Abstract
We previously reported the isolation of novel methanogens by using a new cultivation method, referred to as the coculture method. Here, we extended our coculture method to various anaerobic environmental samples. As a result, we successfully cultivated some uncharacterized methanogens in coculture enrichments and eventually isolated a new methanogen, within the order Methanomicrobiales.
So far, almost all cases of the cultivation and isolation of H2-utilizing methanogenic Archaea (methanogens) have been performed under high-H2 concentrations (e.g., around 100 kPa), even though the concentrations in their natural habitat are far lower (10 to 100 Pa) than in laboratory cultures. This difference between in vitro and in situ physicochemical conditions very likely means that fast-growing methanogens that may prefer high concentrations of H2 will have to be specifically selected; thus, laboratory cultures under such high-H2 conditions result in the growth of a very limited range of species. To avoid this situation, we have proposed a new cultivation method, which we named the coculture method, for cultivating H2-utilizing methanogens (14).
Under anaerobic conditions, methanogens often partner with heterotrophic H2-producing bacteria, which catalyze the oxidation of a variety of substrates (fatty acids, alcohols, and aromatic compounds). The methanogens use the H2 produced by these heterotrophic bacteria, and in return, the bacteria benefit from the removal of the H2 that would otherwise inhibit their growth. This lifestyle is commonly referred to as interspecies H2 transfer, and the heterotrophic H2-producing bacteria relying on H2-utilizing methanogens are called syntrophs (11). In our previous studies, cultivation was performed with propionate as an indirect precursor substrate that is converted to H2 by syntrophs, with the expectation that methanogens would grow as a result of interspecies H2 transfer. Based on this strategy, two novel methanogens representing new genera, Methanocella paludicola strain SANAE and Methanolinea tarda strain NOBI-1, were successfully isolated (7, 13, 14).
In this study, we extended the method to various types of environmental samples to cultivate and isolate uncharacterized methanogens. Moreover, we also extended the H2-supplying substrates to include ethanol and butyrate in addition to propionate, because these substances are also known to be decomposed by a syntrophic association of substrate-oxidizing H2-producing bacteria and H2-utilizing methanogens (15).
Nine anaerobic environmental samples (marine coastal sediment from Kashiwazaki, Niigata, Japan [KO]; freshwater lake sediment from Lake Suwa, Nagano, Japan [SL]; freshwater pond sediment from Shouzuma Pond, Nagano, Japan [SP]; river sediment of the Azusa River, Nagano, Japan [AR]; sediment from a lotus field located in Nagaoka, Niigata, Japan [LF]; rice field soil from Matsumoto, Nagano, Japan [NR]; rice field soil from Nagaoka, Niigata, Japan [SRP]; rice field soil from Tainan, Taiwan [TNR]; and methanogenic granular sludge obtained from a lab-scale upflow anaerobic sludge blanket reactor treating wastewater from the manufacture of palm oil in our laboratory [MP]) were anaerobically incubated with ethanol (10 mM), butyrate (20 mM), or propionate (20 mM) as the sole carbon and energy source. Additionally, we prepared propionate enrichment cultures with the addition of a pure culture of anaerobic syntrophic propionate-oxidizing bacterium Syntrophobacter fumaroxidans strain MPOB (DSMZ 10017) cells (inoculum size, 5% [vol/vol]) to obtain stable cultures (8, 18), except for enrichments from the marine sediment and granular sludge samples because the NaCl resistance of S. fumaroxidans was unknown (6) and the granular sludge was expected to contain a large amount of indigenous syntrophic bacteria (5, 16). Moreover, as control experiments, the same environmental samples were used in enrichments by the canonical cultivation method in the presence of high concentrations of H2 (ca. 150 kPa in headspace) or formate (40 mM). All cultivations were performed anaerobically at 37°C without shaking. In total, 52 primary enrichment cultures were prepared for this study (see Table S1 in the supplemental material).
