Abstract
Background
Hereditary hemorrhagic telangiectasia (HHT), also known as Rendu-Osler-Weber syndrome, is an autosomal dominant disorder which is clinically characterised by recurrent epistaxis, mucocutaneous telangiectasia and visceral arteriovenous malformations. Genetic linkage studies identified two genes primarily related to HHT: endoglin (ENG) on chromosome 9q33-34 and activin receptor-like kinase1 (ACVRL1) on chromosome 12q13. We have screened a total of 41 unselected German patients with the suspected diagnosis of HHT. Mutation analysis for the ENG and ACVRL1 genes in all patients was performed by PCR amplification. Sequences were then compared to the HHT database http://www.hhtmutation.org sequences of the ENG mRNA (accession no. BC014271.2) and the ACVRL1 mRNA (accession no. NM000020.1).
Results
We identified 15 different mutations in 18 cases by direct sequencing. Among these mutations, one novel ENG mutation could be detected which has not yet been described in the literature before. The genotype-phenotype correlation was consistent with a higher frequency of pulmonary arteriovenous malformations in patients with ENG mutations than in patients with ACVRL1 mutations in our collective.
Conclusion
For rapid genotyping of mutations and SNPs (single nucleotide polymorphisms) in ENG and ACVRL1, allele-specific PCR methods with sequence-specific primers (PCR-SSP) were established and their value analysed.
Background
Hereditary hemorrhagic telangiectasia (HHT, MIM#18730), also known as Rendu-Osler-Weber syndrome, is an autosomal dominant disorder of the fibrovascular tissue. Worldwide, HHT has a prevalence of 1 in 10.000 individuals [1,2]. This multi-systemic angiogenic disorder is clinically characterized by severe and recurrent hemorrhages due to epistaxis, mucocutaneous telangiectasia and arteriovenous malformations. Vascular lesions are present in two forms, either as telangiectases on the skin as well as the inner lining of the mouth, nose and gastrointestinal tract (GI-tract) and/or as direct arteriovenous shunts most commonly seen in the lung (PAVM), brain (CAVM) and liver (HAVM) [3-9]. Electron microscopy studies on lesion biopsies revealed that dilated postcapillary venules connect directly to arterioles without intervening capillary bed [10,11]. As a result, these vascular lesions can be the cause for severe complications involving gastrointestinal bleedings, the risk of embolism, stroke and abscesses [12].
Till now, HHT can not be cured. The treatment of HHT patients remains symptomatic or has preventive character. The Curaçao criteria are in clinical use for diagnosis of HHT [13]. However, the clinical manifestations of HHT can be very heterogeneous, both between families and among members of the same family, making the diagnosis of the disease at times very difficult.
In the 90s, genetic linkage studies revealed the two major types of disease, HHT1 and HHT2, caused by mutations in the ENG (endoglin) and ACVRL1 genes. The first locus (HHT type 1) was mapped to chromosome 9q33-34 [14,15], where the 40 kb endoglin (ENG, OMIM 187300) was defined as the affected gene [16], also associated with a high prevalence of pulmonary arteriovenous malformations [17]. The second locus (HHT type 2) mapping to chromosome 12q was identified as the 15 kb activin receptor-like kinase-1 gene (ACVRL1, OMIM 600376) [18,19]. It is characterised by a lower incidence of pulmonary and cerebral arteriovenous malformations than HHT1, but has a higher incidence of liver manifestation [17,20]. The corresponding endoglin and ALK-1 proteins are specific endothelial receptors of the transforming growth factor beta superfamily essential for maintaining vascular integrity [21,22]. Many mutations have been identified in ENG and ACVRL1 genes and support the haploinsufficiency model for HHT [23]. In recent studies, two more genes have been implicated in HHT, which have not yet been identified, the HHT3 gene on chromosome 5 [24], and the HHT4 gene on chromosome 7 [25]. In addition, SMAD4 mutations mapping to chromosome 18 have been observed in patients with a combined syndrome of juvenile polyposis and HHT (JPHT) [26].
In this study, we report mutational analysis of the two main genes ENG and ACVRL1 in a German population affected by HHT. We describe a novel mutation in the ENG gene in HHT patients which has not been described before. Furthermore, we determine the value of allele-specific PCR for rapid genotyping.
