Summary
Dscam is an immunoglobulin (Ig) superfamily protein required for the formation of neuronal connections in Drosophila. Through alternative splicing, Dscam potentially gives rise to 19,008 different extracellular domains linked to one of two alternative transmembrane segments, resulting in 38,016 isoforms. All isoforms share the same domain structure but contain variable amino acid sequences within three Ig domains in the extracellular region. We demonstrate that different isoforms exhibit different binding specificity. Each isoform binds to itself but does not bind or binds poorly to other isoforms. The amino acid sequences of all three variable Ig domains determine binding specificity. Even closely related isoforms sharing nearly identical amino acid sequences exhibit isoform-specific binding. We propose that this preferential homophilic binding specificity regulates interactions between cells and contributes to the formation of complex patterns of neuronal connections.
Introduction
The specificity of neuronal connections is a hallmark of nervous systems from simple invertebrates to humans. On the basis of regeneration experiments in lower vertebrates, Sperry proposed that individual nerve “fibers…must carry some kind of individual identification tags presumably cytochemical in nature” that determine the pattern of connections formed (Sperry, 1963). While molecules involved in cell recognition events in the developing brain have been described, it remains unknown whether a vast repertoire of molecular tags mediating interactions between cells exists. In principle, such molecules could function in multiple steps in the wiring process, including axon guidance, target recognition, and synapse formation.
In many regions of the nervous system, the demand for diverse molecular tags in controlling specificity may be modest. For example, a sequence of guidance steps regulated in a precise spatiotemporal fashion by a small number of signals acting at short and long range to attract or repel extending axons may be sufficient to guide growth cones to regions containing few potential targets (Tessier-Lavigne and Goodman, 1996). In the target region, a relatively small number of cell surface determinants or combinations of them may be sufficient to distinguish between appropriate and inappropriate connections. Genetic experiments suggest that target specificity in the Drosophila neuromuscular system may be regulated in this manner (Winberg et al., 1998).
In some regions of the developing nervous system, the levels of cell surface recognition proteins on different neurons determine wiring patterns. Gradients of cell surface proteins specify the targeting of millions of neurons to form topographic maps. In the vertebrate visual system, for instance, the graded expression of Eph receptors on growth cones and cell surface bound ephrins in the target tissue control the precise targeting of retinal ganglion cell growth cones to specific regions of the superior colliculus or the tectum (Flanagan and Vanderhaeghen, 1998; McLaughlin et al., 2003a). Recent studies indicate that both contact-dependent attraction and contact-dependent repulsion mediated by different levels of Eph receptors and ephrin ligands contribute to topographic map formation (McLaughlin et al., 2003b; Hansen et al., 2004).
It is not clear to what extent large families of cell surface proteins with different recognition properties contribute to specificity. In the mouse olfactory system, molecular diversity plays a crucial role in target specificity. Gain-of-function and loss-of-function studies indicate that odorant receptors, a large family of ∼1000 G protein coupled receptors, play an instructive role in regulating targeting of olfactory receptor neurons to specific glomeruli in the olfactory bulb (Wang et al., 1998). How these receptors regulate this process is not known (Feinstein and Mombaerts, 2004). Other large families of cell surface proteins have been proposed to regulate neuronal connectivity, including classical cadherins (20 in the mouse) (Uemura, 1998), cadherin-related neuronal receptors (52 in the mouse) (Kohmura et al., 1998; Wu and Maniatis, 1999), MHC class I proteins (20 in the mouse) (Huh et al., 2000), and neurexins/neuroligins (∼1000 neurexin isoforms in vertebrates) (Missler and Sudhof, 1998; Ullrich et al., 1995). Whether these families of proteins contribute to specific cellular recognition underlying connection specificity remains to be seen.
The discovery that the Drosophila Dscam locus potentially encodes 38,016 different protein isoforms (Schmucker et al., 2000) raised the intriguing possibility that these isoforms function as molecular tags to regulate neuronal connectivity. Different isoforms are generated by alternative splicing and share the same domain structure with ten immunoglubulin (Ig) domains, six fibronectin type III (FNIII) repeats, a single transmembrane domain, and a C-terminal cytoplasmic tail (see Figure 1). There are four variable domains encoded by blocks of alternative exons. These include the N-terminal halves of Ig2 (12 alternatives) and Ig3 (48 alternatives), all of Ig7 (33 alternatives), and two alternative transmembrane domains. Dscam regulates axon guidance and targeting in many regions of the nervous system (Hummel et al., 2003; Wang et al., 2002). Recent studies have determined that neurons express many different Dscam isoforms, and different classes of neurons express biased sets of them (Neves et al., 2004; Zhan et al., 2004). It remains unclear whether different Dscam isoforms have distinct biochemical properties that contribute to cellular recognition controlling specificity.
Figure 1. Alternative Splicing of Dscam Potentially Generates 38,016 Isoforms.
(A) Schematic representation of Dscam gene, mRNA, and protein. The Dscam protein contains both constant and variable domains. The variable domains are encoded by alternative exons. Each block of alternative exons is indicated by a different color. A transcript contains only one alternative exon from each block. The Dscam gene encodes 12 alternative exons for the N-terminal half of Ig2 (red), 48 alternative exons for the N-terminal half of Ig3 (blue), and 33 alternative exons for Ig7 (green). There are two alternative transmembrane domains (yellow). (B) Schematic representation of Dscam proteins used in this study and explanation of isoform nomenclature. Immunoglobulin domain (Ig), horseshoe; fibronectin type III repeat (FNIII), black rectangle; transmembrane domain (TM), vertical rectangle. The NH2 and COOH termini are indicated. Individual isoforms are denoted by the combination of alternative variable Ig domains. For instance, the isoform comprising Ig2 alternative 7, Ig3 alternative 27, and Ig7 alternative 25 is designated Dscam7.27.25 or simply as 7.27.25. In this study, both full-length proteins and fragments of the extracellular domain were used. Full-length Dscam proteins expressed in COS cells and transgenic flies did not contain an epitope tag. Full-length proteins expressed in Drosophila S2 cells contained an internal FLAG epitope tag in the C-terminal cytoplasmic tail (denoted by a flag icon). For simplicity, the alternative transmembrane domain used in full-length isoforms is not indicated. Purified Dscam proteins containing different regions of the extracellular domain contained either a 6xHis tag (schematic not shown) or the Fc region of human IgG at the C terminus. Fc tags contain the hinge region, which dimerizes the Dscam proteins. 6xHis tags do not dimerize the Dscam proteins.
In this paper, we explore the biochemical properties of different Dscam isoforms. Through a series of binding assays, we demonstrate that each Dscam isoform tested binds to itself. We present data that demonstrates that this binding can promote interactions between cells in vivo. Dscam isoforms show surprising isoform-specific homophilic binding; each isoform binds to itself but poorly, if at all, to other isoforms tested. Based on the binding studies described here and in vivo studies (Zhan et al., 2004), we speculate on the role of Dscam isoforms in regulating neuronal connectivity in the Drosophila brain.
Results
Dscam Can Mediate Interactions between Cells In Vivo
Mammalian Dscam (Agarwala et al., 2000) and its para-log DscamL1 (Agarwala et al., 2001) have been shown to promote aggregation of transfected mouse cells in culture. Both gain- and loss-of-function genetic experiments on Drosophila Dscam are consistent with Dscam-mediated interactions between neurites (Hummel et al., 2003; Schmucker et al., 2000; Wang et al., 2002; Zhan et al., 2004). Based on these observations, we speculated that Drosophila Dscam proteins on opposing neuronal cell surfaces bind to each other.
To address this possibility in vivo, we assessed the consequences of overexpressing a single Dscam isoform in two populations of cells, a subset of interneurons (IN) and midline cells (M) that encounter one another during development. These interneurons extend axons across the midline cells in both the posterior (PC) and anterior (AC) commissures (Figure 2). The targeted expression of a single isoform is superimposed upon a background of many different isoforms of endogenous Dscam protein expressed throughout the neuropil of the embryonic central nervous system (Figures 2D and 2E). Interneuron axons were visualized using a marker specific for these cells (Figure 2B). To generate high levels of Dscam expression in both midline cells and in interneurons, we utilized the Gal4/UAS system. This bipartite expression system, however, precludes expression of one isoform in interneurons and another in midline cells in the same animal.
