Abstract
Human CD36 is a class B scavenger receptor expressed in a variety of cell types such as macrophage and adipocytes. This plasma membrane glycoprotein has a wide range of ligands including oxidized low density lipoprotein and long chain fatty acids which involves the receptor in diseases such as atherosclerosis and insulin resistance. CD36 is heavily modified post-translationally by N-linked glycosylation, and 10 putative glycosylation sites situated in the large extracellular loop of the protein have been identified; however, their utilization and role in the folding and function of the protein have not been characterized. Using mass spectrometry on purified and peptide N-glycosidase F-deglycosylated CD36 and also by comparing the electrophoretic mobility of different glycosylation site mutants, we have determined that 9 of the 10 sites can be modified by glycosylation. Flow cytometric analysis of the different glycosylation mutants expressed in mammalian cells established that glycosylation is necessary for trafficking to the plasma membrane. Minimally glycosylated mutants that supported trafficking were identified and indicated the importance of carboxyl-terminal sites Asn-247, Asn-321, and Asn-417. However, unlike SRBI, no individual site was found to be essential for proper trafficking of CD36. Surprisingly, these minimally glycosylated mutants appear to be predominantly core-glycosylated, indicating that mature glycosylation is not necessary for surface expression in mammalian cells. The data also show that neither the nature nor the pattern of glycosylation is relevant to binding of modified low density lipoprotein.
Human CD36, originally identified in platelets as glycoprotein IV (1), is a class B scavenger receptor localized to the plasma membrane. It is not expressed ubiquitously but is present in a variety of different cells and tissue types including epithelial cells (2), macrophages (3), endothelial cells of the microvasculature (4), and smooth muscle (5). Its function is complex, and its involvement in different disease scenarios, such as cancer (6), atherosclerosis (3, 7, 8), malaria (9), and insulin resistance (10), most likely reflects the interaction of the receptor with a particular ligand in a specific cell type. For example, CD36 expressed in monocytic macrophages functions as a scavenger receptor for the uptake of oxidized LDL2 (3, 11). Under certain physiological conditions, this results in the lipid loading of macrophages at the site of tissue damage in the arterial wall, leading to foam cell formation and plaque development, a key early stage in the pathogenesis of atherosclerosis (8, 12). In fat and muscle cells, CD36 plays an essential role in lipid homeostasis by uptake of long chain fatty acids (13). In this case CD36 deficiency has been linked to disorders in lipid metabolism, giving rise to increased incidences of insulin resistance and cardiomyopathies (11, 14, 15).
Although much is known about the function of CD36, less is known about its structure. CD36 has no bacterial homologues but is a member of a protein family that also includes the mammalian proteins LIMPII (16), CLA-1 (17), SRBI (18), and the Drosophila proteins Croquemort (19) and emp (20). The sequence of 471 amino acids has two short hydrophobic regions at the carboxyl and amino termini separated by a large hydrophilic region (21); however, the topology of the protein is unclear with both ditopic (22) and type I (23) topological models proposed. Both are consistent in predicting that the large hydrophilic region is extracellular, which is clearly supported by epitope mapping studies (24). The protein is heavily modified post-translationally. The six extracellular cysteines, which are highly conserved within the orthologous CD36 subfamily, have been shown to be linked by disulfide bonds in bovine Cd36 (25), and the remaining four cysteines, two at each terminus, are palmitoylated (26), lending credence to the ditopic topological model.
CD36 is also modified by N-linked glycosylation, which accounts for the observation that the protein migrates with an apparent molecular mass of 78–94 kDa on SDS-PAGE (4, 27) despite a theoretical mass for the polypeptide of 53 kDa. N-Linked glycosylation is a common modification of extracellular and secreted proteins, and defects in the glycosylation pathways lead to a wide range of serious diseases known collectively as congenital disorders of glycosylation (28). Glycosylation can be important for correct folding of proteins (29, 30) either by directly inducing and/or stabilizing the tertiary fold of the polypeptide (31) or via an affinity for lectin chaperones such as calnexin or calreticulin (32). Glycosylation has also been shown to be important for the trafficking of certain glycoproteins through affinity for lectin transport machinery (33). The glycosylation status of bovine Cd36 has already been determined with all eight putative sites shown to be glycosylated (34). Human and bovine CD36 are 83% identical (93% when similar residues are included) and share 7 glycosylation sites (human has 10 putative glycosylation sites). In the related mouse SRBI, which is 33% identical (54% similar) to human CD36, there are 11 putative N-linked glycosylation sites, only 3 of which are shared with the human protein. Site-directed mutagenesis of each of the 11 sites independently in SRBI in an otherwise wild type protein indicates that all are glycosylated, with two (Asn-108 and Asn-173) important for either trafficking or folding. Mutagenesis of either of these two residues resulted in very little cell surface expression of the protein (35); however, neither site is conserved in human CD36.
To gain further understanding of the role of glycosylation of CD36, we used mutagenesis and biophysical analysis (mass spectrometry and gel electrophoresis) to identify unequivocally which glycosylation sites are occupied in human CD36. Antibody and ligand binding studies with these mutant proteins also provided insights into the role of glycosylation and site occupancy in the trafficking and function of the protein.
EXPERIMENTAL PROCEDURES
Materials
The detergent n-octyl-β-d-glucopyranoside (OG) was purchased from Merck. The nickel-NTA-agarose was purchased from Qiagen Ltd, UK, and BODIPY Acetylated LDL FL® (BODIPY Ac-LDL) was from Invitrogen. All the protease inhibitors, fatty acid-free BSA, and tunicamycin were purchased from Sigma-Aldrich. Amicon Ultra 15 (molecular weight cutoff 50) centrifugal devices were from Millipore, and peptide N-glycosidase F (PNGase F) and endoglycosidase H (Endo H) were from New England Biolabs. The primary antibodies, mouse mAb1258 (Chemicon International), and rat mAb1955 (R and D Systems), both recognize folded CD36 for use in flow cytometry, but only mAb1955 recognizes the denatured product after SDS-PAGE and Western analysis.
Generation of Expression Vectors Encoding CD36 with a 12-Histidine, Carboxyl-terminal Tag
NcoI and BstEII restriction sites were introduced into the 5′ and 3′ ends, respectively, of the CD36 cDNA (ATCC clone MGC-14530) by mutagenic polymerase chain reaction using oligonucleotides 5′-TTG GTA CAT ACG GTG ACC TTT TAT TGT TTC G-3′ (NcoI) and 5′-CCT GAA CAA GAA CCA TGG GCT GTG ACC-3′ (BstEII). These two restriction enzyme sites were used to subclone CD36 into the baculoviral transfer vector BlueBac4.5 (Invitrogen) containing a 12-histidine tag (36) to generate BlueBac-CD36–12His. The transfer vector was used to engineer a recombinant baculovirus using the Bac-N-Blue system (Invitrogen) as directed. For expression in mammalian cells, the modified CD36 cDNA was subcloned, replacing the rodent sequence in pCI-Cd36–12His (36) using the restriction enzymes NcoI and BstEII to generate pCI-CD36–12His.
Site-directed Mutagenesis of CD36
Site-directed mutagenesis was performed on pCI-CD36–12His using the QuikChange multisite mutagenesis kit (Stratagene) according to the manufacturer's instructions. Primers were designed to replace the asparagine codon in each putative glycosylation site with a codon for glutamine. The entire coding sequence of each mutant was confirmed by DNA sequencing. The sequences of mutagenic primers for the asparagine to glutamine mutants were: N79Q, 5′-CAC AGG AAG TGA TGA TGC AGA GCT CCA ACA TTC AAG TTA AGC-3′; N102Q, 5′-TCG TTT TCT AGC CAA AGA ACA GGT AAC CCA GGA CGC TG-3′; N134Q, 5′-TGG AAC AGA GGC TGA TCA GTT CAC AGT TCT CAA TC-3′; N163Q, 5′-GTT CAA ATG ATC CTC AAT TCA TTA ATT CAG AAG TCA AAA TCT TCT ATG TTC C-3′; N205Q, 5′-GGT CTG TTT TAT CCT TAC CAG AAT ACG GCA GAT GGA GTT TAT AAA G-3′; N220Q, 5′-GTT TTC AAT GGA AAA GAT CAG ATC TCT AAA GTT GCC ATA ATC G-3′; N235Q, 5′-CAT ATA AAG GTA AAA GGC AGC TGT CCT ATT GGG AAA G-3′; N247Q, 5′-CAC TGC GAC ATG ATT CAG GGT ACA GAT GCA GCC-3′; N321Q, 5′-GAA AAA ATT ATC TCA AAG CAA TGT ACA TCA TAT GGT GTG CTA G-3′; N417Q, 5′-TGT GCC TAT TCT TTG GCT CCA GGA GAC TGG GAC CAT TGG-3′.
Mammalian Cell Culture and Protein Expression
HEK293T cells were grown in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum (Invitrogen) at 37 °C in 5% CO2. The cells were transfected transiently with wild type and mutant pCI-CD36–12His or pCI-ABCB1–12His, as described previously (37). 24 h after transfection the cells were treated with 33 μm butyric acid (Sigma) and cultured for a further 24 h before harvesting using trypLETM Express (Invitrogen) for use in flow cytometry and immunoblotting. Tunicamycin (500 ng/ml) was added 5 h post-transfection where indicated.
