Abstract
Background
Sequences and organization of the mitochondrial genome have been used as markers to investigate evolutionary history and relationships in many taxonomic groups. The rapidly increasing mitochondrial genome sequences from diverse insects provide ample opportunities to explore various global evolutionary questions in the superclass Hexapoda. To adequately support such questions, it is imperative to establish an informatics platform that facilitates the retrieval and utilization of available mitochondrial genome sequence data.
Results
The Insect Mitochondrial Genome Database (IMGD) is a new integrated platform that archives the mitochondrial genome sequences from 25,747 hexapod species, including 112 completely sequenced and 20 nearly completed genomes and 113,985 partially sequenced mitochondrial genomes. The Species-driven User Interface (SUI) of IMGD supports data retrieval and diverse analyses at multi-taxon levels. The Phyloviewer implemented in IMGD provides three methods for drawing phylogenetic trees and displays the resulting trees on the web. The SNP database incorporated to IMGD presents the distribution of SNPs and INDELs in the mitochondrial genomes of multiple isolates within eight species. A newly developed comparative SNU Genome Browser supports the graphical presentation and interactive interface for the identified SNPs/INDELs.
Conclusion
The IMGD provides a solid foundation for the comparative mitochondrial genomics and phylogenetics of insects. All data and functions described here are available at the web site http://www.imgd.org/.
Background
The mitochondrial genomes of members of the superclass Hexapoda (generally referred to as the 'insects') are typically approximately 15 kilobases (kb) in length and encode 37 genes, including 13 protein coding genes (PCGs), 2 ribosomal RNA genes (rRNAs), and 22 transfer RNA genes (tRNAs). Owing to its small size, high copy number, and relatively infrequent gene rearrangements, the mitochondrial genome has been extensively used for phylogenetic analyses [1-4]. Phylogenetic analysis based on the mitochondrial gene sequences is often limited to closely related species, due to the high rate of nucleotide substitutions. However, variations in the mitochondrial gene content and order have been utilized to elucidate evolutionary relationships among distantly-related species, on the basis of shared derived characteristics that denote the common ancestry of a given group [5].
Recent years, the number of sequenced mitochondrial genomes has been increasing fast due to rapidly growing sequencing capacity [6]. For example, more than 1,200 metazoan mitochondrial genomes have already been sequenced completely [7,8]. The abundance of available mitochondrial genomes has led to the development of the following web-based relational databases that are specialized for archiving the resulting data: GObase [9], AMiGA [10], Mitome [8], MamMibase [11], OGRe [7], and NCBI Organelle Genome Resources [12]. Some of these resources also provide tools for data analysis and/or viewing: MamMibase provides a web-based phylogenetic analysis tool for studying evolutionary relationships on the basis of the archived mitochondrial genomes [11] and Mitome provides a graphical mitochondrial genome browser [8]. In order to more effectively support uses of the rapidly accumulating mitochondrial genome sequences, an integrated platform that provides a diverse array of analysis tools is necessary.
Single nucleotide polymorphisms (SNPs) in the insect mitochondrial genome sequences can support fine-scale phylogenetic analyses, as illustrated in the following examples. Twenty-four biotypes of Bemisia tabaci (Hemiptera: Aleyrodidae), which could not be distinguished by morphological characteristics, were resolved [13]. SNPs in the cytochrome c oxidase subunits I (COI) locus of Aedes aegypti (Diptera: Cuclicidae) were used to differentiate four strains [14]. Based on the fully sequenced mitochondrial genomes in the genus Flustrellidra (Ctenostomata: Flustrellidridae), a SNP analysis was conducted to identify a suitable gene maker for distinguishing morphologically similar species [15].
Partially sequenced mitochondrial genes from a very large number of species also provide valuable markers for phylogenetic analysis. For example, the COI gene has been used extensively for species identification in the 'DNA barcoding' projects [16,17]. In particular, in Hexapoda, DNA barcoding projects covering multiple orders have been conducted [18-21], resulting in at least 50,000 partial sequences of COI loci in the NCBI. Because the cytochrome c oxidase subunits II (COII) locus is relatively small (approximately 600 bp) and can be amplified well by PCR from diverse species [22,23], many researchers have sequenced this locus, yielding more than 15,000 sequences from Hexapoda. Due to the large number of characterized insect species, sequences of these loci are an excellent resource for comprehensive phylogenetic analyses of insects; however, such data have not yet been archived in the currently available mitochondrial genome databases.
A new integrated platform named the Insect Mitochondrial Genome Database (IMGD; http://www.imgd.org/) was developed to better integrate available mitochondrial gene and genome sequences and to provide bioinformatics tools for efficient data retrieval and utilization. The IMGD archives the sequences of 112 completely sequenced and 20 nearly completed mitochondrial genome sequences, as well as partial sequences of 113,985 mitochondrial genomes (Tables 1, 2, 3) from 25,747 insect species using the standardized framework of the Comparative Fungal Genomics Platform (CFGP; http://cfgp.snu.ac.kr/) [24]. SNPs in the mitochondrial genomes of multiple isolates within eight species were identified via the SNP Analysis Platform (SAP; http://sap.snu.ac.kr/; J. Park et al., unpublished) and presented through the SNU Genome Browser (http://genomebrowser.snu.ac.kr/) [25]. BLAST [26], tRNAScan-SE [27], and mFold [28] were also incorporated into IMGD. Additionally, three phylogenetic analysis tools, including ClustalW, PHYML, and PHYLIP [29-31], were integrated into IMGD to facilitate analyses across multiple species: these tools are available through the web interface supported by Phyloviewer (http://www.phyloviewer.org/; B. Park et al., unpublished). To assist the comparison of these sequences and phylogenetic analysis within selected taxa, a new user interface, termed the Species-driven User Interface (SUI), was designed and implemented. The IMGD provides a highly integrated environment for conducting evolutionary studies of insects using their mitochondrial gene/genome sequences.
Table 1.
List of the number of mitochondrial sequences in Hexapoda archived in the IMGD
| Order | Species | CGa | NGb | PGc |
| Archaeognatha | 10 | 4 | 0 | 15 |
| Blattaria | 274 | 1 | 0 | 991 |
| Coleoptera | 6,594 | 8 | 4 | 25,783 |
| Collembola | 133 | 6 | 2 | 947 |
| Dermaptera | 35 | 0 | 0 | 63 |
| Diplura | 10 | 3 | 0 | 24 |
| Diptera | 3,846 | 24 | 2 | 26,982 |
| Embioptera | 14 | 0 | 0 | 26 |
| Ephemeroptera | 288 | 1 | 0 | 742 |
| Grylloblattodea | 17 | 0 | 1 | 113 |
| Hemiptera | 1,851 | 20 | 5 | 7,299 |
| Hymenoptera | 4,144 | 3 | 2 | 14,737 |
| Isoptera | 647 | 7 | 0 | 2,842 |
| Lepidoptera | 4,556 | 7 | 2 | 19,380 |
| Mantodea | 188 | 1 | 0 | 717 |
| Mantophasmatodea | 17 | 1 | 0 | 194 |
| Mecoptera | 61 | 0 | 0 | 141 |
| Megaloptera | 7 | 2 | 0 | 409 |
| Neuroptera | 143 | 2 | 0 | 437 |
| Odonata | 525 | 0 | 1 | 1,734 |
| Orthoptera | 919 | 14 | 0 | 4,744 |
| Phasmatodea | 64 | 0 | 1 | 482 |
| Phthiraptera | 527 | 3 | 0 | 2,155 |
| Plecoptera | 184 | 1 | 0 | 529 |
| Protura | 2 | 0 | 0 | 6 |
| Psocoptera | 121 | 1 | 0 | 338 |
| Raphidioptera | 4 | 0 | 0 | 5 |
| Siphonaptera | 35 | 0 | 0 | 134 |
| Strepsiptera | 6 | 0 | 0 | 7 |
| Thysanoptera | 154 | 1 | 0 | 857 |
| Trichoptera | 343 | 0 | 0 | 1,100 |
| Zoraptera | 1 | 0 | 0 | 2 |
| Zygentoma | 27 | 2 | 0 | 50 |
| Total | 25,747 | 112 | 20 | 113,985 |
aCompletely sequenced mitochondrial genome, bNearly completely sequenced mitochondrial genome, and cPartially sequenced mitochondrial gene sequences
Table 2.
