Abstract
Background
Chemoreception is vitally important for all animals, yet little is known about the genetics of chemoreception in aquatic organisms. The keystone species Daphnia pulex, a well known crustacean, is the first aquatic invertebrate to have its genome sequenced. This has allowed us the initial investigation of chemoreceptor genes in an aquatic invertebrate, and to begin the study of chemoreceptor evolution across the arthropod phylum.
Results
We describe 58 Grs (gustatory receptors), belonging to the insect chemoreceptor superfamily, which were identified bioinformatically in the draft genome of the crustacean waterflea Daphnia pulex. No genes encoding proteins similar to the insect odorant receptors (Ors) were identified. These 58 Grs form 3 distinctive subfamilies of 37, 12, and 5 genes, as well as a highly divergent singleton (Gr58). In addition, Grs55–57 share distinctive amino acid motifs and cluster with the sugar receptors of insects, and may illuminate the origin of this distinctive subfamily. ESTs, tiling array, and PCR amplification results support 34 predicted gene models, and preliminary expression data comparing the sexes indicates potential female-biased expression for some genes.
Conclusion
This repertoire of 58 chemoreceptors presumably mediates the many chemoperception abilities of waterfleas. While it is always possible that the entire Or gene lineage was lost at some point in the history of Daphnia pulex, we think it more likely that the insect Or lineage is indeed a relatively recently expanded gene lineage concomitant with the evolution of terrestriality in the insects or their hexapod ancestors.
Background
The ability of Daphnia to detect chemical cues released by prey or predator have been glimpsed through studies on feeding behavior and predator avoidance [1,2]. Daphnids reject food particles, adjust feeding currents according to food availability, quality and surrounding chemical cues, and appear to swim and remain in areas where food is abundant [3-6]. However, their presence in food abundant areas can be altered by the presence of predators, and their vertical distribution is often associated with the presence or absence of predators [6,7].
Kairomones from predators, fish or invertebrate, affect Daphnia's swimming patterns, dial vertical migration, and even affect morphology [2,8-11]. Species of Daphnia can develop neck teeth, thicker carapaces, and/or long head spines to reduce their vulnerability to predation [12,13]. Predator chemical signals, both fish and invertebrate kairomones, share some similarities. For example, in the case of Leucaspius delineates and Chaoborus americanus, these kairomones are made up of more than one active component with low-molecular weight and are organic water-soluble molecules with intermediate polarity, that have no primary amines and require hydroxyl groups for activity; they are also heat stable molecules that can be partially destroyed by acid and base digestions [14,15]. When the kairomones of different fish species were compare there was a striking resemblance between both groups, indicating that the signals are very similar if not identical and are found free in solution and not bound to edible particles [15]. However progress is still slow in the identification of the molecular nature of kairomones, and we are yet to isolate any individual chemical that can invoke a robust chemical mediated behavior in aquatic invertebrates [16].
The cues involved in Daphnia mating are not well understood. Unlike copepod males that can trace a female signal in the water column [17], Daphnia males must come into contact with potential mates [18,19]. This inspection is quick and although the cues needed to tell sexes and species apart could be mechanical; it is also possible that a female pheromone is present on the sexual female's carapace, allowing males to quickly identify a mate. The possibility that a chemical cue is involved in mating is hinted at by the fact that males press their antennules against a potential mate, and these antennules are structurally identical to known chemosensors found on other crustaceans and the first antennae of terrestrial insects (also known as chemosensors) [20,21].
In insects a chemoreceptor superfamily of seven-transmembrane domain proteins (TM7) provides the molecular basis for the specificity and sensitivity of both smell and taste (recently reviewed by [22-25]). The superfamily consists of the gustatory receptor (Gr) family [26-28], which contains most of the protein diversity of the superfamily [29], and the odorant receptor (Or) family [30-32], which is a single highly expanded lineage [29]. The Or superfamily has now been described in a variety of insects. These include both endopterygote relatives of the Drosophila melanogaster fruitflies in which they were first discovered, for example, the other 11 Drosophila species with genome sequences [33-36], as well as the mosquitoes Anopheles gambiae [37] and Aedes aegypti [38,39], the silkworm moth Bombyx mori [40,41], the red flour beetle Tribolium castaneum [42,43], and the honey bee Apis mellifera [44]. While this chemoreceptor superfamily is clearly very old with distant relatives of the Grs identified in the Caenorhabditis nematodes, Robertson et al. (2003) suggested that the Ors might be a relatively recent expansion of dedicated odorant receptors from a particular Gr lineage concomitant with the evolution of terrestriality in insects from a crustacean ancestor. The availability of a draft genome sequence for the waterflea Daphnia pulex [45], a representative of the freshwater branchiopod crustaceans thought by some to be the sister group to the terrestrial insects (e.g. [46]), allows a first test of this proposal.
Here we describe the chemoreceptor superfamily revealed by the draft genome sequence for D. pulex, finding six lineages of Grs, including one expanded to 37 genes, for a total of 58 genes. These presumably mediate the many "taste" functions in this freshwater crustacean. Consistent with the prediction of Robertson et al. (2003), we find no evidence of Ors. This includes the basal and highly conserved ortholog of the unusual DmOr83b protein implicated in partnering with each of the specific Ors in individual olfactory sensory neurons [47-52]. While it is always possible that this entire Or gene lineage was lost at some point in the history of Daphnia pulex, we think it more likely that the insect Or lineage is indeed a relatively recently expanded gene lineage concomitant with the evolution of terrestriality in the insects or their hexapod ancestors.
Results
Absence of Ors
Extensive BLASTP searches of the predicted proteins encoded by the v1.0, NCBI GNOMON, and merged v1.1 gene builds provided by the JGI at DOE, as well as TBLASTN searches of the September 2006 draft genome sequence using representative Grs and Ors from all available insects as queries revealed only multiple lineages of Grs. In particular, no homolog of the otherwise highly conserved DmOr83b protein, which has orthologs in all available insect genomes, was identified. It is always possible that a particular gene might be in a region of a genome that cloned poorly in the genomic libraries employed in a genome project, and hence was sequenced too thinly to be assembled. We therefore also searched all 2,724,768 raw traces deposited in the Trace Archive at GenBank using the TBLASTN algorithm for any reads with sequence similarity to all available DmOr83b orthologs from insects, and found none. Similar searches with representative insect Ors similarly revealed no convincing matches. We conclude that the D. pulex genome does not encode a homolog of the DmOr83b protein or any other insect Or homologs and that the entire insect Or gene family is absent from this crustacean genome.
