Abstract
Genome signatures are data vectors derived from the compositional statistics of DNA. The self-organizing map (SOM) is a neural network method for the conceptualisation of relationships within complex data, such as genome signatures. The various parameters of the SOM training phase are investigated for their effect on the accuracy of the resulting output map. It is concluded that larger SOMs, as well as taking longer to train, are less sensitive in phylogenetic classification of unknown DNA sequences. However, where a classification can be made, a larger SOM is more accurate. Increasing the number of iterations in the training phase of the SOM only slightly increases accuracy, without improving sensitivity. The optimal length of the DNA sequence k-mer from which the genome signature should be derived is 4 or 5, but shorter values are almost as effective. In general, these results indicate that small, rapidly trained SOMs are generally as good as larger, longer trained ones for the analysis of genome signatures. These results may also be more generally applicable to the use of SOMs for other complex data sets, such as microarray data.
Keywords: Genome Signature, Self-Organizing Map, Viruses, Phylogeny, Jack-Knife Method, Microarray, Metagenomics, Herpesvirus
Introduction
Molecular evolutionary methodology revolves around the production of sequence alignments and trees. However, as evolutionary distance increases between two homologous molecules, their similarity may decay to the point where they are no longer alignable. Construction of a phylogenetic tree under such circumstances becomes impossible. One method that has been suggested for the study of distant evolutionary relationships is that of genomic signatures or genome signatures† (Karlin and Ladunga, 1994; Karlin and Burge, 1995; Karlin and Mrazek, 1996). At least one reviewer has come to the conclusion that it is the preferred method in cases where evolutionary distance, recombination, horizontal transmission or variable mutation rates may confound traditional alignment-based techniques (Brocchieri, 2001).
The first derivation of genome signatures predates the invention of DNA sequencing. Biochemical studies revealed that the frequencies of nearest-neighbour dinucleotide pairs in DNA were generally consistent within genomes, and often different between genomes. These characteristic nearest neighbour patterns were termed general schemes (Russell et al. 1976; Russell and Subak-Sharpe, 1977), and constitute, in modern terminology, a subset of genome signatures, those of length k = 2.
As long DNA sequences began to be isolated and computers entered the biological laboratory, it became a simple matter to produce nearest-neighbour frequency tables. Indeed, for any DNA sequence of length N, it is theoretically possible to derive frequency tables for all k-mers ranging from 1 to N, within that sequence. The frequency table at k = 1 corresponds to the raw nucleotide content on one strand. On the assumption that DNA is double stranded under most circumstances in most species, the complementary bases are also scored. This reduces the raw count of the four bases to a single value, between zero and one, representing the GC content of that DNA sequence. Correspondingly, at k = 2, the raw count of 16 dinucleotide frequencies, can be reduced to a vector containing 10 values if the count for each dimer on the top strand is added to the count for its complement on the other strand. There are 10 values, not 8, in this vector since GC, CG, AT and TA are self-complementary. This process is called symmetrization (Karlin and Ladunga, 1994). The symmetrized values in the vector are then usually corrected for the frequencies of their component monomers, as follows:
where fXY is the symmetrized frequency of dinucleotide XY, and fX and fY are the symmetrized frequencies of bases X and Y, respectively. The whole vector is referred to as the genome signature at k = 2 or, particularly in the extensive literature of the Karlin group, simply as ρ* XY. For all values of k, the nomenclature GS-k is here adopted.
The vector thus becomes an array of the ratios of observed frequencies of k-mers to their expected frequencies given an underlying zero-order Markov chain model of a DNA sequence. Even though symmetrization will reduce the size of the vector for large values of k, it is apparent that it will still grow in size at the order of 4k for an alphabet of length 4. In practice, most investigators have confined themselves to the study of genome signatures of k = 2, in other words to ρ *XY, symmetrized dinucleotide frequencies corresponding to general schemes, although in recent years the availability of faster computers has undoubtedly contributed to the increasing use of genome signatures up to k = 10 (Deschavanne et al. 1999; Edwards et al. 2002; Abe et al. 2003a; Sandberg et al. 2003; Campanaro et al. 2005; Dufraigne et al. 2005; Wang et al. 2005; Paz et al. 2006).
The length of DNA required to generate a genome signature has conventionally been taken to be around 50 kb, and for this value it has been observed that the Hamming or Euclidean distances between signatures derived from contigs within species are generally considerably smaller than the corresponding average values between species (Karlin and Ladunga, 1994; Karlin and Burge, 1995; Karlin et al. 1997; Abe et al. 2002; Teeling et al. 2004), even when the same-species contigs are on different chromosomes (Gentles and Karlin, 2001). However, recent work has established that genome signatures within species may be stable over lengths as short as 10 kb (Deschavanne et al. 1999; Karlin, 2001; Abe et al. 2002) or less (Sandberg et al. 2001; Jernigan and Baran, 2002; Abe et al. 2003a; Sandberg et al. 2003; McHardy et al. 2007). This has led to their practical application in the detection of pathenogenicity islands (pIs) in pathogenic bacteria. These are sequences originating in horizontal transmission from one bacterium to another, converting a previously innocuous strain into a pathogenic one. Their foreign origin is often reflected in a genome signature closer to their species of origin than their current host genome (Karlin, 1998; Karlin, 2001; Dufraigne et al. 2005).
Phylogenetic conclusions drawn from comparison of genome signatures have sometimes been controversial. For instance, Karlin et al. (1997) found that cyanobacteria do not form a coherent evolutionary group, and that Methanococcus jannaschii is closer to eukaryotes than to other proteobacteria, and Campbell et al. (1999) suggested that archaea do not form a coherent clade. Karlin (1998) posited a wide variety of further revisions of the prokaryotic phylogeny based on genome signature results, as well as a novel origin for mitochondria (Karlin et al. 1999). Edwards et al. (2002) used genome signatures as part of a revision of the phylogeny of birds. Nevertheless, few authors have felt confident enough to draw phylogenetic trees based on genome signature comparisons. Coenye and Vandamme (2004) have shown that dinucleotide content is only a reliable indicator of relatedness for closely related organisms. To visualize genome signature relationships between species, a variety of other representational schemes have been used including histograms (Karlin and Mrázek, 1997), partial ordering graphs (Karlin et al. 1997), chaos games (Deschavanne et al. 1999; Edwards et al. 2002; Wang et al. 2005), and self-organizing maps (Abe et al. 2003b).
This paper uses self-organizing maps (SOMs) as a tool to explore genome signature variability at phylogenetic levels from superkingdom down to genus. The SOM is a neural network method which spreads multi-dimensional data onto a two-dimensional surface (Kohonen, 1997). Its endpoint is therefore similar to multi-dimensional scaling or principal components analysis, and like these other techniques has been extensively used in biology, principally for the analysis of micro-array data but also to a lesser extent for sequence analysis (Arrigo et al. 1991; Giuliano et al. 1993; Andrade et al. 1997; Tamayo et al. 1999; Kanaya et al. 2001; Wang et al. 2001; Abe et al. 2002; Covell et al. 2003; Ressom et al. 2003; Xiao et al. 2003; Mahony et al. 2004; Oja et al. 2005; Abe et al. 2006; Samsonova et al. 2006). The resulting “flat” representation may be a strong aid to intuitive understanding of the structure of complex multidimensional datasets. The SOM is not a clustering technique per se, but the surface may be divided up into zones that are then treated as clusters. Alternatively, cluster boundaries on the surface may be defined more objectively using additional algorithms (Ultsch, 1993). The SOM is also not hierarchical (unlike UPGMA but like K-means clustering, two other commonly used techniques for the analysis of microarrays). This absence of hierarchy means that it is particularly suited to situations where the natural hierarchy of species relationships, reflecting evolutionary descent, may have been violated, e.g. by horizontal gene transfer.
In this paper, the main parameters of the SOM: its size and the number of iterations used in its construction, are investigated for their effects on its classificatory accuracy. These parameters must be chosen at the beginning of each run of SOM building, and there is little guidance in the SOM literature as to their optimal values. As well as the parameters of the SOM, the value of k used in the genome signature is similarly examined. High k genome signatures are extremely long vectors that may present considerable memory problems even on modern computers. Likewise, lengthy iterations in training the SOM, especially if it is a large one, may consume considerable time.
Methods
1. Genome sequences
Complete genome sequences were downloaded from NCBI Taxonomy Browser (http://www.ncbi.nlm.nih.gov/Taxonomy/taxonomyhome.html/). A Perl script was written to divide complete genome sequences into consecutive strings of 10 or 100 kb, as required. Trailing ends, and genomes shorter than the required string length, were discarded. The resulting FASTA-formatted datasets were then processed to calculate their genome signatures.
Table 1 lists the genomes used as the main data set for the paper, that of viruses of the family Herpesviridae. The analyses shown in Figures 3 to 7 use this set. A larger set of genomes with the widest possible phylogenetic range, including all three superkingdoms of cellular life as well as viruses, is given in Table 2. These are used for the “all-life” and superkingdom-level SOMs in Figure 1. Table 3 lists those viral genomes used for the SOM across a wide set of viral genomes, displayed in Figure 2.
Table 1.
Herpesvirus genome sequences used for the analyses shown in Figures 3 to 7. The nomenclature follows the International Committee on Taxonomy of Viruses (Fauquet et al. 2005).
| Name | Accession | Sub-family | Genus |
|---|---|---|---|
| Psittacid herpesvirus 1 | NC_005264 | Alpha | Iltovirus |
| Gallid herpesvirus 2 | NC_002229 | Alpha | Mardivirus |
| Gallid herpesvirus 3 | NC_002577 | Alpha | Mardivirus |
| Meleagrid herpesvirus 1 | NC_002641 | Alpha | Mardivirus |
| Cercopithecine herpesvirus 1 | NC_004812 | Alpha | Simplexvirus |
| Human herpesvirus 1 | NC_001806 | Alpha | Simplexvirus |
| Human herpesvirus 2 | NC_001798 | Alpha | Simplexvirus |
| Bovine herpesvirus 1 | NC_001847 | Alpha | Varicellovirus |
| Bovine herpesvirus 5 | NC_005261 | Alpha | Varicellovirus |
| Cercopithecine herpesvirus 7 | NC_002686 | Alpha | Varicellovirus |
| Equid herpesvirus 1 | NC_001491 | Alpha | Varicellovirus |
| Equid herpesvirus 4 | NC_001844 | Alpha | Varicellovirus |
| Human herpesvirus 3 | NC_001348 | Alpha | Varicellovirus |
| Suid herpesvirus 1 | NC_006151 | Alpha | Varicellovirus |
| Cercopithecine herpesvirus 8 | NC_006150 | Beta | Cytomegalovirus |
| Chimpanzee cytomegalovirus | NC_003521 | Beta | Cytomegalovirus |
| Human herpesvirus 5 (AD169) | NC_001347 | Beta | Cytomegalovirus |
| Human herpesvirus 5 (Merlin) | NC_006273 | Beta | Cytomegalovirus |
| Murid herpesvirus 1 | NC_004065 | Beta | Muromegalovirus |
| Murid herpesvirus 2 | NC_002512 | Beta | Muromegalovirus |
| Human herpesvirus 6 | NC_001664 | Beta | Roseolovirus |
| Human herpesvirus 6B | NC_000898 | Beta | Roseolovirus |
| Human herpesvirus 7 | NC_001716 | Beta | Roseolovirus |
| Tupaia herpesvirus | NC_002794 | Beta | Tupaiavirus |
| Callitrichine herpesvirus 3 | NC_004367 | Gamma | Lymphocryptovirus |
| Cercopithecine herpesvirus 15 | NC_006146 | Gamma | Lymphocryptovirus |
| Human herpesvirus 4 | NC_001345 | Gamma | Lymphocryptovirus |
| Alcelaphine herpesvirus 1 | NC_002531 | Gamma | Rhadinovirus |
| Ateline herpesvirus 3 | NC_001987 | Gamma | Rhadinovirus |
| Bovine herpesvirus 4 | NC_002665 | Gamma | Rhadinovirus |
| Cercopithecine herpesvirus 17 | NC_003401 | Gamma | Rhadinovirus |
| Equid herpesvirus 2 | NC_001650 | Gamma | Rhadinovirus |
| Human herpesvirus 8 | NC_003409 | Gamma | Rhadinovirus |
| Murid herpesvirus 4 | NC_001826 | Gamma | Rhadinovirus |
| Saimiriine herpesvirus 2 | NC_001350 | Gamma | Rhadinovirus |
| Ictalurid herpesvirus 1 | NC_001493 | unassigned | Ictalurivirus |
| Ostreid herpesvirus 1 | NC_005881 | unassigned | unassigned |
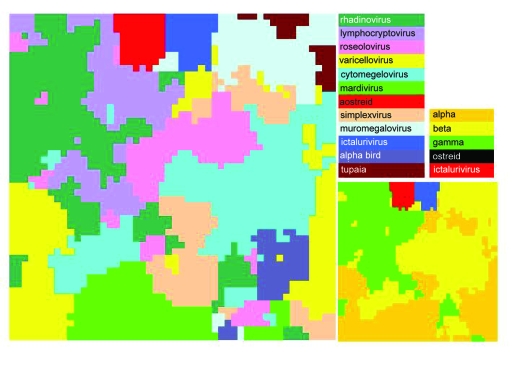
Figure 3.
Dominance maps for GS-2 of 10kb fragments of herpesviruses applied to a 50 ×50 SOM over 500 iterations. The SOM is colored first according to genus membership and then according to family membership (reduced scale inset).
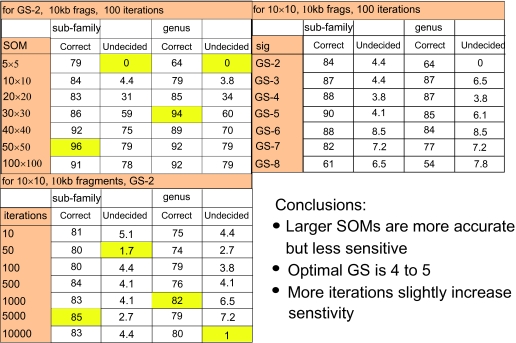
Figure 7.
