and individual spores were tested for growth. Left shows 2 representative tetrads from diploids in which all 15 repeats were removed. Right shows 2 representative tetrads from diploids in which 6 of 15 repeats were removed. The arrows point to the 2 spores (meiotic products) of each tetrad that segregate with the CTR deletion. (B) Deletion of the Spt5 CTR causes sensitivity to elongation inhibitors. Ten-fold serial dilutions of SPT5 and spt5ctrΔ(1–15) strains were tested for growth on plates in the presence of mycophenolic acid (MPA) or 6-AU. (C) Representative chromatin immunoprecipitation (ChIP) showing that elongation by RNAPII is less efficient when the Spt5 CTR is deleted. Chromatin was prepared from SPT5 and spt5 CTRΔ(1–15) strains grown in the presence or absence of 6-AU, as indicated. Chromatin was precipitated with 8WG16 monoclonal antibody that recognizes RNAPII and analyzed using primers specific for the PMA1 locus. A diagram of the PMA1 locus is shown to indicate the location of the ChIP primer pairs (from ref. 56). The asterisks indicate a nontranscribed sequence near the telomere of chromosome 5 that is coamplified in each PCR as a control for nonspecific precipitation. IN, input DNA; IP, precipitated DNA. (D) Quantification of ChIP data like that shown in Fig. 2C, with error bars indicating standard deviations. Relative enrichment was calculated as [(Experimental band IP)/(Control band IP)]/[(Input Experimental band)/(Input Control band)]. Promoter, cds1,2,3, and 3′UTR correspond to primer pairs 1, 2, 3, 4, and 6, respectively.
We also tested wild-type and spt5 CTRΔ strains in the presence of 2 compounds that can inhibit growth, 6-azauracil (6-AU) and mycophenolic acid (MPA). Growth sensitivity in the presence of these drugs, which cause lowered intracellular concentrations of nucleotides used as substrates by RNAPII, is thought to be diagnostic of a defect in transcriptional elongation (38). As shown in Fig. 2B, deletion of the Spt5 CTR created slight sensitivity to MPA and extreme sensitivity to 6-AU. These results are consistent with the hypothesis that the Spt5 CTR promotes elongation by RNAPII. To test this idea more directly, we used ChIP to examine localization of RNAPII in vivo on 2 constitutively expressed genes, PMA1 (Fig. 2C; quantification in Fig. 2D) and ADH1 (Fig. S1), in the presence or absence of 6-AU. In the presence of 6-AU, although less so in its absence, deletion of the CTR caused a significant reduction in polymerase density throughout the coding regions of these genes, but the effect was much more pronounced at the 3′ ends of the genes than at the promoter regions. Similar observations on polymerase density were made by Keogh et al. (10) for temperature-sensitive mutations in the BUR1 gene, suggesting that, like BUR kinase, the Spt5 CTR also promotes elongation by RNAPII.
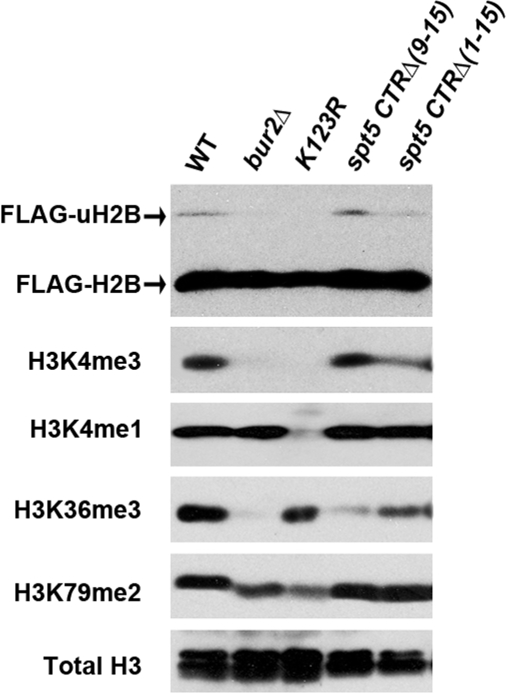
BUR kinase is also required for histone modification in transcribed regions, specifically H2B K123 ubiquitination and H3 trimethylation on K4 and K36 (28, 31, 39, 40), and we hypothesized that some or all of these events could also be at least partly mediated by the Spt5 CTR. To test the effect of the CTR on H2B K123 ubiquitination, strains were constructed in which the Spt5 mutations were transferred to a strain background that contained 1 H2B allele with a FLAG tag (34). As judged by Western blotting with anti-FLAG antibody (Fig. 3 and Fig. S2), the major, higher mobility form of H2B was unchanged in abundance when the 6 C-terminal repeats or all 15 C-terminal repeats were removed from Spt5 (Fig. 3, top gel). However, the lower mobility band in the wild type was greatly reduced when all of the repeats were removed (Fig. 3 and Fig. S2). This band represents the monoubiquitinated form of H2B because it was virtually eliminated when BUR2 was deleted or when the K123 ubiquitinated residue (31) in H2B was mutated to arginine. Therefore, the Spt5 CTR is important for the ubiquitination of H2B, an event that is mediated by Rad6 and Bre1.
