Abstract
Background
Helicobacter pylori has been strongly associated with chronic gastritis, peptic and duodenal ulcers, and it is a risk factor for gastric cancer. Three major virulence factors of H. pylori have been described: the vacuolating toxin (VacA), the cytotoxin-associated gene product (CagA) and the adhesion protein BabA2. Since considerable geographic diversity in the prevalence of H. pylori virulence factors has been reported, the aim of this work was to establish the H. pylori and vacA, cagA and babA2 gene status in 238 adult patients, from a marginal urban area of Mexico, with chronic gastritis.
Methods
H. pylori was identified in cultures of gastric biopsies by nested PCR. vacA and cagA genes were detected by multiplex PCR, whereas babA2 gene was identified by conventional PCR.
Results
H. pylori-positive biopsies were 143 (60.1%). All H. pylori strains were vacA+; 39.2% were cagA+; 13.3% were cagA+ babA2+ and 8.4% were babA2+. Mexican strains examined possessed the vacA s1, m1 (43.4%), s1, m2 (24.5%), s2, m1 (20.3%) and s2, m2 (11.9%) genotypes.
Conclusion
These results show that the Mexican patients suffering chronic gastritis we have studied had a high incidence of infection by H. pylori. Forty four percent (63/143) of the H. pylori strains analyzed in this work may be considered as highly virulent since they possessed two or three of the virulence markers analyzed: vacA s1 cagA babA2 (9.8%, 14/143), vacA s1 babA2 (4.9%, 7/143), and vacA s1 cagA (29.4%, 42/143). However, a statistically significant correlation was not observed between vacAs1, cagA and babA2 virulence markers (χ2 test; P > 0.05).
Background
Helicobacter pylori is a spiral-shaped Gram-negative bacterium that has been strongly associated with chronic gastritis and peptic ulcer disease [1,2], and it is a risk factor for gastric cancer [3-5]
Three major virulence factors of H. pylori have been described: the cytotoxin-associated gene product (CagA), the vacuolating toxin (VacA) and the adhesion protein BabA2. The cytotoxin-associated gene A (CagA) is a protein with a molecular mass of approximately 125–140 kDa, encoded by the cagA gene, [6,7], that is translocated into gastric epithelial cells by a type IV secretion system, encoded by the cag pathogenicity island (cag PAI) [8]. Inside epithelial cells CagA is phosphorylated on tyrosine residues by host cell Src kinases and stimulates cell-signaling pathways [9], which in turn causes elongation of the cell [10] and activation of proto-oncogenes [11].
The vacuolating cytotoxin gene vacA is polymorphic, varying in the signal and middle regions. The main signal region alleles are s1 and s2, whereas the middle region alleles are m1 and m2 [12,13]. VacA is a toxin that binds to several epithelial receptors [14-16] and forms hexameric pores [17], which later are endocytosed and converted in large vacuoles [18].
The BabA adhesin of H. pylori is an outer membrane protein that binds to the fucosylated histo-blood group antigens on the surface of gastric epithelial cells [19,20]. It has been reported that H. pylori strains possessing babA2 gene, which encodes active BabA adhesin, are associated with increased gastric inflammation [21] and increased risk for duodenal ulcer and adenocarcinoma [22].
H. pylori virulence factors frequency varies widely. For instance, vacAs1 prevalence fluctuates from 48% [23] to 100% [24] whereas cagA prevalence fluctuates from 66.9% [23] to 83.6% [25] and 100% [26]. babA2 reported frequencies vary from 46% [27] to 82.3% [28] in South-American countries. Since considerable geographic diversity in the prevalence of H. pylori virulence factors has been reported, the aim of this work was to establish the H. pylori and vacA, cagA and babA2 gene status in 238 adult patients, from a marginal urban area of Mexico, with chronic gastritis.
