A distinctive feature of Trypanosoma brucei and other trypanosomatids is that most of their genes are transcribed into long polycistronic RNAs [1], which are processed into mature monocistronic mRNAs bearing a 39-nucleotide spliced leader (SL) at their 5′ ends that is derived from a capped precursor RNA of ∼140 nucleotides [2; 3]. This trans RNA splicing serves two major functions: in conjunction with 3′ polyadenylation it generates the mature mRNAs from polycistronic primary transcripts, and via the SL it provides a 5′ cap structure for each mRNA [1; 4]. In contrast to cis RNA splicing of introns in other eukaryotes, trans RNA splicing unites exons from two independently transcribed RNAs. trans and cis RNA splicing, however, share several similarities. For example, at least some components of the cis RNA spliceosomes are conserved in T. brucei, and both cis and trans RNA splicing utilize the same general mechanism and require the same general sequence motifs [4]. During an examination of the effect of different 3′ UTRs on gene expression in T. brucei, we noticed differences between bloodstream form (BSF) and procyclic form (PCF) trypanosomes in the addition of SLs to RNAs transcribed from a luciferase reporter gene inserted into the rRNA gene locus. Further inspection revealed that in PCF cells the expected SL addition occurs to precursor luciferase RNA, whereas in BSF cells multiple SL additions occur to the same luciferase RNA that are independent of the 5′ and 3′ UTR sequences.
Fig. 1A depicts the luciferase reporter plasmid, pMP, integrated into the rRNA gene spacer region of the T. brucei genome that was used for the experiments. This plasmid is derived from plasmid pHD496 (kindly provided by C. Clayton), and is used to measure luciferase activity in BSF and PCF cells when various 3′-UTRs are inserted at a BamHI site directly behind the luciferase coding sequence (LUC). An initial northern blot of RNA from BSF cells bearing integrated pMP revealed the presence of multiple RNA species recognized by a LUC probe (Fig 1B). The three bands indicated by arrowheads coincide with the locations of rRNAs and are likely due to entrapment of some luciferase RNA in these rRNAs. The RNA bands indicated by L (long), LUC and S (short) were examined further. To look for potential differential luciferase expression from the same inserted plasmid in BSF and PCF cells of the same transfected cell line, plasmid pMP or its derivatives were first stably transfected into the genome of BSF cells and these cells were then subjected to in vitro differentiation (Fig. 2A) to generate PCF cells bearing the same integrated luciferase reporter construct. The parent pMP plasmid, which contains LUC followed by the ACT intercoding region (called ACT IC and defined as the 3′-UTR, intergenic region and 5′-UTR located between two adjacent actin coding regions in the T. brucei genome), was used as a control since actin is expressed at similar levels in the BSF and PCF stages [5]. A northern blot of luciferase RNA from this integrated parent plasmid (Fig. 2B) again revealed the multi-band pattern of luciferase RNA in BSF cells (lane 3) and showed that it becomes predominantly a single band of luciferase RNA in PCF cells derived from these BSF cells (lane 4).
Fig. 1.
