Abstract
While a monodisperse size distribution is common within one kind of spherical virus, the size of viral shells varies from one type of virus to another. In this article, we investigate the physical mechanisms underlying the size selection among spherical viruses. In particular, we study the effect of genome length and genome and protein concentrations on the size of spherical viral capsids in the absence of spontaneous curvature and bending energy. We find that the coat proteins could well adjust the size of the shell to the size of their genome, which in turn depends on the number of charges on it. Furthermore, we find that different stoichiometric mixtures of proteins and genome can produce virus particles of various sizes, consistent with in vitro experiments.
Introduction
All viruses, from the simplest to the most complicated, are constituted of a protein shell known as the capsid that encloses the genetic material or genome of the virus. The viral genome is either DNA or RNA, which can be in single, partial double, or complete double strand form (1). A capsid is built up of a large number of copies of either one kind of protein subunit or of a small number of similar ones. Under physiological conditions, the coat proteins of many single-stranded (ss) RNA viruses carry ∼6–14 positive charges on a sequence that faces the interior of the capsid (2–5). These positive charges are often located on the amino terminal end of the coat proteins, although sometimes this RNA binding domain or motif is on the C-terminal end. Since in the absence of genome, capsids typically do not form, it is generally accepted that the interaction between the positively charged RNA binding domain and the negatively charged RNA drives the capsid assembly under physiological conditions (6). Approximately half a century ago, Bancroft (7), Bancroft et al. (8,9), Hiebert et al. (10), and Verduin and Bancroft (12) conducted a series of pioneering in vitro experiments, in which they showed that under appropriate solution conditions the protein subunits of many simple RNA viruses readily encapsulate not only their own RNA but also heterologous and nonviral RNAs as well as linear polyanions such as poly(vinyl sulfonate). These experiments confirm that interactions between the positively charged capsid protein subunits and the negatively charged genome are largely nonspecific, that they are electrostatic in origin, and that they constitute the main driving force for virus assembly.
The spontaneous incorporation of genome in the capsid seems to be the distinguishable feature of self-assembly of single-stranded RNA viruses and gives rise to a number of important physical questions. Is there a relation between the length of the encapsidated genome and the capsid radius? What regulates the number of genome molecules that are incorporated? What precisely is the role of the electrostatic interactions in the self-assembly? Does the stoichiometry or ratio of the protein and genome concentrations have any impact on the size of the spherical viral shells that form in the solution? A number of important questions related to virus structure and stability have recently been addressed theoretically (2,13–20). Many questions remain open, however—in particular, those relating to size selection and the role of genome therein.
The focus of this article is on small spherical ss-RNA viruses whose capsids adopt structures with icosahedral symmetry. The number of proteins constituting icosahedral shells equals q = 60 × T, i.e., 60 times the T number, a structural index of viral capsids. It adopts certain integer (“magic”) values 1, 3, 4, 7, 9, 12, and so on, because icosahedral symmetry demands that T = h2 + k2 + hk, with h and k equal to nonnegative integers (21). Recently, it has been shown that the appearance of both icosahedral symmetry and the T-number organization is plausibly a direct consequence of free energy minimization of a very generic interaction that captures the crucial elements of capsid self-assembly: the attraction required for the aggregation and the excluded-volume repulsion due to subunit conformational rigidity (14,22). Still, the mechanism of the selection of one specific capsid size from many possibilities is not very well understood.
The recent model calculations of Zandi et al. (14) indicate that while the energy per protein subunit of the T-structure shells is indeed the lowest among all the other spherical shell types, this energy is almost the same for the smaller icosahedral structures of T = 3, 4, 7, at least in the absence of a strong bending energy and/or a preferred spontaneous curvature. Hence, one would expect that other factors such as the length of genome and/or stoichiometric ratios could provide an answer to the question as to how a virus selects its size, i.e., how it chooses one of the T numbers from all the allowed ones. For the case of ss-RNA viruses, it seems that there is a correlation between genome length and the size of the capsid, as in fact between the total number of positive charges on the RNA binding domains of a capsid and that on the native genome (2).
We note in this context that the genome of T = 1 viruses typically consists of fewer nucleotides than that of T = 3 viruses and should therefore also have a smaller physical extent in free solution. For example, satellite tobacco necrosis virus (STNV) is a T = 1 icosahedral virus with a diameter of 18.4 nm. Its genome consists of as few as 1239 nucleotides (23). Cowpea chlorotic mottle virus (CCMV), on the other hand, is a larger T = 3 icosahedral virus with a diameter of 27.8 nm. The genome of CCMV consists of four RNA molecules. Genomes 1 and 2 are each encapsidated separately, whereas genomes 3 and 4 are packaged together in yet another capsid. RNAs 1, 2, 3, and 4 have 3171, 2774, 2173, and 824 nucleotides, indicating that the total length of encapsidated genome is more or less the same in each capsid. These three types of CCMV capsids are virtually indistinguishable in terms of size and structure, and consist of q = 180 copies of the same capsid protein (1,3,5).
Due to the secondary and tertiary structures, not just the length but also the physical, three-dimensional size of RNA is important for virus assembly. For instance, recent experiments on brome mosaic virus (BMV) show that the alteration of gene order in viral RNA can seriously compromise encapsulation (4,5). While it seems plausible that gene order and nucleotide sequence should impact the size of the RNA in solution, no theory has yet been advanced that predicts the shape or even the radius of gyration of a realistic viral RNA molecule as a function of the length and/or nucleotide sequence. RNA molecules with identical molecular weight but different sequences could well have very different sizes (24), because their secondary and hence tertiary structures are a function of the primary structure. We are not aware of any experiments that shed light on the size and structure of viral RNA as a function of the molecular weight or length. (There is size information on much shorter tRNA structures (15) and on the size of the RNA of tobacco mosaic virus (TMV), a cylindrical virus (25)). The only relevant experimental data we are aware of are recent measurements of the radius of gyration in free solution of RNA 2 of CCMV. By means of radiation scattering, Gopal et al. discovered that the radius of gyration of RNA 2 must be ∼40% larger than the inner radius of the capsid shell (A. Gopal, C. Knobler, and W. M. Gelbart, unpublished).
As is well established, due to the possibility of a multitude of complementary pairing arrangements of the bases, a single-stranded RNA chain can adopt many different secondary structures with energies that are within a few kBTs from each other (27). Recent studies in fact indicate that viral ss-RNAs must have a secondary structure quite different from those of random RNAs or ribosomal RNAs, because the longest path across the secondary structure is smaller by as much as one-third (28). This might lead to a much more compact structure for viral RNAs than others, and may well be due to evolutionary pressures preventing encapsulation of nonviral RNAs. This is, of course, one of the open questions: why in vivo a virus predominantly encapsulates its own RNA in favor of other RNAs present in the cytosol.
Because so much more is known about the properties of synthetic polymers, encapsidation studies in which RNA is replaced by a polyanion or a charged colloid are definitely helpful in addressing some of the issues in hand. For instance, to explore the relation between the capsid diameter and the size of its cargo, Sun et al. (29) studied the packaging of functionalized gold nanoparticles by BMV capsid proteins, and found that the size of formed capsids increased from T = 1 to T = 3 via pseudo T = 2 structures with increasing size of the gold cargos. More recently, Hu et al. (30) investigated the encapsidation of poly(styrene sulfonate) or PSS in solutions containing CCMV coat proteins for molecular weights ranging from 400 kDa to 3.4 MDa. Interestingly, they found that the capsid size jumps from 22 nm (corresponding to a pseudo-icosahedral T = 2 capsid) for PSS of a molecular weight of 1 MDa or below it, to 27 nm (corresponding to a T = 3 capsid) for molecular weights of 2 MDa and higher. No data are available in the crossover region.
