Abstract
The aac(6′)-Ib gene was detected in 86 of 555 (15.5%) Enterobacteriaceae isolates. Among these 86 aac(6′)-Ib-positive isolates, 19 (22.0%) were positive for aac(6′)-Ib-cr: 4 of 31 (12.9%) Enterobacter spp., 7 of 13 (53.8%) Escherichia coli isolates, and 8 of 42 (19.0%) Klebsiella pneumoniae isolates. There was a strong association between aac(6′)-Ib-cr and OXA-1 and CTX-M-1 group β-lactamase genes. One aac(6′)-Ib-positive K. pneumoniae isolate carried both aac(6′)-Ib-cr and qnrS.
Plasmid-mediated quinolone resistance was first identified in a clinical isolate of Klebsiella pneumoniae (7, 12). Recently, a new mechanism of quinolone resistance was identified: transfer from species to species of a plasmid encoding aac(6′)-Ib-cr, a variant of aminoglycoside acetyltransferase that confers reduced susceptibility to ciprofloxacin and norfloxacin by N-acetylation of the amino nitrogen on its piperazinyl substituent (13). Genes responsible for plasmid-mediated quinolone resistance are thought to be linked to extended-spectrum β-lactamase genes (2, 6).
In Korea, Qnr determinants from Enterobacteriaceae have been reported (4, 8), but the presence of aac(6′)-Ib-cr has not been reported. We therefore assessed the prevalence of aac(6′)-Ib-cr genes among clinical isolates of Enterobacteriaceae in Korea.
During the period from January 2005 to December 2006, 555 nonduplicate enterobacterial isolates were collected from blood cultures at Asan Medical Center, a 2,200-bed tertiary care teaching hospital in Seoul, Korea. Screening for aac(6′)-Ib was carried out by PCR amplification with the specific primers 5′-TTGCGATGCTCTATGAGTGGCTA-3′ and 5′-CTCGAATGCCTGGCGTGTTT-3′, to produce a 482-bp product (9). Of the 555 Enterobacteriaceae clinical isolates, 86 (15.5%) were positive for the aac(6′)-Ib gene: 31 of 149 (20.8%) Enterobacter spp., 13 of 204 (6.4%) Escherichia coli isolates, and 42 of 202 (20.8%) Klebsiella pneumoniae isolates. Following digestion of the amplified products with BtsCI, we found that 19 of the 86 (22.0%) aac(6′)-Ib-positive isolates were positive for aac(6′)-Ib-cr: 4 of 31 (12.9%) Enterobacter spp. isolates, 7 of 13 (53.8%) E. coli isolates, and 8 of 42 (19.0%) K. pneumoniae isolates. The rate of aac(6′)-Ib-cr among aac(6′)-Ib-positive E. coli was higher than in aac(6′)-Ib-positive Enterobacter spp. and K. pneumoniae isolates. Random amplified polymorphic DNA analysis was performed by using a 254-decamer primer (5′-CCGCAGCCAA) to assess the clonal diversity (1). The 19 isolates gave 11 different patterns: 2 in Enterobacter cloacae, 3 in E. coli, and 6 in K. pneumoniae (Table 1).
TABLE 1.