Methods
Patients and controls
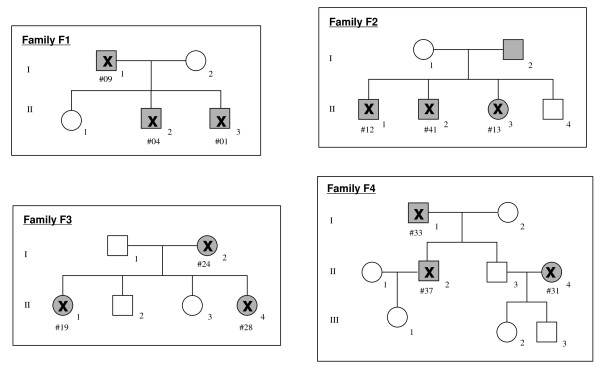
Anti-coagulated peripheral blood samples were obtained from 41 patients of German origin with tentative diagnosis of HHT at the Department of ORL, Head and Neck Surgery, University Hospital Mannheim, and from 768 healthy blood donors as a control group from the Institute of Transfusion Medicine and Immunology, Mannheim (Additional file 1). The HHT patients comprised 4 small-sized families (3 persons each) and 29 single cases (Fig. 1). Information on the clinical data including the medical history, physical examination and diagnostic screening was obtained in personal interviews with the affected individuals and from their patient records with all the data being documented in an HHT database. According to the Curaçao criteria, diagnosis for HHT included the presence of three out of four key symptoms: recurrent epistaxis, telangiectasia, hereditary with an affected first degree relative and visceral involvement [13] (Additional file 2). An informed consent was obtained from each individual and from parents of patients younger than 18 years according to the approval of the Ethic Committee of the University of Mannheim.
Figure 1.
Pedigree of the four small-sized HHT families. White square and white circle = unaffected, grey square and grey circle = affected, X = DNA available, # = Pat.-no.
Isolation of DNA
DNA was extracted from anti-coagulated peripheral blood samples using a commercial DNA isolation kit (QIAamp DNA blood mini kit; Qiagen, Hilden, Germany) according to the standard protocol.
Sequencing of ENG and ACVRL1 exons
All coding exons and flanking intronic sequences of the ENG and ACVRL1 genes were PCR amplified using primers published previously [27,28]. PCR was performed in a total volume of 50 μl including 100 ng DNA, 0.5 μM each primer, 200 μM each dNTP, standard PCR buffer (Qiagen) and 1 U HotStar Taq DNA polymerase (Qiagen). The cycling parameters were as follows: initial denaturation for 15 min at 95°C; 35 cycles with 20 sec. at 94°C, 30 sec. at 60°C and 60 sec. at 72°C; final extension for 5 min at 72°C. After the PCR, products were run on 2% agarose gels. The 15 exons of ENG and the 9 exons of ACVRL1 were sequenced in 41 patients with HHT using Thermosequenase reagents (GE Healthcare Amersham), IRD700 and IRD800 labelled primers and an automated system (NEN® Global IR2 DNA Sequencer; LI-COR inc., Lincoln, NE, USA). Sequences were compared to the HHT database http://www.hhtmutation.org sequences of the ENG mRNA (accession no. BC014271.2) and the ACVRL1 mRNA (accession no. NM000020.1).
PCR-SSP typing of SNPs
For rapid genotyping of the mutations and SNPs in ENG and ACVRL1, allele-specific PCR methods with sequence-specific primers (PCR-SSP) were established. Specific primer sequences and sizes of PCR products are given in Additional file 3. Part of the β-globin locus was amplified as an internal control PCR fragment (540 bp). According to a published PCR-SSP protocol [29,30], 20 ng DNA were subjected to 10 μl PCR reactions including 0.5 μM each allele-specific primers, 0.1 μM each internal control primer, standard PCR buffer (Qiagen), 200 μM each dNTP and 0.5 units Taq DNA polymerase (Qiagen). The cycling conditions were: 2 min initial denaturation at 94°C, followed by 10 cycles with 20 s denaturation at 94°C and 1 min annealing/extension at 65°C, followed by 20 cycles with 20 s duration at 94°C, 1 min annealing at 61°C and 30 s extension at 72°C. Amplification products were separated on 2% agarose gels containing 0.5 ng/ml ethidium bromide in a rapid agarose gel electrophoresis (RAGE; Cascade Biologics, Inc., Portland, OR, USA) chamber for 5 min at 25 V/cm. Results were obtained by visual inspection of the gels and were documented by using a UV documentation device with CCD camera (UVP, Inc., Upland, CA, USA).
Results
The 15 coding exons of the ENG gene and the 9 exons of the ACVRL1 gene including flanking intron sequences were PCR amplified and sequenced from 41 patients with HHT. The sequence data were compared to each other and to the ENG and ACVRL1 sequences published in databases (HHT Mutation Database: http://www.hhtmutation.org). Mutations could be detected in 18 cases (43.9%).