Figure 2. Dscam Promotes Interactions between Cells In Vivo.
To assess whether Dscam can mediate interactions between cells in vivo, a transgene expressing a single isoform of Dscam was expressed in both midline cells and a subclass of interneurons using the Gal4/UAS system (see Experimental Procedures). (A) Schematic representation of the embryonic ventral nerve cord (VNC, gray) at three stages of development showing the spatiotemporal pattern of expression of two Gal4 driver lines used in this assay. The midline cell driver (M; green) is expressed before the interneurons (IN; red) cross the midline. During early stage 13, anterior commissure (AC) axons begin crossing the midline. The posterior commissure (PC) axons develop later and have crossed the midline by late stage 14. By this stage, PC interneuron cell bodies form a tight cluster. Embryos were analyzed at stage 16. Anterior is to the top. (B) Confocal section of a region of the VNC in a control embryo lacking the Dscam transgene. Interneurons (arrows) are labeled in red. (C) Confocal section of an embryo expressing transgenic Dscam in midline cells. Midline transgenic Dscam expression does not affect the trajectory of the interneurons (arrows). (D) Endogenous Dscam (green) is expressed throughout the VNC. (E) Confocal projection of an embryo expressing the Dscam transgene in midline cells (arrows). Strong midline staining is seen in addition to endogenous Dscam. (F) Confocal section of a control embryo lacking the Dscam transgene. Interneuron cell bodies form tight clusters, and axons cross the midline. (G–I) Confocal sections of three different embryos expressing the Dscam transgene only in interneurons (red). About 50% of PC axons show midline crossing phenotypes (arrows). In addition, cell bodies often fail to form tight clusters (arrowheads). (J) Confocal section of a control embryo lacking the Dscam transgene stained with midline (green) and interneuron (red) markers. (K–M) Three examples of embryos expressing the Dscam transgene in both midline cells and interneurons. PC axons rarely cross the midline, and processes remain on the ipsilateral side (arrows). Cell body phenotypes are also observed (arrowheads). (N) Quantification of PC axon midline crossing. The x axis indicates whether transgenic Dscam was expressed in midline cells (M), interneurons (IN), or both. Only 3.5% of PC axons from embryos expressing the Dscam transgene in both cell types crossed the midline. n = number of segments scored.
Midline expression of the Dscam transgene disrupted neither the morphology of midline cells (compare green staining in Figures 2J–2M) nor guidance of the interneurons across it (Figures 2C and 2N). By contrast, expression of the Dscam transgene in only the interneurons gave rise to a weak dominant phenotype (Figures 2F–2I). In about 50% of the segments, PC interneurons did not cross the midline (Figure 2N). These axons were often visible as short stumps (Figure 2H, arrows) or as longer processes that invaded the midline but failed to fasciculate with axons of PC interneurons from the contralateral side (Figure 2I, arrows). The arrangement of PC cell bodies was also abnormal. In wild-type embryos, the four PC cell bodies formed a tight cluster (Figure 2F) (Higashijima et al., 1996), whereas PC interneurons expressing the Dscam transgene remained loosely associated or in some cases completely separated from one another (Figure 2H, arrowheads).
A striking phenotype of PC axons was observed in embryos in which both the interneurons and midline cells expressed transgenic Dscam (Figures 2J–2M). Only 3.5% of the PC axons crossed the midline, a greater than 10-fold reduction over that seen in embryos in which the Dscam transgene was expressed solely in the interneurons (Figure 2N). The AC interneurons were not affected by transgenic Dscam expression. These neurons may not express the molecular machinery required for Dscam signaling. Alternatively, as AC axons cross the midline prior to PC axons, transgenic Dscam may not have accumulated to sufficient levels in these cells to induce a similar response. These data support the notion that Dscam can mediate interactions between opposing cell surfaces in vivo.
Dscam Binds to Itself In Vitro
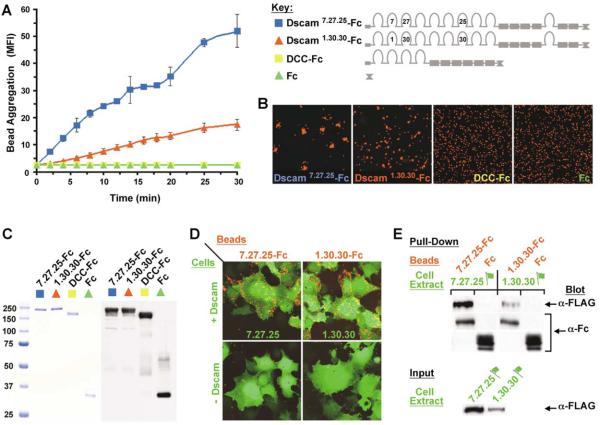
To test whether the extracellular domains of Dscam directly interact, we assessed binding of purified chimeric proteins comprising the entire extracellular domain of Dscam. Different isoforms were tagged at their C termini with the Fc region of human IgG (Figure 1B). For nomenclature of Dscam protein isoforms used in this paper, see Figure 1. Binding was assessed using a bead aggregation assay (De Angelis et al., 1999). Proteins were produced in Drosophila S2 cells, purified by protein A Sepharose affinity chromatography, and then captured via their Fc tags on fluorescent beads (0.4 μm) coated with anti-Fc antibody. We examined the ability of Dscam-Fc proteins to bind to each other by monitoring bead aggregation over a 30 min time course by FACS analysis (Figure 3A; see Experimental Procedures). As beads bind to each other, they form clusters with fluorescence intensity proportional to the number of beads within the aggregate. At various times of incubation, aliquots of beads were removed, and the mean fluorescence intensity (MFI) of 20,000 particles was assessed by FACS analysis. A particle may consist of a single bead or an aggregate of two or more beads. Negative controls included beads with the Fc tag and an Fc fusion protein containing the ectodomain of the Ig superfamily protein DCC. Dscam isoforms with different variable Ig domains were tested for binding (see schematic in Figure 3 for isoforms used). Dscam-Fc beads were largely monomeric (96%; FACS data not shown) at the start of the incubation period. Dscam isoforms promoted robust bead aggregation. By contrast, beads containing Fc and DCC-Fc remained largely monomeric. Beads visualized by confocal microscopy at the 30 min time point revealed aggregates of Dscam-coated beads but not control beads (Figure 3B). Dscam-Fc and control beads contained equivalent levels of purified protein at the end of the aggregation assay as assessed by immunoblotting (Figure 3C). Both Dscam isoforms exhibited distinct binding properties with characteristic rates and extents of aggregation (Figures 3A and 3B).
Figure 3. Dscam Binds to Itself In Vitro.
Three assays were used to assess Dscam binding to itself. (A) Dscam promotes bead aggregation. Fluorescent beads were decorated with the extracellular domain of a Dscam isoform tagged with Fc, Dscam7.27.25-Fc (blue), or Dscam1.30.30-Fc (red). Beads were incubated with agitation, and a time course of aggregation over 30 min was monitored by FACS analysis. Aggregation was not observed for beads decorated with either Fc (green) or Fc fused to DCC (DCC-Fc, yellow), another Ig/FNIII-containing protein. Each time point represents the mean and SD of two independent binding experiments. Two different preparations of each protein were used. (B) Confocal images of beads taken from 30 min time point in (A). Dscam-Fc coated beads form aggregates, while control DCC-Fc and Fc coated beads remained largely monomeric. (C) (Left panel) Coomassie-stained gel of purified Fc proteins used in binding assays. (Right panel) Control beads (Fc and DCC-Fc) and Dscam-Fc beads contained similar levels of protein as assessed by immunoblotting using an anti-Fc antibody following the aggregation assay. (D) Dscam beads bind to cells expressing Dscam. Red fluorescent beads decorated with the indicated isoforms of Dscam-Fc were incubated with COS cells (green) expressing the same Dscam isoform or with control cells. Dscam-Fc beads bound to Dscam-expressing cells but not to control cells. (E) Dscam-Fc proteins bind to full-length Dscam in cell extracts. Dscam-Fc or Fc alone was incubated with extracts of Drosophila S2 cells transfected with full-length FLAG-tagged Dscam. Protein G Sepharose beads were used to bind Fc tags and pull down associated proteins. Dscam-Fc proteins pulled down Dscam-FLAG, but Fc control proteins did not. The same amount of Dscam-Fc or Fc protein (1 μg) was added to each pull-down experiment. The level of FLAG-tagged protein in each extract is shown in the lower panel. The flag icon denotes full-length FLAG-tagged Dscam. For assays used in (A), (D), and (E), see Experimental Procedures.