Insect Cell Culture and Protein Expression
Suspension cultures of Spodoptera frugiperda 21 (Sf21) cells were grown in SF900II serum-free medium (Invitrogen) at 27 °C with shaking at 100 rpm. Cells at a density of 2 × 106 cells/ml were infected with recombinant baculovirus encoding wild type or non-glycosylated CD36 using a multiplicity of infection of at least 3 viruses per cell. After several hours the culture was diluted to a density of 1 × 106 cells/ml with fresh SF900II media.
Insect Cell Membrane Preparation
At 72 h post-infection, the insect cells were harvested by centrifugation at 1000 × g, 4 °C for 10 min and washed in ice-cold buffer 1 (10 mm Tris-HCl, pH 7.5, 250 mm sucrose, 0.2 mm CaCl2, 2 mm benzamidine, 40 μm leupeptin, and 1 μm pepstatin A). The cells were resuspended in 10 ml of buffer 1 and frozen at −20 °C. Once thawed, the cells were homogenized at 4 °C by 5 × 30-s bursts at 24,000 rpm (DI 25 homogenizer; Yellow Line). The sample was centrifuged at 500 × g for 10 min at 4 °C to pellet the large organelles and unbroken cells. The supernatant was recovered and centrifuged at 100,000 × g in a TLA100.3 rotor (Beckman Coulter) for 50 min at 4 °C to obtain pelleted membranes. The crude membrane fraction was resuspended in buffer 2 (buffer 1, minus CaCl2) supplemented with 10% (v/v) glycerol and stored at −80 °C. Total protein concentrations of the membrane fractions were determined by DC Protein Assay (Bio-Rad).
Solubilization and Purification of CD36 from Membrane Fractions
Membrane fractions were pelleted by centrifugation at 100,000 × g for 30 min in a TLA100.3 rotor at 4 °C. The pellets of wild type CD36 were resuspended in solubilization buffer (20 mm Tris-HCl, pH 6.8, 2% (w/v) OG, 150 mm NaCl, 1.5 mm MgCl2, 5% (v/v) glycerol, 2 mm benzamidine, 40 μm leupeptin, and 1 μm pepstatin A) at 5 mg protein/ml, homogenized by extrusion in a 21-gauge needle, and constantly mixed for 90 min at 4 °C. The insoluble fraction was pelleted by ultracentrifugation at 100,000 × g for 30 min in a TLA100.3 rotor at 4 °C. Ni-NTA resin was pre-equilibrated in equilibration buffer (solubilization buffer where 2% OG was replaced with 1% OG in the presence of 20 mm imidazole). Imidazole (20 mm) was added to the solubilized fraction of wild type CD36 membranes and incubated with the Ni-NTA resin using a protein:resin ratio of 8:1 with continuous mixing for 1 h at 4 °C. The resin was washed 4 times with 20 bed volumes and a stepwise gradient of imidazole (60–120 mm) in wash buffer (20 mm Tris-HCl pH 8.0, 150 mm NaCl, 1.5 mm MgCl2, 5% (w/v) glycerol, 1% (w/v) OG, 2 mm benzamidine, 40 μm leupeptin, and 1 μm pepstatin A) to eliminate proteins bound non-specifically to the resin. Wild type CD36 was eluted using equilibration buffer plus 250 mm imidazole. The purification efficiency was visualized by SDS-PAGE stained with colloidal blue. An identical procedure was used for purification of non-glycosylated CD36 (CD36non-g) except 0.6% SDS substituted 2% OG in the solubilization buffer and 0.3% SDS replaced 1% OG in the equilibration and wash solutions. The eluted protein was concentrated using centrifugal devices with a 50-kDa cut off as directed (Amicon Ultra 15, Millipore).
Deglycosylation of Purified Wild Type CD36 Protein
For use in mass spectrometry, ∼10 pmol of wild type CD36 was denatured at 100 °C for 10 min and deglycosylated using PNGase F for 1 h at 37 °C as directed (New England Biolabs).
Mass Spectrometry
Approximately 10 pmol of purified wild type CD36 (pre and post-deglycosylation) were separated by SDS-PAGE and stained with colloidal blue. The protein bands were excised and digested with trypsin using MassPREP Station (Waters) for the liquid chromatography/tandem mass spectroscopy (LC/MS/MS) or BioRobot 3000 (Qiagen) for the Fourier transform ion cyclotron resonance (FT-ICR MS). Peptides were extracted using 0.1% formic acid, and the tryptic peptide mixture was analyzed by automated LC/MS/MS (CapLC, LC Packings, Q-ToF II, Waters) as described (38) or Fourier transform mass spectrometry (LTQ-FT hybrid linear trap/7-T FT-ICR mass spectrometer (Thermo Electron, Bremen, Germany)) as described (39). Data from LC/MS/MS and FT-ICR MS were analyzed in conjunction with the MSDB data base using the software tool Mascot (Matrix Services) and Sequest data base using Bioworks software (Thermo Scientific), respectively.
Flow Cytometry
After harvest, transiently transfected HEK293T cells were washed in FACS buffer (PBS and 1% fatty acid-free BSA) and resuspended at 1 × 107 cells per ml. A saturating concentration (2 μg) of mouse anti-CD36 mAb1258 (or mAb1955) was added to 50 μl of cells and incubated for 30 min at 4 °C (0.5 μg of monoclonal antibody 4E3 (DAKO) was substituted when staining for P-glycoprotein). The cells were centrifuged at 400 × g for 1 min at 4 °C and resuspended in 1 ml of FACS buffer. The wash step was repeated twice more, and the cells were resuspended in 50 μl of FACS buffer. A saturating concentration (4 μg) of goat, anti-mouse IgG secondary antibody conjugated to R-phycoerythrin (DAKO) was added to the cells and incubated in the dark for 30 min at 4 °C. The cells were recovered by centrifugation and washed as before and then resuspended in 400 μl of FACS buffer. During flow cytometry, 10,000 cells of normal size and granularity were analyzed for CD36–12His surface expression measuring R-phycoerythrin fluorescence (Ex 565 nm and Em 578 nm). The cell surface expression of CD36–12His was analyzed by FlowJo (Treestar). The cells typically exhibited a biphasic staining pattern, likely dependent on whether the individual cell was just about to, or had recently divided, before transfection. The heights of these peaks (reflecting cell number) relative to each other sometimes varied, but the fluorescence intensity of each peak (reflecting CD36 density) remained consistent relative to the positive and negative controls (wild type CD36 and CD36non-g, respectively). The maximal expression level of the receptor (the peak with the higher fluorescence intensity) was, therefore, gated, and the median was calculated and compared in all experiments. The expression level of mutant CD36 proteins was always compared with the expression levels of wild type and non-g proteins performed contemporaneously to control for minor variability in transfection efficiency on different days. -Fold reduction in expression was calculated by dividing the percentage surface expression of the wild type protein by the percent surface expression of the mutant. The synergy factor was calculated by dividing the -fold reduction of the multiple mutant by the product of the -fold reduction of the individual mutants, as described (40).
In Vitro Solid Phase Ligand Binding Assay
For purified protein, 1 μg of wild type CD36–12His was added to Ni-NTA-coated plates (Qiagen) in 100 μl of protein binding buffer (20 mm Tris-HCl, pH 6.8, 150 mm NaCl, 1.5 mm MgCl2, 5% (v/v) glycerol, 0.5% (w/v) OG, 2 mm benzamidine, 40 μm leupeptin, and 2 μm pepstatin A) and bound at 4 °C overnight with gentle rocking. Unbound protein was removed, and each well was washed with 2 × 150 μl of ligand binding buffer 1 (LBB1; PBS, 1 mm MgCl2, 1 mm CaCl2, 0.5% (w/v) OG, 0.2% fatty acid-free BSA). Increasing concentrations of BODIPY Ac-LDL were added in LBB1 to a total volume of 100 μl and incubated at room temperature in the dark with gentle rocking for 2 h. Unbound ligand was removed, and the wells were washed with 3 × 200 μl of ice-cold wash buffer 1 (PBS, 1 mm MgCl2, 1 mm CaCl2, 0.5% fatty acid-free BSA). 100 μl of PBS was added per well before determining the bound fluorescence using a fluorescent plate reader (SpectraMax, Gemini EM, Molecular Devices, Ex 485 nm, Em 530 nm). Specific binding of Ac-LDL was calculated after subtraction of the level of BODIPY Ac-LDL bound non-specifically to empty wells.
Cell-based Ligand Binding Assay
For the whole cell assay, 1 × 105 cells were seeded per well of polylysine-coated flat-bottomed 96-well plates and transfected with plasmid DNA, as described above. 48 h after transfection, the cells were washed with 3 × 200 μl wash buffer 2 (WB2; PBS, 1 mm MgCl2, 1 mm CaCl2), then 150 μl of block buffer (WB2 with 1% fatty acid-free BSA) was added to each well and incubated at room temperature for 45 min with gentle rocking. Increasing concentrations of BODIPY Ac-LDL were added in WB2 plus 0.2% fatty acid-free BSA to a total volume of 100 μl and incubated at 4 °C in the dark with gentle rocking for 2 h. Unbound ligand was removed, and the wells were washed with 3 × 200 μl of ice-cold WB2. 100 μl of PBS was added per well before determining the bound fluorescence as described above and calculation of the specific binding by subtraction of the level of BODIPY Ac-LDL bound non-specifically to mock-transfected cells.