It is important to point out that several experiments (29,30,31) indicate that even if CCMV and BMV capsid proteins prefer to meet at a certain angle (i.e., have a preferred curvature or size), the deviation from this preferred curvature cannot be costly enough to prevent the capsids of CCMV and BMV from forming other T numbers in addition to the native T = 3. We also note here that more recent experiments by Dragnea (B. Dragnea, unpublished) show that BMV capsid proteins are not able to form T = 4 or larger capsids even though the cargo inside is sufficiently large for this purpose (B. Dragnea, private communication, 2008). This could indicate that the coat proteins of BMV are less flexible than those of CCMV, and that the final size of BMV capsids can deviate only to certain extent from the one dictated by the spontaneous curvature of the coat proteins. Nevertheless, if somehow one increases the strength of interaction between BMV coat proteins and its cargo (for example by changing the cargo), it might be possible to obtain larger BMV capsids by overcoming the energy cost of the deviation from the preferred curvature.
In addition to the aforementioned studies on the effect of cargo size on the capsid diameter, the influence of stoichiometry on the encapsidation of low molecular weight PSS by CCMV coat protein has recently been studied by Sikkema et al. (33). Following up on the early experiments of Verduin and Bancroft (12), they investigated a change in capsid size as they increased the ratio of the PSS to coat protein concentrations. Remarkably, at constant coat protein concentration, small stoichiometric ratios produced both T = 1 and T = 3 particles whereas at higher ratios only T = 1 particles formed in solution. A similar trend of decreasing capsid size with increasing stoichiometric ratio was in fact found by Verduin and Bancroft (12), using ss-RNA from the rodlike TMV and coat proteins from different spherical bromoviruses. CCMV proteins, for instance, encapsidate TMV RNAs in viral particles of decreasing size from T = 7 to T = 4 for RNA-protein concentration ratio changing from 1:10 to 1:6 (12).
To investigate how the effect of genome size as well as protein and RNA concentrations impact the size of icosahedral viral shells, we apply a Flory theory to estimate the free energy gain of encapsulation and a mass action model to calculate the balance between assembled and disassembled states. In the Flory theory, we presume the interactions between the coat proteins and the genome, and between the parts of the genome, to be predominantly of the screened Coulomb-type. The focus of this article is on those viruses that are able to encapsidate heterologous RNAs as well as synthetic polyelectrolytes. Thus, for our genome models we use both linear and randomly branched chains. We find that for both branched and linear polymers, the optimal size of capsid is set by the size of the genome, which in turn depends on its length and charge density.
With our simple scaling theory, we not only can reproduce the results of the more elaborate calculations preformed for linear chains in the literature (19,30), we are also able to investigate the case of branched polymers, which have not been studied in detail yet, with more sophisticated methods (34). We note that the scaling theories have already been applied to study the conformations of semiflexible chains in elastic tubes by Brochard-Wyart et al. and valuable results were obtained (35). In addition to genome size, we investigate the effect of the stoichiometric ratio of the protein and genome concentrations on the optimal size of the capsid and find that at high concentrations of genome the coat proteins form smaller capsids consistent with the experiments performed in the literature (12,33).
One of the main assumptions of our study is that the solution of capsid proteins and genome is in equilibrium. A recent analysis by Zlotnick indeed reveals that the assembly of a considerable number of viruses follows a reversible path, confirming our assumption of equilibrium here (see (36) and references cited therein). It is also shown in Zlotnick (36) that even if the last step of capsid formation is irreversible, the concentration of individual subunits and capsids still approximately follow the law of mass action at the timescales relevant to the assembly experiments, and therefore this kind of irreversibility does not change the main conclusions of this article.
The outline of the article is as follows. In Genome Encapsulation, we present our scaling equations for the capsid-genome and genome-genome interactions, and show that the free energy of the system goes through a minimum determining the optimal size of the capsid. In Role of Stoichiometry, we investigate the dependencies of the optimum size of capsid on the stoichiometry ratio of the protein to genome concentrations. In Conclusion, we discuss our findings and their implications, and summarize.
Genome Encapsulation
Polymer molecules in confined spaces are found in a wide variety of physical, chemical, and biological contexts, and many aspects have been studied over the last half century (37). What seems unique about viral encapsidation of polymers is the presence of an additional degree of freedom, the flexibility of viral coat proteins in forming discretely sized shells. The other interesting aspect is that ss-RNAs are a special kind of polyelectrolyte because of their secondary and tertiary structures that are not necessarily fixed but might adapt to the conditions that they find themselves in, i.e., they may represent annealed polymeric structures (27,38).
It seems likely that small ss-RNAs are indeed annealed structures in free solution but it is not certain whether this is still true for viral ss-RNAs because they are quite large even for the low T-number viruses. There are indications that series of local stem-loop structures that likely form during the process of replication of RNA are in fact quite long lived, even if they do not actually represent the minimum free energy configurations (39). Inside the virus capsid, the RNA has presumably undergone a considerable restructuring that includes a significant amount of duplexing, but stem-loop structures that interact with the binding domains still remain important. Here we first focus on the experiments of Hu et al. (30) and take a linear polyelectrolyte as our model for the genome. We will then evaluate the effects of random branching on the genome in the subsequence section.
Linear chain
We use a Flory type mean-field theory to calculate the free energy of n linear polymers of M segments each enclosed in a spherical shell of radius R (see Fig. 1). This free energy is the sum of an elastic compression of the chain inside the shell, a self-interaction accounting for self-avoidance and the interaction energy of the chain with the inner wall of the capsid. If because of the interaction with the wall the polymer is confined to a region of thickness a ≤ D ≤ R, and if the segment distribution is more or less uniform in this region, the free energy ΔFn of the n chains must obey the general form (37,40)
| (1) |
in units of thermal energy kBT = β−1. Here, c1, c2, and c3 are numerical constants, a the effective Kuhn length of the chains, υ the mutually excluded volume of the Kuhn segments, and γ the strength of the attraction between a polymer segment and the capsid wall if it is within a range b (b << D ≤ R) of that wall. If the inner capsid wall were smooth and uniformly charged, b would be of the order of the Debye screening length (19). For some viruses such as CCMV this is not an accurate representation of the state of affairs, not least since most (but not all) of the positive charges reside on the disordered RNA binding domain that penetrates the inner cavity (2). Therefore, the quantity b should be seen as some effective interaction length and by the same token γ is an effective interaction strength.
Figure 1.
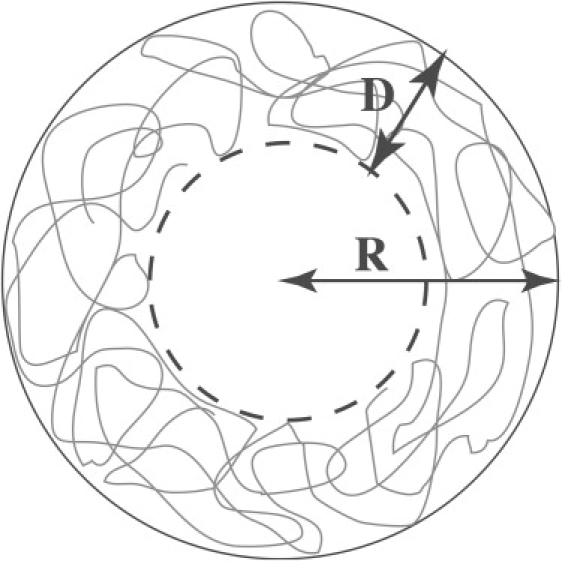
Schematic view of a polymeric molecule inside a capsid. D indicates the thickness of the adsorbed layer, R the radius of the capsid.