List of 56 whole mitochondrial genomes of hexapod species (Part I. 52 holometabolous species) archived in IMGD
| Order | Species | Size (bp) | GC (%) | PCGs | tRNAs | rRNAs |
| Coleoptera | Chaetosoma scaritides* | 15,511 | 20.96 | 13 | 22 | 2 |
| Coleoptera | Crioceris duodecimpunctata | 15,880 | 23.11 | 13 | 22 | 2 |
| Coleoptera | Cyphon sp. BT0012 | 15,919 | 24.83 | 13 | 22 | 2 |
| Coleoptera | Priasilpha obscura* | 16,603 | 23.49 | 13 | 22 | 2 |
| Coleoptera | Pyrocoelia rufa | 17,739 | 22.59 | 13 | 22 | 2 |
| Coleoptera | Pyrophorus divergens | 16,120 | 30.56 | 13 | 22 | 2 |
| Coleoptera | Rhagophthalmus lufengensis | 15,982 | 20.37 | 13 | 22 | 2 |
| Coleoptera | Rhagophthalmus ohbai | 15,704 | 20.85 | 13 | 19 | 2 |
| Coleoptera | Sphaerius sp. BT0074* | 15,121 | 19.28 | 13 | 22 | 2 |
| Coleoptera | Tetraphalerus bruchi | 15,689 | 33.01 | 13 | 22 | 2 |
| Coleoptera | Trachypachus holmbergi* | 15,722 | 20.54 | 13 | 22 | 2 |
| Coleoptera | Tribolium castaneum | 15,881 | 28.32 | 13 | 22 | 2 |
| Diptera | Anopheles funestus* | 15,354 | 21.84 | 7 | 22 | 2 |
| Diptera | Anopheles gambiae | 15,363 | 22.44 | 13 | 22 | 2 |
| Diptera | Anopheles quadrimaculatus A Orlando | 15,455 | 22.64 | 13 | 22 | 2 |
| Diptera | Bactrocera oleae Italy | 15,815 | 27.41 | 13 | 22 | 2 |
| Diptera | Bactrocera oleae Portugal | 15,815 | 27.37 | 13 | 22 | 2 |
| Diptera | Ceratitis capitata | 15,980 | 22.52 | 13 | 22 | 2 |
| Diptera | Chrysomya putoria | 15,837 | 23.30 | 13 | 23 | 2 |
| Diptera | Cochliomyia hominivorax | 16,022 | 23.10 | 13 | 22 | 2 |
| Diptera | Cydistomyia duplonotata | 16,247 | 22.07 | 13 | 23 | 2 |
| Diptera | Drosophila ananassae | 14,920 | 22.59 | 13 | 22 | 2 |
| Diptera | Drosophila erecta | 14,952 | 22.77 | 13 | 22 | 2 |
| Diptera | Drosophila grimshawi | 14,874 | 23.24 | 13 | 22 | 2 |
| Diptera | Drosophila mauritiana G52 | 14,964 | 22.29 | 13 | 22 | 2 |
| Diptera | Drosophila melanogaster | 19,517 | 17.84 | 13 | 22 | 2 |
| Diptera | Drosophila mojavensis | 14,904 | 23.54 | 13 | 22 | 2 |
| Diptera | Drosophila simulans KY007 | 14,946 | 22.33 | 13 | 22 | 2 |
| Diptera | Drosophila simulans KY045 | 14,946 | 22.36 | 13 | 22 | 2 |
| Diptera | Drosophila simulans KY201 | 14,946 | 22.32 | 13 | 22 | 2 |
| Diptera | Drosophila simulans KY215 | 14,946 | 22.33 | 13 | 22 | 2 |
| Diptera | Drosophila persimilis | 14,930 | 22.70 | 13 | 22 | 2 |
| Diptera | Drosophila virilis | 14,949 | 23.25 | 13 | 22 | 2 |
| Diptera | Drosophila willistoni | 14,915 | 22.76 | 13 | 22 | 2 |
| Diptera | Drosophila yakuba | 16,019 | 21.41 | 13 | 22 | 2 |
| Diptera | Simosyrphus grandicornis | 16,141 | 19.16 | 13 | 22 | 2 |
| Diptera | Stomoxys calcitrans* | 16,790 | 21.07 | 12 | 23 | 2 |
| Diptera | Trichophthalma punctata | 16,396 | 26.04 | 13 | 21 | 2 |
| Hymenoptera | Abispa ephippium | 16,953 | 19.39 | 13 | 26 | 2 |
| Hymenoptera | Apis mellifera | 16,343 | 15.14 | 13 | 22 | 2 |
| Hymenoptera | Bombus ignites | 16,434 | 13.22 | 13 | 22 | 2 |
| Hymenoptera | Vanhornia eucnemidarum* | 16,574 | 19.86 | 13 | 18 | 2 |
| Hymenoptera | Xenos vesparum* | 14,519 | 20.68 | 13 | 23 | 1 |
| Lepidoptera | Adoxophyes honmai | 15,680 | 19.61 | 13 | 22 | 2 |
| Lepidoptera | Bombyx mandarina | 15,928 | 18.32 | 13 | 22 | 2 |
| Lepidoptera | Bombyx mori C-108 | 15,656 | 18.64 | 13 | 22 | 2 |
| Lepidoptera | Coreana raphaelis | 15,314 | 17.34 | 13 | 23 | 2 |
| Lepidoptera | Manduca sexta | 15,516 | 18.21 | 13 | 23 | 2 |
| Lepidoptera | Ochrogaster lunifer | 15,593 | 22.16 | 13 | 22 | 2 |
| Lepidoptera | Ostrinia furnacalis* | 14,536 | 19.62 | 13 | 22 | 2 |
| Lepidoptera | Ostrinia nubilalis* | 14,535 | 19.84 | 13 | 22 | 2 |
| Lepidoptera | Saturnia boisduvalii | 15,360 | 19.38 | 13 | 22 | 2 |
| Megaloptera | Corydalus cornutus | 15,687 | 25.10 | 13 | 22 | 2 |
| Megaloptera | Protohermes concolorus | 15,851 | 24.17 | 13 | 22 | 2 |
| Neuroptera | Ascaloptynx appendiculatus | 15,877 | 24.43 | 13 | 22 | 2 |
| Neuroptera | Polystoechotes punctatus | 16,036 | 21.04 | 12 | 22 | 2 |
*Nearly completely sequenced mitochondrial genome.
Table 3.