Jack-knifing experiments to determine effects of SOM size (top left), GS number (top right) and number of iterations (lower left). The “undecided” column indicates the percentage of sequences in the test set that could not be assigned to a sub-family or genus. The “correct” column indicates the percentage of assignable sequences that were correctly assigned. Optimal values are highlighted in yellow.
Table 2.
Genomes used for the analysis shown in Figure 1. In total there are 79 eukaryotic, 156 eubacterial, 30 archaeal and 122 viral genomes with more than 100kb of sequence.
| Name | Superkingdom | Accession |
|---|---|---|
| Aeropyrum pernix K1 | archaea | NC_000854 |
| Archaeoglobus fulgidus DSM 4304 | archaea | NC_000917 |
| cf. Archaea SAR-1 | archaea | NS_000019 |
| Ferroplasma acidarmanus Type I | archaea | NS_000030 |
| Ferroplasma sp. Type II | archaea | NS_000029 |
| Haloarcula marismortui ATCC43049 chromosome I | archaea | NC_006396 |
| Haloarcula marismortui ATCC43049 chromosome II | archaea | NC_006397 |
| Halobacterium sp. NRC-1 | archaea | NC_002607 |
| Halobacterium sp. NRC-1 plasmid pNRC100 | archaea | NC_001869 |
| Methanocaldococcus jannaschii DSM2661 | archaea | NC_000909 |
| Methanococcus maripaludis S2 | archaea | NC_005791 |
| Methanopyrus kandleri AV19 | archaea | NC_003551 |
| Methanosarcina acetivorans C2A | archaea | NC_003552 |
| Methanosarcina barkeri str. fusaro chromosome 1 | archaea | NC_007355 |
| Methanosarcina mazei Go1 | archaea | NC_003901 |
| Methanothermobacter thermautotrophicus str. DeltaH | archaea | NC_000916 |
| Nanoarchaeum equitans Kin4-M | archaea | NC_005213 |
| Natronomonas pharaonis DSM2160 | archaea | NC_007426 |
| Picrophilus torridus DSM9790 | archaea | NC_005877 |
| Pyrobaculum aerophilum str. IM2 | archaea | NC_003364 |
| Pyrococcus abyssi GE5 | archaea | NC_000868 |
| Pyrococcus furiosus DSM3638 | archaea | NC_003413 |
| Pyrococcus horikoshii OT3 | archaea | NC_000961 |
| Sulfolobus acidocaldarius DSM639 | archaea | NC_007181 |
| Sulfolobus solfataricus P2 | archaea | NC_002754 |
| Sulfolobus tokodaii str. 7 | archaea | NC_003106 |
| Thermococcus kodakaraensis KOD1 | archaea | NC_006624 |
| Thermoplasma acidophilum DSM1728 | archaea | NC_002578 |
| Thermoplasma volcanium GSS1 | archaea | NC_002689 |
| Thermoplasmatales archaeon Gpl | archaea | NS_000033 |
| Agrobacterium tumefaciens str. C58 | eubacteria | NC_003062 |
| Anabaena variabilis ATCC 29413 | eubacteria | NC_007413 |
| Aquifex aeolicus VF5 | eubacteria | NC_000918 |
| Azoarcus sp. EbN1 | eubacteria | NC_006513 |
| Bacillus cereus ATCC 10987 | eubacteria | NC_003909 |
| Bacillus cereus E33L | eubacteria | NC_006274 |
| Bacillus subtilis sub sp. subtilis str. 168 | eubacteria | NC_000964 |
| Bacteroides fragilis NCTC9343 | eubacteria | NC_003228 |
| Bacteroides fragilis YCH46 | eubacteria | NC_006347 |
| Bartonella henselae str. Houston-1 | eubacteria | NC_005956 |
| Bartonella quintana str. Toulouse | eubacteria | NC_005955 |
| BBUR Borrelia burgdorferi B31 | eubacteria | NC_001318 |
| Bifidobacterium longum NCC2705 | eubacteria | NC_004307 |
| Bordetella parapertussis 12822 | eubacteria | NC_002928 |
| Bordetella pertussis TohamaI | eubacteria | NC_002929 |
| Bradyrhizobium japonicum USDA110 | eubacteria | NC_004463 |
| Brucella abortus biovar 1 str. 9-941 chromosome I | eubacteria | NC_006932 |
| Brucella abortus biovar 1 str. 9-941 chromosome II | eubacteria | NC_006933 |
| Brucella suis 1330 chromosome I | eubacteria | NC_004310 |
| Buchnera aphidicola str. APS (Acyrthosiphonpisum) | eubacteria | NC_002528 |
| Buchnera aphidicola str. Sg (Schizaphisgraminum) | eubacteria | NC_004061 |
| Burkholderia mallei ATCC23344 chromosome 1 | eubacteria | NC_006348 |
| Burkholderia mallei ATCC23344 chromosome 2 | eubacteria | NC_006349 |
| Burkholderia pseudomallei 1710b chromosome I | eubacteria | NC_007434 |
| Burkholderia pseudomallei 1710b chromosome II | eubacteria | NC_007435 |
| Burkholderia pseudomallei K96243 chromosome 1 | eubacteria | NC_006350 |
| Burkholderia sp. 383 chromosome 1 | eubacteria | NC_007510 |
| Burkholderia sp. 383 chromosome 2 | eubacteria | NC_007511 |
| Burkholderia sp. 383 chromosome 3 | eubacteria | NC_007509 |
| Candidatus Blochmannia pennsylvanicus str. BPEN | eubacteria | NC_007292 |
| Candidatus Pelagibacter ubique HTCC1062 | eubacteria | NC_007205 |
| Carboxydothermus hydrogenoformans Z-2901 | eubacteria | NC_007503 |
| Caulobacter crescentus CB15 | eubacteria | NC_002696 |
| Chlamydia trachomatis A/HAR-13 | eubacteria | NC_007429 |
| Chlamydia trachomatis D/UW-3/CX | eubacteria | NC_000117 |
| Chlamydophila caviae GPIC | eubacteria | NC_003361 |
| Chlamydophila pneumoniae AR39 | eubacteria | NC_002179 |
| Chlamydophila pneumoniae CWL029 | eubacteria | NC_000922 |
| Chlamydophila pneumoniae J138 | eubacteria | NC_002491 |
| Chlorobium chlorochromatii CaD | eubacteria | NC_007514 |
| Clostridium acetobutylicum ATCC824 | eubacteria | NC_003030 |
| Clostridium tetani E88 | eubacteria | NC_004557 |
| Colwellia psychrerythraea 34H | eubacteria | NC_003910 |
| Corynebacterium glutamicum ATCC13032 | eubacteria | NC_003450 |
| Corynebacterium jeikeium K411 | eubacteria | NC_007164 |
| Coxiella burnetii RSA493 | eubacteria | NC_002971 |
| Dechloromonas aromatica RCB | eubacteria | NC_007298 |
| Dehalococcoides sp. CBDB1 | eubacteria | NC_007356 |
| Deinococcus radiodurans R1 chromosome 1 | eubacteria | NC_001263 |
| Deinococcus radiodurans R1 chromosome 2 | eubacteria | NC_001264 |
| Desulfovibrio vulgaris sub sp. vulgaris str. Hildenborough | eubacteria | NC_002937 |
| Desulfovibriode sulfuricans G20 | eubacteria | NC_007519 |
| Ehrlichia canis str. Jake | eubacteria | NC_007354 |
| Erwinia carotovora sub sp. atrosepticaSCRI1043 | eubacteria | NC_004547 |
| Escherichia coli CFT073 | eubacteria | NC_004431 |
| Escherichia coli K12 | eubacteria | NC_000913 |
| Escherichia coli O157:H7EDL933 | eubacteria | NC_002655 |
| Francisella tularensis sub sp. tularensis Schu4 | eubacteria | NC_006570 |
| Geobacter metallireducens GS-15 | eubacteria | NC_007517 |
| Haemophilus ducreyi 35000HP | eubacteria | NC_002940 |
| Haemophilus influenzae 86-028NP | eubacteria | NC_007146 |
| Haemophilus influenzae RdKW20 | eubacteria | NC_000907 |
| Helicobacter pylori 26695 | eubacteria | NC_000915 |
| Helicobacter pylori J99 | eubacteria | NC_000921 |
| Leifsoniaxyli sub sp. xyli str. CTCB07 | eubacteria | NC_006087 |
| Leptospira interrogans serovar Copenhageni chromosome I | eubacteria | NC_005823 |
| Leptospira interrogans serovar Copenhageni chromosome II | eubacteria | NC_005824 |
| Leptospira interrogans serovar Lai str. 56601 chromosome I | eubacteria | NC_004342 |
| Mannheimia succiniciproducens MBEL55E | eubacteria | NC_006300 |
| Mesoplasma florum L1 | eubacteria | NC_006055 |
| Mesorhizobium loti MAFF303099 | eubacteria | NC_002678 |
| Methylococcus capsulatus str. Bath | eubacteria | NC_002977 |
| Mycobacterium avium sub sp. paratuberculosis K-10 | eubacteria | NC_002944 |
| Mycobacterium bovis AF2122/97 | eubacteria | NC_002945 |
| Mycobacterium leprae TN | eubacteria | NC_002677 |
| Mycobacterium tuberculosis H37Rv | eubacteria | NC_000962 |
| Mycoplasma genitalium G-37 | eubacteria | NC_000908 |
| Mycoplasma hyopneumoniae 7448 | eubacteria | NC_007332 |
| Mycoplasma hyopneumoniae J | eubacteria | NC_007295 |
| Mycoplasma synoviae 53 | eubacteria | NC_007294 |
| Neisseria gonorrhoeae FA1090 | eubacteria | NC_002946 |
| Neisseria meningitidis MC58 | eubacteria | NC_003112 |
| Neisseria meningitidis Z2491 | eubacteria | NC_003116 |
| Nitrobacter winogradskyi Nb-255 | eubacteria | NC_007406 |
| Nitrosococcus oceani ATCC 19707 | eubacteria | NC_007484 |
| Nitrosomonas europaea ATCC 19718 | eubacteria | NC_004757 |
| Nocardia farcinicaI FM10152 | eubacteria | NC_006361 |
| Oceanobacillus iheyensis HTE831 | eubacteria | NC_004193 |
| Parachlamydia sp. UWE25 | eubacteria | NC_005861 |
| Pasteurella multocida sub sp. multocida str. Pm70 | eubacteria | NC_002663 |
| Pelobacter carbinolicus DSM2380 | eubacteria | NC_007498 |
| Pelodictyon luteolum DSM273 | eubacteria | NC_007512 |
| Photobacterium profundum SS9 chromosome 1 | eubacteria | NC_006370 |
| Photobacterium profundum SS9 chromosome 2 | eubacteria | NC_006371 |
| Photorhabdus luminescens sub sp. laumondii TTO1 | eubacteria | NC_005126 |
| Prochlorococcus marinus str. NATL2A | eubacteria | NC_007335 |
| Prochlorococcus marinus sub sp. pastoris str. CCMP1986 | eubacteria | NC_005072 |
| Propionibacterium acnes KPA171202 | eubacteria | NC_006085 |
| Pseudoalteromonas haloplanktis TAC125 chromosome I | eubacteria | NC_007481 |
| Pseudoalteromonas haloplanktis TAC125 chromosome II | eubacteria | NC_007482 |
| Psuedomonas fluorescens Pf-5 | eubacteria | NC_004129 |
| Psuedomonas fluorescens PfO-1 | eubacteria | NC_007492 |
| Psuedomonas putida KT2440 | eubacteria | NC_002947 |
| Psuedomonas syringae pv. phaseolicola 1448A | eubacteria | NC_005773 |
| Psuedomonas syringae pv. syringae B728a | eubacteria | NC_007005 |
| Psuedomonas syringae pv. tomato str. DC3000 | eubacteria | NC_004578 |
| Psychrobacter arcticus 273-4 | eubacteria | NC_007204 |
| Ralstonia eutropha JMP134 chromosome 1 | eubacteria | NC_007347 |
| Ralstonia eutropha JMP134 chromosome 2 | eubacteria | NC_007348 |
| Ralstonia solanacearum GMI1000 | eubacteria | NC_003295 |
| Rhodobacter sphaeroides 2.1 chromosome 1 | eubacteria | NC_007493 |
| Rhodobacter sphaeroides 2.1 chromosome 2 | eubacteria | NC_007494 |
| Rickettsia conorii str. Malish 7 | eubacteria | NC_003103 |
| Rickettsia felis URRWXCal2 | eubacteria | NC_007109 |
| Rickettsia prowazekii str. MadridE | eubacteria | NC_000963 |
| Rickettsia typhi str. Wilmington | eubacteria | NC_006142 |
| Salmonella enterica serovar Choleraesuis str. SC-B67 | eubacteria | NC_006905 |
| Salmonella enterica serovar Typhi str. CT18 | eubacteria | NC_003198 |
| Shewanella oneidensis MR-1 | eubacteria | NC_004347 |
| Shigella flexneri 2a str. 2457T | eubacteria | NC_004741 |
| Shigella flexneri 2a str. 301 | eubacteria | NC_004337 |
| Shigella sonnei Ss046 | eubacteria | NC_007384 |
| Sinorhizobium meliloti 1021 | eubacteria | NC_003047 |
| Staphylococcus aureus sub sp. Aureus Mu50 | eubacteria | NC_002758 |
| Staphylococcus haemolyticus JCSC143 | eubacteria | NC_007168 |
| Staphylococcus saprophyticus sub sp. saprophyticus | eubacteria | NC_007350 |
| Streptococcus agalactiae A909 | eubacteria | NC_007432 |
| Streptococcus pyogenes MGAS10394 | eubacteria | NC_006086 |
| Streptococcus pyogenes MGAS315 | eubacteria | NC_004070 |
| Streptococcus pyogenes MGAS500 | eubacteria | NC_007297 |
| Streptococcus pyogenes MGAS6180 | eubacteria | NC_007296 |
| Streptococcus pyogenes SSI-1 | eubacteria | NC_004606 |
| Streptococcus thermophilus CNRZ1066 | eubacteria | NC_006449 |
| Streptococcus thermophilus LMG18311 | eubacteria | NC_006448 |
| Streptomyces avermitilis MA-4680 | eubacteria | NC_003155 |
| Streptomyces coelicolor A3(2) | eubacteria | NC_003888 |
| Synechococcus sp. CC9605 | eubacteria | NC_007516 |
| Synechococcus sp. CC9902 | eubacteria | NC_007513 |
| Thermobifida fusca YX | eubacteria | NC_007333 |
| Thermus thermophilus HB8 | eubacteria | NC_006461 |
| Thiobacillus denitrificans ATCC2525 | eubacteria | NC_007404 |
| Thiomicrospira crunogena XCL-2 | eubacteria | NC_007520 |
| Tropheryma whipplei str. Twist | eubacteria | NC_004572 |
| Vibrio cholerae O1 biovar eltor str. N16961 chromosome I | eubacteria | NC_002505 |
| Vibrio vulnificus CMCP6 chromosome I | eubacteria | NC_004459 |
| Vibrio vulnificus CMCP6 chromosome II | eubacteria | NC_004460 |