The third term in Eq. 1 represents the attraction between the wall and the chain only if the interaction is highly screened. Indeed, as noted in the introduction, the experiments of Hu et al. (30) revealed that linear polymers as long as 4900 monomers (1 MDa) and 16,500 monomers (3.4 MDa) were encapsidated in T = 2 and T = 3 structures, respectively. We note that the total number of positive charges on a T = 3 structure is ∼1800 while for a T = 2 one is ∼1200. Assuming that each monomer is charged and given the number of monomers per capsid, we conclude there must be a considerable overcharging happening. The fact that 1 MDa polymers choose T = 2 structures over T = 3 ones, and that 3.4 MDa polymers are enclosed in T = 3 structures, proves that the Coulomb interaction between CCMV capsid proteins and PSS must be highly screened, justifying our assumption for the form of the interaction.
Finally, the last term in Eq. 1 represents a reference free energy of n free chains in solution, where the quantity B is free energy of a single free chain and hence does not depend on R and D. We ignore this term because within a Flory theory it scales as (υa−3)2/5M1/5 and should therefore be subdominant. We also ignore the contribution of a surface tension between the adsorbed polymer layer and the core of the capsid devoid of polymer segments, which is reasonable if the density in the adsorption layer is not very high (40).
It is possible, at least in principle, to extract functional dependencies on, e.g., the ionic strength of the parameters υ, γ, a, and b from the various polyelectrolyte theories. A simple estimate of these quantities based on Debye-Hückel approximation is given in van der Schoot and Bruinsma (19). We postpone a detailed description of theses parameters to the end of the next section and first analyze the predictions of the model as it stands. Order of magnitude estimates are the excluded volume υ ≈ 1 nm3, the strength of the attractive interaction between a polymer segment and the capsid wall γ ≈ 1, the effective Kuhn length of the genome a ≈ 1 nm and the range of interaction b ≈ 1 nm, at least near physiological conditions (19).
If we minimize Eq. 1 with respect to the adsorption layer thickness D, we obtain
| (2) |
provided that a ≤ D ≤ R or γ ≥ c2υ nM/c3bR2. Estimates for the numerical constants are c1 ≈ π2 (41) from a ground-state calculation for an ideal chain in a spherical annulus, and 1/4π ≲ c2 ≲ 3/4π and 1/4π ≲ c3 ≲ 3/4π from simple geometry. Because of the approximate nature of the theory we shall not use these estimates here and calculate all the relevant quantities as a function of these parameters. If we insert Eq. 2 into Eq. 1, we get for the free energy of encapsulation
| (3) |
again provided D ≤ R. Interestingly, Eq. 3 suggests that the optimal thickness sets itself such that the overall free energy gain of encapsulation becomes exactly equal to the free energy loss due to elastic compression.
The question arises as to what the optimal number of chains n∗ absorbed by the capsid is in case it has a fixed radius R. If we optimize the free energy with respect to n for fixed M, we find
| (4) |
implying that the optimal number of segments n∗M encapsulated is an invariant of the molecular weight of the polymer. The reason for this behavior is the subdominant contribution from the reference state, the last term in Eq. 1, for long enough chains. It is straightforward to verify that the reference state increases the optimal number of encapsidated chains by a relative amount of ∼M−4/5. Interestingly, the recent measurements by Ren et al. on the encapsulation of poly(styrene sulfonic acid) by the coat protein of hibiscus chlorotic ringspot virus HCRV are consistent with our results (42). They found within the experimental error that the loading efficiency was constant for molecular weights of 13, 75, 200, and 990 kDa. The number of encapsidated polymers dropped from ∼100 to 1 on increasing the molecular weight over that range. It also agrees with the variation of the number of RNAs in the different CCMV particles discussed in the Introduction. Note that if n∗ is not an integer, for example if n∗ < 1 and the capsid size R is determined by the properties of the proteins in hand, we expect a suboptimal number of chains n∗ = 1 to be encapsulated.
If the protein subunits can form capsids of different sizes, in other words, if R could vary freely, then an interesting question is what the optimal size of the capsid is, given that a fixed amount of polymeric material nM is encapsulated. It is important to note that in this context encapsidation sets in if the chemical potential of the solution of free protein subunits and genome exceeds that of bound proteins and genome in a filled capsid. If the capsid consists of q protein subunits, to obtain the free energy per protein subunit associated with the protein-genome interaction , we assume that the area of each subunit is equal to πr02 ≡ 4πR2/q, and thus we find (30).
After optimizing fn, we obtain for the optimal radius
| (5) |
which is consistent with the previous result obtained by optimizing the number of encapsulated chains at fixed radius. According to Eq. 5, the optimal capsid radius increases with increasing genome length. As noted in the Introduction, the simulations of Zandi et al. (14) reveal that proteins in icosahedral structures have quite similar binding energies, at least if the proteins do not possess a strong intrinsic preference of a radius of curvature or T number. This implies that any additional size-determining mechanisms need only to provide a fairly weak dependence on the radius R to produce a nearly monodisperse solution of one T number in favor of the others.
We illustrate the importance of the genome length M on optimal capsid size in Fig. 2, where we plot the scaled free energy per capsid protein, fn/|f∗n|, as a function of R where f∗n is the free energy at the optimal radius. The degree of polymerization of the dashed curve is 1.6 times that of the solid curve. All the other physical parameters for both curves are the same and are chosen such that for the shorter chain the free energy goes through a minimum at ∼R = 17 nm while for the longer one this happens at ∼22 nm. The radius of a T = 4 virus is almost equal to RT=4 = 16 nm while that of a T = 7 structure is RT=7 = 21 nm. Although the interactions between the coat proteins in a T = 7 capsid have been predicted to be stronger than those in a T = 4 structure (14), the energy associated with the protein-genome interaction such as the one presented in Fig. 2 for the case of the shorter genome can easily bring the energy of a T = 4 structure significantly below that of a T = 7 structure. This would explain the effect of genome size in determining the size of a capsid, if coat proteins are flexible to form different T structures, such as CCMV. Note that if the strength of protein-genome interaction is very high, nonicosahedral capsids may in fact also form (14).
Figure 2.
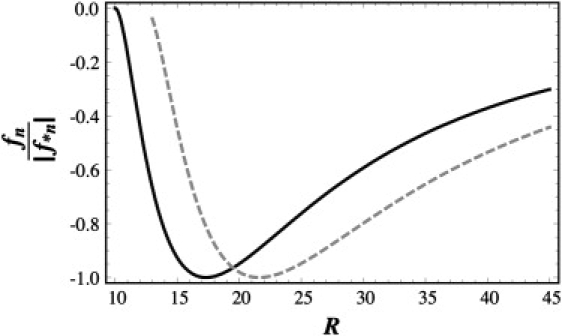
Free energy per protein subunit fn scaled to its minimum value |f∗n| versus the capsid radius R in nanometers. The values of the combination of parameters (nMν/γb) for the corresponding optimal radius, Eq. 5, were chosen such as to obtain the radii corresponding to typical T = 4 and T = 7 viruses. We set the molecular weight corresponding to the dashed curve for the T = 4 structure 1.6 times higher than that of the solid curve for the T = 7 structure, while keeping all the other parameters constant.