Two additional assays were used to assess Dscam binding. As discussed in later sections, these assays were crucial for testing binding between different Dscam isoforms. In the first assay, red fluorescent beads decorated with Dscam-Fc were tested for binding to COS cells expressing full-length Dscam and GFP (Figure 3D). A slurry of beads was layered over cells transfected with Dscam or control cells, incubated for 2 hr, and washed to remove unbound beads. Bead binding to cells was visualized by confocal microscopy. Beads bound to Dscam-expressing COS cells but not to control cells. In the second assay, Dscam-Fc proteins captured on protein G Sepharose beads were used to test binding of Dscam to itself in extracts of cultured Drosophila S2 cells transfected with full-length FLAG-tagged Dscam (Figure 3E, also see Figure 1B for schematic of Dscam-FLAG). Proteins pulled down were analyzed by immunoblots probed with anti-FLAG antibody. Dscam-FLAG was precipitated by Dscam-Fc but not by Fc alone. In addition to these two isoforms, nine others were tested for binding using these assays (see below). All exhibited homophilic binding (Table 1). These 11 isoforms included five of the 12 alternative versions for Ig2, five of the 48 alternative versions for Ig3, and seven of the 33 alternative versions for Ig7. Based on these observations, we conclude that homophilic binding is a general property of Dscam proteins. As there are literally tens of thousands of different Dscam isoforms, it remains possible that some isoforms will not exhibit homophilic binding. We explore below whether different Dscam isoforms bind to each other.
Table 1.
Summary of Dscam Isoform Binding
| Binding between same isoforms |
Binding between different isoforms |
|||||||
|---|---|---|---|---|---|---|---|---|
| Beads | Cells | Binding | Beads | Cells | Binding | Beads | Cells | Binding |
| 1.30.30 | 1.30.30 | + | 1.30.30 | 7.27.25 | − | 7.27.25 | 7.27.13 | − |
| 7.1.25 | 7.1.25 | + | 1.30.30 | 11.31.25a | − | 7.27.25 | 10.27.25 | − |
| 7.27.13 | 7.27.13 | + | 1.30.30 | 12.20.19a | − | 7.27.25 | 11.31.25 | − |
| 7.27.19 | 7.27.19 | + | 7.1.25 | 7.27.25 | − | 7.27.25 | 12.20.19a | − |
| 7.27.20 | 7.27.20 | + | 7.1.25 | 10.27.25a | − | 10.27.25 | 7.1.25a | − |
| 7.27.21 | 7.27.21 | + | 7.27.13 | 7.27.25 | − | 10.27.25 | 7.27.25 | − |
| 7.27.25 | 7.27.25 | + | 7.27.19 | 7.27.25 | − | 11.31.25 | 1.30.30a | − |
| 7.27.26 | 7.27.26 | + | 7.27.20 | 7.27.25 | − | 11.31.25 | 7.27.25 | − |
| 10.27.25 | 10.27.25 | + | 7.27.21 | 7.27.25 | − | 12.20.19 | 1.30.30a | − |
| 11.31.25 | 11.31.25 | + | 7.27.25 | 1.30.30 | − | 12.20.19 | 7.27.25a | − |
| 12.20.19 | 12.20.19a | + | 7.27.25 | 7.1.25 | − | 7.27.26 | 7.27.25 | +b |
Denotes isoform binding was only tested in bead binding-to-cells assay.
Binding observed in bead binding-to-cells assay; no binding observed in pull-down assay from S2 cell extract.
Dscam Binding Maps to a Region Containing the Variable Ig Domains
As a first step toward assessing whether different isoforms bind to each other, we tested whether the region of the extracellular domain required for binding contained the variable Ig domains. A series of C-terminal truncations of the extracellular domain of Dscam tagged with Fc (Figure 4A) was examined for bead aggregation (Figures 4B and 4C), bead binding to cells (Figure 4E), and the ability to pull down full-length Dscam from cell extracts (Figure 4F). Following the aggregation assay, immunoblotting confirmed that the level of purified protein on beads was equivalent (Figure 4D). Proteins containing the eight N-terminal Ig domains (i.e., EC8 and EC10) exhibited binding largely indistinguishable from the entire extracellular domain (EC16) in all three assays (Figures 4B, 4C, 4E, and 4F). By contrast, smaller fragments containing three, four, or six N-terminal Ig domains (i.e., EC3, EC4, and EC6) did not bind (Figures 4B, 4C, 4E, and 4F). Of the deletion mutants we tested, the smallest protein that exhibited binding contained all three variable Ig domains. This raised the possibility that variable Ig domains may influence binding specificity.
Figure 4. Dscam Binding Maps to a Region Containing the Variable Ig Domains.
A C-terminal deletion series of the extracellular domain was tested for bead aggregation and binding to full-length Dscam. (A) Schematic representation of Dscam-Fc deletions used and summary of binding data. Red, blue, and green lines indicate variable regions in Ig2, Ig3, and Ig7, respectively. EC indicates extracellular domain. The number following EC denotes the number of domains (i.e., Ig [horseshoe] and FNIII [rectangle]) in each protein. The icon at the C terminus symbolizes the Fc tag. (B) Beads containing the eight N-terminal Ig domains promote aggregation. Fluorescent beads were decorated with different Dscam-Fc deletions, and a time course of bead aggregation over 30 min was monitored by FACS analysis. Proteins containing the N-terminal eight Ig domains (red) show binding indistinguishable from the entire extracellular domain. Proteins containing six or fewer N-terminal Ig domains (blue) did not support bead aggregation. Beads were decorated using equimolar amounts of protein. Each time point represents the mean and SD of two independent binding experiments. Two different preparations of each protein were used. (C) Confocal images of beads containing the proteins indicated at 30 min time point in (B). Beads coated with proteins containing the N-terminal eight Ig domains form aggregates similar in size to beads containing the entire extracellular domain, while beads decorated with proteins containing six Ig domains remained monomeric. (D) (Left panel) Coomassie-stained gel of purified Fc-tagged proteins used in binding assays. (Right panel) Immunoblots show that beads used in the aggregation assay were coated with equivalent levels of the indicated proteins. (E) Beads containing the eight N-terminal Ig domains bind to Dscam-expressing cells. Beads coated with different Dscam-Fc deletions were incubated with Dscam-transfected COS cells. Beads containing eight or more N-terminal Ig domains bound, whereas beads containing six N-terminal Ig domains did not. (F) Dscam-Fc proteins containing the N-terminal eight Ig domains bind to full-length Dscam in cell extracts. Dscam-Fc proteins were incubated with extracts of S2 cells transfected with full-length FLAG-tagged Dscam, and protein complexes were pulled down using protein G Sepharose beads. The ability of Dscam deletions to pull down full-length Dscam-FLAG was assessed by immunoblotting with anti-FLAG antibody. Proteins containing the eight N-terminal Ig domains bound to full-length Dscam, whereas proteins with six or fewer N-terminal Ig domains did not. The flag icon denotes full-length FLAG-tagged Dscam. The level of FLAG-tagged protein in each pull-down is shown in the lower panel.
Different Dscam Isoforms Exhibit Isoform-Specific Binding
To test whether different Dscam isoforms interact with each other, we assessed binding between two isoforms differing in all three variable Ig domains (Figure 5A). These isoforms, chosen arbitrarily from cDNAs isolated from Drosophila S2 cells and embryos, were shown to exhibit homophilic binding (Figure 3). They differed significantly from one another, sharing 56%, 37%, and 37% amino acid identity within the N-terminal halves of Ig2 and Ig3 and all of Ig7, respectively. Beads decorated with the entire extracellular domain of each isoform were tested for binding to transfected COS cells or Drosophila neurons in culture from transgenic animals expressing full-length versions of these isoforms. Surprisingly, only binding of beads to cells expressing the same isoform was detected (Figure 5B). This specificity was also seen in binding experiments using extracts prepared from S2 cells expressing full-length FLAG-tagged Dscam isoforms (Figure 5C). For bead binding to protein in extracts or to proteins expressed on the surface of cells, the thresholds of detection are less than 10% (data not shown) and 5% (see below and Figure 8D) of the level of isoforms binding to themselves, respectively.