Materials and methods
Subjects and clinical samples
Two hundred and thirty eight patients, endoscopic diagnosed with chronic gastritis (154 women and 84 men) with an average age of 52.24 years (range 16 to 83), who had undergone endoscopy in Hospital General Regional 72 of the Instituto Mexicano del Seguro Social at Tlalnepantla, Estado de Mexico, Mexico, were included in this study. Written informed consent for participation was obtained from every patient before the study. The ethics committee at Hospital General Regional 72 approved the study protocol in advance. Antral biopsy specimens were evaluated for the presence of H. pylori by culture. The genotype profiles of H. pylori isolates were determined by PCR.
H. pylori culture
For bacterial culture, biopsy specimens were macerated and homogenized in Brucella Broth and a 100 μL aliquot was inoculated on Casman Agar (Difco) containing 5% horse blood and H. pylori selective supplement (Oxoid-SR 147E). Agar plates were incubated in 6% CO2, for up to four days. Colonies were identified as H. pylori according to standard criteria including negative Gram staining, typical cell morphology, and positive reactions to catalase, oxidase, and urease.
Identification of H. pylori by nested PCR
H. pylori DNA was extracted from colonies collected in microcentrifuge tubes containing 125 μL of sterile phosphate-buffered saline. Suspensions were vortexed vigorously for 2 min; the tubes were boiled in a water bath for 15 min, cooled in ice, and centrifuged at 13000 × g for 1 min. DNA in supernatant was stored at -20°C until used as template in PCR.
H. pylori was detected by nested PCR. First PCR run was done as described by Li et al., [29] with primers EHC-U (5'-CCCTCACGCCATCAGTCCCAAAAA-3') and EHC-L (5'-AAGAAGTCAAAAACGCCCCAAAAC-3'). Amplification was performed in 25 μL reaction volume containing 1 μL (25 pmol) of each primer (EHC-U and EHC-L, Sigma-Genosys), 2.5 μL 10× Buffer Solution, 17.5 μL nuclease-free water, 3 μL template DNA, 1.5 mmol MgCl2, 0.5 U AmpliTaq polimerase and 100 mmol dNTPs (PuRetaqTM Ready-To-GoTM PCR beads). Products were amplified under the following conditions: 5 min at 95°C for initial denaturation followed by 40 cycles of 45 s at 94°C, 45 s at 59°C, and 30 s at 72°C with a final round of 10 min at 72°C in a Corbett Research CGI-96 Thermocycler. A 417 bp product was obtained by this procedure. Second PCR run was done as described by Song et al., [30] with primers ET-5U (5'-GCCAAATCATAAGTCCGCAGAA-3') and ET-5L (5'-TGAGACTTTCCTAGAAGCGGTGTT-3') complementary to an internal fragment of the amplicon obtained with EHC-U and EHC-L primers. Amplification conditions were identical to those of the first run, except that 0.2 μL of the first PCR run product as template, and 25 cycles, were used. A 230 bp amplicon was obtained.
In each experiment, both positive and negative controls, with DNA from H. pylori ATCC 43629 and without template DNA, were included.
Detection of cagA, vacA, and babA2 by PCR
In order to detect cagA and vacA alleles, primers and multiplex PCR amplification conditions described by Chattopadhyay, et al., [31] and Atherton et al., [13,32] were used. These PCR protocols detect cagA (350 bp amplicon) and distinguish vacA s1 (259 bp amplicon) from vacA s2 (286 bp amplicon), and vacA m1 (567 bp amplicon) from vacA m2 (642 bp amplicon). PCR detection of babA2 was done as described by Gerhard et al., [22]. PCR products were analyzed by agarose gel electrophoresis at 120 V, 94 mA for 120 min. Gels were stained with ethidium bromide and photographed under UV illumination with Gel Logic 100 system (Kodak).