Randomly branched chain
Because of basepairing or duplexing, an RNA molecule has secondary structures and behaves differently from a linear chain. This implies that, e.g., the excluded volume υ and the interaction energy γ with the capsid wall is not only different from that of a linear chain but in fact should depend on the kind of RNA under consideration. As noted previously, there is no accurate theory available that even addresses the relation between the viral RNA size and shape on the one hand and the nucleotide length and/or sequence on the other. A complete theory of the more complicated issue of the adsorption of a single-stranded RNA to an oppositely charged wall with or without explicit model for the binding domain seems very remote indeed. Even if the important issue of electrostatics is ignored, the theoretical problem is quite formidable (34).
A crude way to account for the complex structure of the RNA is to treat it as a randomly branched chain. We need then only replace the first, elastic term in Eq. 1 by c4nMa4/D4 (37,40,43,44), with c4 a numerical constant. (See also the scaling argument in the Appendix.) If we follow the same procedure as we did for the linear chain, we find
| (6) |
which implies that
| (7) |
if we keep all system parameters for branched and linear chains of the same number of segments the same. (If c1 = 2c4 the approximate relation will be exact within the model.) Note that since D/a >> 1 the adsorbed layer thickness is much reduced if the same chain is branched instead of linear.
Cryo-TEM studies of RNA viruses, indeed, report that RNA is uniformly distributed inside the capsid (6), except for a region in the center devoid of RNA (2,3,5,46). Quite interestingly, the experiments of Hu et al. (30) show that if the RNA2 of CCMV is replaced by PSS of equal or larger number of monomers than the number of nucleotides in RNA2, the size of the polymer-free region in the middle of the capsid becomes smaller than that observed for capsids filled with RNA2, in agreement with our conclusion.
For the randomly branched chain model the optimal free energy becomes
| (8) |
which is different from what we obtained for linear chains, so branching has a large impact on the free energy of encapsulation. Remarkably, however, the optimal number of encapsulated chains n∗ given at a fixed capsid radius R turns out to be identical to what we found for the linear chain,
| (9) |
apart from a factor of 9/7 ≈ 1.3, implying that according to the model branching very slightly increases the optimal loading. This is of course only true if all physical parameters γ (the strength of the interaction), b (the range of the interaction), R (the radius of the capsid) and υ (the excluded volume interaction of the chains) are the same in both cases. Here, we again dropped the last term in the free energy Eq. 1 stemming from the reference state, because for branched chains B scales as (υa−3)2/5M1/2 giving a relative correction of ∼M−1/2 to the optimal number of encapsulated chains.
Note that we find the optimal capsid radius at fixed loading must be larger for a linear chain by a factor . This is a rather expected result because the radius of gyration of an branched polymer is smaller than that of an linear chain of equal degree of polymerization.
Even if the loading of the capsid is fixed for instance because the molecular weight M is large enough so that at most one chain finds itself absorbed in the capsid, the free energy difference between linear and randomly branched chains is quite large. In fact, for fixed nM we find
| (10) |
Because for linear chains D/a >> 1, we deduce that random branching enhances the encapsidation of a polymer. The reason is presumably that due to its topology a branched chain is inherently denser than a linear chain, leading to an enhanced interaction with the inner surface of the capsid. This could explain the observation mentioned in the Introduction that changing the gene order in a viral RNA can be enough to suppress encapsidation: a change in the sequence of nucleic acids can significantly modify the degree of branching, which in turn has as a strong impact on the free energy and therefore on the critical concentration of encapsulation that we will discuss in more detail in the following section. We note that within our simple Flory theory, Eq. 10 can be generalized straightforwardly (45) to any pair of differently branched polymers (43,44), and our conclusion is not restricted to the two extremes, i.e., linear and random branching, which we focus on in this work.
Model versus physical parameters
At the end of this section, we seek to connect the model parameters a (Kuhn length of the genome), γ (strength of the interaction between genome and coat protein), b (the range of this interaction), and υ (the excluded volume of segments of the genome) to the physical properties of the polyelectrolyte chains and the capsid proteins. These parameters are dominated by the effects of Coulomb interactions between the various species. Several studies have addressed the impact of electrostatics on the assembly of empty (17,20) and filled capsids (19) albeit at the level of the linearized Poisson-Boltzmann theory applied to idealized polyelectrolytes and idealized viral capsids. The issue remains quite contentious, however, because both RNA and viral capsids are in fact highly charged objects (47). Our goal is, here, to compare our Flory theory with the earlier, more detailed descriptions. Even if the linearized theory turns out inaccurate to describe viral assembly, the Flory theory should still hold.
Let α be the effective number of charges per monomer length of the polyelectrolyte chain. Within a Debye-Hückel approximation, we have for the excluded volume υ ≈ υ0 + 4πα2λBλD2 where under good-solvent conditions υ0 ≈ 4πa03/3 is the bare, hard-core excluded volume of a segment of Kuhn length a0 (48), and λB ≈ 0.7 nm the Bjerrum length and nm the Debye screening length if the (monovalent) salt concentration is cS M, and the solution at room temperature. If we model the inner capsid wall as a smooth surface with a uniform charge distribution, and let σ denote the number of positive charges per unit area, we have at the same level of approximation the range of interaction between wall and genome b ≈ λD and a strength of interaction γ ≈ 4πσαλBλD, if λD << R and the concentration of salt is not very low. (For a detailed discussion, see (19).) One key assumption here is that only segments that are within a Debye length of the wall actually interact with it.
Finally, the effective segment length a depends on the concentration of salt too. This remains a highly controversial and unresolved issue even for linear chains let alone branched ones (48). Often used is the expression by Odijk (49), and Skolnick and Fixman (50), a ≈ a0 + α2λBλD2/2a02 with a0 again the bare step length of the chain (half the bare persistence length), originally derived for semiflexible (persistent) chains (37). For highly charged polymers we expect α ≈ λB/a0 because of counterion condensation (48).
If we insert these estimates in Eq. 4 and absorb all numerical prefactors into a constant of proportionality c5, we get
| (11) |
with Q = 4πR2σ the total number of charges on the binding domains of the capsid. Two regimes emerge.
-
1.
If then Eq. 11 reduces to n∗M ≈ Q/α if we drop the constant of proportionality. This implies that the optimal number of polymers in a capsid does not depend on the salt concentration. This result is consistent with previous more elaborate calculations (2,19). The degree of branching potentially affects only the constant of proportionality, not the scaling with the number of charges Q on the capsid. A linear relationship between the M and Q has indeed been found for a whole range of viruses for which n∗ = 1 (2).
-
2.
If , Eq. 11 simplifies to n∗M ≈ QαλBλD2/υ0. To account for solvent quality we may put υ0 ≈ a03τ, where τ ≡ (T – Tθ)/Tθ is a relative temperature scale with Tθ the so-called Flory temperature (37,40). In this regime, the absorbed amount depends not only on the solvent quality but also on the concentration of salt through the Debye length λD. This implies that less material is encapsulated the higher the concentration of added salt. If we reduce the solvent quality, υ0 decreases and hence, according to Eq. 11, n∗M increases to the maximal value of Q/α.
We emphasize that even though n∗ depends only weakly on the chain architecture, the free energy of binding depends very strongly on it. Indeed, according to our studies, if given a choice, the capsid proteins would thermodynamically favor encapsulating a branched chain for any given degree of polymerization if all the other parameters, i.e., the quantities a, v, b, and γ in Eqs. 3 and 8, are the same.
