Abstract
Transcription activation by RNA polymerase II is a complicated process driven by combined, precisely coordinated action of a wide array of coactivator complexes, which carry out chromatin-directed activities and nucleate the assembly of the preinitiation complex on the promoter. Using various techniques, we have shown the existence of a stable coactivator supercomplex consisting of the chromatin-remodeling factor Brahma (SWI/SNF) and the transcription initiation factor TFIID, named BTFly (Brahma and TFIID in one assembly). The coupling of Brahma and TFIID is mediated by the SAYP factor, whose evolutionarily conserved activation domain SAY can directly bind to both BAP170 subunit of Brahma and TAF5 subunit of TFIID. The integrity of BTFly is crucial for its ability to activate transcription. BTFly is distributed genome-wide and appears to be a means of effective transcription activation.
Keywords: coactivators, protein complex
Activation of transcription by eukaryotic RNA polymerase II (Pol II) requires different groups of coactivators (for reviews, see refs. 1 and 2). The primary function of coactivators is to remodel and modify the chromatin template. Thus, chromatin remodelers of the Brahma (SWI/SNF-related) family play a genome-wide role in activation of Pol II-transcribed genes (3, 4). One more function of coactivators is to further recruit general transcription factors (GTFs) to form the Pol II preinitiation complex. The TFIID coactivator performs this function for most of Pol II-dependent genes (5, 6).
Different coactivators recruited to the promoter assist each other and interact in a highly organized gene-specific manner (for a review, see ref. 7). However, this important regulatory step is still poorly understood. The best studied model is that of successive one-by-one recruitment of coactivators, which, in particular, is confirmed by the fact that the recruitment of chromatin-remodeling complexes is usually a prerequisite for the efficient recruitment of GTFs to the promoter (8, 9). The opposite model proposes one-time recruitment of preexisting supercomplex of several coactivators (10–12), although the composition of such supercomplexes described to date appears to be either ambiguous or incomplete.
We have described the coactivator SAYP in Drosophila (13). SAYP is present at numerous sites on polytene chromosomes and colocalizes with Pol II in transcriptionally active euchromatin. SAYP homologs in various metazoans have an evolutionarily conserved core containing the SAY domain, which is involved in transcription activation, and 2 PHD fingers (13). Recently, SAYP was found to be associated with the chromatin-remodeling Brahma complex of the PBAP subfamily (14). Here, we show that SAYP interacts both with Brahma and with TFIID, assembling them into a stable supercomplex named BTFly (Brahma and TFIID in one assembly). The presence of all BTFly components is crucial for its function in transcription activation. An important fact is that highly purified BTFly contains the full set of TFIID and Brahma subunits and, therefore, is an example of a stably integrated full-set coactivator complex functioning at 2 consecutive stages of transcription activation.
Results
SAYP Is Present in the High-Molecular-Weight Complex Containing Brahma and TFIID.
To study the mechanism of SAYP action, a SAYP-containing complex was purified from the embryonic nuclear extract. During preliminary size fractionation of the nuclear extract from Drosophila embryos on a Superose 6 column, SAYP migrated as a sharp peak in fractions 16 and 17 containing high-molecular-weight complexes of at least 2 MDa (Fig. 1A). However, SAYP was eluted markedly later than the void volume and subsequent fraction 15, which could contain large nonspecific aggregates. Treating the initial extract with DNase I did not change the SAYP migration profile, which excluded the possibility of association through DNA fragments.
Fig. 1.
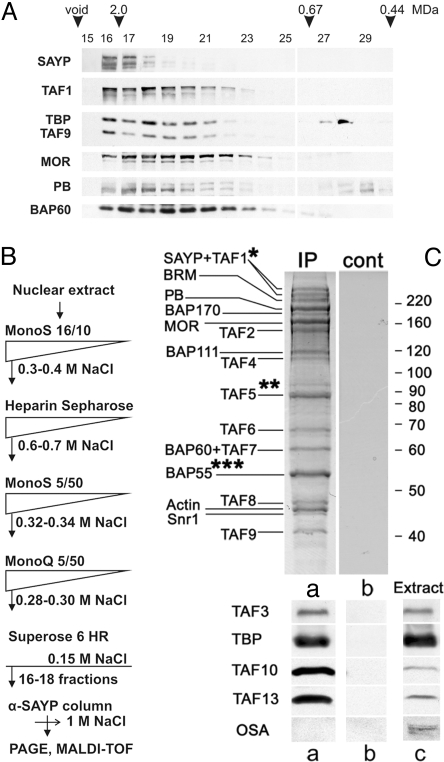
Purification of the SAYP-containing complex. (A) Gel filtration of a DNase I-treated Drosophila embryo nuclear extract on Superose 6. The fractions were analyzed for the presence of the proteins indicated. (B) Scheme of purification of the SAYP-containing complex. (C Upper) The preparation of the SAYP-containing complex was resolved by 9% SDS/PAGE and Coomassie stained to identify the SAYP-associated proteins by mass spectrometry. The staining of the control immunoprecipitate obtained with a preimmune serum is shown on the right. Band (*) TAF1 and SAYP both migrated in 2 adjacent bands. Band (**) contained the Hpr1 subunit of the THO complex (26), which was often detected in minor amounts in our SAYP-containing preparations. Band (***) additionally contained pontin, which is associated with the Brahma complex (27). (Lower) Shown are Western blot analysis of the same preparation (Ca), control IP with preimmune IgG (Cb), and nuclear extract for the presence of OSA and several components of TFIID (Cc).