A diversity of Grs
We identified fifty eight genes encoding proteins belonging to the Gr family (Table 1 and Figure 1). About half of these genes are found in tandem arrays across 21 scaffolds in the sequenced genome (Table 1). While genes within tandem arrays are usually phylogenetic close to each other in the tree, there has been considerable gene movement within the genome. For example, although Grs1–9 cluster together in the tree, they are in three tandem arrays spaced across 2 Mbp on scaffold 4. Grs47–52 form a phylogenetic cluster, and most are in a tandem array on scaffold 2, but Gr47 is on scaffold 58.
Table 1.
Daphnia pulex gustatory receptor (Gr) gene model support.
|
DpuGr |
Location |
JGI V1.1 gene model |
Protein ID |
New Protein ID |
Comments |
| 1 | scaffold_4:272236-273762 | fgenesh1_pg.C_scaffold_4000034 | 346811 | NA | Same |
| 2 | scaffold_4:278009-279502 | NCBI_GNO_0400033 | 311261 | 346819 | truncated 1st exon |
| 3 | scaffold_4:279988-281469 | NCBI_GNO_0400034 | 311262 | 346813 | missing final exon |
| 4 | scaffold_4:341660-343135 | fgenesh1_pg.C_scaffold_4000053 | 95937 | 346911 | truncated 1st exon |
| 5 | scaffold_4:339828-341493 | PASA_GEN_0400197 | 305579 | NA | Same |
| 6 | scaffold_4:2188983-2190473 | NCBI_GNO_0400391 | 311617 | NA | Same |
| 7 | scaffold_4:2190837-2192326 | NCBI_GNO_0400392 | 311618 | 346837 | 4th exon too long & missing 5th exon |
| 8 | scaffold_4:2192733-2194232 | NCBI_GNO_0400393 | 311619 | NA | Same |
| 9 | scaffold_4:2194646-2196117 | NCBI_GNO_0400394 | 311620 | 346838 | match on all exons but NCBI model has extras at 3' end |
| 10 | scaffold_4:2634693-2636319 | NCBI_GNO_0400515 | 311740 | 346840 | truncated 1st exon missing last exon |
| 11 | scaffold_145:138652-140036 | fgenesh1_pg.C_scaffold_145000046 | 115102 | 346841 | missing 1st intron & last (5th) exon |
| 12 | scaffold_87:435725-437154 | NCBI_GNO_8700117 | 327171 | NA | Same |
| 13 | scaffold_87:437579-438974 | SNAP_00023793 | 255735 | 346842 | 1st exon and truncated 2nd exon |
| 14 | scaffold_87:441383-442931 | fgenesh1_pg.C_scaffold_87000145 | 111713 | 346843 | 4th intron too long |
| 15 | scaffold_40:105929-107556 | NCBI_GNO_4000025 | 321270 | 346844 | last (5th) exon missing |
| 16N | scaffold_40:103864-105444 | NCBI_GNO_4000024 | 321269 | NA | Same |
| 17 | scaffold_87:211272-212862 | NCBI_GNO_8700061 | 327115 | 346847 | 5' end missing 9 bp |
| 18 | scaffold_87:213286-214940 | NCBI_GNO_8700062 | 327116 | 346848 | 5' end missing 9 bp |
| 19 | scaffold_87:196838-198317 | NCBI_GNO_8700054 | 327108 | 346849 | 5th exon too long |
| 20 | scaffold_87:193344-194997 | NCBI_GNO_8700053 | 327107 | 346850 | truncated 1st exon & missing 3rd exon |
| 21 | scaffold_87:191113-192511 | NCBI_GNO_8700052 | 327106 | NA | Same |
| 22 | scaffold_87:187203-188649 | SNAP_00023720 | 255662 | NA | Same |
| 23FIX | scaffold_87:185158-186582 | NCBI_GNO_8700050 | 327104 | 442586 | missing last 3 exons |
| 24 | scaffold_87:30125-31504 | NCBI_GNO_8700006 | 327060 | NA | Same |
| 25 | scaffold_87:31824-33484 | SNAP_00023664 | 255606 | 346855 | truncated 1st exon & missing 5th exon |
| 26P | scaffold_328:49769-50798 | NCBI_GNO_32800005 | 334296 | 442583 | missing last 2 exons of our model |
| 27 | scaffold_4:2218191-2219707 | SNAP_00002848 | 234790 | 346857 | 5th exon mismatch |
| 28 | scaffold_4:2216102-2217599 | SNAP_00002847 | 234789 | 346858 | missing 1st exon |
| 29 | scaffold_4:2213168-2214628 | NCBI_GNO_0400395 | 311621 | 346859 | 5th intron is longer |
| 30 | scaffold_51:492193-493689 | NCBI_GNO_5100060 | 323020 | 346860 | extra intron within 2nd exon |
| 31 | scaffold_86:355128-356818 | SNAP_00023611 | 255553 | 346861 | missing 4th exon & truncated 6th exon |
| 32 | scaffold_66:753423-755119 | NCBI_GNO_6600115 | 325025 | 346862 | missing last 3 exons |
| 33 | scaffold_117:358469-360263 | fgenesh1_pg.C_scaffold_117000028 | 113818 | 346863 | truncated 1st exon |
| 34FIX | scaffold_29:299592-300898 | no hit | NA | 442578 | |
| 35FIX | scaffold_123:44710-46019 | NCBI_GNO_12300006 | 329587 | 442580 | truncated 1st exon & missing 6th exon |
| 36 | scaffold_123:46645-48216 | NCBI_GNO_12300007 | 329588 | 346866 | truncated 1st exon |
| 37 | scaffold_187:187229-188772 | NCBI_GNO_18700047 | 332335 | 346867 | truncated 1st & 2nd exons |
| 38 | scaffold_187:180574-182181 | NCBI_GNO_18700046 | 332334 | 346875 | 1st exon missing & longer 4th exon |
| 39 | scaffold_187:182801-184413 | PASA_GEN_18700024 | 302748 | 346876 | truncated 1st exon and 6th exon too long |
| 40P | scaffold_187:184972-186472 | fgenesh1_pg.C_scaffold_187000047 | NA | NA | |
| 41 | scaffold_187:177577-179164 | no hit | NA | 346878 | |
| 42 | scaffold_4:2636875-2638477 | NCBI_GNO_0400516 | 311741 | 346879 | truncated 1st exon & 3 exons instead of 2 |
| 43 | scaffold_87:433963-435377 | NCBI_GNO_8700116 | 327170 | NA | same |
| 44N | scaffold_6:1830849-1832318 | NCBI_GNO_0600407 | 312608 | 442555 | JGI – 5' 1st exon missing |