Fig. 3.
Cotranscriptional histone modification depends on the Spt5 CTR. All strains contained FLAG-tagged histone H2B and were derived from strain YZS246 (34). Whole-cell extracts were Western-blotted with either M2 anti-FLAG antibody (top gel) or the indicated antibodies.
Because trimethylation of H3K4 and H3K36 are also reduced when the BUR kinase is mutated (28, 31, 36, 39), we also used antibodies specific for various methylated forms of histone H3 in Western blotting experiments to test whether the Spt5 CTR is also needed for these events. Although monomethylation (Fig. 3 and Fig. S2) of H3K4 and dimethylation of H3K79 (Fig. 3 and Fig. S2) are unaffected by deletion of the Spt5 CTR, we found that deletion of the Spt5 CTR reduced trimethylation of both H3K4 and H3K36 (Fig. 3). Both H3K4 and H3K36 trimethylation are markers of active transcription (41, 42) and were eliminated when BUR2 was deleted. These observations indicated again that the BUR kinase and the Spt5 CTR function in overlapping pathways, consistent with our finding that BUR kinase is needed for the phosphorylation of Spt5.
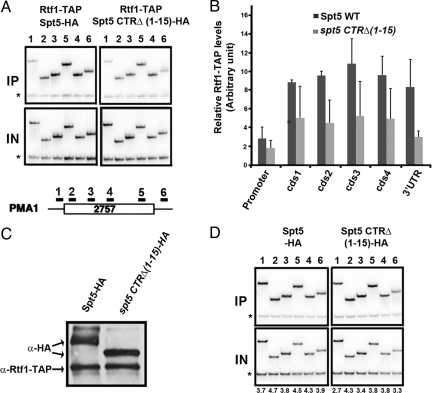
Ubiquitination of H2B and trimethylation of H3K4 and H3K36 require not only the BUR kinase and the Spt5 CTR but also the PAF complex that associates with RNAPII (18, 35, 40). Also, like the BUR kinase and the Spt5 CTR, the PAF complex is important for elongation (43). Because the BUR kinase is required for the recruitment of the PAF complex to transcribed regions (28), the possibility that the Spt5 CTR might also be important for recruitment of the PAF complex was raised. To test this idea, we used ChIP with antibody against TAP-tagged Rtf1, a subunit of the PAF complex, to monitor the effect of the Spt5 CTR on the association of the PAF complex with transcribed regions. As shown in Fig. 4A and Fig. S3 and quantified in Fig. 4B, the PAF complex was found throughout the coding regions (44, 45) of the constitutively transcribed PMA1 and ADH1 loci in a wild-type strain, and this association was reduced when the Spt5 CTR was deleted. As shown in the Western blot experiment of Fig. 4C, this reduced recruitment of Paf1 cannot be accounted for by reduced expression of Rtf1 in the Spt5 CTR mutant. Therefore, reduced recruitment of the PAF complex could account for reduced RNAPII elongation efficiency, H2B ubiquitination, and H3K4 and H3K36 trimethylation when the Spt5 CTR is deleted. An Spt5 requirement for recruitment of the PAF complex is consistent with previous observations that SPT5 and the PAF complex interact genetically and physically (46) and that the Spt4 subunit of yeast DSIF is also required for the recruitment of the PAF complex (47).
Fig. 4.
The Spt5 C-terminal domain is required for optimal recruitment of the PAF complex. (A) Recruitment of the PAF complex to PMA1is reduced in the spt5 CTRΔ(1–15) mutant. ChIP was performed with anti-TAP antibody on the indicated strains containing TAP tags on the Rtf1 subunit of the PAF complex. A diagram of the PMA1 locus is shown to indicate location of the ChIP primer pairs (from ref. 56). The asterisks indicate a nontranscribed sequence near the telomere of chromosome 5 that is coamplified in each PCR as a control for nonspecific precipitation. IN, input DNA; IP, precipitated DNA. (B) Quantification of ChIP data like that shown in Fig. 2B, with error bars indicating standard deviations. Promoter, cds1,2,3,4, and 3′UTR correspond to primer pairs 1–6, respectively. (C) The Spt5 CTR is not required for the expression or stability of Rtf1. Yeast whole cell extracts from the indicated strains containing HA tags on Spt5 were Western blotted with anti-HA antibody, which also binds to the protein A moiety of the TAP tag on Rtf1 in these strains. (D) The Spt5 CTR is not required for recruitment of Spt5. ChIP was performed with an anti-HA antibody on chromatin prepared from yeast strains with SPT5-HA or spt5 CTRΔ(1–15)-HA. Calculated enrichment scores are shown below each lane.