List of 76 whole mitochondrial genomes of hexapod species (Part II. 73 species excluding holometabolous orders) archived in IMGD
| Order | Species | Size (bp) | GC (%) | PCGs | tRNAs | rRNAs |
| Archaeognatha | Nesomachilis australica | 15,474 | 31.17 | 13 | 21 | 2 |
| Archaeognatha | Pedetontus silvestrii | 15,879 | 25.66 | 13 | 22 | 2 |
| Archaeognatha | Petrobius brevistylis | 15,698 | 32.12 | 13 | 22 | 2 |
| Archaeognatha | Trigoniophthalmus alternatus | 16,197 | 28.59 | 13 | 22 | 2 |
| Zygentoma | Thermobia domestica | 15,152 | 33.01 | 13 | 22 | 2 |
| Zygentoma | Tricholepidion gertschi | 15,267 | 31.40 | 13 | 22 | 2 |
| Collembola | Cryptopygus antarcticus | 15,297 | 29.10 | 13 | 23 | 2 |
| Collembola | Gomphiocephalus hodgsoni | 15,075 | 25.92 | 13 | 22 | 2 |
| Collembola | Friesea grisea | 15,425 | 27.73 | 13 | 22 | 2 |
| Collembola | Onychiurus orientalis* | 12,984 | 30.89 | 13 | 20 | 1 |
| Collembola | Orchesella villosa | 14,924 | 27.82 | 13 | 22 | 2 |
| Collembola | Podura aquatica* | 13,809 | 34.21 | 13 | 20 | 1 |
| Collembola | Sminthurus viridis | 14,817 | 30.56 | 13 | 22 | 2 |
| Collembola | Tetrodontophora bielanensis | 15,455 | 27.32 | 13 | 22 | 2 |
| Diplura | Campodea fragilis | 14,965 | 27.44 | 13 | 22 | 2 |
| Diplura | Campodea lubbocki | 14,974 | 25.19 | 13 | 22 | 2 |
| Diplura | Japyx solifugus | 15,785 | 35.18 | 13 | 22 | 2 |
| Ephemeroptera | Parafronurus youi | 15,481 | 33.62 | 13 | 23 | 2 |
| Odonata | Orthetrum triangulare melania* | 14,033 | 26.09 | 13 | 19 | 2 |
| Grylloblattodea | Grylloblatta sculleni* | 15,595 | 29.71 | 12 | 19 | 2 |
| Blattaria | Periplaneta fuliginosa | 14,996 | 24.85 | 13 | 22 | 2 |
| Isoptera | Reticulitermes flavipes IS13 | 16,565 | 33.82 | 13 | 22 | 2 |
| Isoptera | Reticulitermes flavipes IS57 | 16,569 | 33.76 | 13 | 22 | 2 |
| Isoptera | Reticulitermes flavipes IS58 | 16,567 | 33.78 | 13 | 22 | 2 |
| Isoptera | Reticulitermes hageni | 16,590 | 34.45 | 13 | 22 | 2 |
| Isoptera | Reticulitermes santonensis IS54 | 16,567 | 33.91 | 13 | 22 | 2 |
| Isoptera | Reticulitermes virginicus IS59 | 16,513 | 34.12 | 13 | 22 | 2 |
| Isoptera | Reticulitermes virginicus IS60 | 15,966 | 34.37 | 13 | 22 | 2 |
| Mantodea | Tamolanica tamolana | 16,055 | 24.73 | 13 | 22 | 2 |
| Mantophasmatodea | Sclerophasma paresisense | 15,500 | 24.94 | 13 | 22 | 2 |
| Orthoptera | Acrida willemsei | 15,601 | 23.78 | 13 | 22 | 2 |
| Orthoptera | Anabrus simplex | 15,766 | 30.56 | 13 | 22 | 2 |
| Orthoptera | Calliptamus italicus | 15,675 | 26.74 | 13 | 22 | 2 |
| Orthoptera | Chorthippus chinensis | 15,599 | 24.89 | 13 | 22 | 2 |
| Orthoptera | Deracantha onos | 15,650 | 30.76 | 13 | 22 | 2 |
| Orthoptera | Gryllotalpa orientalis | 15,521 | 29.51 | 13 | 22 | 2 |
| Orthoptera | Gryllotalpa pluvialis | 15,525 | 27.80 | 13 | 22 | 2 |
| Orthoptera | Locusta migratoria | 15,722 | 24.67 | 13 | 22 | 2 |
| Orthoptera | Myrmecophilus manni | 15,323 | 29.82 | 13 | 22 | 2 |
| Orthoptera | Oxya chinensis | 15,443 | 24.11 | 13 | 22 | 2 |
| Orthoptera | Ruspolia dubia | 14,971 | 29.14 | 13 | 22 | 2 |
| Orthoptera | Gastrimargus marmoratus | 15,924 | 24.82 | 13 | 22 | 2 |
| Orthoptera | Gampsocleis gratiosa | 15,929 | 34.69 | 13 | 22 | 2 |
| Orthoptera | Troglophilus neglectus | 15,810 | 26.63 | 13 | 23 | 2 |
| Phasmatodea | Timema californicum* | 14,387 | 27.86 | 13 | 19 | 1 |
| Plecoptera | Pteronarcys princeps | 16,004 | 28.54 | 13 | 22 | 2 |
| Hemiptera | Aeschyntelus notatus* | 14,532 | 24.29 | 13 | 22 | 2 |
| Hemiptera | Aleurochiton aceris | 15,388 | 22.10 | 13 | 21 | 2 |
| Hemiptera | Aleurodicus dugesii | 15,723 | 13.67 | 13 | 20 | 2 |
| Hemiptera | Bemisia tabaci | 15,322 | 24.32 | 13 | 22 | 2 |
| Hemiptera | Coptosoma bifaria | 16,179 | 28.67 | 13 | 22 | 2 |
| Hemiptera | Dysdercus cingulatus | 16,249 | 22.31 | 13 | 22 | 2 |
| Hemiptera | Geocoris pallidipennis* | 14,592 | 24.14 | 13 | 22 | 2 |
| Hemiptera | Hydaropsis longirostris | 16,521 | 24.54 | 13 | 22 | 2 |
| Hemiptera | Macroscytus subaeneus* | 14,620 | 26.21 | 13 | 22 | 2 |
| Hemiptera | Malcus inconspicuus | 15,575 | 22.20 | 13 | 22 | 2 |
| Hemiptera | Neomaskellia andropogonis | 14,496 | 18.73 | 13 | 18 | 2 |
| Hemiptera | Neuroctenus parus | 15,354 | 31.14 | 13 | 22 | 2 |
| Hemiptera | Nezara viridula | 16,889 | 23.12 | 13 | 22 | 2 |
| Hemiptera | Orius niger* | 14,494 | 23.47 | 13 | 22 | 2 |
| Hemiptera | Pachypsylla venusta | 14,711 | 25.00 | 13 | 22 | 2 |
| Hemiptera | Phaenacantha marcida* | 14,540 | 26.54 | 13 | 22 | 2 |
| Hemiptera | Philaenus spumarius | 16,324 | 23.01 | 13 | 22 | 2 |
| Hemiptera | Physopelta gutta | 14,935 | 25.49 | 13 | 22 | 2 |
| Hemiptera | Riptortus pedestris | 17,191 | 23.41 | 13 | 22 | 2 |
| Hemiptera | Saldula arsenjevi | 15,324 | 25.39 | 13 | 22 | 2 |
| Hemiptera | Schizaphis graminum | 15,721 | 16.06 | 13 | 22 | 2 |
| Hemiptera | Tetraleurodes acacia | 15,080 | 28.02 | 13 | 19 | 2 |
| Hemiptera | Trialeurodes vaporariorum | 18,414 | 27.70 | 13 | 22 | 2 |
| Hemiptera | Triatoma dimidiate | 17,019 | 30.43 | 13 | 22 | 2 |
| Hemiptera | Yemmalysus parallelus | 15,747 | 22.82 | 13 | 22 | 2 |
| Phthiraptera | Bothriometopus macrocnemis | 15,564 | 29.20 | 13 | 25 | 2 |
| Phthiraptera | Campanulotes bidentatus | 14,804 | 29.88 | 13 | 22 | 2 |
| Phthiraptera | Heterodoxus macropus | 14,670 | 20.72 | 13 | 22 | 2 |
| Psocoptera | Lepidopsocid sp. RS2001 | 16,924 | 20.98 | 13 | 22 | 2 |
| Thysanoptera | Thrips imaginis | 15,407 | 23.43 | 13 | 23 | 2 |
*Nearly completely sequenced mitochondrial genome.
Construction and content
System architecture and design
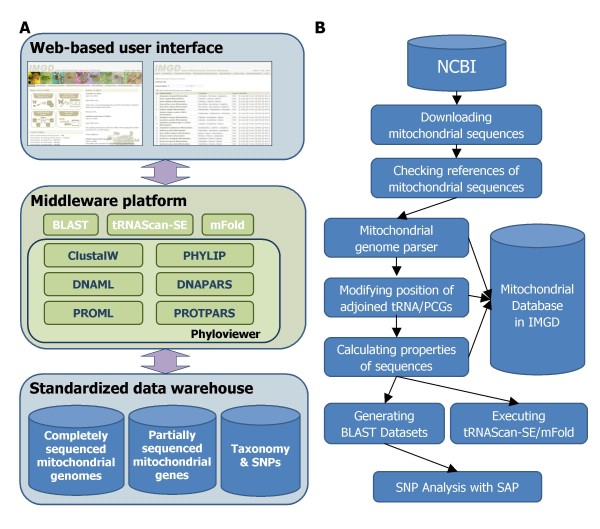
The IMGD consists of three integrated layers: i) a standardized data warehouse that is supported by CFGP [24], ii) the middleware platform for the integration of various bioinformatics programs via standardized input and output interfaces, and iii) the web-based user interface, including the Species-driven User Interface (Figure 1A). In order to support the efficient archiving and analysis of a very large number of heterogeneous mitochondrial gene sequences (Table 2 and Table 3), a standardized structure for sequence data was required: this requirement was solved using CFGP [24], which has demonstrated its reliability and expandability via several published databases [32-37].
Figure 1.
The system architecture and pipeline of IMGD. (A) Each rectangular box shows three layers. In the standardized data warehouse, diverse databases are placed. The middleware platform manages not only BLAST, tRNAScan-SE, and mFold but also six phylogenetic tools managed by Phyloviewer (http://www.phyloviewer.org/). The web-based user interface supports browsing all information deposited in IMGD. (B) The pipeline for archiving hexapod mitochondrial sequences and calculating their properties was presented as a flowchart diagram.