When primary enrichment was made using high concentrations of H2 and formate, the growth of methanogen-like microbes was confirmed within 3 to 5 days of incubation (as examples, photomicrographs of the enrichment cultures from TNR are shown in Fig. S1A to D in the supplemental material). After three consecutive transfers, 16S rRNA gene-based clone analysis was performed using an archaeal universal primer pair, Ar109f/1490R (14). Twenty-nine phylotypes were detected and were closely related to previously isolated methanogens (Fig. 1 and 2A and B; see Table S1 in the supplemental material). Among them, 26 phylotypes were classified into the genus Methanobacterium, two phylotypes in the genus Methanospirillum, and one in the genus Methanogenium. Moreover, 20 phylotypes showed high similarities (>97%) with the 16S rRNA genes of previously isolated methanogens, whereas the remaining nine phylotypes showed 95 to 96% similarities with the 16S rRNA genes of previously characterized methanogens (see Table S1 in the supplemental material). The methanogens possessing these sequences may be taxonomically novel at least at the species level (17), but they were all affiliated with the well-studied genera Methanobacterium and Methanospirillum.
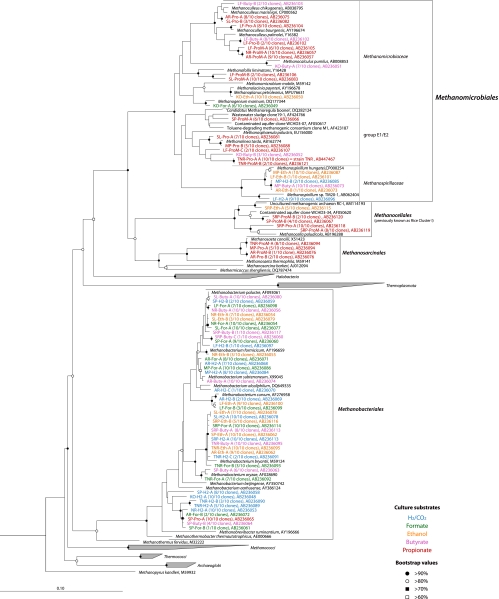
FIG. 1.
Phylogenetic tree showing the placement of 16S rRNA gene sequences/clones obtained in this study. The colored phylotypes were obtained in this study. The difference in color among phylotypes indicates the different substrates used for the enrichment cultures (blue, hydrogen; green, formate; orange, ethanol; pink, butyrate; red, propionate and propionate plus S. fumaroxidans strain MPOB). The name of each phylotype is composed of the sample name, an abbreviation of the substrate for cultivation (H2, hydrogen; For, formate; Eth, ethanol; Buty, butyrate; Pro, propionate; ProM, propionate plus S. fumaroxidans), and the phylotype (for example, SRP-Pro-A is phylotype A recovered from the propionate enrichment culture cultivated from the environmental sample SRP). The number in parentheses indicates the number of identical clones obtained per number of clones analyzed for each phylotype. The accession numbers are also shown after each phylotype name. The phylotypes indicated by the same accession numbers have the same sequences (e.g., SP-For-A and SRP-Buty-C, AR-ProM-A and NR-ProM-A). All of the clonal sequences were greater than 1,000 nucleotides in length, with the exception of Methanospirillum sp. TM20-1 (GenBank acc. no. AB062404; 789 bp). Therefore, the initial tree was constructed with sequences greater than 1,000 nucleotides using the neighbor-joining method. Subsequently, the Methanospirillum sp. TM20-1 sequence was inserted into the tree by using the parsimony insertion tool of the ARB program. The scale bar indicates the estimated number of base changes per nucleotide sequence position. The symbols at the branch nodes indicate bootstrap values.
FIG. 2.
Phylogenetic affiliation of the identified phylotypes based on their cultivation substrates. The panels indicate the results of enrichment cultures with the following substrates: H2 (A), formate (B), ethanol (C), butyrate (D), propionate (E), and propionate with the addition of the pure culture of S. fumaroxidans (F). The identified phylotypes were classified into their respective genera according to their 16S rRNA gene similarity with previously characterized methanogens. Phylotypes possessing sequence similarity greater than 92% were treated as the same genus. The number of phylotypes for each group is indicated in parentheses.