| 45 | scaffold_6:1833035-1834297 | NCBI_GNO_0600408 | 312609 | 346880 | truncated 5' end |
| 46 | scaffold_8:1391176-1392681 | fgenesh1_pg.C_scaffold_8000220 | 98040 | NA | same |
| 47 | scaffold_58:302684-304219 | NCBI_GNO_5800046 | 323957 | 346882 | 1st & 2nd exons missing/3rd |
| 48 | scaffold_2:711166-709624 | no hit | NA | NA | exon truncated |
| 49 | scaffold_2:705282-706818 | NCBI_GNO_0200131 | 310197 | 346895 | partial, last 4 exons only |
| 50 | scaffold_2:702774-704369 | NCBI_GNO_0200130 | 310196 | 346897 | extra intron within 1st exon |
| 51 | scaffold_2:700887-702432 | NCBI_GNO_0200129 | 310195 | NA | same |
| 52N | scaffold_2:399077-400562 | NCBI_GNO_0200074 | 310142 | 442581 | truncated 1st intron |
| 53 | scaffold_13:642296-644073 | NCBI_GNO_1300117 | 315056 | NA | same |
| 54 | scaffold_138:252456-255386 | SNAP_00028520 | 260462 | 346908 | 1st & 4th exon missing |
| 55 | scaffold_6:842460-843909 | SNAP_00003790 | 235732 | 346901 | 4th exon missing & truncated 5th exon |
| 56 | scaffold_6:840584-842029 | NCBI_GNO_0600186 | 312387 | NA | same |
| 57 | scaffold_4:2311538-2313083 | NCBI_GNO_0400416 | 311642 | 346902 | same |
| 58 | scaffold_24:135381-137169 | NCBI_GNO_2400021 | 318197 | NA | same |
The location and protein ID plus the newly annotated protein ID for each gene model found in the Daphnia genome V1.1 is given, along with annotation comments. Genes in the first column followed by the letter P indicates Pseudogene, N indicates predicted models needed revision, and FIX indicates gene models that were not initially predicted and were manually curated.
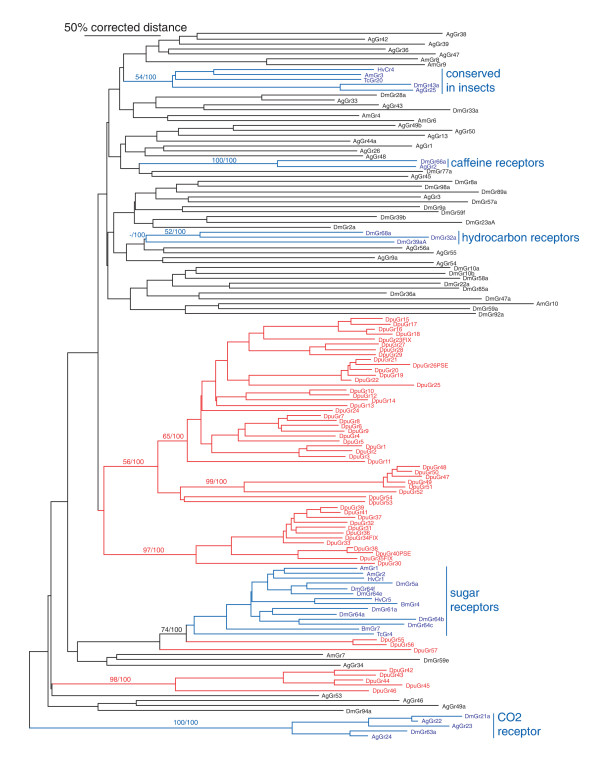
Figure 1.
Phylogenetic relationships of the 58 Daphnia pulex Grs to each other and a representative set of insect Grs. This is a corrected distance tree, with the highly conserved CO2 receptor lineage designated as the outgroup to root the tree. Bootstrap values from 10,000 replications of uncorrected distance analysis are shown on major branches, followed by Bayesian posterior probabilities. DpuGr (D. pulex) lineages are highlighted in red. Major groups of insect Grs whose ligands are known or which are mentioned in the text are highlighted in blue (Ag – Anopheles gambiae, Am – Apis mellifera, Bm-Bombyx mori, Hv- Heliothis virescens, and Tc- Tribolium castaneum).
These Grs are fairly easily recognized through their somewhat conserved TM7 regions near the C-terminus, which includes a TYhhhhhQF motif in TM7. Almost all Gr genes in D. pulex have a phase-0 intron six codons before this motif, an intron that is present in most Gr genes in insects as well as their nematode relatives, the gur genes [29]. The only exceptions are the divergent Gr42–46 subfamily (see below). These Daphnia Grs align fully with the insect Grs, including a cluster of hydrophobic amino acids at the N-terminus that includes a few conserved amino acids. We note that several fragmentary or highly degenerate pseudogenes also exist in this genome which we have not named or included in our analyses.
There are three well-conserved and distinctive lineages within the insect Grs that one might anticipate finding in the D. pulex set. The first lineage is the carbon dioxide receptors, exemplified by the heterodimeric pair Gr21a and Gr63a in Drosophila melanogaster [53,54] and the heterotrimeric set Gr22–24 in Anopheles gambiae [37,55], which is present in moths and beetles as well [55,56]. Remarkably this otherwise highly conserved lineage is absent from all other available more basal insect and arthropod genomes, including D. pulex [56].
The second lineage is the sugar receptors, consisting of eight Grs in D. melanogaster (Gr5a, 61a, and 64a-f) [57-60], nine Grs in the three available mosquitoes [39], five in the silkmoth Bombyx mori [41], sixteen in the flour beetle Tribolium castaneum [43], and two in the honey bee Apis mellifera [44]. This highly divergent set of proteins has several amino acids that are distinctive, most prominently a glutamic acid (E) residue immediately after the conserved TY pair in TM7, although the functional significance of these residues is unknown. Three DpuGrs have such a residue, Grs55–57, and they cluster with the insect sugar receptors near the base of the tree in our phylogenetic analysis, although there is only bootstrap support for Gr55 and 56 clustering with the insect sugar receptors (Figure 1). Bayesian analysis actually suggests that these two proteins cluster within this sugar subfamily, internal to TcGr4 and BmGr7. These insect sugar receptors have a distinctive set of intron locations [29], and only the last two are shared with Gr55–57, number 2 and 3 in [29]. These last two introns are shared across the entire superfamily and hence are not diagnostic of the sugar receptors. We propose that at least DpuGr55 and 56 are functional sugar receptors, perhaps representing the origins of this sensory specificity in arthropods, from which the insect sugar receptors evolved with considerable sequence and gene structure evolution.