Results and discussion
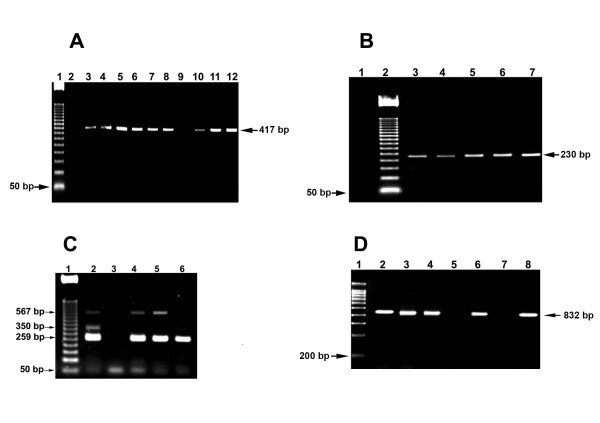
It is known that more than 50% of the world's human population is colonized by H. pylori [33,34]. We report here that H. pylori was cultured from 60.1% biopsy samples (143/238) and identified by nested PCR, which rendered the expected 417 bp and 230 bp amplicons (Fig 1A, B) as reported by Li et al., [29] and Song et al., [30]. This result is in agreement with previously reported H. pylori prevalence in Mexican people. A community-based national seroprevalence survey of H. pylori infection in Mexico showed an overall prevalence of 66%. Twenty-percent of one-year-old children had antibodies against H. pylori, with an increased seropositivity of up to 50% in children who were 10 years of age [35]. Variations in prevalence have been reported among particular regions with a prevalence of 86.1% in southeastern Mexico [36] and 47.1% in children from northwestern Mexico [37].
Figure 1.
Identification of H. pylori isolated from gastric biopsy samples and genotyping of its main virulence genes by PCR. Images shown are from representative gel electrophoresis. A: Lane 1, MWM 50 bp-ladder; Lanes 2 and 9 negative control without DNA; Lanes 3–8, 10 and 11, H. pylori isolated from gastric biopsy samples (417-bp amplicon); Lane 12, reference strain H. pylori ATCC 43629. B: Lane 1, negative control without DNA, Lane 2, MWM 50 bp-ladder; Lanes 3–6, H. pylori isolated from gastric biopsy samples (230-bp amplicon); Lane 7, H. pylori ATCC 43629. C: Lane 1, MWM 50-bp-ladder; Lane 2, H. pylori isolated from gastric biopsy sample (vacA s1, m1 cagA); Lane 3, negative control without DNA; Lanes 4–5, H. pylori isolated from gastric biopsy samples (vacA s1, m1); Lane 6, ATCC 43629. D: Lane 1, MWM 200 bp-ladder; Lane 2, ATCC 43629; Lanes 3,4,6 and 8, babA2-positive H. pylori isolated from gastric biopsy samples; Lane 5, control without DNA; Lane 7, babA2-negative H. pylory isolated from gastric biopsy sample.
All H. pylori strains were positive for the vacA gene (Table 1), as evidenced by PCR product sizes, which enabled to differentiate s and m alleles (Fig. 1C). s1 m1 was the most frequent vacA allelic combination in the H. pylori strains examined, followed by s1 m2, s2 m1 and s2 m2 (Table 1). These results suggest that two thirds of these strains are virulent, as it has been reported that H. pylori isolates with vacAs1m1 and vacAs1m2 allelic combinations exhibit high and low vacuolating activity, respectively, whereas those with vacAs2 fails to induce cell vacuolation in vitro [13].
Table 1.
Frequency of vacA H. pylori genotypes in Mexican patients with chronic gastritis
| Genotype | Number (%) |
| s1m1 | 62 (43.4) |
| s1m2 | 35 (24.5) |
| s2m1 | 29 (20.3) |
| s2m2 | 17 (11.9) |
| 143 (100) | |
Amplicons of the cagA and babA2 genes were detected in agarose gels as 350 bp and 832 bp bands, respectively, (Fig. 1C and 1D). Fifty two percent of the H. pylori isolates were cagA-positive, prevalence less than that reported in other studies from South-American [28,38,39] and Asian countries [40]. In Mexico, prevalence of H. pylori infection with CagA-positive strains varies from 47.6% to 63.4% [41]. cagA-positive H. pylori strains have been associated with the severe mucosa inflammation that underlies peptic ulcer, atrophic gastritis and gastric carcinoma [42-44].