Ligand Binding Analysis
Data were analyzed using Graphpad Prism software Version 4.0, and saturation binding data were best fitted by Langmuir adsorption isotherm (Equation 1), which describes binding of ligand to a single class of binding site as
 |
where B is bound ligand (relative fluorescent units), [L] is concentration of ligand (μg/ml), and Kd is the concentration of ligand giving half-maximal binding and a measure of the affinity of ligand-receptor interaction.
Immunoblotting
HEK293T cell lysates (50 μg) (untreated or treated with 1 unit of Endo H or 1 unit of PNGase F as directed (New England Biolabs)) were separated by SDS-PAGE and transferred to polyvinylidene difluoride membranes (Millipore). Western blots were probed with rat anti-CD36 primary mAb1955 (or mouse anti-P-glycoprotein primary C219) and rabbit anti-rat (or goat anti-mouse as appropriate) secondary antibody-conjugated to horseradish peroxidase (DAKO) before visualization by ECL chemiluminescent detection system (Amersham Biosciences) as directed.
RESULTS
Overexpression, Purification, and Functional Integrity of Human CD36 in Insect Cells
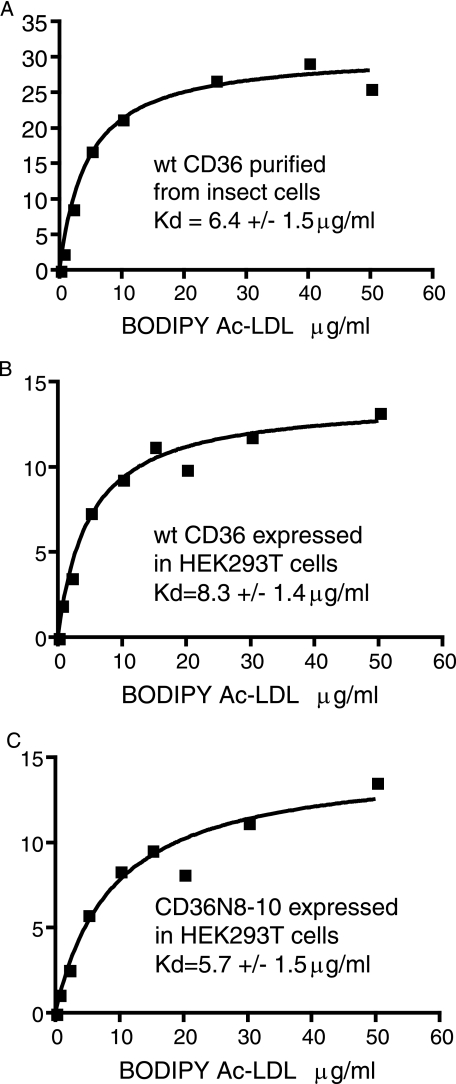
Using the consensus sequence N-X-Ser/Thr (where X is any amino acid except proline), asparagines 79, 102, 134, 163, 205, 220, 235, 247, 321 and 417 (henceforth designated N1—N10 for convenience) were identified as possible sites of modification. Mass spectrometric analysis of purified protein can be used to determine the occupancy of these sites; therefore, recombinant wild type CD36 was engineered with a 12 histidine, carboxyl-terminal tag and expressed in Sf21 insect cells using a baculoviral system. Proteins were solubilized in OG from insect cell membrane fractions, and the recombinant CD36 was purified by nickel-NTA affinity chromatography (Fig. 1A). Although insect cells faithfully recognize the mammalian glycosylation sites (41), they do not elaborate on the core glycan paucimannose (42). It was, therefore, important to establish that the CD36 produced in the heterologous host was functional and, thus, suitable for analysis. The purified protein was immobilized on plates coated with Ni-NTA and its affinity for Ac-LDL was determined (Fig. 2A; Table 1). The mean Kd ± S.E. = 6.4 ± 1.5 μg/ml (n = 3) for Ac-LDL of the receptor purified from insect cells was not significantly different from the affinity measured for the recombinant protein expressed on the surface of mammalian cells (Kd = 8.3 ± 1.4 μg/ml (n = 4); Fig. 2B; Table 1). Thus, the purified protein retains its native fold and ligand binding function, and the specific nature of the glycan is not important for ligand interaction.
FIGURE 1.
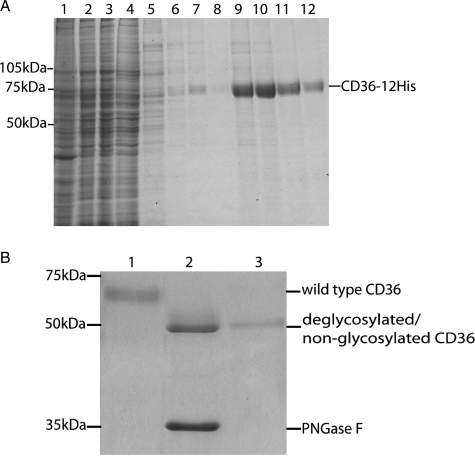
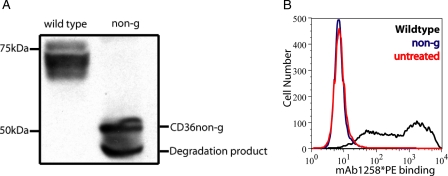
Purification of CD36–12His by affinity chromatography and deglycosylation by PNGase F. A, SDS-polyacrylamide gel stained with colloidal blue showing protein fractions during the purification of wild type CD36–12His from baculoviral-infected Sf21 cells. Lane 1, 50 μg of crude membrane fraction (0.1% of starting material); lane 2, OG-solubilized membrane proteins (0.1% by volume); lane 3, proteins failing to bind to the Ni-NTA (0.1% by volume); lanes 4–8, washes 60–120 mm imidazole of the Ni-NTA resin to remove proteins non-specifically bound (2% by volume); lanes 9–12, elution fractions in 250 mm imidazole (2% by volume). B, the electrophoretic mobility of purified recombinant wild type CD36 (lane 1) was compared with CD36 after deglycosylation by PNGase F (lane 2) and non-glycosylatable CD36non-g purified from protein aggregates (lane 3).
FIGURE 2.
The affinity of CD36 for ligand is not affected directly by the nature or pattern of glycosylation. Interaction of CD36 proteins with Ac-LDL was measured using increasing concentrations of BODIPY Ac-LDL to core-glycosylated, wild type (wt) CD36 purified from Sf21 insect cells and immobilized on a solid phase (A), mature glycosylated, wild type CD36 on the surface of HEK293T cells (B), and CD36N8–10 on the surface of HEK293T cells (C). A representative graph from each is shown and is plotted according to Equation 1; the Kd quoted represents the mean ± S.E. (μg/ml) from at least three independent experiments.
TABLE 1.
Binding affinity of BODIPY Ac-LDL to CD36 proteins
| Protein | Kd(mean ± S.E.) |
|---|---|
| μg/ml | |
| Wild type CD36 (purified from Sf21) | 6.35 ± 1.47 n = 3 |
| Wild type CD36 (expressed in HEK293T) | 8.26 ± 1.38 n = 4 |
| CD36N8–10 (expressed in HEK293T) | 5.73 ± 1.54 n = 4 |
| CD36N1–7,9,10 (expressed in HEK293T) | 6.21 ± 0.03 n = 2 |
| CD36N1–8,10 (expressed in HEK293T) | 4.12 ± 0.32 n = 2 |
| CD36N1–9 (expressed in HEK293T) | 4.24 ± 0.03 n = 2 |
Identification of N-Linked Glycosylation Site Occupancy of CD36 by Mass Spectrometry, Mutagenesis, and Electrophoretic Mobility
Digestion of the CD36 purified from insect cells with PNGase F increased the mobility of the protein during SDS-PAGE (Fig. 1B). The apparent mass difference of 20 kDa suggests several of the 10 putative glycosylation sites are modified. The PNGase F-treated CD36 migrated with similar mobility to a non-glycosylatable version of the protein (CD36non-g, described below) consistent with complete N-linked deglycosylation of wild type CD36. Deglycosylation of a protein by PNGase F removes the N-linked oligosaccharide from the peptide chain and converts the previously glycosylated asparagine to an aspartic acid. This results in a mass shift of +1 for each deamidation event as compared with the predicted molecular mass of the protein backbone containing a non-utilized asparagine (the molecular mass of a glycopeptide is difficult to predict and to measure because of the heterogeneity of the sugar chains and increased mass to charge ratio). We used this mass shift associated with the deglycosylation to assess the glycosylation status of the protein by Q-ToF mass spectrometry after tryptic digestion. Complete tryptic digest of wild type CD36 generates 46 fragments, of which 7 contain the putative N-linked glycosylation sites. In Q-ToF MS, peptides are ionized to form, among others, different y-series carboxyl-terminal and b-series amino-terminal ions, and analysis of the mass and charge of these ions determines the amino acid sequence of the peptide (Fig. 3A).