The Western blot of Fig. 4C shows that a form of Spt5 that migrates more slowly in an SDS gel disappears when the CTR is deleted (Fig. 4C), perhaps because hyperphosphorylation of its CTR retards the gel mobility of Spt5. As well, the expression of Spt5 may be slightly reduced when its CTR is deleted, perhaps because the large Spt5 protein is encoded by a long gene and so reduced elongation efficiency caused by deletion of the Spt5 CTR may reduce the synthesis of Spt5 itself. Because reduced recruitment of Spt5 caused by deletion of its CTR could account for reduced recruitment of the PAF complex, we performed ChIP with an anti-HA antibody to monitor recruitment of either Spt5-HA or Spt5 CTRΔ-HA to the PMA1 gene (Fig. 4D). This result demonstrated that deletion of the Spt5 CTR causes no detectable difference in the recruitment of Spt5 to the coding regions or 3′UTR of PMA1. Therefore, it is possible that the CTR itself acts as a platform for the assembly of the PAF complex, which subsequently influences the histone modification enzymes and elongation by RNAPII.
Discussion
In higher eukaryotes, P-TEFb is responsible for phosphorylating both RNAPII on S2 of the heptapeptide repeats of the CTD of its largest subunit RPB1 and the negatively acting elongation factors DSIF (SPT4/SPT5) and NELF (2). Phosphorylation of NELF causes its dissociation from the elongation complex (2), whereas phosphorylation of DSIF on threonine of GS[R/Q]TP repeats in the SPT5 CTR converts DSIF from a negatively acting elongation factor to a positively acting one (16). Although the equivalent of NELF appears not to exist in S. cerevisiae, we have shown here that yeast Spt5 is also phosphorylated in vivo, and that its CTR, which contains 15 repeats of a hexapeptide whose consensus sequence is S[T/A]WGG[Q/A], can be phosphorylated by BUR kinase in vitro. Our data also indicate that the CTR of yeast Spt5 has a positive role during transcription elongation, much like that of the mammalian SPT5. One possibility is that the Spt5 CTR binds one or more factors that create a chromatin environment conducive to transcription elongation.
The 2 yeast CDKs most closely related to CDK9, the catalytic subunit of P-TEFb, are Ctk1 and Bur1 (37% and 40% identity and 59% and 59% similarity, respectively). Deletion of CTK1 eliminates all or almost all of the S2 phosphorylation of the yeast Rpb1 CTD, whereas BUR kinase seems to have little, if any, role in S2 phosphorylation of the CTD in vivo (10). We have shown here that phosphorylation of Spt5 on both serine and threonine requires BUR kinase in vivo, whereas deletion of the CTK1 gene has little or no effect. Because purified BUR kinase can phosphorylate the recombinant Spt5 CTR in vitro; whereas similarly purified CTDK1(whose catalytic subunit is Ctk1) and TFIIH (whose catalytic subunit is Kin28) do not, although all 3 yeast CDKs nonspecifically phosphorylate histones in vitro to similar extents; it seems likely that BUR kinase is responsible for the direct phosphorylation of the Spt5 CTR in vivo. Therefore, 2 of the roles of the CDK9 subunit of P-TEFb in higher eukaryotes, namely S2 phosphorylation of the RPB1 CTD and phosphorylation of the SPT5 CTR repeats, have apparently been devolved in S. cerevisiae to 2 separate CDKs, Ctk1 and Bur1. Presumably, gene duplication followed by further evolution led to subfunctionalization of the roles of higher eukaryotic CDK9 in S. cerevisiae.
P-TEFb has a critical role in regulating elongation by RNAPII in higher eukaryotes (2). In particular, association of P-TEFb with a complex containing 7SK RNA and the Hexim proteins inhibits P-TEFb function, whereas recruitment of P-TEFb to the elongation complex by the activator protein Tat, of HIV, which interacts with both HIV transactivation-response element RNA located downstream of the HIV promoter and the cyclin subunits of P-TEFb, stimulates elongation by RNAPII. Given that a large fraction of the inactive genes in higher eukaryotes contain molecules of RNAPII that have paused downstream of their promoters (48), it is likely that recruitment of P-TEFb plays an important role in the activation of many genes, perhaps even most genes, in these organisms.