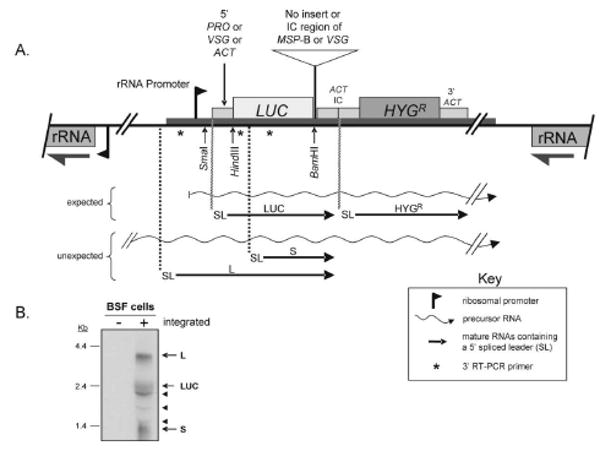
(A) Diagram of the orientation and placement of the luciferase reporter plasmid, pMP, in the rRNA spacer region. The plasmid contains a rRNA promoter (depicted by a flag) followed in succession by the 5′-UTR and upstream poly(Y) tract of the EP-procyclin gene (5′ PRO), the luciferase coding region (LUC), the intercoding (IC) region between two tandem actin coding regions (ACT IC) including the actin 3′-UTR, intergenic region and 5′-UTR, and the hygromycin resistance gene (HYGR) followed by the actin 3′ UTR (3′ ACT). The NotI linearized plasmid integrates into an rRNA spacer region in the genome so that LUC and HYGR are transcribed in the opposite direction than the rRNA genes. PCR-amplified MSP-B IC or 10.1 VSG IC regions were inserted at the indicated unique BamHI site. For the IC region of the 10.1 VSG gene, the region from the stop codon to the first telomeric repeat was used. The 5′-PRO preceding LUC in plasmid pMP was also replaced with the 5′-UTR and upstream poly(Y) tract of 10.1 VSG or ACT using unique SmaI and HindIII restriction sites. As noted in the key, the diagram shows the potential precursor RNAs and the expected and unexpected RNAs containing the 5′-spliced leader (SL). Asterisks (*) indicate the placement of three 3′ primers used with a 5′ spliced-leader (SL) primer to reverse transcriptase-PCR amplify the 5′ ends of RNAs bearing a 5′ SL. Unexpected SL-containing L and S RNAs correspond to the ∼4.0-kb and ∼1.4-kb bands seen in the northern blots shown in Fig. 1B and Fig. 2. L = long luciferase RNA and S = short luciferase RNA. (B) A northern blot probed with LUC showing a multi-band pattern of luciferase RNA in BSF cells bearing the integrated reporter plasmid. The L, LUC, and S RNAs are indicated. The three arrowheads indicate bands thought to result from entrapment of variable amounts of luciferase RNA in the abundant ribosomal RNAs. These bands were repeatedly seen in northern blots of BSF RNA, but varied in intensity among BSF RNA preparations. Similar bands occur in the northern blots shown in Fig. 2.
Fig. 2.
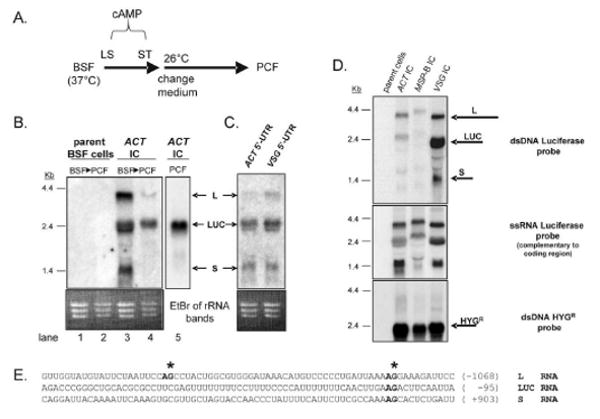
Processing of luciferase RNA is different in BSF and PCF cells. (A) Diagram showing the procedure used for in vitro differentiation of BSF cells to PCF cells. Differentiation is initiated by adding pelleted BSF cells to BSF medium supplemented with 0.4 mM 8-(4-chlorophenylthio)-cAMP, a membrane-permeable derivative of cAMP that induces differentiation from long slender (LS) BSF to short stumpy (ST) BSF [20]. After 48 hr, the medium is changed to PCF medium supplemented with cis-aconitate and citrate, and the BSF-growth temperature of 37°C is dropped to the PCF-growth temperature of 26°C [21]. After 72 hr of growth, the PCF cells are harvested. Northern and western blots not shown demonstrate that the PCF cells have lost VSG mRNA/protein and acquired procyclin mRNA/protein. For unknown reasons these in vitro differentiated PCF cells typically do not survive under the standard PCF growth conditions for more than 2 – 3 weeks. (B) Northern blot probed with the LUC showing that a triple-banding pattern of luciferase RNA from the reporter plasmid in BSF cells becomes predominately a single RNA band in differentiated PCF cells. The ethidium bromide (EtBr) stained gel is shown as a loading control. The parent BSF cell line [22] does not contain the integrated test plasmid (lanes 1 and 2). “ACT IC” refers to parent cells bearing an integrated pMP plasmid with the ACT IC region immediately downstream of LUC (lanes 3 and 4). For comparison, lane 5 contains RNA from a stably transfected PCF culture cell line bearing the same integrated ACT IC region downstream of LUC. The three bands corresponding to the L, LUC, and S RNAs depicted in Fig. 