The third conserved lineage of insect Grs is the DmGr43a protein and relatives in other species (AgGr25, AaGr34, HvCr4, BmGr9/10, TcGr20–28 and 183, and AmGr3), however there is no obvious ortholog in D. pulex. Nor are there obvious orthologs for the DmGr66a protein implicated in bitter taste in Drosophila e.g[61], or the candidate hydrocarbon receptors DmGr68a, 32a, and the 39a protein set [22,62]. Neither of the latter two observations is surprising as these receptors are only conserved in flies, indeed the latter three only in drosophilid flies.
Instead, most of the remaining D. pulex Grs form three distinctive gene subfamilies without obvious relatives in the available insect genomes. The first consists of 37 proteins in the middle of Figure 1 in two well-supported clusters, specifically Grs1–29, and 47–54. A second subfamily of 12 genes, Grs30–41, share a gene structure with the above subfamily, with three phase-0 introns at the C-terminus, called 1–3 in [29], that are shared by all the insect chemoreceptor lineages (the only exception is Gr53, which lost the first of these three). All 49 of these genes also share a phase-0 intron about half way along the genes, which may be unique to these Daphnia Grs (it also appears to be present in Grs55–58, however the alignment is less definitive in them).
A third highly divergent subfamily consists of Grs42–46, which have a completely different gene structure, having lost all three of the ancestral phase-0 introns near the C-terminus. Grs42–44 appear to have phase-1 introns near their N-termini, Gr45 is intronless in its coding region, and Gr46 has two internal phase-2 introns.
Finally, Gr58 is a particularly highly divergent protein with a long branch hence was not included in Figure 1, nevertheless it has all the hallmarks of a Gr, including the TYhhhhhQF motif in TM7 with a phase-0 intron immediately before the final exon encoding this motif (as well as two internal phase-0 introns and one phase-2 intron). There are two fragmentary and highly degenerate pseudogene copies of Gr58 in the genome, one immediately downstream of it in scaffold_24 and another in scaffold_21. Similarly highly degenerate pseudogene copies exist for other Grs, such as Gr27 and 47.
Expression of Grs in Daphnia
Insect Grs are generally expressed at low levels in only a few gustatory or olfactory sensory neurons and studies in insects are largely limited to Drosophila melanogaster where promoter::LacZ or promoter::GFP fusion transgenes have allowed visualization of their expression patterns (e.g. [63-65]). Transformation techniques are not yet available for Daphnia, so we examined the only available large study of Daphnia gene expression, an unpublished Nimblegen genome tiling array experiment comparing males and females using whole bodies, performed in conjunction with the genome project (J. Colbourne personal communication). This reveals generally low but convincing levels of expression for 27 of these genes (Figure 2). Gr11, 13, 15, 45, and 53 show particularly high levels of expression, of which all but Gr11, have female-biased expression. Only one slightly male-biased receptor was identified (Gr6). PCR amplification of a subset of Grs from female and male cDNA supported expression for 11 genes and some showing negligible expression on the tiling array were also verified using qRT/PCR amplification from whole bodies. This investigation revealed that 7 genes having negligible expression on the tiling array, are indeed expressed (Figure 2). There is no obvious pattern of expression level with clustering of genes in the phylogenetic tree (data not shown).
Figure 2.
Daphnia pulex Gr expression. The bars represent tiling array results which where qualitatively analyzed; expression differences were assessed based on average height of signal for each gene between the sexes. We also indicate other types of expression support from ESTs, cDNA amplification, and qRT/PCR. Black bars – female support; gray bars – male support; Black filled diamond – Genes that were successfully amplified using standard PCR techniques; black filled star – genes with EST support; and black filled circle- genes amplified through qRT/PCR.
Discussion
We describe the 58 Grs we found encoded by the draft Daphnia genome sequence. We believe these constitute the entirety of the "insect" chemoreceptor superfamily in D. pulex. This superfamily of odorant and gustatory receptors was identified originally in D. melanogaster and has been identified in all other insects with sequenced genomes, and it was anticipated that it would also be present in other arthropods. The absence of the Or family, a single particularly highly divergent and expanded lineage within the superfamily, is consistent with the prediction of Robertson et al. (2003) that the insect Or family evolved with terrestriality in insects or their immediate hexapod ancestors, although sequences of additional crustaceans, other arthropods, and basal hexapods, will be required to test this hypothesis further. We have undertaken several steps to identify all members of the Gr family, including highly sensitive TBLASTN searches using only the somewhat conserved TM7 region of these proteins, and HMMER searches of all available predicted proteins using all available Grs in the model set. Grs can sometimes be extraordinarily divergent, however, so it remains possible that some have been missed. For example, Kent et al. (2008) report five new Gr genes in the Anopheles gambiae genome that were missed by Hill et al. (2002) because they are so highly divergent and automated gene models for them were not sufficiently well built to find them using PSI-BLASTP searches.
The only Daphnia Grs with a clear relationship to particular insect Gr lineages are Gr55 and 56, and perhaps Gr57, which cluster with the sugar receptor subfamily. This indicates that Daphnia likely can sense some sugars, presumably dissolved in water and perhaps indicating food sources [66]. Despite extensive searches we find no orthologs of the other well-known and highly conserved Gr lineage in insects, the carbon dioxide heterotrimeric receptors, represented by DmGr21a and 63a [53-56]. This is perhaps not surprising given that Daphnia are not known to be able to sense carbon dioxide, although it appears that Daphnia epphipia (or resting eggs), do respond and at times require a carbon dioxide signal to hatch (see [67]). The only other relatively well conserved Gr lineage in insects is that of DmGr43a, AgGr25, HvCr4, and AmGr3, however the conservation here is insufficient to expect to find this lineage in Daphnia (Figure 1). The remaining insect Grs for which ligands are known, DmGr66 for caffeine [61] and DmGr68a and 32a for cuticular hydrocarbons [22,62], are dipteran-specific lineages, hence were not expected to have Daphnia orthologs.
Instead we believe there are only three other major Gr subfamilies in Daphnia, all expansions within crustaceans, consisting of 37, 12, and 5 genes. The highly divergent Gr58 might represent another subfamily that may be more evident in other crustaceans.
An interesting feature of some of these Daphnia Grs, e.g. 31–34, 36, 37, 39, and 41, is that they end immediately after the conserved TYhhhhhQF motif which forms the core of TM7. These are the shortest versions of Grs known, and indicate that the C-terminus of these proteins is unlikely to be involved in any important interactions with other proteins. This situation is compatible with recent findings that the insect chemoreceptors likely have the opposite membrane topology to the TM7 GPCRs [50,52,68], because the C-terminus would be external to the cell where no significant interactions with proteins in any signaling transduction machinery would be expected. They therefore support the hypothesis that these chemoreceptors are not coupled to G-proteins and instead function as ligand-gated ion channels [69,70].