Characteristics of aac(6′)-Ib-cr-positive isolates
| Species and isolate | CIP MIC (μg/ml) | Mutation(s) in QRDRs
|
RAPD patterna | β-Lactamases | Resistance to antibioticsb | |
|---|---|---|---|---|---|---|
| gyrA | parC | |||||
| E. cloacae | ||||||
| 37 | 4 | Ser83Ile | Ser80Arg | I | CTX-M-3, OXA-1 | TOB, GEN, CTX, SXT |
| 101 | 8 | Ser83Ile | Ser80Ile | II | CTX-M-3, OXA-1 | TOB, GEN, CTX, CAZ, SXT |
| 153 | 8 | Ser83Ile | Ser80Ile | II | CTX-M-3, OXA-1 | TOB, GEN, CTX, CAZ, SXT |
| 165 | 8 | Ser83Ile | Ser80Ile | II | CTX-M-3, OXA-1 | CTX, CAZ, SXT |
| E. coli | ||||||
| 17 | >16 | Ser83Leu, Asp87Asn | Ser80Ile, Glu84Val | III | CTX-M-15, OXA-1, TEM-1 | TOB, GEN, CTX, CAZ, SXT |
| 24 | >16 | Ser83Leu, Asp87Asn | Ser80Ile, Glu84Val | III | CTX-M-15, OXA-1, TEM-1 | TOB, CTX, CAZ |
| 47 | >16 | Ser83Leu, Asp87Asn | Ser80Ile | IV | CTX-M-15, OXA-1 | TOB, GEN, CTX, CAZ, SXT |
| 48 | >16 | Ser83Leu, Asp87Asn | Ser80Ile | V | CTX-M-15, OXA-1 | TOB, CTX, CAZ |
| 125 | >16 | Ser83Leu, Asp87Asn | Ser80Ile, Glu84Val | III | CTX-M-15, OXA-1, TEM-1 | TOB, GEN, CTX, CAZ |
| 130 | >16 | Ser83Leu, Asp87Asn | Ser80Ile | IV | CTX-M-15, OXA-1 | TOB, CTX, CAZ |
| 189 | >16 | Ser83Leu, Asp87Asn | Ser80Ile, Glu84Val | III | CTX-M-15, OXA-1, TEM-1 | TOB, GEN, CTX, CAZ |
| K. pneumoniae | ||||||
| 78 | 1 | Ser83Tyr | VI | CTX-M-15, OXA-1, SHV | TOB, GEN, CTX, CAZ, SXT | |
| 80 | 1 | Ser83Tyr | VI | CTX-M-15, OXA-1, SHV | TOB, GEN, CTX, CAZ, SXT | |
| 110 | >16 | Ser83Ile | Ser80Ile | VII | OXA-1, SHV | TOB, GEN, CAZ, SXT |
| 132 | 0.125 | VIII | CTX-M-15, OXA-1, SHV | TOB, GEN, CTX, CAZ | ||
| 135 | >16 | Ser83Ile | Ser80Ile | IX | CTX-M-15, OXA-1, TEM-1, SHV | TOB, GEN, CTX, CAZ, SXT |
| 185 | 1 | Ser83Tyr | X | CTX-M-15, OXA-1, SHV | TOB, GEN, CTX, SXT | |
| 194 | 16 | Ser83Tyr | VI | CTX-M-15, OXA-1, SHV | TOB, GEN, CTX, CAZ, SXT | |
| 202 | >16 | Ser83Ile | Ser80Ile | XI | CTX-M-3, OXA-1, SHV | TOB, GEN, CTX, CAZ |
RAPD, randomly amplified polymorphic DNA.
CIP, ciprofloxacin; TOB, tobramycin; GEN, gentamicin; CTX, cefotaxime; CAZ, ceftazidime; SXT, trimethoprim-sulfamethoxazole.
We also assessed whether these Enterobacteriaceae clinical isolates possessed the three qnr genes by PCR, as described previously (14). Ten Enterobacter spp. were positive for qnrA; 3 Enterobacter spp. and 8 K. pneumoniae isolates were positive for qnrB; and 1 Enterobacter spp., 1 E. coli isolate, and 4 K. pneumoniae isolates were positive for qnrS. One or more qnr genes were present in 27 of the 555 (4.9%) isolates: 14 of 149 Enterobacter spp. (9.4%), 1 of 204 E. coli isolates (0.5%), and 12 of 202 K. pneumoniae isolates (5.9%). The rate of qnr carriage among Enterobacter spp. was higher than in E. coli and K. pneumoniae isolates. One K. pneumoniae isolate (no. 135) contained both the aac(6′)-Ib-cr and qnrS genes; to our knowledge, this is the first such finding in a clinical isolate from Korea. The genetic structure between the aac(6′)-Ib-cr and qnrS genes in the K. pneumoniae no. 135 clinical isolate was determined by sequencing on plasmid DNA. Results showed that the β-lactamase-encoding genes blaOXA-1, blaCTX-M-15, and blaTEM-1 were detected between the aac(6′)-Ib-cr and qnrS genes, along with other genes (Fig. 1).