To support phylogenetic studies using the archived hexapod mitochondrial sequences, ClustalW (Version 1.83), PHYLIP (Version 3.68), and PHYML (Version 3.0) [29-31], which support the Neighbour Joining (NJ), Maximum Parsimony (MP), and Maximum Likelihood (ML) methods, respectively, were incorporated. The visualization and management of the resulting phylogenetic data are supported by the Phyloviewer (http://www.phyloviewer.org/), which has been successfully employed in other platforms [24,34,35]. BLAST [26] was integrated with datasets containing mitochondrial gene sequences and hexapod taxonomy information, and tRNAscan-SE (Version 1.23) [27] and mFold (Version 3.2) [28] were embedded to allow for the display and comparison of secondary structures of tRNAs and anticodon sequences.
The user interface of IMGD provides the Mitochondrial Genome Browser, which is founded on the SNU Genome Browser (http://genomebrowser.snu.ac.kr/) [25], to support the browsing and comparison of mitochondrial genome sequences in both the text and graphical forms via an interactive interface, and the Partial Sequence Browser to allow for the browsing of partially sequenced mitochondrial sequences. The IMGD also provides the Object Browser, which can collect and move selected sequences in IMGD into the Favorite, a personalized virtual storage space, for further data analyses using the analysis tools in both IMGD and CFGP [24]. The IMGD archives sequences and taxonomical information from more than 25,000 hexapod species. To facilitate the organization and presentation of data according to the taxonomic position/grouping of the species of origin, a new interface named the Species-driven User Interface (SUI) was designed and implemented in IMGD.
Pipeline for updating the IMGD data warehouse
To support periodic updating of the IMGD data warehouse, the following automatic analysis pipeline was developed (Figure 1B). In the first step, completely and partially sequenced mitochondrial genome sequences are downloaded from NCBI using proper keywords after filtering out unpublished sequences. The downloaded sequences are subsequently filtered using several stop words in order to remove non-mitochondrial sequences. Secondly, the mitochondrial genome parsers, which were written in PERL, parse and store the filtered data into the data warehouse. Thirdly, adjoined stop codons at the 3'-end of the PCGs that overlap with neighboring PCGs or tRNAs in the mitochondrial genome [38], are manually checked to determine whether they are correct or not. Lastly, certain properties of the genome, including the CG content, AT skew, and codon usage, are calculated for graphical representations via SNU Genome Browser, and various cache tables are updated. In the final step, BLAST datasets, tRNA annotation information via both tRNAScan-SE [27] and mFold [28], and SNP databases are updated.
Taxonomic origins of the sequences data archived in IMGD
The IMGD archives 132 hexapod mitochondrial genomes and 113,985 GenBank accessions of partially sequenced mitochondrial genes, consisting of 102,430 PCGs, 19,452 rRNAs, and 17,944 tRNAs, from 25,747 species belonging to 33 orders (Table 1). More than 10,000 mitochondrial gene sequences were derived from >1,000 species in the orders Coleoptera, Lepidoptera, Hymenoptera, and Diptera. In particular, members of Diptera and Coleoptera account for 26 (20.00%) and 12 (9.23%) mitochondrial genomes, respectively, reflecting active researches on these orders [39,40]. In contrast, the following 13 orders (39.39%) are represented only by less than 50 species in total: Dermaptera, Siphonaptera, Zygentoma, Grylloblattodea, Mantophasmatodea, Embioptera, Diplura, Archaeognatha, Strepsiptera, Megaloptera, Raphidioptera, Protura, and Zoraptera (Table 1). The underrepresentation of mitochondrial gene sequences from many orders suggests that to adequately support the analysis of evolutionary relationships within the Hexapoda, these underrepresented orders require more attention.
Notable features in hexapod mitochondrial genomes
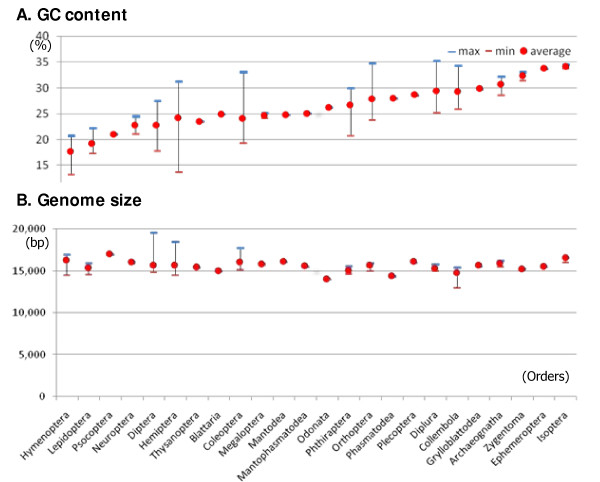
The genome size, GC content, and the number of PCGs, tRNAs, and rRNAs of the 132 mitochondrial genomes archived in IMGD (Table 2 and Table 3) were assessed (Figure 2). The GC content ranges from 13.22% to 35.18% with an average of 25.09%, showing the association at the order level (Figure 2A). The genome sizes vary from 12,984 bp to 19,517 bp, with an average of 15,617 bp with no clear correlation at any taxon levels (Figure 2B). Analyses of gene order in the 112 completely sequenced mitochondrial genomes revealed several interesting features. In 42 genomes (37.50%), which represent 12 orders, at least 222 gene insertions, deletions, inversions, and translocations were identified relative to the gene arrangement of the ancestral insect Drosophila yakuba [3,41] (Lee et al., in preparation). Gene translocations and inversions were detected in the following 12 orders: Collembola, Archaeognatha, Zygentoma, Hemiptera, Thysanoptera, Psocoptera, Phthiraptera, Neuroptera, Hymenoptera, Orthoptera, Lepidoptera, and Diptera. Gene insertions and deletions were detected in Collembola, Ephemeroptera, Orthoptera, Hemiptera, Phthiraptera, Diptera, and Lepidoptera.
Figure 2.
Estimates of the GC content and genome size of the 132 hexapod mitochondrial genomes. (A) The ranges of the GC content in the nearly completely and the completely sequenced mitochondrial genomes are shown. The closed red circle indicates the average GC content, and the blue and red bars present the maximum and minimum GC contents, respectively. (B) The distribution of mitochondrial genome sizes in different hexapod orders is shown. The closed red circle indicates the average mitochondrial genome size and the blue and red bars present the largest and smallest genome sizes, respectively (see also Table 2 and Table 3).
Examples of phylogenetic analyses results using insectmitochondrial genomes
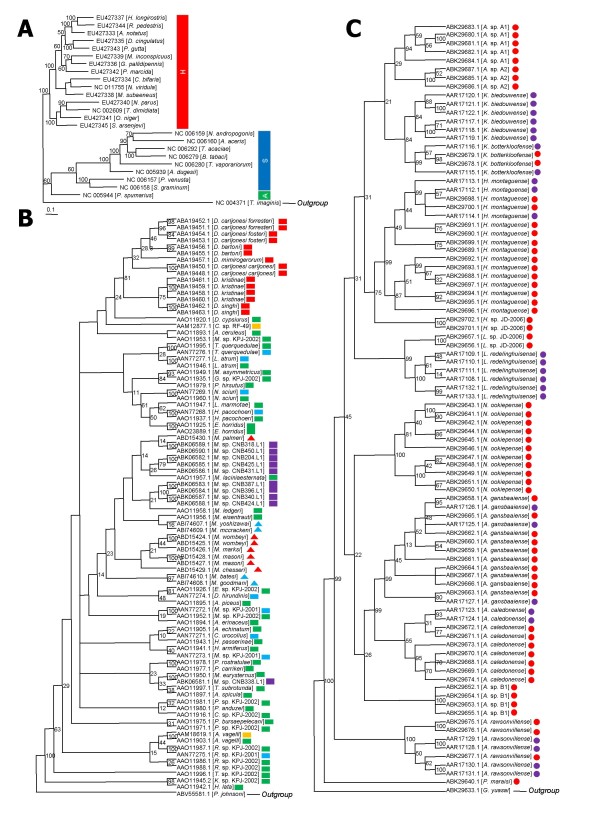
To demonstrate the utility of IMGD for phylogenetic analysis and also to test the system, many phylogenetic analyses using the data archived in IMGD have been conducted (e.g., Figure 3). Figure 3A shows an ML phylogenetic tree based on 19 completely sequenced and 5 nearly completed mitochondrial genomes in the order Hemiptera, which clearly shows two major suborder clades (Sternorrhyncha + Auchenorrhyncha and Heteroptera). The MP trees based on the COI gene sequences (Figure 3B and 3C) revealed more comprehensive phylogenetic relationships than those derived from previous studies in the orders Phthiraptera [42-44] and Mantophasmatodea [45,46].