In S. cerevisiae, BUR kinase also has a key role in promoting elongation by RNAPII (10). Chromatin assembly factors like Spt16-Pob3 (FACT) and Spt6 cause nucleosome reassembly behind elongating yeast RNAPII (49). These nucleosomes are methylated on K36 by Set2 (50), which interacts with S2-phosphorylated elongating RNAPII (18–20), and deacetylated by the Rpd3S complex, which is recruited by the H3 K36 trimethylation mark generated by Set2 (21–23). This process prevents initiation by RNAPII at cryptic TATA boxes in coding regions (22), but it should also generate closed chromatin as a barrier to elongation by subsequent molecules of RNAPII. Perhaps as a consequence, the BUR kinase is required for elongation unless SET2 or a gene encoding a subunit of Rpd3S is deleted (21). Because we have shown that BUR kinase can phosphorylate the Spt5 CTR, and the Spt5 CTR promotes elongation by RNAPII, our results suggest that the effect of BUR kinase on elongation is mediated, at least in part, by phosphorylation of the Spt5 CTR. However, the negative effect on cell growth of deleting the Spt5 CTR is not suppressed by deleting the SET2 gene (Fig. S4). Thus, although the BUR kinase in S. cerevisiae seems to promote elongation by phosphorylating the Spt5 CTR, just as P-TEFb promotes elongation in higher eukaryotes by phosphorylating the SPT5 subunit of DSIF, it would appear to be a different activity of BUR kinase that overcomes the negative effects on elongation of Set2 and Rpd3S. Because SET2 deletion did not fully suppress the growth defect caused by deleting the BUR kinase (21), it is likely that BUR kinase stimulates elongation by at least 2 different mechanisms, only one of which overcomes the negative effects on elongation of the pathway involving Set2 and Rpd3S.
Consistent with this hypothesis, the BUR kinase is essential for viability whereas the CTR of yeast Spt5 is not, indicating again that the Spt5 CTR is not the only target of BUR kinase whose phosphorylation promotes elongation by RNAPII. BUR kinase also phosphorylates Rad6, leading to the ubiquitination of histone H2B on K123 (31, 51). Because K123 ubiquitination also has a positive effect on elongation by RNAPII (52), it could be that it is H2B ubiquitination rather than Spt5 CTR phosphorylation that helps overcome the inhibiting effects of Set2 and Rpd3S on elongation. In this scenario, BUR kinase might be essential because it phosphorylates both Rad6 and the Spt5 CTR, neither of which is essential on its own. Indeed, we did find that combining deletion of the Spt5 CTR with a C-terminal TAP tag on RAD6 causes synthetic sickness (Fig. S5), suggesting that BUR kinase may indeed be essential because it phosphorylates both Rad6 and the Spt5 CTR.
Another possibility is that failure to phosphorylate the Spt5 CTR may itself be lethal. In this scenario, deletion of the Spt5 CTR, as we have done, would not be equivalent to a failure to phosphorylate it. For example, the unphosphorylated Spt5 CTR could have a negative effect on elongation, whereas Spt5 without its CTR may have no effect on elongation. Phosphorylation of the Spt5 CTR might then both eliminate the negative effect of the unphosphorylated CTR and cause an additional positive effect on elongation. This idea could be tested by replacing the wild-type CTR of the SPT5 gene with one that cannot be phosphorylated, then examining the effects of the mutation on both cell viability and elongation by RNAPII.
Because the CTR of yeast Spt5 contains 15 repeats of a hexapeptide whose consensus sequence is S[T/A]WGG[Q/A], our results suggest that the Spt5 CTR, like the CTD of the Rpb1 subunit of RNAPII (4), is likely to be a “platform” for the association of regulatory factors. In the case of the Spt5 CTR, our data suggest that these factors would control both transcriptional elongation and cotranscriptional histone modification. The unphosphorylated CTR might interact directly with RNAPII to inhibit elongation, or else bind a negatively acting elongation factor yet to be identified. Conversely, the phosphorylated CTR should bind factors that positively affect elongation and histone modification. The most obvious candidate for interaction with the phosphorylated CTR is the PAF complex, which can enhance elongation and is important for both histone H2B K123 ubiquitination and histone H3 trimethylation on K4 and K36. Consistent with this idea, both the BUR kinase (28) and the Spt5 CTR are needed for the recruitment of the PAF complex to transcribed regions. Although the phosphorylated Spt5 CTR may be important for association of Spt5 with the PAF complex within a transcription complex, we found that coimmunoprecipitation of the PAF complex with Spt5 from soluble yeast cell extracts was not affected by deletion of the Spt5 CTR (Fig. S6). Of course, other positively acting elongation factors whose identities are as yet unknown may also interact with the Spt5 CTR. As well, even if the PAF complex interacts with the Spt5 CTR, it is possible that the CTR also interacts directly with the BUR kinase to stimulate Rad6 phosphorylation or with one or more histone modifying enzymes whose activity is stimulated by the CTR (e.g., COMPASS). Therefore, much remains to be done to determine precisely how the Spt5 CTR influences transcriptional elongation and histone modification.
Methods
Yeast Strains and Strain Construction.