Role of Stoichiometry
After analyzing the role of genome size in determining the size of viral capsid, we focus on how the stoichiometric ratio of the concentrations of protein and cargo molecules, i.e., real or ersatz genome, influences the size selection. For simplicity we shall restrict ourselves to dealing with the competition between structures corresponding to two neighboring T numbers. We assume that the solution is dilute and consists of free cargo molecules, single protein subunits, and fully formed viral particles with two different T numbers. Because partially formed capsids are stable only in extremely small amounts, we ignore them altogether (51,52). We also ignore empty capsids, as typically these do not tend to form under near-neutral pH conditions. Hence, each of the viruslike particles is presumed to contain one or more negatively charged cargo molecules that we do not describe in detail. We also do not specify the kind of protein building blocks: depending on the species of virus they may be monomers, dimers or even pentamers or hexamers of the actual coat proteins (6).
Mass action equations
Within a mean-field approximation, the Helmholtz free energy of our model system can be written as (53)
| (12) |
where Nα denotes the number and ρα = Nα/V the number density of the various species in the volume V of the system. Here, the subscript g refers to the free cargo molecules, subscript p refers to the free protein subunits, and α = q1, q2 refers to the two different T structures in the solution. Without loss of generality, we presume that q1 is always smaller than q2. Therefore, the index 1 represents the smaller of the two T numbers considered. The quantity ω is an interaction volume that we presume to be approximately the size of the solvent molecules. This makes the product of the number density and interaction volume to a good approximation equal to a mole fraction, xα ≡ ραω. Notice that since the solution is dilute, we assume that xα << 1 for all species.
The quantity is the effective binding free energy of a single subunit part of a capsid of species i = 1, 2, which includes the contributions of the protein-protein interactions, , and the genome-protein interactions, . The binding free energy is an averaged quantity over all qi subunits of a fully formed capsid, where we recall that in icosahedral capsids the coat proteins do not have identical but quasiequivalent local environments (6). Depending on the genome size and overall charge, the solution conditions such as pH and ionic strength and the type of virus coat protein, can be smaller or bigger than
The equilibrium distribution of proteins over the assembled and disassembled states follows by minimization of the free energy subject to the conservation of the mass of protein subunits (53). Let the overall mole fraction of protein subunits be Xp. We then have to demand that . If Xg is the overall genome concentration in mole fraction units, and n1 and n2 are the number of genome molecules in the i = 1 and 2 structures, respectively, we have . To keep our presentation simple, we assume that n1 and n2 are given quantities. They can represent either the optimal number of encapsulated cargo molecules in each type of capsid as calculated in the previous section, or kinetically determined ones.
Setting and , we obtain
| (13) |
| (14) |
where we for convenience have expressed number densities in terms of mole fractions. Equations 13 and 14 establish the relation between the equilibrium concentrations of the viruses with two different T numbers. The phase diagram implicit in these two coupled mass action equations is quite complex and depends on many variables indeed. We shall be focusing below on those regimes relevant to the conditions of recent experiments of Sikkema et al. (33) and Ren et al. (42), and also the experiments of Bancroft (7) and Verduin and Bancroft (12) that were performed more than half a century ago (7,12).
To evaluate the effective binding free energy of a single protein subunit, , for the capsids with the two T numbers corresponding to i = 1, 2, it is necessary to obtain the protein-protein interaction free energies and the cargo-protein interaction free energy . The latter can be estimated using the prescription of the previous section, although there is of course the issue of the unknown numerical constants. Ultimately, the unknown parameters could be obtained by a systematic comparison of in vitro assembly studies and the theories similar to the one presented in this article. As for the protein-protein interaction energies, Monte Carlo simulations of coarse-grained models do provide some insight in the energy gain of coat protein in icosahedral and nonicosahedral structures (14,22).
In what follows, we first analyze some relevant consequences of the coupled mass action equations, and next discuss the separate cases of high and low molecular weight polymers (12,30,33).
Mass action, assembly, and size competition
Mass action acts on the distribution of protein molecules over the two species as well as on the distribution over assembled and disassembled states of the proteins. Let us deal with the latter problem first. For this purpose it is useful to suppress one of the capsid sizes, say species 2, by setting the effective binding free energy of a single protein subunit in the species 2 . For simplicity we also fix n1 = 1, so only one cargo molecule is encapsulated by the coat protein. The highly nonlinear mass action equation that results can be solved explicitly for different ranges of the stoichiometric ratio r ≡ q1Xg/Xp (with Xg the overall genome and Xp the overall protein concentrations). It turns out practical to introduce two new variables, being the fraction of proteins in capsids, , and the fraction of bound genome molecules, . Three regimes emerge:
-
1.If the genome concentration is relatively low and r << 1, we find that , and
with a critical concentration of protein molecules. Because the total number of protein subunits in the species 1, q1 >> 1, the transition from the unassembled to assembled state is very sharp indeed, and may be approximated by η = 0 and ς = 0 for Xp < X1,∗, and η = r and ς = 1 for Xp > X1,∗. So, beyond the critical protein concentration X1,∗ almost all of the genome is encapsulated.(15) -
2.Under conditions of perfect stoichiometry of r = 1, we obtain
for Xp > X1,∗ and η = ς ∼ 0 for Xp < X1,∗. Again, we have used the fact that q1 >> 1 to find a sharp transition between assembled and disassembled states, albeit that now the fraction of genome encapsulated rises much more slowly with increasing protein concentration. Exactly the same behavior is found in models for the self-assembly of empty capsids (17,49,50).(16) -
3.In the presence of large quantities of genome, so r >> 1, the critical concentration is renormalized by the presence of the genome. In the limit q1 >> 1, we have
if . For , we have η = rς ∼ 0. Because q1 >> 1, the renormalized critical concentration depends only very weakly on the concentration of genome. Notice that because of the large stoichiometric ratio and ς = η/r < 1/r << 1, only a small fraction of genome finds itself encapsulated.(17)
To investigate whether or not mass action can change the preference from one T number to another, let us no longer presume that the effective binding free energy of a single protein subunit in the species 2 is but, instead, demand that , allowing for a more manageable analysis. If we in addition for simplicity presume that n1 = n2, we deduce from Eqs. 13 and 14 that the mole fraction of species 1 to 2 is . Obviously, the smaller capsids should be the dominant species if the density of free protein in solution obeys , and the larger ones predominate if . The situation turns out a little bit more complex because the concentration of genome modifies this initial conclusion.
Let us focus on the case n1 = n2 = 1. The fraction of protein in capsids reads while the relative prevalence of proteins in species 2 to species 1 capsids is indicated by the variable . From the ratio of Eqs. 14 and 13 we then find that
| (18) |
while Eq. 13 reduces to
| (19) |
where, as before, r = q1Xg/Xp and
The last expression can be simplified considerably. According to Eq. 18, there is an appreciable number of capsids in the solution only if Xp >> X∗. In that case, η → 1, which allows the reduction of Eq. 19 to
| (20) |
in the limit r >> 1.
We conclude that the relative concentration of species 2 capsids grows with increasing ratio of the overall protein to genome concentrations, Xp/Xg, and as a result, the smaller the quantity Xg, the more strongly the larger species is preferred, and vice versa. This is in agreement with experimental observation (33). Even if , this effect survives, as we shall show next. We shall also see that this conclusion is not restricted to the case n1 = n2 = 1.
Long versus short genome
As already mentioned, in their experiments, Bancroft (7) and Bancroft et al. (9) mixed coat proteins of CCMV with TMV RNA, which consists of ∼6000 nucleotides. Because TMV RNA is more than twice as long as the RNAs 1 and 2 of CCMV, one expects that considerably larger capsids form. However, the experiments also show that in addition to genome size, the ratio of coat protein to genome concentration plays an important role in determining the size of the capsids. The size of a CCMV capsid that encapsulates the large TMV RNA can change from a T = 4 or T = 7 structure to a T = 3 structure (the native structure of the coat protein) if the ratio of protein to genome concentration is sufficiently small.