Conclusion
This repertoire of 58 Grs presumably underlies the many abilities of Daphnia to sense their external chemical environment, which they do using both a classic "taste" mode involving physical contact with objects, as well as what might be considered a "smell" mode in which they sense dissolved chemicals in the water. As elaborated in the Introduction, these include food, potential mating partners, and potential threats like fish. Therefore, we suspect that these genes will be expressed in identified chemosensors, such as the first antennule and feeding appendages [71]. Our preliminary assessment of expression levels of these chemoreceptors comparing males and females reveals apparent female-biased expression for a few of them, but no clearly male-specific receptors that might perceive sex differences. The next obvious step in studies of these Daphnia Grs will be to determine their expression patterns more precisely. Initially this will be achieved by RT/PCR studies of surgically separated structures, like the antennules, although this is technically challenging but achievable for such tiny animals. While in situ hybridization might allow more refined studies of their expression patterns, in D. melanogaster at least, Grs typically are expressed at too low levels for reliable in situ hybridization. Ultimately studies using promoter::GFP fusion transgenes might be required to establish confident expression patterns once transgenic techniques are developed for Daphnia. It will be of particular interest to determine whether any of these six gene lineages, for example perhaps the most highly expanded 37 and 12 gene subfamilies, is exclusively expressed in the antennules or swimming antennae, in which case these might constitute the effective "olfactory" receptors of Daphnia.
Methods
Known insect chemoreceptors whose sequences have been entered in to GENBANK (National Center for Biotechnology Information) were used to search for similar genes in the Daphnia genome sequence. Protein sequences were used to perform TBLASTN [72] searches of assembled scaffolds available through two websites: Joint Genome Institute (JGI) Daphnia pulex V 1.0 and V 1.1 [73] and Daphnia Genome BLAST [74]. Genomic scaffold sequences were used to constructgenes manually in the PAUP*v4 [75] and MEGAv4 [76] text editors, using comparisons with known exons and online programs to predict exon/intron splice sites [77,78]. Divergent Daphnia proteins were used in iterative rounds of TBLASTN searches to find additional genes. In three cases genes were truncated by the ends of contigs, but in each case the complete gene sequence could be assembled with the aid of raw reads, and these are indicated by the suffix FIX after their names. Two genes in the named set are clear pseudogenes, with internal frameshifting deletions, and are indicated by the suffix PSE. All proteins were aligned using CLUSTALX [79], and gene models refined to fix apparent alignment difficulties. Intron locations and phases were located in the alignment in the text editor of PAUP to assist gene model refinement and subfamily analysis. All proteins are available as a FASTA file [see additional file 1].
Our manually curated gene models were compared with the set of 30,907 gene models generated by the JGI known as v1.1. They were also validated through nr, SwissPro and Pfam hits. In summary, 13 gene models were identical, 13 needed minor revisions, and 29 needed modification, and 3 (Grs 34, 41, 48) were completely unannotated. 44 genes genes where supported by nr, SwissPro and Pfam hits, with the drosophilid Gr64 sugar receptor family supporting DpuGr 55 and 56 as potential sugar receptors. We also compared our gene models with preliminary tiling array expression (NimbleGen, Madison, WI) results to see if expressed exons agreed with our predicted models, and 27 gene models gained additional support thereby.
For phylogenetic analysis, representative insect Grs, primarily from Drosophila melanogaster, Anopheles gambiae, with a few from Bombyx mori, Heliothis virescens, Tribolium castaneum, and Apis mellifera, were included in the alignment for comparison. The length-divergent N- and C-terminal regions, as well as internal regions with major alignment gaps, were removed, leaving 328 aligned amino acid positions. For the main phylogenetic analysis, corrected distance was performed in PAUP*v4 using the heuristic search with tree-bisection-and-reconnection branch swapping. Distances were corrected for multiple amino acid replacements in the past using the maximum likelihood model, the BLOSUM62 amino acid exchange matrix, and default settings in TREE-PUZZLE v5.0 [80]. Additional Bayesian analysis was performed using MrBayes v3.1 [81] with the JTT substitution model, four chains, 1 million generations, and two runs. Trees were sampled every 100 generations, discarding a burnin of 250,000 generations.
Using the polymerase chain reaction (PCR) technique we designed primers for assessing expression of a subset of our gene models. This subset included genes having EST and tiling support as well as those lacking any type of support. Primers were designed and tested on both genomic DNA and cDNA of Daphnia pulex male and female clones. Quantitative real-time PCR (qRT/PCR) was run on a few models to assess differences between the sexes and to investigate whether lack of support was due to low levels of expression which standard PCR cannot amplify to detectable levels on a gel.
Abbreviations
Grs: gustatory receptors; Ors: olfactory receptors; ESTs: expressed sequence tags; PCR: polymerase chain reaction; qRT/PCR: quantitative real-time PCR; JGI: Joint Genome Institute; DOE: Department of Energy; TM7: seven transmembrane domain protein.
Authors' contributions
HMR and DCPA annotated all genes and wrote the manuscript. DCPA performed all expression analyses. HMR performed the phylogenetic analyses. ML supervised and helped edit the manuscript.
Supplementary Material
Daphnia pulex Gustatory receptor gene models. FASTA format of all 58 DpuGr gene models.
Acknowledgments
Acknowledgements
The sequencing and portions of the analyses were performed at the DOE Joint Genome Institute under the auspices of the U.S. Department of Energy's Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Livermore National Laboratory under Contract No. W-7405-Eng-48, Lawrence Berkeley National Laboratory under Contract No. DE-AC02-05CH11231, Los Alamos National Laboratory under Contract No. W-7405-ENG-36 and in collaboration with the Daphnia Genomics Consortium (DGC) http://daphnia.cgb.indiana.edu. Additional analyses were performed by wFleaBase, developed at the Genome Informatics Lab of Indiana University with support to Don Gilbert from the National Science Foundation and the National Institutes of Health. Coordination infrastructure for the DGC is provided by The Center for Genomics and Bioinformatics at Indiana University, which is supported in part by the METACyt Initiative of Indiana University, funded in part through a major grant from the Lilly Endowment, Inc. Our work benefits from, and contributes to the Daphnia Genomics Consortium. HMR was funded by NIH grant AI56081 and ML was funded by NSF grant EF0328516.
Contributor Information
D Carolina Peñalva-Arana, Email: dcpenalv@indiana.edu.