The last 399 or 177 bp of the SPT5 gene were replaced with DNA encoding the epitope tag HA to create C-terminal deletions lacking all 15 C-terminal repeats or 6 of 15 repeats, respectively. The HA tag was PCR-amplified from the plasmid pFA6a-3xHA-HIS3MX6 (53) and integrated into the diploid BY4743 strain by standard lithium acetate transformation. Transformants were selected on synthetic dextrose medium without His. The integration of the epitope tag was verified by PCR and Western blotting by using α-HA (Open Biosystems). The correct strains were then sporulated at room temperature in liquid medium (1% potassium acetate/0.1% yeast extract/0.05% glucose), and tetrads were dissected to identify haploid spores with the C-terminal deletion marked by the HA tag. To construct strains containing TAP tags, the TAP cassette was amplified from the plasmid pBS1539 (Euro Scarf) and transformed into various deletion strains. The yeast strains used in this work are listed in Table S1.
Growth Tests.
Overnight cultures of yeast strains were grown in yeast extract/peptone/dextrose (YPD) or synthetic dextrose media without uracil (SD-Ura) medium at 30 °C and diluted into fresh medium the next day. The cultures were grown to mid-log phase and then diluted to OD600 = 0.2. Cells were spotted on plates in 10-fold serial dilutions. Plates were incubated at 30 °C and photographed each day to monitor growth. For drug tests, 6-AU was added to SD-Ura plates to a concentration of 100 μg/mL, and mycophenolic acid was added to a concentration of 15 μg/mL.
Protein Extraction and Western Blotting.
Trichloroacetic acid extraction was performed as described in ref. 54. To examine states of histone modification, the following antibodies were used in Western blotting: α-H3 (Lake Placid Biologicals); α-H3K4 trimethylation (Abcam); α-H3K4 monomethylation (Abcam); α-H3K4 dimethylation (Abcam); α-H3K36 trimethylation (Abcam); and α-FLAG (Sigma: M2).
Protein Immunoprecipitation.
Yeast were grown overnight to saturation at 30 °C, then diluted into fresh YPD medium at OD600 = 0.2 and grown to mid-log phase. The immunoprecipitation protocol was as described in ref. 10. To examine the phosphorylation states of precipitated proteins, α-phosphoserine (Q5 from Qiagen) and α-phosphothreonine (Q7 from Qiagen) were used.
ChIP.
For most ChIP assays, overnight cultures were grown in YPD, diluted to OD600 = 0.2, and grown until OD600 = 0.5–1.0 at 30 °C. For ChIP assays in the presence of 6-AU, cells were grown in SD-Ura, and at mid-log phase 100 μg/mL 6-AU was added to the culture for 2 h. The ChIP protocol was performed as described in ref. 56.
Yeast Protein Purification.
Yeast protein kinase complexes were purified by growing 1.5-L cultures in YPD to OD600 < 1.0 at 30 °C. Kinases were C-terminally tagged with the TAP tag module in the BY4741 background. The purification procedure was as detailed in ref. 55.
Preparation of GST-Spt5-CTR.
The DNA sequence corresponding to the Spt5 CTR was amplified from pMS4 (a gift of G. Hartzog, University of California, Santa Cruz, CA) by using the Pfx polymerase (Invitrogen). The PCR product was then cloned into the vector pGEX-6p-1 (GE Healthcare) digested with BamHI and NotI. The GST-fused CTR was subsequently expressed in E. coli BL21 by adding IPTG to a concentration of 1 mM. An overnight culture was grown at 37 °C in LB plus ampicillin at 100 μg/mL, diluted 1:500 in 500 mL of LB plus ampicillin, and grown to OD600 = 0.6. After 2–3 h of induction, the culture was collected by centrifugation, washed with ice-cold H2O, and then pellet-frozen at −80 °C in a 50-mL Falcon tube before being thawed and resuspended in 20 mL of GST buffer (20 mM Tris, pH 7.4/1 M NaCl/0.2 mM EDTA/1 mM DTT). The suspension was then lysed by sonication with the Branson Sonifier 450 (4 × 20-s pulses, 50% output, 40 duty cycle). Triton X-100 was added to 1%. The lysate was then centrifuged at 14,000 rpm in a Sorvall SS-34 rotor for 20 min at 4 °C and the supernatant was transferred to a Falcon tube and kept on ice. Five-hundred microliters of 50% equilibrated glutathione-Sepharose (Amersham) resin was added, and the mixture was rotated at 4 °C for 1 h. The Sepharose was then washed 4 times with ice-cold GST buffer (twice with 1 M NaCl and twice with 0.1 M NaCl) and transferred to an Eppendorf tube. To elute the GST fusion protein, 1.5 volumes of glutathione elution buffer (50 mM Tris·HCl, pH 8.0/15 mM reduced glutathione) was used at 4° C for 2 h and the eluate was frozen at −80 °C.
In Vitro Kinase Assays.
The kinase reaction protocol was essentially as described in ref. 10.
Supplementary Material
Acknowledgments.