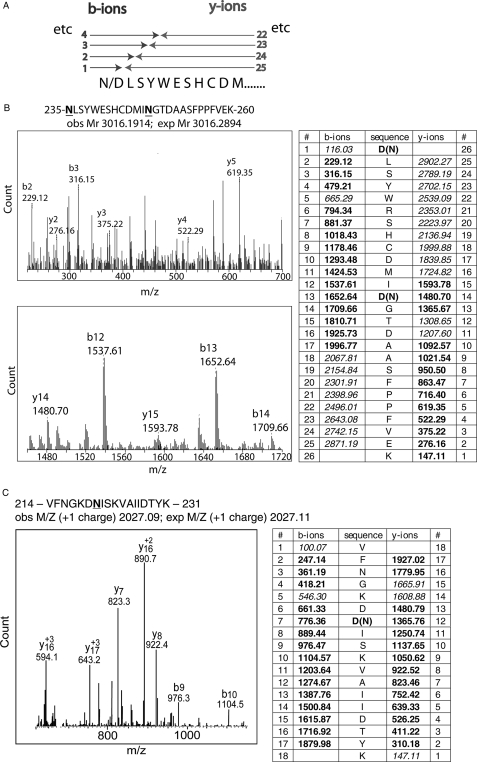
FIGURE 3.
Mass spectrometry showing evidence of glycosylation of sites N6, N7, and N8 of CD36 purified from Sf21 insect cells. After tryptic digestion, protein fragments are ionized into related y- and b-series protonated ions (as shown by the schematic in A) with unique mass/charge (m/z) detectable by MS. B, selected portions of Q-ToF MS spectra of the PNGase F and trypsin-digested wild type CD36. The spectra show the diagnostic ions, which demonstrate that N7 and N8 are deglycosylated and deamidated to aspartic acids by treatment with PNGase F, result in a mass shift of +1 each. C, selected portion of FT-ICR MS spectrum of similarly treated wild type CD36 showing that N6 can be glycosylated. The sequence of the tryptic digest fragments with the putative glycosylated asparagine underlined and the complete y and b-series ions generated by the ionization of the PNGase-F-deglycosylated and -deamidated fragment are shown. For Q-ToF MS, those single-charged ions detected by MS are shown in bold. Fragments not detected are shown in italics. Note in the FT-ICR MS spectrum, some of the y- and b-series ions have more than one charge. The sequence of the fragments and the observed and expected Mr of the monoisotopic parent ion (accounting for the deamidation of the underlined Asn to Asp) are given above the spectra. For the full spectra, tables of ions detected and deviations from the expected ion mass, see supplemental Fig. 1.
For example, tryptic fragment 235NLSYWESHCDMINGTDAASFPPFVEK260 contains 2 putative glycosylation sites, N7 and N8, with N7 at the amino terminus of the fragment. As N7 is at the amino terminus, the b-ions determine whether N7 has been deglycosylated and deamidated to an aspartic acid (Fig. 3B, upper panel). The b-ions with m/z 229.12 (b2) and m/z 316.15 (b3) are shown on the spectrum, and both are diagnostic for aspartic acid at position 235, indicative of modification of the wild type Asn (N7) by glycosylation. Putative glycosylation site N8 is located in the middle of the tryptic fragment; therefore both b-ions and y-ions can determine whether this site is occupied. The b-ion with m/z 1709.66 (b14) is diagnostic for aspartic acids in both positions 235 and 247, and the y-ions with m/z 1593.78 (y15) and 1480.70 (y14) indicate an aspartic acid in position 247, indicative that N8 is glycosylated in wild type CD36 (Fig. 3B, lower panel). Fragments with asparagines that are not within consensus glycosylation sites show little evidence of similar mass change, indicating that spontaneous deamidation is rare (data not shown).
Q-ToF MS on tryptic digest fragments from wild type CD36, in the absence of PNGase F treatment, failed to detect any fragments containing putative glycosylation sites, although other fragments of CD36, not predicted to be glycosylated, were detected. This too, is consistent with glycosylation of the relevant fragment, as the sensitivity of glycopeptide detection is often low due to signal suppression and the likely heterogeneity of glycosylation. Q-ToF mass spectrometry of the deglycosylated protein identified five of the seven tryptic fragments sought and was able to establish that N1, N5, N7, N8, N9, and N10 were utilized as glycosylation sites (full spectra, data tables of the ions detected, and their mass errors from the calculated values are shown in supplemental Fig. 1, A–E).
The remaining four putative glycosylation sites were located in tryptic digest fragments too large (6953 mass units for N2, N3, N4) or too small (577 mass units for N6) to be detected by Q-ToF MS. We, therefore, utilized FT-ICR MS to attempt to determine the glycosylation status of these putative sites. FT-ICR MS achieved 42% coverage of the protein and identified that N6 can also be glycosylated (Fig. 3C shows the spectrum from a partial tryptic digest product containing N6). However, not all fragments exhibited this mass shift, indicating that N6 modification is unlikely to be comprehensive (the full spectra of both these fragments are recorded in supplemental Fig. 1, F and G). Again, the fragment containing N2, N3, and N4 was not detected. This is most likely due to a low charge state for the peptide, which has only one internal histidine, precluding an m/z value lower than 2000. As digestion with other proteases did not result in fragments suitable for detection by mass spectrometry, electrophoretic mobility of CD36 isoforms with and without the test site was used to determine the glycosylation status of these putative sites and corroboration of the incomplete utilization of N6.
Glycosylation Status of Sites N2, N 3, N4, and N6
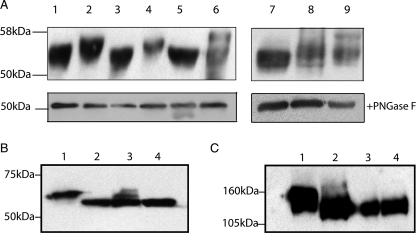
Reasoning that the difference in electrophoretic mobility would be more evident in a less complex CD36 template with fewer glycosylation sites, we first established that CD36N8–10, with only the carboxyl-terminal three sites intact, trafficked to the cell surface (see below for characterization of this mutant). CD36N8–10 migrated with an apparent molecular mass of ∼55 kDa on SDS-PAGE (Fig. 4A). A series of mutants were generated based on the CD36N8–10 template to test the utilization of putative glycosylation sites N2, N3, N4, and N6. These sites were reintroduced individually into the minimally glycosylated template to generate CD36N2,8–10, CD36N3,8–10, CD36N4,8–10, and CD36N6,8–10 (Table 2). These mutants, together with CD36N8–10, were expressed in HEK293T cells, and the electrophoretic mobility of the proteins was compared by SDS-PAGE and immunoblotting by probing with mAb1955. It was hypothesized that if the introduced putative glycosylation site is occupied in any of the mutants, the electrophoretic mobility of this protein is likely to be slower than that of CD36N8–10. As a positive control, CD36N7–10 (N7 was shown by Q-ToF MS to be glycosylated in the wild type protein) was compared with CD36N8–10 and found to migrate more slowly through the gel (Fig. 4A, upper panel). The immunoblot suggests that N2, N3, and N6 are glycosylated as CD36N2,8–10, CD36N3,8–10, and CD36N6,8–10 migrate more slowly than CD36N8–10. After treatment with PNGase F, all of the CD36 proteins migrate with the same mobility (Fig. 4A, lower panel), confirming that the glycans on N2, N3, and N6 were responsible for the electrophoretic difference observed in Fig. 4A. In repeated experiments, inclusion of site N6 in the CD36N8–10 consistently resulted in partial glycosylation of the protein as judged by the significant level of multiple protein forms in lane 6 of the upper panel of Fig. 4A, which are resolved into a single species in the lower panel by treatment with PNGase F. Although we cannot rule out the possibility that inclusion of N6 may influence the occupancy of sites N8–N10, the simplest interpretation, and one that is consistent with the FT-ICR data on the wild type protein, is that N6 itself is recognized inefficiently as a glycosylation site. In these experiments we found no evidence to suggest N4 is modified by glycosylation. Taken together with the mass spectrometry data, it is evident that 9 of the 10 putative glycosylation sites can be modified by the glycosyl transferases.
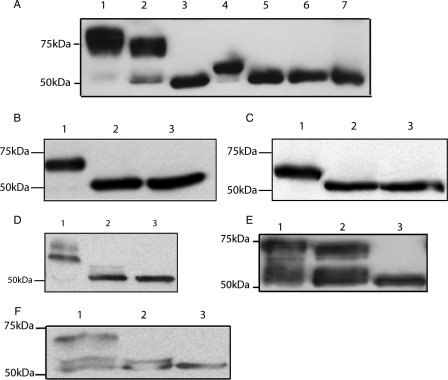
FIGURE 4.