1 are indicated. (C) Northern blot of BSF RNA probed with LUC showing that the three-band pattern of luciferase RNA in stably transfected BSF cells is present when the PRO 5′-UTR upstream of LUC (Figs. 1 and 2B) has been replaced by the 5′-UTR of either the actin (ACT) or 10.1 VSG gene. (D) Northern blot of RNA from BSF cells bearing an integrated plasmid in which the MSP-B IC or 10.1 VSG IC has been inserted at the BamHI site shown in Fig. 1. The top panel shows a northern blot probed with the double-stranded (dsDNA) luciferase coding region. The middle panel shows that when the same RNAs are probed with a single-stranded (ss) RNA luciferase probe complementary to the coding sequence, the same three-band pattern is present. The complementary ssRNA luciferase probe does not detect any of these three RNAs (not shown). The bottom panel shows that a single HYGR RNA of the expected 2.4-kb size is detected when a dsDNA HYGR probe is used. (E) Sequences of the regions surrounding the SL-addition sites that yield the L, Luc, and S RNAs. The SL-addition sites (asterisks), AG dinucleotides involved in splicing (bold) and positions of the sequences relative to the LUC start codon (number) are indicated.
In T. brucei the polyadenylation site at the 3′ end of a 3′-UTR is predominately determined by the location of the active downstream SL acceptor site in the same IC region [6; 7]. The distance between the polyadenylation and SL addition sites is often about 100 - 300 nucleotides, although variation occurs among genes and/or trypanosomatid organisms. A pyrimidine-rich [poly(Y)] tract upstream of the SL addition site in the IC region has been shown to be important in determining the 3′ SL splice site in trypanosomatids [8]. In the case of the well-studied procyclin genes of T. brucei, two IC poly(Y) tracts between the coding regions are used for splicing, suggesting there could be a hierarchy to processing in which one splice site dominates, whereas other(s) are used less frequently [9]. Segments within the 5′-UTR have also been shown in some cases to influence efficiency of trans RNA splicing [10; 11]. These findings suggest that splicing efficiency is influenced by the context of the splice site.
Thus, in the system described here, one possibility is that the multiple luciferase RNAs found specifically in BSF cells might be due to the presence of the PRO 5′-UTR in front of LUC in the pMP plasmid (Fig.1A). PRO mRNA is much more abundant in PCF cells than BSF cells [12]. Using unique restriction sites within the pMP plasmid, the PRO 5′-UTR was replaced with either a VSG-specific or ACT-specific 5′-UTR sequence (Fig. 1A). VSG mRNA is predominately in BSF cells and actin mRNA is in both BSF and PCF cells. A northern blot of RNA from BSF cells containing the integrated pMP plasmid with the ACT and VSG 5′-UTRs showed the presence of the same three LUC RNAs, indicating that the 5′-UTR is not fully responsible for the multiple luciferase RNAs in BSF cells (Fig. 2C). Quantification of the relative abundance of the three BSF luciferase RNA species within each lane (not shown) on the northern blots illustrates that the upper L band was reduced from about 30% of the combined intensities of the three main bands in the lane when the PRO 5′-UTR was present (Fig. 1B and Fig. 2B, lane 3) to 14% when the 5′-UTR of a VSG or actin gene was present (Fig. 2C). In contrast, the L luciferase RNA is only 7% of the total luciferase RNA in the in vitro differentiated PCF cells (Fig. 2B, lane 4). This 7% may indicate incomplete differentiation of the BSF cells to PCF since this L RNA cannot be detected in the stably transfected cultured PCF cell line (Fig 2B, lane 5). The S luciferase RNA was not detected in either the differentiated or cultured PCF cells (lanes 4 and 5).