Figure 3.
Examples of phylogenetic analyses conducted using data and tools in IMGD. (A) ML tree of the 24 Hemipteran species (19 completely and 5 nearly completed mitochondrial genomes) with Thrips imaginis (Thysanoptera) as an outgroup was constructed using DNAML. S, Sternorrhyncha; A, Auchenorrhyncha; H, Heteroptera. (B) MP tree built based on 88 COI sequences from 70 Phthirapteran species using DNAPARS, is shown. Ptycta johnsoni (Psocoptera) was used as an outgroup. The blue square indicates the sequences originated from Johnson and Whiting (2002) [42]; green square, Johnson et al. (2003) [43]; blue triangle, Price and Johnson (2006) [44]; red, violet and yellow squares, and red triangle present unpublished mitochondrial gene sequences. (C) MP tree using 90 COI sequences from 14 Mantophasmatodean species, with Galloisiana yuasai (Grylloblattodea) as an outgroup, was drawn using DNAPARS. The red circle indicates the mitochondrial sequences reported by Damgaard et al. (2008) [46] and violet circle presents the sequences from the study of Klass et al. (2003) [45]. The numbers on individual nodes of the trees in A, B, and C indicate bootstrap values with 10, 100, and 100 repeats, respectively, and the names of the species used and NCBI accession numbers are shown at the end of individual branches.
Single Nucleotide Polymorphisms among 9 insect mitochondrial genomes
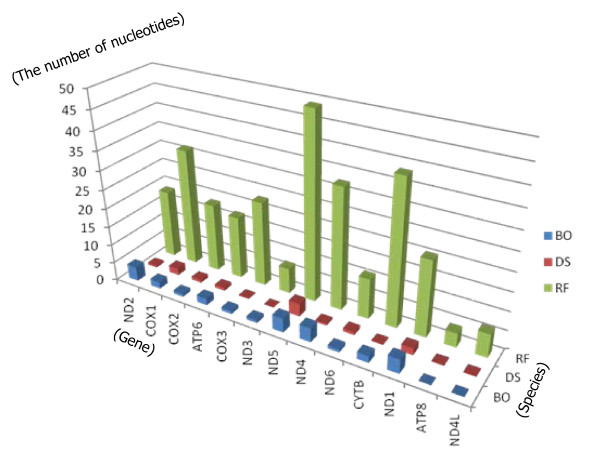
Single nucleotide polymorphisms (SNPs) in eight species with more than one mitochondrial genome having been sequenced (Table 4), were analyzed via the SNP Analysis Platform (http://sap.snu.ac.kr/), which is based on BLAST. A total of 856 SNPs and 30 insertion and deletions (INDELs) were identified (Table 4) from 187 kbp of aligned mitochondrial genome sequences (6 pair-wise comparisons of mitochondrial genomes). Among these, 621 SNPs (72.55%) were identified in 13 PCGs and designated as cSNPs. Figure 4 shows the average number of cSNPs in each species, order and PCG. Bactrocera oleae (BO), Drosophila simulans (DS), and Reticulitermes flavipes (RF) exhibited the highest frequency of cSNPs, similar to the results from previous genome sequence analyses [47-49]. Among the 13 PCGs, the COI, NADH dehydrogenase subunit 4 (ND4), and/or NADH dehydrogenase subunit 5 (ND5) genes showed the highest frequency of SNPs in Diptera (COI and ND5) and Isoptera (cytochrome b, ND4, and ND5) (Figure 4). These regions can serve as potential molecular markers in population genetic studies of these three orders.
Table 4.
List of mitochondrial genome comparisons for SNP analysis
| Order | Source/Target Species | Size (bp) | Aligned (bp) | SNPs | INDELs |
| Diptera | Bactrocera oleae Italy vs | 15,815 | 15,815 | 31 | 0 |
| Bactrocera oleae portugal | 15,815 | 15,815 | |||
| Drosophila simulans KY007 | 14,946 | 14,946 | 25 | 2 | |
| Drosophila simulans KY045 | 14,946 | 14,946 | |||
| Drosophila simulans KY007 | 14,946 | 14,946 | 17 | 2 | |
| Drosophila simulans KY201 | 14,946 | 14,946 | |||
| Drosophila simulans KY007 | 14,946 | 14,946 | 6 | 0 | |
| Drosophila simulans KY215 | 14,946 | 14,946 | |||
| Isoptera | Reticulitermes flavipes IS13 vs | 16,565 | 16,561 | 393 | 14 |
| Reticulitermes flavipes IS57 | 16,569 | 16,565 | |||
| Reticulitermes flavipes IS13 vs | 16,565 | 16,561 | 384 | 12 | |
| Reticulitermes flavipes IS58 | 16,567 | 16,563 | |||
| Total | 6 pair-wise comparisons | 187,572 | 187,556 | 856 | 30 |
Figure 4.
Distribution of SNPs in 13 PCGs in 9 mitochondrial genomes. The bar graph displays the distribution of SNPs in 13 PCGs of three insect species: BO, Bactrocera oleae; DS, Drosophila simulans; RF, Reticulitermes flavipes. ATP6 and 8 (ATP synthase subunit 6 and 8); COX1–3 (cytochrome c oxidase subunits I–III); CYTB (cytochrome b); ND1–6 (NADH dehydrogenase subunits 1–6); ND4L (NADH dehydrogenase subunit 4L) (see also Table 4).
Utilities and discussion
Species-driven User Interface (SUI)
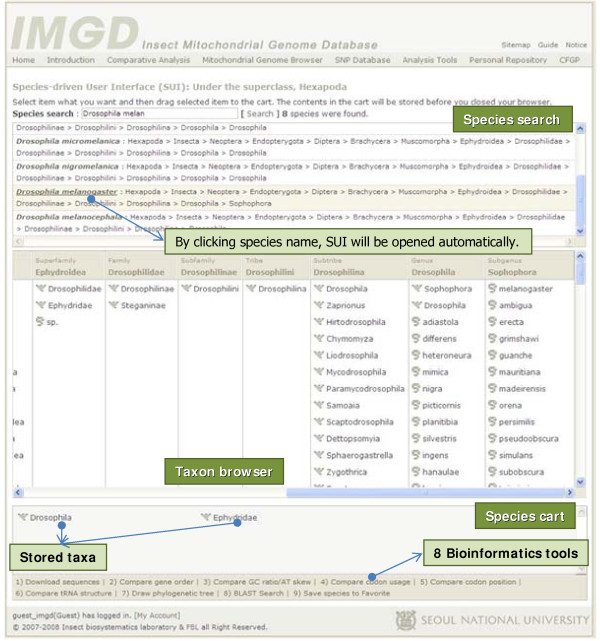
The SUI of IMGD supports efficient data retrieval and analysis at multi-taxon levels. The SUI was developed using Ajax technology, which supports faster performance than other methods (e.g. JavaScript and Java applets). The SUI helps the users of IMGD search hexapod taxa using the 'Species search' and supports the addition and deletion of selected insect species via the 'Species cart' function, which is similar to the cart functions commonly used on online shopping sites (Figure 3). After placing the taxa of interest in the cart, they can be analyzed in the following ways: i) downloading nucleotide and protein sequences and/or storing them into the Favorite with various options, ii) comparing gene orders, GC content/AT skew, codon usage and position among mitochondrial genomes, iii) displaying tRNA secondary structures predicted by tRNAScan-SE [27] and mFold [28], iv) executing ClustalW for multiple sequence alignment and calculating phylogenetic trees based on three methods, including NJ, MP, and ML, with a bootstrapping option, v) executing a BLAST search against the selected taxa, and vi) saving species information into the Favorite for future analyses (Figure 5). Since SUI was designed using a standardized application programming interface (API), additional programs can be easily incorporated into SUI.
Figure 5.
Species-driven User Interface (SUI) optimized for collecting data based on taxa. The Species-driven User Interface (SUI) consists of three parts: i) Species search, ii) Taxon browser, and iii) Species cart. The Species search function supports the search of species by species name. The Taxon browser provides the interface for browsing taxa in a hierarchical manner. The Species cart can store selected taxa, bridging the data from them to nine bioinformatics tools.