Electrophoretic mobility shift analysis to test glycosylation status of CD36. A, whole cell lysates (50 μg of protein) from transiently transfected HEK293T cells were analyzed by immunoblotting probing with mAb1955 before (upper panel) and after (lower panel) deglycosylation with PNGase F. Lane 1, CD36N8–10; lane 2, CD36N8–10+N7; lane 3, CD36N8–10; lane 4, CD36N8–10+N3; lane 5, CD36N8–10+N4; lane 6, CD36N8–10+N6; lane 7, CD36N8–10; lane 8, CD36N8–10+N7; lane 9, CD36N8–10+N2. B and C, analysis of CD36N8–10 probed with mAb1955 (B) and P-glycoprotein probed with mAbC219 (C) after tunicamycin treatment of the transfected cells to inhibit glycosylation: lane 1, untreated sample; lane 2, treated with PNGase F; lane 3, treated with tunicamycin; lane 4, treated with tunicamycin and PNGase F.
TABLE 2.
CD36 mutants generated, and details of asparagines that have been mutated to glutamines
Glycosylation Is Necessary for Trafficking of CD36 to the Cell Membrane
To assess the role of glycosylation in the biology of CD36, we first removed all 10 putative glycosylation sites, replacing the conserved asparagines with glutamines by site-directed mutagenesis to generate CD36non-g. The mutant cDNA was expressed in the human cell line HEK293T. CD36non-g is translated by HEK293T cells as shown by Western blotting, probing with the anti-CD36 monoclonal mAb1955 (Fig. 5A), but when intact cells were analyzed by flow cytometry for surface expression using mAb1258 (or mAb1955, data not shown), it is clear that CD36non-g fails to traffic to the plasma membrane (Fig. 5B).
FIGURE 5.
Non-glycosylatable CD36non-g fails to traffic to the plasma membrane. A, immunoblot analysis probing with mAb1955 confirms CD36non-g is translated when expressed transiently in HEK293T cells; the lower molecular weight species in the non-g lane is not always observed and is likely to be a degradation product. B, whole cell flow cytometry analysis using an antibody (mAb1258) that recognizes an epitope in the extracellular loop of CD36 indicates that CD36non-g fails to traffic to the plasma membrane. Wild type CD36 is shown in black, CD36non-g shown in blue, and untransfected cells shown in red. PE, R-Phycoerythrin.
Determination of the Minimal Glycosylation to Support Trafficking of CD36
To determine which N-linked glycosylation sites are required for CD36 to reach the cell surface, we reintroduced glycosylation sites back into CD36non-g (for details of all mutants generated, see Table 2). Flow cytometry was used to analyze the level of cell surface expression of the mutated proteins expressed in HEK293T cells. Saturating levels (data not shown) of primary mAb1258 and secondary antibodies were used to allow quantification of surface expression. FACS analysis of two mutants with an overlapping pattern of intact glycosylation sites N1–7 (CD36N1–7) or N7–10 (CD36N7–10) (Fig. 6A) showed that both reached the plasma membrane but were expressed to very different levels (1.6 and 11% of wild type CD36 (wtCD36), respectively; the top section of Table 3) (although only 1.6% of the level of wild type, antibody binding to cells expressing CD36N1–7 is significant, being 20-fold greater than the background binding associated with CD36non-g). This indicated two things; first, it was likely that there was both an amino-terminal and a carboxyl-terminal set of glycosylation sites that supported trafficking (unless N7 proved to be the key site); second, the low level of expression of both mutants suggested that sites in both the amino- and carboxyl-terminal sets were likely to be necessary to achieve high level expression of the receptor.
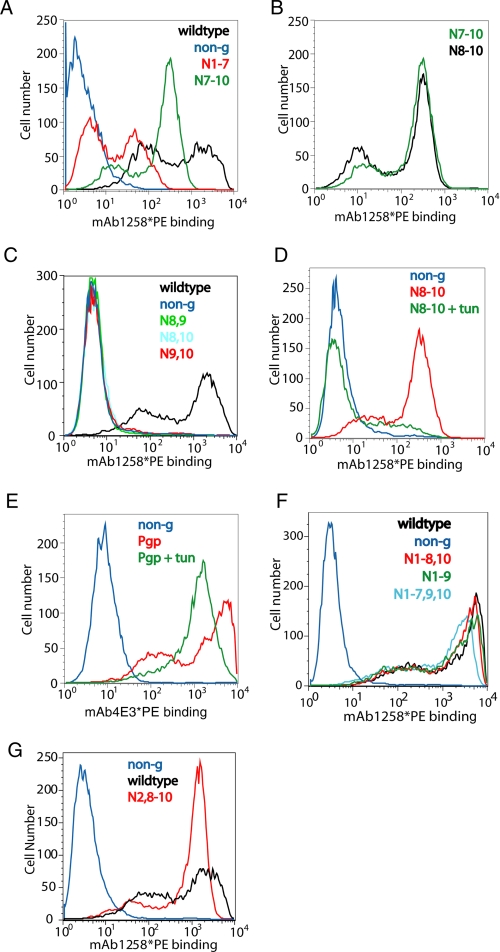
FIGURE 6.
Cell surface expression of CD36 mutants. Wild type and mutated CD36 and P-glycoprotein were expressed transiently in HEK293T cells, and flow cytometry was used to measure the cell surface expression using saturating amounts of antibody. Wild type CD36, which traffics efficiently, was used as the positive control (black trace, where shown), and CD36non-g, which fails to traffic, was used as the negative control (blue trace, where shown) for all CD36 experiments. PE, R-Phycoerythrin. A, cell surface expression of CD36N7–10 (green) and CD36N1–7 (red). B, CD36N7–10 (green) and CD36N8–10 (black). C, CD36N8,9 (green), CD36N8,10 (light blue), and CD36N9,10 (red). D, surface expression of CD36N8–10 in untreated (red) and tunicamycin (tun)-treated cells (green). E, surface expression of P-glycoprotein (Pgp) in untreated (red) and tunicamycin-treated cells (green). F, cell surface expression of CD36N1–8,10 (red), CD36N1–9 (green), and CD36N1–7,9,10 (light blue). G, cell surface expression of CD36N2,8–10.
TABLE 3.
The cell surface expression of mutant CD36 compared to wild type, measured by flow cytometry
Surface expression values are calculated for the cell subpopulations with the highest fluorescence and normalized to 100% for the wild type protein and 0% for CD36non-g (i.e. the same fluorescence as background binding to mock-transfected cells).
| CD36 mutant | Surface expression % ± S.D. (n =3) |
|---|---|
| CD36 wild-type | 100 |
| CD36non-g | 0 |
| CD36N1–7 | 1.6 ± 0.3 |
| CD36N7–10 | 11 ± 1.7 |
| CD36N1–7 | 1.6 ± 0.3 |
| CD36N1–6 | 1.3 ± 0.6 |
| CD36N1–5 | 0 |
| CD36N2–7 | 0.3 ± 0.2 |
| CD36N3–7 | 0 |
| CD36N1,2,6,7 | 0 |
| CD36N7–10a | 11 ± 1.7 |
| CD36N8–10 | 12 ± 2.0 |
| CD36N7,9,10 | 0 |
| CD36N7,8,10 | 0 |
| CD36N7–9 | 0 |
| CD36N8,9 | 0 |
| CD36N8,10 | 0 |
| CD36N9,10 | 0 |
| CD36N1–7,8 | 34 ± 2.0 |
| CD36N1–7,9 | 30 ± 1.7 |
| CD36N1–7,10 | 24 ± 2.1 |
| CD36N1–7,9,10 | 63 ± 14 |
| CD36N1–8,10 | 78 ± 5.5 |
| CD36N1–9 | 75 ± 4.0 |
| CD36N8–10 | 12 ± 2.0 |
| CD36N1,8–10 | 31 ± 4.2 |
| CD36N2,8–10 | 59 ± 5.6 |
| CD36N3,8–10 | 31 ± 3.2 |
| CD36N4,8–10 | 13 ± 1.5 |
| CD36N5,8–10 | 60 ± 5.0 |
| CD36N6,8–10 | 31 ± 3.2 |
| CD36N7,8–10a | 11 ± 1.7 |
a These are the same mutant.
Characterization of the Amino-terminal Glycosylation Set
To further assess which of the seven amino-terminal glycosylation sites N1–7 of CD36 (six of which are modified by N-linked glycosylation) were necessary for the protein to reach the cell membrane, sites were removed systematically from each end (the second panel of Table 3). The expression level of CD36N1–6 was essentially the same as CD36N1–7, but CD36N1–5 failed to traffic to the cell surface, whereas CD36N2–7 caused a 5-fold reduction in trafficking (as compared with CD36N1–7), and CD36N3–7 failed to reach the plasma membrane (all of the mutants generated were analyzed by Western blotting using the monoclonal antibody mAb1955, which recognizes both the wild type and the non-glycosylated version of the protein to confirm that the glycosylation site mutants were synthesized by the HEK293T cells irrespective of surface representation (data not shown)). These data could suggest the importance of N1, N2, and N6 in the amino-terminal set; however, a mutant with only sites N1, N2, N6, and N7 present failed to traffic to the cell surface, indicating that these sites can only exert an effect in the context of a more heavily glycosylated protein. Indeed, given that N4 does not appear to be modified by N-linked glycosylation, there is a correlation between glycosylation site density and trafficking within the amino-terminal set. Thus, all mutants with five or more of the utilized amino-terminal glycosylation sites irrespective of combination appear on the cell surface, whereas mutants with fewer than five utilized sites are unable to support trafficking.