To eliminate protein contaminants from the preparation of the SAYP-containing complex, we developed a multistep purification procedure (Fig. 1B and Fig. S1). At the last step, the SAYP-containing material (high-molecular-weight fractions from the Superose 6 column) was loaded onto an immunosorbent with antibodies against the SAYP N end, washed with a buffer containing 1 M NaCl, eluted with acidic glycin, and resolved by SDS/PAGE, yielding ≈20 bands upon Coomassie staining (Fig. 1C).
MALDI-TOF MS of individual bands revealed the complete set of the subunits of the Brahma chromatin-remodeling complex, including Brahma (BRM), Moira (MOR), Polybromo (PB), BAP170, BAP111, BAP60, BAP55, actin, and Snr1. The presence of PB and BAP170 but not of OSA (Fig. 1C Lower) indicated that SAYP was associated only with the PBAP subfamily of the Brahma complex (3). This is in agreement with the results reported by (14).
However, in addition to Brahma, all subunits of the TFIID complex were also found to copurify with SAYP. TAF1, TAF2, and TAF4–TAF9 were identified by MALDI-TOF MS. The presence of low-molecular-weight subunits TAF10 and TAF13 was confirmed by Western blot analysis (Fig. 1C Lower). TAF3 and TBP, which had been shown to occur in purified TFIID in substoichiometric amounts (L. Tora, personal communication) (15), were also revealed in our preparation by western blot analysis (Fig. 1C Lower). SAYP was found in the same 2 bands as TAF1 by MALDI-TOF MS.
Thus, the large protein assembly isolated as described was tripartite, comprising the SAYP and complete Brahma and TFIID complexes. Importantly, no components of other transcription-related complexes were found in comparable amounts, providing evidence for the specificity of purification. In agreement with the results of size exclusion chromatography (Fig. 1A), the molecular mass of this assembly, calculated as the sum of molecular mass of its components, was 2.5 MDa (1 MDa for PBAP, 1.3 MDa for TFIID, and 0.2 MDa for SAYP). Importantly, the material obtained at the end of the purification procedure had almost the same SAYP profile (Fig. 1A and Fig. S1), suggesting a high stability of the assembly.
To confirm the direct interaction of Brahma, TFIID, and SAYP, we performed coimmunoprecipitation experiments with the pooled SAYP-containing fraction 16 and 17 (Fig. 2A). Antibodies against PB and MOR coprecipitated SAYP as well as TAF4 and TAF5. Conversely, antibodies against TAF1 and TAF8 coprecipitated significant amounts of SAYP, PB, and MOR. No cross-precipitation was observed in the same assay with fractions eluted later (fractions 20 and 21 in Fig. 1A), which contained Brahma and TFIID but no SAYP (Fig. S2), indicating that Brahma and TFIID did not aggregate nonspecifically.
Fig. 2.
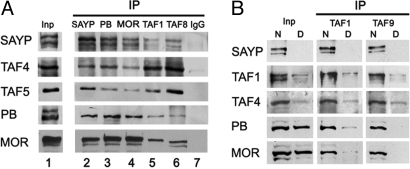
SAYP and a portion of Brahma and TFIID are assembled into 1 complex. (A) TFIID, Brahma, and SAYP are coimmunoprecipitated with each other. Fractions 16 and 17 from Superose 6 were used for IP with antibodies against SAYP, PB, MOR, TAF1, and TAF8 or with preimmune IgG. Equal parts of the input (Inp) and precipitate (IP) were tested for the proteins indicated on the left. (B) Coimmunoprecipitation of TFIID and Brahma is strongly affected by depletion of SAYP. Fractions 16 and 17 from Superose 6 (Inp) after depletion with preimmune IgG (N) or anti-SAYP antibodies (D) were taken for IP with antibodies against TAF1 and TAF9, and the precipitates (IP) were analyzed for the proteins indicated on the left. 20% of Input and 20% of IP were loaded.
Finally, the role of SAYP as a signature subunit of the Brahma-TFIID assembly was confirmed by immunodepletion of SAYP from the fraction 16 and 17 with anti-SAYP antibodies. Subsequent coimmunoprecipitation with antibodies against TAF1 or TAF9 (Fig. 2B) showed that SAYP depletion strongly suppressed the Brahma-TFIID interaction.
We then estimated the proportions of TFIID and Brahma incorporated into the assembly. The elution profile of SAYP upon gel filtration overlapped with those of TFIID and Brahma only in fraction 16 and 17 (Fig. 1A). In the immunodepletion experiment (Fig. 2B), treatment with anti-SAYP antibodies almost completely depleted fraction 16 and 17 of TAF1 and TAF9, indicating that nearly all TFIID in this fraction was SAYP-bound. In contrast, the content of Brahma subunits in the fraction was reduced relatively slightly. On this basis, we estimated that ≈20% of TFIID and only a few percent of Brahma contained in the nuclear extract were incorporated into the assembly.
SAYP Interacts with TFIID and Brahma Components in Drosophila Development.
We have described a SAYP gene mutation, e(y)3u1, which reduces the level of SAYP (13). Close analysis of males hemizygous for e(y)3u1 has shown that the viability of hemizygous males is decreased by 20%. A characteristic manifestation of e(y)3u1 is a femur maldevelopment (13) (bent femur, Fig. S3A), observed in 10% of hemizygous males (Fig. 3A).