Although vacAs1 cagA+ H. pylori strains had been considered as virulent, in a study of H. pylori isolates from Mexican patients it was reported that vacAs1b and cagA+ strains were found at similar frequencies in adults with and without peptic ulcers [45].
H. pylori BabA adhesin, encoded by the babA2 gene, participates in adhesion of H. pylori to Leb antigens on human gastric epithelial cells [19]. The babA2 gene was found in only 21.7% of the H. pylori isolates (Table 2). This frequency is considerably lesser than babA2 reported frequencies, which vary from 46% [27] to 82.3% [28] in South-American countries. It is important to note that PCR detection of babA2 in H. pylori do not always correlates with its adhesive properties and, conversely, failure to detect babA2 by PCR does not mean that the strain is not adherent, as there is substantial allelic variation in babA2 gene [46-48].
Table 2.
Frequency of cagA and babA2 genes in H. pylori isolates
| Genotype | Number (%) | ||
| vacA | cagA | babA2 | |
| + | + | + | 19 (13.3) |
| + | - | + | 12 (8.4) |
| + | + | - | 56 (39.2) |
| + | - | - | 56 (39.2) |
| 143 (100) | |||
Forty four percent of the H. pylori strains analyzed in this work (63/143) possessed two or three of the virulence markers analyzed (Table 3): vacA s1 cagA babA2 (9.8%, 14/143), vacA s1 babA2 (4.9%, 7/143), and vacA s1 cagA (29.4%, 42/143).
Table 3.
Frequency of cagA an babA2 genes in vacAs1-positive H. pylori isolates
| Genotype | Number (%) | ||
| vacAs1 | cagA | babA2 | |
| + | + | + | 14 (14.4) |
| + | - | + | 7 (7.2) |
| + | + | - | 42 (43.3) |
| + | - | - | 34 (35.0) |
| 97 (100) | |||
Conclusion
Results presented here show that the Mexican patients suffering chronic gastritis we have studied had a high incidence of infection by H. pylori, and suggest that 44% of the H. pylori strains examined may be considered virulent, since they possessed two or three of the virulence markers analyzed. However, a statistically significant correlation was not observed between vacAs1, cagA and babA2 virulence markers (χ2 test; P > 0.05).
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
GP, EM and EN carried out the molecular studies; RR, JC and AC, obtained the gastric biopsy samples; SA and CR carried out the microbiological procedures; SV conceived of the study, and participated in its design and coordination, and drafted the manuscript. All authors read and approved the final manuscript.
Acknowledgments
Acknowledgements
Project supported by PAPIIT IN216508, DGAPA, UNAM.
Contributor Information
Gloria Luz Paniagua, Email: mya@unam.mx.
Eric Monroy, Email: mopi@unam.mx.
Raymundo Rodríguez, Email: rodmoc@prodigy.net.mx.
Salvador Arroniz, Email: chinpato@prodigy.net.mx.
Cristina Rodríguez, Email: cristy@unam.mx.
José Luis Cortés, Email: coel63@hotmail.com.
Ausencio Camacho, Email: dipatci@hotmail.com.
Erasmo Negrete, Email: negretee@yahoo.com.
Sergio Vaca, Email: vsergio@servidor.unam.mx.