To investigate in more detail the impact of the stoichiometric ratio of the protein and genome concentrations on the capsid size and structure, we need to carefully examine the mass action equations, Eqs. 13 and 14, for the two competing sizes 1 and 2. Because the TMV genome is ∼6000 nucleotides long, we plausibly presume that only one genome molecule is encapsidated by each capsid type, and set n1 = n2 = 1. (We relax this assumption below.) This scenario would also apply to the experiments of Hu et al. on the encapsulation of high-molecular weight poly(styrene sulfonate) by CCMV coat proteins (30).
Now, if the genome concentration increases at a fixed protein concentration, more capsids potentially form and hence more genome can be encapsulated if the capsids become smaller. Arguably, the loss in the free energy of binding should then be compensated for by an increase in translational entropy. Equations 13 and 14 support this scenario as is shown in Fig. 3. To produce this figure, we set the values of the aggregation numbers q1 and q2 equal to the number of capsomers in the T = 4 and T = 7 capsids, and also assumed that the genome-coat protein interaction prefers a T = 7 structure over that of T = 4. Therefore, we set the binding free energy per single protein subunit in species 2, , slightly more negative than the one in species 1, . Because the quantities and have not been assessed exactly in any experiments, and the energy landscape is too large to be completely explored numerically, the values of and used for the figures are only chosen to show the proof of principle. The figure illustrates that for a given choice of parameters and for a fixed protein concentration, the dominant structure changes from a T = 4 to a T = 7 structure if the concentration of cargo molecules is decreased, consistent with the TMV RNA encapsidation experiments by CCMV coat proteins (9,12).
Figure 3.
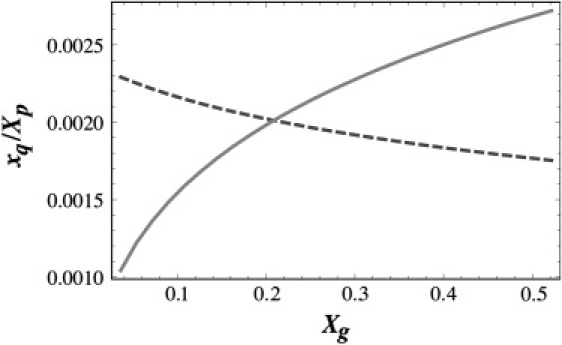
Plotted is the scaled fraction of capsids xq/Xp versus the mole fraction of genome, Xg. The concentration of T = 4 structures (solid curve) increases while that of the T = 7 ones (dashed curve) decreases as the concentration of genome, Xg, increases. To calculate the curves, we set q1 = 42, q2 = 72, ɛq1 = −3 kBT, and ɛq2 = −3.075 kBT. The mole fraction of capsid proteins was fixed at Xp = 0.054.
According to the figure, for a fixed concentration of protein, T = 7 structures prevail at low RNA concentrations as in fact is expected from the analysis of the preceding subsection. However, as the concentration of genome increases, the T = 4 structures become the increasingly dominant species. So, even if for a specific genome length T = 7 structures are energetically more favorable than the structures of T = 4, the number of the free coat proteins in solution can be so low that T = 4 structures become entropically more favorable compared to T = 7 structures. The mass action equations also show that in the opposite case where the length of the genome prefers a T = 4 structure, T = 7 structures will nevertheless form if the genome concentration is low and the protein concentration is high. These results follow the experimental observations referred to above (12).
Next, we investigate the role of stoichiometric conditions on capsid size relevant to more recent experiments on mixtures of the coat protein of CCMV and of low-molecular weight PSS (33). Similar to the experiments on the encapsulation of large RNAs, a change in capsid size was observed with varying stoichiometric ratio of the polyelectrolyte and coat protein concentrations. At small stoichiometric ratios of 0.4 or 4, two types of particles form that were attributed to T = 1 and T = 3 icosahedral structures. However, upon increasing the stoichiometric ratio to 40 and 400, only T = 1 particles form. Higher stoichiometric ratios than 400 inhibit particle formation altogether, an observation attributed to polyelectrolyte effects (33).
In the experiments of Sikkema et al. (33), the length of the chain is so small that more than one chain can be encapsidated in each capsid, so the result of previous section on the correlation between capsid size and genome length cannot be applied here. Under physiological conditions, the Coulomb interactions are largely (but not completely) screened, so we expect from the theory of the previous section that the optimal number of polymers in each capsid is related to the number of charges on the inner capsid wall. We expect the number of positive charges on amino acid residues in the capsid inner surface to be proportional to the number of negative charges on the segments of encapsidated PSS. (Note that Belyi and Muthukumar (2) and van der Schoot and Bruinsma (19) predict complete or almost complete charge inversion to occur, which would explain why CCMV particles are negatively charged (8).) Hence, we consider only the case in which the number of anionic polymers in a T = 3 capsid is larger than in a T = 1 capsid, i.e., n1 < n2.
Because the native CCMV capsid is a T = 3 structure, it is reasonable to assume that the energy per protein subunit of a T = 3 particle is lower than that of a T = 1 particle. However, Fig. 4 clearly shows that for a fixed coat protein concentration, the molar concentration of T = 1 particles increases much faster than that of T = 3 particles with increasing polyelectrolyte concentration. Once again we observe the effect of entropy on capsid assembly: the structures with smaller T numbers prevail while larger T number structures are energetically more favorable. A direct comparison of the results presented in this article and the relevant experiments will provide valuable information about the viral protein binding energies.
Figure 4.
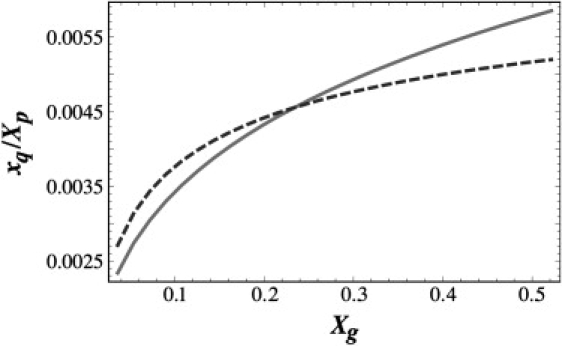
The scaled fraction of capsids versus the mole fraction of genome. The concentration of T = 1 structures (solid line) increases faster than that of T = 3 (dashed curve) ones as the concentration of RNA increases. For the T = 1 structures, q1 = 12 and ɛq1 = −2 kBT. For the T = 3 structures, q2 = 32 and ɛq2 = −2.37 kBT. The number of encapsulated chains for the two T numbers was set to g1 = 1 and g2 = 2. The protein concentration was kept constant at Xp = 0.05.
Conclusion
In conclusion, we explored the physical principles underlying the remarkable fact that CCMV protein subunits assemble into icosahedral structures with different sizes. To obtain insight in the contribution of the genome length to the free energy of formation of a virus, we used a simple Flory model that captures the essential ingredients of genome-capsid interaction. These ingredients are the electrostatic attraction between the polyelectrolyte and the surface, and the increased self-repulsion and the elastic entropy loss of the encapsulated genome upon adsorption onto the capsid wall. Although simple, our model rationalizes many observations on the size selection of viruses both in terms of the length of genome and its structure and can be experimentally tested. We note that the focus of this article is on the aspects of virus assembly that are common among many viruses. Viruses in which particular interactions such as sequence-specific interactions play important role in their assembly are subject to another study.