| Wolbachia endosymbiont strain TRS of Brugiamalayi | eubacteria | NC_006833 |
| Wolinella succinogenes DSM1740 | eubacteria | NC_005090 |
| Xanthomonas axonopodis pv. citri str. 306 | eubacteria | NC_003919 |
| Xanthomonas campestris pv. campestris str. 8004 | eubacteria | NC_007086 |
| Xanthomonas campestris pv. campestris str. ATCC33913 | eubacteria | NC_003902 |
| Xanthomonas campestris pv. vesicatoria str. 85-10 | eubacteria | NC_007508 |
| Xanthomonas oryzae pv. oryzae KACC10331 | eubacteria | NC_006834 |
| Xylella fastidiosa 9a5c | eubacteria | NC_002488 |
| Xylella fastidiosa Temecula 1 | eubacteria | NC_004556 |
| Yersinia pseudotuberculosis IP32953 | eubacteria | NC_006155 |
| Bos taurus genome 12 | eukaryote | NC_007310 |
| Bos taurus genome 13 | eukaryote | NC_007311 |
| Bos taurus genome 14 | eukaryote | NC_007312 |
| Bos taurus genome 15 | eukaryote | NC_007313 |
| Bos taurus genome 16 | eukaryote | NC_007314 |
| Bos taurus genome 17 | eukaryote | NC_007315 |
| Bos taurus genome 18 | eukaryote | NC_007316 |
| Bos taurus genome 19 | eukaryote | NC_007317 |
| Bos taurus genome 20 | eukaryote | NC_007318 |
| Bos taurus genome 21 | eukaryote | NC_007319 |
| Bos taurus genome 22 | eukaryote | NC_007320 |
| Bos taurus genome 23 | eukaryote | NC_007324 |
| Bos taurus genome 24 | eukaryote | NC_007325 |
| Bos taurus genome 25 | eukaryote | NC_007326 |
| Bos taurus genome 26 | eukaryote | NC_007327 |
| Bos taurus genome 27 | eukaryote | NC_007328 |
| Bos taurus genome 28 | eukaryote | NC_007329 |
| Bos taurus genome 29 | eukaryote | NC_007330 |
| Bos taurus genome X | eukaryote | NC_007331 |
| Candida albicans genomic DNA, genome 7 | eukaryote | NC_007436 |
| Cryptococcus neoformans genome 1 | eukaryote | NC_006670 |
| Cryptococcus neoformans genome 10 | eukaryote | NC_006679 |
| Cryptococcus neoformans genome 11 | eukaryote | NC_006680 |
| Cryptococcus neoformans genome 12 | eukaryote | NC_006681 |
| Cryptococcus neoformans genome 13 | eukaryote | NC_006682 |
| Cryptococcus neoformans genome 14 | eukaryote | NC_006683 |
| Cryptococcus neoformans genome 2 | eukaryote | NC_006684 |
| Cryptococcus neoformans genome 3 | eukaryote | NC_006685 |
| Cryptococcus neoformans genome 4 | eukaryote | NC_006686 |
| Cryptococcus neoformans genome 5 | eukaryote | NC_006687 |
| Cryptococcus neoformans genome 6 | eukaryote | NC_006691 |
| Cryptococcus neoformans genome 7 | eukaryote | NC_006692 |
| Cryptococcus neoformans genome 8 | eukaryote | NC_006693 |
| Cryptococcus neoformans genome 9 | eukaryote | NC_006694 |
| Cryptosporidium parvum genome 1 | eukaryote | NC_006980 |
| Cryptosporidium parvum genome 2 | eukaryote | NC_006981 |
| Cryptosporidium parvum genome 3 | eukaryote | NC_006982 |
| Cryptosporidium parvum genome 4 | eukaryote | NC_006983 |
| Cryptosporidium parvum genome 5 | eukaryote | NC_006984 |
| Cryptosporidium parvum genome 6 | eukaryote | NC_006985 |
| Cryptosporidium parvum genome 7 | eukaryote | NC_006986 |
| Cryptosporidium parvum genome 8 | eukaryote | NC_006987 |
| Drosophila melanogaster genome 2L | eukaryote | NT_033779 |
| Drosophila melanogaster genome 2R | eukaryote | NT_033778 |
| Drosophila melanogaster genome 3L | eukaryote | NT_037436 |
| Drosophila melanogaster genome 3R | eukaryote | NT_033777 |
| Drosophila melanogaster genome 4 | eukaryote | NC_004353 |
| Drosophila melanogaster genome X | eukaryote | NC_004354 |
| Leishmania major strain Friedlin genome 27 | eukaryote | NC_007268 |
| Leishmania major strain Friedlin genome 29 | eukaryote | NC_007270 |
| Leishmania major strain Friedlin genome 4 | eukaryote | NC_007245 |
| Saccharomyces cerevisiae genome I | eukaryote | NC_001133 |
| Saccharomyces cerevisiae genome II | eukaryote | NC_001134 |
| Saccharomyces cerevisiae genome III | eukaryote | NC_001135 |
| Saccharomyces cerevisiae genome IV | eukaryote | NC_001136 |
| Saccharomyces cerevisiae genome IX | eukaryote | NC_001141 |
| Saccharomyces cerevisiae genome V | eukaryote | NC_001137 |
| Saccharomyces cerevisiae genome VI | eukaryote | NC_001138 |
| Saccharomyces cerevisiae genome VII | eukaryote | NC_001139 |
| Saccharomyces cerevisiae genome VIII | eukaryote | NC_001140 |
| Saccharomyces cerevisiae genome X | eukaryote | NC_001142 |
| Saccharomyces cerevisiae genome XI | eukaryote | NC_001143 |
| Saccharomyces cerevisiae genome XII | eukaryote | NC_001144 |
| Saccharomyces cerevisiae genome XIII | eukaryote | NC_001145 |
| Saccharomyces cerevisiae genome XIV | eukaryote | NC_001146 |
| Saccharomyces cerevisiae genome XV | eukaryote | NC_001147 |
| Saccharomyces cerevisiae genome XVI | eukaryote | NC_001148 |
| Trypanosoma brucei TREU927 genome 1 | eukaryote | NC_007334 |
| Trypanosoma brucei TREU927 genome 10 | eukaryote | NC_007283 |
| Trypanosoma brucei TREU927 genome 11 scaffold 1 | eukaryote | NT_165288 |
| Trypanosoma brucei TREU927 genome 2 | eukaryote | NC_005063 |
| Trypanosoma brucei TREU927 genome 3 | eukaryote | NC_007276 |
| Trypanosoma brucei TREU927 genome 4 | eukaryote | NC_007277 |
| Trypanosoma brucei TREU927 genome 5 | eukaryote | NC_007278 |
| Trypanosoma brucei TREU927 genome 6 | eukaryote | NC_007279 |
| Trypanosoma brucei TREU927 genome 7 | eukaryote | NC_007280 |
| Trypanosoma brucei TREU927 genome 8 | eukaryote | NC_007281 |
| Trypanosoma brucei TREU927 genome 9 | eukaryote | NC_007282 |
| Trypansomabrucei TREU927 genome 11 scaffold 2 | eukaryote | NT_165287 |
| Acanthamoeba polyphaga mimivirus | virus | NC_006450 |
| Adoxophyes honmai nucleopolyhedrovirus | virus | NC_004690 |
| Aeromonas phage 31 | virus | NC_007022 |
| African swine fever virus | virus | NC_001659 |
| Agrotis segetum granulovirus | virus | NC_005839 |
| Alcelaphine herpesvirus 1 | virus | NC_002531 |
| Ambystoma tigrinum virus | virus | NC_005832 |
| Amsacta moorei entomopoxvirus | virus | NC_002520 |
| Ateline herpesvirus 3 | virus | NC_001987 |
| Autographa californica nucleopolyhedrovirus | virus | NC_001623 |
| bacteriophage 44 RR2.8t | virus | NC_005135 |
| bacteriophage Aeh1 | virus | NC_005260 |
| bacteriophage G1 | virus | NC_007066 |
| bacteriophage KVP40 | virus | NC_005083 |
| bacteriophage RM378 | virus | NC_004735 |
| bacteriophage SPBc2 | virus | NC_001884 |
| bacteriophage S-PM2 virion | virus | NC_006820 |
| bacteriophage T5 virion | virus | NC_005859 |
| Bombyx mori nucleopolyhedrovirus | virus | NC_001962 |
| Bovine herpesvirus 1 | virus | NC_001847 |
| Bovine herpesvirus 4 | virus | NC_002665 |
| Bovine herpesvirus 5 | virus | NC_005261 |
| Bovine papular stomatitis virus | virus | NC_005337 |
| Callitrichine herpesvirus 3 | virus | NC_004367 |
| Camelpoxvirus | virus | NC_003391 |
| Canarypoxvirus | virus | NC_005309 |
| Cercopithecine herpesvirus 1 | virus | NC_004812 |
| Cercopithecine herpesvirus 15 | virus | NC_006146 |
| Cercopithecine herpesvirus 17 | virus | NC_003401 |
| Cercopithecine herpesvirus 2 | virus | NC_006560 |
| Cercopithecine herpesvirus 7 | virus | NC_002686 |
| Cercopithecine herpesvirus 8 | virus | NC_006150 |
| Chimpanzee cytomegalovirus | virus | NC_003521 |
| Choristoneura fumiferana defective nucleopolyhedrovirus | virus | NC_005137 |
| Choristoneura fumiferana MNPV | virus | NC_004778 |
| Chrysodeixis chalcites nucleopolyhedrovirus | virus | NC_007151 |
| Cowpox virus | virus | NC_003663 |
| Cryptophlebia leucotreta granulovirus | virus | NC_005068 |
| Culex nigripalpus baculovirus | virus | NC_003084 |
| Cyanophage P-SSM2 | virus | NC_006883 |
| Cyanophage P-SSM4 | virus | NC_006884 |
| Cydia pomonella granulovirus | virus | NC_002816 |
| Ectocarpus siliculosus virus | virus | NC_002687 |
| Ectromelia virus | virus | NC_004105 |
| Emiliania huxleyi virus 86 | virus | NC_007346 |
| Enterobacteria phage RB43 | virus | NC_007023 |
| Enterobacteria phage RB49 | virus | NC_005066 |
| Enterobacteria phage RB69 | virus | NC_004928 |
| Enterobacteria phage T4 | virus | NC_000866 |
| Epiphyas postvittana nucleopolyhedrovirus | virus | NC_003083 |
| Equid herpesvirus 1 | virus | NC_001491 |
| Equid herpesvirus 2 | virus | NC_001650 |
| Equid herpesvirus 4 | virus | NC_001844 |
| Fowlpox virus | virus | NC_002188 |
| Frogvirus 3 | virus | NC_005946 |
| Gallid herpesvirus 1 | virus | NC_006623 |
| Gallid herpesvirus 2 | virus | NC_002229 |
| Gallid herpesvirus 3 | virus | NC_002577 |
| Goatpox virus | virus | NC_004003 |
| Helicoverpa armigera nuclearpolyhedrosisvirus | virus | NC_003094 |
| Helicoverpa zea single nucleocapsid nucleopolyhedrovirus | virus | NC_003349 |
| Heliocoverpa armigera nucleopolyhedrovirus G4 | virus | NC_002654 |
| Heliothis zea virus 1 | virus | NC_004156 |
| Human herpesvirus 1 | virus | NC_001806 |
| Human herpesvirus 2 | virus | NC_001798 |
| Human herpesvirus 3 (strain Dumas) | virus | NC_001348 |
| Human herpesvirus 4 | virus | NC_001345 |
| Human herpesvirus 5 (laboratory strain AD169) | virus | NC_001347 |
| Human herpesvirus 5(wildtype strain Merlin) | virus | NC_006273 |
| Human herpesvirus 6 | virus | NC_001664 |
| Human herpesvirus 6B | virus | NC_000898 |
| Human herpesvirus 7 | virus | NC_001716 |
| Human herpesvirus 8, genome | virus | NC_003409 |
| Ictalurid herpesvirus 1 | virus | NC_001493 |
| Infectious spleen and kidney necrosis virus | virus | NC_003494 |
| Invertebrate iridescent virus 6 | virus | NC_003038 |
| Lactobacillus plantarum bacteriophage LP65virion | virus | NC_006565 |
| Lumpy skin disease virus | virus | NC_003027 |
| Lymantria dispar nucleopolyhedrovirus | virus | NC_001973 |
| Lymphocystis disease virus 1 | virus | NC_001824 |
| Lymphocystis disease virus-isolate China | virus | NC_005902 |
| Macaca fuscata rhadinovirus | virus | NC_007016 |
| Mamestra configurata NPV-A | virus | NC_003529 |
| Mamestra configurata nucleopolyhedrovirus B | virus | NC_004117 |
| Melanoplus sanguinipes entomopoxvirus | virus | NC_001993 |
| Meleagrid herpesvirus 1 | virus | NC_002641 |
| Molluscum contagiosum virus | virus | NC_001731 |
| Monkeypox virus | virus | NC_003310 |
| Muledeerpox virus | virus | NC_006966 |
| Murid herpesvirus 1 | virus | NC_004065 |
| Murid herpesvirus 2 | virus | NC_002512 |
| Murid herpesvirus 4 | virus | NC_001826 |
| Mycobacteriophage Bxz1 virion | virus | NC_004687 |
| Mycobacteriophage Omega virion | virus | NC_004688 |
| Myxoma virus | virus | NC_001132 |
| Orf virus | virus | NC_005336 |
| Orgyia pseudotsugata multicapsid nucleopolyhedrovirus | virus | NC_001875 |
| Ostreid herpesvirus 1 | virus | NC_005881 |
| Paramecium bursaria Chlorellavirus 1 | virus | NC_000852 |
| Phthorimaea operculella granulovirus | virus | NC_004062 |
| Plutella xylostella granulovirus | virus | NC_002593 |
| Psittacid herpesvirus 1 | virus | NC_005264 |
| Psuedomonas phage phiKZ | virus | NC_004629 |
| Rabbit fibroma virus | virus | NC_001266 |
| Rachiplusia ou multiple nucleopolyhedrovirus | virus | NC_004323 |
| Saimiriine herpesvirus 2 | virus | NC_001350 |
| Sheeppox virus | virus | NC_004002 |
| Shrimp whitespot syndrome virus | virus | NC_003225 |
| Singapore grouper iridovirus | virus | NC_006549 |
| Spodoptera exigua nucleopolyhedrovirus | virus | NC_002169 |
| Spodoptera litura nucleopolyhedrovirus | virus | NC_003102 |
| Staphylococcus phage K virion | virus | NC_005880 |
| Staphylococcus phage Twort | virus | NC_007021 |
| Suid herpesvirus 1 | virus | NC_006151 |
| Swinepox virus | virus | NC_003389 |
| Trichoplusia ni SNPV virus | virus | NC_007383 |
| Tupaia herpesvirus | virus | NC_002794 |
| Vaccinia virus | virus | NC_001559 |
| Variola virus | virus | NC_001611 |
| Xestiac-nigrum granulovirus | virus | NC_002331 |
| Yaba monkey tumorvirus | virus | NC_005179 |
| Yaba-like disease virus | virus | NC_002642 |
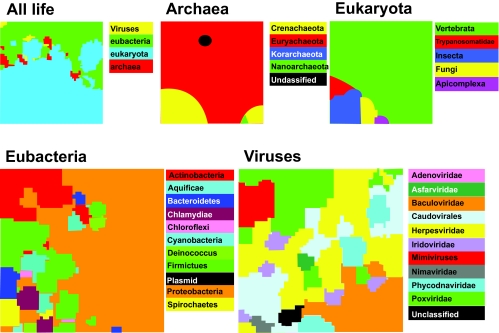
Figure 1.