References
- Marshall BJ, Warrren JR. Unidentified curved bacilli in the stomach of patients with gastritis and peptic ulceration. Lancet . 1984;1:1311–1315. doi: 10.1016/s0140-6736(84)91816-6. [DOI] [PubMed] [Google Scholar]
- Petersen WL. Helicobacter pylori and peptic ulcer disease. N Engl J Med. 1991;324:1043–1048. doi: 10.1056/NEJM199104113241507. [DOI] [PubMed] [Google Scholar]
- NIH Consensus Development Panel Helicobacter pylori in peptic ulcer disease. J Am Med Assoc. 1994;272:65–69. doi: 10.1001/jama.272.1.65. [DOI] [PubMed] [Google Scholar]
- International Agency for Research on Cancer . Schistosomes, liver flukes and Helicobacter pylori. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. Vol. 61. World Health Organization, Geneva, Switzerland; 1994. [PMC free article] [PubMed] [Google Scholar]
- Isaacson PG. Gastric lymphoma and Helicobacter pylori. N Engl J Med. 1994;330:1310–1311. doi: 10.1056/NEJM199405053301812. [DOI] [PubMed] [Google Scholar]
- Covacci A, Censini S, Bugnoli M, Petracca R, Burroni D, Macchia G, Massone A, Papini E, Xiang Z, Figura N, Rappuoli R. Molecular characterization of the 128-kDa immunodominant antigen of Helicobacter pylori associated with cytotoxicity and duodenal ulcer. Proc Natl Acad Sci USA. 1993;90:5791–5795. doi: 10.1073/pnas.90.12.5791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tummuru MK, Cover TL, Blaser MJ. Cloning and expression of a high-molecular-mass major antigen of Helicobacter pylori: evidence of linkage to cytotoxin production. Infect Immun. 1993;61:1799–1809. doi: 10.1128/iai.61.5.1799-1809.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Odenbreit S, Puls J, Sedlmaier B, Gerland E, Fischer W, Hass R. Translocation of Helicobacter pylori CagA into gastric epithelial cells by type IV secretion. Science. 2000;287:1497–1500. doi: 10.1126/science.287.5457.1497. [DOI] [PubMed] [Google Scholar]
- Selbach M, Moese S, Hauck CR, Meyer TF, Backert S. Src is the kinase of the Helicobacter pylori CagA protein in vitro and in vivo. J Biol Chem. 2002;277:6775–6778. doi: 10.1074/jbc.C100754200. [DOI] [PubMed] [Google Scholar]
- Segal ED, Cha J, Lo J, Falkow S, Tompkins LS. Altered states: involvement of phosphorylated CagA in the induction of host cellular growth changes by Helicobacter pylori. Proc Natl Acad Sci USA. 1999;96:14559–14564. doi: 10.1073/pnas.96.25.14559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer-ter-Vehn T, Covacci A, Kist M, Pahl HL. Helicobacter pylori activates mitogen-activated protein kinase cascades and induces expression of the proto-oncogenes c-fos and c-jun. J Biol Chem. 2000;275:16064–16072. doi: 10.1074/jbc.M000959200. [DOI] [PubMed] [Google Scholar]
- Cover TL, Tummuru MK, Cao P, Thompson SA, Blaser MJ. Divergence of genetic sequences for the vacuolating cytotoxin among Helicobacter pylori strains. J Biol Chem. 1994;269:10566–10573. [PubMed] [Google Scholar]
- Atherton JC, Cao P, Peek RM, Tummuru MK, Blaser MJ, Cover TL. Mosaicism in vacuolating cytotoxin alleles of Helicobacter pylori. Association of specific vacA types with cytotoxin production and peptic ulceration. J Biol Chem. 1995;270:17771–17777. doi: 10.1074/jbc.270.30.17771. [DOI] [PubMed] [Google Scholar]