Gene order browser for graphical presentation of elements on the mitochondrial genome
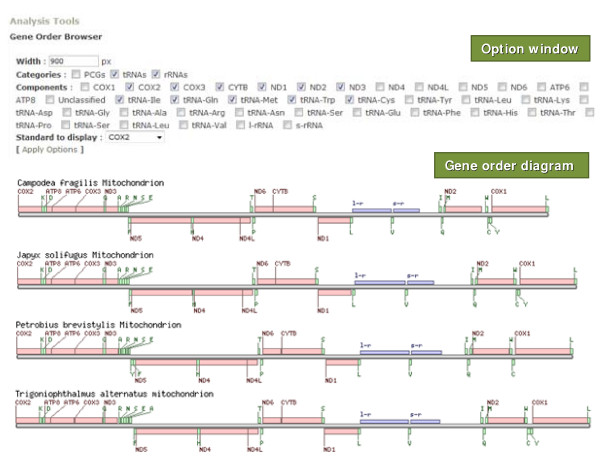
Gene rearrangement events in the mitochondrial genomes can be used for tracing the evolutionary history of the mitochondrial genomes in Hexapoda (Lee et al., in preparation). The gene order browser implemented in IMGD was designed for efficient graphical presentation of PCGs, tRNAs, and rRNAs in the mitochondrial genome. To display different features on the genome graphically, the browser uses three different colors for PCGs, tRNAs, and rRNAs, and presents names of individual units (Figure 6). Moreover, the gene order browser displays the gene organizations using a specific gene as the start site for the linear genome diagrams regardless of the arbitrary start position given to individual mitochondrial genomes. Users can choose the number of mitochondrial genomes to be displayed by selecting them via SUI.
Figure 6.
Gene order browser for graphical presentation of the mitochondrial gene order. The gene order browser consists of two parts: one is the option window and the other is the gene order diagram. In the option window, three options, including width, categories, and components, are displayed. After clicking 'Apply Options,' a gene order diagram based on the chosen option will be displayed. To indicate the nature of specific genetic elements on displayed mitochondrial genomes, the following abbreviations were used: A, tRNA-Ala;C, tRNA-Cys; D, tRNA-Asp; E, tRNA-Glu; F, tRNA-Phe; G, tRNA-Gly; H, tRNA-His; I, tRNA-Ile; K, tRNA-Lys; L, tRNA-Leu; M, tRNA-Met; N, tRNA-Asn; P, tRNA-Pro; Q, tRNA-Gln; R, tRNA-Arg; S, tRNA-Ser; T, tRNA-Thr; V, tRNA-Val; W, tRNA-Trp; Y, tRNA-Tyr; COX1–3, cytochrome c oxidase subunits I–III; CYTB, cytochrome b; ATP6 and ATP8, subunits 6 and 8 of the F0ATPase; ND1–6 and nad4L, NADH dehydrogenase subunits 1–6 and 4L; l-r and s-r, large and small subunit of ribosomal RNA genes; PCGs, protein coding genes; rRNAs, ribosomal RNA genes; tRNAs, transfer RNA genes.
Integrated platform for phylogenetic analyses supported by Phyloviewer
The Phyloviewer (http://www.phyloviewer.org/) provides four phylogenetic analysis programs (ClustalW, DNAPARS/PROTPARS, DNAML/PROML, and PHYML [29-31]) via a common interface to support phylogenetic studies based on the mitochondrial gene/genome sequences archived in IMGD. Three different methods of drawing phylogenetic trees (NJ, MP, and ML) are currently available. In addition, the interactive capability of graphical presentation of sequence alignments and selecting and storing all sequences under a selected node in the resulting phylogenetic tree by clicking the node is also provided.
Comparative mitochondrial genomics via the SNU Genome Browser
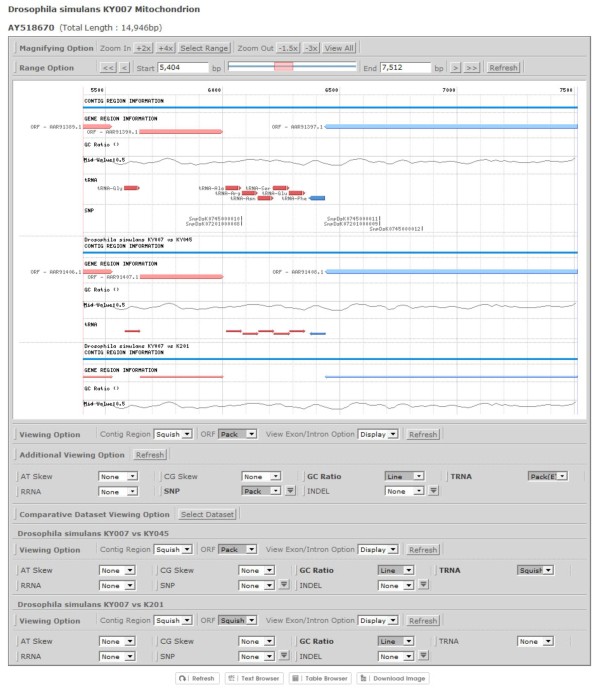
To support intuitive visualization of sequences, SNPs, and INDELs between two mitochondrial genomes, the SNU Genome Browser (http://genomebrowser.snu.ac.kr/)[25] was implemented. This recently developed genome browser provides an interactive user interface that supports visualization of the alignment region between genomes with the capability of comparing multiple genomes simultaneously (Figure 7). It also supports the text browser function for displaying nucleotide sequences of a selected region, supporting the confirmation of SNPs and INDELs. The table browser provides a list of individual elements present in the selected region with their positional information in a tabular form.
Figure 7.
Interactive graphical interface for visualizing aligned mitochondrial genomes via the SNU Genome Browser. The SNU Genome Browser displays SNPs/INDELs as well as PCGs, tRNAs, GC contents among the aligned genomes of Drosophila simulans KY007, KY045, and K201 strains.
Favorite, a personalized virtual space for storing data and conducting further analysis
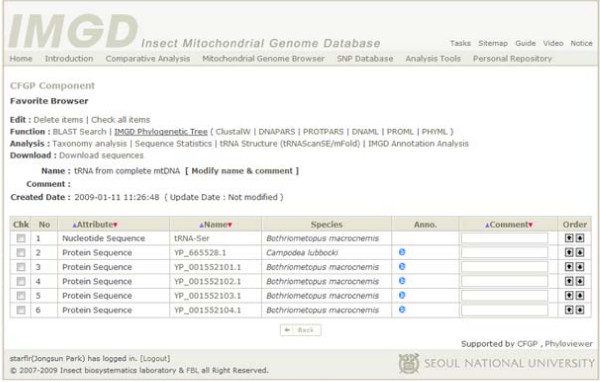
Most of the data analysis and/or retrieval interfaces in IMGD provide the Object Browser, which allows users to save all or selected sequences and/or species shown into Favorite. This will help users manage their own datasets via IMGD. Through the interface of Favorite, BLAST, six different phylogenetics programs, and four data analysis tools are available for further analyses (Figure 8). The Favorite is linked to CFGP (http://cfgp.snu.ac.kr/), which provides not only diverse bioinformatic tools but also a data warehouse containing complete sequences of 19 insect nuclear genomes [24], so that further analyses with diverse resources can be conducted easily via this web interface.
Figure 8.
Favorite, a personalized virtual space for data storage and further analyses. The browser in Favorite provides four options: 'Edit,' 'Function,' 'Analysis,' and 'Download.' Any sequences listed at the bottom part can be selected by users for analyzing the selected sequences using seven programs and four analysis tools via the web.
Conclusion
We developed IMGD to support versatile comparative analyses of hexapod mitochondrial gene/genome sequences. In IMGD, 132 completely or nearly-completely sequenced mitochondrial genomes and 113,985 mitochondrial gene sequences from 25,747 species were archived. The IMGD provides a variety of phylogenetic analysis tools via Phyloviewer, which supports the interactive graphical presentation of resultant phylogenetic trees. The IMGD, supported by the SNP analysis platform and the SNU Genome Browser, provides a graphical view of mitochondrial genome comparisons. In the near future, additional analysis tools, such as PAML [50] for the determination of positive/negative selection based on dS/dN values, will be integrated into IMGD. Moreover, based on the database of widely sequenced mitochondrial genes, an insect species identification system based on multiple loci can be developed. The IMGD is expected to significantly enhance evolutionary studies on the superclass Hexapoda using rapidly accumulating insect mitochondrial genome sequences.