Fig. 3.
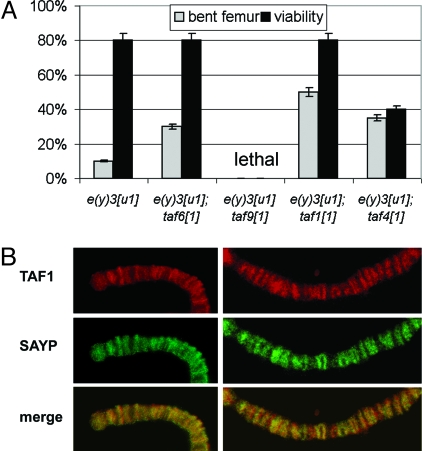
SAYP and TFIID interact in development. (A) Mutations in components of the TFIID complex strongly enhance the penetrance of the e(y)3[u1] mutant phenotype. The viability and frequency of the bent femur phenotype are shown for the genotypes indicated. The e(y)3[u1] taf9[1] compound was lethal. (B) Immunostaining of polytene chromosomes with antibodies against TAF1 and SAYP shows significant colocalization of these factors.
The e(y)3u1 flies were crossed with flies bearing TFIID mutations, which displayed no mutant phenotype or decreased viability in heterozygotes. Examination of males hemizygous for e(y)3u1 and heterozygous for the second mutation (Fig. 3A) showed that the proportion of flies with the bent femur phenotype significantly increased in all combinations. The viability of mutant flies was markedly decreased in combinations of e(y)3u1 and taf4 mutations. Moreover, the compound of weak mutations e(y)3u1 taf91 was lethal. Thus, mutations of TFIID components, even when heterozygous, aggravated the weak mutation of SAYP.
Chalkley et al. (14) provided evidence for genetic interactions of SAYP with subunits of PBAP (but not BAP). We confirmed their findings and also revealed a decrease in the viability of flies carrying a combination of e(y)3u1 and brm2 (Fig. S3B). These genetic results show that the activity of SAYP in fly development strongly depends on both Brahma and TFIID.
We also compared the distribution patterns of SAYP and TFIID on the polytene chromosomes of Drosophila larval salivary glands (Fig. 3B) and found that these patterns overlap considerably. Together with the data by (14), this fact demonstrates that SAYP shares its chromosomal sites with both TFIID and Brahma.
The SAY Domain Activates Transcription upon Interaction with TFIID and Brahma.
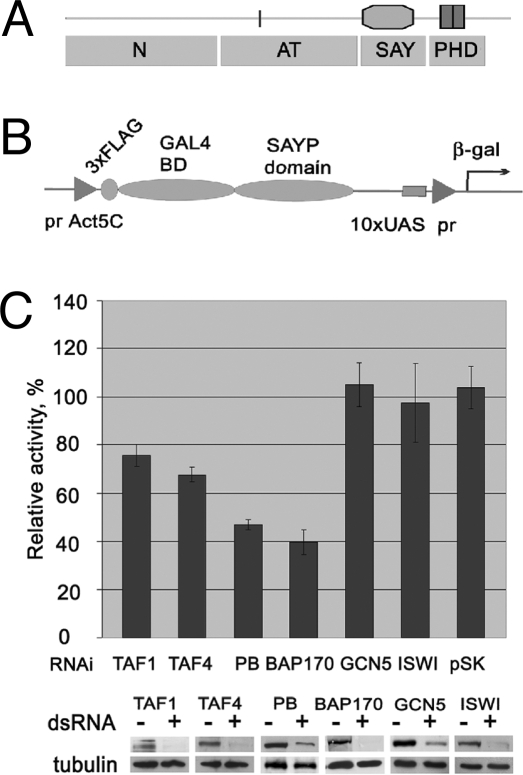
SAYP has a Drosophila-specific N-proximal part (N-terminal and central domain with an AT-hook) and the evolutionarily conserved C-proximal part with the SAY domain and 2 PHD fingers (Fig. 4A). SAY was shown to be the minimal domain activating transcription in yeast (13). To ascertain the role of particular SAYP domains in the natural environment, we designed the pTrAssay construct (Fig. 4B) harboring the lacZ reporter under the control of 10 binding sites for GAL4 BD (UAS) and the minimal TATA-containing hsp70 promoter together with the sequence encoding the GAL4 BD tagged with N-terminal 3×FLAG under the constitutive actin gene promoter. DNA fragments encoding different domains of SAYP were then inserted into the construct to be expressed in fusion with GAL4 BD. The effect of each domain on transcription was measured as β-galactosidase activity in Schneider (S2) cells.
Fig. 4.
Transcriptional activity of SAY depends on Brahma and TFIID. (A) Domain structure of SAYP: the N-terminal domain N, AT-hook-containing domain (AT), SAY, and PHD fingers. (B) Coding region of the pTrAssay construct used for transcription assays. (C Upper) Effects of knockdown of Brahma and TFIID components, ISWI, and GCN5 on the reporter gene transcription activated by SAY. In an RNAi control, cells were treated with the pSK II vector. In each case, lacZ activity was normalized to the level of SAY expression and to SAY activity in intact cells. (Lower) The knockdown efficiency was tested by Western blot analysis. The level of protein expression was analyzed in wild-type S2 cells (−) or cells harvested after 5 days of RNAi treatment (+). Tubulin was used as a loading control.