Sequence analysis of ENG
ENG mutations
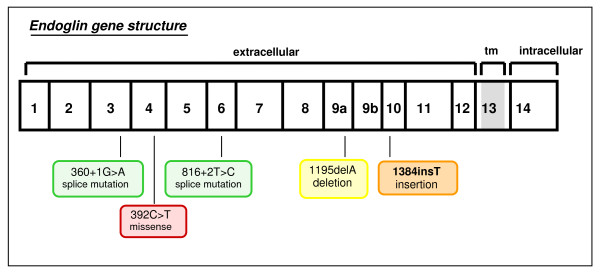
We identified 5 different ENG mutations in 6 out of 41 HHT patients. The 5 different ENG mutations represented: 1 deletion, 1 insertion, 2 splice site mutations and 1 missense mutation (Additional file 4). In our study, mutation c.816+2T>C was detected in 2 related cases. One novel ENG mutation was detected in exon 10 (c.1384insT) causing a frameshift (p.Gln462fs) which has not yet been listed in the HHT gene database before. Altogether, all of the mutations found in ENG were detected in exons coding for the extracellular protein-domain. No mutations were detected in the region coding for the intracellular domain (Fig. 2).
Figure 2.
Distribution of mutations in the ENG gene. Novel mutation is given in bold.
Polymorphisms in the ENG gene
The 5 different polymorphisms and their relative frequencies identified in the ENG gene are presented in Additional file 5. A previously reported G to A substitution at position 207 of exon 2 (c.207G>A) was observed in 12 out of 41 HHT patients (29.3%). Two patients carried a c.1844C>T polymorphism in exon 13 (4.8%). At position 1029 (c.1029C>T) of exon 8, a C to T change was observed in 7 HHT patients (17%). A c.1347A>G polymorphism in exon 10 and a c.1771G>A polymorphism in exon 13 was observed in one HHT patient (2.4%).
Sequence analysis of ACVRL1
ACVRL1 mutations
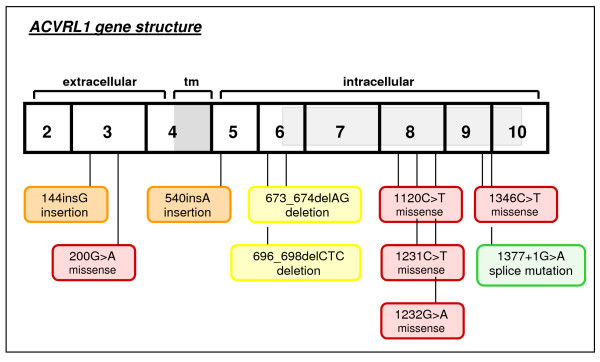
Ten different ACVRL1 mutations could be identified in 12 out of 41 HHT patients, including 2 deletions, 2 insertions, 1 splice site mutation and 5 missense mutations (Additional file 6). The insertion c.144_145insG (p.Ala49fs) was detected in 2 members of the same family, whereas the missense mutation c.1120C>T (p.Arg374Trp) was detected in 2 unrelated cases. Half of the HHT2 associated ACVRL1 mutations consisted of single base substitutions, leading to amino acid changes. More than 75% of all ACVRL1 mutations were detected in exons coding for the intracellular domain (Fig. 3). Especially four out of five missense mutations (80%) were found in the receptor domain or the kinase domain, mainly affecting exon 8. No mutations were detected in the region coding for the transmembrane domain.
Figure 3.
Distribution of mutations in the ACVRL1 gene.
PCR-SSP typing for ENG and ACVRL1 mutations
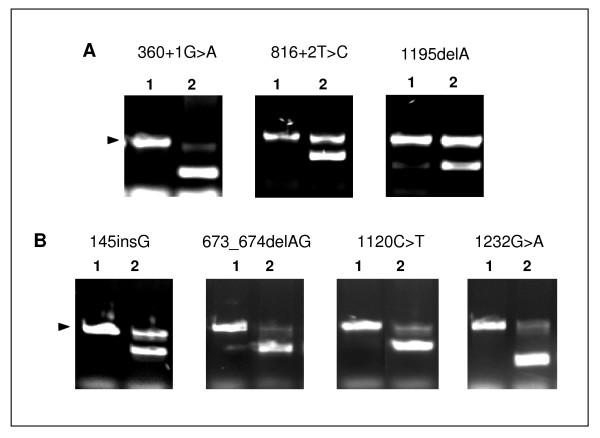
Further investigation on allele and genotype frequencies focused on the mutations identified in our cohort of HHT patients. PCR-SSP systems could be established for all ACVRL1 mutations and all ENG mutations. A panel of DNA samples from HHT patients with different mutations was selected on the basis of the sequencing data and was used to validate all 15 PCR-SSP systems. The genotypes were estimated on the basis of the presence or the absence of the allele-specific PCR products (Fig. 4). All of the 41 HHT samples were retyped by PCR-SSP and data were compared to sequencing data. We could achieve 100% concordance of data, i.e. DNA samples with mutation revealed a positive PCR signal in the corresponding PCR-SSP system and DNA samples without mutation were negative in all 15 PCR-SSP systems. In addition, the 15 PCR-SSP systems were applied for genotyping of the ENG and ACVRL1 mutations in 768 healthy blood donor controls. None of the healthy control individuals showed an ENG or ACVRL1 mutation indicating the specificity of the gene variants for the HHT phenotype.
Figure 4.