Michael Lynch, Email: milynch@indiana.edu.
Hugh M Robertson, Email: hughrobe@uiuc.edu.
References
- Pijanowska J. Alarm Signals in Daphnia? Oecologia. 1997;112:12–16. doi: 10.1007/s004420050277. [DOI] [PubMed] [Google Scholar]
- Roozen F, Lurling M. Behavioural response of Daphnia to olfactory cues from food, competitors and predators. Journal of Plankton Research. 2001;23:797–808. doi: 10.1093/plankt/23.8.797. [DOI] [Google Scholar]
- Burns CW. Direct observations of mechanisms regulating feeding behaviour of Daphnia in lakewater. Internationale Revue der gesamten Hydrobiologia. 1968;53:83–100. doi: 10.1002/iroh.19680530104. [DOI] [Google Scholar]
- De Meester L. Genotype, fish-mediated chemicals, and phototactic behavior in Daphnia magna. Ecology. 1993;74:1467–1474. doi: 10.2307/1940075. [DOI] [Google Scholar]
- Peñalva-Arana DC, Moore PA, Feinberg BA, DeWall J, Strickler JR. Studying Daphnia feeding behavior as a black box: a novel electrochemical approach. Hydrobiologia. 2007;594:153–163. doi: 10.1007/s10750-007-9080-7. [DOI] [Google Scholar]
- Kleiven OT, Larsson P, Hobaek A. Direct distributional response in Daphnia pulex to a predator kairomone. Journal of Plankton Research. 1996;18:1341–1348. doi: 10.1093/plankt/18.8.1341. [DOI] [Google Scholar]
- Rautio M, Korhola A, Zellmer ID. Vertical distribution of Daphnia longispina in a shallow subarctic pond: Does the interaction of ultraviolet radiation and Chaoborus predation explain the pattern? Polar Biology. 2003;26:659–665. doi: 10.1007/s00300-003-0533-9. [DOI] [Google Scholar]
- Stabell OB, Ogbebo F, Primicerio R. Inducible defenses in Daphnia depend on latent alarm signals from conspecific prey activated in predators. Chemical Senses. 2003;28:141–153. doi: 10.1093/chemse/28.2.141. [DOI] [PubMed] [Google Scholar]
- Young JD, Riessen HP. The interaction of Chaoborus size and vertical distribution determines predation effects on Daphnia. Freshwater Biology. 2005;50:993–1006. doi: 10.1111/j.1365-2427.2005.01381.x. [DOI] [Google Scholar]
- Ordemann A, Balazsi G, Moss F. Pattern formation and stochastic motion of the zooplankton Daphnia in a light field. Physica a-Statistical Mechanics and Its Applications. 2003;325:260–266. doi: 10.1016/S0378-4371(03)00204-8. [DOI] [Google Scholar]
- Stirling G. Daphnia Behavior as a Bioassay of Fish Presence or Predation. Functional Ecology. 1995;9:778–784. doi: 10.2307/2390252. [DOI] [Google Scholar]
- Laforsch C, Beccara L, Tollrian R. Inducible defenses: The relevance of chemical alarm cues in Daphnia. Limnology and Oceanography. 2006;51:1466–1472. [Google Scholar]
- Hebert PD, Grewe PM. Chaoborus-Induced Shifts in the Morphology of Daphnia-Ambigua. Limnology and Oceanography. 1985;30:1291–1297. [Google Scholar]
- Parejko K, Dodson SI. Progress towards characterization of a prey/predator kairomone: Daphnia pulex and Chaoborus. Hydrobiologia. 1990;198:51–59. doi: 10.1007/BF00048622. [DOI] [Google Scholar]
- Von Elert E, Loose CJ. Predator-induced diel vertical migration in Daphnia: Enrichment and preliminary chemical characterization of a fish kairomone exuded by fish. Journal of Chemical Ecology. 1996;22:885–895. doi: 10.1007/BF02029942. [DOI] [PubMed] [Google Scholar]
- Pohnert G, Steinke M, Tollrian R. Chemical cues, defence metabolites and the shaping of pelagic interspecific interactions. Trends in Ecology & Evolution. 2007;22:198–204. doi: 10.1016/j.tree.2007.01.005. [DOI] [PubMed] [Google Scholar]
- Yen J, Weissburg M, Doall M. The fluid physics of signal perception by mate-tracking copepods. Philosophical Transactions of the Royal Society of London Series B-Biological Sciences. 1998;353:787–804. doi: 10.1098/rstb.1998.0243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brewer MC. Mating behaviours of Daphnia pulicaria, a cyclic parthenogen: comparisons with copepods. Philosophical Transactions of the Royal Society of London B. 1998;353:805–815. doi: 10.1098/rstb.1998.0244. [DOI] [Google Scholar]
- Winsor GL, Innes DJ. Sexual reproduction in Daphnia pulex (Curstacea: Cladocera): observations on male mating behaviour and avoidance of inbreeding. Freshwater Biology. 2002;47:441–450. doi: 10.1046/j.1365-2427.2002.00817.x. [DOI] [Google Scholar]
- Hallberg E, Johansson KUI, Elofsson R. The Aesthetasc concept: Structural variations of putative olfactory receptor cell complexes in Crustacea. Microscopy Research and Technique. 1992;22:325–335. doi: 10.1002/jemt.1070220403. [DOI] [PubMed] [Google Scholar]
- Dodson S, Frey D. Cladocera and other Branchiopoda. In: Thorpe JH, Covich AP, editor. Ecology and classification of North American freshwater invertebrates. New York: Academic Press; 1991. pp. 723–786. [Google Scholar]
- Ebbs ML, Amrein H. Taste and pheromone perception in the fruit fly Drosophila melanogaster. Pflügers Archiv European Journal of Physiology. 2007;454:735–747. doi: 10.1007/s00424-007-0246-y. [DOI] [PubMed] [Google Scholar]
- Hallem E, Dahanukar A, Carlson JR. Insect odor and taste receptors. Annual Review of Entomology. 2006;51:113–135. doi: 10.1146/annurev.ento.51.051705.113646. [DOI] [PubMed] [Google Scholar]
- Rützler M, Zwiebel LJ. Molecular biology of insect olfaction: recent progress and conceptual models. Journal of Comparative Physiology A. 2005;191:777–790. doi: 10.1007/s00359-005-0044-y. [DOI] [PubMed] [Google Scholar]
- Vosshall LB, Stocker RF. Molecular Architecture of Smell and Taste in Drosophila. Annual Review of Neuroscience. 2007;30:505. doi: 10.1146/annurev.neuro.30.051606.094306. [DOI] [PubMed] [Google Scholar]
- Clyne PJ, Warr CG, Carlson JR. Candidate Taste Receptors in Drosophila. Science. 2000;287:1830. doi: 10.1126/science.287.5459.1830. [DOI] [PubMed] [Google Scholar]