As expected, the isolated SAY domain proved capable of appreciably activating transcription in S2 cells (Fig. S4) whereas no activation was observed with any other separate domain of SAYP. However, the total evolutionarily conserved SAY-PHD region was 3 times more efficient than SAY alone; that is, the PHD fingers were important for the full-level activator function of SAYP.
For subsequent experiments, several S2 cell lines were generated that stably carried the constructs with SAY, SAY-PHDs, PHDs, or GAL4 BD alone, with the first 2 lines being designated SAYline and SPline.
To identify the factors required for the SAY domain to exhibit its activator function, we examined how the reporter gene expression in the SAYline was affected by an RNAi knockdown of several well known coactivator components (Fig. 4C). A significant reduction of transcriptional activity was observed on a knockdown of TFIID and Brahma subunits whereas a knockdown of GCN5 histone acetyltransferase (6) or ISWI ATPase of the ISWI remodeler group (4) lacked any effect.
The cooperation of SAY with TFIID and Brahma was further corroborated by immunoprecipitation of extracts from the above cell lines. The SAY domain proved to coprecipitate with TFIID and Brahma subunits but not with GCN5 or ISWI (Fig. 5A and Fig. S5A). Separate PHD fingers failed to interact with Brahma or TFIID subunits.
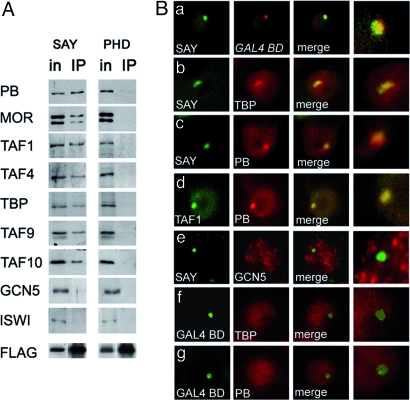
Fig. 5.
SAY domain interacts with Brahma and TFIID. (A) SAY domain is coimmunoprecipitated with Brahma and TFIID subunits. IP with antibodies against FLAG was performed with an extract of S2 cells stably expressing FLAG-GAL4 BD fusions with SAY and PHD. The precipitate was tested for the proteins indicated on the left. Five percent of Input and 25% of IP were loaded. (B) SAY colocalizes with Brahma and TFIID subunits on a transgene in the nuclei of SAYline cells. Immunostaining with antibodies against FLAG was used to detect FLAG-GAL4 BD-SAY. (a) Colocalization of SAY and the transgene was demonstrated by combined immunostaining-FISH. (b–d) TBP, PB, and TAF1 and SAY are concentrated in 1 speckle whereas GCN5 (e) is lacking. (f and g) TBP and PB do not form speckle in the control cell line stably carrying the construct with FLAG-GAL4 BD alone. Magnification: 1,000×; rightmost images (merges), 3,000×.
The association of SAY, TFIID, and Brahma was further confirmed by immunostaining of the SAYline. In stably transformed Drosophila cell lines, a transgene integrates into the genome in multiple copies, forming 1 or a few loci in the host cell chromosomes (16). First, combined DNA FISH and anti-FLAG staining demonstrated exclusive colocalization of SAY and the transgenic insert in the nucleus (Fig. 5Ba), confirming that SAY was indeed recruited to the transgenic construct. Next, simultaneous immunostaining for SAY and TBP (Fig. 5Bb) or SAY and PB (Fig. 5Bc) revealed strong accumulation of PB and TBP in the speckle containing SAY. This experiment additionally demonstrated that TBP was reliably associated with SAY in the cell. Moreover, TAF1 occurred in the same speckle as did PB (Fig. 5Bd). However, SAY did not colocalize with GCN5 (Fig. 5Be) or ISWI (Fig. S5B). Thus, SAY is the key domain in the natural activation-supporting function of SAYP and is sufficient for its interaction with TFIID and Brahma.
SAY Domain Can Directly Bind both BAP170 and TAF5.
To identify the direct partner of SAY in the Brahma complex, we reasoned that, because SAYP is associated exclusively with the PBAP subfamily of Brahma, the most likely candidates were PBAP signature subunits PB and BAP170 (3). We performed RNAi knockdown of either PB or BAP170 in the SAYline and tested whether SAY could still be coprecipitated with the Brahma core components (Fig. 6A). Knockdown of PB affected neither the total content of BAP170 or MOR, as already reported (17), nor the association of SAY with MOR and BAP170. On the contrary, knockdown of BAP170 strongly decreased the association of SAY with MOR (with the overall level of MOR remaining unchanged), partly reducing the overall PB level and strongly affecting the PB association with SAY. Thus, BAP170 but not PB is crucial for the association of SAY with Brahma.
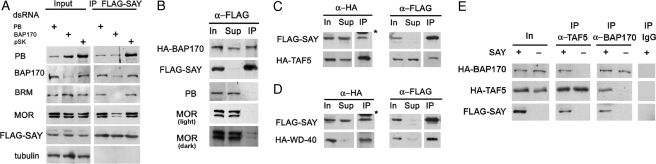
Fig. 6.
