Abstract
Differentiation of the Drosophila compound eye from the eye imaginal disc is a progressive process: columns of cells successively differentiate in a posterior to anterior sequence, clusters of cells form at regularly spaced intervals within each column, and individual photoreceptors differentiate in a defined order within each cluster. The progression of differentiation across the eye disc is driven by a positive autoregulatory loop of expression of the secreted molecule Hedgehog, which is temporally delayed by the intercalation of a second signal, Spitz. Hedgehog refines the spatial position at which each column initiates its differentiation by inducing secondary signals that act over different ranges to control the expression of positive and negative regulators. The position of clusters within each column is controlled by secreted inhibitory signals from clusters in the preceding column, and a single founder neuron, R8, is singled out within each cluster by Notch-mediated lateral inhibition. R8 then sequentially recruits surrounding cells to differentiate by producing a short-range signal, Spitz, which induces a secondary short-range signal, Delta. Intrinsic transcription factors act in combination with these two signals to produce cell-type diversity within the ommatidium. The Hedgehog and Spitz signals are transported along the photoreceptor axons and reused within the brain as long-range and local cues to trigger the differentiation and assembly of target neurons.
Keywords: photoreceptor, autoregulatory loop, Hedgehog, Spitz, Delta
1. Introduction
The Drosophila compound eye is a highly organized structure that constitutes an excellent developmental system in which to address the molecular and cellular mechanisms of pattern formation (Wolff and Ready, 1993). The retina is composed of 750–800 identical units called ommatidia, which are organized into a regular hexagonally packed array. Each ommatidial unit contains eight photoreceptors (R1–R8), four cone cells and two primary pigment cells arranged in a stereotypic pattern, and these units are surrounded by a lattice of secondary and tertiary pigment cells. The photoreceptors project axons into the optic lobes of the brain, where they form retinotopic projections in two separate ganglia, the lamina and the medulla (Clandinin and Zipursky, 2002).
The transformation leading from an unpatterned epithelial monolayer within the larval eye imaginal disc to the highly ordered adult eye has been extensively studied during the three decades since it was first introduced as an experimental system (Ready et al., 1976). The eye disc is specified in the embryonic and early larval stages through the action of a network of transcription factors known as the retinal determination genes (Silver and Rebay, 2005). Pattern formation and ommatidial differentiation begin in the third larval instar with the appearance of a groove called the morphogenetic furrow (MF) at the posterior margin of the eye disc (Ready et al., 1976). This indentation in the epithelium, which results from an apical constriction and apical-basal contraction of the cells, sweeps progressively across the eye disc from posterior to anterior over a 2-day period. Cells anterior to the MF are undifferentiated and proliferate asynchronously, whereas cells posterior to the MF are organized into columns of regularly spaced clusters within which photoreceptor differentiation occurs in a defined sequence (Tomlinson and Ready, 1987, Wolff and Ready, 1991). An average of 30 columns, each initiated every 90–120 minutes, will form the entire retinal field. Unlike other imaginal discs, which are patterned by organizers formed at stable boundaries between cellular territories, the eye disc has a progressive pattern of differentiation controlled by signals that are constantly changing their spatial positions.
One critical signal driving the initiation and progression of the MF is the secreted protein Hedgehog (Hh) (Heberlein et al., 1995, Heberlein et al., 1993b, Ma et al., 1993). Hh is expressed at the posterior margin of the eye disc prior to MF initiation, and induces differentiation of anterior cells; as they differentiate into photoreceptors, these cells also begin to express hh and can therefore act on cells anterior to them (Heberlein and Moses, 1995). One target of Hh signaling is the Bone Morphogenetic Protein (BMP) family member Decapentaplegic (Dpp), which functions redundantly with Hh in MF progression (Burke and Basler, 1996, Curtiss and Mlodzik, 2000, Greenwood and Struhl, 1999, Heberlein et al., 1993b, Wiersdorff et al., 1996). Another Hh target is the basic Helix-Loop-Helix (bHLH) transcription factor Atonal (Ato), which is required for the specification of R8 cells, the first photoreceptors to differentiate in each cluster. Ato is initially expressed in a broad stripe just anterior to the MF, and its expression is gradually refined to single R8 cells within the MF in a process requiring lateral inhibition mediated by the Notch receptor (Baker and Zitron, 1995, Dokucu et al., 1996, Jarman et al., 1994, Jarman et al., 1995).
R8 orchestrates subsequent ommatidial development by recruiting surrounding uncommitted cells to differentiate into other photoreceptors, cone cells and pigment cells. Secretion of the Epidermal growth factor receptor (EGFR) ligand Spitz (Spi) from R8 and subsequently recruited cells promotes the sequential differentiation of photoreceptors R2/R5, R3/R4, R1/R6, R7, the cone cells and the primary pigment cells, as well as the survival of secondary and tertiary pigment cells (Freeman, 1996, Miller and Cagan, 1998). The Notch ligand Delta (Dl) acts as a critical secondary signal for several of the later differentiating cell types (Flores et al., 2000, Nagaraj and Banerjee, 2007, Tomlinson and Struhl, 2001, Tsuda et al., 2002). The founder role of R8 cells makes their specification and spacing critical for the organization of the adult eye. Both Hh and Spi are also transported along the photoreceptor axons and act on photoreceptor target cells in the brain (Huang and Kunes, 1996, Huang et al., 1998), coordinating the development of the two tissues. We will describe the current understanding of the mechanistic basis of pattern formation in this relatively simple system.
2. Progression of the morphogenetic furrow: a delayed autoregulatory loop
The MF initiates in the third larval instar at the dorso-ventral midpoint of the posterior margin of the eye disc. hh expression is activated at the center of the posterior margin in second instar eye discs (Cavodeassi et al., 1999, Cho et al., 2000) by members of the odd-skipped family of genes (Bras-Pereira et al., 2006). Notch, which is activated at the dorsoventral midline by asymmetric distribution of its ligands and the glycosyltransferase Fringe, also contributes to MF initiation, as do Dpp, the JAK/STAT pathway ligand Unpaired, and the EGFR (Burke and Basler, 1996, Cavodeassi et al., 1999, Cho and Choi, 1998, Dominguez and de Celis, 1998, Ekas et al., 2006, Kumar and Moses, 2001, Papayannopoulos et al., 1998, Tsai et al., 2007, Wiersdorff et al., 1996). The temporal control of initiation is not fully understood; it may be triggered by the hormone ecdysone (Niwa et al., 2004), or by growth of the eye disc that brings the posterior margin out of the range of anterior inhibitory signals (Kenyon et al., 2003, Ma and Moses, 1995, Treisman and Rubin, 1995).
While MF initiation is a unique developmental event, its progression occurs as a repeated sequence of events driven by an autoregulatory feedback loop. Cells that receive the Hh signal in and anterior to the MF are induced to differentiate as photoreceptors; as they differentiate posterior to the MF, these photoreceptors themselves begin to express hh, allowing them to drive the differentiation of more anterior cells (Dominguez and Hafen, 1997, Heberlein et al., 1995, Heberlein et al., 1993b, Ma et al., 1993, Strutt and Mlodzik, 1995). This cyclical induction of Hh during MF progression depends on an enhancer element of the hh gene that reproduces hh expression specifically in photoreceptor cells; deletions of this element result in loss of hh expression in the photoreceptors and arrest MF progression (Rogers et al., 2005). This element is directly regulated by Pointed (Pnt), the transcription factor downstream of EGFR signaling (O’Neill et al., 1994, Rogers et al., 2005). Cells that receive the Hh signal from photoreceptors posterior to them turn on atonal expression, leading to the specification of R8 cells. R8 then secretes Spi, which acts through the EGFR to activate Pnt in the neighboring cells. Pnt promotes the differentiation of these cells into photoreceptors (O’Neill et al., 1994), and simultaneously activates hh expression. This regulation of hh by Pnt, which is itself an indirect target of Hh signaling, creates a positive feedback loop that drives anterior propagation of hh expression and the MF (Figure 1).
Figure 1. A delayed autoregulatory loop of Hh expression drives MF progression.
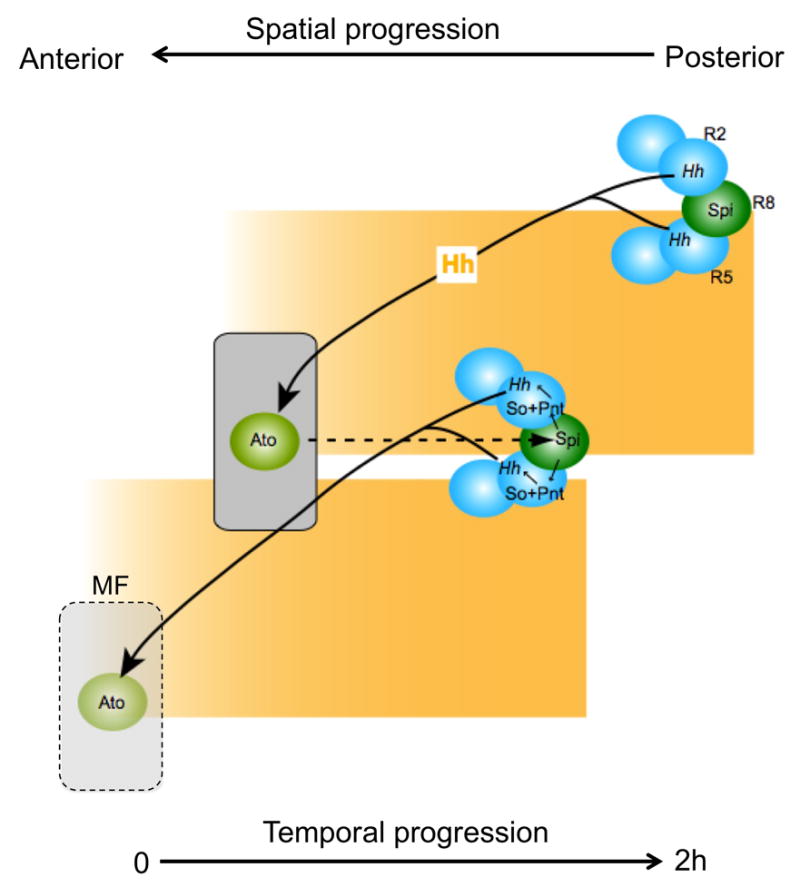
Hh is secreted by differentiating photoreceptors, primarily R2 and R5, and diffuses anteriorly to activate ato expression in the MF. Ato then promotes R8 specification and expression of the proteases Rho and Ru, which cleave the Spi precursor to produce active Spi. Spi acts locally to enhance the activity of the transcription factor Pnt, which together with So activates an eye-specific enhancer of the hh gene. The resulting Hh expression reiterates the cycle, which repeats approximately every 2 h in a spatial progression from posterior to anterior.