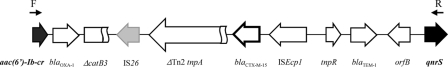
FIG. 1.
Genetic organization of the 13-kb region between the aac(6′)-Ib-cr and qnrS genes in the K. pneumoniae no. 135 clinical isolate. The genes and their transcription orientations are represented by horizontal arrows. For sequencing of plasmid DNA, PCR was done with primers F (5′-TTGCGATGCTCTATGAGTGGCTA-3′) and R (5′-TAAATTGGCACCCTGTAGGC-3′). Both strands of the PCR product were used for DNA sequencing.
In Enterobacteriaceae isolates, aac(6′)-Ib-cr is linked to the extended-spectrum β-lactamase genes (2, 6). Using PCR and DNA sequencing as described previously (3, 5, 10), we determined whether β-lactamase genes were present in our isolates and analyzed whether they were TEM, SHV, CTX-M, or OXA types (Table 1). All of the aac(6′)-Ib-cr-positive E. cloacae isolates produced CTX-M-3 and OXA-1. Most of the aac(6′)-Ib-cr-positive E. coli and K. pneumoniae isolates produced CTX-M-15 and OXA-1, except for K. pneumoniae no. 202, which produced CTX-M-3 and OXA-1. Some isolates also produced TEM-1 as well as CTX-M-15 and OXA-1. One aac(6′)-Ib-cr-positive K. pneumoniae isolate (no. 110) produced only OXA-1. These results showed that aac(6′)-Ib-cr is simultaneously associated with OXA-1 and CTX-M-1 (CTX-M-3 or CTX-M-15). No SHV-type gene was detected in any of the aac(6′)-Ib-cr-positive E. cloacae or E. coli isolates.
The transferability of the aac(6′)-Ib-cr gene was determined by conjugation experiments using azide-resistant E. coli J53 Azir as the recipient. The aac(6′)-Ib-cr gene was successfully transferred from eight isolates, and its presence was confirmed in all eight transconjugants by PCR (Table 2). Most of the aac(6′)-Ib-cr-positive isolates were resistant to gentamicin, tobramycin, and nalidixic acid, as well as ciprofloxacin. The MICs of ciprofloxacin and norfloxacin against transconjugants were two- to fourfold higher than for the recipient E. coli J53, indicating that aac(6′)-Ib-cr contributed to the decrease in ciprofloxacin susceptibility. The transconjugant for the aac(6′)-Ib-cr-positive K. pneumoniae isolate 135, which carried both aac(6′)-Ib-cr and qnrS, showed a 32-fold increase in MIC for ciprofloxacin (1 μg/ml), the clinical breakpoint for susceptibility, compared with the MIC shown by the recipient E. coli J53. Thus, when both the aac(6′)-Ib-cr and qnr genes are present in the same cells, the level of resistance is much higher than that conferred by aac(6′)-Ib-cr alone.
TABLE 2.
Susceptibility profiles of aac(6′)-Ib-cr-positive isolates and their transconjugants
| Isolatea | MIC (μg/ml)b
|
|||||||
|---|---|---|---|---|---|---|---|---|
| CIP | NOR | MXF | NAL | TOB | GEN | CAZ | CTX | |
| E. cloacae 37 | 4 | >16 | 2 | >256 | >16 | >32 | 8 | 128 |
| Tc Ecl 37 | 0.06 | 0.25 | 0.06 | 8 | >16 | >32 | 2 | 64 |
| E. cloacae 101 | 8 | >16 | 4 | >256 | >16 | >32 | >32 | 256 |
| Tc Ecl 101 | 0.06 | 0.25 | 0.06 | 8 | 2 | 0.25 | 2 | 32 |
| E. cloacae 153 | 8 | >16 | 4 | >256 | >16 | >32 | >32 | 256 |
| Tc Ecl 153 | 0.06 | 0.25 | 0.06 | 8 | >16 | >32 | 1 | 32 |
| E. coli 17 | >16 | >16 | >16 | >256 | >16 | >32 | 32 | 256 |
| Tc Eco 17 | 0.06 | 0.25 | 0.06 | 8 | 16 | 32 | 16 | 256 |
| E. coli 48 | >16 | >16 | 16 | >256 | 16 | 0.25 | >32 | >256 |
| Tc Eco 48 | 0.06 | 0.25 | 0.06 | 8 | 8 | 0.25 | >32 | >256 |
| K. pneumoniae 132 | 0.125 | 0.5 | 0.125 | 8 | >16 | >32 | 32 | >256 |
| Tc Kp 132 | 0.03 | 0.125 | 0.06 | 8 | 16 | 32 | 16 | >256 |
| K. pneumoniae 135 | >16 | >16 | >16 | >256 | >16 | >32 | >32 | 256 |
| Tc Kp 135 | 1 | 8 | 1 | 32 | >16 | >32 | >32 | 256 |
| K. pneumoniae 202 | >16 | >16 | 16 | >256 | >16 | >32 | >32 | >256 |
| Tc Kp 202 | 0.03 | 0.125 | 0.06 | 8 | >16 | >32 | 1 | 16 |
| J53, recipient | 0.016 | 0.06 | 0.06 | 8 | 0.25 | 0.25 | 0.25 | 0.06 |
Tc, transconjugant.