SAY directly interacts with BAP170 and WDR domain of TAF5. (A) BAP170 is crucial for the association of Brahma with SAYP. RNAi knockdown of either PB or BAP170 (indicated at the top) in SAYline cells was performed. Control cells were treated with pSK dsRNA. Cell extracts (Input) and proteins coimmunoprecipitated with SAY (IP FLAG-SAY) were tested by Western blotting for the subunits of Brahma as indicated. Tubulin was used as a loading control. (B) SAY directly binds BAP170. HA-BAP170 was coexpressed with FLAG-SAY. IP from cell extract was performed with anti-FLAG antibodies. Aliquots of the extract (In, 5%), supernatant fluid (Sup, 5%), and immunoprecipitate (IP, 25%) were analyzed for the overexpressed proteins using antibodies against HA and FLAG and Brahma subunits. On longer exposure, precipitation of a small amount of MOR subunit was revealed. (C and D) SAY directly binds WDR domain of TAF5. Either HA-TAF5 (C) or HA-WDR (D) domain of TAF5 were coexpressed with FLAG-SAY. IP from cell extract was performed with anti-HA or anti-FLAG antibodies. Aliquots of the cell extract (In, 5%), supernatant fluid (Sup, 5%), and immunoprecipitate (IP, 25%) were analyzed for TAF5, the WDR domain, and SAY using antibodies against HA or FLAG. *, IgG heavy chain. (E) TAF5 and BAP170 interact with each other in presence of SAY domain. HA-TAF5 and HA-BAP170 were coexpressed either in S2 cells (SAY−) or in SAYline (SAY+). IP from cell extract was performed with specific anti-TAF5 or anti-BAP170 antibodies. Aliquots of the extract (In, 5%) and immunoprecipitate (IP, 25%) were analyzed using antibodies against HA or FLAG tags. IP on preimmune IgG was used as a control.
Next, we checked whether SAY could directly bind BAP170. SAYline cells were transfected with a construct expressing HA-tagged BAP170. Immunoprecipitation with anti-FLAG antibodies revealed a strong interaction of SAY with BAP170 (Fig. 6B). Notably, SAY association with the endogenous subunits of Brahma was strongly reduced (compare Figs. 5A and 6B) but not completely abolished. Most probably, overexpressed HA-BAP170 competitively replaced the endogenous Brahma, which is evidence for a major role of BAP170 in mediating the SAY-Brahma interaction. In a control experiment, no coprecipitation of FLAG-GAL4 BD and HA-BAP170 was detected (Fig. S6A).
To identify the SAYP partners in TFIID, we performed the RNAi knockdown of several TAFs (TAF1, TAF2, TAF4, and TAF5) in SAYline cells and estimated the association of SAY with other TFIID subunits by coimmunoprecipitation. Knockdown of TAF5 affected the association of SAY with all other TAFs. However, TAF5 is a key structural component of TFIID and knockdown of TAF5 impairs the integrity of the complex (18). Hence, we directly checked whether TAF5 and SAY interacted in the cell.
SAYline cells were transfected to express HA-tagged TAF5. Reciprocal immunoprecipitation demonstrated a significant association of HA-TAF5 and FLAG-SAY (Fig. 6C). TAF5 contains an evolutionarily conserved WDR domain, which is thought to mediate the interaction of TFIID with other complexes (19). We tested the WDR domain of TAF5 for interaction with SAY. The HA-tagged WDR domain was expressed in the SAYline and displayed almost complete coprecipitation with FLAG-SAY (Fig. 6D). Controls showed no coprecipitation of FLAG-GAL4 BD and HA-TAF5 or HA-WDR (Fig. S6 B and C). Importantly, the separate WDR domain of TAF5 is not incorporated into the TFIID complex in the cell (19, 20). Therefore, the whole TFIID did not contribute to the observed interaction of SAY and the WDR domain.
Finally, we reconstituted the triple TAF5-SAY-BAP170 protein complex. All 3 subunits were coexpressed in S2 cells, and immunoprecipitation revealed association of TAF5 with BAP170. This interaction was not detected in the absence of SAY domain (Fig. 6E).
The above data do not allow us to completely exclude the existence of other mediators of TFIID-SAYP-Brahma interactions and may not exactly reflect interactions in the endogenous assembly. Nevertheless, they provide a sound basis for the model that SAYP couples TFIID and Brahma via the direct interaction of its SAY domain with TAF5 and BAP170.
SAYP, TFIID, and Brahma Are Interdependently Recruited to Promoters.
To investigate the mode of the assembly recruitment to chromatin in vivo, we chose 3 genes (CG11395, CG11400, and globin1) whose transcription was found to drop by a factor of 7–10 upon SAYP depletion by RNAi in S2 cells. Chromatin immunoprecipitation (ChIP) with TFIID-, SAYP-, and Brahma-specific antibodies confirmed that all 3 factors were stably associated with the promoters and much less with the coding regions of these genes (Fig. S7 A and B).
Next, we knocked down by RNAi single proteins of the assembly (SAYP, BAP170, or TAF4) in S2 cells and examined the abundance of SAYP, Brahma, and TFIID subunits, as well as Pol II, on the promoters by ChIP. The occupancies of all 3 promoters proved to change in a very similar manner (Fig. 7A and Fig. S7 C and D).
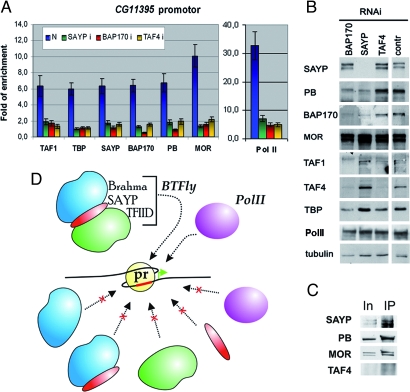
Fig. 7.