A second possibility is that the multiple luciferase RNAs in BSF cells are due to the 394-bp ACT IC sequence downstream of LUC. So, two other IC regions were inserted at the BamHI site immediately after LUC in plasmid pMP. One was the 585-bp IC region between two identical MSP-B coding regions in the T. brucei genome. MSP-B encodes the Major Surface Protease of T. brucei and its mRNA occurs with lower abundance than actin mRNA in both BSF and PCF cells [13]. The other was the 307-bp IC region between the 10.1 VSG coding region in its telomere-linked expression site and the first telomere repeat [14]. The highly abundant VSG mRNA occurs exclusively in BSF cells. Fig. 2D shows northern blots of RNA from BSF cells containing the integrated pMP plasmid with either the ACT IC, MSP-B IC or VSG IC following LUC. The blot in the top panel of Fig. 2D was probed with the same LUC probe as was used in the blots of Fig. 1B, 2B and C. The same three main BSF luciferase RNAs were detected, and the overall abundance of these three RNAs varied with the IC, i.e., the three RNAs from the plasmid with a VSG IC were more abundant than the three RNAs from the ACT IC plasmid, which were more abundant than the three RNAs from the MSP-B IC plasmid. This result is consistent with previous demonstrations that the 3′-UTR often influences mRNA stability and abundance in T. brucei [1]. The different sizes of the three luciferase RNAs in each of the three BSF cell lines are consistent with the corresponding sizes of the ACT, MSP-B and VSG 3′-UTRs in each of the three ICs, indicating that polyadenylation at their 3′-ends has occurred correctly. In addition, since these three main luciferase RNAs occur in all three BSF cells lines, the IC sequence appears not to influence their presence. The northern blot in the middle panel of Fig. 2D showed that the three luciferase RNAs are all derived from the LUC and HYGR coding strand, and use of the complementary single-strand LUC probe in a northern blot confirmed that none of these RNAs is derived from the other strand, i.e., the rRNA coding strand (not shown). Finally, the bottom panel in Fig. 2D showed that HYGR mRNA in these BSF cells is present as a predominately single species of the expected 2.4-kb size, indicating that it has been correctly processed at both the 5′- and 3′-ends (see Fig 1A for the location of the HYGR gene downstream of the LUC gene).
The presence of the integrated plasmid in the rRNA spacer region opposite in direction to the transcription of the rRNA gene clusters makes the existence of L luciferase RNA surprising. The integrated plasmid contains an rRNA promoter of its own to drive transcription through the plasmid (Fig. 1A). However, because the HYGR probe does not detect the 4.0-kb L luciferase RNA (Fig. 2D), this L RNA must be derived from RNA whose transcription begins upstream of the plasmid insertion site. In contrast, S luciferase RNA is too small to encode the luciferase protein. To determine the splice sites of both the S and L luciferase RNAs, three 3′ PCR primers were designed to hybridize (i) upstream of the rRNA promoter of the pMP plasmid, (ii) near the 5′ start site of LUC, and (iii) in the middle of LUC (indicated by asterisks in Fig. 1A). Reverse-transcriptase (RT) PCR was performed on RNA from BSF cells (see Fig. 2B, lane 3) using these three 3′ primers and a 5′ SL primer to amplify only mature luciferase RNAs bearing the SL. The PCR amplification products were purified on gels, eluted and cloned in a plasmid, and their sequences determined for each amplification product. These sequences were used to identify the major SL addition sites for the L, LUC and S RNAs (Fig. 2E). For L and S RNA, two or more SL addition sites within about 70 nucleotides of each other were found. Fig. 2E shows two of these sites for L RNA and one site for S RNA. For the LUC RNA, a single predominant SL addition site was found. For all three luciferase RNAs in BSF cells, inspection of the DNA sequence upstream of the SL addition sites revealed the presence of at least one poly(Y) tract, consistent with the observation that trans splicing in T. brucei depends on the presence of an upstream poly(Y) tract [11]. Similar RT-PCR amplification of luciferase RNA in stably transfected PCF cells (see Fig. 2B, lane 5) and subsequent sequencing of the amplification product revealed the same SL addition site for LUC RNA in these cells as in the BSF cells. L and S RNAs were not detected by RT-PCR of RNA in these stably transfected PCF cells.