This interdependence of glycosylation sites made assessing the contribution of the amino-terminal sites to the trafficking efficiency difficult; however, their influence on the trafficking when combined with the carboxyl-terminal set suggested that sites 2 and 5 were particularly important (see below). Nevertheless, the demonstration that CD36N1–6 can be expressed at the cell surface and the knowledge that it shares no common glycosylation site with CD36N7–10 indicate that none of the glycosylation sites is individually essential for CD36 to traffic to the cell surface.
Characterization of the Carboxyl-terminal Glycosylation Set
Generation and analysis of four mutants CD36N7,9,10, CD36N8–10, CD36N7,8,10, and CD36N7–9 (third panel of Table 3) confirmed that N7 was not required for trafficking of CD36 to the plasma membrane, because the surface expression of CD36N7–10 was essentially the same as CD36N8-10 (Fig. 6B). The failure of CD36N7,9,10, CD36N7,8,10, and CD36N7–9 to express at the cell surface suggested that sites N8, N9, and N10 (all of which were shown by Q-ToF MS to be modified by N-linked glycosylation) were essential for expression and that the three intact glycosylation sites in CD36N8–10 were the minimal number in the carboxyl terminus that would allow cell surface expression. Indeed, mutant proteins with only two of these glycosylation sites, CD36N8,9, CD36N8,10 and CD36N9,10 (third panel of Table 3), were unable to traffic to the cell membrane (Fig. 6C).
Carboxyl-terminal Glycosylation Sites Contribute Significantly to the Cell Surface Expression in the Presence of the Amino-terminal Set
Although the paired carboxyl-terminal glycosylation sites were unable to support trafficking of an otherwise non-glycosylated CD36, the individual sites do have a strong influence on the level of trafficking when amino-terminal glycosylation sites are present. This became evident when each of the three sites was restored individually to CD36N1–7. CD36N1–7,8, CD36N1–7,9, and CD36N1–7,10 trafficked to the cell membrane with 34, 30, and 24% efficiency of the wild type protein (Table 3, fourth panel).
Generation of the reciprocal set of mutants (removing each of the carboxyl-terminal three sites individually from an otherwise wild type CD36) allowed us to assess whether these three sites functioned additively or synergistically to influence the trafficking of the protein. CD36N1–7,9,10, CD36N1–8,10, and CD36N1–9, trafficked to the cell membrane at, on average, 63, 78, and 75% that of the level of wild type CD36 (Fig. 6F; fifth panel of Table 3). When compared with the trafficking efficiency of CD36N1–7, from which all three carboxyl-terminal sites are absent and which is present at the cell surface at 1% that of the level of the wild type protein, there is a strong suggestion that the three carboxyl-terminal sites function synergistically. This is because the -fold reduction of the individual mutants compared with the wild type range between 1.3 and 1.6, but the surface expression of the triple mutant is reduced 62.5-fold. The synergy factor, i.e. the -fold reduction of the triple mutant, divided by the product of the -fold reduction of the three individual mutants, should approximate 1 if the mutants are additive. Here the synergy factor of 23.1 suggests a strong synergistic linkage between the three sites and trafficking of the protein (Table 3).
Glycosylated Sites in the Amino Terminus Contribute Significantly to the Cell Surface Expression in the Presence of the Carboxyl-terminal Set
To assess the contribution of amino-terminal glycosylation sites, we restored each, individually, to CD36N8–10 and assessed their ability to traffic to the plasma membrane (bottom panel of Table 3). Only site N7 and site N4 had non-significant effects on the trafficking of the protein, entirely consistent with our earlier observations that N7 had no effect on trafficking when part of the amino-terminal set and that N4 is not utilized as a glycosylation site. Each of the other amino-terminal sites significantly improve the trafficking of CD36N8–10, with sites N2 and N5 having the greatest effect, achieving ∼60% of the cell surface expression level of the wild type protein (to illustrate, the surface expression of CD36N2,8–10 is shown in Fig. 6G).
Glycosylation, and Not the Reintroduction of the Asparagines, Determines Trafficking Efficiency
The data indicate the presence of at least two minimal sets of glycosylation sites that support trafficking of CD36 proteins. To rule out the possibility that it is the reintroduced asparagines that determines trafficking rather than their modification by glycosylation, we used tunicamycin (an inhibitor of glycan precursor synthesis) to prevent modification of the asparagines. Western analysis showed that glycosylation of the CD36N8–10 mutant (Fig. 4B) and also of the multidrug resistance P-glycoprotein (Fig. 4C), which can traffic and function in the absence of glycosylation (43), was efficiently, albeit incompletely, inhibited in the tunicamycin-treated cells. The ability of these proteins to reach the cell surface in tunicamycin-treated and untreated cells was then compared by flow cytometry (Fig. 6, D and E). CD36N8–10 in cells treated with tunicamycin failed to reach the cell surface (Fig. 6D), but P-glycoprotein did (Fig. 6E). Repetition of the experiment with the CD36N1–7 mutant (data not shown) gave a very similar result to CD36N8–10. The simplest interpretation of these data is that it is the N-linked glycosylation of CD36 that enables the protein to fold and traffic correctly.
Mature Glycosylation Is Not Necessary for Cell Surface Expression of CD36
Western analysis of the mutants generated (probed with mAb1955) confirmed that all proteins including those that failed to traffic to the plasma membrane were synthesized by HEK293T cells. Those proteins with utilized glycosylation sites that fail to reach the cell surface do, also, become glycosylated, indicative of translation at, and translocation into, the endoplasmic reticulum. For example, the electrophoretic mobility of CD36N7,9,10 is increased not only by digestion with PNGase F but also by endoglycosidase H, which cleaves immature, high mannose, glycans from the protein (Fig. 7A). The presence of glycosylation in proteins that do not appear on the cell surface indicates that the mutant is stuck in the trafficking pathway. A fraction of wild type CD36 also appears sensitive to endoglycosidase H (judging by the slight increase in mobility of the main protein band in Fig. 7A, lane 2); however, whether high mannose glycans occupy the same glycosylation site in each molecule or whether the slight endoglycosidase H sensitivity reflects incomplete maturation of glycans over the population as a whole remains to be established.
FIGURE 7.
Some trafficking-competent CD36 isoforms are only core-glycosylated. 50 μg of whole cell protein lysates from transiently transfected HEK293T cells treated with either 1 unit of Endo H or PNGase F were analyzed by immunoblotting probing with mAb1955. A, wild type CD36: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F. CD36N7,9,10: lane 4, untreated; lane 5, treated with Endo H; lane 6, treated with PNGase F; lane 7, CD36 non-g. B, CD36N1–7: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F. C, CD36N8–10: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F. D, CD36N2,8–10: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F. E, CD36N1–7,9,10: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F. F, CD36N1–7,9: lane 1, untreated; lane 2, treated with Endo H; lane 3, treated with PNGase F.
We also analyzed the nature of the glycan in CD36 isoforms that trafficked to the cell surface and were surprised to find that a number of mutants were sensitive to endoglycosidase H. For example, CD36N1–7 and CD36N8–10, which share no glycosylation sites in common, are sensitive to deglycosylation by EndoH (Fig. 7, B and C, respectively). This does not appear to be related to the efficiency of trafficking, as CD36N2,8–10, which is expressed on the cell surface at 59% that of wild type levels (Fig. 6G), can be efficiently deglycosylated by Endo H (Fig. 7D). On the other hand, CD36N1–7,9,10, which lacks site N8 but is expressed at the same level as CD36N2,8–10, is substantially more resistant to deglycosylation by Endo H, indicating that its glycans are, predominantly, of a mature form (Fig. 7E). The glycans on CD36N1–7,8,10 and CD36N1–9, each of which is missing one of the three carboxyl-terminal sites, also have predominantly mature glycans. However, each of the three double site mutants (CD36N1–7,8, CD36N1–7,9, and CD36N1–7,10) is largely sensitive to endoglycosidase H (CD36N1–7,9 is shown in Fig. 7F to illustrate).
Glycosylation of Specific Sites on CD36 Is Not Necessary for Ligand Binding
Solid phase ligand binding assay was used to establish whether the minimally glycosylated mutant CD36 proteins retained ligand binding functionality. Equilibrium binding over a range of ligand concentrations was used to measure the affinity of CD36N8–10 and CD36N1–7 for Ac-LDL. The Kd values of wild type CD36 and CD36N8–10 are not significantly different, as determined by unpaired Student's t test (>95% confidence level) (Fig. 2C; Table 1); however, we failed to detect binding to CD36N1–7 (data not shown). The absence of binding to CD36N1–7 may have meant that N8–10 are essential for binding of Ac-LDL; however, affinity is retained in the CD36N1–7,9,10, CD36N1–8,10, and CD36N1–9 (Table 1), each of which is missing one of the three carboxyl-terminal sites. It is likely, therefore, that no individual glycan is important for ligand binding, and the failure to detect binding to CD36N1–7 is due to the very low surface expression level of this mutant.