Mode of SAYP, TFIID, and Brahma recruitment to SAYP-dependent genes. (A) SAYP, TFIID, and Brahma are interdependently recruited and are all crucial for gene activity. The levels of SAYP, the subunits of TFIID and Brahma, and Pol II on the CG11395 promoter were assayed in normal cell (N, blue) and after knockdown of SAYP (green), BAP170 (red), and TAF4 (yellow). ChIP results are given as an enrichment relative to 28S rDNA. (B) Total levels of SAYP and the subunits of TFIID and Brahma after knockdown of SAYP, BAP170, or TAF4 and in cells treated with pSK dsRNA. (C) SAYP is coimmunoprecipitated with Brahma in the absence of TFIID. IP with antibodies against SAYP was performed with an extract of S2 cells after knockdown of TAF4. Input (25%) and IP (50%) were loaded. (D) Model of the joint recruitment of SAYP, TFIID, and Brahma in 1 BTFly supercomplex to the promoter (pr) contained in a chromatin template. Partial variants of BTFly are not recruited; neither do they activate transcription.
Knockdown of SAYP led to a significant loss of both TFIID and Brahma from the promoters. However, the total level of PB and BAP170 was also affected, which, in turn, should affect the recruitment of PBAP to chromatin (17). The total level of TAFs and TBP was not affected (Fig. 7B). Thus, we can state that TFIID could not be recruited to the promoters examined in the absence of SAYP and Brahma. As a negative control, we used the hsp70 promoter, which is not occupied by SAYP in normal cells (Fig. S7E). The recruitment of TFIID and Brahma to the hsp70 promoter was not impaired after knockdown of SAYP.
Likewise, knockdown of BAP170 strongly impaired the binding of all factors under study with the promoters. However, a noticeable effect was also observed for the total level of both PBAP and TFIID subunits, but not for the level of SAYP, which should exist in a free state under these conditions. These findings indicate that free SAYP cannot be recruited to the SAYP-dependent promoters.
Knockdown of TAF4 also had a strong negative effect on the binding of Brahma and SAYP to the promoters. As expected, TAF4 knockdown led to degradation of the whole TFIID (18) without affecting the total level of SAYP or Brahma. Significantly, SAYP still remains to be associated with Brahma (Fig. 7C). However, this partial assembly (SAYP-Brahma without TFIID) is incapable of stable association with the promoter.
Moreover, knockdown of SAYP, BAP170, or TAF4 significantly decreased the level of Pol II on the promoters examined (Fig. 7 A and B and Fig. S7 C and D), indicating that the simultaneous presence of SAYP, TFIID, and Brahma on the promoters is important for their function in transcription activation.
In summary, these results testify to a mode of gene activation based on the interdependent recruitment of Brahma, SAYP, and TFIID on the gene promoter. This observation further supports the idea of SAYP-dependent association of TFIID and Brahma in 1 supercomplex.
Discussion
In the study, we report the discovery of a large supercomplex integrating 2 main players in transcription activation, Brahma and TFIID, which we named BTFly. Brahma and TFIID are united by coactivator SAYP, which we have described in ref. 13. BTFly includes all subunits of TFIID and Brahma (PBAP subfamily), but not comparably abundant subunits of other coactivators, and is stable in the absence of a chromatin template according to biochemical evidence. Functional cooperation of SAYP, TFIID, and Brahma in development has been verified in genetic experiments.
We estimate that approximately 20% of TFIID and a few percent of Brahma are embodied into BTFly in embryonic nuclear extracts. Apparently, BTFly-mediated transcription activation is widely used in the Drosophila genome because SAYP has been found in ≈150 euchromatin sites on polytene chromosomes, all containing Pol II (13).
Chalkley et al. (14) describe SAYP as a Brahma-associated protein and did not report the presence of TFIID subunits in preparations of the Brahma complex purified using antibodies against BRM and PB. In our opinion, a probable explanation is that these preparations contained a manifold excess of SAYP- (and TFIID)-free Brahma, the more so that the amounts of SAYP in them were barely traceable. In our study, we have shown that SAYP directly unites Brahma and TFIID, with a relatively small proportion of Brahma being incorporated into this assembly. Also we have shown that SAYP-associated Brahma (i.e., its form is considered in ref. 14) is unfit for stable recruitment to the promoters.
The results of experiments with recombinant proteins suggest a structural model with the SAY domain of SAYP taken to be the linchpin of the BTFly complex. SAY directly interacts with the TAF5 subunit of TFIID and the BAP170 subunit of Brahma, assembling them into one complex. Importantly, SAY is evolutionarily conserved, suggesting a conservation of the coupling of TFIID and Brahma in other metazoans.
By means of ChIP, we have revealed BTFly on the promoters of SAYP-dependent genes. The presence of all components of BTFly is crucial for its recruitment and gene activation (Fig. 7D). We have shown that the recruitment of SAYP, TFIID, or Brahma in the free state is impaired. SAYP-associated Brahma in the absence of TFIID is not recruited to the SAYP-dependent promoters, although no impediments are expected in this case according to the model of a sequential recruitment of remodeling complexes and TFIID. We may conclude that BTFly functions as a single entity in transcription activation.
The coupling of TFIID and Brahma by BTFly may serve to increase the efficiency of transcription activation of a definite gene. Indeed, chromatin remodeling is crucial for transcription initiation to occur (8), and TFIID binding is a rate-limiting step of transcription initiation in vivo (21).