- Dunipace L, Meister S, McNealy C, Amrein H. Spatially restricted expression of candidate taste receptors in the Drosophila gustatory system. Current Biology. 2001;11:822–835. doi: 10.1016/S0960-9822(01)00258-5. [DOI] [PubMed] [Google Scholar]
- Scott K, Brady R, Cravchik A, Morozov P, Rzhetsky A, Zuker C, Axel R. A Chemosensory Gene Family Encoding Candidate Gustatory and Olfactory Receptors in Drosophila. Cell. 2001;104:661–673. doi: 10.1016/S0092-8674(01)00263-X. [DOI] [PubMed] [Google Scholar]
- Robertson HM, Warr CG, Carlson JR. Molecular evolution of the insect chemoreceptor gene superfamily in Drosophila melanogaster. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:14537–14542. doi: 10.1073/pnas.2335847100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clyne PJ, Warr CG, Freeman MR, Lessing D, Kim J, Carlson JR. A novel family of divergent seven-transmembrane proteins: candidate odorant receptors in Drosophila. Neuron. 1999;22:327–338. doi: 10.1016/S0896-6273(00)81093-4. [DOI] [PubMed] [Google Scholar]
- Vosshall LB. Olfaction in Drosophila. Current Opinion in Neurobiology. 2000;10:498–503. doi: 10.1016/S0959-4388(00)00111-2. [DOI] [PubMed] [Google Scholar]
- Vosshall LB, Amrein H, Morozov PS, Rzhetsky A, Axel R. A spatial map of olfactory receptor expression in the Drosophila Antenna. Cell. 1999;96:725–736. doi: 10.1016/S0092-8674(00)80582-6. [DOI] [PubMed] [Google Scholar]
- Guo S, Kim J. Molecular Evolution of Drosophila Odorant Receptor Genes. Molecular Biology and Evolution. 2007;24:1198–1207. doi: 10.1093/molbev/msm038. [DOI] [PubMed] [Google Scholar]
- McBride CS, Arguello JR, O'Meara BC. Five Drosophila genomes reveal nonneutral evolution and the signature of host specialization in the chemoreceptor superfamily. Genetics. 2007;177:1395–1416. doi: 10.1534/genetics.107.078683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nozawa M, Nei M. Evolutionary dynamics of olfactory receptor genes in Drosophila species. Proceedings of the National Academy of Sciences. 2007;104:7122. doi: 10.1073/pnas.0702133104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robertson HM. The insect chemoreceptor superfamily in Drosophila pseudoobscura : molecular evolution of ecologically-relevant genes over 25 million years. Journal of Insect Science. 2008. [DOI] [PMC free article] [PubMed]
- Hill CA, Fox AN, Pitts RJ, Kent LB, Tan PL, Chrystal MA, Cravchik A, Collins FH, Robertson HM, Zwiebel LJ. G protein coupled receptors in Anopheles gambiae. Science. 2002;298:176–178. doi: 10.1126/science.1076196. [DOI] [PubMed] [Google Scholar]
- Bohbot J, Pitts RJ, Kwon HW, Rutzler M, Robertson HM, Zwiebel LJ. Molecular characterization of the Aedes aegypti odorant receptor gene family. Insect Molecular Biology. 2007;0:525–537. doi: 10.1111/j.1365-2583.2007.00748.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kent LB, Walden KKO, Robertson HM. The Gr Family of Candidate Gustatory and Olfactory Receptors in the Yellow-Fever Mosquito Aedes aegypti. Chem Senses. 2007. p. bjm067. [DOI] [PubMed]
- Wanner KW, Anderson AR, Trowell SC, Theilmann DA, Robertson HM, Newcomb RD. Female-biased expression of odourant receptor genes in the adult antennae of the silkworm, Bombyx mori. Insect Molecular Biology. 2007;16:107–119. doi: 10.1111/j.1365-2583.2007.00708.x. [DOI] [PubMed] [Google Scholar]
- Wanner KW, Robertson HM. The gustatory receptors (Gr) family of candidate chemoreceptors from the silkworm moth, Bombyx mori. Molecular Biology. 2008. [DOI] [PubMed]
- Engsontia P, Sanderson AP, Cobb M, Walden KKO, Robertson HM, Brown S. The red flour beetle's large nose: An expanded odorant receptor gene family in Tribolium castaneum. Insect Biochemistry and Molecular Biology. 2008;38:387–397. doi: 10.1016/j.ibmb.2007.10.005. [DOI] [PubMed] [Google Scholar]
- Consortium TGS. The genome model of the beetle and the prest Tribolium castaneum. Nature. 2008;452:949–955. doi: 10.1038/nature06784. [DOI] [PubMed] [Google Scholar]
- Robertson HM, Wanner KW. The chemoreceptor superfamily in the honey bee, Apis mellifera: Expansion of the odorant, but not gustatory, receptor family. Genome Research. 2006;16:1395–1403. doi: 10.1101/gr.5057506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colbourne J, Singan V, Gilbert D. wFleaBase: the Daphnia genome database. BMC Bioinformatics. 2005;6 doi: 10.1186/1471-2105-6-45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glenner H, Thomsen PF, Hebsgaard MB, Sorensen MV, Willerslev E. EVOLUTION: The Origin of Insects. Science. 2006;314:1883. doi: 10.1126/science.1129844. [DOI] [PubMed] [Google Scholar]
- Jones WD, Nguyen TAT, Kloss B, Lee KJ, Vosshall LB. Functional conservation of an insect odorant receptor gene across 250 million years of evolution. Current Biology. 2005;15:R119–R121. doi: 10.1016/j.cub.2005.02.007. [DOI] [PubMed] [Google Scholar]
- Krieger J, Klink O, Mohl C, Raming K, Breer H. A candidate olfactory receptor subtype highly conserved across different insect orders. Journal of Comparative Physiology A: Sensory, Neural, and Behavioral Physiology. 2003;189:519–526. doi: 10.1007/s00359-003-0427-x. [DOI] [PubMed] [Google Scholar]
- Larsson MC, Domingos AI, Jones WD, Chiappe ME, Amrein H, Vosshall LB. Or83b Encodes a Broadly Expressed Odorant Receptor Essential for Drosophila Olfaction. Neuron. 2004;43:703–714. doi: 10.1016/j.neuron.2004.08.019. [DOI] [PubMed] [Google Scholar]
- Lundin C, Käll L, Kreher SA, Kapp K, Sonnhammer EL, Carlson JR, Heijne G, Nilsson IM. Membrane topology of the Drosophila OR83b odorant receptor. FEBS Letters. 2007. [DOI] [PMC free article] [PubMed]