We thank N. Laribee (University of North Carolina, Chapel Hill, NC) for providing strain YNL001, G. Hartzog (University of California, Santa Cruz, CA) for plasmid pMS4, M. Keogh for technical advice, and Joyce Li and Guoquing Zhong for excellent technical assistance. This work was supported by a National Cancer Institute of Canada grant with funds from the Canadian Cancer Society and Canadian Institutes of Health Research Grant MT-6092. W.H.W.K was supported by a National Sciences and Engineering Research Council of Canada Alexander Graham Bell Graduate Scholarship.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/cgi/content/full/0806302106/DCSupplemental.
References
- 1.Saunders A, Core LJ, Lis JT. Breaking barriers to transcription elongation. Nat Rev Mol Cell Biol. 2006;7:557–567. doi: 10.1038/nrm1981. [DOI] [PubMed] [Google Scholar]
- 2.Peterlin BM, Price DH. Controlling the elongation phase of transcription with P-TEFb. Mol Cell. 2006;23:297–305. doi: 10.1016/j.molcel.2006.06.014. [DOI] [PubMed] [Google Scholar]
- 3.Price DH. Poised polymerases: On your mark… get set… go! Mol Cell. 2008;30:7–10. doi: 10.1016/j.molcel.2008.03.001. [DOI] [PubMed] [Google Scholar]
- 4.Phatnani HP, Greenleaf AL. Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev. 2006;20:2922–2936. doi: 10.1101/gad.1477006. [DOI] [PubMed] [Google Scholar]
- 5.Komarnitsky P, Cho EJ, Buratowski S. Different phosphorylated forms of RNA polymerase II and associated mRNA processing factors during transcription. Genes Dev. 2000;14:2452–2460. doi: 10.1101/gad.824700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schroeder SC, Schwer B, Shuman S, Bentley D. Dynamic association of capping enzymes with transcribing RNA polymerase II. Genes Dev. 2002;14:2435–2440. doi: 10.1101/gad.836300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Krishnamurthy S, He X, Reyes-Reyes M, Moore C, Hampsey M. Ssu72 is an RNA polymerase II CTD phosphatase. Mol Cell. 2004;14:387–394. doi: 10.1016/s1097-2765(04)00235-7. [DOI] [PubMed] [Google Scholar]
- 8.Nedea E, et al. Organization and function of APT, a subcomplex of the yeast cleavage and polyadenylation factor involved in the formation of mRNA and small nucleolar RNA 3′-ends. J Biol Chem. 2003;278:33000–33010. doi: 10.1074/jbc.M304454200. [DOI] [PubMed] [Google Scholar]
- 9.Patturajan M, Conrad NK, Bregman DB, Corden JL. Yeast carboxyl-terminal domain kinase I positively and negatively regulates RNA polymerase II carboxyl-terminal domain phosphorylation. J Biol Chem. 1999;274:27823–27828. doi: 10.1074/jbc.274.39.27823. [DOI] [PubMed] [Google Scholar]
- 10.Keogh MC, Podolny V, Buratowski S. Bur1 kinase is required for efficient transcription elongation by RNA polymerase II. Mol Cell Biol. 2003;23:7005–7018. doi: 10.1128/MCB.23.19.7005-7018.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kobor MS, et al. An unusual eukaryotic protein phosphatase required for transcription by RNA polymerase II and CTD dephosphorylation in S. cerevisiae. Mol Cell. 1999;4:55–62. doi: 10.1016/s1097-2765(00)80187-2. [DOI] [PubMed] [Google Scholar]
- 12.Ahn SH, Kim M, Buratowski S. Phosphorylation of serine 2 within the RNA polymerase II C-terminal domain couples transcription and 3′ end processing. Mol Cell. 2004;13:67–76. doi: 10.1016/s1097-2765(03)00492-1. [DOI] [PubMed] [Google Scholar]
- 13.Ni Z, Schwartz BE, Werner J, Suarez JR, Lis JT. Coordination of transcription, RNA processing, and surveillance by P-TEFb kinase on heat shock genes. Mol Cell. 2004;13:55–65. doi: 10.1016/s1097-2765(03)00526-4. [DOI] [PubMed] [Google Scholar]
- 14.Price DH. P-TEFb, a cyclin-dependent kinase controlling elongation by RNA polymerase II. Mol Cell Biol. 2000;20:2629–2634. doi: 10.1128/mcb.20.8.2629-2634.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Marshall NF, Peng J, Xie Z, Price DH. Control of RNA polymerase II elongation potential by a novel carboxyl-terminal domain kinase. J Biol Chem. 1996;271:27176–27183. doi: 10.1074/jbc.271.43.27176. [DOI] [PubMed] [Google Scholar]
- 16.Yamada T, et al. P-TEFb-mediated phosphorylation of hSpt5 C-terminal repeats is critical for processive transcription elongation. Mol Cell. 2006;21:27–37. doi: 10.1016/j.molcel.2005.11.024. [DOI] [PubMed] [Google Scholar]