- Seto K, Hayashi-Kuwabara Y, Yoneta T, Suda H, Tamaki H. Vacuolation induced by cytotoxin from Helicobacter pylori is mediated by EGF receptor in HeLa cells. FEBS Lett. 1998;431:347–350. doi: 10.1016/S0014-5793(98)00788-1. [DOI] [PubMed] [Google Scholar]
- Padilla PI, Wada A, Yahiro K, Kimura M, Niidome T, Aoyagi H, Kumatori A, Anami M, Hayashi T, Fujisawa J, Saito H, Moss J, Hirayama T. Morphologic differentiation of HL-60 cells is associated with appearance of RPTP beta and induction of Helicobacter pylori VacA sensitivity. J Biol Chem. 2000;275:15200–15206. doi: 10.1074/jbc.275.20.15200. [DOI] [PubMed] [Google Scholar]
- Yahiro K, Wada A, Nakayama M, Kimura T, Ogushi K, Niidome T, Aoyagi H, Yoshino K, Yonezawa K, Moss J, Hirayama T. Protein-tyrosine phosphatase alpha, RPTP alpha, is a Helicobacter pylori VacA receptor. J Biol Chem. 2003;278:19183–19189. doi: 10.1074/jbc.M300117200. [DOI] [PubMed] [Google Scholar]
- Czajkowsky DM, Iwamoto H, Cover TL, Shao ZF. The vacuolating toxin from Helicobacter pylori forms hexameric pores in lipid bilayers at low pH. Proc Natl Acad Sci USA. 1999;96:2001–2006. doi: 10.1073/pnas.96.5.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papini E, de Bernard M, Milia E, Bugnoli M, Zerial M, Rappuoli R, Montecucco C. Cellular vacuoles induced by Helicobacter pylori originate from late endosomal compartments. Proc Natl Acad Sci USA. 1994;91:9720–9724. doi: 10.1073/pnas.91.21.9720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borén T, Falk P, Roth KA, Larson G, Normark S. Attachment of Helicobacter pylori to human gastric epithelium mediated by blood group antigens. Science. 1993;262:1892–1895. doi: 10.1126/science.8018146. [DOI] [PubMed] [Google Scholar]
- Ilver D, Arnqvist A, Ogren J, Frick IM, Dangeruta K, Incecik ET, Berg DE, Covacci A, Engstrand L, Borén T. Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science. 1998;279:373–376. doi: 10.1126/science.279.5349.373. [DOI] [PubMed] [Google Scholar]
- Prinz C, Schoniger M, Rad R, Becker I, Keiditsch E, Wagenpfeil S, Classen M, Rosch T, Schepp W, Gerhard M. Key importance of the Helicobacter pylori adherence factor blood group antigen binding adhesin during chronic gastric inflammation. Cancer Res. 2001;61:1903–1909. [PubMed] [Google Scholar]
- Gerhard M, Lehn N, Neumayer N, Borén T, Rad R, Schepp W, Miehlke S, Classen M, Prinz C. Clinical relevance of the Helicobacter pylori gene for blood-group antigen-binding adhesin. Proc Natl Acad Sci USA. 1999;96:12778–12783. doi: 10.1073/pnas.96.22.12778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alarcón T, Domingo D, Martinez MJ, López-Brea M. cagA gene and vacA alleles in Spanish Helicobacter pylori clinical isolates from patients of different ages. FEMS Immunol Med Microbiol. 1999;24:215–219. doi: 10.1111/j.1574-695X.1999.tb01285.x. [DOI] [PubMed] [Google Scholar]
- Chomvarin C, Namwat W, Chaicumpar K, Mairiang P, Sangchan A, Sripa B, Tor-Udom S, Vilaichone RK. Prevalence of Helicobacter pylori vacA, cagA, iceA and babA2 genotypes in Thai dyspeptic patients. Int J Infect Dis . 2008;12:30–36. doi: 10.1016/j.ijid.2007.03.012. [DOI] [PubMed] [Google Scholar]