A temporal delay is introduced into this loop by the indirect effect of Hh on Spi production. An inactive precursor form of Spi is produced in all cells, but secretion of the active form requires the activities of the chaperone protein Star and the proteases Rhomboid (Rho) or Roughoid/Rhomboid-3 (Ru) (Freeman et al., 1992a, Shilo, 2003, Wasserman et al., 2000). rho and ru are required in the R8 cell (Wasserman et al., 2000), where their expression is regulated by Ato, a direct target of Hh signaling (Baonza et al., 2001, Dominguez, 1999). Activation of the hh enhancer by Pnt thus requires reception and transduction of the Hh signal, transcription and translation of Ato, transcription and translation of Rho and Ru, processing and secretion of Spi, and reception and transduction of the Spi signal. An additional level of regulation ensures that this mechanism is specific to the eye disc. Pnt binding sites alone are not sufficient to drive hh expression in photoreceptors (Rogers et al., 2005); the enhancer also contains essential binding sites for the retinal determination protein Sine oculis (So) (Pauli et al., 2005). So is specifically expressed in the eye field (Cheyette et al., 1994), restricting the control of hh expression by EGFR signaling to this developmental system.
Superimposed on this basic autoregulatory loop are a variety of other mechanisms that constrain the pace of MF progression. Several repressors of photoreceptor differentiation present in the region anterior to the MF are controlled by long-range and short-range ligands that are themselves targets of Hh signaling (Figure 2). Dpp, which is expressed in a stripe of cells within the MF in a Hh-dependent manner (Heberlein et al., 1993b, Masucci et al., 1990), acts at a long range to repress the expression of Homothorax (Hth), a transcription factor that prevents retinal differentiation in the anterior of the eye disc (Bessa et al., 2002). This repression of hth allows cells to enter a preproneural state in which they are able to respond to Hh. Dpp also activates the expression of hairy in a stripe anterior to the MF (Greenwood and Struhl, 1999); Hairy is a repressor of ato and acts in combination with another anteriorly expressed HLH protein, Extramacrochaetae, to prevent cells from initiating the differentiation process prematurely (Brown et al., 1995). Since Hth has been shown to repress hairy (Bessa et al., 2002), it is possible that the effect of Dpp on hairy is mediated indirectly through hth repression.
Figure 2. Hh induces long-range and short-range secondary signals that control the precise position of the MF.
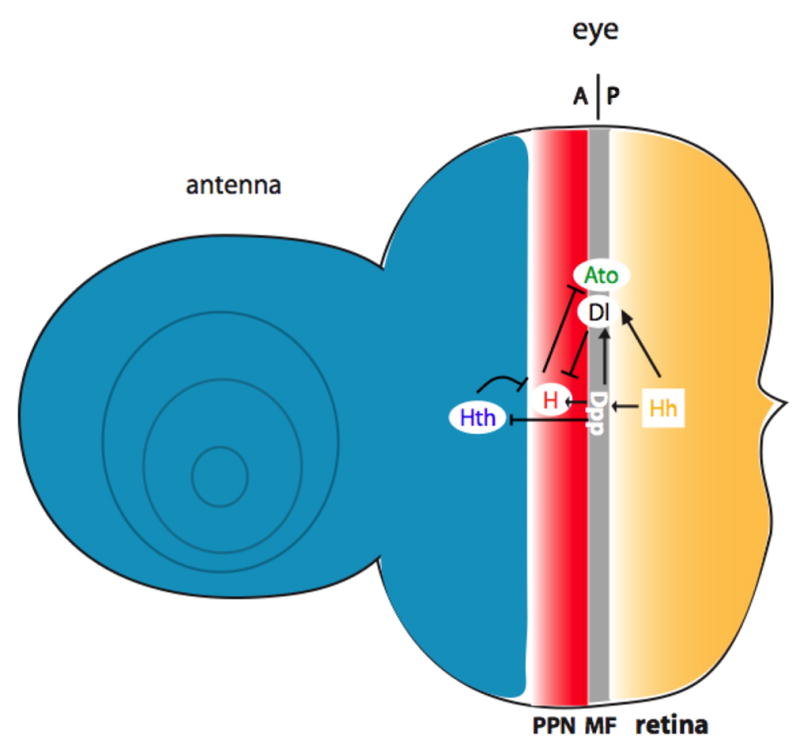
Hh acts over a short range to induce the expression of Dpp, which diffuses over a long range to turn off hth and turn on hairy, establishing a preproneural domain (PPN). Hh and Dpp also induce the expression of Delta, a transmembrane ligand that acts on adjacent cells to turn off hairy and allow ato expression, initiating photoreceptor differentiation.
Down-regulation of hairy expression, which relieves the repression of ato, requires a short-range signal that is provided by Delta (Dl), a transmembrane ligand for the Notch receptor expressed in the MF under the redundant control of Hh and Dpp (Baonza and Freeman, 2001, Baonza and Freeman, 2005, Parks et al., 1995). Misexpression of Dpp does not result in ectopic photoreceptor differentiation in most regions of the eye disc (Chanut and Heberlein, 1997, Pignoni and Zipursky, 1997); however, coexpression of Dpp with Dl can induce neural differentiation anywhere anterior to the MF, indicating that these signals are sufficient to induce a preproneural state that will progress to a proneural state (Baonza and Freeman, 2001). Through its activation of Dpp and Dl, Hh thus activates hairy expression at a long range and represses it at a shorter range, allowing precise spatial control of the initiation of ato expression. Hh also appears to be capable of repressing hairy by another mechanism in the absence of Notch signaling (Fu and Baker, 2003). The restriction of ato expression to single R8 photoreceptors is likewise Hh-dependent; Hh induces three negative regulators of ato expression, rough, Bar, and daughterless, in cells adjacent to R8 (Dokucu et al., 1996, Dominguez, 1999, Lim and Choi, 2003, Lim and Choi, 2004, Lim et al., 2008). The onset of expression of these genes is delayed relative to ato because like hh itself, they are indirectly induced by EGFR signaling (Dominguez et al., 1998, Lim and Choi, 2004).
It is interesting that the mechanisms controlling Hh signaling in the eye disc differ from those used in the wing disc. The transcription factor Cubitus interruptus (Ci) is processed into its repressor form (Ci75) in anterior wing disc cells that fail to receive the Hh signal, and represses hh expression in these cells (Aza-Blanc et al., 1997, Dominguez et al., 1996, Methot and Basler, 1999). This suggests that positive autoregulation through inhibition of Ci75 production is intrinsic to the Hh pathway. This mechanism must be actively suppressed in the cells at the anterior-posterior boundary of the wing disc, which receive Hh signaling but do not express hh (Apidianakis et al., 2001). However, Ci75 is not involved in hh autoregulation in the eye disc; ci mutant clones do not result in premature photoreceptor differentiation (Fu and Baker, 2003, Pappu et al., 2003), although Hh misexpression does (Heberlein et al., 1995), and ectopic Ci75 cannot repress hh in the eye disc (Lee et al., 2002). In the wing disc, Hh limits its own diffusion by upregulating the expression of Patched, a receptor that binds and sequesters Hh (Chen and Struhl, 1996). In the eye disc, Hh transport seems instead to be limited by apical constriction of cells in the MF, a shape change that is also Hh-dependent (Benlali et al., 2000, Corrigall et al., 2007, Escudero et al., 2007, Heberlein et al., 1995, Schlichting and Dahmann, 2008). Finally, Hh-expressing cells in the posterior compartment of the wing disc fail to respond to Hh signaling because the transcription factor Engrailed represses the expression of ci (Tabata et al., 1995). In the eye disc, responsiveness to Hh is downregulated posterior to the MF by degradation of the Ci protein following its ubiquitination by an SCF complex containing Cullin 3 and Roadkill/Hh-induced MATH and BTB-containing protein, the same mechanism used in cells receiving high levels of Hh at the anterior-posterior boundary of the wing disc (Kent et al., 2006, Ou et al., 2002, Ou et al., 2007, Zhang et al., 2006). The eye disc has integrated several secondary signals downstream of Hh to allow controlled movement of the boundary of Hh expression, which is stable throughout development in the wing disc.
3. Spacing of photoreceptor clusters is achieved through inhibitory mechanisms
To achieve a precisely ordered array of ommatidia in the adult eye, clusters within each column must initiate at regularly spaced intervals, and each column must also be offset from the preceding column. Each cluster initiates with the specification of its R8 cell, which depends on the proneural gene ato (Frankfort and Mardon, 2002). Tight control of ato expression is thus essential to guarantee the precision of the final lattice. ato is initially expressed in all cells in a stripe just ahead of the MF, and becomes restricted first to regularly spaced intermediate groups of about twenty cells, then to R8 equivalence groups of two to three cells, and finally to single cells that are the future R8 cells of each ommatidium (Figure 3)(Baker et al., 1996, Dokucu et al., 1996, Jarman et al., 1995).
Figure 3. Inhibitory signals control cluster spacing and R8 specification.