Dominance maps for GS-2 applied to a 50 ×50 SOM over 100 iterations. The eubacterial and viral SOMs are shown at a larger scale owing to their greater detail. Dominance areas are color coded.
Table 3.
Viral genomes used for the SOM covering a wide range of viruses, shown in Figure 2. 579 viral genomes have at least 10kb of sequence. This is approximately 35% of all fully sequenced viral genomes available at the time of the analysis.
| Name | Accession |
|---|---|
| Bovine adenovirus 2 | AC_000001 |
| Bovine adenovirus 3 | AC_000002 |
| Canine adenovirus type 1 | AC_000003 |
| Duck adenovirus 1 | AC_000004 |
| Human adenovirus type 12 | AC_000005 |
| Human adenovirus type 17 | AC_000006 |
| Human adenovirus type 2 | AC_000007 |
| Human adenovirus type 5 | AC_000008 |
| Porcine adenovirus 5 | AC_000009 |
| Simian adenovirus 21 | AC_000010 |
| Simian adenovirus 25 | AC_000011 |
| Murine adenovirus 1 | AC_000012 |
| Fowl adenovirus 9 | AC_000013 |
| Fowl adenovirus 1 | AC_000014 |
| Human adenovirus type 11 | AC_000015 |
| Turkey adenovirus 3 | AC_000016 |
| Human adenovirus type 1 | AC_000017 |
| Human adenovirus type 7 | AC_000018 |
| Human adenovirus type 35 | AC_000019 |
| Canine adenovirus type 2 | AC_000020 |
| Paramecium bursaria Chlorella virus 1 | NC_000852 |
| Viral hemorrhagic septicemia virus | NC_000855 |
| Enterobacteria phage T4 | NC_000866 |
| Alteromonas phage PM2 | NC_000867 |
| Streptococcus thermophilus bacteriophage Sfi19 | NC_000871 |
| Streptococcus thermophilus bacteriophage Sfi21 | NC_000872 |
| Lactobacillus bacteriophage phi adh | NC_000896 |
| Human herpesvirus 6B | NC_000898 |
| Fowl adenovirus D | NC_000899 |
| Bacteriophage VT2-Sa | NC_000902 |
| Snakehead rhabdovirus | NC_000903 |
| Bacteriophage 933W | NC_000924 |
| Enterobacteria phage Mu | NC_000929 |
| Acyrthosiphon pisum bacteriophage APSE-1 | NC_000935 |
| Murine adenovirus A | NC_000942 |
| Murray Valley encephalitis virus | NC_000943 |
| Myxomavirus | NC_001132 |
| Rabbit fibromavirus | NC_001266 |
| Bacteriophage phi YeO3-12 | NC_001271 |
| Enterobacteria phage 186 | NC_001317 |
| Mycobacterium phage L5 | NC_001335 |
| Sulfolobus spindle-shaped virus 1 | NC_001338 |
| Human herpesvirus 4 | NC_001345 |
| Human herpesvirus 5 (laboratory strain AD169) | NC_001347 |
| Human herpesvirus 3 (strain Dumas) | NC_001348 |
| Saimiriine herpesvirus 2 | NC_001350 |
| Simian foamy virus | NC_001364 |
| Human adenovirus C | NC_001405 |
| Bacteriophage lambda | NC_001416 |
| Enterobacteria phage PRD1 | NC_001421 |
| Bacillus phage PZA | NC_001423 |
| Japanese encephalitis virus | NC_001437 |
| Achole plasmaphage L2 | NC_001447 |
| Venezuelan equine encephalitis virus | NC_001449 |
| Avian infectious bronchitis virus | NC_001451 |
| Human adenovirus F | NC_001454 |
| Human adenovirus A | NC_001460 |
| Bovine viral diarrheavirus 1 | NC_001461 |
| Dengue virus type 2 | NC_001474 |
| Dengue virus type 3 | NC_001475 |
| Dengue virus type 1 | NC_001477 |
| Equid herpesvirus 1 | NC_001491 |
| Cryphonectria hypovirus 1 | NC_001492 |
| Ictalurid herpesvirus 1 | NC_001493 |
| Measles virus | NC_001498 |
| O’nyong-nyong virus | NC_001512 |
| Rabies virus | NC_001542 |
| Ross River virus | NC_001544 |
| Sindbis virus | NC_001547 |
| Sendai virus | NC_001552 |
| Vaccinia virus | NC_001559 |
| Vesicular stomatitis Indiana virus | NC_001560 |
| West Nile virus | NC_001563 |
| Cell fusing agent virus | NC_001564 |
| Beet yellows virus | NC_001598 |
| Enterobacteria phage T7 | NC_001604 |
| Lake Victoria marburg virus | NC_001608 |
| Bacteriophage P4 | NC_001609 |
| Variola virus | NC_001611 |
| Sonchus yellow net virus | NC_001615 |
| Autographa californica nucleopolyhedrovirus | NC_001623 |
| Rice tungro spherical virus | NC_001632 |
| Equid herpesvirus 2 | NC_001650 |
| Infectious hematopoietic necrosis virus | NC_001652 |
| African swine fever virus | NC_001659 |
| Citrus tristeza virus | NC_001661 |
| Human herpesvirus 6 | NC_001664 |
| Tick-borne encephalitis virus | NC_001672 |
| Haemophilus phage HP1 | NC_001697 |
| Lactococcus phage c2 | NC_001706 |
| Human herpesvirus 7 | NC_001716 |
| Fowl adenovirus A | NC_001720 |
| Human immunodeficiency virus 2 | NC_001722 |
| Snakehead retrovirus | NC_001724 |
| Molluscum contagiosum virus | NC_001731 |
| Canine adenovirus | NC_001734 |
| Human foamy virus | NC_001736 |
| Human respiratory syncytial virus | NC_001781 |
| Papaya ringspot virus | NC_001785 |
| Barmah Forest virus | NC_001786 |
| Human spuma retrovirus | NC_001795 |
| Human parainfluenza virus 3 | NC_001796 |
| Human herpesvirus 2 | NC_001798 |
| Respiratory syncytial virus | NC_001803 |
| Human herpesvirus 1 | NC_001806 |
| Louping ill virus | NC_001809 |
| Duck adenovirus A | NC_001813 |
| Lymphocystis disease virus 1 | NC_001824 |
| Streptococcus phage Cp-1 | NC_001825 |
| Murid herpesvirus 4 | NC_001826 |
| Bovine foamy virus | NC_001831 |
| Bacteriophage sk1 | NC_001835 |
| Little cherry virus 1 | NC_001836 |
| Sweet potato feathery mottle virus | NC_001841 |
| Equid herpesvirus 4 | NC_001844 |
| Murine hepatitis virus strain A59 | NC_001846 |
| Bovine herpesvirus 1 | NC_001847 |
| Walleye dermal sarcoma virus | NC_001867 |
| Simian-Human immunodeficiency virus | NC_001870 |
| Feline foamy virus | NC_001871 |
| Rhopalosiphum padi virus | NC_001874 |
| Orgyia pseudotsugata nucleopolyhedrovirus | NC_001875 |
| Bovine adenovirus B | NC_001876 |
| Bacteriophage SPBc2 | NC_001884 |
| Enterobacteria phage P2 | NC_001895 |
| Mycobacteriophage D29 | NC_001900 |
| Bacteriophage N15 | NC_001901 |
| Methanobacterium phage psiM2 | NC_001902 |
| Hendra virus | NC_001906 |
| Bacteriophage bIL170 | NC_001909 |
| Canine distemper virus | NC_001921 |
| Igbo Ora virus | NC_001924 |
| Mycoplasma arthritidis bacteriophage MAV1 | NC_001942 |
| Hemorrhagic enteritis virus | NC_001958 |
| Porcine reproductive and respiratory syndrome virus | NC_001961 |
| Bombyx mori nucleopolyhedrovirus | NC_001962 |
| Lymantria dispar nucleopolyhedrovirus | NC_001973 |
| Bacteriophage phi-C31 | NC_001978 |
| Ateline herpesvirus 3 | NC_001987 |
| Bovine respiratory syncytial virus | NC_001989 |
| Melanoplus sanguinipes entomopox virus | NC_001993 |
| Yellow fever virus | NC_002031 |
| Bovine viral diarrhea virus genotype 2 | NC_002032 |
| Human adenovirus D | NC_002067 |
| Streptococcus thermophilus bacteriophage DT1 | NC_002072 |
| Bovine parainfluenza virus3 | NC_002161 |
| Enterobacteria phage HK022 | NC_002166 |
| Bacteriophage HK97 | NC_002167 |
| Spodoptera exigua nucleopolyhedrovirus | NC_002169 |
| Streptococcus thermophilus bacteriophage 7201 | NC_002185 |
| Fowlpox virus | NC_002188 |
| Tupaia paramyxovirus | NC_002199 |
| Mumps virus | NC_002200 |
| Equine foamy virus | NC_002201 |
| Streptococcus thermophilus bacteriophage Sfi11 | NC_002214 |
| Gallid herpesvirus 2 | NC_002229 |
| Northern cereal mosaic virus | NC_002251 |
| Transmissible gastroenteritis virus | NC_002306 |
| Staphylococcus aureus bacteriophage PVL | NC_002321 |
| Xestiac-nigrum granulovirus | NC_002331 |
| Enterobacteria phage P22 | NC_002371 |
| Pseudomonas phage D3 | NC_002484 |
| Staphylococcus aureus prophage phiPV83 | NC_002486 |
| Frog adenovirus | NC_002501 |
| Murid herpesvirus 2 | NC_002512 |
| Ovine adenovirus A | NC_002513 |
| Mycoplasma virus P1 | NC_002515 |
| Roseophage SIO1 | NC_002519 |
| Amsacta moorei entomopox virus | NC_002520 |
| Bovine ephemeral fever virus | NC_002526 |
| Alcelaphine herpesvirus 1 | NC_002531 |
| Equine arteritis virus | NC_002532 |
| Lactate dehydrogenase-elevating virus | NC_002534 |
| Zaire ebola virus | NC_002549 |
| Gallid herpesvirus 3 | NC_002577 |
| Plutella xylostella granulovirus | NC_002593 |
| Newcastle disease virus | NC_002617 |
| Methanothermobacter wolfeii prophage psiM100 | NC_002628 |
| Dengue virus type 4 | NC_002640 |
| Meleagrid herpesvirus 1 | NC_002641 |
| Yaba-like disease virus | NC_002642 |
| Human coronavirus 229E | NC_002645 |
| Bacillus phage GA-1 | NC_002649 |
| Heliocoverpa armigera nucleopolyhedrovirus G4 | NC_002654 |
| Mycobacteriophage Bxb1 | NC_002656 |
| Classical swine fever virus | NC_002657 |
| Staphylococcus aureus temperate phage phi SLT | NC_002661 |
| Bovine herpesvirus 4 | NC_002665 |
| Bacteriophage bIL285 | NC_002666 |
| Bacteriophage bIL286 | NC_002667 |
| Bacteriophage bIL309 | NC_002668 |
| Bacteriophage bIL310 | NC_002669 |
| Bacteriophage bIL311 | NC_002670 |
| Bacteriophage bIL312 | NC_002671 |
| Bovine adenovirus D | NC_002685 |
| Cercopithecine herpesvirus 7 | NC_002686 |
| Ectocarpus siliculosus virus | NC_002687 |
| Porcine adenovirus C | NC_002702 |
| Bacteriophage Tuc2009 | NC_002703 |
| Nipah virus | NC_002728 |
| Bacteriophage HK620 | NC_002730 |
| Lactococcus lactis bacteriophage TP901-1 | NC_002747 |
| Tupaia herpesvirus | NC_002794 |
| Lactococcus phage BK5-T | NC_002796 |
| Spring viremia of carp virus | NC_002803 |
| Cydia pomonella granulovirus | NC_002816 |
| Taura syndrome virus | NC_003005 |
| Lumpy skin disease virus | NC_003027 |
| Invertebrate iridescent virus 6 | NC_003038 |
| Avian paramyxovirus 6 | NC_003043 |
| Bovine coronavirus | NC_003045 |
| Streptococcus pneumoniae bacteriophage MM1 | NC_003050 |
| Epiphyas postvittana nucleopolyhedrovirus | NC_003083 |
| Culex nigripalpus baculovirus | NC_003084 |
| Bacteriophage Mx8 | NC_003085 |
| Simian hemorrhagic fever virus | NC_003092 |
| Helicoverpa armigera nuclearpolyhedrosis virus | NC_003094 |
| Spodopteralitura nucleopolyhedrovirus | NC_003102 |
| Temperate phage PhiNIH1.1 | NC_003157 |
| Sulfolobus islandicus filamentous virus | NC_003214 |
| Semliki forest virus | NC_003215 |
| Bacteriophage A118 | NC_003216 |
| Shrimp white spot syndrome virus | NC_003225 |
| Australian bat lyssa virus | NC_003243 |
| Human adenovirus E | NC_003266 |
| Bacteriophage phiCTX | NC_003278 |
| Bacteriophage phiETA | NC_003288 |
| Bacteriophage PSA | NC_003291 |
| Bacteriophage T3 | NC_003298 |
| Bacteriophage phiE125 | NC_003309 |
| Monkeypox virus | NC_003310 |
| Bacteriophage K139 | NC_003313 |
| Haemophilus phage HP2 | NC_003315 |
| Sinorhizobium meliloti phage PBC5 | NC_003324 |
| Halovirus HF2 | NC_003345 |
| Helicoverpa zea nucleopolyhedrovirus | NC_003349 |
| Bacteriophage P27 | NC_003356 |
| Mycobacteriophage TM4 | NC_003387 |
| Swinepox virus | NC_003389 |