Availability and requirements
All data described in this paper can be browsed and downloaded through the IMGD web site at http://www.imgd.org/.
Authors' contributions
WL, JP, SL, and YHL designed and managed this project, JC, JP, and WL construct the IMGD sequence database and WL, JL, JC, and JP curated all sequences in IMGD. KJ designed the IMGD web site and developed SUI. BP developed the interface for Phyloviewer, JP integrated SAP to IMGD and DH conducted mitochondrial genome alignments. JP, JC, BP, KJ, KA, DH and WS constructed and tested the whole web pages. WL, JP and SK wrote the manuscript.
Acknowledgments
Acknowledgements
This research was supported by Korea Ministry of Environment as "The Eco-technopia 21 project", a grant from by Agricultural R&D Promotion Center, and a grant from Biogreen21 Project (20070301034032) to S.L. It was also supported by a grant from Crop Functional Genomics Center (CG1141), Korean Research Foundation Grant (KRF-2006-005-J04701), Biogreen21 Project (20080401034044) funded by Rural Development Administration and the Korea Science and Engineering Foundation (KOSEF) grant funded by the Korea government (MEST) (R11-2008-062-03001-0) to Y.H.L. J.P. acknowledges a graduate fellowship provided by the Ministry of Education through the Brain Korea 21 Project.
Contributor Information
Wonhoon Lee, Email: won9@snu.ac.kr.
Jongsun Park, Email: starflr@snu.ac.kr.
Jaeyoung Choi, Email: amethyst1016@gmail.com.
Kyongyong Jung, Email: lulupon0@snu.ac.kr.
Bongsoo Park, Email: pbsfree@gmail.com.
Donghan Kim, Email: hoppang1234@gmail.com.
Jaeyoung Lee, Email: toughcd1@snu.ac.kr.
Kyohun Ahn, Email: dksrygns00@snu.ac.kr.
Wonho Song, Email: james1201@snu.ac.kr.
Seogchan Kang, Email: sxk55@psu.edu.
Yong-Hwan Lee, Email: yonglee@snu.ac.kr.
Seunghwan Lee, Email: seung@snu.ac.kr.
References
- Rawlings TA, Collins TM, Bieler R. A major mitochondrial gene rearrangement among closely related species. Mol Biol Evol. 2001;18:1604–1609. doi: 10.1093/oxfordjournals.molbev.a003949. [DOI] [PubMed] [Google Scholar]
- Dowton M. Relationships among the cyclostome braconid (Hymenoptera: Braconidae) subfamilies inferred from a mitochondrial tRNA gene rearrangement. Mol Phylogenet Evol. 1999;11:283–287. doi: 10.1006/mpev.1998.0580. [DOI] [PubMed] [Google Scholar]
- Boore JL. Animal mitochondrial genomes. Nucleic Acids Res. 1999;27:1767–1780. doi: 10.1093/nar/27.8.1767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boore JL, Brown WM. Big trees from little genomes: mitochondrial gene order as a phylogenetic tool. Curr Opin Genet Dev. 1998;8:668–674. doi: 10.1016/S0959-437X(98)80035-X. [DOI] [PubMed] [Google Scholar]
- Boore JL, Lavrov DV, Brown WM. Gene translocation links insects and crustaceans. Nature. 1998;392:667–668. doi: 10.1038/33577. [DOI] [PubMed] [Google Scholar]
- Ellegren H. Sequencing goes 454 and takes large-scale genomics into the wild. Mol Ecol. 2008;17:1629–1631. doi: 10.1111/j.1365-294X.2008.03699.x. [DOI] [PubMed] [Google Scholar]
- Jameson D, Gibson AP, Hudelot C, Higgs PG. OGRe: a relational database for comparative analysis of mitochondrial genomes. Nucleic Acids Res. 2003;31:202–206. doi: 10.1093/nar/gkg077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee YS, Oh J, Kim YU, Kim N, Yang S, Hwang UW. Mitome: dynamic and interactive database for comparative mitochondrial genomics in metazoan animals. Nucleic Acids Res. 2008;36:D938–942. doi: 10.1093/nar/gkm763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Brien EA, Zhang Y, Yang L, Wang E, Marie V, Lang BF, Burger G. GOBASE–a database of organelle and bacterial genome information. Nucleic Acids Res. 2006;34:D697–699. doi: 10.1093/nar/gkj098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feijao PC, Neiva LS, de Azeredo-Espin AM, Lessinger AC. AMiGA: the arthropodan mitochondrial genomes accessible database. Bioinformatics. 2006;22:902–903. doi: 10.1093/bioinformatics/btl021. [DOI] [PubMed] [Google Scholar]
- Vasconcelos AT, Guimaraes AC, Castelletti CH, Caruso CS, Ribeiro C, Yokaichiya F, Armoa GR, Pereira Gda S, da Silva IT, Schrago CG, Fernandes AL, da Silveira AR, Carneiro AG, Carvalho BM, Viana CJ, Gramkow D, Lima FJ, Correa LG, Mudado Mde A, Nehab-Hess P, Souza R, Correa RL, Russo CA. MamMiBase: a mitochondrial genome database for mammalian phylogenetic studies. Bioinformatics. 2005;21:2566–2567. doi: 10.1093/bioinformatics/bti326. [DOI] [PubMed] [Google Scholar]
- Wolfsberg TG, Schafer S, Tatusov RL, Tatusov TA. Organelle genome resource at NCBI. Trends Biochem Sci. 2001;26:199–203. doi: 10.1016/S0968-0004(00)01773-4. [DOI] [PubMed] [Google Scholar]
- Boykin LM, Shatters RG, Jr, Rosell RC, McKenzie CL, Bagnall RA, De Barro P, Frohlich DR. Global relationships of Bemisia tabaci (Hemiptera: Aleyrodidae) revealed using Bayesian analysis of mitochondrial COI DNA sequences. Mol Phylogenet Evol. 2007;44:1306–1319. doi: 10.1016/j.ympev.2007.04.020. [DOI] [PubMed] [Google Scholar]
- Morlais I, Severson DW. Complete mitochondrial DNA sequence and amino acid analysis of the cytochrome C oxidase subunit I (COI) from Aedes aegypti. DNA Seq. 2002;13:123–127. doi: 10.1080/10425170290030051. [DOI] [PubMed] [Google Scholar]
- Waeschenbach A, Telford MJ, Porter JS, Littlewood DT. The complete mitochondrial genome of Flustrellidra hispida and the phylogenetic position of Bryozoa among the Metazoa. Mol Phylogenet Evol. 2006;40:195–207. doi: 10.1016/j.ympev.2006.03.007. [DOI] [PubMed] [Google Scholar]
- Kress WJ, Erickson DL. DNA barcodes: genes, genomics, and bioinformatics. Proc Natl Acad Sci USA. 2008;105:2761–2762. doi: 10.1073/pnas.0800476105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ratasingham S, Hebert PDN. BOLD: The Barcode of Life Data System http://www.barcodinglife.org. Mol Ecol Notes. 2007;7:355–364. doi: 10.1111/j.1471-8286.2007.01678.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajibabaei M, Janzen DH, Burns JM, Hallwachs W, Hebert PD. DNA barcodes distinguish species of tropical Lepidoptera. Proc Natl Acad Sci USA. 2006;103:968–971. doi: 10.1073/pnas.0510466103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stahls G, Savolainen E. MtDNA COI barcodes reveal cryptic diversity in the Baetis vernus group (Ephemeroptera, Baetidae) Mol Phylogenet Evol. 2008;46:82–87. doi: 10.1016/j.ympev.2007.09.009. [DOI] [PubMed] [Google Scholar]
- Pfenninger M, Nowak C, Kley C, Steinke D, Streit B. Utility of DNA taxonomy and barcoding for the inference of larval community structure in morphologically cryptic Chironomus (Diptera) species. Mol Ecol. 2007;16:1957–1968. doi: 10.1111/j.1365-294X.2006.03136.x. [DOI] [PubMed] [Google Scholar]
- Nelson LA, Wallman JF, Dowton M. Using COI barcodes to identify forensically and medically important blowflies. Med Vet Entomol. 2007;21:44–52. doi: 10.1111/j.1365-2915.2007.00664.x. [DOI] [PubMed] [Google Scholar]