Thus, the basic experimental observation described here is that precursor luciferase RNA from integrated plasmid pMP is processed into three major SL-containing RNAs in BSF cells and into one predominant SL-containing RNA in PCF cells (Fig. 2B, lanes 3 and 4). The difference in this 3′ splice site choice between BSF and PCF cells is independent of the 5′ and 3′ UTRs flanking LUC, suggesting that BSF cells can recognize some sequences in precursor RNA as a SL-addition site that PCF cells do not. Computer programs have been reported for identifying potential splicing sites and specific mRNA processing signals in T. brucei, which have been useful for genome-wide analysis of trypanosome genes [15], but have not addressed possible trans splicing differences in BSF and PCF cells. Alternative splicing of introns has been shown to be important in generating different gene products from the same gene in various cell types of higher eukaryotes [16] and alternative splicing has been described in trypanosomatids [7; 9; 17-19]. In addition, trans-splicing has been suggested as a potential mechanism for posttranscriptional gene control via differences in trans-splicing efficiency [11], which may be applicable in the system described here. There is also evidence that fully processed non-coding RNAs in T. brucei can be produced by splicing in the intergenic region between open reading frames [9] and whose function, if any, is unknown. Thus, the SL addition sites that generate the L and S RNAs may be more efficiently utilized in BSF cells than in PCF cells, but they are predicted to produce non-functional RNAs. An alternative, less likely, possibility is that these RNAs are made in PCF cells, but are too unstable to detect on northern blots. The luciferase enzymatic activity does not appear to be influenced by the multiple splice sites in BSF cells since its RNA abundances and enzymatic values mirror regulation control by the 3′-UTRs inserted downstream of the luciferase reporter gene (Fig. 2D and not shown). Thus, additional SL addition sites in BSF cells could be aberrant, or they might be utilized by BSF cells to expand the number of gene products from individual genes, similar to alternative intron splicing events in higher eukaryotes. Further research using T. brucei endogenous genes, rather than the exogenously added luciferase gene, is necessary to examine the latter possibility.
Acknowledgments
This work was supported by N.I.H. grants AI059451 and AI007511.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Clayton C, Shapira M. Post-transcriptional regulation of gene expression in trypanosomes and leishmaniasis. Mol Biochem Parasitol. 2007;156:93–101. doi: 10.1016/j.molbiopara.2007.07.007. [DOI] [PubMed] [Google Scholar]
- 2.Laird PW. Trans splicing in trypanosomes--archaism or adaptation? Trends Genet. 1989;5:204–8. doi: 10.1016/0168-9525(89)90082-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Ullu E, Matthews KR, Tschudi C. Temporal order of RNA-processing reactions in trypanosomes: rapid trans splicing precedes polyadenylation of newly synthesized tubulin transcripts. Mol Cell Biol. 1993;13:720–5. doi: 10.1128/mcb.13.1.720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Liang XH, Haritan A, Uliel S, Michaeli S. trans and cis splicing in trypanosomatids: mechanism, factors, and regulation. Eukaryot Cell. 2003;2:830–40. doi: 10.1128/EC.2.5.830-840.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Garcia-Salcedo JA, Perez-Morga D, Gijon P, Dilbeck V, Pays E, Nolan DP. A differential role for actin during the life cycle of Trypanosoma brucei. Embo J. 2004;23:780–9. doi: 10.1038/sj.emboj.7600094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Schurch N, Hehl A, Vassella E, Braun R, Roditi I. Accurate polyadenylation of procyclin mRNAs in Trypanosoma brucei is determined by pyrimidine-rich elements in the intergenic regions. Mol Cell Biol. 1994;14:3668–75. doi: 10.1128/mcb.14.6.3668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vassella E, Braun R, Roditi I. Control of polyadenylation and alternative splicing of transcripts from adjacent genes in a procyclin expression site: a dual role for polypyrimidine tracts in trypanosomes? Nucleic Acids Res. 1994;22:1359–64. doi: 10.1093/nar/22.8.1359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Huang J, Van der Ploeg LH. Requirement of a polypyrimidine tract for trans-splicing in trypanosomes: discriminating the PARP promoter from the immediately adjacent 3′ splice acceptor site. Embo J. 1991;10:3877–85. doi: 10.1002/j.1460-2075.1991.tb04957.