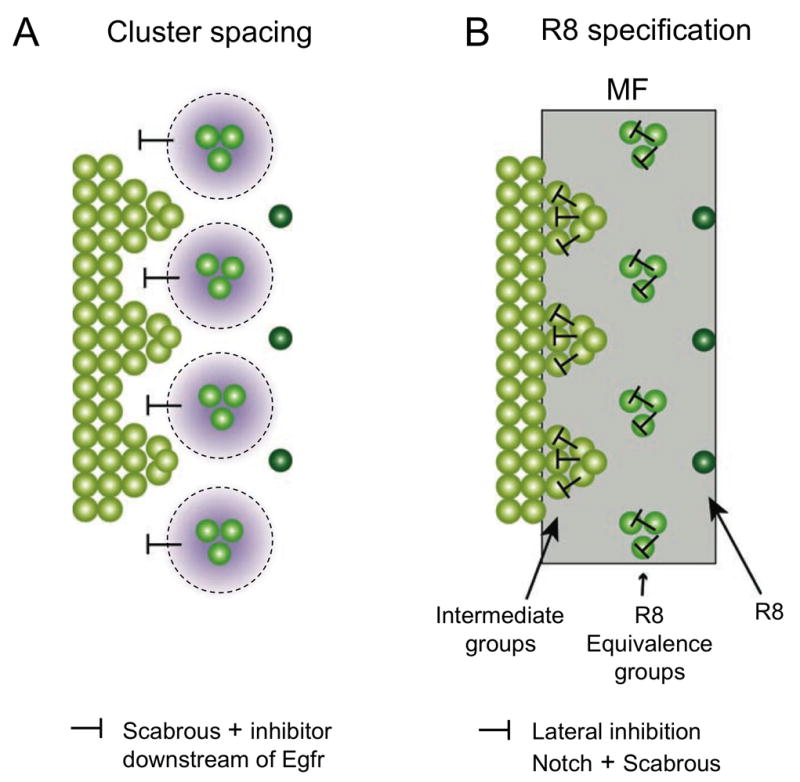
(A) shows that spacing of intermediate groups is controlled by inhibitory signals from clusters in the preceding column, including Sca and a second factor downstream of EGFR signaling. (B) shows the process of restriction of Ato expression within the MF from a continuous stripe of cells to intermediate groups, 3-cell R8 equivalence groups, and single R8 cells. R8 selection requires local lateral inhibition mediated by Notch and Sca.
Spacing of the intermediate groups is thought to involve an inhibitory signal from clusters in the preceding column that prevents new groups from forming directly anterior to existing clusters (Figure 3A). The secreted glycoprotein Scabrous (Sca), which is expressed in a subset of cells in the intermediate groups and reaches its highest level in R8 (Baker et al., 1990, Baker et al., 1996, Baker and Zitron, 1995, Mlodzik et al., 1990a), is a candidate for this signal. Sca secreted by R8 cells in posterior clusters is thought to diffuse anteriorly to repress ato expression between the forming intermediate groups; in sca mutants, R8 cells form too close together (Baker and Zitron, 1995, Lee et al., 1996). The receptor for this activity of Sca is unknown. Although Sca interacts with Notch in other contexts (Baker and Zitron, 1995, Li et al., 2003, Powell et al., 2001), intermediate group spacing is unaffected by inactivation of Notch (Baker and Zitron, 1995, Lee et al., 1996). The formation of intermediate groups coincides with organization of cells into clusters, in part because Ato activates the transcription of DE-cadherin (Baker and Yu, 1998, Brown et al., 2006). Sca has been shown to alter DE-Cadherin localization and cell adhesion in the notum (Renaud and Simpson, 2001), suggesting another possible mechanism for its effect on intermediate group spacing.
There may be another signal important for intermediate group spacing that is induced by EGFR signaling. Some studies have shown that EGFR mutant clones have a non-autonomous effect on spacing, suggesting that EGFR affects the secretion of a secondary factor involved in the restriction of ato expression (Chen and Chien, 1999, Spencer et al., 1998). Since Ato activates rho expression (Baonza et al., 2001), leading to secretion of Spi and activation of the EGFR pathway, this mechanism would function as a delayed feedback inhibitory loop that transmits spacing information from one column to the next, ensuring that successive columns are out of phase. The secreted signal involved has not been identified. One candidate is Argos (Aos), which is expressed in response to EGFR signaling and diffuses over a long range to bind the EGFR ligand Spi and antagonize its activity (Klein et al., 2004, Schweitzer et al., 1995, Spencer et al., 1998). However, aos mutant clones show normal spacing unless the clones are very large (Baonza et al., 2001, Yang and Baker, 2001), indicating that Aos does not transmit precise positional information about the location of each cluster. The molecule downstream of EGFR signaling must act in parallel to Sca in spacing intermediate groups, as EGFR sca double mutant clones have a more severe spacing defect than either single mutant (Baonza et al., 2001). Since sca expression also depends on ato (Lee et al., 1996), both signals would operate through the same regulatory logic. However, the role of EGFR in the spacing process has been disputed (Rodrigues et al., 2005).
Restriction of ato expression to a single R8 precursor cell within each intermediate group depends on lateral inhibition through the Notch receptor. The process of lateral inhibition has been well described during embryonic neurogenesis (Campos-Ortega and Jan, 1991). Slight differences between neighboring cells in the level of expression of the Notch ligand Dl are amplified because cells that receive a stronger Notch signal express less Dl. This process allows a single cell to be selected as the Dl-expressing neuron from a group of cells with equal potential (Bray, 1998). This mechanism appears to be conserved in the eye; when either Notch or Dl is inactivated using a temperature-sensitive allele, all cells in the intermediate groups continue to express ato and differentiate into extra R8 cells (Baker et al., 1996, Baker and Yu, 1998, Baker and Zitron, 1995, Cagan and Ready, 1989a, Frankfort and Mardon, 2002) (Figure 3B). This change from a positive effect of Notch and Dl on the early stripe of ato expression, described in section 2, to a negative effect on ato expression in the intermediate groups coincides with a switch in the enhancer element used to regulate ato (Baker et al., 1996, Baker and Yu, 1997, Sun et al., 1998). The 5′ enhancer element used in the intermediate groups requires autoregulation by Ato, and Notch signaling appears to interfere with this autoregulatory process (Baker et al., 1996, Sun et al., 1998).
Precise spacing of R8 cells could not be achieved if R8 were randomly selected within the proneural cluster, suggesting that the choice must be biased in some way. However, examination of Dl expression did not reveal obvious differences between cells in the intermediate groups (Baker and Yu, 1998). One possible source of such a bias is Sca, which is secreted by the R8 precursor and can interact with Notch to sustain Notch signaling in surrounding cells (Baker and Zitron, 1995, Li et al., 2003, Powell et al., 2001). Sca appears to act within endosomes, perhaps preventing endocytic down-regulation of Notch (Li et al., 2003). Another factor restricting the choice of R8 is Rough, a transcription factor that represses ato expression and is expressed in R2, 5, 3 and 4 under the control of EGFR signaling (Dokucu et al., 1996, Dominguez et al., 1998). Rough is excluded from the R8 precursor due to repression by Senseless, a transcription factor induced by Ato to lock in the R8 fate (Frankfort et al., 2001). Specification of R8 thus proceeds sequentially; the pattern of intermediate groups is imposed by signals from the preceding column, and a single R8 is selected within each group by biased lateral inhibition.
4. Sequential differentiation within each cluster depends on signaling range
Spatial and temporal patterning of cluster formation is achieved by controlling the differentiation of a single photoreceptor in each cluster, R8. R8 then initiates the sequential recruitment of the other ommatidial cell types. First R2/R5 and then R3/R4 are recruited to complete the five-cell precluster; the surrounding undifferentiated cells undergo a final cell division, the second mitotic wave, before the remaining cells are recruited in the order R1/R6, R7, anterior and posterior cone cells, and equatorial and polar cone cells (Tomlinson and Ready, 1987, Wolff and Ready, 1993) (Figure 4). During pupal development, two primary pigment cells are recruited to surround each ommatidium, and the lattice of secondary and tertiary pigment cells is formed by death of the surplus cells (Cagan and Ready, 1989b). All these cell fates except R8 are directly or indirectly dependent on Spi signaling through the EGFR. Mosaic clones lacking spi or EGFR function contain only R8 cells (Dominguez et al., 1998, Freeman, 1994, Tio et al., 1994), and activation of the pathway can induce additional recruitment of all cell types except R8 (Dominguez et al., 1998, Freeman, 1996).
Figure 4. Sequential recruitment of ommatidial cells is controlled by the short-range ligands Spi and Dl.
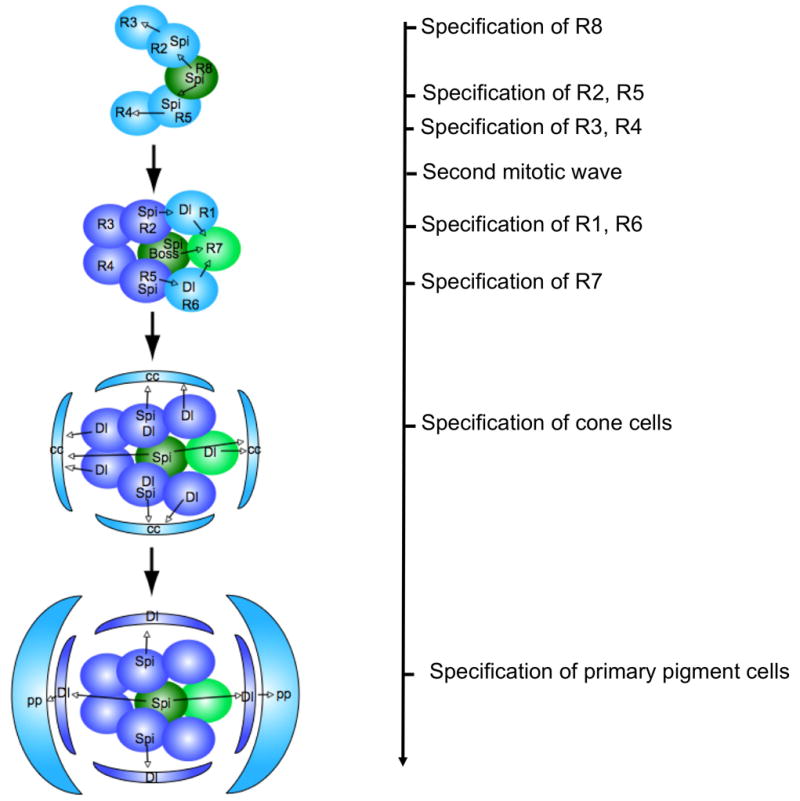
Spi promotes the differentiation of photoreceptors R2, 5, 3, 4, 1 and 6. It also induces the expression of Dl, which acts together with Spi and Boss to promote R7 differentiation, together with Spi to promote cone cell differentiation, and alone to promote primary pigment cell differentiation. Cells are added sequentially because Spi and Dl can only act over a short distance.