In the coculture enrichments, substrate degradation concomitant with methane formation was confirmed after 1 week and more than 1 to 3 months of incubation in ethanol enrichment cultures and butyrate and propionate enrichment cultures, respectively. In particular, the growth of microbes in the propionate enrichments without the addition of S. fumaroxidans cells, except for the enrichments constructed from the RF and SRP samples, was very slow and unstable; the growth and methane production stopped unexpectedly and often made successive passages to fresh medium difficult. Additionally, two propionate enrichments in the absence of S. fumaroxidans inoculated from the KO and NR samples did not show methane production after a year of incubation. On the other hand, all of the propionate enrichments in the presence of S. fumaroxidans cells showed stable growth. During the incubation of the coculture enrichments, the H2 partial pressures in the cultures were kept at <100 Pa in the ethanol enrichments and at <30 Pa in the butyrate and propionate cultures. Methane, H2, short-chain fatty acids, and ethanol were measured as described previously (14). Microscopic observation after three to four transfers showed that those enrichments were comprised mainly of F420-autofluorescent methanogen-like cells and oval- or rod-shaped bacterial cells, possibly syntrophs (see Fig. S1E to L in the supplemental material). These observations suggested that ethanol, butyrate, and propionate degradation were carried out by syntrophic association between syntrophic substrate-oxidizing H2-producing bacteria and H2-utilizing methanogens. To identify the methanogens present in those enrichments, archaeal 16S rRNA gene-based clone analyses were performed. A total of 52 phylotypes were obtained (Fig. 1 and 2C to F; see Table S1 in the supplemental material). Of these, 23 phylotypes were classified into the genera Methanobacterium (19 phylotypes) and Methanospirillum (4 phylotypes), which were very similar to those obtained from the H2 and formate enrichments. On the other hand, the remaining 29 phylotypes were comprised of the orders Methanomicrobiales (20 phylotypes), Methanocellales (5 phylotypes), and Methanosarcinales (4 phylotypes, all belonging to the genus Methanosaeta). Within the order Methanomicrobiales, some phylotypes were affiliated with the genera Methanoculleus (nine phylotypes), Methanofollis (two phylotypes), Methanocalculus (one phylotype), and Methanoplanus (one phylotype). Additionally, sequences very closely related to the recently isolated methanogens Methanolinea tarda (six phylotypes) (7, 14) and “Candidatus Methanoregula boonei” (one phylotype) (1) were also obtained. Both M. tarda and “Ca. Methanoregula boonei” represent a family-level clade, which had long been recognized as an uncultured archaeal lineage called the group E1/E2 (1) (Fig. 1). Regarding the five phylotypes within the order Methanocellales, all were obtained from SRP and SP enrichments. Though the order Methanocellales had been recognized as the clone cluster rice cluster I, one strain has been isolated very recently (13, 14) and the rice cluster I methanogens are now being unveiled.
Of the 52 phylotypes obtained from the coculture enrichments, 38 phylotypes (73% of the total phylotypes) were >97% similar to the 16S rRNA genes of the previously characterized (cultivated) methanogens. In contrast, 14 phylotypes (27%) had <96% sequence similarity with those of known methanogens. The organisms represented by these phylotypes were considered to be taxonomically novel at the species or even the genus level. Most of these phylotypes were affiliated with the orders Methanomicrobiales and Methanocellales with 92 to 96% sequence similarity (see Table S1 in the supplemental material). According to the 16S rRNA gene-based clone analysis, taxonomically novel methanogens were found in abundance in one ethanol, two butyrate, and eight propionate enrichments (from the KO, SP, SL, TNR, LF, and SRP samples). Especially, the ethanol and six propionate enrichments (from the SP, TNR, LF, and SRP samples) contained novel methanogens belonging to the group E1/E2 and/or the order Methanocellales (formerly known as rice cluster I), both of which contain only a few cultivated representatives so far. Therefore, we attempted to isolate these methanogens from the enrichments. After several attempts were performed over a year, a novel methanogen, designated strain TNR, was successfully isolated from the propionate enrichment culture (TNR) by serial dilution in liquid medium with H2 (ca. 150 kPa) as the substrate.
Strain TNR was a nonmotile, rod-shaped methanogen, which utilized H2/CO2 and formate for growth and methane production (see Fig. S2 in the supplemental material). The doubling time was 1.2 days at 37°C and pH 7. The most closely related methanogen cultivated so far was Methanolinea tarda that we have recently isolated (7), but the similarity of the 16S rRNA genes between the two was only 95% (Fig. 1). On the other hand, the isolation of methanogens from the other enrichments was not successful, i.e., when the coculture enrichments were inoculated into the serial dilution cultures with high concentrations of H2 or formate, nontargeted methanogens, almost all of which had >97% sequence similarities to the 16S rRNA genes of known Methanobacterium and Methanoculleus species, outgrew in the cultures. The conventional method for final purification (i.e., using high concentrations of H2 or formate as a direct substrate) has, therefore, a clear limitation, and new methods to overcome this will be needed.