Representative results from PCR-SSP analysis of different ENG (A) and ACVRL1 (B) mutations. Three of the 5 PCR-SSP systems for ENG mutations and 4 of the 10 PCR-SSP systems for ACVRL1 mutations are shown as examples. The specificity of PCR-SSP for the corresponding mutations was given by the absence of the specific PCR product in healthy controls (each lane 1) and presence of the PCR product in the HHT patient (each lane 2). The internal control fragment of 540 bp in size (arrow head) was present in each reaction (or at least where the PCR-SSP product was absent).
Phenotype-Genotype Correlation
Diagnosis of HHT in an individual required that they met three or more of the following four Curaçao criteria: 1) spontaneous and recurrent epistaxis, 2) mucocutaneous telangiectasias, 3) organ manifestation with arteriovenous malformations (AVMs) including pulmonary, cerebral, hepatic, gastrointestinal and/or 4) hereditary with a first-degree relative with HHT [13]. All patients included in this study either met the criteria for manifest clinical diagnosis or suspected clinical diagnosis of HHT with at least three or two criteria present. Altogether, 71% of the patients (29 out of 41) had a manifest HHT diagnosis. All of the patients had as main clinical symptom recurrent epistaxis and a positive family history of HHT, followed by telangiectasias in 65.8% and AVMs of any type in 56% of the cases. Of the 41 patients tested, 18 patients were found to have sequence alterations, with 6 patients showing mutations for ENG and 12 patients revealing mutations for ACVRL1. The genotype-phenotype correlation of all HHT patients with confirmed mutation is given in Additional file 7.
Discussion
Presently, ENG (endoglin) on chromosome 9q33-q34 and ACVRL1 (activin-receptor-like kinase) on chromosome 12q13 are the two genes primarily implicated in the development of Hereditary Hemorrhagic Telangiectasia (HHT). In literature, frequent mutations in both genes have been detected in different case studies indicating a common occurrence of ENG and ACVRL1 DNA changes in patients with HHT [27,28].
In a consecutive series of 41 patients, we performed clinical genetic testing for the two major types of disease, HHT1 and HHT2, caused by mutations in the ENG and ACVRL1 gene sequence for HHT. The series included 29 patients with clinical diagnosis of HHT who met three or more clinical features and 12 patients with suspected diagnosis with two clinical features according to the Curaçao criteria. The mutation detection rate in our series of 41 German patients was 44%. Based on the Curaçao criteria, 15 of the 18 patients (83%) with a mutation had a manifest clinical HHT diagnosis, with the remaining 3 patients (17%) fulfilling two criteria. This rate is similar to other studies which have been consistently described in literature [17,27,28]. Most of the mutations were found in the ACVRL1 gene with a total of 10 different mutations in 12 HHT patients whereas a total of 5 mutations were identified in the coding sequence of the ENG gene in 6 individuals. Most of the ENG mutations were widely distributed throughout the gene, not indicating any special hot spot areas within the gene. Out of five detected DNA changes, four mutations had been described previously as mutations in the HHT mutation database or recently been published [2,5,20,27,31-35]. The alterations included mainly nucleotide substitutions such as missense, splice site and frameshift mutations. In spite of the fact that only a small series of HHT patients was investigated in this study, one novel ENG mutation c.1384insT (p.Gln462fs) could be detected which had not been described before in the HHT mutation database. Unlike ENG, 50% of the mutations in the ACVRL1 gene were missense substitutions. Similar results had been published by Abdalla et al. who analysed all 123 till then known mutations of ACVRL1. In their study overview they could also register more than half (53%) of the mutations in the ACVRL1 gene as missense mutations [36,37]. As previously described in other HHT families, the frequency of ACVRL1 mutations was highest in exon 8 which might be an indicator for mutation hotspots. The other mutations c.144_145insG (p.Ala49fs), c.200G>A (p.Arg67Gln), c.540_541insA (p.Asp181fs), c.673_674delAG (p.Ser225fs), c.696_698delCTC (p.Ser233del), c.1346C>T (p.Pro449Leu) and c.1377+1G>A in ACVRL1 were already described by other workgroups, mostly located in the intracellular kinase domain of the gene [28,35,37-40]. Only two already known mutations could be detected in the extracellular domain of the gene. However, large deletions and insertions in ENG and ACVRL1 were not analyzed which could explain the low detection rate of mutations in our study compared to other studies described in literature [28,32,36].
As far as the genotype-phenotype correlation is concerned, most of the pulmonary arteriovenous malformations (PAVMs) in our study were detected in HHT patients with mutations for ENG. This phenomenon has already been described by previous studies given in literature, stating that mutations of ENG are mainly observed in HHT type 1 with an incidence of up to 40% for PAVMs, whereas mutations of ALK1 are primarily observed in HHT type 2 with an incidence of only 14% for PAVMs, which clinically distinguishes these two types of mutation [3,7,28,36]. Liver manifestations were more frequent in patients with ACVRL1 mutations, whereas manifestations of the skin and the GI-tract were seen equally distributed amongst HHT patients with ENG and ACVRL1 mutations.