DISCUSSION
The extracellular domain of the class B scavenger receptor CD36 is heavily modified by glycosylation. Ten consensus glycosylation sites (asparagines 79 (N1), 102 (N2), 134 (N3), 163 (N4), 205 (N5), 220 (N6), 235 (N7), 247 (N8), 321 (N9), and 417 (N10)) are present in this domain of the human form of the protein. Having established that the protein produced heterologously in insect cells and purified to near homogeneity is functionally equivalent to that expressed in mammalian cells, we used mass spectrometry to establish unequivocally that seven sites (N1, N5, N7, N8, N9, N10) were modified in wild type CD36. Evidence of glycosylation of N6 was observed in some but not all of the fragments detected. The central tryptic digest fragment containing the other three putative glycosylation sites was not detected by mass spectrometry, and a different analysis was required to determine the utilization of sites N2, N3, and N4. Reasoning that a molecular mass difference associated with an individual glycosylation event would be more reliably measured in a less complex glycoprotein with a smaller molecular mass, we first determined that N8, N 9, and N10 were the minimum number of carboxyl-terminal sites of glycosylation required for trafficking of CD36 to the plasma membrane.
Adding each of the three test sites (N2, N3, N4) individually to this template and, also N6, because the mass spectrometry suggested incomplete utilization of this site, produced a modest but significant reduction in the rate of migration in proteins containing N2, N3, or N6. These sites are therefore exposed to, and can be modified by, the glycosyltransferases on transit through the folding and trafficking pathway. However, again the data suggested that N6 is not always modified. No evidence was found for modification of N4. Thus, 9 of the 10 consensus glycosylation sites in human CD36 are likely to be modified by glycosylation, and of the 7 glycosylation sites conserved between human CD36 and bovine Cd36 (in which all 8 are reported to be utilized (34)), all were shown to be glycosylated in human CD36.
Non-glycosylatable CD36, with all 10 consensus sites mutated, fails to reach the cell surface, indicating that glycosylation is necessary for correct trafficking of the protein. CD36non-g protein is pelleted at 100,000 × g from mechanically disrupted insect cells along with cellular membranes. This suggests that CD36non-g is retained in an intracellular fraction with a high molecular weight. When combined with the insolubility of CD36non-g in all but the strongest ionic detergent (SDS), it seems likely that the protein is shunted from the folding and trafficking pathway to form protein aggregates, perhaps similar to bacterial inclusion bodies.
The total absence of trafficking of CD36non-g can be rescued by restoring glycosylation sites to the amino-terminal half (CD36N1–7) or to the carboxyl-terminal half (CD36N8–10) of the protein, demonstrating a level of redundancy in the glycan pattern needed for cell surface expression. Neither of these proteins is expressed highly, achieving 1.6 and 12% that of the wild type CD36 level, respectively. However, after restoration of N2 or N5 to CD36N8–10, a dramatic increase in expression can be attained (up to 60% that of the wild type level). Similarly, each of the carboxyl-terminal three sites is able to improve trafficking of the N1–7 mutant by up to 20-fold, suggestive of a synergistic relationship between the amino- and carboxyl-terminal glycosylation events for folding and trafficking of the protein.
CD36N8–10 retained its protein fold, being able to bind ligand with the same affinity as the wild type protein. However, none of these three carboxyl-terminal glycans was necessary for ligand binding because mutant proteins devoid of any one of the three sites retain affinity for ligand. In contrast, CD36N1–7 was not shown to bind ligand. This most likely reflects the low level expression of the N1–7 form of the protein. At only 1% of the level of the wild type protein, it may be impossible to distinguish specific binding to the CD36N1–7 in HEK293T cells. We cannot formally rule out the possibility that the carboxyl-terminal sites are specifically required for proper folding of the ligand binding domain, but this seems unlikely because all three of the single mutants, missing only N8 or N9 or N10, retain high affinity for ligand.
A major surprise was the finding that the glycans on these partially glycosylated proteins remained immature and sensitive to deglycosylation by Endo H. This is true of proteins that share no sites in common (e.g. CD36N1–7 and CD36N8–10), which together include all nine utilized glycosylation sites in the protein, some or all of which must be occupied by mature glycans in the wild type protein. It is also true of some proteins that are highly expressed (e.g. CD36N2,8–10). Although it is clear from our analysis of the wild type protein sourced from insect cells which are unable to elaborate core glycosylation that the nature of the glycan is not absolutely required for appropriate folding, trafficking, and ligand binding activity of CD36, it is not clear why the immature glycans on CD36N2,8–10 fail to become matured in the Golgi compartment of the mammalian cells. However, although our data are not conclusive in this matter, there is a correlation between site density on the protein and maturation of the glycans. The glycans on, for example, the single site mutants devoid of either N8, N9, or N10 (each of which contains eight glycosylation sites) become matured, yet those on the three double mutants CD36N1–7,8, CD36N1–7,9 and CD36N1–7,10 (each with seven glycosylation sites), CD36N1–7 (six glycosylation sites), or CD36N2,8–10 (four glycosylation sites) remain immature.
Our data are most consistent with a role for glycosylation in the folding and trafficking of the receptor. Potentially, proper folding of the protein could have been due to induction by intramolecular interaction of the amino acid side chains with the added glycans, as hypothesized in the folding of Erythrina corallodendron lectin (31, 44). However, although CD36N8–10 and CD36N1–7 do not share a common glycosylation site, they are both able to traffic to the cell membrane. None of N8, N9, or N10 was shown to be essential because single site mutants of each were expressed on the cell surface and retained affinity for Ac-LDL so it seems unlikely that the CD36 polypeptide folds around a specific glycan. The requirement for glycosylation of CD36 for trafficking, but the redundancy observed with regard to the number and position of these sites, is more consistent with the involvement of lectin-like molecular chaperones. Lectin-like chaperones have been shown to recognize the glycans of, for example, the human organic anion transporter hOAT4 (45) and influenza hemagglutinin (46) and to facilitate protein folding in the ER before trafficking to the Golgi. This would also explain why the receptor cannot be functionally expressed in bacteria.3 That no individual glycosylation event is essential for trafficking of CD36 contrasts with the situation in the related class B scavenger receptor SRBI in which glycosylation of two individual sites (neither of which are conserved in CD36) were shown to be necessary (33).
In summary, we report that 9 of the 10 putative glycosylation sites of CD36 can be modified. We show that glycosylation of CD36 is necessary for trafficking of the protein to the cell surface but not for ligand recognition. There is redundancy in both the chemical nature and the pattern of glycosylation, suggesting that it does not directly induce the folding of the protein but, rather, may ensure contact with the appropriate chaperones within the secretory pathway. Furthermore, mutant CD36 with fewer glycosylation sites can fold, traffic to the cell surface, and bind ligand. These less complex forms of the protein may prove more suitable for structural studies.
Supplementary Material
Acknowledgments
We are grateful to Geeta Patel, Dinah Rahman, and Justin Lock of the MRC proteomics facility for Q-ToF MS and to Cleidiane Zampronio, Antony Jones, and Helen Cooper of the Functional Genomics Laboratory, Birmingham University, for FT-ICR MS (the Functional Genomics Laboratory is funded by Biotechnology and Biological Sciences Research Council, Swindon, United Kingdom Grant 6/JIF13209).
This work was supported by the Medical Research Council UK and also by British Heart Foundation Project Grant PG/03/044/15328 (to C. A. M. and K. J. L.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Fig. 1.
S. J. Hoosdally, E. J. Andress, C. Wooding, C. A. Martin, and K. J. Linton, unpublished data.
- LDL
- low density lipoprotein
- SRBI
- scavenger receptor class B, type I
- OG
- n-octyl-β-d-glucopyranoside
- PNGase F
- peptide N-glycosidase F
- Endo H
- endoglycosidase H
- Sf21
- Spodoptera frugiperda 21
- non-g
- non-glycosylated
- Ni-NTA
- nickel-nitrilotriacetic acid
- BSA
- bovine serum albumin
- FT-ICR
- Fourier transform ion cyclotron resonance
- MS
- mass spectroscopy
- FACS
- fluorescence-activated cell sorter
- PBS
- phosphate-buffered saline
- mAb
- monoclonal antibody
- Q-Tof
- quadrupole-time of flight.