- Salehi Z, Jelodar MH, Rassa M, Ahaki M, Mollasalehi H, Mashayekhi F. Helicobacter pylori cagA status and peptic ulcer disease in Iran. Dig Dis Sci. 2009;54:608–613. doi: 10.1007/s10620-008-0378-8. [DOI] [PubMed] [Google Scholar]
- Pan ZJ, Hulst RW van der, Feller M, Xiao SD, Tytgat GN, Dankert J, van der Ende A. Equally high prevalence of infection with cagA-positive Helicobacter pylori in Chinese patients with peptic ulcer disease and those with chronic gastritis-associated dyspepsia. J Clin Microbiol. 1997;35:1344–1347. doi: 10.1128/jcm.35.6.1344-1347.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliveira AG, Santos A, Guerra JB, Rocha GA, Rocha AM Oliveira CA, Cabral MM, Nogueira AM, Queiroz DM. babA2- and cagA-positive Helicobacter pylori strains are associated with duodenal ulcer and gastric carcinoma in Brazil. J Clin Microbiol. 2003;41:3964–3966. doi: 10.1128/JCM.41.8.3964-3966.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torres LE, Melián K, Moreno A, Alonso J, Sabatier CA, Hernández M, Bermúdez L, Rodríguez BL. Prevalence of vacA, cagA and babA2 genes in Cuban Helicobacter pylori isolates. World J Gastroenterol. 2009;15:204–210. doi: 10.3748/wjg.15.204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li C, Musich PR, Ha T, Ferguson DA, Jr, Patel NR, Chi DS, Thomas E. High prevalence of Helicobacter pylori in saliva demonstrated by a novel PCR assay. J Clin Pathol. 1995;48:662–666. doi: 10.1136/jcp.48.7.662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song Q, Haller B, Schmid RM, Adler G, Bode G. Helicobacter pylori in dental plaque: a comparison of different PCR primer sets. Dig Dis Sci. 1999;44:479–484. doi: 10.1023/A:1026680618122. [DOI] [PubMed] [Google Scholar]
- Chattopadhyay S, Patra R, Ramamurthy T, Chowdhury A, Santra A, Dhali GK, Bhattacharya SK, Berg DE, Nair GB, Mukhopadhyay AK. Multiplex PCR assay for rapid detection and genotyping of Helicobacter pylori directly from biopsy specimens. J Clin Microbiol. 2004;42:2821–2824. doi: 10.1128/JCM.42.6.2821-2824.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atherton JC, Cover TL, Twells RJ, Morales MR, Hawkey CJ, Blaser MJ. Simple and accurate PCR-based system for typing vacuolating cytotoxin alleles of Helicobacter pylori. J Clin Microbiol. 1999;37:2979–2982. doi: 10.1128/jcm.37.9.2979-2982.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsonnet J. Incidence of Helicobacter pylori infection. Aliment Pharmacol Ther. 1995;9:45–51. [PubMed] [Google Scholar]
- Atherton JC. The pathogenesis of Helicobacter pylori-induced gastro-duodenal diseases. Annu Rev Pathol Mech Dis. 2006;1:63–96. doi: 10.1146/annurev.pathol.1.110304.100125. [DOI] [PubMed] [Google Scholar]
- Torres J, Leal-Herrera Y, Perez-Perez G, Gomez A, Camorlinga-Ponce M, Cedillo-Rivera R, Tapia-Conyer R, Muñoz O. A community-based seroepidemiological study of Helicobacter pylori infection in Mexico. J Infect Dis. 1998;178:1089–1094. doi: 10.1086/515663. [DOI] [PubMed] [Google Scholar]
- Gaurner J, Mohar A, Parsonnet J, Halperin D. The association of Helicobacter pylori with gastric cancer and preneoplastic gastric lesions in Chiapas, Mexico. Cancer. 1993;71:297–301. doi: 10.1002/1097-0142(19930115)71:2<297::AID-CNCR2820710205>3.0.CO;2-9. [DOI] [PubMed] [Google Scholar]