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hug M, Hotz HR, Hartmann C, Clayton C. Hierarchies of RNA-processing signals in a trypanosome surface antigen mRNA precursor. Mol Cell Biol. 1994;14:7428–35. doi: 10.1128/mcb.14.11.7428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lopez-Estrano C, Tschudi C, Ullu E. Exonic sequences in the 5′ untranslated region of alpha-tubulin mRNA modulate trans splicing in Trypanosoma brucei. Mol Cell Biol. 1998;18:4620–8. doi: 10.1128/mcb.18.8.4620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Siegel TN, Tan KS, Cross GA. Systematic study of sequence motifs for RNA trans splicing in Trypanosoma brucei. Mol Cell Biol. 2005;25:9586–94. doi: 10.1128/MCB.25.21.9586-9594.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hotz HR, Hartmann C, Huober K, Hug M, Clayton C. Mechanisms of developmental regulation in Trypanosoma brucei: a polypyrimidine tract in the 3′-untranslated region of a surface protein mRNA affects RNA abundance and translation. Nucleic Acids Res. 1997;25:3017–26. doi: 10.1093/nar/25.15.3017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.LaCount DJ, Gruszynski AE, Grandgenett PM, Bangs JD, Donelson JE. Expression and function of the Trypanosoma brucei major surface protease (GP63) genes. J Biol Chem. 2003;278:24658–64. doi: 10.1074/jbc.M301451200. [DOI] [PubMed] [Google Scholar]
- 14.LaCount DJ, El-Sayed NM, Kaul S, Wanless D, Turner CM, Donelson JE. Analysis of a donor gene region for a variant surface glycoprotein and its expression site in African trypanosomes. Nucleic Acids Res. 2001;29:2012–9. doi: 10.1093/nar/29.10.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Benz C, Nilsson D, Andersson B, Clayton C, Guilbride DL. Messenger RNA processing sites in Trypanosoma brucei. Mol Biochem Parasitol. 2005;143:125–34. doi: 10.1016/j.molbiopara.2005.05.008. [DOI] [PubMed] [Google Scholar]
- 16.Matlin AJ, Clark F, Smith CW. Understanding alternative splicing: towards a cellular code. Nat Rev Mol Cell Biol. 2005;6:386–98. doi: 10.1038/nrm1645. [DOI] [PubMed] [Google Scholar]
- 17.Revelard P, Lips S, Pays E. Alternative splicing within and between alleles of the ATPase gene 1 locus of Trypanosoma brucei. Mol Biochem Parasitol. 1993;62:93–101. doi: 10.1016/0166-6851(93)90181-v. [DOI] [PubMed] [Google Scholar]
- 18.Chamond N, Goytia M, Coatnoan N, Barale JC, Cosson A, Degrave WM, Minoprio P. Trypanosoma cruzi proline racemases are involved in parasite differentiation and infectivity. Mol Microbiol. 2005;58:46–60. doi: 10.1111/j.1365-2958.2005.04808.x. [DOI] [PubMed] [Google Scholar]
- 19.Colasante C, Robles A, Li CH, Schwede A, Benz C, Voncken F, Guilbride DL, Clayton C. Regulated expression of glycosomal phosphoglycerate kinase in Trypanosoma brucei. Mol Biochem Parasitol. 2007;151:193–204. doi: 10.1016/j.molbiopara.2006.11.003. [DOI] [PubMed] [Google Scholar]
- 20.Breidbach T, Ngazoa E, Steverding D. Trypanosoma brucei: in vitro slender-to-stumpy differentiation of culture-adapted, monomorphic bloodstream forms. Exp Parasitol. 2002;101:223–30. doi: 10.1016/s0014-4894(02)00133-9. [DOI] [PubMed] [Google Scholar]
- 21.Overath P, Czichos J, Haas C. The effect of citrate/cis-aconitate on oxidative metabolism during transformation of Trypanosoma brucei. Eur J Biochem. 1986;160:175–82. doi: 10.1111/j.1432-1033.1986.tb09955.x. [DOI] [PubMed] [Google Scholar]
- 22.Wirtz E, Leal S, Ochatt C, Cross GA. A tightly regulated inducible expression system for conditional gene knock-outs and dominant-negative genetics in Trypanosoma brucei. Mol Biochem Parasitol. 1999;99:89–101. doi: 10.1016/s0166-6851(99)00002-x. [DOI] [PubMed] [Google Scholar]