The dependence of all cell types on the same inductive signal raises the question of how their differentiation is ordered in a precise temporal sequence. This seems to be due in part to the short range of action of the ligand Spi. Spi is initially secreted by R8 due to Ato-dependent expression of the proteases Rho and Ru (Baonza et al., 2001, Wasserman et al., 2000). Spi then induces the cells immediately neighboring R8 to differentiate into R2 and R5. These cells also express Rho, under the control of the transcription factor Rough (Freeman et al., 1992a), as well as the chaperone protein Star (Heberlein et al., 1993a), and can therefore produce Spi to induce the differentiation of their neighbors in the arc-shaped cluster into R3 and R4 (Freeman, 1997, Wolff and Ready, 1991). It has been proposed that an autoregulatory loop expands the expression of Rho and Ru to additional photoreceptors as they differentiate, allowing subsequent waves of recruitment (Freeman, 1997, Shilo, 2005). However, mosaic analysis has shown little evidence of a requirement for rho, ru, Star or spi in any photoreceptors other than R8, 2 and 5 (Freeman, 1994, Heberlein and Rubin, 1991, Tio et al., 1994, Wasserman et al., 2000). Indeed, R1, 6 and 7 differentiate adjacent to R8, 2 and 5 (Tomlinson and Ready, 1987), suggesting that they are responding to Spi produced by these cells. Spi secreted from other photoreceptors may nevertheless contribute to the recruitment of cone and pigment cells, which were not examined in the mosaic studies.
One reason for the sequential, rather than simultaneous, recruitment of photoreceptors is likely to be the very short range of Spi action, which is limited to 1–2 cell diameters by several mechanisms (Miura et al., 2006, Schlesinger et al., 2004, Wasserman et al., 2000). Palmitoylation of the cleaved extracellular domain of Spi tethers it to the plasma membrane, restricting its diffusion (Miura et al., 2006). Small wing, a phospholipase Cγ, prevents the secretion of any Spi that is cleaved in the endoplasmic reticulum by Ru (Schlesinger et al., 2004). Finally, Spi signaling induces the expression of a secreted feedback inhibitor, Aos (Golembo et al., 1996), which binds to Spi and prevents it from binding to the EGFR (Klein et al., 2004). aos mutant eyes contain extra photoreceptors, cone cells and pigment cells (Freeman et al., 1992b). Aos may prevent Spi from reaching distant cells, restricting its activity to the region where its local concentration exceeds that of Aos (Freeman, 1997). Although mosaic studies suggest that Aos can act over a range of about 10 cell diameters (Freeman et al., 1992b), mathematical modeling of Aos function in the embryo indicates that Aos could exert a long-range effect by acting as a ligand sink, without itself diffusing over a long distance (Reeves et al., 2005). Interestingly, a second negative feedback loop involving the intracellular inhibitor Sprouty subsequently turns off aos expression in R8, 2 and 5; aos is maintained in other photoreceptors because sprouty expression is inhibited by the transcription factor Seven-up (Svp) expressed in these cells (Iwanami et al., 2005). This reduction in Aos production may allow Spi levels to increase sufficiently to induce the later differentiating cell types.
The use of Spi signaling to recruit all ommatidial cell types raises the question of how diversity is generated among these cells. Cell types are not simply determined by the level of EGFR activity, since ectopic activation of the EGFR pathway induces different cell types at different stages of development (Freeman, 1996, Hayashi and Saigo, 2001). One possible explanation is that Spi signaling acts in combination with other developmental signals. For example, R7 differentiation requires the activity of a second receptor tyrosine kinase, Sevenless (Sev), as well as the EGFR (Zipursky and Rubin, 1994). Bride of sevenless (Boss), the ligand for Sev, is a transmembrane protein specifically expressed in R8, which acts on the adjacent undifferentiated cell to induce its differentiation into R7 (Zipursky and Rubin, 1994). The combination of Sev and EGFR signaling is thought to increase signal transduction through their common downstream pathway, resulting in high-level expression of target genes such as prospero (Freeman, 1996, Tio and Moses, 1997, Xu et al., 2000).
Another signal that interacts with Spi to promote cell fate diversity is the Notch ligand Dl (Figure 4). Notch and Dl are required for the differentiation of R7, cone cells and primary pigment cells (Flores et al., 2000, Nagaraj and Banerjee, 2007, Tomlinson and Struhl, 2001, Tsuda et al., 2002). Dl is itself a target of EGFR signaling, which induces Dl transcription by promoting nuclear export and degradation of a corepressor for the Suppressor of Hairless transcription factor (Tsuda et al., 2002). This relationship between the two signals introduces another temporally delayed autoregulatory loop. Spi signaling induces R1-6 to differentiate and to express Dl; Dl subsequently acts in combination with Spi to induce differentiation of R7 and the cone cells (Flores et al., 2000, Tomlinson and Struhl, 2001, Tsuda et al., 2002). As EGFR activation in the cone cells increases during the pupal stages, they too express Dl, which is sufficient to induce differentiation of adjacent cells into primary pigment cells (Daga et al., 1996, Nagaraj and Banerjee, 2007). These cells require EGFR signaling only indirectly as an activator of Dl expression in cone cells (Nagaraj and Banerjee, 2007). Dl is a transmembrane molecule that can only signal to immediately adjacent cells (Artavanis-Tsakonas et al., 1999); this limit on its range of action again contributes to the ordered sequence of cell recruitment.
A third contribution to cell fate diversity comes from intrinsic properties of the responding cells. The transcription factor Lozenge (Lz) is expressed in undifferentiated cells posterior to the MF and in cells that differentiate after the second mitotic wave: R1, R6, R7, and the cone and pigment cells (Flores et al., 1998). Its misexpression in R3 and R4 transforms these cells into R7 cells, due to its positive effect on R7-specific genes such as prospero and its negative effect on genes such as seven-up that promote the identity of R3 and R4 (Daga et al., 1996, Flores et al., 1998, Xu et al., 2000). Lz also positively regulates the Bar genes, which are expressed in R1 and R6 (Daga et al., 1996, Higashijima et al., 1992). Expression of lz itself requires the transcription factor Glass, which is expressed in all cells posterior to the MF, and the retinal determination protein So, but it is unknown how lz is excluded from R8, 2, 5, 3 and 4 (Moses and Rubin, 1991, Yan et al., 2003). Other transcription factors expressed in different subsets of photoreceptors include Rough (R2, 5, 3, 4), Spalt (R3, 4), and Svp (R3, 4, 1, 6) (Barrio et al., 1999, Kimmel et al., 1990, Mlodzik et al., 1990b). These specific expression patterns are likely to result from similar combinatorial control mechanisms as well as cross-regulatory interactions (Hayashi and Saigo, 2001, Heberlein et al., 1991). In summary, differentiation of distinct cell types in an ordered sequence appears to be due to the combination of temporally delayed autoregulatory loops of the short-range ligands Spi and Dl with intrinsic prepatterning information in the form of transcription factor expression.
5. Photoreceptors coordinate the differentiation of their target cells in the lamina
As photoreceptor cells progressively differentiate in the eye disc, they extend axons into the brain, where R1-6 terminate in the lamina and R7 and R8 project through the lamina to terminate in the medulla (Clandinin and Zipursky, 2002). Within the lamina, the bundle of photoreceptor axons from each ommatidium associates with a cartridge composed of five lamina neurons, and these fascicles terminate in a retinotopic pattern, recreating a map of the visual field in the brain (Clandinin and Zipursky, 2002). Along the anterior-posterior axis, this organization arises because signals from the photoreceptor axons induce the formation of their target cells. Hh, which drives the propagation of the MF within the eye disc, is also transported along the photoreceptor axons and released to induce the final division of lamina precursor cells (LPCs) and their expression of the differentiation marker Dachshund (Dac) (Huang and Kunes, 1996) (Figure 5). Interestingly, this axonal transport requires a targeting signal that lies within the C-terminal protease domain of Hh (Chu et al., 2006), although LPCs respond to the N-terminal secreted domain through the canonical Hh signaling pathway (Huang and Kunes, 1998). Hh also induces the expression of the transcription factor Single-minded, which directs developing LPCs to associate with photoreceptor axons (Umetsu et al., 2006). Each column of ommatidia in the eye disc thus induces the formation of a corresponding column of target cells, allowing progressive posterior to anterior differentiation to be coordinated between the eye and the brain.
Figure 5. Hh and Spi signals are transported down the photoreceptor axons to organize the target region.