- Jiménez-Guerra F, Shetty P, Kurpad A. Prevalence of and risk factors for Helicobacter pylori infection in school children in Mexico. Ann Epidemiol. 2000;10:474. doi: 10.1016/S1047-2797(00)00109-5. [DOI] [PubMed] [Google Scholar]
- Faundez G, Troncoso M, Figueroa G. cagA and vacA in strains of Helicobacter pylori from ulcer and non-ulcerative dyspepsia patients. BMC Gastroenterol. 2002;2:20. doi: 10.1186/1471-230X-2-20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattar R, dos Santos AF, Eisig JN, Rodrigues TN, Silva FM, Lupinacci RM, Iriya K, Carrilho FJ. No correlation of babA2 with vacA and cagA genotypes of Helicobacter pylori and grading of gastritis from peptic ulcer disease patients in Brazil. Helicobacter. 2005;10:601–608. doi: 10.1111/j.1523-5378.2005.00360.x. [DOI] [PubMed] [Google Scholar]
- Maeda S, Ogura K, Yoshida H, Kanai F, Ikenoue T, Kato N, Shiratori Y, Omata M. Major virulence factors, VacA and CagA, are commonly positive in Helicobacter pylori isolates in Japan. Gut. 1998;42:338–343. doi: 10.1136/gut.42.3.338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torres J, Lopez L, Lazcano E, Camorlinga M, Flores L, Muñoz O. Trends in Helicobacter pylori infection and gastric cancer in Mexico. Cancer Epidemiol Biomarkers Prev. 2005;14:1874–1877. doi: 10.1158/1055-9965.EPI-05-0113. [DOI] [PubMed] [Google Scholar]
- Kuipers EJ, Pérez-Pérez GI, Meuwissen SG, Blaser MJ. Helicobacter pylori and atrophic gastritis: importance of the cagA status. J Natl Cancer Inst. 1995;87:1777–1780. doi: 10.1093/jnci/87.23.1777. [DOI] [PubMed] [Google Scholar]
- Blaser MJ, Pérez-Pérez GI, Kleanthous H, Cover TL, Peek RM, Chyou PH, Stemmermann GN, Nomura A. Infection with Helicobacter pylori strains possessing cagA is associated with an increased risk of developing adenocarcinoma of the stomach. Cancer Res. 1995;55:2111–2115. [PubMed] [Google Scholar]
- Yamaoka Y, Kodama T, Gutierrez O, Kim JG, Kashima K, Graham DY. Relationship between Helicobacter pylori iceA, cagA, and vacA status and clinical outcome: studies in four different countries. J Clin Microbiol. 1999;37:2274–2279. doi: 10.1128/jcm.37.7.2274-2279.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- González-Valencia G, Atherton JC, Muñoz O, Dehesa M, Madrazo-de la Garza A, Torres J. Helicobacter pylori vacA and cagA genotypes in Mexican adults and children. J Infect Dis. 2000;182:1450–1454. doi: 10.1086/315864. [DOI] [PubMed] [Google Scholar]
- Hennig EE, Mernaugh R, Edl J, Cao P, Cover TL. Heterogeneity among Helicobacter pylori strains in expression of the outer membrane protein BabA. Infect Immun. 2004;72:3429–3435. doi: 10.1128/IAI.72.6.3429-3435.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olfat FO, Zheng Q, Oleastro M, Voland P, Borén T, Karttunen R, Engstrand L, Rad R, Prinz C, Gerhard M. Correlation of the Helicobacter pylori adherence factor BabA with duodenal ulcer disease in four European countries. FEMS Immunol Med Microbiol. 2005;44:151–156. doi: 10.1016/j.femsim.2004.10.010. [DOI] [PubMed] [Google Scholar]
- Colbeck JC, Hansen LM, Fong JM, Solnick JV. Genotyping profile of the outer membrane proteins BabA and BabB in clinical isolates of Helicobacter pylori. Infect Immun. 2006;74:4375–4378. doi: 10.1128/IAI.00485-06. [DOI] [PMC free article] [PubMed] [Google Scholar]