We show that there is a correlation between the optimal radius of a capsid and the length of its encapsidated genome. In principle, longer genomes require larger capsids if the coat proteins do not exhibit a preference for a particular radius of curvature. Long genomes can be encapsulated by a small capsid, if the radius of gyration of genome (which, in turn, depends on the RNA secondary and tertiary structures and its charge density), allows for it. This might explain why the RNA of turnip yellow mosaic virus TYMV, although ∼6000 nucleotides long, and that of CCMV at ∼3000 nucleotides, are both encapsulated in a shell of ∼30 nm. Experiments also reveal that the ratio of protein/genome concentration can modify this conclusion (7,33). This is due to entropy effects not yet considered in the literature.
While the level of branching of the genome does not seem to have a large effect on the optimal number of monomers encapsulated if the capsid size is largely fixed by the properties of the coat proteins, the level of branching of genome does seem to have a significant impact on the free energy of encapsulation and hence on the critical encapsulation concentration. This explains why a mere change of gene order has such a considerable effect on the encapsidation of RNA: it shifts the critical concentration as it depends exponentially on this free energy.
We have shown that while electrostatic interactions drive viral self-assembly around the genome, other factors such as the conformations of the genome and/or the entropic factor associated with mass action must be at least as important in determining the final size of the capsid. Hence, a description that only involves the length of the genome and/or the number of charges on the RNA binding domain cannot explain many viral experiments (2,19).
A quantitative comparison between experiments and the theory presented in this article will result in a better knowledge of the protein-protein and the protein-genome interactions. These quantities could be obtained by a set of systematic in vitro studies of virus assembly, along the lines of the work of Ceres and Zlotnick on hepatitis B virus capsids (54). A comprehensive investigation of the physico-chemical parameters that impact capsid formation could have great potential in the development of antiviral therapies and a systematic treatment of viral infection.
Uncited references
Appendix
A simple scaling estimate of the free energy cost FN(R, D) of putting an ideal chain of arbitrary connectivity in a spherical annulus of radius R and width D is easily derived. Starting point is the free energy cost of putting a chain of N segments and fractal dimension d in a spherical confinement much smaller than its unperturbed radius of gyration. The number of segments between two collisions must be equal to g ≈ (R/a)d, so the free energy of confinement scales as the number of collisions times the thermal energy, FN(R, 0) ≈ kBT N/g ≈ kBTNad/Rd. The number blobs Ng of size D3 in the annulus is given by Ng ≈ R2D/D3 = R2/D2 and the number of segments g in each blob is given by N/Ng. Hence, the free energy of confinement must be equal to the number of blobs times the free energy of confinement of each blob, FN(R,D) ≈ Ng(D,0) ≈ kBT Nad/Dd. For linear chains d = 2, and for randomly branched ones, d = 4. For d = 2 we retrieve the scaling relation obtained in the explicit ground-state calculation of Yaman et al. (41).
Acknowledgments
The authors acknowledge helpful discussions with Chuck Knobler, William M. Gelbart, Robijn Bruinsma, and Jeroen Cornelissen. We are grateful to Marloes van Beek and Aviva Shackell for critically reading the manuscript.
R.Z. acknowledges support by the National Science Foundation through grant No. DMR-06-45668.
References
- 1.Flint S.J., Enquist L.W., Krug R.M., Racaniello V.R., Skalka A.M. ASM Press; Washington, DC: 2000. Principles of Virology: Molecular Biology, Pathogenesis, and Control. [Google Scholar]
- 2.Belyi V.A., Muthukumar M. Electrostatic origin of the genome packing in viruses. Proc. Natl. Acad. Sci. USA. 2006;103:17174–17178. doi: 10.1073/pnas.0608311103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fox J.M., Wang G., Speir J.A., Olson N.H., Johnson J.E. Comparison of the native CCMV virion with in vitro assembled CCMV virions by cryoelectron microscopy and image reconstruction. Virology. 1998;244:212–218. doi: 10.1006/viro.1998.9107. [DOI] [PubMed] [Google Scholar]
- 4.Choi Y.G., Rao A.L.N. Packaging of brome mosaic virus RNA3 is mediated through a bipartite signal. J. Virol. 2003;77:9750–9757. doi: 10.1128/JVI.77.18.9750-9757.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rao A.L.N. Genome packaging by spherical plant viruses. Annu. Rev. Phytopathol. 2006;44:61–87. doi: 10.1146/annurev.phyto.44.070505.143334. [DOI] [PubMed] [Google Scholar]
- 6.Chiu W., Burnett R.M., Garcea R.L. Oxford University Press; Oxford: 1997. Structural Biology of Viruses. [Google Scholar]
- 7.Bancroft J.B. The self-assembly of spherical plant viruses. Adv. Virus Res. 1970;16:99–134. doi: 10.1016/s0065-3527(08)60022-6. [DOI] [PubMed] [Google Scholar]
- 8.Bancroft J.B., Hiebert E., Rees M.W., Markham R. Properties of cowpea chlorotic mottle virus, its protein and nucleic acid. Virology. 1968;34:224–239. doi: 10.1016/0042-6822(68)90232-8. [DOI] [PubMed] [Google Scholar]
- 9.Bancroft J.B., Hiebert E., Bracker C.E. The effects of various polyanions on shell formation of some spherical viruses. Virology. 1969;39:924–930. doi: 10.1016/0042-6822(69)90029-4. [DOI] [PubMed] [Google Scholar]
- 10.Hiebert E., Bancroft J.B., Bracker C.E. The assembly in vitro of some small spherical viruses, hybrid viruses and other nucleoproteins. Virology. 1968;34:492–508. doi: 10.1016/0042-6822(68)90069-x. [DOI] [PubMed] [Google Scholar]
- 11.Reference deleted in proof.