| Cyanophage P60 | NC_003390 |
| Camelpox virus | NC_003391 |
| Cercopithecine herpesvirus 17 | NC_003401 |
| Human herpesvirus 8 | NC_003409 |
| Mayaro virus | NC_003417 |
| Sleeping disease virus | NC_003433 |
| Porcine epidemic diarrhea virus | NC_003436 |
| Human parainfluenza virus 2 | NC_003443 |
| Shigella flexneri bacteriophage V | NC_003444 |
| Human parainfluenza virus 1 strain Washington/1964 | NC_003461 |
| Infectious spleen and kidney necrosis virus | NC_003494 |
| Chimpanzee cytomegalovirus | NC_003521 |
| Bacteriophage phi3626 | NC_003524 |
| Stx2 converting bacteriophage I | NC_003525 |
| Mamestra configurata NPV-A | NC_003529 |
| Cryphonectria hypovirus | NC_003534 |
| Dasheen mosaic virus | NC_003537 |
| Lettuce mosaic virus | NC_003605 |
| Maize chlorotic dwarf virus | NC_003626 |
| Modoc virus | NC_003635 |
| Cowpox virus | NC_003663 |
| Rio Bravo virus | NC_003675 |
| Apoi virus | NC_003676 |
| Pestivirus Reindeer-1 | NC_003677 |
| Pestivirus Giraffe-1 | NC_003678 |
| Border disease virus 1 | NC_003679 |
| Powassan virus | NC_003687 |
| Langat virus | NC_003690 |
| Rice yellow stunt virus | NC_003746 |
| Acyrthosiphon pisum virus | NC_003780 |
| Sweet potato mild mottle virus | NC_003797 |
| Eastern equine encephalitis virus | NC_003899 |
| Aura virus | NC_003900 |
| Vibriophage VpV262 | NC_003907 |
| Western equine encephalomyelitis virus | NC_003908 |
| Salmon pancreas disease virus | NC_003930 |
| Tamana bat virus | NC_003996 |
| Human adenovirus B | NC_004001 |
| Sheeppox virus | NC_004002 |
| Goatpox virus | NC_004003 |
| Leek yellow stripe virus | NC_004011 |
| Ovine adenovirus 7 | NC_004037 |
| Phthorimaea operculella granulovirus | NC_004062 |
| Murid herpesvirus 1 | NC_004065 |
| Lactococcus lactisbacteriophageul36 | NC_004066 |
| Tiomanvirus | NC_004074 |
| VirusPhiCh1 | NC_004084 |
| Sulfolobus islandicus rod-shaped virus 2 | NC_004086 |
| Sulfolobus islandicus rod-shaped virus 1 | NC_004087 |
| Ectromelia virus | NC_004105 |
| Lactobacillus casei bacteriophage A2 | NC_004112 |
| Mamestra configurata nucleopolyhedrovirus B | NC_004117 |
| Montana myotis leukoencephalitis virus | NC_004119 |
| Human metapneumovirus | NC_004148 |
| Heliothis zea virus 1 | NC_004156 |
| Dugbe virus segment L | NC_004159 |
| Reston Ebola virus | NC_004161 |
| Chikungunya virus | NC_004162 |
| Bacteriophage B103 | NC_004165 |
| Bacteriophage SPP1 | NC_004166 |
| Bacteriophage phi-105 | NC_004167 |
| Bacteriophage r1t | NC_004302 |
| Streptococcus thermophilus bacteriophage O1205 | NC_004303 |
| Bacteriophage phig1e | NC_004305 |
| Salmonella typhimurium phage ST64B | NC_004313 |
| Rachiplusia ou multiple nucleopolyhedrovirus | NC_004323 |
| Burkholderia cepacia phage Bcep781 | NC_004333 |
| Salmonella typhimurium bacteriophage ST64T | NC_004348 |
| Alkhurma virus | NC_004355 |
| Callitrichine herpesvirus 3 | NC_004367 |
| Treeshrew adenovirus | NC_004453 |
| Vibrio harveyi bacteriophage VHML | NC_004456 |
| Bacteriophage IN93 | NC_004462 |
| Pseudomonas aeruginosa phage PaP3 | NC_004466 |
| Streptococcus pyogenes phage 315.1 | NC_004584 |
| Streptococcus pyogenes phage 315.2 | NC_004585 |
| Streptococcus pyogenes phage 315.3 | NC_004586 |
| Streptococcus pyogenes phage 315.4 | NC_004587 |
| Streptococcus pyogenes phage 315.5 | NC_004588 |
| Streptococcus pyogenes phage 315.6 | NC_004589 |
| Staphylococcus aureus phage phi11 | NC_004615 |
| Staphylococcus aureus phage phi12 | NC_004616 |
| Staphylococcus aureus phage phi13 | NC_004617 |
| Pseudomonas phage phiKZ | NC_004629 |
| Bacteriophage phi-BT1 | NC_004664 |
| Pseudomonas phage gh-1 | NC_004665 |
| Grapevine leaf roll-associated virus 3 | NC_004667 |
| Staphylococcus phage 44AHJD | NC_004678 |
| Staphylococcus aureus phage phiP68 | NC_004679 |
| Mycobacteriophage Che8 | NC_004680 |
| Mycobacteriophage CJW1 | NC_004681 |
| Mycobacteriophage Bxz2 | NC_004682 |
| Mycobacteriophage Che9c | NC_004683 |
| Mycobacteriophage Rosebush | NC_004684 |
| Mycobacteriophage Corndog | NC_004685 |
| Mycobacteriophage Che9d | NC_004686 |
| Mycobacteriophage Bxz1 | NC_004687 |
| Mycobacteriophage Omega | NC_004688 |
| Mycobacteriophage Barnyard | NC_004689 |
| Adoxophyes honmai nucleopolyhedrovirus | NC_004690 |
| SARS coronavirus | NC_004718 |
| Grapevine rootstock stem lesion associated virus | NC_004724 |
| Bacteriophage RM378 | NC_004735 |
| Staphylococcus phage phiN315 | NC_004740 |
| Bacteriophage L-413C | NC_004745 |
| Lactococcus phage P335 | NC_004746 |
| Enterobacteria phage epsilon15 | NC_004775 |
| Yersinia pestis phage phiA1122 | NC_004777 |
| Choristoneura fumiferana MNPV | NC_004778 |
| Cercopithecine herpesvirus 1 | NC_004812 |
| Phage phi4795 | NC_004813 |
| Streptococcus phage C1 | NC_004814 |
| Bacteriophage phBC6A51 | NC_004820 |
| Bacteriophage phBC6A52 | NC_004821 |
| Bacteriophage Aaphi23 | NC_004827 |
| Deformed wing virus | NC_004830 |
| Enterobacteria phage SP6 | NC_004831 |
| Xanthomonas oryzae bacteriophage Xp10 | NC_004902 |
| Stx1 converting bacteriophage | NC_004913 |
| Stx2 converting bacteriophage II | NC_004914 |
| Halovirus HF1 | NC_004927 |
| Enterobacteria phage RB69 | NC_004928 |
| Streptococcus mitis phage SM1 | NC_004996 |
| Papaya leaf-distortion mosaic potyvirus | NC_005028 |
| Onion yellow dwarf virus | NC_005029 |
| Goose paramyxovirus SF02 | NC_005036 |
| Adoxophyes orana granulovirus | NC_005038 |
| Yokose virus | NC_005039 |
| Bacteriophage phiKMV | NC_005045 |
| Bacteriophage WPhi | NC_005056 |
| Omsk hemorrhagic fever virus | NC_005062 |
| Kamiti River virus | NC_005064 |
| Little cherry virus 2 | NC_005065 |
| Enterobacteria phage RB49 | NC_005066 |
| Cryptophlebia leucotreta granulovirus | NC_005068 |
| Bacteriophage PY54 | NC_005069 |
| Bacteriophage KVP40 | NC_005083 |
| Fer-de-lance virus | NC_005084 |
| Burkholderia cepacia phage BcepNazgul | NC_005091 |
| Hirame rhabdovirus | NC_005093 |
| Bacteriophage 44RR2.8t | NC_005135 |
| Choristoneura fumiferana nucleopolyhedrovirus | NC_005137 |
| Human coronavirus OC43 | NC_005147 |
| Bacteriophage D3112 | NC_005178 |
| Yaba monkey tumor virus | NC_005179 |
| Bacillus thuringiensis bacteriophage Bam35c | NC_005258 |
| Mycobacteriophage PG1 | NC_005259 |
| Bacteriophage Aeh1 | NC_005260 |
| Bovine herpesvirus 5 | NC_005261 |
| Burkholderia cepacia phage Bcep22 | NC_005262 |
| Burkholderia cenocepacia phage Bcep1 | NC_005263 |
| Psittacid herpesvirus 1 | NC_005264 |
| Sulfolobus spindle-shaped virus 2 | NC_005265 |
| Bacteriophage Felix01 | NC_005282 |
| Dolphin morbillivirus | NC_005283 |
| Bacteriophage phi1026b | NC_005284 |
| Bacteriophage EJ-1 | NC_005294 |
| Crimean-Congo hemorrhagic fever virus segment L | NC_005301 |
| Canarypox virus | NC_005309 |
| Orfvirus | NC_005336 |
| Bovine papularstomatitis virus | NC_005337 |
| Mossman virus | NC_005339 |
| Bacteriophage PSP3 | NC_005340 |
| Burkholderia cepacia phage Bcep43 | NC_005342 |
| Enterobacteria phage Sf6 | NC_005344 |
| Bacteriophage VWB | NC_005345 |
| Lactobacillus johnsonii prophage Lj928 | NC_005354 |
| Lactobacillus johnsonii prophage Lj965 | NC_005355 |
| Bacteriophage 77 | NC_005356 |
| Bordetella phage BPP-1 | NC_005357 |
| Sulfolobus spindle-shaped virus Ragged Hills | NC_005360 |
| Sulfolobus spindle-shaped virus Kamchatka-1 | NC_005361 |
| Bordetella phage BMP-1 | NC_005808 |
| Bordetella phage BIP-1 | NC_005809 |
| Bacteriophage phiLC3 | NC_005822 |
| Acidianus filamentus virus 1 | NC_005830 |
| Human coronavirus NL63 | NC_005831 |
| Ambystoma tigrinum virus | NC_005832 |
| Enterobacteria phage T1 | NC_005833 |
| Agrotis segetum granulovirus | NC_005839 |
| Salmonella typhimurium bacteriophage ST104 | NC_005841 |
| Enterobacteria phage P1 | NC_005856 |
| Bacteriophage phiKO2 | NC_005857 |
| Bacteriophage T5 | NC_005859 |
| Porcine adenovirus A | NC_005869 |
| Pyrobaculum spherical virus | NC_005872 |
| Kakugo virus | NC_005876 |
| Vibriophage VP2 | NC_005879 |
| Staphylococcus phage K | NC_005880 |
| Ostreid herpesvirus 1 | NC_005881 |
| Burkholderia cenocepacia phage BcepMu | NC_005882 |
| Pseudomonas aeruginosa bacteriophage PaP2 | NC_005884 |
| Actinoplanes phage phiAsp2 | NC_005885 |
| Burkholderia cenocepacia phage BcepB1A | NC_005886 |
| Burkholderia cepacia complex phage BcepC6B | NC_005887 |
| Vibriophage VP5 | NC_005891 |
| Sulfolobus turreted icosahedral virus | NC_005892 |
| Bacteriophage phiAT3 | NC_005893 |
| Lymphocystis disease virus-isolate China | NC_005902 |
| Neodiprion sertifer nucleopolyhedrovirus | NC_005905 |
| Neodiprion lecontei NPV | NC_005906 |
| Frog virus 3 | NC_005946 |
| Bacteriophage phiMFV1 | NC_005964 |
| Maize fine streak virus | NC_005974 |
| Maize mosaic virus | NC_005975 |
| Simian adenovirus A | NC_006144 |
| Cercopithecine herpesvirus 15 | NC_006146 |
| Cercopithecine herpesvirus 8 | NC_006150 |
| Suid herpesvirus 1 | NC_006151 |
| Watermelon mosaic virus | NC_006262 |
| Sulfolobus tengchongensis spindle-shaped virus STSV1 | NC_006268 |
| Human herpesvirus 5 (wildtype strain Merlin) | NC_006273 |
| Rinderpest virus | NC_006296 |
| Bovine adenovirus A | NC_006324 |
| Bacteriophage 11b | NC_006356 |
| Peste-des-petits-ruminants virus | NC_006383 |
| Simian parainfluenza virus 41 | NC_006428 |
| Mokola virus | NC_006429 |
| Simian parainfluenza virus5 | NC_006430 |
| Sudan ebola virus | NC_006432 |
| Acanthamoeba polyphaga mimivirus | NC_006450 |
| Varroa destructor virus 1 | NC_006494 |
| Bacteriophage B3 | NC_006548 |
| Singapore grouper iridovirus | NC_006549 |
| Usutu virus | NC_006551 |
| Pseudomonas aeruginosa phage F116 | NC_006552 |
| Thermoproteus tenax spherical virus 1 | NC_006556 |
| Bacillus clarkii bacteriophage BCJA1c | NC_006557 |
| Getah virus | NC_006558 |
| Cercopithecine herpesvirus 2 | NC_006560 |
| Lactobacillus plantarum bacteriophage LP65 | NC_006565 |
| Human coronavirus HKU1 | NC_006577 |
| Pneumonia virus of mice J3666 | NC_006579 |
| Gallid herpesvirus 1 | NC_006623 |
| Cotesia congregata virus segment Circle 1 | NC_006633 |
| Cotesia congregata virus segment Circle 2 | NC_006634 |
| Cotesia congregata virus segment Circle 3 | NC_006635 |
| Cotesia congregata virus segment Circle 4 | NC_006636 |
| Cotesia congregata virus segment Circle 5 | NC_006637 |
| Cotesia congregata virus segment Circle 6 | NC_006638 |
| Cotesia congregata virus segment Circle 7 | NC_006639 |