- 17.Murray S, Udupa R, Yao S, Hartzog G, Prelich G. Phosphorylation of the RNA polymerase II carboxy-terminal domain by the Bur1 cyclin-dependent kinase. Mol Cell Biol. 2001;21:4089–4096. doi: 10.1128/MCB.21.13.4089-4096.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Krogan NJ, et al. Methylation of histone H3 by Set2 in Saccharomyces cerevisiae is linked to transcriptional elongation by RNA polymerase II. Mol Cell Biol. 2003;23:4207–4218. doi: 10.1128/MCB.23.12.4207-4218.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Xiao T, et al. Phosphorylation of RNA polymerase II CTD regulates H3 methylation in yeast. Genes Dev. 2003;17:654–663. doi: 10.1101/gad.1055503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li B, Howe L, Anderson S, Yates JR, III, Workman JL. The Set2 histone methyltransferase functions through the phosphorylated carboxyl-terminal domain of RNA polymerase II. J Biol Chem. 2003;278:8897–8903. doi: 10.1074/jbc.M212134200. [DOI] [PubMed] [Google Scholar]
- 21.Keogh MC, et al. Cotranscriptional set2 methylation of histone H3 lysine 36 recruits a repressive Rpd3 complex. Cell. 2005;123:593–605. doi: 10.1016/j.cell.2005.10.025. [DOI] [PubMed] [Google Scholar]
- 22.Carrozza MJ, et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell. 2005;123:581–592. doi: 10.1016/j.cell.2005.10.023. [DOI] [PubMed] [Google Scholar]
- 23.Joshi AA, Struhl K. Eaf3 chromodomain interaction with methylated H3–K36 links histone deacetylation to Pol II elongation. Mol Cell. 2005;20:971–978. doi: 10.1016/j.molcel.2005.11.021. [DOI] [PubMed] [Google Scholar]
- 24.Kao CF, et al. Rad6 plays a role in transcriptional activation through ubiquitylation of histone H2B. Genes Dev. 2004;18:184–195. doi: 10.1101/gad.1149604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wood A, et al. Bre1, an E3 ubiquitin ligase required for recruitment and substrate selection of Rad6 at a promoter. Mol Cell. 2003;11:267–274. doi: 10.1016/s1097-2765(02)00802-x. [DOI] [PubMed] [Google Scholar]
- 26.Hwang WW, et al. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol Cell. 2003;11:261–266. doi: 10.1016/s1097-2765(02)00826-2. [DOI] [PubMed] [Google Scholar]
- 27.Wood A, Schneider J, Dover J, Johnston M, Shilatifard A. The Paf1 complex is essential for histone monoubiquitination by the Rad6-Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J Biol Chem. 2003;278:34739–34742. doi: 10.1074/jbc.C300269200. [DOI] [PubMed] [Google Scholar]
- 28.Laribee RN, et al. BUR kinase selectively regulates H3 K4 trimethylation and H2B ubiquitylation through recruitment of the PAF elongation complex. Curr Biol. 2005;15:1487–1493. doi: 10.1016/j.cub.2005.07.028. [DOI] [PubMed] [Google Scholar]
- 29.Henry KW, et al. Transcriptional activation via sequential histone H2B ubiquitylation and deubiquitylation, mediated by SAGA-associated Ubp8. Genes Dev. 2003;17:2648–2663. doi: 10.1101/gad.1144003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Daniel JA, et al. Deubiquitination of histone H2B by a yeast acetyltransferase complex regulates transcription. J Biol Chem. 2004;279:1867–1871. doi: 10.1074/jbc.C300494200. [DOI] [PubMed] [Google Scholar]
- 31.Wood A, Schneider J, Dover J, Johnston M, Shilatifard A. The Bur1/Bur2 complex is required for histone H2B monoubiquitination by Rad6/Bre1 and histone methylation by COMPASS. Mol Cell. 2005;20:589–599. doi: 10.1016/j.molcel.2005.09.010. [DOI] [PubMed] [Google Scholar]
- 32.Ng HH, Robert F, Young RA, Struhl K. Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol Cell. 2003;11:709–719. doi: 10.1016/s1097-2765(03)00092-3. [DOI] [PubMed] [Google Scholar]
- 33.Dover J, et al. Methylation of histone H3 by COMPASS requires ubiquitination of histone H2B by Rad6. J Biol Chem. 2002;277:28368–28371. doi: 10.1074/jbc.C200348200. [DOI] [PubMed] [Google Scholar]
- 34.Sun ZW, Allis CD. Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature. 2002;418:104–108. doi: 10.1038/nature00883. [DOI] [PubMed] [Google Scholar]
- 35.Krogan NJ, et al. The Paf1 complex is required for histone H3 methylation by COMPASS and Dot1p: Linking transcriptional elongation to histone methylation. Mol Cell. 2003;11:721–729. doi: 10.1016/s1097-2765(03)00091-1. [DOI] [PubMed] [Google Scholar]