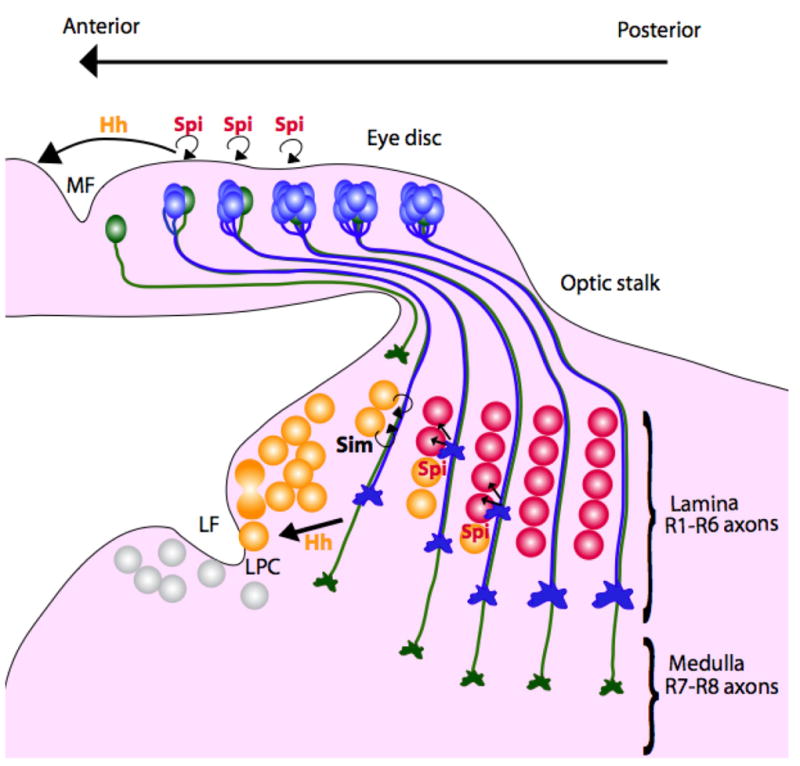
Hh promotes the final division of lamina precursor cells (LPC) posterior to the lamina furrow (LF), and through Sim, their association with photoreceptor axons. Hh also induces the LPCs to express Dac, which activates expression of the EGFR, allowing them to respond to Spi. Cells that respond to Hh by expressing Dac and EGFR are colored yellow. Spi then promotes LPC differentiation adjacent to retinal axons. Cells that respond to Spi by expressing Elav are colored red. Release of the same signals from the cell body and the axons coordinates morphogenesis of the eye and the lamina.
In addition, Hh acts through Dac to induce LPCs to express the EGFR, making them responsive to a second signal transported down the axons (Chotard et al., 2005, Huang et al., 1998). Spi, the signaling molecule that recruits photoreceptors to each ommatidial cluster, is also essential to direct the assembly of each lamina cartridge (Huang et al., 1998). Spi is necessary and sufficient for the neuronal differentiation, indicated by Elav expression, of the five lamina neurons associated with each ommatidial fascicle (Huang et al., 1998). The very short-range activity of Spi presumably ensures that lamina neurons differentiate only in the immediate vicinity of retinal axons. Coordination of eye and brain development achieved by using the same signals in both tissues is likely to be important to establish the spatial precision necessary for accurate vision.
6. Conclusions
The progressive development of identical units within a monolayer epithelium makes the Drosophila eye disc an excellent system in which to address the regulatory mechanisms of pattern formation. Positive autoregulatory loops play an important role in driving the progression of differentiation, and a temporal delay can be introduced into such loops by requiring the production of a second signal dependent on the first. The range over which signals are distributed is also an important factor in controlling the sequence of differentiation. Although columns differentiate sequentially, they can also interact with each other through secreted signals that determine the placement of clusters. Finally, the long processes formed by neurons allow them to act on different target cell populations by releasing the same signals from their cell bodies and axon terminals. Several aspects of this differentiation process have now been described at a level at which mathematical modeling could be applied to test and extend our understanding of the mechanisms involved.
Acknowledgments
Funding for the authors’ work on pattern formation in the eye disc is provided by the National Institutes of Health (EY13777). The manuscript was improved by the critical comments of Ines Carrera, Kevin Legent and Josefa Steinhauer.
References
- APIDIANAKIS Y, GRBAVEC D, STIFANI S, DELIDAKIS C. Groucho mediates a ci-independent mechanism of hedgehog repression in the anterior wing pouch. Development. 2001;128:4361–70. doi: 10.1242/dev.128.21.4361. [DOI] [PubMed] [Google Scholar]
- ARTAVANIS-TSAKONAS S, RAND MD, LAKE RJ. Notch signaling: Cell fate control and signal integration in development. Science. 1999;284:770–6. doi: 10.1126/science.284.5415.770. [DOI] [PubMed] [Google Scholar]
- AZA-BLANC P, RAMIREZ-WEBER FA, LAGET MP, SCHWARTZ C, KORNBERG TB. Proteolysis that is inhibited by Hedgehog targets Cubitus interruptus protein to the nucleus and converts it to a repressor. Cell. 1997;89:1043–53. doi: 10.1016/s0092-8674(00)80292-5. [DOI] [PubMed] [Google Scholar]
- BAKER NE, MLODZIK M, RUBIN GM. Spacing differentiation in the developing Drosophila eye: A fibrinogen-related lateral inhibitor encoded by scabrous. Science. 1990;250:1370–7. doi: 10.1126/science.2175046. [DOI] [PubMed] [Google Scholar]
- BAKER NE, YU S, HAN D. Evolution of proneural atonal expression during distinct regulatory phases in the developing Drosophila eye. Curr Biol. 1996;6:1290–301. doi: 10.1016/s0960-9822(02)70715-x. [DOI] [PubMed] [Google Scholar]
- BAKER NE, YU SY. Proneural function of neurogenic genes in the developing Drosophila eye. Curr Biol. 1997;7:122–32. doi: 10.1016/s0960-9822(06)00056-x. [DOI] [PubMed] [Google Scholar]
- BAKER NE, YU SY. The R8-photoreceptor equivalence group in Drosophila: Fate choice precedes regulated Delta transcription and is independent of Notch gene dose. Mech Dev. 1998;74:3–14. doi: 10.1016/s0925-4773(98)00054-9. [DOI] [PubMed] [Google Scholar]
- BAKER NE, ZITRON AE. Drosophila eye development: Notch and Delta amplify a neurogenic pattern conferred on the morphogenetic furrow by Scabrous. Mech Dev. 1995;49:173–89. doi: 10.1016/0925-4773(94)00314-d. [DOI] [PubMed] [Google Scholar]
- BAONZA A, CASCI T, FREEMAN M. A primary role for the epidermal growth factor receptor in ommatidial spacing in the Drosophila eye. Curr Biol. 2001;11:396–404. doi: 10.1016/s0960-9822(01)00125-7. [DOI] [PubMed] [Google Scholar]
- BAONZA A, FREEMAN M. Notch signalling and the initiation of neural development in the Drosophila eye. Development. 2001;128:3889–98. doi: 10.1242/dev.128.20.3889. [DOI] [PubMed] [Google Scholar]
- BAONZA A, FREEMAN M. Control of cell proliferation in the Drosophila eye by Notch signaling. Dev Cell. 2005;8:529–39. doi: 10.1016/j.devcel.2005.01.019. [DOI] [PubMed] [Google Scholar]
- BARRIO R, DE CELIS JF, BOLSHAKOV S, KAFATOS FC. Identification of regulatory regions driving the expression of the Drosophila spalt complex at different developmental stages. Dev Biol. 1999;215:33–47. doi: 10.1006/dbio.1999.9434. [DOI] [PubMed] [Google Scholar]
- BENLALI A, DRASKOVIC I, HAZELETT DJ, TREISMAN JE. Act up controls actin polymerization to alter cell shape and restrict Hedgehog signaling in the Drosophila eye disc. Cell. 2000;101:271–81. doi: 10.1016/s0092-8674(00)80837-5. [DOI] [PubMed] [Google Scholar]
- BESSA J, GEBELEIN B, PICHAUD F, CASARES F, MANN RS. Combinatorial control of Drosophila eye development by eyeless, homothorax, and teashirt. Genes Dev. 2002;16:2415–27. doi: 10.1101/gad.1009002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- BRAS-PEREIRA C, BESSA J, CASARES F. odd-skipped genes specify the signaling center that triggers retinogenesis in Drosophila. Development. 2006;133:4145–9. doi: 10.1242/dev.02593. [DOI] [PubMed] [Google Scholar]
- BRAY S. Notch signalling in Drosophila: Three ways to use a pathway. Semin Cell Dev Biol. 1998;9:591–7. doi: 10.1006/scdb.1998.0262. [DOI] [PubMed] [Google Scholar]
- BROWN KE, BAONZA A, FREEMAN M. Epithelial cell adhesion in the developing Drosophila retina is regulated by Atonal and the EGF receptor pathway. Dev Biol. 2006;300:710–21. doi: 10.1016/j.ydbio.2006.08.003. [DOI] [PubMed] [Google Scholar]
- BROWN NL, SATTLER CA, PADDOCK SW, CARROLL SB. hairy and emc negatively regulate morphogenetic furrow progression in the Drosophila eye. Cell. 1995;80:879–87. doi: 10.1016/0092-8674(95)90291-0. [DOI] [PubMed] [Google Scholar]
- BURKE R, BASLER K. Hedgehog-dependent patterning in the Drosophila eye can occur in the absence of Dpp signaling. Dev Biol. 1996;179:360–8. doi: 10.1006/dbio.1996.0267. [DOI] [PubMed] [Google Scholar]
- CAGAN RL, READY DF. Notch is required for successive cell decisions in the developing Drosophila retina. Genes Dev. 1989a;3:1099–112. doi: 10.1101/gad.3.8.1099. [DOI] [PubMed] [Google Scholar]
- CAGAN RL, READY DF. The emergence of order in the Drosophila pupal retina. Dev Biol. 1989b;136:346–62. doi: 10.1016/0012-1606(89)90261-3. [DOI] [PubMed] [Google Scholar]
- CAMPOS-ORTEGA JA, JAN YN. Genetic and molecular bases of neurogenesis in Drosophila melanogaster. Annu Rev Neurosci. 1991;14:399–420. doi: 10.1146/annurev.ne.14.030191.002151. [DOI] [PubMed] [Google Scholar]
- CAVODEASSI F, DIEZ DEL CORRAL R, CAMPUZANO S, DOMINGUEZ M. Compartments and organising boundaries in the Drosophila eye: The role of the homeodomain Iroquois proteins. Development. 1999;126:4933–42. doi: 10.1242/dev.126.22.4933. [DOI] [PubMed] [Google Scholar]
- CHANUT F, HEBERLEIN U. Role of decapentaplegic in initiation and progression of the morphogenetic furrow in the developing Drosophila retina. Development. 1997;124:559–67. doi: 10.1242/dev.124.2.559. [DOI] [PubMed] [Google Scholar]