| Cotesia congregata virus segment Circle 9 | NC_006641 |
| Cotesia congregata virus segment Circle 10 | NC_006642 |
| Cotesia congregata virus segment Circle 11 | NC_006643 |
| Cotesia congregata virus segment Circle 12 | NC_006644 |
| Cotesia congregata virus segment Circle 13 | NC_006645 |
| Cotesia congregata virus segment Circle 14 | NC_006646 |
| Cotesia congregata virus segment Circle 17 | NC_006648 |
| Cotesia congregata virus segment Circle 18 | NC_006649 |
| Cotesia congregata virus segment Circle 19 | NC_006650 |
| Cotesia congregata virus segment Circle 20 | NC_006651 |
| Cotesia congregata virus segment Circle 22 | NC_006653 |
| Cotesia congregata virus segment Circle 23 | NC_006654 |
| Cotesia congregata virus segment Circle 25 | NC_006655 |
| Cotesia congregata virus segment Circle 26 | NC_006656 |
| Cotesia congregata virus segment Circle 30 | NC_006657 |
| Cotesia congregata virus segment Circle 31 | NC_006658 |
| Cotesia congregata virus segment Circle 32 | NC_006659 |
| Cotesia congregata virus segment Circle 33 | NC_006660 |
| Cotesia congregata virus segment Circle 35 | NC_006661 |
| Cotesia congregata virus segment Circle 36 | NC_006662 |
| Bacteriophage S-PM2 | NC_006820 |
| Murine hepatitis virus strain JHM | NC_006852 |
| Simian adenovirus 1 | NC_006879 |
| Cyanophage P-SSP7 | NC_006882 |
| Cyanophage P-SSM2 | NC_006883 |
| Cyanophage P-SSM4 | NC_006884 |
| Lactobacillus plantarum bacteriophage phiJL-1 | NC_006936 |
| Bacteriophage phiJL001 | NC_006938 |
| Bacteriophage KS7 | NC_006940 |
| Taro vein chlorosis virus | NC_006942 |
| Mint virus 1 | NC_006944 |
| Bacillus thuringiensis phage GIL16c | NC_006945 |
| Karshi virus | NC_006947 |
| Salmonella typhimurium bacteriophage ES18 | NC_006949 |
| Listonella pelagia phage phiHSIC | NC_006953 |
| Muledeerpox virus | NC_006966 |
| Vaccinia virus | NC_006998 |
| Macaca fuscata rhadinovirus | NC_007016 |
| Streptococcus thermophilus bacteriophage 2972 | NC_007019 |
| Tupaia rhabdovirus | NC_007020 |
| Staphylococcus phage Twort | NC_007021 |
| Aeromonas phage 31 | NC_007022 |
| Enterobacteria phage RB43 | NC_007023 |
| Xanthomonas campestris pv. pelargonii phage Xp15 | NC_007024 |
| Feline coronavirus | NC_007025 |
| Microplitis demolitor bracovirus segment G | NC_007034 |
| Microplitis demolitor bracovirus segment H | NC_007035 |
| Microplitis demolitor bracovirus segment J | NC_007036 |
| Microplitis demolitor bracovirus segment K | NC_007037 |
| Microplitis demolitor bracovirus segment M | NC_007038 |
| Microplitis demolitor bracovirus segment N | NC_007039 |
| Microplitis demolitor bracovirus segment L | NC_007040 |
| Microplitis demolitor bracovirus segment I | NC_007041 |
| Microplitis demolitor bracovirus segment O | NC_007044 |
| Bacteriophage PT1028 | NC_007045 |
| Bacteriophage 66 | NC_007046 |
| Bacteriophage 187 | NC_007047 |
| Bacteriophage 69 | NC_007048 |
| Bacteriophage 53 | NC_007049 |
| Bacteriophage 85 | NC_007050 |
| Bacteriophage 2638A | NC_007051 |
| Bacteriophage 42e | NC_007052 |
| Bacteriophage 3A | NC_007053 |
| Bacteriophage 47 | NC_007054 |
| Bacteriophage 37 | NC_007055 |
| Bacteriophage EW | NC_007056 |
| Bacteriophage 96 | NC_007057 |
| Bacteriophage ROSA | NC_007058 |
| Bacteriophage 71 | NC_007059 |
| Bacteriophage 55 | NC_007060 |
| Bacteriophage 29 | NC_007061 |
| Bacteriophage 52A | NC_007062 |
| Bacteriophage 88 | NC_007063 |
| Bacteriophage 92 | NC_007064 |
| Bacteriophage X2 | NC_007065 |
| Bacteriophage G1 | NC_007066 |
| Phytophthora endorna virus 1 | NC_007069 |
| Burkholderia pseudomallei phage phi52237 | NC_007145 |
| Vibriophage VP4 | NC_007149 |
| Chrysodeixis chalcites nucleopolyhedrovirus | NC_007151 |
| Bacteriophage SH1 | NC_007217 |
| Bacteriophage JK06 | NC_007291 |
| Emiliania huxleyi virus 86 | NC_007346 |
| Trichoplusia ni SNPV virus | NC_007383 |
| Acidianus two-tailed virus | NC_007409 |
| Shallot yellow stripe virus | NC_007433 |
| Breda virus | NC_007447 |
| Grapevine leaf roll-associated virus 2 | NC_007448 |
| Enterobacteria phage L17 | NC_007449 |
| Enterobacteria phage PR3 | NC_007450 |
| Enterobacteria phage PR4 | NC_007451 |
| Enterobacteria phage PR5 | NC_007452 |
| Enterobacteria phage PR772 | NC_007453 |
| J-virus | NC_007454 |
| Coliphage K1F | NC_007456 |
| Bacillus anthracis phage Cherry | NC_007457 |
| Bacillus anthracis phage Gamma | NC_007458 |
| Burkholderia cepacia phage Bcep176 | NC_007497 |
| Bacteriophage Lc-Nu | NC_007501 |
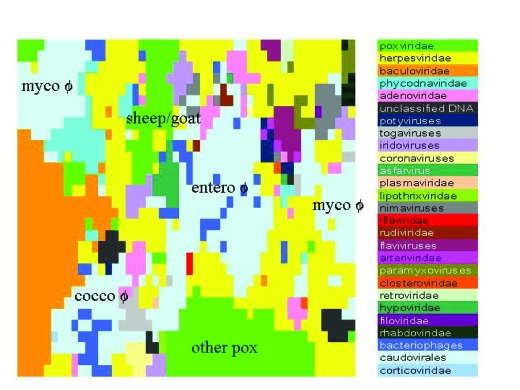
Figure 2.
Dominance map for GS-2 of 10kb fragments of viruses applied to a 50 ×50 SOM over 1000 iterations. The category “Bacteriophages” refers to unclassified phages. Most phages are members of the family Caudovirales. The text added to the dominance map shows the general divisions of the Poxviridae and Caudovirales which form more than one well defined dominance area.
2. Calculation of genome signatures
A Perl script was written to derive raw k-mer counts on FASTA-formatted databases of input sequences, using the SeqWords.pm module from BioPerl (http://www.bioperl.org/Pdoc-mirror/bioperl-live/Bio/Tools/SeqWords.html). The raw k-mer frequencies were then symmetrized, as follows:
where fν and fν-comp are the raw frequencies of a k-mer ν and its complement ν-comp.
Symmetrization means that a sequence and its complement will generate the same answer. The symmetrized frequencies are then corrected for the 1-mer content. For instance for a 2-mer XY, where X and Y can each represent any nucleotide base {A, C, T, G}:
where fsXY is the symmetrized frequency for dimer XY and fsX and fsY are the symmetrized frequencies of its component 1-mers. For a 3-mer XYZ, the correction would be for the 1-mers, X, Y and Z and so on.
The genome signature vector for length k, is thus composed of a series of ratios of observed to expected values of its component k-mers, where the expected values are determined by a zero-order Markov chain (Bernouilli series) model. Genome signatures are therefore not distorted by gross base compositional differences between genomes, which would otherwise be the dominant factor.
3. Self-organizing map
Self-organizing maps (SOMs) were run following Tamayo et al. (1999), using a Perl script. Input consisted of an array of the genome signatures generated as described above. The dimensions of the SOM and the number of iterations in training were variables entered by the user. Euclidean distances were used when comparing vectors.
Once the dimensions of the SOM were set, x columns by y rows, weight vectors initializing each of the xy cells of the SOM were selected at random from the entire set of genome signature data vectors. The SOM is thus initially simply filled with a random subset of the data. Training then commences, for nominated t iterations. At each iteration m, each data vector in turn was compared to each weight vector, and the closest weight vector for each data vector designated the winning weight vector of that data vector in that iteration. Each time a winning weight vector is identified, the winning weight vector, and the weight vectors of cells within a spatial range ℜ on the SOM, were then trained by the data vector as follows.
Each value c in the winning weight vector w is altered, so that its value at iteration, m, becomes at the next iteration m+1:
where wcm – vc represents the difference between the winning weight vector and the data vector for each value c along the vectors. In other words, one simply aligns the data vector and the winning weight vector and subtracts them. Each value of the winning weight vector is then altered to bring it closer to the data vector by a factor of τ, the training effect, which is derived as follows:
τ changes at each iteration of the process, and is the ratio of two other values α and γ.
α is calculated for each iteration m as follows:
where m is the number of the current iteration, and t the number of total iterations requested. There-fore, the number of iterations of the SOM, a parameter chosen at the start of the process, determines the gradient at which α will decrease as the iterations progress.
Whereas α is the same for all cells in the SOM and changes according to the iteration number only, γ is the Euclidean distance on the SOM from the weight vector being trained within range ℛ of the winning weight vector.
τ can therefore be seen to decrease as the SOM progresses, since α decreases, and also to decrease the further one goes away from the winning weight vector, since γ increases.
The range within which weight vectors are trained at each iteration is calculated:
where S is the length or breadth of the SOM, whichever is the smaller. The area of the SOM being trained therefore also shrinks as α decreases with increasing iterations.
Once each data vector has found its winning weight vector and trained it, also training the weight vectors within range ℜ of the winning weight vector, then one iteration is completed. New values of α,τ and ℜ are then calculated, and the second iteration can commence. It can be intuitively grasped that there is a great deal of “churn” in initial iterations of the SOM. When α is close to 1, data vectors will effectively change their winning weight vector to copies of themselves. Only at the limits of the trained area R will the effect be subtler. However, as the number of iterations mounts, α will decrease and each data vector will have a relatively weaker effect on its winning weight vector and even less on those weight vectors in its vicinity. Observation (data not shown) of distribution of a simple data set over a SOM through the iterative process shows that a relatively chaotic process dominates until approximately halfway through the nominated number of iterations, at which point structure rapidly builds in the SOM. The final 10% or so of iterations consist mostly of fine-tuning of the final weight vector values. Training SOMs can also be time consuming, especially for large data sets of high dimensionality vectors trained over large numbers of iterations. The longest run presented here (that in Fig. 2) took in excess of 3 weeks on a single 2.8 GHz Intel processor under a Linux operating system. One of the major motivations of this paper was to define ways to reduce SOM training time without losing accuracy or sensitivity.