Thus, we consider that the direct coupling of different activities may be an important way of controlling gene expression, which is as yet poorly understood. BTFly as a probable example of a relatively simple nuclear supercomplex appears to be a useful tool for further research in this field.
Materials and Methods
Antibodies.
Details about the antibodies are given in SI Text and Fig. S8.
Purification of BTFly.
We used 0.4 g (by protein) of a nuclear extract prepared from 0- to 12-h Drosophila embryos by 0.42 M ammonium sulfate extraction (22) and heparin Sepharose, MonoS HR 16/10, MonoS 5/50 GL, MonoQ 5/50 GL, and Superose 6 HR 10/30 columns (GE Healthcare). Details of the purification procedure are shown in Fig. 1B. The columns were equilibrated with the HEMG buffer [25 mM Hepes-KOH (pH 7.6), 12.5 mM MgCl2, 0.1 mM EDTA, 10% glycerol, 1 mM DTT, and complete protease inhibitor mixture (Roche)] containing 150 mM NaCl (HEMG-150). Immunoaffinity purification was performed on a column prepared by coupling anti-SAYP antibodies to protein A Sepharose (Sigma) by DMP (Sigma) according to a published protocol (23). We used HEMG-150 with 0.1% Nonidet P-40 for loading, HEMG-1000 with 0.1% Nonidet P-40 for washing, and 0.1 M glycine (pH 2.5) for elution. Eluted proteins were resolved by SDS/PAGE and stained with Coomassie. The protein bands were cut out and subjected for in-gel trypsin digestion. MALDI-TOF MS was performed using an Ultraflex II mass-spectrometer (Bruker Daltonics). Protein spectra were internally calibrated using trypsin autolysis products, and the resulting peptide weights were searched against the nonredundant database maintained by the National Center for Biotechnology Information, using the Mascot search engine.
The Superose 6 column was calibrated with an HMW Calibration Kit (GE Healthcare). The void volume of the column was 7.0 mL, and the volume of each fraction was 0.5 mL.
Experiments with Cell Culture.
Drosophila Schneider line 2 (S2) cells were maintained at 25 °C in Schneider's insect medium (Sigma) containing 10% FBS (HyClone). To extract proteins, S2 cells were lyzed in 10 mM Hepes (pH 7.9) containing 5 mM MgCl2, 0.5% Nonidet P-40, 0.45 M NaCl, 1 mM DTT, and complete protease inhibitor mixture (Roche). IP was performed as described in ref. 24. Before IP, the extract was treated with DNase I (USB, 0.6 units/mL) and RNase (Stratagene, 10 units/mL).
Description of transfection, RNAi, and immunostaining experiments are given in SI Text.
DNA Constructs and Reporter Gene Assay.
Details are given in SI Text. The transcription activation level was estimated as the ratio of β-galactosidase activity to the amount of the FLAG-tagged protein in the cell lysate.
ChIP and Quantitative (q) PCR Analysis.
ChIP was performed according to published procedures (25). DNA was sheared to size of ≈300 bp. Cells (3 × 106) and 10 μg of an antibody were taken for one experiment. After ChIP, the recovered DNA was analyzed by qPCR with MiniOpticon (Bio-Rad). The primers for analysis are described in SI Text. The qPCR results were normalized to 28S rDNA. All experiments were repeated at least 3 times.
Supplementary Material
Acknowledgments.
We thank N.A. Gorgoluk, T.I. Tkacheva, and A.V. Galkin for their critical reading of the manuscript; P. Verrijzer and Y. Moshkin (Erasmus University Medical Center, Rotterdam) for antibodies and microarray data; J. Kadonaga and L. Tora (Institut de Genetique et de Biologie Moleculaire et Cellulaire, Strasbourg, France) for antibodies; and I. Toropygin for his assistance with MALDI experiments. This work was supported by the “Molecular and Cell Biology” program of the Russian Academy of Sciences and a Scientific School Support Grant 2994.2008.4 and Russian Foundation for Basic Research Grants 07-04-00723, 08-04-91976, and 08-04-12216. Y.S. acknowledges a fellowship from University of Oslo, Centre for Medical Studies in Russia and was supported by the Young Scientists Support Program of the President of the Russian Federation (project No. MK-4106.2007.4).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/cgi/content/full/0901801106/DCSupplemental.