By using the coculture method, we successfully enriched methanogens that were absent in previous cultivation attempts and were only detected as environmental clones. In addition, we were able to isolate a methanogen belonging to the group E1/E2 of the order Methanomicrobiales. Our study clearly demonstrated that the coculture method is an effective way to cultivate hitherto uncharacterized methanogens. Interestingly, the taxonomic compositions of the phylotypes were clearly different depending on the substrates used in the coculture method (Fig. 1 and 2). When conventional cultivation was employed using high concentrations of H2 and formate, only very limited phylotypes were obtained, namely, Methanobacterium- and Methanospirillum-related phylotypes in the H2 cultures and Methanobacterium- and Methanogenium-related phylotypes in the formate cultures. When using the coculture method with ethanol or butyrate, Methanobacterium-related phylotypes were also dominant, accounting for 64.3% of the total phylotypes, whereas more diverse methanogen phylotypes than those in the H2 and formate cultures were retrieved. Contrary to these results, propionate (with and without S. fumaroxidans) enrichments allowed quite a different pattern of methanogen phylotypes to become established. The most abundant phylotypes obtained from the propionate enrichments belonged to the orders Methanocellales and Methanomicrobiales, accounting for 72.8 and 84.7% of the clones examined. The addition of S. fumaroxidans cells into the propionate enrichments seemed to have no significant effect on the methanogenic community compositions that emerged, but it helped the stability of the whole community and the capability of the propionate degradation. The theoretical ranges of H2 partial pressure that allow the anaerobic oxidation of ethanol, butyrate, and propionate to occur are 0.5 to 27,000 Pa; 0.5 to 60 Pa; and 0.5 to 28 Pa, respectively. These values were calculated based on the review on energy conservation by Thauer et al. (19), in which the concentrations of products and reactants were 0.35 atm  , 0.65 atm
, 0.65 atm  , and 20 mM substances at 37°C and pH 7. For the calculation, a temperature correction was made using the van't Hoff equation. Theoretically, the H2 partial pressures in the various cultures differ depending on the substrates used, becoming lower in the order of substrates: ethanol > butyrate > propionate. Actually, the H2 partial pressures measured during substrate degradation in the coculture enrichments remained within these theoretical ranges (data not shown). Given the above theoretical values, the apparent H2 partial pressure that could be generated from a particular substrate would be the crucial factor affecting the change in the compositions of H2-utilizing methanogens in the community. In fact, the relative abundance of members of the genera Methanobacterium and Methanospirillum increased as the given H2 partial pressure became higher (propionate → butyrate → ethanol → H2), and conversely, the relative abundance of members of the orders Methanomicrobiales (except for the genus Methanospirillum) and Methanocellales increased as the H2 partial pressure became lower (Fig. 1 and 2). We assume that Methanocellales spp. and Methanomicrobiales spp. (except for Methanospirillum spp.) have higher affinities for H2 than Methanobacterium spp. and Methanospirillum spp. Several previous studies also support this prediction. Lu et al. reported that Methanocellales methanogens incorporated 13C when rice roots were incubated in a low-H2 atmosphere in the presence of 13CO2, while Methanobacteriales and Methanosarcinales methanogens preferentially incorporated 13C in a high-H2 atmosphere (10). Also, Methanocellales phylotypes were detected from methanogenic environments, usually with a low concentration of H2, such as rice fields, fens, and peat bogs (e.g., see references 3, 4, and 9). In addition to the Methanocellales methanogens, members of the order Methanomicrobiales were frequently found in abundance in low-H2-concentration methanogenic environments, such as peat bogs, fens, lake sediments, and rice fields (e.g., see references 2, 12, and 20). Detailed substrate affinity information will provide insight into the relevance between the population structures of methanogens and the H2 concentrations of their habitats.