Many studies report a mutation detection rate of up to 92% [27,28,33]. However, in our study we were able to identify only 43.9% of disease-causing mutations in a series of 41 HHT patients. The difference in sensitivity may result from several factors, including a very heterogeneous cohort of patients in terms of locations and initial physician involved, and because patients with only 2 Curaçao criteria were included in the study. In addition, only ENG and ACVRL1 genes were screened for mutations, but in some cases, mutations in SMAD4, HHT3 and HHT4 loci may be responsible for the clinical symptoms [24-26].
In daily clinical routine, the diagnosis of HHT is primarily based on the clinical findings according to the Curaçao criteria. Nevertheless, in many cases clinical diagnostic is at times very difficult due to the heterogeneous outcome and manifestation of the disease. This fact asks for additional diagnostic tools which facilitate the daily clinical work. Already in the past, many studies could show that the molecular characterization of HHT enables to predict the diagnosis before the manifestation of clinical symptoms. The results of the study reflect the very high and heterogenic distribution and allocation of mutations in HHT. With an allele-specific PCR technique (PCR-SSP) we could establish a diagnostic testing device for rapid genotyping of mutations. The exon sequencing of ENG and ACVRL1 confirmed the previously described polymorphisms and outlined one novel mutation Gln462fs in exon 10 of ENG gene. The PCR-SSP technique can easily be performed with standard laboratory equipment and may help to minimize costs for genetic testing in the routine diagnosis of family members of HHT patients with known mutations.
Conclusion
In summary, this study is a further contribution to the determination of already known ENG and ACVRL1 mutations as well as the detection of one novel ENG mutation in HHT. Next to the Curaçao criteria, genetic analysis certainly contributes as an essential tool to a reliable diagnosis of clinically affected HHT patients and clinically un-symptomatic HHT patients, thus helping to take early preventive measures even before the occurrence of first clinical symptoms. The PCR-SSP technique could facilitate this high task of genetic analysis in routine HHT diagnostics and underlines the importance of using molecular diagnosis for early identification of individuals carrying mutations and being at risk of vascular complications.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
All the authors have read and approved the final manuscript. A large part of the mutational analysis was done by HS, JH under the supervision of PB. The rest of the mutational analysis was done by UG, JS, KH completing the analysis of mutations. Bank data collection, writing of the manuscript and the direction of all the work was done under HS, the main author.
Pre-publication history
The pre-publication history for this paper can be accessed here:
Supplementary Material
Table 1. Overview on the detailed clinical data of all HHT patients.
Table 2. Diagnostic criteria according to the Curaçao criteria for the diagnosis of HHT.
Table 3. Primers used in PCR-SSP genotyping for mutations in the ENG and ACVRL1 genes.
Table 4. Summary of mutations identified in the ENG gene.
Table 5. Polymorphisms in ENG.
Table 6. Summary of mutations identified in the ACVRL1 gene.
Table 7. Genotype-phenotype correlation in HHT patients with confirmed mutation.
Contributor Information
Haneen Sadick, Email: haneen.sadick@web.de.
Johanna Hage, Email: haneen.sadick@umm.de.
Ulrich Goessler, Email: ulrich.goessler@hno.ma.uni-heidelberg.de.
Jens Stern-Straeter, Email: jens.stern-straeter@hno.ma.uni-heidelberg.de.
Frank Riedel, Email: f_riedel@gmx.de.
Karl Hoermann, Email: karl.hoermann@umm.de.
Peter Bugert, Email: peter.bugert@umm.de.
Acknowledgements
We are indebted to the patients and their families and relatives for their participation. We would like to thank Petra Prohaska and Gabi Rink for excellent technical assistance.