REFERENCES
- 1.Tandon N. N., Kralisz U., Jamieson G. A. ( 1989) J. Biol. Chem. 264, 7576– 7583 [PubMed] [Google Scholar]
- 2.Greenwalt D. E., Watt K. W., So O. Y., Jiwani N. ( 1990) Biochemistry 29, 7054– 7059 [DOI] [PubMed] [Google Scholar]
- 3.Endemann G., Stanton L. W., Madden K. S., Bryant C. M., White R. T., Protter A. A. ( 1993) J. Biol. Chem. 268, 11811– 11816 [PubMed] [Google Scholar]
- 4.Greenwalt D. E., Lipsky R. H., Ockenhouse C. F., Ikeda H., Tandon N. N., Jamieson G. A. ( 1992) Blood 80, 1105– 1115 [PubMed] [Google Scholar]
- 5.Harmon C. M., Abumrad N. A. ( 1993) J. Membr. Biol. 133, 43– 49 [DOI] [PubMed] [Google Scholar]
- 6.Rutella S., Rumi C., Di Mario A., Leone G. ( 1997) Eur. J. Histochem. 41, Suppl. 2, 53– 54 [PubMed] [Google Scholar]
- 7.Ma X., Bacci S., Mlynarski W., Gottardo L., Soccio T., Menzaghi C., Iori E., Lager R. A., Shroff A. R., Gervino E. V., Nesto R. W., Johnstone M. T., Abumrad N. A., Avogaro A., Trischitta V., Doria A. ( 2004) Hum. Mol. Genet. 13, 2197– 2205 [DOI] [PubMed] [Google Scholar]
- 8.Febbraio M., Podrez E. A., Smith J. D., Hajjar D. P., Hazen S. L., Hoff H. F., Sharma K., Silverstein R. L. ( 2000) J. Clin. Invest. 105, 1049– 1056 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Serghides L., Smith T. G., Patel S. N., Kain K. C. ( 2003) Trends Parasitol. 19, 461– 469 [DOI] [PubMed] [Google Scholar]
- 10.Aitman T. J., Glazier A. M., Wallace C. A., Cooper L. D., Norsworthy P. J., Wahid F. N., Al-Majali K. M., Trembling P. M., Mann C. J., Shoulders C. C., Graf D., St Lezin E., Kurtz T. W., Kren V., Pravenec M., Ibrahimi A., Abumrad N. A., Stanton L. W., Scott J. ( 1999) Nat. Genet. 21, 76– 83 [DOI] [PubMed] [Google Scholar]
- 11.Love-Gregory L., Sherva R., Sun L., Wasson J., Schappe T., Doria A., Rao D. C., Hunt S. C., Klein S., Neuman R. J., Permutt M. A., Abumrad N. A. ( 2008) Hum. Mol. Genet. 17, 1695– 1704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yamashita S., Hirano K., Kuwasako T., Janabi M., Toyama Y., Ishigami M., Sakai N. ( 2007) Mol. Cell Biochem. 299, 19– 22 [DOI] [PubMed] [Google Scholar]
- 13.Abumrad N. A., el-Maghrabi M. R., Amri E. Z., Lopez E., Grimaldi P. A. ( 1993) J. Biol. Chem. 268, 17665– 17668 [PubMed] [Google Scholar]
- 14.Koonen D. P., Febbraio M., Bonnet S., Nagendran J., Young M. E., Michelakis E. D., Dyck J. R. ( 2007) Circulation 116, 2139– 2147 [DOI] [PubMed] [Google Scholar]
- 15.Hwang E. H., Taki J., Yasue S., Fujimoto M., Taniguchi M., Matsunari I., Nakajima K., Shiobara S., Ikeda T., Tonami N. ( 1998) J. Nucl. Med. 39, 1681– 1684 [PubMed] [Google Scholar]
- 16.Vega M. A., Seguí-Real B., García J. A., Calés C., Rodríguez F., Vanderkerckhove J., Sandoval I. V. ( 1991) J. Biol. Chem. 266, 16818– 16824 [PubMed] [Google Scholar]
- 17.Calvo D., Vega M. A. ( 1993) J. Biol. Chem. 268, 18929– 18935 [PubMed] [Google Scholar]
- 18.Acton S. L., Scherer P. E., Lodish H. F., Krieger M. ( 1994) J. Biol. Chem. 269, 21003– 21009 [PubMed] [Google Scholar]
- 19.Franc N. C., Dimarcq J. L., Lagueux M., Hoffmann J., Ezekowitz R. A. ( 1996) Immunity 4, 431– 443 [DOI] [PubMed] [Google Scholar]
- 20.Hart K., Wilcox M. ( 1993) J. Mol. Biol. 234, 249– 253 [DOI] [PubMed] [Google Scholar]
- 21.Oquendo P., Hundt E., Lawler J., Seed B. ( 1989) Cell 58, 95– 101 [DOI] [PubMed] [Google Scholar]
- 22.Gruarin P., Thorne R. F., Dorahy D. J., Burns G. F., Sitia R., Alessio M. ( 2000) Biochem. Biophys. Res. Commun. 275, 446– 454 [DOI] [PubMed] [Google Scholar]
- 23.Pearce S. F., Wu J., Silverstein R. L. ( 1994) Blood 84, 384– 389 [PubMed] [Google Scholar]
- 24.Asch A. S., Liu I., Briccetti F. M., Barnwell J. W., Kwakye-Berko F., Dokun A., Goldberger J., Pernambuco M. ( 1993) Science 262, 1436– 1440 [DOI] [PubMed] [Google Scholar]
- 25.Rasmussen J. T., Berglund L., Rasmussen M. S., Petersen T. E. ( 1998) Eur. J. Biochem. 257, 488– 494 [DOI] [PubMed] [Google Scholar]
- 26.Tao N., Wagner S. J., Lublin D. M. ( 1996) J. Biol. Chem. 271, 22315– 22320 [DOI] [PubMed] [Google Scholar]
- 27.Alessio M., Ghigo D., Garbarino G., Geuna M., Malavasi F. ( 1991) Cell. Immunol. 137, 487– 500 [DOI] [PubMed] [Google Scholar]
- 28.Freeze H. H. ( 2006) Nat. Rev. Genet. 7, 537– 551 [DOI] [PubMed] [Google Scholar]
- 29.O'Connor S. E., Imperiali B. ( 1996) Chem. Biol. 3, 803– 812 [DOI] [PubMed] [Google Scholar]
- 30.Helenius A. ( 1994) Mol. Biol. Cell 5, 253– 265 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mitra N., Sinha S., Ramya T. N., Surolia A. ( 2006) Trends Biochem. Sci. 31, 156– 163 [DOI] [PubMed] [Google Scholar]
- 32.Trombetta E. S., Helenius A. ( 1998) Curr. Opin. Struct. Biol. 8, 587– 592 [DOI] [PubMed] [Google Scholar]
- 33.Nufer O., Guldbrandsen S., Degen M., Kappeler F., Paccaud J. P., Tani K., Hauri H. P. ( 2002) J. Cell Sci. 115, 619– 628 [DOI] [PubMed] [Google Scholar]
- 34.Berglund L., Petersen T. E., Rasmussen J. T. ( 1996) Biochim. Biophys. Acta 1309, 63– 68 [DOI] [PubMed] [Google Scholar]
- 35.Viñals M., Xu S., Vasile E., Krieger M. ( 2003) J. Biol. Chem. 278, 5325– 5332 [DOI] [PubMed] [Google Scholar]
- 36.Martin C. A., Longman E., Wooding C., Hoosdally S. J., Ali S., Aitman T. J., Gutmann D. A., Freemont P. S., Byrne B., Linton K. J. ( 2007) Protein Sci. 16, 2531– 2541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Dixon P. H., Weerasekera N., Linton K. J., Donaldson O., Chambers J., Egginton E., Weaver J., Nelson-Piercy C., de Swiet M., Warnes G., Elias E., Higgins C. F., Johnston D. G., McCarthy M. I., Williamson C. ( 2000) Hum. Mol. Genet. 9, 1209– 1217 [DOI] [PubMed] [Google Scholar]
- 38.Gavin A. C., Bösche M., Krause R., Grandi P., Marzioch M., Bauer A., Schultz J., Rick J. M., Michon A. M., Cruciat C. M., Remor M., Höfert C., Schelder M., Brajenovic M., Ruffner H., Merino A., Klein K., Hudak M., Dickson D., Rudi T., Gnau V., Bauch A., Bastuck S., Huhse B., Leutwein C., Heurtier M. A., Copley R. R., Edelmann A., Querfurth E., Rybin V., Drewes G., Raida M., Bouwmeester T., Bork P., Seraphin B., Kuster B., Neubauer G., Superti-Furga G. ( 2002) Nature 415, 141– 147 [DOI] [PubMed] [Google Scholar]
- 39.Peterman S. M., Dufresne C. P., Horning S. ( 2005) J. Biomol. Tech. 16, 112– 124 [PMC free article] [PubMed] [Google Scholar]
- 40.Klinkenberg L. G., Webb T., Zitomer R. S. ( 2006) Eukaryot. Cell 5, 1007– 1017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hsieh P., Robbins P. W. ( 1984) J. Biol. Chem. 259, 2375– 2382 [PubMed] [Google Scholar]
- 42.Jarvis D. L. ( 2003) Virology 310, 1– 7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Schinkel A. H., Kemp S., Dollé M., Rudenko G., Wagenaar E. ( 1993) J. Biol. Chem. 268, 7474– 7481 [PubMed] [Google Scholar]
- 44.Kulkarni K. A., Srivastava A., Mitra N., Sharon N., Surolia A., Vijayan M., Suguna K. ( 2004) Proteins 56, 821– 827 [DOI] [PubMed] [Google Scholar]
- 45.Zhou F., Xu W., Hong M., Pan Z., Sinko P. J., Ma J., You G. ( 2005) Mol. Pharmacol. 67, 868– 876 [DOI] [PubMed] [Google Scholar]
- 46.Hebert D. N., Zhang J. X., Chen W., Foellmer B., Helenius A. ( 1997) J. Cell Biol. 139, 613– 623 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.