- 36.Mulder KW, Brenkman AB, Inagaki A, van den Broek NJ, Timmers HT. Regulation of histone H3K4 tri-methylation and PAF complex recruitment by the Ccr4-Not complex. Nucleic Acids Res. 2007;35:2428–2439. doi: 10.1093/nar/gkm175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Rigaut G, et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol. 1999;17:1030–1032. doi: 10.1038/13732. [DOI] [PubMed] [Google Scholar]
- 38.Reines D. Use of RNA yeast polymerase II mutants in studying transcription elongation. Methods Enzymol. 2003;371:284–292. doi: 10.1016/S0076-6879(03)71021-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chu Y, Sutton A, Sternglanz R, Prelich G. The BUR1 cyclin-dependent protein kinase is required for the normal pattern of histone methylation by SET2. Mol Cell Biol. 2006;26:3029–3038. doi: 10.1128/MCB.26.8.3029-3038.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Chu Y, Simic R, Warner MH, Arndt KM, Prelich G. Regulation of histone modification and cryptic transcription by the Bur1 and Paf1 complexes. EMBO J. 2007;26:4646–4656. doi: 10.1038/sj.emboj.7601887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Gerber M, Shilatifard A. Transcriptional elongation by RNA polymerase II and histone methylation. J Biol Chem. 2003;278:26303–26306. doi: 10.1074/jbc.R300014200. [DOI] [PubMed] [Google Scholar]
- 42.Hampsey M, Reinberg D. Tails of intrigue: Phosphorylation of RNA polymerase II mediates histone methylation. Cell. 2003;113:429–432. doi: 10.1016/s0092-8674(03)00360-x. [DOI] [PubMed] [Google Scholar]
- 43.Rondón AG, Gallardo M, García-Rubio M, Aguilera A. Molecular evidence indicating that the yeast PAF complex is required for transcription elongation. EMBO Rep. 2004;5:47–53. doi: 10.1038/sj.embor.7400045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Krogan NJ, et al. RNA polymerase II elongation factors of Saccharomyces cerevisiae: A targeted proteomics approach. Mol Cell Biol. 2002;22:6979. doi: 10.1128/MCB.22.20.6979-6992.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Pokholok DK, Hannett NM, Young RA. Exchange of RNA polymerase II initiation and elongation factors during gene expression in vivo. Mol Cell. 2002;9:799–809. doi: 10.1016/s1097-2765(02)00502-6. [DOI] [PubMed] [Google Scholar]
- 46.Squazzo SL, et al. The Paf1 complex physically and functionally associates with transcription elongation factors in vivo. EMBO J. 2002;21:1764–1774. doi: 10.1093/emboj/21.7.1764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Qiu H, Hu C, Wong CM, Hinnebusch AG. The Spt4p subunit of yeast DSIF stimulates association of the Paf1 complex with elongating RNA polymerase II. Mol Cell Biol. 2006;26:3135–3148. doi: 10.1128/MCB.26.8.3135-3148.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wade JT, Struhl K. The transition from transcriptional initiation to elongation. Curr Opin Genet Dev. 2008;18:130–136. doi: 10.1016/j.gde.2007.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kaplan CD, Laprade L, Winston F. Transcription elongation factors repress transcription initiation from cryptic sites. Science. 2003;301:1096. doi: 10.1126/science.1087374. [DOI] [PubMed] [Google Scholar]
- 50.Strahl BD, et al. Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol Cell Biol. 2002;22:1298–1306. doi: 10.1128/mcb.22.5.1298-1306.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wood A, Shilatifard A. Bur1/Bur2 and the Ctk complex in yeast: The split personality of mammalian P-TEFb. Cell Cycle. 2006;5:1066–1068. doi: 10.4161/cc.5.10.2769. [DOI] [PubMed] [Google Scholar]
- 52.Xiao T, et al. Histone H2B ubiquitylation is associated with elongating RNA polymerase II. Mol Cell Biol. 2005;25:637–651. doi: 10.1128/MCB.25.2.637-651.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Longtine MS, et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast. 1998;14:953–961. doi: 10.1002/(SICI)1097-0061(199807)14:10<953::AID-YEA293>3.0.CO;2-U. [DOI] [PubMed] [Google Scholar]
- 54.Keogh MC, et al. The Saccharomyces cerevisiae histone H2A variant Htz1 is acetylated by NuA4. Genes Dev. 2006;20:660–665. doi: 10.1101/gad.1388106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Kim M, et al. Transitions in RNA polymerase II elongation complexes at the 3′ ends of genes. EMBO J. 2004;23:354–364. doi: 10.1038/sj.emboj.7600053. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.