CIP, ciprofloxacin; NOR, norfloxacin; MXF, moxifloxacin; NAL, nalidixic acid; TOB, tobramycin; GEN, gentamicin; CAZ, ceftazidime; CTX, cefotaxime.
Almost all aac(6′)-Ib-cr-positive isolates contained a CTX-M-1 group β-lactamase gene, except for K. pneumoniae no. 110, which expressed only OXA-1. The cefotaxime MICs for such CTX-M-1-producing isolates were higher than those of ceftazidime, and this result was also found in their transconjugants (Table 2). The MICs of ceftazidime and cefotaxime for isolates producing CTX-M-3 were lower than those for isolates producing CTX-M-15 (Table 2). One amino acid difference at position 240 in CTX-M-15 was found to confer increased catalytic activity compared to that of CTX-M-3 (11).
The MICs of ciprofloxacin in aac(6′)-Ib-cr-positive isolates were much higher than those for the corresponding transconjugants, with MICs of 1 to ≥ 32 μg/ml, except for K. pneumoniae no. 110 (0.125 μg/ml). To determine if any target modification occurred in aac(6′)-Ib-cr-positive isolates, their quinolone resistance-determining regions (QRDRs) were sequenced. All aac(6′)-Ib-cr-positive isolates had point mutations in the QRDRs of the gyrA gene, at codon 83 and/or codon 87, except for K. pneumoniae no. 110, which did not have mutations in the QRDRs of the gyrA or parC genes. All aac(6′)-Ib-cr-positive E. cloacae and E. coli and three aac(6′)-Ib-cr-positive K. pneumoniae isolates had mutations in the QRDRs of the parC gene, at codon 80 and/or codon 84.
In conclusion, aac(6′)-Ib-cr was detected in three genera of Enterobacteriaceae (E. cloacae [four isolates], E. coli [seven isolates], and K. pneumoniae [eight isolates]), indicating horizontal transfer among the Enterobacteriaceae. The aac(6′)-Ib-cr gene showed a high association with β-lactamase genes, including OXA-1, CTX-M-3 or -15, and TEM-1, in isolates from Korea.
Acknowledgments
This work was supported by grant 2007-348 from the Asan Institute for Life Sciences, Seoul, Korea.
E.S.K. and J.-Y.J. contributed equally to this work.
Footnotes
Published ahead of print on 16 March 2009.