- CHEN CK, CHIEN CT. Negative regulation of atonal in proneural cluster formation of Drosophila R8 photoreceptors. Proc Natl Acad Sci U S A. 1999;96:5055–60. doi: 10.1073/pnas.96.9.5055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CHEN Y, STRUHL G. Dual roles for Patched in sequestering and transducing Hedgehog. Cell. 1996;87:553–63. doi: 10.1016/s0092-8674(00)81374-4. [DOI] [PubMed] [Google Scholar]
- CHEYETTE BN, GREEN PJ, MARTIN K, GARREN H, HARTENSTEIN V, ZIPURSKY SL. The Drosophila sine oculis locus encodes a homeodomain-containing protein required for the development of the entire visual system. Neuron. 1994;12:977–96. doi: 10.1016/0896-6273(94)90308-5. [DOI] [PubMed] [Google Scholar]
- CHO KO, CHERN J, IZADDOOST S, CHOI KW. Novel signaling from the peripodial membrane is essential for eye disc patterning in Drosophila. Cell. 2000;103:331–42. doi: 10.1016/s0092-8674(00)00124-0. [DOI] [PubMed] [Google Scholar]
- CHO KO, CHOI KW. Fringe is essential for mirror symmetry and morphogenesis in the Drosophila eye. Nature. 1998;396:272–276. doi: 10.1038/24394. [DOI] [PubMed] [Google Scholar]
- CHOTARD C, LEUNG W, SALECKER I. Glial cells missing and Gcm2 cell autonomously regulate both glial and neuronal development in the visual system of Drosophila. Neuron. 2005;48:237–51. doi: 10.1016/j.neuron.2005.09.019. [DOI] [PubMed] [Google Scholar]
- CHU T, CHIU M, ZHANG E, KUNES S. A C-terminal motif targets Hedgehog to axons, coordinating assembly of the Drosophila eye and brain. Dev Cell. 2006;10:635–46. doi: 10.1016/j.devcel.2006.03.003. [DOI] [PubMed] [Google Scholar]
- CLANDININ TR, ZIPURSKY SL. Making connections in the fly visual system. Neuron. 2002;35:827–41. doi: 10.1016/s0896-6273(02)00876-0. [DOI] [PubMed] [Google Scholar]
- CORRIGALL D, WALTHER RF, RODRIGUEZ L, FICHELSON P, PICHAUD F. Hedgehog signaling is a principal inducer of Myosin-II-driven cell ingression in Drosophila epithelia. Dev Cell. 2007;13:730–42. doi: 10.1016/j.devcel.2007.09.015. [DOI] [PubMed] [Google Scholar]
- CURTISS J, MLODZIK M. Morphogenetic furrow initiation and progression during eye development in Drosophila: The roles of decapentaplegic, hedgehog and eyes absent. Development. 2000;127:1325–36. doi: 10.1242/dev.127.6.1325. [DOI] [PubMed] [Google Scholar]
- DAGA A, KARLOVICH CA, DUMSTREI K, BANERJEE U. Patterning of cells in the Drosophila eye by Lozenge, which shares homologous domains with AML1. Genes Dev. 1996;10:1194–205. doi: 10.1101/gad.10.10.1194. [DOI] [PubMed] [Google Scholar]
- DOKUCU ME, ZIPURSKY SL, CAGAN RL. Atonal, Rough and the resolution of proneural clusters in the developing Drosophila retina. Development. 1996;122:4139–47. doi: 10.1242/dev.122.12.4139. [DOI] [PubMed] [Google Scholar]
- DOMINGUEZ M. Dual role for hedgehog in the regulation of the proneural gene atonal during ommatidia development. Development. 1999;126:2345–53. doi: 10.1242/dev.126.11.2345. [DOI] [PubMed] [Google Scholar]
- DOMINGUEZ M, BRUNNER M, HAFEN E, BASLER K. Sending and receiving the Hedgehog signal: Control by the Drosophila Gli protein Cubitus interruptus. Science. 1996;272:1621–5. doi: 10.1126/science.272.5268.1621. [DOI] [PubMed] [Google Scholar]
- DOMINGUEZ M, DE CELIS JF. A dorsal/ventral boundary established by Notch controls growth and polarity in the Drosophila eye. Nature. 1998;396:276–8. doi: 10.1038/24402. [DOI] [PubMed] [Google Scholar]
- DOMINGUEZ M, HAFEN E. Hedgehog directly controls initiation and propagation of retinal differentiation in the Drosophila eye. Genes Dev. 1997;11:3254–64. doi: 10.1101/gad.11.23.3254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DOMINGUEZ M, WASSERMAN JD, FREEMAN M. Multiple functions of the EGF receptor in Drosophila eye development. Curr Biol. 1998;8:1039–48. doi: 10.1016/s0960-9822(98)70441-5. [DOI] [PubMed] [Google Scholar]
- EKAS LA, BAEG GH, FLAHERTY MS, AYALA-CAMARGO A, BACH EA. JAK/STAT signaling promotes regional specification by negatively regulating wingless expression in Drosophila. Development. 2006;133:4721–9. doi: 10.1242/dev.02675. [DOI] [PubMed] [Google Scholar]
- ESCUDERO LM, BISCHOFF M, FREEMAN M. Myosin II regulates complex cellular arrangement and epithelial architecture in Drosophila. Dev Cell. 2007;13:717–29. doi: 10.1016/j.devcel.2007.09.002. [DOI] [PubMed] [Google Scholar]
- FLORES GV, DAGA A, KALHOR HR, BANERJEE U. Lozenge is expressed in pluripotent precursor cells and patterns multiple cell types in the Drosophila eye through the control of cell-specific transcription factors. Development. 1998;125:3681–7. doi: 10.1242/dev.125.18.3681. [DOI] [PubMed] [Google Scholar]
- FLORES GV, DUAN H, YAN H, NAGARAJ R, FU W, ZOU Y, NOLL M, BANERJEE U. Combinatorial signaling in the specification of unique cell fates. Cell. 2000;103:75–85. doi: 10.1016/s0092-8674(00)00106-9. [DOI] [PubMed] [Google Scholar]
- FRANKFORT BJ, MARDON G. R8 development in the Drosophila eye: A paradigm for neural selection and differentiation. Development. 2002;129:1295–306. doi: 10.1242/dev.129.6.1295. [DOI] [PubMed] [Google Scholar]
- FRANKFORT BJ, NOLO R, ZHANG Z, BELLEN H, MARDON G. Senseless repression of rough is required for R8 photoreceptor differentiation in the developing Drosophila eye. Neuron. 2001;32:403–14. doi: 10.1016/s0896-6273(01)00480-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- FREEMAN M. The spitz gene is required for photoreceptor determination in the Drosophila eye where it interacts with the EGF receptor. Mech Dev. 1994;48:25–33. doi: 10.1016/0925-4773(94)90003-5. [DOI] [PubMed] [Google Scholar]
- FREEMAN M. Reiterative use of the EGF receptor triggers differentiation of all cell types in the Drosophila eye. Cell. 1996;87:651–60. doi: 10.1016/s0092-8674(00)81385-9. [DOI] [PubMed] [Google Scholar]
- FREEMAN M. Cell determination strategies in the Drosophila eye. Development. 1997;124:261–70. doi: 10.1242/dev.124.2.261. [DOI] [PubMed] [Google Scholar]
- FREEMAN M, KIMMEL BE, RUBIN GM. Identifying targets of the rough homeobox gene of Drosophila: Evidence that rhomboid functions in eye development. Development. 1992a;116:335–46. doi: 10.1242/dev.116.2.335. [DOI] [PubMed] [Google Scholar]
- FREEMAN M, KLAMBT C, GOODMAN CS, RUBIN GM. The argos gene encodes a diffusible factor that regulates cell fate decisions in the Drosophila eye. Cell. 1992b;69:963–75. doi: 10.1016/0092-8674(92)90615-j. [DOI] [PubMed] [Google Scholar]
- FU W, BAKER NE. Deciphering synergistic and redundant roles of hedgehog, decapentaplegic and Delta that drive the wave of differentiation in Drosophila eye development. Development. 2003;130:5229–39. doi: 10.1242/dev.00764. [DOI] [PubMed] [Google Scholar]
- GOLEMBO M, SCHWEITZER R, FREEMAN M, SHILO BZ. argos transcription is induced by the Drosophila EGF receptor pathway to form an inhibitory feedback loop. Development. 1996;122:223–30. doi: 10.1242/dev.122.1.223. [DOI] [PubMed] [Google Scholar]
- GREENWOOD S, STRUHL G. Progression of the morphogenetic furrow in the Drosophila eye: The roles of hedgehog, decapentaplegic and the raf pathway. Development. 1999;126:5795–808. doi: 10.1242/dev.126.24.5795. [DOI] [PubMed] [Google Scholar]
- HAYASHI T, SAIGO K. Diversification of cell types in the Drosophila eye by differential expression of prepattern genes. Mech Dev. 2001;108:13–27. doi: 10.1016/s0925-4773(01)00466-x. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, HARIHARAN IK, RUBIN GM. Star is required for neuronal differentiation in the Drosophila retina and displays dosage-sensitive interactions with ras1. Dev Biol. 1993a;160:51–63. doi: 10.1006/dbio.1993.1285. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, MLODZIK M, RUBIN GM. Cell-fate determination in the developing Drosophila eye: Role of the rough gene. Development. 1991;112:703–12. doi: 10.1242/dev.112.3.703. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, MOSES K. Mechanisms of Drosophila retinal morphogenesis: The virtues of being progressive. Cell. 1995;81:987–90. doi: 10.1016/s0092-8674(05)80003-0. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, RUBIN GM. Star is required in a subset of photoreceptor cells in the developing Drosophila retina and displays dosage sensitive interactions with rough. Dev Biol. 1991;144:353–61. doi: 10.1016/0012-1606(91)90427-5. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, SINGH CM, LUK AY, DONOHOE TJ. Growth and differentiation in the Drosophila eye coordinated by hedgehog. Nature. 1995;373:709–11. doi: 10.1038/373709a0. [DOI] [PubMed] [Google Scholar]