After the final iteration, each data vector is again compared to each weight vector and assigned to the closest. This results in partition of each data vector to one cell in the SOM, thus spreading the multi-dimensional data across the two-dimensional surface of the SOM. Conversely, each final weight vector in the SOM is assigned to its closest data vector, the centroid nearest neighbour (cnn). If the data vectors belong to several categories, each cell in the SOM can be colored according to the origin of its cnn, which is then said to dominate that cell in the SOM. This allows the production of color-coded dominance maps indicating the general spread of the data vector set over the SOM. NCBI taxonomic categories were used throughout, except for herpesviruses where the International Committee on the Taxonomy of Viruses (ICTV) usage is followed (Davison 2002; Davison et al. 2005; Fauquet et al. 2005).
4. Availability of scripts
All Perl scripts, for processing genomes, calculating genome signatures, and running SOMs are available on request from the author ( d.gatherer@mrcvu.gla.ac.uk).
Results
1. SOMs on large sequence datasets
The ability of SOMs to distinguish the origin of fragments of DNA based on their genome signatures, was initially tested using GS-2 (see Methods, section 2, above) measured over fragments of 100 kb. At the time of analysis there were 79 eukaryotic, 156 eubacterial, 30 archaeal and 122 viral genomes with more than 100 kb of sequence each (Table 2). The dimension of the SOM was 50 × 50 and 100 iterations were used. At the end of the iterations, dominance areas (see Methods, section 3, above), were used to color the SOM. For the entire data set, “all life” in Figure 1, the superkingdoms of archaea, eubacteria and eukaryota were chosen, along with the unranked category of viruses. Within each of the SOMs applied to the superkingdoms and the viruses, the next level down was used for coloring dominance maps. This is the phylum level in the archaea and eubacteria, and the family level in the viruses. In the eukaryota, the relative scarcity of completely sequenced genomes required a more ad hoc classification.
When all input sets are pooled, GS-2 produces a SOM in which eubacterial sequences cluster together (Fig. 1; “All life”, green). Archaeal sequences are split into several groups that are situated along the boundary between the eubacteria and the eukaryotes. Likewise, viral sequences are split into one group in the top left corner and other clusters along the eubacterial-eukaryotic border. It is evident that this “all life” SOM does not contribute to the issue of the phylogeny of the three superkingdoms, except to underline that archaea are not derivatives of either eukaryotes or eubacteria.
When the SOM is confined to archaeal sequences (Fig. 1; “Archaea”), those genomes designated “unclassified” by NCBI, are located well within the territory of the Euryarchaeota, strongly suggesting that they belong to this phylum. In general the archaeal inter-phylum boundaries are clear, although the Crenarchaeaota are split into two clusters. The predominance of Euryarchaeota in terms of area is a reflection of the larger number of complete genomes in that phylum.
Likewise, in the eukaryotes (Fig. 1; “Eukaryota”), the large size of the human genome contributes to a large area dominated by the Vertebrata. It should be remembered that the classification in the eukaryotes is ad hoc owing to the relatively small number of complete genomes. However, it is interesting that the boundaries between the dominance areas are as distinct as those in the archaea.
The situation is considerably more complicated within the eubacteria (Fig. 1; “Eubacteria”), being the superkingdom with the greatest number of completely sequenced genomes. Some eubacterial phyla are rather fragmented in their dominance areas. For instance, the phylum Firmicutes occupies several partly adjacent areas. The phylum Deinococcus has two small and rather distant dominance areas, and the Bacteroidetes and Spirochaetes both have small outlying fragments. The Proteobacteria dominate the right side of the SOM and penetrate between the various groups on the left side. The overall impression is of less clear-cut differences in GS-2 between phyla in eubacteria than in eukaryotes or archaea.
A similar situation is observed in the SOM on viral sequences (Fig. 1; “Viruses”). A few viral families, such as the Baculoviridae, the family Mimivirus and the Nimaviridae do manage coherent dominance areas, but all others are extensively mixed. The Baculoviridae are the only family of any size than maintain a distinctive dominance area.
This basic illustration of the SOM in action demonstrates that for a single parameter set, namely 50 × 50 SOM and 100 iterations, different phylogenetic groups exhibit variable degrees of partition across the SOM.
2. Increased resolution SOM on viruses
To increase the resolution of the SOM against viral sequences, GS-2 was reapplied to viral sequences only using 10 kb fragments. This enables a larger number of viral genomes to be analysed, up from 122 to 579, as genomes of 10 kb or more can be included (Table 3). The number of iterations was increased to 1000. The resulting dominance map is shown in Figure 2.
When viral sequences alone are considered at higher resolution, the SOM becomes very complex. The family level classification is maintained for the dominance map but there are now more families, since viruses as small as 10 kb are eligible. Perhaps the most salient feature is that Poxviridae are divisible into sheep/goat pox viruses and others (Fig. 2: “sheep/goat” and “other pox”). Additionally phages, within the family Caudovirales, tend to be differentially located on the SOM in four major areas, one of which, mycophages, accounts for two of these areas (Fig. 2: “myco-ϕ”, “entero-ϕ” and “cocco-ϕ”). Again the Baculoviridae form a noticeably large and coherent cluster. Herpesviridae, by contrast, are spread across the entire map.
Herpesviridae (Table 1) are next considered alone under the same conditions as in Figure 2. Dominance maps for this narrower selection are shown in Figure 3.
Figure 3 shows that when family-level taxonomy is considered within herpesviruses, GS-2 distinguishes the ostreid herpesviruses and the ictaluriviruses as two fairly homogenous blocs distinct from the Alloherpesviridae (Davison, 2002), comprising the alpha, beta and gamma families. At the genus-level, Muromegalovirus alone forms a nearly contiguous bloc although Mardivirus nearly does so. The remaining genera, like the families, are considerably mixed across the SOM. Like the wide spread of herpesvirus signatures across the viral SOM, this is a reflection of the degree of sequence heterogeneity with the Herpesviridae.
The three figures presented above demonstrate that the SOM is an intriguing tool for the conceptualisation of relationships between genome signatures. However, the evident complexity of some of the topographical arrangements raises serious questions concerning its utility as a diagnostic tool for phylogeny.
Therefore, some experiments are described which address this issue in a quantitative way.
3. Effect of length of k-mer used to generate genome signature
In order to investigate if genome signatures of longer k give better resolution than k = 2, 10 kb herpesvirus sequences were processed into genome signature of GS-2 to GS-6 and the SOM was trained for 100 iterations (Fig. 4). On first inspection, it does not appear that a higher genome signature provides any better resolution than a lower one. The GS-3 SOM was also run on a 20 × 20 map, but again this produces no major change to the overall pattern. In all cases, ostreid herpesvirus and ictalurivirus have coherent dominance areas on the SOM. At GS-5, alpha herpes-viruses also have a coherent dominance area, but this disappears again at GS-6. In order to further investigate this apparent lack of improvement at higher values of k, the density of sequences of each family was plotted onto the SOM (Fig. 5). Instead of the dominance map approach, in which each cell is colored according to the affiliation of its cnn (Fig. 1–4 are all of this type), cells in which more than 95% of allocated sequences are of a single type are colored red, and those with fewer than 5% of that type are white. Cells between these two extremes are colored yellow. A ratio is then produced of red-to-yellow in each SOM. A perfectly partitioned SOM will therefore have a ratio of infinity, indicating no mixed cells, or more accurately no cells with greater than 5% mixture of the “wrong” family.
Figure 4.
Dominance maps herpesvirus families, illustrating the effect of varying GS values using 10kb herpesvirus sequences, on a 10 ×10 SOM (except for GS-3 at 20 ×20) over 100 iterations.
Figure 5.
The density of herpesviral sequences, classed by family, on a 10 ×10 SOM after 100 iterations. >95% density: red; 5%–95% density: yellow; <5% density: white. The figure in each box is the ratio of sequences in red to yellow areas of the SOM.
Figure 5 demonstrates that family level taxonomy is better determined at higher GS in all five families of herpesviruses. The ratio of high alpha-density (>95%, red) to medium alpha-density (5% to 95%, yellow) increases from 0.88 to 2.83 as the GS increases from 2 to 4. The corresponding increases for the beta and gamma families are from 0.52 to 2.33 and from 1.91 to 2.11 respectively. For the ostreid herpesviral sequences, perfect partition is reached at GS-4 and for the ictalurid viruses at GS-3. This is probably a reflection of the presence of a single virus in each of these categories with a correspondingly lower number of sequences analysed.
4. Effect of length of training phase of SOM
It is therefore apparent that genome signature of longer values of k produce some improvement in the accuracy of the final partition on the SOM. However, longer k results in longer data vectors, increasing at order 4k and therefore much slower training of the SOM. One way to speed training of the SOM is simply to reduce the number of training cycles. The effect of the number of iterations on density of each family is displayed in Figure 6.
Figure 6.
The density of herpesviral sequences, classed by family, on a 10 ×10 SOM of GS-2, run over a varying number of iterations, i. >95% density: red; 5%–95% density: yellow; <5% density: white. The figure in each box is the ratio of sequences in red to yellow areas of the SOM.
Figure 6 shows that increasing the number of iterations has a mixed effect on the density of family sequences. The alpha herpesviral sequences increase in density from 0.92 to 1.35 as the number of iterations increases from 10 to 1000, and the beta herpesviruses from 0.52 to 0.83. The ostreid herpesviral sequences are also perfectly clustered at 100 iterations. However, the gamma and ictalurid sequences are more poorly partitioned at higher numbers of iterations.
5. Jack-knifing analysis
Figures 1–6 provide a largely qualitative impression of the effectiveness of SOMs in correctly assigning the origins of DNA sequences based on their genome signature. To provide a further more quantitative assessment of the parameters of the process, a jack-knifing analysis was carried out. All herpesviral sequences were divided randomly into two groups. Genome signatures and SOMs were constructed as appropriate using one half. Then the remaining half was applied to the SOM to predict their origin at the family and genus level. To make a prediction concerning the origin of a data vector, the Euclidean distances between that vector and all of the weight vectors of the preconstructed SOM, are calculated. The origin of the nearest weight vector is taken to be the classification of the data vector being tested. Where a data vector falls into a cell on the SOM containing none of the original data vectors used to construct the SOM, its origin is deemed to be “undecided” (Fig. 7).
When SOM size is varied for GS-2 at 100 iterations (Fig. 7, top left table), SOMs of greater than 10 × 10 introduce considerably uncertainty into the assignment. However, for those sequences that can be assigned, 95% accuracy at the subfamily level is achieved in a 50 × 50 SOM. Likewise, a 30 × 30 SOM gives 94% accuracy at the genus level. When SOM size is held at 10 × 10 and the signature length at GS-2 and the number of iterations is varied (Fig. 7, lower left table), there is little effect on the sensitivity. At the subfamily level, there are never more than 4.4% of sequences that cannot be assigned, and never more than 7.2% at the genus level. Where sequences can be assigned, optimal accuracy is achieved at 1000 or 5000 iterations, but the variation in accuracy is low. Increasing the iterations from 10 to 5000 only gives a 4% increase in accuracy of assignment at the sub-family level. When 100 iterations are used and the SOM size is held at 10 × 10 (Fig. 7, top right table), GS-4 or GS-5 appear to be optimal.
Discussion
Genome signatures provide a summary of the k-mer content of a genome, corrected for compositional bias. Various studies in a wide range of species have revealed that genome signatures are generally constant within genomes and similar in related genomes (Karlin and Ladunga, 1994; Karlin et al. 1998; Gentles and Karlin, 2001). The extent to which this is a phenomenon of neutral drift or one of active conservation is unknown. It is intuitively obvious that two identical genomes will have identical genome signatures, and that as they diverge the genome signatures will also diverge. Indeed this is the basis of a least one bioinformatical tool that assesses sequence relatedness (Li et al. 2001; Li et al. 2002). However, various suggestions have been made for conservative selection pressures which would act to maintain genome signature similarity in related organisms, including dinucleotide stacking energies, curvature, methylation, superhelicity, context-dependent mutation biases and effects deriving from related replication machinery (Karlin and Burge, 1995; Blaisdell et al. 1996). If these factors are similar within a clade, they might act as a brake on genome signature divergence. The conservation of genome signatures within genomes (which is what originally gave rise to the term “signature” in this context) would tend to suggest that signatures do not drift neutrally, at least within genomes.
Figure 1 demonstrates that at the phylum level within the three superkingdoms of cellular life, satisfactory partition of GS-2 can be obtained by the SOM. However, this is less true for eubacteria than it is for eukaryotes and archaea. At the family level in viruses the picture is considerably more confused, with only the Baculoviridae demonstrating anything like territorial coherence on the SOM at GS-2 (Fig. 1 and 2). This may well be a reflection of speed of substitution in viral genomes. However at the species level, the same coherence within genomes as found in cellular organisms may well be the norm. For instance, when the ostreid and ictalurid herpes-virus families are included in a SOM with the Alloherpesviridae, these two families, both represented by a single viral genome, have strongly discrete areas on the SOM (Fig. 3 and 4).
This does not mean that genome signatures are not diagnostic tools for phylogenetic assignment at the family and sub-family level in herpesviruses, merely that the results should be interpreted with caution. The use of higher values of k appears to have a marginal effect on improving the discrete distribution of family-level herpesviral signatures on the SOM (Fig. 5) but jack-knifing indicates that this does not improve above k = 5 (Fig. 7). The effects of larger dimension SOMs and increased iterations are ambiguous at best. Optimal values appear to be around GS-4 or GS-5 with 500 to 1000 iterations of the SOM. The size of the SOM might be varied, with an initial run at high dimension (e.g. 50 × 50) followed by a lower dimension run (e.g. 10 × 10) for sequences unassigned by the first run (Fig. 7).
The use of genome signatures in the identification of pathogenicity islands is by now well established (Karlin, 1998; Karlin, 2001; Dufraigne et al. 2005). They are valuable in this context in that they indicate regions within genomes that have characteristics different to the rest of the genome. However, it is apparent from the present work that it is difficult on the basis of genome signatures to accurately identify the origin of the exogenous DNA. A BLAST search is more likely to generate informative hits in this context. Nevertheless for sequences that cannot be precisely identified on the basis of alignment-based methods such as BLAST, genome signatures with SOMs holds out the prospect of identification of origin to a reasonable level.