- Neuhaus EM, Gisselmann G, Zhang W, Dooley R, Stoertkuhl K, Hatt H. Odorant receptor heterodimerization in the olfactory system of Drosophila melanogaster. Nature Neuroscience. 2005;8:15–17. doi: 10.1038/nn1371. [DOI] [PubMed] [Google Scholar]
- Benton R, Sachse S, Michnick SW, Vosshall LB. Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol. 2006;4:e20. doi: 10.1371/journal.pbio.0040020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones WD, Cayirlioglu P, Kadow IG, Vosshall LB. Two chemosensory receptors together mediate carbon dioxide detection in Drosophila. Nature. 2007;445:86–90. doi: 10.1038/nature05466. [DOI] [PubMed] [Google Scholar]
- Kwon JY, Dahanukar A, Weiss LA, Carlson JR. The molecular basis of CO2 reception in Drosophila. Proceedings of the National Academy of Sciences. 2007;104:3574. doi: 10.1073/pnas.0700079104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu T, Qiu YT, Wang G, Kwon JY, Rutzler M, Kwon HW, Pitts RJ, van Loon JJA, Takken W, Carlson JR. Odor Coding in the Maxillary Palp of the Malaria Vector Mosquito Anopheles gambiae. Current Biology. 2007;17:1533–1544. doi: 10.1016/j.cub.2007.07.062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robertson HM, Kent LB. Evolution of the gene lineage encoding the carbon dioxide heterodimeric receptor in insects. Journal of Insect Science. 2008. [DOI] [PMC free article] [PubMed]
- Chyb S. Drosophila gustatory receptors: from gene identification to functional expression. Journal of Insect Physiology. 2004;50:469–477. doi: 10.1016/j.jinsphys.2004.03.012. [DOI] [PubMed] [Google Scholar]
- Dahanukar A, Hallem EA, Carlson JR. Insect chemoreception. Curr Opin Neurobiol. 2005;15:423–430. doi: 10.1016/j.conb.2005.06.001. [DOI] [PubMed] [Google Scholar]
- Jiao Y, Moon SJ, Montell C. From the Cover: A Drosophila gustatory receptor required for the responses to sucrose, glucose, and maltose identified by mRNA tagging. Proceedings of the National Academy of Sciences. 2007;104:14110. doi: 10.1073/pnas.0702421104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slone J, Daniels J, Amrein H. Sugar Receptors in Drosophila. Current Biology. 2007;17:1809–1816. doi: 10.1016/j.cub.2007.09.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moon SJ, Köttgen M, Jiao Y, Xu H, Montell C. A Taste Receptor Required for the Caffeine Response In Vivo. Current Biology. 2006;16:1812–1817. doi: 10.1016/j.cub.2006.07.024. [DOI] [PubMed] [Google Scholar]
- Bray S, Amrein H. A Putative Drosophila Pheromone Receptor Expressed in Male-Specific Taste Neurons Is Required for Efficient Courtship. Neuron. 2003;39:1019–1029. doi: 10.1016/S0896-6273(03)00542-7. [DOI] [PubMed] [Google Scholar]
- Marella S, Fischler W, Kong P, Asgarian S, Rueckert E, Scott K. Imaging Taste Responses in the Fly Brain Reveals a Functional Map of Taste Category and Behavior. Neuron. 2006;49:285–295. doi: 10.1016/j.neuron.2005.11.037. [DOI] [PubMed] [Google Scholar]
- Thorne N, Amrein H. Atypical expression of Drosophila gustatory receptor genes in Sensory and Central neurons. The Journal of Comparative Neurology. 2008;506:548–568. doi: 10.1002/cne.21547. [DOI] [PubMed] [Google Scholar]
- Thorne N, Chromey C, Bray S, Amrein H. Taste perception and coding in Drosophila. Current Biology. 2004;14:1065–1079. doi: 10.1016/j.cub.2004.05.019. [DOI] [PubMed] [Google Scholar]
- Nishide E, Anzai H, Uchida N, Nisizawa K. Sugar constituents of fucose-containing polysaccharides from various Japanese brown algae. Hydrobiologia. 1990;204:573–576. doi: 10.1007/BF00040289. [DOI] [Google Scholar]
- Stross RG. Photoperiod control of diapause in Daphnia. IV. Light and CO2-sensitive phases within the cycle of activation. MBL. 1971;140:137–155. doi: 10.2307/1540033. [DOI] [PubMed] [Google Scholar]
- Wistrand M, Kall L, Sonnhammer ELL. A general model of G protein-coupled receptor sequences and its application to detect remote homologs. 2006;15:509–521. doi: 10.1110/ps.051745906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato K, Pellegrino M, Nakagawa T, Nakagawa T, Vosshall LB, Touhara K. Insect olfactory receptors are heteromeric ligand-gated ion channels. Nature. 2008;452:1002–1006. doi: 10.1038/nature06850. [DOI] [PubMed] [Google Scholar]
- Wicher D, Schafer R, Bauernfeind R, Stensmyr MC, Heller R, Heinemann SH, Hansson BS. Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature. 2008;452:1007–1011. doi: 10.1038/nature06861. [DOI] [PubMed] [Google Scholar]
- Hallberg E, Johansson K, Wallen R. Olfactory sensilla in crustaceans: morphology, sexual dimorphism, and distribution patterns. International Journal of Insect Morphology and Embryology. 1997;26:173–180. doi: 10.1016/S0020-7322(97)00019-6. [DOI] [Google Scholar]
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research. 25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- JGI Genome Portal http://genome.jgi-psf.org/Dappu1/Dappu1.home.html
- wFleaBase http://wfleabase.org/blast/index.html
- Swofford DL. PAUP* 4.0: Phylogenetic Analysis Using Parsimony. Sinauer Associates; 2002. [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- BDGPSpliceSitePredictionByNeuralNetwork http://www.fruitfly.org/seq_tools/splice.html
- SplicePredictor http://deepc2.psi.iastate.edu/cgi-bin/sp.cgi
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research. 1997;25:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt HA, Strimmer K, Vingron M, von Haeseler A. TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics. 2002;18:502–504. doi: 10.1093/bioinformatics/18.3.502. [DOI] [PubMed] [Google Scholar]
- Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19:1572–1574. doi: 10.1093/bioinformatics/btg180. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Daphnia pulex Gustatory receptor gene models. FASTA format of all 58 DpuGr gene models.