- HEBERLEIN U, WOLFF T, RUBIN GM. The TGF beta homolog dpp and the segment polarity gene hedgehog are required for propagation of a morphogenetic wave in the Drosophila retina. Cell. 1993b;75:913–26. doi: 10.1016/0092-8674(93)90535-x. [DOI] [PubMed] [Google Scholar]
- HIGASHIJIMA S, KOJIMA T, MICHIUE T, ISHIMARU S, EMORI Y, SAIGO K. Dual Bar homeo box genes of Drosophila required in two photoreceptor cells, R1 and R6, and primary pigment cells for normal eye development. Genes Dev. 1992;6:50–60. doi: 10.1101/gad.6.1.50. [DOI] [PubMed] [Google Scholar]
- HUANG Z, KUNES S. Hedgehog, transmitted along retinal axons, triggers neurogenesis in the developing visual centers of the Drosophila brain. Cell. 1996;86:411–22. doi: 10.1016/s0092-8674(00)80114-2. [DOI] [PubMed] [Google Scholar]
- HUANG Z, KUNES S. Signals transmitted along retinal axons in Drosophila: Hedgehog signal reception and the cell circuitry of lamina cartridge assembly. Development. 1998;125:3753–64. doi: 10.1242/dev.125.19.3753. [DOI] [PubMed] [Google Scholar]
- HUANG Z, SHILO BZ, KUNES S. A retinal axon fascicle uses Spitz, an EGF receptor ligand, to construct a synaptic cartridge in the brain of Drosophila. Cell. 1998;95:693–703. doi: 10.1016/s0092-8674(00)81639-6. [DOI] [PubMed] [Google Scholar]
- IWANAMI M, HIROMI Y, OKABE M. Cell-type specific utilization of multiple negative feedback loops generates developmental constancy. Genes Cells. 2005;10:743–52. doi: 10.1111/j.1365-2443.2005.00871.x. [DOI] [PubMed] [Google Scholar]
- JARMAN AP, GRELL EH, ACKERMAN L, JAN LY, JAN YN. atonal is the proneural gene for Drosophila photoreceptors. Nature. 1994;369:398–400. doi: 10.1038/369398a0. [DOI] [PubMed] [Google Scholar]
- JARMAN AP, SUN Y, JAN LY, JAN YN. Role of the proneural gene, atonal, in formation of Drosophila chordotonal organs and photoreceptors. Development. 1995;121:2019–30. doi: 10.1242/dev.121.7.2019. [DOI] [PubMed] [Google Scholar]
- KENT D, BUSH EW, HOOPER JE. Roadkill attenuates Hedgehog responses through degradation of Cubitus interruptus. Development. 2006;133:2001–10. doi: 10.1242/dev.02370. [DOI] [PubMed] [Google Scholar]
- KENYON KL, RANADE SS, CURTISS J, MLODZIK M, PIGNONI F. Coordinating proliferation and tissue specification to promote regional identity in the Drosophila head. Dev Cell. 2003;5:403–14. doi: 10.1016/s1534-5807(03)00243-0. [DOI] [PubMed] [Google Scholar]
- KIMMEL BE, HEBERLEIN U, RUBIN GM. The homeo domain protein Rough is expressed in a subset of cells in the developing Drosophila eye where it can specify photoreceptor cell subtype. Genes Dev. 1990;4:712–27. doi: 10.1101/gad.4.5.712. [DOI] [PubMed] [Google Scholar]
- KLEIN DE, NAPPI VM, REEVES GT, SHVARTSMAN SY, LEMMON MA. Argos inhibits epidermal growth factor receptor signalling by ligand sequestration. Nature. 2004;430:1040–4. doi: 10.1038/nature02840. [DOI] [PubMed] [Google Scholar]
- KUMAR JP, MOSES K. The EGF receptor and Notch signaling pathways control the initiation of the morphogenetic furrow during Drosophila eye development. Development. 2001;128:2689–97. doi: 10.1242/dev.128.14.2689. [DOI] [PubMed] [Google Scholar]
- LEE EC, HU X, YU SY, BAKER NE. The scabrous gene encodes a secreted glycoprotein dimer and regulates proneural development in Drosophila eyes. Mol Cell Biol. 1996;16:1179–88. doi: 10.1128/mcb.16.3.1179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LEE JD, AMANAI K, SHEARN A, TREISMAN JE. The ubiquitin ligase Hyperplastic discs negatively regulates hedgehog and decapentaplegic expression by independent mechanisms. Development. 2002;129:5697–706. doi: 10.1242/dev.00159. [DOI] [PubMed] [Google Scholar]
- LI Y, FETCHKO M, LAI ZC, BAKER NE. Scabrous and gp150 are endosomal proteins that regulate Notch activity. Development. 2003;130:2819–27. doi: 10.1242/dev.00495. [DOI] [PubMed] [Google Scholar]
- LIM J, CHOI KW. Bar homeodomain proteins are anti-proneural in the Drosophila eye: Transcriptional repression of atonal by Bar prevents ectopic retinal neurogenesis. Development. 2003;130:5965–74. doi: 10.1242/dev.00818. [DOI] [PubMed] [Google Scholar]
- LIM J, CHOI KW. Induction and autoregulation of the anti-proneural gene Bar during retinal neurogenesis in Drosophila. Development. 2004;131:5573–80. doi: 10.1242/dev.01426. [DOI] [PubMed] [Google Scholar]
- LIM J, JAFAR-NEJAD H, HSU YC, CHOI KW. Novel function of the class I bHLH protein Daughterless in the negative regulation of proneural gene expression in the Drosophila eye. EMBO Rep. 2008 doi: 10.1038/embor.2008.166. (epub ahead of print) [DOI] [PMC free article] [PubMed] [Google Scholar]
- MA C, MOSES K. Wingless and Patched are negative regulators of the morphogenetic furrow and can affect tissue polarity in the developing Drosophila compound eye. Development. 1995;121:2279–89. doi: 10.1242/dev.121.8.2279. [DOI] [PubMed] [Google Scholar]
- MA C, ZHOU Y, BEACHY PA, MOSES K. The segment polarity gene hedgehog is required for progression of the morphogenetic furrow in the developing Drosophila eye. Cell. 1993;75:927–38. doi: 10.1016/0092-8674(93)90536-y. [DOI] [PubMed] [Google Scholar]
- MASUCCI JD, MILTENBERGER RJ, HOFFMANN FM. Pattern-specific expression of the Drosophila decapentaplegic gene in imaginal disks is regulated by 3′ cis-regulatory elements. Genes Dev. 1990;4:2011–23. doi: 10.1101/gad.4.11.2011. [DOI] [PubMed] [Google Scholar]
- METHOT N, BASLER K. Hedgehog controls limb development by regulating the activities of distinct transcriptional activator and repressor forms of Cubitus interruptus. Cell. 1999;96:819–31. doi: 10.1016/s0092-8674(00)80592-9. [DOI] [PubMed] [Google Scholar]
- MILLER DT, CAGAN RL. Local induction of patterning and programmed cell death in the developing Drosophila retina. Development. 1998;125:2327–35. doi: 10.1242/dev.125.12.2327. [DOI] [PubMed] [Google Scholar]
- MIURA GI, BUGLINO J, ALVARADO D, LEMMON MA, RESH MD, TREISMAN JE. Palmitoylation of the EGFR ligand Spitz by Rasp increases Spitz activity by restricting its diffusion. Dev Cell. 2006;10:167–76. doi: 10.1016/j.devcel.2005.11.017. [DOI] [PubMed] [Google Scholar]
- MLODZIK M, BAKER NE, RUBIN GM. Isolation and expression of scabrous, a gene regulating neurogenesis in Drosophila. Genes Dev. 1990a;4:1848–61. doi: 10.1101/gad.4.11.1848. [DOI] [PubMed] [Google Scholar]
- MLODZIK M, HIROMI Y, WEBER U, GOODMAN CS, RUBIN GM. The Drosophila seven-up gene, a member of the steroid receptor gene superfamily, controls photoreceptor cell fates. Cell. 1990b;60:211–24. doi: 10.1016/0092-8674(90)90737-y. [DOI] [PubMed] [Google Scholar]
- MOSES K, RUBIN GM. glass encodes a site-specific DNA-binding protein that is regulated in response to positional signals in the developing Drosophila eye. Genes Dev. 1991;5:583–93. doi: 10.1101/gad.5.4.583. [DOI] [PubMed] [Google Scholar]
- NAGARAJ R, BANERJEE U. Combinatorial signaling in the specification of primary pigment cells in the Drosophila eye. Development. 2007;134:825–31. doi: 10.1242/dev.02788. [DOI] [PubMed] [Google Scholar]
- NIWA N, HIROMI Y, OKABE M. A conserved developmental program for sensory organ formation in Drosophila melanogaster. Nat Genet. 2004;36:293–7. doi: 10.1038/ng1308. [DOI] [PubMed] [Google Scholar]
- O’NEILL EM, REBAY I, TJIAN R, RUBIN GM. The activities of two Ets-related transcription factors required for Drosophila eye development are modulated by the Ras/MAPK pathway. Cell. 1994;78:137–47. doi: 10.1016/0092-8674(94)90580-0. [DOI] [PubMed] [Google Scholar]
- OU CY, LIN YF, CHEN YJ, CHIEN CT. Distinct protein degradation mechanisms mediated by Cul1 and Cul3 controlling Ci stability in Drosophila eye development. Genes Dev. 2002;16:2403–14. doi: 10.1101/gad.1011402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- OU CY, WANG CH, JIANG J, CHIEN CT. Suppression of Hedgehog signaling by Cul3 ligases in proliferation control of retinal precursors. Dev Biol. 2007;308:106–19. doi: 10.1016/j.ydbio.2007.05.008. [DOI] [PubMed] [Google Scholar]