The optimization of SOM parameters reported here may also extend to other applications of SOMs. Of particular interest in bioinformatics is their use for the analysis of microarray data. The experimental design would be the same, with a standard microarray data set (e.g. the breast cancer data provided by Reid et al. 2005) substituting for the genome signature arrays. Dominance mapping would be done by clinical outcome, and jack-knife analysis could test the accuracy and sensitivity of assignment of that outcome.
Footnotes
Both genome signature and genomic signature are used interchangeably in the field, including by their originators. However, the term genome signature is to be preferred, since genomic signature is used in the field of molecular diagnostics to refer to a genotype correlated with medical symptoms or prognosis (e.g. Russo et al. 2005)
Please note that this article may not be used for commercial purposes. For further information please refer to the copyright statement at http://www.la-press.com/copyright.htm
References
- Abe T, Kanaya S, Kinouchi M, Ichiba Y, Kozuki T, Ikemura T. Informatics for unveiling hidden genome signatures. Genome. Res. 2003a;13:693–702. doi: 10.1101/gr.634603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abe T, Kanaya S, Kinouchi M, Ichiba Y, Kozuki T, Ikemura T. A novel bioinformatic strategy for unveiling hidden genome signatures of eukaryotes: Self-organizing map of oligonucleotide frequency. Genome Informatics. 2002;13:13–20. [PubMed] [Google Scholar]
- Abe T, Kanaya S, Kinouchi M, Ishihara N, Kosaka Y, Kozuki T, Ohyama A, Ikemura T. Human Genome Mapping 2003. Human Genome Organization; 2003b. Self-organizing maps reveal hidden genome characteristics on a single map analysing 90 genomes of prokaryotes and eukaryotes. [Google Scholar]
- Andrade MA, Casari G, Sander C, Valencia A. Classification of protein families and detection of the determinant residues with an improved self-organizing map. Biol Cybern. 1997;76:441–50. doi: 10.1007/s004220050357. [DOI] [PubMed] [Google Scholar]
- Arrigo P, Giuliano F, Scalia F, Rapallo A, Damiani G. Identification of a new motif on nucleic acid sequence data using Kohonen’s self-organizing map. Comput. Appl. Biosci. 1991;7:353–7. doi: 10.1093/bioinformatics/7.3.353. [DOI] [PubMed] [Google Scholar]
- Blaisdell B, Campbell A, Karlin S. Similarities and dissimilarities of phage genomes. Proc. Natl. Acad. Sci., U.S.A. 1996;93:5854–9. doi: 10.1073/pnas.93.12.5854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brocchieri L. Phylogenetic inferences from molecular sequences: review and critique. Theoretical Population Biology. 2001;59:27–41. doi: 10.1006/tpbi.2000.1485. [DOI] [PubMed] [Google Scholar]
- Campanaro S, Vezzi A, Vitulo N, Lauro F, D’Angelo M, Simonato F, Cestaro A, Malacrida G, Bertoloni G, Valle G, et al. Laterally transferred elements and high pressure adaptation in Photobacterium profundum strains. BMC Genomics. 2005;6:122–36. doi: 10.1186/1471-2164-6-122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Campbell A, Mrazek J, Karlin S. Genome signature comparisons among prokaryotic, plasmid and mitochondrial DNA. Proc. Natl. Acad. Sci., U.S.A. 1999;96:9184–9. doi: 10.1073/pnas.96.16.9184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coenye T, Vandamme P. Use of the genomic signature in bacterial classification and identification. Systematic and Applied Microbiology. 2004;27:175–85. doi: 10.1078/072320204322881790. [DOI] [PubMed] [Google Scholar]
- Covell DG, Wallqvist A, Rabow AA, Thanki N. Molecular classification of cancer: unsupervised self-organizing map analysis of gene expression microarray data. Mol. Cancer Ther. 2003;2:317–32. [PubMed] [Google Scholar]
- Davison A. Evolution of the herpesviruses. Veterinary Microbiology. 2002;86:69–88. doi: 10.1016/s0378-1135(01)00492-8. [DOI] [PubMed] [Google Scholar]
- Davison A, Eberle R, Ehlers B, Hayward G, McGeoch D, Minson A, Pellett P, Roizman B, Studdert M, Thiry E. ICTV. San Francisco: 2005. A planned order: Herpesvirales. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deschavanne P, Giron A, Vilain J, Fagot F, Fertil B. Genomic signature: Characterization and classification of species assessed by chaos game representation of sequences. Mol. Biol. Evol. 1999;16:1391–9. doi: 10.1093/oxfordjournals.molbev.a026048. [DOI] [PubMed] [Google Scholar]
- Dufraigne C, Fertil B, Lespinats S, Giron A, Deschavanne P. Detection and characterization of horizontal transfers in prokaryotes using genomic signature. Nucl. Acids Res. 2005;33:e6. doi: 10.1093/nar/gni004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards S, Fertil B, Giron A, Deschavanne P. A genomic schism in birds revealed by phylogenetic analysis of DNA strings. Syst Biol. 2002;51:599–613. doi: 10.1080/10635150290102285. [DOI] [PubMed] [Google Scholar]
- Fauquet C, Mayo M, Maniloff M, Desselburger U, Ball L. Virus Taxonomy: The Eighth Report of the International Committee on Taxonomy of Viruses. Elsevier; 2005. [Google Scholar]
- Gentles A, Karlin S. Genome-scale compositional comparisons in eukaryotes. Genome Res. 2001;11:540–6. doi: 10.1101/gr.163101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giuliano F, Arrigo P, Scalia F, Cardo PP, Damiani G. Potentially functional regions of nucleic acids recognized by a Kohonen’s self-organizing map. Comput Appl Biosci. 1993;9:687–93. doi: 10.1093/bioinformatics/9.6.687. [DOI] [PubMed] [Google Scholar]
- Jernigan R, Baran R. Pervasive properties of the genomic signature. BMC Genomics. 2002;3:23–32. doi: 10.1186/1471-2164-3-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanaya S, Kinouchi M, Abe T, Kudo Y, Yamada Y, Nishi T, Mori H, Ikemura T. Analysis of codon usage diversity of bacterial genes with a self-organizing map (SOM): characterization of horizontally transferred genes with emphasis on the E. coli O157 genome. Gene. 2001;276:89–99. doi: 10.1016/s0378-1119(01)00673-4. [DOI] [PubMed] [Google Scholar]
- Karlin S. Detecting anomalous gene clusters and pathogenicity islands in diverse bacterial genomes. Trends in Microbiology. 2001;9:335–43. doi: 10.1016/s0966-842x(01)02079-0. [DOI] [PubMed] [Google Scholar]
- Karlin S. Global dinucleotide signatures and analysis of genomic heterogeneity. Current Opinion in Microbiology. 1998;1:598–610. doi: 10.1016/s1369-5274(98)80095-7. [DOI] [PubMed] [Google Scholar]
- Karlin S, Brocchieri L, Mrazek J, Campbell A, AM S. A chimeric prokaryotic ancestry of mitochondrial and primitive eukaryotes. Proc. Natl. Acad. Sci., U.S.A. 1999;96:9190–95. doi: 10.1073/pnas.96.16.9190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S, Burge C. Dinucleotide relative abundance extremes: a genomic signature. Trends in Genetics. 1995;11:283–90. doi: 10.1016/s0168-9525(00)89076-9. [DOI] [PubMed] [Google Scholar]
- Karlin S, Campbell A, Mrázek J. Comparative DNA analysis across diverse genomes. Annual. Review. of Genetics. 1998;32:185–225. doi: 10.1146/annurev.genet.32.1.185. [DOI] [PubMed] [Google Scholar]
- Karlin S, Ladunga I. Comparisons of eukaryotic genomes sequences. Proc. Natl. Acad. Sci. U.S.A. 1994;91:12832–6. doi: 10.1073/pnas.91.26.12832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S, Mrazek J. What drives codon choices in human genes? J. Mol. Biol. 1996;262:459–72. doi: 10.1006/jmbi.1996.0528. [DOI] [PubMed] [Google Scholar]
- Karlin S, Mrázek J. Compositional differences within and between eukaryotic genomes. Proc. Natl. Acad. Sci., U.S.A. 1997;94:10227–32. doi: 10.1073/pnas.94.19.10227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlin S, Mrázek J, Campbell A. Compositional biases of bacterial genomes and evolutionary implications. Journal of Bacteriology. 1997;179:3899–913. doi: 10.1128/jb.179.12.3899-3913.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohonen T. Self-Organizing Maps. 2 edn. vol 30. Berlin: Springer; 1997. [Google Scholar]
- Li W, Jaroszewski L, Godzik A. Clustering of highly homologous sequences to reduce the size of large protein database. Bioinformatics. 2001;17:282–3. doi: 10.1093/bioinformatics/17.3.282. [DOI] [PubMed] [Google Scholar]
- Li W, Jaroszewski L, Godzik A. Tolerating some redundancy significantly speeds up clustering of large protein databases. Bioinformatics. 2002;18:77–82. doi: 10.1093/bioinformatics/18.1.77. [DOI] [PubMed] [Google Scholar]
- Mahony S, McInerney JO, Smith TJ, Golden A. Gene prediction using the Self-Organizing Map: automatic generation of multiple gene models. BMC Bioinformatics. 2004;5:23. doi: 10.1186/1471-2105-5-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McHardy AC, Martin HG, Tsirigos A, Hugenholtz P, Rigoutsos I. Accurate phylogenetic classification of variable-length DNA fragments. Nature Methods. 2007;4:63–72. doi: 10.1038/nmeth976. [DOI] [PubMed] [Google Scholar]
- Oja M, Sperber GO, Blomberg J, Kaski S. Self-organizing map-based discovery and visualization of human endogenous retroviral sequence groups. Int. J. Neural. Syst. 2005;15:163–79. doi: 10.1142/S0129065705000177. [DOI] [PubMed] [Google Scholar]
- Paz A, Kirzhner V, Nevo E, Korol A. Coevolution of DNA-interacting proteins and genome “dialect”. Mol. Biol. Evol. 2006;23:56–64. doi: 10.1093/molbev/msj007. [DOI] [PubMed] [Google Scholar]
- Reid JF, Lusa L, De Cecco L, Coradini D, Veneroni S, Daidone MG, Gariboldi M, Pierotti MA. Limits of predictive models using microarray data for breast cancer clinical treatment outcome. J. Natl. Cancer Inst. 2005;97:927–30. doi: 10.1093/jnci/dji153. [DOI] [PubMed] [Google Scholar]
- Ressom H, Wang D, Natarajan P. Clustering gene expression data using adaptive double self-organizing map. Physiol. Genomics. 2003;14:35–46. doi: 10.1152/physiolgenomics.00138.2002. [DOI] [PubMed] [Google Scholar]
- Russell G, Subak-Sharpe J. Similarity of the general designs of protochordates and invertebrates. Nature. 1977;266:533–5. doi: 10.1038/266533a0. [DOI] [PubMed] [Google Scholar]
- Russell G, Walker P, Elton R, Subak-Sharpe J. Doublet frequency analysis of fractionated vertebrate nuclear DNA. J. Mol. Biol. 1976;108:1–23. doi: 10.1016/s0022-2836(76)80090-3. [DOI] [PubMed] [Google Scholar]
- Russo J, Moral R, Balogh G, Mailo D, Russo I. The protective role of pregnancy in breast cancer. Breast Cancer Research. 2005;7:131–42. doi: 10.1186/bcr1029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samsonova EV, Kok JN, Ijzerman AP. TreeSOM: Cluster analysis in the self-organizing map. Neural Networks. 2006;19:935–49. doi: 10.1016/j.neunet.2006.05.003. [DOI] [PubMed] [Google Scholar]
- Sandberg R, Bränden C-I, Ernberg I, Cöster J. Quantifying the species-specificity in genomic signatures, synonymous codon choice, amino acid usage and G + C content. Gene. 2003;311:35–42. doi: 10.1016/s0378-1119(03)00581-x. [DOI] [PubMed] [Google Scholar]
- Sandberg R, Winberg G, Bränden C-I, Kaske A, Ernberg I, Cöster J. Capturing whole-genome characteristics in short sequences using a naive Bayesian classifier. Genome Res. 2001;11:1404–9. doi: 10.1101/gr.186401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamayo P, Slonim D, Mesirov J, Zhu Q, Kitareewan S, Dmitrovsky E, Lander E, Golub T. Interpreting patterns of gene expression with self-organizing maps: methods and application to hematopoietic differentiation. Proc. Natl. Acad. Sci., U.S.A. 1999;96:2907–12. doi: 10.1073/pnas.96.6.2907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teeling H, Meyerdierks A, Bauer M, Amann R, Glockner FO. Application of tetranucleotide frequencies for the assignment of genomic fragments. Environ. Microbiol. 2004;6:938–47. doi: 10.1111/j.1462-2920.2004.00624.x. [DOI] [PubMed] [Google Scholar]
- Ultsch A. Self-organized feature maps for monitoring and knowledge acquisition of a chemical process. In: Gielen S, Kappen B, editors. Proc. Intl. Conf. on Artificial Neural Networks (ICANN) Amsterdam: Springer-Verlag; 1993. 1993. pp. 864–7. [Google Scholar]
- Wang HC, Badger J, Kearney P, Li M. Analysis of codon usage patterns of bacterial genomes using the self-organizing map. Mol. Biol. Evol. 2001;18:792–800. doi: 10.1093/oxfordjournals.molbev.a003861. [DOI] [PubMed] [Google Scholar]
- Wang Y, Hill K, Singh S, Kan L. The spectrum of genomic signatures: from dinucleotides to chaos game representation. Gene. 2005;346:173–85. doi: 10.1016/j.gene.2004.10.021. [DOI] [PubMed] [Google Scholar]
- Xiao L, Wang K, Teng Y, Zhang J. Component plane presentation integrated self-organizing map for microarray data analysis. FEBS Lett. 2003;538:117–24. doi: 10.1016/s0014-5793(03)00156-x. [DOI] [PubMed] [Google Scholar]