- Roe AD, Sperling FA. Patterns of evolution of mitochondrial cytochrome c oxidase I and II DNA and implications for DNA barcoding. Mol Phylogenet Evol. 2007;44:325–345. doi: 10.1016/j.ympev.2006.12.005. [DOI] [PubMed] [Google Scholar]
- Caterino MS, Cho S, Sperling FA. The current state of insect molecular systematics: a thriving Tower of Babel. Annu Rev Entomol. 2000;45:1–54. doi: 10.1146/annurev.ento.45.1.1. [DOI] [PubMed] [Google Scholar]
- Park J, Park B, Jung K, Jang S, Yu K, Choi J, Kong S, Park J, Kim S, Kim H, Kim S, Kim J, Blair J, Lee K, Kang S, Lee Y-H- CFGP: A Web-based, Comparative Fungal Genomics Platform. Nucleic Acids Res. 2008;36:D562–D571. doi: 10.1093/nar/gkm758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung K, Park J, Choi J, Park B, Kim S, Ahn K, Choi J, Choi D, Kang S, Lee Y-H. SNUGB: a versatile genome browser supporting comparative and functional fungal genomics. BMC Genomics. 2008;9:585. doi: 10.1186/1471-2164-9-586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGinnis S, Madden TL. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res. 2004;32:W20–25. doi: 10.1093/nar/gkh435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25:955–964. doi: 10.1093/nar/25.5.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003;31:3406–3415. doi: 10.1093/nar/gkg595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Felsenstein J. PHYLIP (PHYLogeny Inference Package) 3.6a2. Seattle: Distributed by the author, Department of Genetics, University of Washington, Seattle, WA; 1993. [Google Scholar]
- Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guindon S, Lethiec F, Duroux P, Gascuel O. PHYML Online–a web server for fast maximum likelihood-based phylogenetic inference. Nucleic Acids Res. 2005;33:W557–559. doi: 10.1093/nar/gki352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi J, Park J, Jeon J, Chi MH, Goh J, Yoo SY, Park J, Jung K, Kim H, Park SY, Rho HS, Kim S, Kim BR, Han SS, Kang S, Lee YH. Genome-wide analysis of T-DNA integration into the chromosomes of Magnaporthe oryzae. Molecular Microbiology. 2007;66:371–382. doi: 10.1111/j.1365-2958.2007.05918.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeon J, Park SY, Chi MH, Choi J, Park J, Rho HS, Kim S, Goh J, Yoo S, Choi J, Park JY, Yi M, Yang S, Kwon MJ, Han SS, Kim BR, Khang CH, Park B, Lim SE, Jung K, Kong S, Karunakaran M, Oh HS, Kim H, Kim S, Park J, Kang S, Choi WB, Kang S, Lee YH. Genome-wide functional analysis of pathogenicity genes in the rice blast fungus. Nat Genet. 2007;39:561–565. doi: 10.1038/ng2002. [DOI] [PubMed] [Google Scholar]
- Park J, Park B, Veeraraghavan N, Jung K, Lee YH, Blair J, Geiser DM, Isard S, Mansfield MA, Nikolaeva E, Park SY, Russo J, Kim SH, Greene M, Ivors KL, Balci Y, Peiman M, Erwin DC, Coffey MD, Rossman A, Farr D, Cline E, Crünwald NJ, Luster DG, Schrandt J, Martin F, Ribeiro OK, Makalowska I, Kang S. Phytophthora Database: A Forensic Database Supporting the Identification and Monitoring of Phytophthora. Plant Dis. 2008;92:966–972. doi: 10.1094/PDIS-92-6-0966. [DOI] [PubMed] [Google Scholar]
- Park J, Park J, Jang S, Kim S, Kong S, Choi J, Ahn K, Kim J, Lee S, Kim S, Park B, Jung K, Kim S, Kang S, Lee YH. FTFD: An Informatics Pipeline Supporting Phylogenomic Analysis of Fungal Transcription Factors. Bioinformatics. 2008;24:1024–1025. doi: 10.1093/bioinformatics/btn058. [DOI] [PubMed] [Google Scholar]
- Park J, Lee S, Choi J, Ahn K, Park B, Park J, Kang S, Lee YH. Fungal Cytochrome P450 Database. BMC Genomics. 2008;9:402. doi: 10.1186/1471-2164-9-402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xi H, Park J, Ding G, Lee YH, Li Y. SysPIMP: the web-based systematical platform for identifying human disease-related mutated sequences from mass spectrometry. Nucleic Acids Res. 2008;37:D913–D920. doi: 10.1093/nar/gkn848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong MY, Lee EM, Jo YH, Park HC, Kim SR, Hwang JS, Jin BR, Kang PD, Kim KG, Han YS, Kim I. Complete nucleotide sequence and organization of the mitogenome of the silk moth Caligula boisduvalii (Lepidoptera: Saturniidae) and comparison with other lepidopteran insects. Gene. 2008;413:49–57. doi: 10.1016/j.gene.2008.01.019. [DOI] [PubMed] [Google Scholar]
- Sheffield NC, Song H, Cameron SL, Whiting MF. A comparative analysis of mitochondrial genomes in Coleoptera (Arthropoda: Insecta) and genome descriptions of six new beetles. Mol Biol Evol. 2008;25:2499–2509. doi: 10.1093/molbev/msn198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron SL, Lambkin CL, Barker SC, Whiting MF. A mitochondrial genome phylogeny of Diptera: whole genome sequence data accurately resolve relationships over broad timescales with high precision. Syst Entomol. 2007;32:40–59. doi: 10.1111/j.1365-3113.2006.00355.x. [DOI] [Google Scholar]
- Boore JL, Collins TM, Stanton D, Daehler LL, Brown WM. Deducing the pattern of arthropod phylogeny from mitochondrial DNA rearrangements. Nature. 1995;376:163–165. doi: 10.1038/376163a0. [DOI] [PubMed] [Google Scholar]
- Johnson KP, Whiting MF. Multiple genes and the monophyly of Ischnocera (Insecta: Phthiraptera) Mol Phylogenet Evol. 2002;22:101–110. doi: 10.1006/mpev.2001.1028. [DOI] [PubMed] [Google Scholar]
- Johnson KP, Cruickshank RH, Adams RJ, Smith VS, Page RD, Clayton DH. Dramatically elevated rate of mitochondrial substitution in lice (Insecta: Phthiraptera) Mol Phylogenet Evol. 2003;26:231–242. doi: 10.1016/S1055-7903(02)00342-1. [DOI] [PubMed] [Google Scholar]
- Price RD, Johnson KP. Four new species of Myrsidea Waterston chewing lice (Phthiraptera: Menoponidae) from the Malagasy warblers (Passeriformes) Zootaxa. 2006;1297:47–55. [Google Scholar]
- Klass K-D, Picker MD, Damgaard J, van Noort S, Tojo K. The Taxonomy, Genitalic Morphology, and Phylogenetic Relationships of South African Mantophasmatodea (Insecta) Entomol Abh. 2003;61:3–67. [Google Scholar]
- Damgaard J, Klass KD, Picker MD, Buder G. Phylogeny of the Heelwalkers (Insecta: Mantophasmatodea) based on mtDNA sequences, with evidence for additional taxa in South Africa. Mol Phylogenet Evol. 2008;47:443–462. doi: 10.1016/j.ympev.2008.01.026. [DOI] [PubMed] [Google Scholar]
- Cameron SL, Whiting MF. Mitochondrial genomic comparisons of the subterranean termites from the Genus Reticulitermes (Insecta: Isoptera: Rhinotermitidae) Genome. 2007;50:188–202. doi: 10.1139/G06-148. [DOI] [PubMed] [Google Scholar]
- Yukuhiro K, Sezutsu H, Itoh M, Shimizu K, Banno Y. Significant levels of sequence divergence and gene rearrangements have occurred between the mitochondrial genomes of the wild mulberry silkmoth, Bombyx mandarina, and its close relative, the domesticated silkmoth, Bombyx mori. Mol Biol Evol. 2002;19:1385–1389. doi: 10.1093/oxfordjournals.molbev.a004200. [DOI] [PubMed] [Google Scholar]
- Solignac M. Mitochondrial DNA in the Drosophila melanogaster complex. Genetica. 2004;120:41–50. doi: 10.1023/B:GENE.0000017628.52056.97. [DOI] [PubMed] [Google Scholar]
- Yang Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 2007;24:1586–1591. doi: 10.1093/molbev/msm088. [DOI] [PubMed] [Google Scholar]