References
- 1.Naar AM, Lemon BD, Tjian R. Transcriptional coactivator complexes. Annu Rev Biochem. 2001;70:475–501. doi: 10.1146/annurev.biochem.70.1.475. [DOI] [PubMed] [Google Scholar]
- 2.Taatjes DJ, Marr MT, Tjian R. Regulatory diversity among metazoan co-activator complexes. Nat Rev Mol Cell Biol. 2004;5:403–410. doi: 10.1038/nrm1369. [DOI] [PubMed] [Google Scholar]
- 3.Mohrmann L, Verrijzer CP. Composition and functional specificity of SWI2/SNF2 class chromatin remodeling complexes. Biochim Biophys Acta - Gene Structure and Expression. 2005;1681:59–73. doi: 10.1016/j.bbaexp.2004.10.005. [DOI] [PubMed] [Google Scholar]
- 4.Neely KE, Workman JL. The complexity of chromatin remodeling and its links to cancer. Biochim Biophys Acta. 2002;1603:19–29. doi: 10.1016/s0304-419x(02)00067-7. [DOI] [PubMed] [Google Scholar]
- 5.Albright SR, Tjian R. TAFs revisited: More data reveal new twists and confirm old ideas. Gene. 2000;242:1–13. doi: 10.1016/s0378-1119(99)00495-3. [DOI] [PubMed] [Google Scholar]
- 6.Muller F, Tora L. The multicoloured world of promoter recognition complexes. EMBO J. 2004;23:2–8. doi: 10.1038/sj.emboj.7600027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cosma MP. Ordered recruitment: Gene-specific mechanism of transcription activation. Mol Cell. 2002;10:227–236. doi: 10.1016/s1097-2765(02)00604-4. [DOI] [PubMed] [Google Scholar]
- 8.Morse RH. Transcription factor access to promoter elements. J Cell Biochem. 2007;102:560–570. doi: 10.1002/jcb.21493. [DOI] [PubMed] [Google Scholar]
- 9.Narlikar GJ, Fan HY, Kingston RE. Cooperation between complexes that regulate chromatin structure and transcription. Cell. 2002;108:475–487. doi: 10.1016/s0092-8674(02)00654-2. [DOI] [PubMed] [Google Scholar]
- 10.Parvin JD, Young RA. Regulatory targets in the RNA polymerase II holoenzyme. Curr Opin Genet Dev. 1998;8:565–570. doi: 10.1016/s0959-437x(98)80012-9. [DOI] [PubMed] [Google Scholar]
- 11.Nakamura T, et al. ALL-1 is a histone methyltransferase that assembles a supercomplex of proteins involved in transcriptional regulation. Mol Cell. 2002;10:1119–1128. doi: 10.1016/s1097-2765(02)00740-2. [DOI] [PubMed] [Google Scholar]
- 12.Saurin AJ, et al. A Drosophila Polycomb group complex includes Zeste and dTAFII proteins. Nature. 2001;412:655–660. doi: 10.1038/35088096. [DOI] [PubMed] [Google Scholar]
- 13.Shidlovskii YV, et al. A novel multidomain transcription coactivator SAYP can also repress transcription in heterochromatin. EMBO J. 2005;24:97–107. doi: 10.1038/sj.emboj.7600508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chalkley GE, et al. The transcriptional coactivator SAYP is a trithorax group signature subunit of the PBAP chromatin remodeling complex. Mol Cell Biol. 2008;28:2920–2929. doi: 10.1128/MCB.02217-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sanders SL, Garbett KA, Weil PA. Molecular characterization of Saccharomyces cerevisiae TFIID. Mol Cell Biol. 2002;22:6000–6013. doi: 10.1128/MCB.22.16.6000-6013.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Echalier G. Drosophila Cells in Culture. San Diego: Academic; 1997. [Google Scholar]
- 17.Moshkin YM, Mohrmann L, van Ijcken WF, Verrijzer CP. Functional differentiation of SWI/SNF remodelers in transcription and cell cycle control. Mol Cell Biol. 2007;27:651–661. doi: 10.1128/MCB.01257-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wright KJ, Marr MT, Tjian R. TAF4 nucleates a core subcomplex of TFIID and mediates activated transcription from a TATA-less promoter. Proc Natl Acad Sci USA. 2006;103:12347–12352. doi: 10.1073/pnas.0605499103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Dubrovskaya V, et al. Distinct domains of hTAFII100 are required for functional interaction with transcription factor TFIIF beta (RAP30) and incorporation into the TFIID complex. EMBO J. 1996;15:3702–3712. [PMC free article] [PubMed] [Google Scholar]
- 20.Kokubo T, et al. Molecular cloning, expression, and characterization of the Drosophila 85-kilodalton TFIID subunit. Mol Cell Biol. 1993;13:7859–7863. doi: 10.1128/mcb.13.12.7859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wu SY, Chiang CM. TATA-binding protein-associated factors enhance the recruitment of RNA polymerase II by transcriptional activators. J Biol Chem. 2001;276:34235–34243. doi: 10.1074/jbc.M102463200. [DOI] [PubMed] [Google Scholar]
- 22.Sandaltzopoulos R, Quivy JP, Becker PB. Analysis of protein/DNA interactions by solid-phase footprinting. Methods Mol Cell Biol. 1995;5:176–181. [Google Scholar]
- 23.Harlow E, Lane D. Antibodies: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1988. [Google Scholar]
- 24.Georgieva S, et al. Two novel Drosophila TAF(II)s have homology with human TAF(II)30 and are differentially regulated during development. Mol Cell Biol. 2000;20:1639–1648. doi: 10.1128/mcb.20.5.1639-1648.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Boehm AK, Saunders A, Werner J, Lis JT. Transcription factor and polymerase recruitment, modification, and movement on dhsp70 in vivo in the minutes after heat shock. Mol Cell Biol. 2003;23:7628–7637. doi: 10.1128/MCB.23.21.7628-7637.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rehwinkel J, et al. Genome-wide analysis of mRNAs regulated by the THO complex in Drosophila melanogaster. Nat Struct Mol Biol. 2004;11:558–566. doi: 10.1038/nsmb759. [DOI] [PubMed] [Google Scholar]
- 27.Diop SB, et al. Reptin and Pontin function antagonistically with PcG and TrxG complexes to mediate Hox gene control. EMBO Rep. 2008;9:260–266. doi: 10.1038/embor.2008.8. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.