Figure 5. Dscam Exhibits Isoform-Specific Binding.
Two Dscam isoforms were tested for binding to each other. (A) Schematic representation of the two isoforms used in this experiment showing amino acid sequence identity between variable Ig domains. The percent identity is indicated. The icon at the C terminus indicates the Fc tag. (B) Beads decorated with one isoform of Dscam-Fc bind only to cells expressing the same isoform. Dscam7.27.25-Fc and Dscam1.30.30-Fc beads (red) were incubated with transfected COS cells and neurons from transgenic flies (green) overexpressing a single Dscam isoform as indicated. (C) Dscam-Fc proteins bind only to the same isoform of full-length Dscam in cell extracts. Dscam7.27.25-Fc and Dscam1.30.30-Fc were incubated with extracts of transfected S2 cells expressing full-length FLAG-tagged versions of these isoforms. The Fc-tagged proteins were captured by protein G Sepharose, and their association with FLAG-tagged Dscam proteins was assessed by immunoblotting with anti-FLAG antibody. The same amount of Dscam-Fc protein (1 μg) was added to each pull-down experiment. The flag icon denotes full-length FLAG-tagged Dscam. The level of FLAG-tagged protein in each extract is shown in the right panel.
Figure 8. Closely Related Isoforms Differing by 7–12 Amino Acids Exhibit Preferential Isoform-Specific Binding.
Binding specificity of a series of nearly identical isoforms was examined. (A) Schematic representation of isoforms tested for binding. All share the same Ig2 and Ig3 domains but differ in Ig7. The alternative Ig7 domains tested are highly related. Ig7.25 and Ig7.26 share the highest sequence identity of any two Dscam isoforms differing in only seven amino acids. The icon at the C terminus indicates the Fc tag. (B) Beads (red) decorated with Dscam-Fc isoforms, as indicated, bind to COS cells (green) expressing the same isoform. (C) Beads with isoforms differing from Dscam7.27.25 show reduced or no binding to COS cells expressing Dscam7.27.25. (D) Quantification of bead particle binding on COS cells. A bead particle may consist of one bead or an aggregate of them. Error bars depict SEM. n, number of cells counted. (E) (Top panel) Each Dscam-Fc isoform binds to full-length versions of the same isoform tagged with FLAG in pull-downs from extracts of transfected S2 cells. (Bottom panel) Dscam-Fc isoforms differing from Dscam7.27.25-Fc do not bind to FLAG-tagged Dscam7.27.25. Equivalent levels of Dscam-Fc protein (1 μg) and extract containing Dscam7.27.25-FLAG were added to each pull-down. Arrowheads indicate variable Ig7 domains. Flag icon denotes full-length FLAG-tagged Dscam.
To further investigate the specificity of isoform binding, we utilized a competition assay. A time course of Dscam bead aggregation for these two different isoforms was monitored by FACS analysis in the presence of increasing amounts of the same or a different isoform. Competitor proteins contained the N-terminal eight Ig domains, were monomeric, and were produced in a baculovirus expression system with a 6x-histidine tag at their C termini (Dscam EC8-6xHis). A time course of bead aggregation for both Dscam isoforms in the presence of the same and a different Dscam competitor was determined. Increasing amounts of the same competitor isoform interfered with the ability of Dscam beads to aggregate in a dose-dependent manner (Figure 6). Inhibition was first detected between 30 and 100 μg/ml and was complete by 3 mg/ml. Conversely, no inhibition was detected with up to 3 mg/ml of a different isoform (Figure 6). Each competitor was shown to be specific, as it inhibited binding of beads decorated with the same isoform but did not affect aggregation of beads containing a different isoform.
Figure 6. Competition Assays Reveal Isoform-Specific Binding.
Aggregation of Dscam EC16-Fc-coated beads was monitored in the presence of increasing amounts of soluble competitor, Dscam EC8-6xHis protein, of the same or a different isoform, as indicated. Beads were incubated with agitation, and a time course of aggregation over 45 min was monitored by FACS analysis. Bead aggregation for both Dscam isoforms was inhibited by the presence of the same competing isoform in a dose-dependent manner. Inhibition of bead aggregation was first detected between 30 and 100 μg/ml competitor and was complete by 3 mg/ml competitor. Bead aggregation for both Dscam isoforms was unaffected by competition with up to 3 mg/ml of the different isoform. Each time point represents the mean and SD of two independent binding experiments. Two different preparations of each Dscam-Fc protein were used.
In summary, two different Dscam isoforms exhibited isoform-specific binding. Each isoform bound to itself but not to the other isoform. In subsequent experiments, described below, we tested interactions between additional isoforms that share greater amino acid sequence identity to each other than the two isoforms tested above. These studies allowed us to identify the regions of Dscam that confer specificity. In all cases tested, each isoform bound to itself but showed little if any binding to other isoforms. A summary of interactions between different isoforms is presented in Table 1.
Variable Domains Ig2, Ig3, and Ig7 Determine Binding Specificity
Binding specificity may be determined by identity within all three of the variable Ig domains. Alternatively, identity within two or even a single variable domain may be sufficient for binding. To assess whether each variable domain is necessary for binding specificity, we tested binding between isoforms that differed from each other in only one variable Ig domain. Different isoforms of Dscam-Fc (Figure 7) were tested for binding to the same or different isoforms of full-length Dscam expressed on the surface of COS cells and in S2 cell lysates. No binding was detected between isoforms differing only in the N-terminal half of Ig2 (Figure 7A). The variable Ig2 domains in these isoforms share 50% amino acid sequence identity. Similarly, no binding was detected between isoforms differing only in the N-terminal half of Ig3 (Figure 7B). These variable Ig3 domains share 45% amino acid sequence identity. Finally, no binding was detected between isoforms differing only in Ig7 (Figure 7C). These Ig7 domains share 33% sequence identity. Hence, binding studies conducted with a set of isoforms differing only in one variable Ig domain demonstrated that all three variable Ig domains contribute to isoform-specific binding.
Figure 7. Each Variable Ig Domain Contributes to Binding Specificity.
To investigate which variable Ig domains mediate binding specificity, a series of swapping experiments was conducted. (A) Variable domain Ig2 contributes to isoform-specific binding. (Top panel) Schematic representation of isoforms tested for binding. These isoforms have different N-terminal halves of Ig2 (50% amino acid sequence identity) but are otherwise identical. The icon at the C terminus indicates the Fc tag. (Middle panel) Red fluorescent beads decorated with Dscam-Fc proteins were incubated with transfected COS cells (green) expressing full-length versions of the same or a different Dscam isoform, as indicated. Identical isoforms bound, while isoforms differing in Ig2 did not bind to each other in this assay. (Bottom panel) Dscam-Fc proteins were incubated with extracts of S2 cells expressing the same or a different isoform of full-length FLAG-tagged Dscam. The association of full-length Dscam-FLAG with Dscam-Fc in protein G Sepharose pull-downs was assessed by immunoblotting with anti-FLAG antibody. The same amount of Dscam-Fc protein (1 μg) was added to each pull-down experiment. Only identical Dscam isoforms bound to each other in these assays. (B and C) Variable domains Ig3 and Ig7 contribute to isoform-specific binding. Isoforms differing in Ig3 (B) and isoforms differing in Ig7 (C) were analyzed as described in (A). Identical isoforms bound to each other but did not bind to isoforms differing in either Ig3 or Ig7. The gray arrowheads indicate the variable Ig domain swapped in each experiment. The flag icon denotes full-length FLAG-tagged Dscam.