References
- Guttmacher AE, Marchuk DA, White RI Jr. Hereditary hemorrhagic telangiectasia. N Engl J Med. 1995;333(14):918–24. doi: 10.1056/NEJM199510053331407. [DOI] [PubMed] [Google Scholar]
- Abdalla SA, Cymerman U, Rushlow D, Chen N, Stoeber GP, Lemire EG, Letarte M. Novel mutations and polymorphisms in genes causing hereditary hemorrhagic telangiectasia. Hum Mutat. 2005;25(3):320–1. doi: 10.1002/humu.9312. [DOI] [PubMed] [Google Scholar]
- Kjeldsen A, Kjeldsen J. Gastrointestinal bleeding in patients with hereditary hemorrhagic telangiectasia. Am J Gastroenterol. 2000;95:415–418. doi: 10.1111/j.1572-0241.2000.01792.x. [DOI] [PubMed] [Google Scholar]
- Reilly PJ, Nostrant TT. Clinical manifestations of hereditary hemorrhagic telangiectasia. Am J Gastroenterol. 1984;79:363–367. [PubMed] [Google Scholar]
- Kjeldsen AD, Oxhøj H, Andersen PE, Green A, Vase P. Prevalence of pulmonary arteriovenous malformations (PAVMs) and occurrence of neurologic symptoms in patients with hereditary haemorrhagic telangiectasia (HHT) J Int Med. 2000;248:255–262. doi: 10.1046/j.1365-2796.2000.00725.x. [DOI] [PubMed] [Google Scholar]
- Shovlin CL, Letarte M. Hereditary haemorrhagic telangiectasia and pulmonary arteriovenous malformations: issues in clinical management and review of pathogenic mechanisms. Thorax. 1999;54:714–729. doi: 10.1136/thx.54.8.714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernard G, Mion F, Henry L, Plauchu H, Paliard P. Hepatic involvement in hereditary haemorrhagic telangiectasia: clinical, radiological, and hemodynamic studies of 11 cases. Gastroenterology. 1993;105:482–487. doi: 10.1016/0016-5085(93)90723-p. [DOI] [PubMed] [Google Scholar]
- Garcia-Tsao G, Korzenik JR, Young L, Henderson KJ, Jain D, Byrd B, Pollak JS, White RI Jr. Liver disease in patients with hereditary hemorrhagic telangiectasia. N Engl J Med. 2000;343(13):931–936. doi: 10.1056/NEJM200009283431305. [DOI] [PubMed] [Google Scholar]
- Sadick H, Sadick M, Götte K, Naim R, Riedel F, Bran G, Hörmann K. Hereditary hemorrhagic telangiectasia: an update on clinical manifestations and diagnostic measures. Wien Klin Wochenschr. 2006;118(3–4):72–80. doi: 10.1007/s00508-006-0561-x. [DOI] [PubMed] [Google Scholar]
- Braverman IM, Keh A, Jacobson BS. Ultrastructure and three-dimensional organization of the teleangiectases of hereditary hemorrhagic teleangiectasia. J Invest Dermatol. 1990;95:422–427. doi: 10.1111/1523-1747.ep12555569. [DOI] [PubMed] [Google Scholar]
- Jahnke V. Ultrastructure of hereditary hemorrhagic telangiectasia. Arch Otolaryngol Head and Neck Surg. 1970;91:262–265. [PubMed] [Google Scholar]
- White RI Jr. Pulmonary arteriovenous malformations: how do we diagnose them and why is it important to do so? Radiology. 1992;182(3):633–5. doi: 10.1148/radiology.182.3.1535872. [DOI] [PubMed] [Google Scholar]
- Shovlin CL, Guttmacher AE, Buscarini E, Faughnan ME, Hyland RH, Westermann CJ, Kjeldsen AD, Plauchu H. Diagnostic criteria for hereditary hemorrhagic telangiectasia (Rendu-Osler-Weber syndrome) Am J Med Genet. 2000;91(1):66–7. doi: 10.1002/(SICI)1096-8628(20000306)91:1<66::AID-AJMG12>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- McDonald MT, Papenberg KA, Ghosh S, Glatfelter AA, Biesecker BB, Helmbold EA, Markel DS, Zolotor A, McKinnon WC, Vanderstoep JL, Jackson CE, Iannuzzi M, Collins FS, Boehnke M, Porteous ME, Guttmacher AE, Marchuk DA. A disease locus for hereditary haemorrhagic telangiectasia maps to chromosome 9q33-34. Nat Genet. 1994;6(2):197–204. doi: 10.1038/ng0294-197. [DOI] [PubMed] [Google Scholar]
- Shovlin CL, Hughes JM, Tuddenham EG, Temperley I, Perembelon YF, Scott J, Seidman CE, Seidman JG. A gene for hereditary haemorrhagic telangiectasia maps to chromosome 9q3. Nat Genet. 1994;6(2):205–9. doi: 10.1038/ng0294-205. [DOI] [PubMed] [Google Scholar]
- McAllister KA, Grogg KM, Johnson DW, Gallione CJ, Baldwin MA, Jackson CE, Helmbold EA, Markel DS, McKinnon WC, Murrell J. Endoglin, a TGF-beta binding protein of endothelial cells, is the gene for hereditary haemorrhagic telangiectasia type 1. Nat Genet. 1994;8(4):345–51. doi: 10.1038/ng1294-345. [DOI] [PubMed] [Google Scholar]