, and 20 mM substances at 37°C and pH 7. For the calculation, a temperature correction was made using the van't Hoff equation. Theoretically, the H2 partial pressures in the various cultures differ depending on the substrates used, becoming lower in the order of substrates: ethanol > butyrate > propionate. Actually, the H2 partial pressures measured during substrate degradation in the coculture enrichments remained within these theoretical ranges (data not shown). Given the above theoretical values, the apparent H2 partial pressure that could be generated from a particular substrate would be the crucial factor affecting the change in the compositions of H2-utilizing methanogens in the community. In fact, the relative abundance of members of the genera Methanobacterium and Methanospirillum increased as the given H2 partial pressure became higher (propionate → butyrate → ethanol → H2), and conversely, the relative abundance of members of the orders Methanomicrobiales (except for the genus Methanospirillum) and Methanocellales increased as the H2 partial pressure became lower (Fig. 1 and 2). We assume that Methanocellales spp. and Methanomicrobiales spp. (except for Methanospirillum spp.) have higher affinities for H2 than Methanobacterium spp. and Methanospirillum spp. Several previous studies also support this prediction. Lu et al. reported that Methanocellales methanogens incorporated 13C when rice roots were incubated in a low-H2 atmosphere in the presence of 13CO2, while Methanobacteriales and Methanosarcinales methanogens preferentially incorporated 13C in a high-H2 atmosphere (10). Also, Methanocellales phylotypes were detected from methanogenic environments, usually with a low concentration of H2, such as rice fields, fens, and peat bogs (e.g., see references 3, 4, and 9). In addition to the Methanocellales methanogens, members of the order Methanomicrobiales were frequently found in abundance in low-H2-concentration methanogenic environments, such as peat bogs, fens, lake sediments, and rice fields (e.g., see references 2, 12, and 20). Detailed substrate affinity information will provide insight into the relevance between the population structures of methanogens and the H2 concentrations of their habitats.
Nucleotide sequence accession numbers.
Sequence data obtained in this study were deposited in the DDBJ/EMBL/GenBank databases under the accession numbers AB236048 to AB236107, AB236113 to AB236121, and AB447467.
Supplementary Material
Acknowledgments
We thank Hironori Okano, Kotaro Horisawa, and Eiji Tasumi for their technical assistance.
This study was financially supported by grants from the Japan Society for the Promotion of Science (Grant-in-Aid for Young Scientists [Start-Up]); the Ministry of Education, Culture, Sports, Science and Technology, Japan; and the Institute for Fermentation, Osaka, Japan.
Footnotes
Published ahead of print on 22 May 2009.
Supplemental material for this article may be found at http://aem.asm.org/.
REFERENCES
- 1.Bräuer, S. L., H. Cadillo-Quiroz, E. Yashiro, J. B. Yavitt, and S. H. Zinder. 2006. Isolation of a novel acidiphilic methanogen from an acidic peat bog. Nature 442:192-194. [DOI] [PubMed] [Google Scholar]
- 2.Cadillo-Quiroz, H., E. Yashiro, J. B. Yavitt, and S. H. Zinder. 2008. Characterization of the archaeal community in a minerotrophic fen and terminal restriction fragment length polymorphism-directed isolation of a novel hydrogenotrophic methanogen. Appl. Environ. Microbiol. 74:2059-2068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Conrad, R., C. Erkel, and W. Liesack. 2006. Rice cluster I methanogens, an important group of Archaea producing greenhouse gas in soil. Curr. Opin. Biotechnol. 17:262-267. [DOI] [PubMed] [Google Scholar]