Closely Related Isoforms Exhibit Preferential Isoform-Specific Binding
We tested interactions between nearly identical isoforms of Dscam to assess how closely related isoforms have to be to bind to one another. One Dscam isoform, referred to as Dscam7.27.25, was tested for binding to a panel of four isoforms that share the highest amino acid sequence identity, 86%–92%, in variable domain Ig7 (Figure 8A). This corresponds to a difference in seven to twelve amino acids in Ig7; all other domains are identical. Different Dscam-Fc isoforms were tested for binding to Dscam7.27.25 on COS cells and in detergent-solubilized S2 cell extracts (Figure 8). Robust binding of each isoform to COS cells expressing a full-length version of the same isoform was observed (Figure 8B and 8D white bars; 47–56 beads/cell, n = 46–77 cells analyzed). Similarly, each isoform bound to itself in S2 cell extracts (Figure 8E, top panel). Conversely, isoforms sharing 86%–90% amino acid sequence identity in Ig7 to Dscam7.27.25 did not bind to this isoform in either assay (Figures 8C and 8E, bottom panel, and Figure 8D, black bars; ∼1 bead/cell, n = 78–106 cells analyzed). Although reduced bead binding to cells was observed for an isoform sharing 92% identity (Figures 8C and 8D, gray bar; 26 beads/25 cells; 17 beads/cell, n = 107), no binding between these isoforms was observed in S2 cell extracts (Figure 8E, bottom panel). In summary, even highly related isoforms exhibit homophilic binding specificity. Each isoform tested binds to itself but poorly, if at all, to other isoforms.
Discussion
We previously reported that alternative splicing at the Drosophila Dscam locus potentially generates 38,016 Ig-containing cell surface proteins sharing the same domain structure. This includes 19,008 distinct extracellular domains linked to one of two alternative transmembrane segments. Here we demonstrate that a set of 11 different Dscam isoforms show surprising homophilic binding specificity. Each isoform preferentially binds to itself over different isoforms. Should this binding property extend to the entire spectrum of Dscam isoforms, this would provide enormous potential for regulating interactions between neurites during the establishment of neuronal connections.
The Variable Ig Domains of Dscam Determine Binding Specificity
Several lines of evidence support the view that Dscam proteins on opposing cell surfaces bind to each other. Mammalian Dscams have been shown to promote cell aggregation when transfected into cultured mouse cells (Agarwala et al., 2000, 2001). Here, we show that Dscam mediates interactions between cells in vivo. The trajectory of interneurons overexpressing a single isoform of Dscam is disrupted upon encountering midline cells that also overexpress the same Dscam isoform. That this reflects direct interactions between Dscam proteins on opposing cell surfaces is supported by the biochemical experiments presented in this paper. Dscam binding was localized to the N-terminal eight Ig domains. As this region contains the three variable Ig domains, it raised the possibility that differences within these domains could modulate interactions between isoforms.
All Dscam isoforms we tested exhibited preferential binding to self over other isoforms. Isoforms differing in any one of the three variable Ig domains did not bind to each other, or showed marked differences in binding. While binding between different isoforms was undetectable in the pull-down assay from S2 cell extracts, the possibility remains that weak interactions between different isoforms exist that are below the limit of detection in this assay. Quantification of the sensitivity of the pull-down assay demonstrated that 10-fold less protein on the Western blots would not have been reliably detected. Therefore, if heterophilic binding occurs between the isoforms we tested, it is significantly weaker than the isoform-specific homophilic interaction. That interactions between different isoforms do occur under milder conditions (e.g., no detergent) is underscored by the binding of two isoforms of Dscam differing in seven amino acids in the bead binding-to-cells assay. Hence, we speculate that, while each isoform preferentially binds to itself, isoforms also exhibit a range of weaker binding interactions with other isoforms.
All three variable Ig domains played a crucial role in binding specificity; swapping any one resulted in a marked reduction or a complete loss of binding. It is anticipated that future biochemical and structural studies will provide insights into the molecular basis of isoform-specific recognition. In the meantime, the simplest model for the “matching” of alternative Ig domains is that each variable Ig domain interacts with the same variable Ig domain in an opposing molecule (Figure 9A). The binding of all three Ig domains is likely required to stabilize otherwise weak interactions between individual variable domains.
Figure 9. A Model for Dscam-Mediated Interactions.
(A) Schematic representation of homophilic binding mediated by the three variable Ig domains. The eight N-terminal Ig domains (circles) are shown. Constant Ig domains are gray, and variable Ig domains are in color. The remainder of the protein is represented by a gray rectangle. Isoforms sharing identical variable Ig domains (represented by matching colors) bind to each other, while isoforms differing in only one variable Ig domain (gray arrowheads) do not. We propose that each variable Ig domain binds to the same variable Ig domain in an opposing molecule. As isoforms sharing any two identical variable Ig domains do not bind or exhibit reduced binding relative to binding between identical isoforms, it is likely that the binding of all three variable Ig domains is required to stabilize otherwise weak interactions between individual variable Ig domains. (B) Schematic representation of Dscam-mediated interactions between neurites. We propose that differences in levels of Dscam signaling influence the nature of the interactions between neurites. High levels of signaling between neurites expressing the same array of Dscam isoforms result in contact-dependent repulsion (left panel), while low levels of signaling between neurites expressing some of the same isoforms or isoforms that bind weakly to each other result in adhesion (middle panel). Some neurites may express isoforms of Dscam that do not interact (right panel). Dscam-mediated contact-dependent repulsion and adhesion may regulate interactions between neurites during axon guidance, targeting, or synapse formation.
Interactions between Isoforms of Dscam May Play an Important Role in Regulating the Patterning of Axonal Projections
How might isoform-specific binding contribute to wiring the fly brain? While the notion that each neuron may express one or only a few isoforms that specify interactions with other neurons is attractive, recent studies argue that single neurons express multiple isoforms and that even neurons of the same class express different and largely nonoverlapping sets of them (Neves et al., 2004; Zhan et al., 2004). Hence, it is highly unlikely that any two neurons will express an identical set of isoforms. This ensures that the only neurites that express an identical set of isoforms are those from the same neuron.
Studies on developing mushroom body (MB) neurons provide support for the view that interactions between identical isoforms play a crucial role in mediating interactions between two neurites of the same cell (Zhan et al., 2004). MB neurons extend axons that bifurcate at a common branch point, and the resulting sister branches segregate to different pathways (Wang et al., 2002). A prominent feature of the loss-of-function phenotype in these neurons is a failure of sister branches to segregate. A simple model to account for this is that identical isoforms of Dscam on sister branches bind to each other and induce a contact-dependent repulsive interaction (Figure 9B) perhaps analogous to repulsive interactions between Eph receptors and ephrin ligands (Hattori et al., 2000). Prior to bifurcation, MB axons project together within a fascicle, and, following bifurcation, each sister branch also extends within a fascicle with other MB axon branches. To allow fasciculation, it is likely important that the array of Dscam isoforms expressed on each MB axon is different from its neighbors. Indeed, expression analysis reveals that, as with other neuronal subclasses (Neves et al., 2004), individual MB neurons express multiple isoforms and largely nonoverlapping arrays of them (Zhan et al., 2004). Hence, while the specific isoforms of Dscam expressed in MB neurons may be unimportant, it may be crucial that neighboring MB axons express different isoforms. In support of this view, expression of a single Dscam isoform in multiple MB neurons induces a dominant phenotype characterized by defasciculation of MB axons (Zhan et al., 2004), whereas expression of a single isoform in a single mutant neuron rescues the defect in the segregation of sister branches (Zhan et al., 2004; Wang et al., 2004). The notion that interactions between identical isoforms induces a repellent response is consistent with other loss- and gain-of-function studies (Hummel et al., 2003; Schmucker et al., 2000; Wang et al., 2002).
It seems unlikely that Dscam acts only in a cell-autonomous fashion to mediate interactions between processes of the same neuron. In the absence of Dscam, defasciculation of axons has been observed both in the developing mushroom body (Zhan et al., 2004) and in Bolwig's nerve (Schmucker et al., 2000). Perhaps weaker signals resulting from interactions between different isoforms or between a small fraction of identical isoforms expressed on different neurons may promote adhesive interactions leading to fasciculation (Figure 9B). Interestingly, recent studies have argued that different levels of Eph/ephrin signaling result in qualitatively different responses; high levels induce contact-dependent repulsion, while lower levels promote contact-dependent attraction (Hansen et al., 2004; McLaughlin et al., 2003b).