- Letteboer TG, Mager JJ, Snijder RJ, Koeleman BP, Lindhout D, Ploos van Amstel JK, Westermann CJ. Genotype-phenotype relationship in hereditary haemorrhagic telangiectasia. J Med Genet. 2006;43(4):371–7. doi: 10.1136/jmg.2005.035451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg JN, Gallione CJ, Stenzel TT, Johnson DW, Allen WP, Schwartz CE, Jackson CE, Porteous ME, Marchuk DA. The activin receptor-like kinase 1 gene. genomic structure and mutations in hereditary hemorrhagic telangiectasia type 2. Am J Hum Genet. 1997;61(1):60–7. doi: 10.1086/513903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson DW, Berg JN, Baldwin MA, Gallione CJ, Marondel I, Yoon SJ, Stenzel TT, Speer M, Pericak-Vance MA, Diamond A, Guttmacher AE, Jackson CE, Attisano L, Kucherlapati R, Porteous ME, Marchuk DA. Mutations in the activin receptor-like kinase 1 gene in hereditary haemorrhagic telangiectasia type 2. Nat Genet. 1996;13:189–95. doi: 10.1038/ng0696-189. [DOI] [PubMed] [Google Scholar]
- Kuehl HK, Caselitz M, Hasenkamp S, Wagner S, El-Harith el-HA, Manns MP, Stuhrmann M. Hepatic manifestation is associated with ALK1 in hereditary hemorrhagic telangiectasia: identification of five novel ALK1 and one novel ENG mutations. Hum Mutat. 2005;25(3):320. doi: 10.1002/humu.9311. [DOI] [PubMed] [Google Scholar]
- Arthur HM, Ure J, Smith AJ, Renforth G, Wilson DI, Torsney E, Charlton R, Parums DV, Jowett T, Marchuk DA, Burn J, Diamond AG. Endoglin, an ancillary TGFbeta receptor, is required for extraembryonic angiogenesis and plays a key role in heart development. Dev Biol. 2000;217:42–53. doi: 10.1006/dbio.1999.9534. [DOI] [PubMed] [Google Scholar]
- Oh SP, Seki T, Goss KA, Imamura T, Yi Y, Donahoe PK, Li L, Miyazono K, ten Dijke P, Kim S, Li E. Activin receptor-like kinase 1 modulates transforming growth factor-beta 1 signaling in the regulation of angiogenesis. Proc Natl Acad Sci USA. 2000;97:2626–31. doi: 10.1073/pnas.97.6.2626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pece-Barbara N, Cymerman U, Vera S, Marchuk DA, Letarte M. Expression analysis of four endoglin missense mutations suggests that haploinsufficiency is the predominant mechanism for hereditary hemorrhagic telangiectasia type 1. Hum Mol Genet. 1999;8(12):2171–81. doi: 10.1093/hmg/8.12.2171. [DOI] [PubMed] [Google Scholar]
- Cole SG, Begbie ME, Wallace GM, Shovlin CL. A new locus for hereditary haemorrhagic telangiectasia (HHT3) maps to chromosome 5. J Med Genet. 2005;42(7):577–82. doi: 10.1136/jmg.2004.028712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bayrak-Toydemir P, McDonald J, Akarsu N, Toydemir RM, Calderon F, Tuncali T, Tang W, Miller F, Mao R. A fourth locus for hereditary hemorrhagic telangiectasia maps to chromosome 7. Am J Med Genet A. 2006;140:2155–62. doi: 10.1002/ajmg.a.31450. [DOI] [PubMed] [Google Scholar]
- Gallione CJ, Repetto GM, Legius E, Rustgi AK, Schelley SL, Tejpar S, Mitchell G, Drouin E, Westermann CJ, Marchuk DA. A combined syndrome of juvenile polyposis and hereditary haemorrhagic telangiectasia associated with mutations in MADH4 (SMAD4) Lancet. 2004;363:852–9. doi: 10.1016/S0140-6736(04)15732-2. [DOI] [PubMed] [Google Scholar]
- Abdalla SA, Letarte M. Hereditary haemorrhagic telangiectasia: current views on genetics and mechanisms of disease. J Med Genet. 2006;43(2):97–110. doi: 10.1136/jmg.2005.030833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulte C, Geisthoff U, Lux A, Kupka S, Zenner HP, Blin N, Pfister M. High frequency of ENG and ALK1/ACVRL1 mutations in German HHT patients. Hum Mutat. 2005;25(6):595. doi: 10.1002/humu.9345. [DOI] [PubMed] [Google Scholar]
- Bugert P, Hoffmann MM, Winkelmann BR, Vosberg M, Jahn J, Entelmann M, Katus HA, März W, Mansmann U, Boehm BO, Goerg S, Klüter H. The variable number of tandem repeat polymorphism in the P-selectin glycoprotein ligand-1 gene is not associated with coronary heart disease. J Mol Med. 2003;81(8):495–501. doi: 10.1007/s00109-003-0459-2. [DOI] [PubMed] [Google Scholar]