REFERENCES
- 1.Aslam, M., F. Nattress, G. Greer, C. Yost, C. Gill, and L. McMullen. 2003. Origin of contamination and genetic diversity of Escherichia coli in beef cattle. Appl. Environ. Microbiol. 69:2794-2799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cordeiro, N. F., L. Robino, J. Medina, V. Seija, I. Bado, V. García, M. Berro, J. Pontet, L. López, C. Bazet, G. Rieppi, G. Gutkind, J. A. Ayala, and R. Vignoli. 2008. Ciprofloxacin-resistant enterobacteria harboring the aac(6′)-Ib-cr variant isolated from feces of inpatients in an intensive care unit in Uruguay. Antimicrob. Agents Chemother. 52:806-807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ho, P. L., R. H. Shek, K. H. Chow, R. S. Duan, G. C. Mak, E. L. Lai, W. C. Yam, K. W. Tsang, and W. M. Lai. 2005. Detection and characterization of extended-spectrum beta-lactamases among bloodstream isolates of Enterobacter spp. in Hong Kong, 2000-2002. J. Antimicrob. Chemother. 55:326-332. [DOI] [PubMed] [Google Scholar]
- 4.Jeong, J. Y., H. J. Yoon, E. S. Kim, Y. Lee, S. H. Choi, N. J. Kim, J. H. Woo, and Y. S. Kim. 2005. Detection of qnr in clinical isolates of Escherichia coli from Korea. Antimicrob. Agents Chemother. 49:2522-2524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kim, J., and Y. M. Lim. 2005. Prevalence of derepressed AmpC mutants and extended-spectrum β-lactamase producers among clinical isolates of Citrobacter freundii, Enterobacter spp., and Serratia marcescens in Korea: dissemination of CTX-M-3, TEM-52, and SHV-12. Antimicrob. Agents Chemother. 43:2452-2455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Machado, E., T. M. Coque, R. Cantón, F. Baquero, J. C. Sousa, and L. Peixe. 2006. Dissemination in Portugal of CTX-M-15-, OXA-1-, and TEM-1-producing Enterobacteriaceae strains containing the aac(6′)-Ib-cr gene, which encodes an aminoglycoside- and fluoroquinolone-modifying enzyme. Antimicrob. Agents Chemother. 50:3220-3221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Martinez-Martinez, L., A. Pascual, and G. A. Jacoby. 1998. Quinolone resistance from a transferable plasmid. Lancet 351:797-799. [DOI] [PubMed] [Google Scholar]
- 8.Pai, H., M. R. Seo, and T. Y. Choi. 2007. Association of QnrB determinants and production of extended-spectrum beta-lactamases or plasmid-mediated AmpC beta-lactamases in clinical isolates of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 51:366-368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Park, C. H., A. Robicsek, G. A. Jacoby, D. Sahm, and D. C. Hooper. 2006. Prevalence in the United States of aac(6′)-Ib-cr encoding a ciprofloxacin-modifying enzyme. Antimicrob. Agents Chemother. 50:3953-3955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Park, Y. J., S. Y. Park, E. J. Oh, J. J. Park, K. Y. Lee, G. J. Woo, and K. Lee. 2005. Occurrence of extended-spectrum β-lactamases among chromosomal AmpC-producing Enterobacter cloacae, Citrobacter freundii, and Serratia marcescens in Korea and investigation of screening criteria. Diagn. Microbiol. Infect. Dis. 51:265-269. [DOI] [PubMed] [Google Scholar]
- 11.Poirel, L., M. Gniadkowski, and P. Nordmann. 2002. Biochemical analysis of the ceftazidime-hydrolysing extended-spectrum beta-lactamase CTX-M-15 and of its structurally related beta-lactamase CTX-M-3. J. Antimicrob. Chemother. 50:1031-1034. [DOI] [PubMed] [Google Scholar]
- 12.Robicsek, A., G. A. Jacoby, and D. C. Hooper. 2006. The worldwide emergence of plasmid-mediated quinolone resistance. Lancet Infect. Dis. 6:629-640. [DOI] [PubMed] [Google Scholar]
- 13.Robicsek, A., J. Strahilevitz, G. A. Jacoby, M. Macielag, D. Abbanat, C. H. Park, K. Bush, and D. C. Hooper. 2006. Fluoroquinolone modifying enzyme: a new adaptation of a common aminoglycoside acetyltransferase. Nat. Med. 12:83-88. [DOI] [PubMed] [Google Scholar]
- 14.Wu, J. J., W. C. Ko, H. M. Wu, and J. J. Yan. 2008. Prevalence of Qnr determinants among bloodstream isolates of Escherichia coli and Klebsiella pneumoniae in a Taiwanese hospital, 1999-2005. J. Antimicrob. Chemother. 61:1234-1239. [DOI] [PubMed] [Google Scholar]