In summary, we propose that the nature of the interactions between Dscam isoforms on the surface of neurites produces qualitatively or quantitatively different intracellular signals influencing the development of neurites. Signaling may be modulated by the number of identical isoforms shared by two neurites, the level of expression of each isoform, and the binding affinity or avidity of different isoforms. For instance, high signaling levels produced by interactions between neurites of the same neuron expressing an identical array of Dscam isoforms would induce repulsion (Figure 9B). Conversely, lower signals produced by weaker interactions between neurites of different cells expressing few or no identical Dscam isoforms would promote growth along one another, thereby allowing fasciculation (Figure 9B). As other Ig superfamily proteins have been shown to interact with multiple proteins, it remains possible that other Dscam phenotypes may reflect interactions with additional cell surface or soluble ligands that may or may not exhibit isoform-specific interactions.
Alternative Splicing of Dscam Is Conserved in Insects but Not in Vertebrates
Alternative splicing of Dscam has been highly conserved over some 250 million years separating the fly, the mosquito (Holt, 2002), and the bee (Graveley et al., 2004). This observation, combined with the biochemistry reported here and genetic data establishing a role for Dscam in neuronal connectivity (Hummel et al., 2003; Schmucker et al., 2000; Wang et al., 2002; Zhan et al., 2004), supports the hypothesis that Dscam isoforms function as molecular tags contributing to the formation of precise patterns of neuronal connections.
While Dscam diversity has been highly conserved during insect evolution, the mouse and human Dscam genes do not undergo extensive alternative splicing (Agarwala et al., 2001; Yamakawa et al., 1998). In addition to Dscam, there are a number of genes in the fly genome with arrays of three or more alternatives for a given exon that encode related amino acid sequences. It is striking, however, that no mammalian genes appear to share a similar arrangement, although genes containing only two alternatives for a given exon are common in the mammalian genome (B. Graveley, personal communication). This suggests that diversification of gene function in the mammalian genome has not occurred through the massive cassette-like strategy utilized to generate biochemically distinct isoforms of Drosophila Dscam. Other mechanisms may have evolved in mammals to generate comparable diversity in neuronal cell surface proteins. These may include the use of large families of related proteins encoded by separate genes (e.g., odorant receptors [Wang et al., 1998]), smaller families of proteins used in a combinatorial fashion (e.g., CNRs [Wu and Maniatis, 1999], MHC class I [Huh et al., 2000], classical cadherins [Price et al., 2002]), gradients of receptors and ligands (e.g., Ephrins and Eph receptors [Flanagan and Vanderhaeghen, 1998]), or a combination of multiple genes, alternative transcription start sites, and alternative splicing, as in the case of neurexins (Ullrich et al., 1995; Missler and Sudhof, 1998).
Conclusion
Dscam plays a widespread role in regulating the formation of neuronal connections in Drosophila. Recent expression studies revealed that different neurons express different combinations of Dscam isoforms endowing each neuron with a discrete molecular identity. The biochemical studies described here demonstrate that different Dscam isoforms have striking differences in binding specificity. We propose that a general function of Dscam diversity is to promote repellent interactions between neurites from the same cell expressing the same array of Dscam isoforms in a cell-autonomous fashion. Differences in the arrays of isoforms expressed in different neurons may also contribute to the patterning of neuronal connections. Whether Dscam diversity is indeed crucial to patterning neuronal connections in flies awaits additional analyses in which the number and type of Dscam isoforms expressed in different neurons are systematically manipulated.
Experimental Procedures
Genetics and Histology
eg-Gal4, sim-Gal4, and eg-kinesin lacZ lines were generously provided by John Thomas (Salk Institute). Since GFP expression in axons of eg interneurons was difficult to score when GFP was also present in midline cells, the eg-kinesin lacZ marker was used to visualize these neurons.
Expression of Dscam in Midline Cells
To test whether the trajectory of eg interneurons was affected by Dscam expression at the midline, sim-Gal4/CyO Wg-lacZ; sim-Gal4/sim-Gal4 flies were crossed to w; UAS-Dscam; eg-kinesin lacZ/TM2 flies. Embryos were stained with mouse anti-lacZ (1:200; Promega) and rabbit anti-Dscam (1:1000) (gift of C. Worby and J. Dixon). These embryos were then stained with anti-mouse Cy3 (1:400; Jackson Labs) and anti-rabbit 488 Alexaflour (1:400; Molecular Probes) secondary antibodies. Stage 16 embryos lacking Wg-lacZ were analyzed.
Expression of Dscam in eg Interneurons
To generate embryos expressing Dscam in eg interneurons, flies carrying w; Bl/CyO Wg-lacZ; eg-Gal4 UAS mCD8 GFP eg-kinesin lacZ/TM2 were crossed to w; UAS-Dscam; TM2/TM6b flies. Embryos were stained with anti-lacZ as above.
Expression of Dscam in Both eg Interneurons and Midline Cells
To generate embryos expressing Dscam in both eg interneurons and midline cells, flies carrying w; UAS-Dscam/CyO Wg-lacZ; eg-Gal4 UAS mCD8 GFP eg-kinesin lacZ/TM2 were crossed to a line carrying sim-Gal4/CyO Wg-lacZ; sim-Gal4/sim-Gal4. Embryos were stained with anti-lacZ as above and with rabbit anti-GFP (1:2000; Molecular Probes). Embryos lacking Wg-lacZ and carrying GFP were analyzed for midline crossing.
Plasmid Construction
Dscam expression constructs were derived from the original full-length cDNA encoding alternative exons 4.1, 6.30, 9.30, and 17.2 or 4.7, 6.27, 9.25, and 17.1. Dscam constructs containing different alternative sequences were generated by replacing the alternative sequence in the cDNA with the desired alternative sequence from a library of Dscam RT-PCR clones. Replacement of a KpnI/SgrAI fragment was used to exchange an alternative exon 4 and 6 pair; alternative exon 9 was varied by swapping XhoI fragments and a NarI fragment for alternative exon 17.
FLAG-tagged versions of full-length Dscam were created by inserting two tandem repeats of two annealed oligonucleotides (5′-CTAGGGATTACAAGGATGACGACGATAAGG and 5′-CTAGCCTTATCGTCGTCATCCTTGTAATCC) into an engineered XbaI site that was introduced after the 101st nucleotide of exon 22. Full-length Dscam cDNAs were inserted into pUAST for Schneider2 (S2) cell expression and pcDNA3 for COS cell expression.
Dscam-Fc constructs for S2 cell expression were prepared in the pIB/Fc vector. pIB/Fc was generated by inserting a PCR fragment containing the Fc region of human IgG into XhoI/XbaI of the pIB/V5-His vector in which the SpeI polylinker site had been previously removed. The Fc PCR fragment inserted into pIB/V5-His contained an engineered SpeI site, internal to the XhoI site, for in-frame upstream subcloning of Dscam extracellular domain regions. Some Dscam-Fc expression constructs were derived from full-length Dscam constructs, described above, using PCR to amplify the entire extracellular domain or smaller regions and subsequent subcloning into pIB/Fc via NotI/SpeI sites. Alternatively, some Dscam-Fc constructs were prepared by replacement of fragments in pIB/Dscam7.27.25-Fc, as described above. A detailed description of how each construct was generated is available upon request.
Protein Production
Dscam-Fc proteins were purified as follows from Drosophila S2 cells. Cells were seeded in 15 cm culture dishes at a density of 5 × 107 per dish in serum-free medium (GIBCO Invitrogen Corporation) and transfected with 5 μg pIB/Dscam-Fc using Effectene transfection reagent (Qiagen). Cells were incubated at 25°C and chimeric proteins accumulated in the medium for 4 days, at which time the medium was harvested, spun 40 min at 16,000 rpm (4°C), and filtered (0.2 μm, low protein binding). DCC-Fc protein was purified as follows from 293T HEK cells: cells were plated in 15 cm culture dishes at a density of 1 × 107 per dish in DMEM/10% FCS and incubated overnight at 37°C and 5% CO2. Cells were transfected with 10 μg pCEP4/DCC-Fc (gift of Elke Stein) using FuGENE 6 transfection reagent (Roche Diagnostics Corporation). After 72 hr, the medium was harvested as above.