- 12.Verduin B.J.M., Bancroft J.B. The infectivity of tobacco mosaic virus RNA in coat protein from spherical viruses. Virology. 1969;37:501–506. doi: 10.1016/0042-6822(69)90240-2. [DOI] [PubMed] [Google Scholar]
- 13.Bruinsma R.F., Gelbart W.M., Reguera D., Rudnick J., Zandi R. Viral self-assembly as a thermodynamic process. Phys. Rev. Lett. 2003;90:248101–248104. doi: 10.1103/PhysRevLett.90.248101. [DOI] [PubMed] [Google Scholar]
- 14.Zandi R., Reguera D., Bruinsma R.F., Gelbart W.M., Rudnick J. Origin of icosahedral symmetry in viruses. Proc. Natl. Acad. Sci. USA. 2004;101:15556–15560. doi: 10.1073/pnas.0405844101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hyeon C., Dima R.I., Thirumalai D. Size, shape and flexibility of RNA structures. J. Chem. Phys. 2006;125:1–10. doi: 10.1063/1.2364190. [DOI] [PubMed] [Google Scholar]
- 16.Lidmar J., Mirny L., Nelson D.R. Virus shapes and buckling transitions in spherical shells. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2003;68:051910. doi: 10.1103/PhysRevE.68.051910. [DOI] [PubMed] [Google Scholar]
- 17.Kegel W.K., van der Schoot P. Competing hydrophobic and screened-Coulomb interactions in hepatitis B virus capsid assembly. Biophys. J. 2004;86:3905–3913. doi: 10.1529/biophysj.104.040055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hagan M.F., Chandler D. Dynamic pathways for viral capsid assembly. Biophys. J. 2006;91:42–54. doi: 10.1529/biophysj.105.076851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.van der Schoot P., Bruinsma R. Electrostatics of an RNA virus. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2005;70:1–12. doi: 10.1103/PhysRevE.71.061928. [DOI] [PubMed] [Google Scholar]
- 20.Šiber A., Podgornik R. Role of electrostatic interactions in the assembly of empty spherical viral particles. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2007;76:061906. doi: 10.1103/PhysRevE.76.061906. [DOI] [PubMed] [Google Scholar]
- 21.Caspar D.L.D., Klug A. Physical principles in the construction of regular viruses. Quant. Biol. 1962;27:1–24. doi: 10.1101/sqb.1962.027.001.005. [DOI] [PubMed] [Google Scholar]
- 22.Zandi R., Reguera D. Mechanical properties of viral capsids. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2005;72:1–12. doi: 10.1103/PhysRevE.72.021917. [DOI] [PubMed] [Google Scholar]
- 23.Ysebaert M., van Emmelo J., Fiers W. Total nucleotide sequence of a nearly full-size DNA copy of satellite tobacco necrosis virus RNA. J. Mol. Biol. 1980;143:273–287. doi: 10.1016/0022-2836(80)90190-4. [DOI] [PubMed] [Google Scholar]
- 24.Higgs P.G. RNA secondary structure: a comparison of real and random sequences. J. Phys. I Fr. 1993;3:43–59. [Google Scholar]
- 25.Muroga Y., Sano Y., Tagawa H., Shimizu S. Studies on the conformation of a polyelectrolyte in solution: local conformation of Cucumber Green Mottle Mosaic Virus RNA compared with Tobacco Mosaic Virus RNA. J. Phys. Chem. B. 2007;111:8619–8625. doi: 10.1021/jp068944j. [DOI] [PubMed] [Google Scholar]
- 26.Reference deleted in proof.
- 27.Gutin A., Grosberg A., Shakhnovich E. Conformational entropy of a branched polymer. Macromolecules. 1995;28:3718–3727. [Google Scholar]
- 28.Yoffe A., Gelbart W.M., Ben-Shaul A. Secondary structure statistics of random vs. viral RNA. Biophys. J. 2005;88:573A. [Google Scholar]
- 29.Sun J., DuFord C., Daniel M.-C., Murali A., Chen C. Core-controlled polymorphism in virus-like particles. Proc. Natl. Acad. Sci. USA. 2007;104:1354–1359. doi: 10.1073/pnas.0610542104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hu Y., Zandi R., Anavitarte A., Knobler C.M., Gelbart W.M. Packaging of a polymer by a viral capsid: the interplay between polymer length and capsid size. Biophys. J. 2008;94:1428–1436. doi: 10.1529/biophysj.107.117473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Chang C.B., Knobler C.M., Gelbart W.M., Mason T.G. Viral protein structures on encapsidated nanoemulsion droplets. ACS Nano. 2008;2:281–286. doi: 10.1021/nn700385z. [DOI] [PubMed] [Google Scholar]
- 32.Reference deleted in proof.
- 33.Sikkema F.D., Cornellas-Aragnones M., Fokkink R.G., Verduin B.J.M., Cornelissen J.J.L.M. Monodisperse polymer-virus hybrid nanoparticles. Org. Biomol. Chem. 2007;5:54–57. doi: 10.1039/b613890j. [DOI] [PubMed] [Google Scholar]
- 34.Nguyen T.T., Bruinsma R.F. RNA condensation and the wetting transition. Phys. Rev. Lett. 2006;87:108102. doi: 10.1103/PhysRevLett.97.108102. [DOI] [PubMed] [Google Scholar]
- 35.Brochard-Wyart F., Tanaka T., Borghi N., de Gennes P.-G. Semiflexible polymers confined in soft tubes. Langmuir. 2005;21:4144–4148. doi: 10.1021/la0474114. [DOI] [PubMed] [Google Scholar]
- 36.Zlotnick A. Distinguishing reversible from irreversible virus capsid assembly. J. Mol. Biol. 2007;366:14–18. doi: 10.1016/j.jmb.2006.11.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Rubinstein M., Coby R.H. Oxford University Press; Oxford, UK: 2003. Polymer Physics. [Google Scholar]
- 38.Gutin A.M., Grosberg A.Yu., Shakhnovitch E.I. Polymers with annealed and quenched branchings belong to different universality classes. Macromolecules. 1993;26:1293–1295. [Google Scholar]
- 39.Kuznetsov Y.G., Daijogo S., Zhou J., Semler B.L., McPherson A. Atomic force microscopy analysis of icosahedral virus RNA. J. Mol. Biol. 2005;347:41–52. doi: 10.1016/j.jmb.2005.01.006. [DOI] [PubMed] [Google Scholar]
- 40.Grosberg A.Yu., Khokhlov A.R. The American Institute of Physics; New York: 1994. Statistical Physics of Macromolecules. [Google Scholar]
- 41.Yaman K., Pincus P., Solis F., Witten T.A. Polymers in curved boxes. Macromolecules. 1997;30:1173–1178. [Google Scholar]
- 42.Ren Y., Wong S.-M., Lim L.-Y. In vitro reassembled plant virus-like particles for loading of polyacids. J. Gen. Virol. 2006;87:2749–2754. doi: 10.1099/vir.0.81944-0. [DOI] [PubMed] [Google Scholar]
- 43.Vilgis T.A. Polymer theory: path integrals and scaling. Phys. Rep. 2000;336:167–254. [Google Scholar]
- 44.Sakaue T., Rafael E. Polymer chains in confined spaces and flow-injection problems: some remarks. Macromolecules. 2006;39:2621–2628. [Google Scholar]
- 45.Reference deleted in proof.
- 46.Zlotnick A., Cheng N., Stahl S.J., Conway J.F., Steven A.C. Localization of the C-terminus of the assembly domain of hepatitis B virus capsid protein: implications for morphogenesis and organization of encapsidated RNA. Proc. Natl. Acad. Sci. USA. 1997;94:9556–9561. doi: 10.1073/pnas.94.18.9556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hu, T., R. Zhang, and B.I. Shklovskii. Electrostatic theory of viral self-assembly: a toy model. arXiv:q-bio/0610009.
- 48.Netz R., Andelman D. Neutral and charged polymers at interfaces. Phys. Rep. 2003;380:1–95. [Google Scholar]
- 49.Odijk T. Polyelectrolytes near the rod limit. J. Polym. Sci. 1977;15:477. [Google Scholar]
- 50.Skolnick J., Fixman M. Electrostatic persistence length of a wormlike polyelectrolyte. Macromolecules. 1977;10:944. [Google Scholar]
- 51.Zlotnick A. To build a virus capsid. An equilibrium model of the self-assembly of polyhedral protein complexes. J. Mol. Biol. 1994;241:59–67. doi: 10.1006/jmbi.1994.1473. [DOI] [PubMed] [Google Scholar]
- 52.Zandi R., van der Schoot P., Reguera D., Kegel W., Reiss H. Classical nucleation theory of virus capsid assembly. Biophys. J. 2006;90:1939–1948. doi: 10.1529/biophysj.105.072975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.van der Schoot P. Theory of supramolecular polymerization. In: Ciferri A., editor. Supramolecular Polymers. 2nd Ed. CRC Press; Boca Raton, FL: 2005. [Google Scholar]
- 54.Ceres P., Zlotnick A. Hepatitis B virus capsid assembly is driven by weak intersubunit contacts. Biochemistry. 2002;41:11525–11531. doi: 10.1021/bi0261645. [DOI] [PubMed] [Google Scholar]