- Bugert P, Lese A, Meckies J, Zieger W, Eichler H, Klüter H. Optimized sensitivity of allele-specific PCR for prenatal typing of human platelet alloantigen single nucleotide polymorphisms. Biotechniques. 2003;35(1):170–4. doi: 10.2144/03351md05. [DOI] [PubMed] [Google Scholar]
- Bossler AD, Richards J, George C, Godmilow L, Ganguly A. Novel mutations in ENG and ACVRL1 identified in a series of 200 individuals undergoing clinical genetic testing for hereditary hemorrhagic telangiectasia HHT): correlation of genotype with phenotype. Hum Mutat. 2006;27(7):667–75. doi: 10.1002/humu.20342. [DOI] [PubMed] [Google Scholar]
- Lesca G, Burnichon N, Raux G, Tosi M, Pinson S, Marion MJ, Babin E, Gilbert-Dussardier B, Rivière S, Goizet C, Faivre L, Plauchu H, Frébourg T, Calender A, Giraud S. French Rendu-Osler Network. Distribution of ENG and ACVRL1 (ALK1) mutations in French HHT patients. Hum Mutat. 2006;27(6):598. doi: 10.1002/humu.9421. [DOI] [PubMed] [Google Scholar]
- Lesca G, Olivieri C, Burnichon N, Pagella F, Carette MF, Gilbert-Dussardier B, Goizet C, Roume J, Rabilloud M, Saurin JC, Cottin V, Honnorat J, Coulet F, Giraud S, Calender A, Danesino C, Buscarini E, Plauchu H, French-Italian-Rendu-Osler Network. Genotype-phenotype correlations in hereditary hemorrhagic telangiectasia: data from the French-Italian HHT network. Genet Med. 2007;9(1):14–22. doi: 10.1097/GIM.0b013e31802d8373. [DOI] [PubMed] [Google Scholar]
- Fernandez-L A, Sanz-Rodriguez F, Zarrabeitia R, Perez-Molino A, Morales C, Restrepo CM, Ramirez JR, Coto E, Lenato GM, Bernabeu C, Botella LM. Mutation study of Spanish patients with hereditary hemorrhagic telangiectasia and expression analysis of Endoglin and ALK1. Hum Mutat. 2006;27(3):295. doi: 10.1002/humu.9413. [DOI] [PubMed] [Google Scholar]
- Fontalba A, Fernandez-L A, García-Alegria E, Albiñana V, Garrido-Martin EM, Blanco FJ, Zarrabeitia R, Perez-Molino A, Bernabeu-Herrero ME, Ojeda ML, Fernandez-Luna JL, Bernabeu C, Botella LM. Mutation study of Spanish patients with hereditary hemorrhagic telangiectasia. BMC Med Genet. 2008;9:75. doi: 10.1186/1471-2350-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wehner LE, Folz BJ, Argyriou L, Twelkemeyer S, Teske U, Geisthoff UW, Werner JA, Engel W, Nayernia K. Mutation analysis in hereditary haemorrhagic telangiectasia in Germany reveals 11 novel ENG and 12 novel ACVRL1/ALK1 mutations. Clin Genet. 2006;69(3):239–45. doi: 10.1111/j.1399-0004.2006.00574.x. [DOI] [PubMed] [Google Scholar]
- Abdalla SA, Letarte M. Hereditary haemorrhagic telangiectasia: current views on genetics and mechanisms of disease. J Med Genet. 2006;43(2):97–110. doi: 10.1136/jmg.2005.030833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klaus DJ, Gallione CJ, Anthony K, Yeh EY, Yu J, Lux A, Johnson DW, Marchuk DA. Novel missense and frameshift mutations in the activin receptor-like kinase-1 gene in hereditary hemorrhagic telangiectasia. Mutations in brief no. 164. Hum Mutat. 1998;12(2):137. doi: 10.1002/(SICI)1098-1004(1998)12:2<137::AID-HUMU15>3.0.CO;2-M. [DOI] [PubMed] [Google Scholar]
- Abdalla SA, Cymerman U, Johnson RM, Deber CM, Letarte M. Disease-associated mutations in conserved residues of ALK-1 kinase domain. Eur J Hum Genet. 2003;11(4):279–87. doi: 10.1038/sj.ejhg.5200919. [DOI] [PubMed] [Google Scholar]
- Lesca G, Plauchu H, Coulet F, Lefebvre S, Plessis G, Odent S, Rivière S, Leheup B, Goizet C, Carette MF, Cordier JF, Pinson S, Soubrier F, Calender A, Giraud S. French Rendu-Osler Network. Molecular screening of ALK1/ACVRL1 and ENG genes in hereditary hemorrhagic telangiectasia in France. Hum Mutat. 2004;23(4):289–99. doi: 10.1002/humu.20017. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table 1. Overview on the detailed clinical data of all HHT patients.
Table 2. Diagnostic criteria according to the Curaçao criteria for the diagnosis of HHT.
Table 3. Primers used in PCR-SSP genotyping for mutations in the ENG and ACVRL1 genes.
Table 4. Summary of mutations identified in the ENG gene.
Table 5. Polymorphisms in ENG.
Table 6. Summary of mutations identified in the ACVRL1 gene.
Table 7. Genotype-phenotype correlation in HHT patients with confirmed mutation.