For all proteins, cleared medium was incubated overnight at 4°C with protein A Sepharose beads (RepliGen Corporation) to capture Fc-tagged protein. Protein A Sepharose beads were washed with 1X PBS (pH 7.4) and applied to a chromatography column, and Fc-tagged proteins were eluted with 100 mM glycine (pH 2.8) and subsequently neutralized using 1M Tris (pH 8). Protein concentrations were estimated using the Bio-Rad protein microassay and SpectraMax 340 PC microplate spectrophotometer (Molecular Devices). Proteins were analyzed by Coomassie blue staining of SDS-polyacrylamide gels.
Dscam EC8 proteins containing a 6xHis tag (Dscam EC8-6xHis) were produced using a baculovirus expression system and obtained from Peter Snow of the Protein Expression Facility at the Beckman Institute at Caltech.
Bead Aggregation Assay
Mouse anti-human IgG Fc (Chemicon International) was passively adsorbed to red fluorescent sulfate polystyrene microspheres (beads, 0.39 μm, Duke Scientific Corporation) with slight modifications to a manufacturer protocol (Bangs Laboratories, Inc., Technote #204). Two preparations of antibody-coupled beads were used in these experiments. Briefly, 125 μl of beads (2% solids) were washed with PBS (pH 7.4) and incubated with 0.2 mg antibody overnight at 4°C in the dark on a Nutator. Beads were subsequently washed with PBS/5% FCS and incubated 2 hr at 4°C with gentle mixing. Final washing was performed with PBS/0.1% BSA, and beads were stored in PBS/0.1% BSA at 4°C.
For the bead aggregation assay, 1 μl of IgG-coupled beads was coated with purified Fc-tagged protein (750 nM in a volume of 100 μl) in PBS/0.1% BSA overnight at 4°C with gentle agitation. Following capture of chimeric Fc proteins, beads were dispersed into monomeric particles by sonication for 5 min in an ice bath cup horn in polystyrene tubes. The beads were incubated at 25°C with shaking (1400 rpm) for 30 min. At various time points, 2 μl aliquots of beads was removed into 150 μl ice cold PBS, and 20,000 events were analyzed by FACS. Competition bead aggregation assays were performed as above with the following modifications: IgG-coupled beads were coated with 2 μg purified Dscam EC16-Fc protein, and increasing amounts of purified Dscam EC8-6xHis (30 μg/ml–3 mg/ml) were added to the reactions.
All FACS analyses were performed on a Beckton Dickinson FACS-can apparatus with CellQuest software. At the 30 min time point in Figures 3 and 4, 5 μl of beads were removed, spotted on a glass microslide, and visualized by confocal microscopy. Following FACS analysis, beads were pelleted, washed twice with PBS/0.1% BSA, boiled in SDS sample buffer, and analyzed by SDS-PAGE with subsequent immunoblotting using an anti-Fc antibody to assess levels of Fc-tagged proteins on beads.
Bead Binding to Cell Assay
COS-7 cells were seeded in 35 mm glass-bottom microwell dishes (MatTek Corporation) at a density of 1 × 105 per dish in DMEM/10% FCS and incubated overnight at 37°C and 5% CO2. Cells were transfected with 1 μg DNA (1:1 pGreenLantern1 and pcDNA3/Dscam) using FuGENE 6 transfection reagent.
To generate cultured neurons overexpressing a single isoform of Dscam, a mushroom body driver, OK107-Gal4, was used in combination with UAS mCD8 GFP and UAS-Dscam. Two copies of UAS mCD8 GFP were required to get efficient GFP fluorescence. The stock w; Bl/CyO; UAS mCD8 GFP; OK107 was crossed to w; UAS Dscam; UAS mCD8 GFP to generate third instar larvae expressing GFP and Dscam in the brain. The brain and ventral nerve cord were dissected from these animals in L-15 media containing 5% FBS and 1X penicillin/streptomycin. Cells were dissociated by incubation of 10 brains in 500 μl media containing 10 μl of liberase (Roche) for 15 min at room temperature and then triturated with a pulled Pasteur pipet until tissue clumps disappeared. Cells were centrifuged at 4000 rpm for 4 min, media was removed, and cells were resus-pended in 10 μl of media per brain. Ten brain equivalent of culture was plated onto Conconavalin A-coated 35 mm glass-bottom microwell dishes. Cells were incubated at 25°C for 48 hr before bead binding (see below).
COS-7 cells (48–72 hr following transfection) or larval brain neurons (∼48 hr in culture) were incubated with Dscam-Fc-coated red fluorescent beads. Beads were sonicated to disperse them before adding to the media. Beads were incubated with cells for 2 hr in culture media. Media was subsequently removed, and cells were washed with media to remove unbound beads. Bead binding to cells was visualized by confocal microscopy.
Pull-Down Experiments from S2 Extracts
S2 cells (0.5 × 106 in 2.2 ml) were cotransfected with full-length FLAG-tagged Dscam isoforms under the control of UAS using Effectene (Qiagen) and a CuSO4 inducible Gal4 construct (pRMHA3). Protein expression was induced 24 hr posttransfection. Cells were harvested 72 hr postinduction and lysed in RIPA buffer (100 μl) (Clemens et al., 1996). Lysate was then sonicated for 5 s using a microtip sonicator. In the case of experimental samples, 1 μg of purified Dscam-Fc protein was added to 40 μl lysate. For control samples, Fc protein was added. Purified protein was incubated with lysate for 1 hr on ice. Protein G Sepharose (10 μl; Pierce) was added to both experimental and control samples. Samples were incubated with gentle rocking for 1 hr at 4°C. Protein G Sepharose beads were then washed using RIPA buffer (3 × 1 ml), resuspended in SDS-PAGE loading buffer (10 μl), and boiled (10 min). Samples (5 μl) were run on 6% SDS-PAGE gel. Bound protein was detected on immunoblots using a mouse monoclonal anti-FLAG (1:5000; Sigma) primary antibody and a goat anti-mouse (1:5000; Jackson Immuno-Research Laboratories) secondary antibody.
Acknowledgments
We thank Richard Axel, Eddy De Robertis, Ralf Landgraf, Gerry Weinmaster, Owen Witte, and members of the Zipursky lab for comments on an early version of the manuscript and helpful discussion. We thank Tzumin Lee for helpful discussions, for communication of data prior to publication, and for coordinating publication of his in vivo studies (see Neuron, Sept. 5, 2004). We acknowledge Owen Witte for use of the FACScan analyzer, Jon Saxe and Jing Huang for help with protein purification, and Elke Stein for the generous gift of DCC-Fc expression construct. We would like to especially acknowledge Peter Snow for his contribution of Dscam EC8-6xHis protein, and we are saddened by his untimely death. This work was supported by an NIH predoctoral training grant (USPHS National Research Service Award GM07185, W.M.W.), an individual NRSA predoctoral fellowship (F31NS044582 from NINDS, J.J.F.), an NIH postdoctoral training grant in Cellular Neurobiology (5 T32 NS07101, S.S.M.), and a fellowship from the Helen Hay Whitney Foundation (J.C.C.). S.L.Z. is an investigator of the Howard Hughes Medical Institute.