- 4.Großkopf, R., S. Stubner, and W. Liesack. 1998. Novel euryarchaeotal lineages detected on rice roots and in the anoxic bulk soil of flooded rice microcosms. Appl. Environ. Microbiol. 64:4983-4989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Harmsen, H. J. M., A. D. L. Akkermans, A. J. M. Stams, and W. M. de Vos. 1996. Population dynamics of propionate-oxidizing bacteria under methanogenic and sulfidogenic conditions in anaerobic granular sludge. Appl. Environ. Microbiol. 62:2163-2168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Harmsen, H. J. M., B. L. M. Van Kuijk, C. M. Plugge, A. D. L. Akkermans, W. M. de Vos, and A. J. M. Stams. 1998. Syntrophobacter fumaroxidans sp. nov., a syntrophic propionate-degrading sulfate-reducing bacterium. Int. J. Syst. Bacteriol. 48:1383-1387. [DOI] [PubMed] [Google Scholar]
- 7.Imachi, H., S. Sakai, Y. Sekiguchi, S. Hanada, Y. Kamagata, A. Ohashi, and H. Harada. 2008. Methanolinea tarda gen. nov., sp. nov., a methane-producing archaeon isolated from a methanogenic digester sludge. Int. J. Syst. Evol. Microbiol. 58:294-301. [DOI] [PubMed] [Google Scholar]
- 8.Imachi, H., Y. Sekiguchi, Y. Kamagata, A. Ohashi, and H. Harada. 2000. Cultivation and in situ detection of a thermophilic bacterium capable of oxidizing propionate in syntrophic association with hydrogenotrophic methanogens in a thermophilic methanogenic granular sludge. Appl. Environ. Microbiol. 66:3608-3615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Juottonen, H., P. E. Galand, E.-S. Tuittila, J. Laine, H. Fritze, and K. Yrjälä. 2005. Methanogen communities and Bacteria along an ecohydrological gradient in a northern raised bog complex. Environ. Microbiol. 7:1547-1557. [DOI] [PubMed] [Google Scholar]
- 10.Lu, Y., T. Lueders, M. W. Friedrich, and R. Conrad. 2005. Detecting active methanogenic populations on rice roots using stable isotope probing. Environ. Microbiol. 7:326-336. [DOI] [PubMed] [Google Scholar]
- 11.McInerney, M. J., C. G. Struchtemeyer, J. Sieber, H. Mouttaki, A. J. M. Stams, B. Schink, L. Rohlin, and R. P. Gunsalus. 2008. Physiology, ecology, phylogeny, and genomics of microorganisms capable of syntrophic metabolism. Ann. N. Y. Acad. Sci. 1125:58-72. [DOI] [PubMed] [Google Scholar]
- 12.Nüsslein, B., K.-J. Chin, W. Eckert, and R. Conrad. 2001. Evidence for anaerobic syntrophic acetate oxidation during methane production in the profundal sediment of subtropical Lake Kinneret (Israel). Environ. Microbiol. 3:460-470. [DOI] [PubMed] [Google Scholar]
- 13.Sakai, S., H. Imachi, S. Hanada, A. Ohashi, H. Harada, and Y. Kamagata. 2008. Methanocella paludicola gen. nov., sp. nov., a methane-producing archaeon, the first isolate of the lineage “rice cluster I,” and proposal of the new archaeal order Methanocellales ord. nov. Int. J. Syst. Evol. Microbiol. 58:929-936. [DOI] [PubMed] [Google Scholar]
- 14.Sakai, S., H. Imachi, Y. Sekiguchi, A. Ohashi, H. Harada, and Y. Kamagata. 2007. Isolation of key methanogens for global methane emission from rice paddy fields: a novel isolate affiliated with the clone cluster rice cluster I. Appl. Environ. Microbiol. 73:4326-4331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Schink, B. 1997. Energetics of syntrophic cooperation in methanogenic degradation. Microbiol. Mol. Biol. Rev. 61:262-280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sekiguchi, Y., K. Syutsubo, A. Ohashi, H. Harada, Y. Kamagata, and K. Nakamura. 1998. Phylogenetic diversity of mesophilic and thermophilic granular sludges determined by 16S rRNA gene analysis. Microbiology 144:2655-2665. [DOI] [PubMed] [Google Scholar]
- 17.Stackebrandt, E., and B. Goebel. 1994. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Bacteriol. 44:846-849. [Google Scholar]
- 18.Stams, A. J. M., K. C. F. Grolle, C. T. M. Frijters, and J. B. Van Lier. 1992. Enrichment of thermophilic propionate-oxidizing bacteria in syntrophy with Methanobacterium thermoautotrophicum or Methanobacterium thermoformicicum. Appl. Environ. Microbiol. 58:346-352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Thauer, R. K., K. Jungermann, and K. Decker. 1977. Energy conservation in chemotrophic anaerobic bacteria. Bacteriol. Rev. 41:100-180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Watanabe, T., S. Asakawa, A. Nakamura, K. Nagaoka, and M. Kimura. 2004. DGGE method for analyzing 16S rDNA of methanogenic archaeal community in paddy field soil. FEMS Microbiol. Lett. 232:153-163. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.