References
- Agarwala KL, Nakamura S, Tsutsumi Y, Yamakawa K. Down syndrome cell adhesion molecule DSCAM mediates homophilic intercellular adhesion. Brain Res. Mol. Brain Res. 2000;79:118–126. doi: 10.1016/s0169-328x(00)00108-x. [DOI] [PubMed] [Google Scholar]
- Agarwala KL, Ganesh S, Tsutsumi Y, Suzuki T, Amano K, Yamakawa K. Cloning and functional characterization of DSCAML1, a novel DSCAM-like cell adhesion molecule that mediates homophilic intercellular adhesion. Biochem. Biophys. Res. Commun. 2001;285:760–772. doi: 10.1006/bbrc.2001.5214. [DOI] [PubMed] [Google Scholar]
- Clemens JC, Ursuliak Z, Clemens KK, Price JV, Dixon J. A Drosophila protein-tyrosine phosphatase associates with an adapter protein required for axonal guidance. J. Biol. Chem. 1996;271:17002–17005. doi: 10.1074/jbc.271.29.17002. [DOI] [PubMed] [Google Scholar]
- De Angelis E, MacFarlane J, Du DS, Yeo G, Hicks R, Rathjen FG, Kenwrick S, Brummendorf T. Pathological missence mutations of neural cell adhesion molecule L1 affect homophilic and heterophilic binding activities. EMBO J. 1999;18:4744–4753. doi: 10.1093/emboj/18.17.4744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinstein P, Mombaerts P. A contextual model for axonal sorting into glomeruli in the mouse olfactory system. Cell. 2004;117:817–831. doi: 10.1016/j.cell.2004.05.011. [DOI] [PubMed] [Google Scholar]
- Flanagan JG, Vanderhaeghen P. The ephrins and Eph receptors in neural development. Annu. Rev. Neurosci. 1998;21:309–345. doi: 10.1146/annurev.neuro.21.1.309. [DOI] [PubMed] [Google Scholar]
- Graveley BR, Kaur A, Gunning D, Zipursky SL, Rowen L, Clemens JC. The organization and evolution of the Dipteran and Hymenopteran Dscam genes. RNA. 2004 doi: 10.1261/rna.7105504. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen MJ, Dallal GE, Flanagan JG. Retinal axon response to ephrin-as shows a graded, concentration-dependent transition from growth promotion to inhibition. Neuron. 2004;42:717–730. doi: 10.1016/j.neuron.2004.05.009. [DOI] [PubMed] [Google Scholar]
- Hattori M, Osterfield M, Flanagan JG. Regulated cleavage of a contact-mediated axon repellent. Science. 2000;289:1360–1365. doi: 10.1126/science.289.5483.1360. [DOI] [PubMed] [Google Scholar]
- Higashijima S, Shishido E, Matsuzaki M, Saigo K. eagle, a member of the steroid receptor gene superfamily, is expressed in a subset of neuroblasts and regulates the fate of their putative progeny in the Drosophila CNS. Development. 1996;122:527–536. doi: 10.1242/dev.122.2.527. [DOI] [PubMed] [Google Scholar]
- Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR, Wincker P, Clark AG, Ribeiro JM, Wides R, et al. The genome sequence of the malaria mosquito Anopheles gambiae. Science. 2002;298:129–149. doi: 10.1126/science.1076181. [DOI] [PubMed] [Google Scholar]
- Huh GS, Boulanger LM, Du H, Riquelme PA, Brotz TM, Shatz CJ. Functional requirement for class I MHC in CNS development and plasticity. Science. 2000;290:2155–2159. doi: 10.1126/science.290.5499.2155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hummel T, Vasconcelos ML, Clemens JC, Fishilevich Y, Vosshall LB, Zipursky SL. Axonal targeting of olfactory receptor neurons in Drosophila is controlled by dscam. Neuron. 2003;37:221–231. doi: 10.1016/s0896-6273(02)01183-2. [DOI] [PubMed] [Google Scholar]
- Kohmura N, Senzaki K, Hamada S, Kai N, Yasuda R, Watanabe M, Ishii H, Yasuda M, Mishina M, Yagi T. Diversity revealed by a novel family of cadherins expressed in neurons at a synaptic complex. Neuron. 1998;20:1137–1151. doi: 10.1016/s0896-6273(00)80495-x. [DOI] [PubMed] [Google Scholar]
- McLaughlin T, Hindges R, O'Leary DD. Regulation of axial patterning of the retina and its topographic mapping in the brain. Curr. Opin. Neurobiol. 2003a;13:57–69. doi: 10.1016/s0959-4388(03)00014-x. [DOI] [PubMed] [Google Scholar]
- McLaughlin T, Hindges R, Yates PA, O'Leary DD. Bifunctional action of ephrin-B1 as a repellent and attractant to control bidirectional branch extension in dorsal-ventral retinotopic mapping. Development. 2003b;130:2407–2418. doi: 10.1242/dev.00467. [DOI] [PubMed] [Google Scholar]
- Missler M, Sudhof TC. Neurexins: three genes and 1001 products. Trends Genet. 1998;14:20–26. doi: 10.1016/S0168-9525(97)01324-3. [DOI] [PubMed] [Google Scholar]
- Neves G, Zucker J, Daly M, Chess A. Stochastic yet biased expression of multiple Dscam splice variants by individual cells. Nat. Genet. 2004;36:240–246. doi: 10.1038/ng1299. [DOI] [PubMed] [Google Scholar]
- Price SR, De Marco Garcia NV, Ranscht B, Jessell TM. Regulation of motor neuron pool sorting by differential expression of type II cadherins. Cell. 2002;109:205–216. doi: 10.1016/s0092-8674(02)00695-5. [DOI] [PubMed] [Google Scholar]
- Schmucker D, Clemens JC, Shu H, Worby CA, Xiao J, Muda M, Dixon JE, Zipursky SL. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell. 2000;101:671–684. doi: 10.1016/s0092-8674(00)80878-8. [DOI] [PubMed] [Google Scholar]
- Sperry RW. Chemoaffinity in the orderly growth of nerve fiber patterns and connections. Proc. Natl. Acad. Sci. USA. 1963;50:703–710. doi: 10.1073/pnas.50.4.703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tessier-Lavigne M, Goodman CS. The molecular biology of axon guidance. Science. 1996;274:1123–1133. doi: 10.1126/science.274.5290.1123. [DOI] [PubMed] [Google Scholar]
- Uemura T. The cadherin superfamily at the synapse: more members, more missions. Cell. 1998;93:1095–1098. doi: 10.1016/s0092-8674(00)81452-x. [DOI] [PubMed] [Google Scholar]
- Ullrich B, Ushkaryov YA, Sudhof TC. Cartography of neurexins: more than 1000 isoforms generated by alternative splicing and expressed in distinct subsets of neurons. Neuron. 1995;14:497–507. doi: 10.1016/0896-6273(95)90306-2. [DOI] [PubMed] [Google Scholar]
- Wang F, Nemes A, Mendelsohn M, Axel R. Odorant receptors govern the formation of a precise topographic map. Cell. 1998;93:47–60. doi: 10.1016/s0092-8674(00)81145-9. [DOI] [PubMed] [Google Scholar]
- Wang J, Zugates CT, Liang IH, Lee CH, Lee T. Drosophila Dscam is required for divergent segregation of sister branches and suppresses ectopic bifurcation of axons. Neuron. 2002;33:559–571. doi: 10.1016/s0896-6273(02)00570-6. [DOI] [PubMed] [Google Scholar]
- Wang J, Ma X, Yang JS, Zheng X, Zugates CT, Lee C-HJ, Lee T. Transmembrane/juxtamembrane domain-dependent Dscam distribution and function during mushroom body neuronal morphogenesis. Neuron. 2004;43:663–672. doi: 10.1016/j.neuron.2004.06.033. [DOI] [PubMed] [Google Scholar]
- Winberg ML, Mitchell KJ, Goodman CS. Genetic analysis of the mechanisms controlling target selection: complementary and combinatorial function of netrins, semaphorins, and IgCAMs. Cell. 1998;93:581–591. doi: 10.1016/s0092-8674(00)81187-3. [DOI] [PubMed] [Google Scholar]
- Wu Q, Maniatis T. A striking organization of a large family of human neural cadherin-like cell adhesion genes. Cell. 1999;97:779–790. doi: 10.1016/s0092-8674(00)80789-8. [DOI] [PubMed] [Google Scholar]
- Yamakawa K, Huot YK, Haendelt MA, Hubert R, Chen XN, Lyons GE, Korenberg JR. DSCAM: a novel member of the immunoglobulin superfamily maps in a Down syndrome region and is involved in the development of the nervous system. Hum. Mol. Genet. 1998;7:227–237. doi: 10.1093/hmg/7.2.227. [DOI] [PubMed] [Google Scholar]
- Zhan X-L, Clemens JC, Neves G, Hattori D, Flanagan JJ, Hummel T, Vasconcelos ML, Chess A, Zipursky SL. Analysis of Dscam diversity in regulating axon guidance in Drosophila mushroom bodies. Neuron. 2004;43:673–686. doi: 10.1016/j.neuron.2004.07.020. [DOI] [PubMed] [Google Scholar]