- PAPAYANNOPOULOS V, TOMLINSON A, PANIN VM, RAUSKOLB C, IRVINE KD. Dorsal-ventral signaling in the Drosophila eye. Science. 1998;281:2031–4. doi: 10.1126/science.281.5385.2031. [DOI] [PubMed] [Google Scholar]
- PAPPU KS, CHEN R, MIDDLEBROOKS BW, WOO C, HEBERLEIN U, MARDON G. Mechanism of Hedgehog signaling during Drosophila eye development. Development. 2003;130:3053–62. doi: 10.1242/dev.00534. [DOI] [PubMed] [Google Scholar]
- PARKS AL, TURNER FR, MUSKAVITCH MA. Relationships between complex Delta expression and the specification of retinal cell fates during Drosophila eye development. Mech Dev. 1995;50:201–16. doi: 10.1016/0925-4773(94)00336-l. [DOI] [PubMed] [Google Scholar]
- PAULI T, SEIMIYA M, BLANCO J, GEHRING WJ. Identification of functional Sine oculis motifs in the autoregulatory element of its own gene, in the eyeless enhancer and in the signalling gene hedgehog. Development. 2005;132:2771–82. doi: 10.1242/dev.01841. [DOI] [PubMed] [Google Scholar]
- PIGNONI F, ZIPURSKY SL. Induction of Drosophila eye development by decapentaplegic. Development. 1997;124:271–8. doi: 10.1242/dev.124.2.271. [DOI] [PubMed] [Google Scholar]
- POWELL PA, WESLEY C, SPENCER S, CAGAN RL. Scabrous complexes with Notch to mediate boundary formation. Nature. 2001;409:626–30. doi: 10.1038/35054566. [DOI] [PubMed] [Google Scholar]
- READY DF, HANSON TE, BENZER S. Development of the Drosophila retina, a neurocrystalline lattice. Dev Biol. 1976;53:217–40. doi: 10.1016/0012-1606(76)90225-6. [DOI] [PubMed] [Google Scholar]
- REEVES GT, KALIFA R, KLEIN DE, LEMMON MA, SHVARTSMAN SY. Computational analysis of EGFR inhibition by Argos. Dev Biol. 2005;284:523–35. doi: 10.1016/j.ydbio.2005.05.013. [DOI] [PubMed] [Google Scholar]
- RENAUD O, SIMPSON P. Scabrous modifies epithelial cell adhesion and extends the range of lateral signalling during development of the spaced bristle pattern in Drosophila. Dev Biol. 2001;240:361–76. doi: 10.1006/dbio.2001.0482. [DOI] [PubMed] [Google Scholar]
- RODRIGUES AB, WERNER E, MOSES K. Genetic and biochemical analysis of the role of EGFR in the morphogenetic furrow of the developing Drosophila eye. Development. 2005;132:4697–707. doi: 10.1242/dev.02058. [DOI] [PubMed] [Google Scholar]
- ROGERS EM, BRENNAN CA, MORTIMER NT, COOK S, MORRIS AR, MOSES K. Pointed regulates an eye-specific transcriptional enhancer in the Drosophila hedgehog gene, which is required for the movement of the morphogenetic furrow. Development. 2005;132:4833–43. doi: 10.1242/dev.02061. [DOI] [PubMed] [Google Scholar]
- SCHLESINGER A, KIGER A, PERRIMON N, SHILO BZ. Small wing PLCgamma is required for ER retention of cleaved Spitz during eye development in Drosophila. Dev Cell. 2004;7:535–45. doi: 10.1016/j.devcel.2004.09.001. [DOI] [PubMed] [Google Scholar]
- SCHLICHTING K, DAHMANN C. Hedgehog and Dpp signaling induce cadherin Cad86c expression in the morphogenetic furrow during Drosophila eye development. Mech Dev. 2008;125:712–28. doi: 10.1016/j.mod.2008.04.005. [DOI] [PubMed] [Google Scholar]
- SCHWEITZER R, HOWES R, SMITH R, SHILO BZ, FREEMAN M. Inhibition of Drosophila EGF receptor activation by the secreted protein Argos. Nature. 1995;376:699–702. doi: 10.1038/376699a0. [DOI] [PubMed] [Google Scholar]
- SHILO BZ. Signaling by the Drosophila Epidermal growth factor receptor pathway during development. Exp Cell Res. 2003;284:140–9. doi: 10.1016/s0014-4827(02)00094-0. [DOI] [PubMed] [Google Scholar]
- SHILO BZ. Regulating the dynamics of EGF receptor signaling in space and time. Development. 2005;132:4017–27. doi: 10.1242/dev.02006. [DOI] [PubMed] [Google Scholar]
- SILVER SJ, REBAY I. Signaling circuitries in development: Insights from the retinal determination gene network. Development. 2005;132:3–13. doi: 10.1242/dev.01539. [DOI] [PubMed] [Google Scholar]
- SPENCER SA, POWELL PA, MILLER DT, CAGAN RL. Regulation of EGF receptor signaling establishes pattern across the developing Drosophila retina. Development. 1998;125:4777–90. doi: 10.1242/dev.125.23.4777. [DOI] [PubMed] [Google Scholar]
- STRUTT DI, MLODZIK M. Ommatidial polarity in the Drosophila eye is determined by the direction of furrow progression and local interactions. Development. 1995;121:4247–56. doi: 10.1242/dev.121.12.4247. [DOI] [PubMed] [Google Scholar]
- SUN Y, JAN LY, JAN YN. Transcriptional regulation of atonal during development of the Drosophila peripheral nervous system. Development. 1998;125:3731–40. doi: 10.1242/dev.125.18.3731. [DOI] [PubMed] [Google Scholar]
- TABATA T, SCHWARTZ C, GUSTAVSON E, ALI Z, KORNBERG TB. Creating a Drosophila wing de novo, the role of engrailed, and the compartment border hypothesis. Development. 1995;121:3359–69. doi: 10.1242/dev.121.10.3359. [DOI] [PubMed] [Google Scholar]
- TIO M, MA C, MOSES K. Spitz, a Drosophila homolog of Transforming growth factor-alpha, is required in the founding photoreceptor cells of the compound eye facets. Mech Dev. 1994;48:13–23. doi: 10.1016/0925-4773(94)90002-7. [DOI] [PubMed] [Google Scholar]
- TIO M, MOSES K. The Drosophila TGF alpha homolog Spitz acts in photoreceptor recruitment in the developing retina. Development. 1997;124:343–51. doi: 10.1242/dev.124.2.343. [DOI] [PubMed] [Google Scholar]
- TOMLINSON A, READY DF. Neuronal differentiation in the Drosophila ommatidium. Dev Biol. 1987;120:366–376. doi: 10.1016/0012-1606(87)90239-9. [DOI] [PubMed] [Google Scholar]
- TOMLINSON A, STRUHL G. Delta/Notch and Boss/Sevenless signals act combinatorially to specify the Drosophila R7 photoreceptor. Mol Cell. 2001;7:487–95. doi: 10.1016/s1097-2765(01)00196-4. [DOI] [PubMed] [Google Scholar]
- TREISMAN JE, RUBIN GM. Wingless inhibits morphogenetic furrow movement in the Drosophila eye disc. Development. 1995;121:3519–27. doi: 10.1242/dev.121.11.3519. [DOI] [PubMed] [Google Scholar]
- TSAI YC, YAO JG, CHEN PH, POSAKONY JW, BAROLO S, KIM J, SUN YH. Upd/JAK/STAT signaling represses wg transcription to allow initiation of morphogenetic furrow in Drosophila eye development. Dev Biol. 2007;306:760–71. doi: 10.1016/j.ydbio.2007.04.011. [DOI] [PubMed] [Google Scholar]
- TSUDA L, NAGARAJ R, ZIPURSKY SL, BANERJEE U. An EGFR/Ebi/Sno pathway promotes Delta expression by inactivating Su(H)/SMRTER repression during inductive Notch signaling. Cell. 2002;110:625–37. doi: 10.1016/s0092-8674(02)00875-9. [DOI] [PubMed] [Google Scholar]
- UMETSU D, MURAKAMI S, SATO M, TABATA T. The highly ordered assembly of retinal axons and their synaptic partners is regulated by Hedgehog/Single-minded in the Drosophila visual system. Development. 2006;133:791–800. doi: 10.1242/dev.02253. [DOI] [PubMed] [Google Scholar]
- WASSERMAN JD, URBAN S, FREEMAN M. A family of rhomboid-like genes: Drosophila rhomboid-1 and roughoid/rhomboid-3 cooperate to activate EGF receptor signaling. Genes Dev. 2000;14:1651–63. [PMC free article] [PubMed] [Google Scholar]
- WIERSDORFF V, LECUIT T, COHEN SM, MLODZIK M. Mad acts downstream of Dpp receptors, revealing a differential requirement for Dpp signaling in initiation and propagation of morphogenesis in the Drosophila eye. Development. 1996;122:2153–62. doi: 10.1242/dev.122.7.2153. [DOI] [PubMed] [Google Scholar]
- WOLFF T, READY DF. The beginning of pattern formation in the Drosophila compound eye: The morphogenetic furrow and the second mitotic wave. Development. 1991;113:841–50. doi: 10.1242/dev.113.3.841. [DOI] [PubMed] [Google Scholar]
- WOLFF T, READY DF. Pattern formation in the Drosophila retina. In: BATE M, MARTINEZ-ARIAS A, editors. The development of Drosophila melanogaster. II. Cold Spring Harbor Laboratory Press, Cold Spring Harbor; 1993. pp. 1277–1316. [Google Scholar]
- XU C, KAUFFMANN RC, ZHANG J, KLADNY S, CARTHEW RW. Overlapping activators and repressors delimit transcriptional response to receptor tyrosine kinase signals in the Drosophila eye. Cell. 2000;103:87–97. doi: 10.1016/s0092-8674(00)00107-0. [DOI] [PubMed] [Google Scholar]
- YAN H, CANON J, BANERJEE U. A transcriptional chain linking eye specification to terminal determination of cone cells in the Drosophila eye. Dev Biol. 2003;263:323–9. doi: 10.1016/j.ydbio.2003.08.003. [DOI] [PubMed] [Google Scholar]
- YANG L, BAKER NE. Role of the EGFR/Ras/Raf pathway in specification of photoreceptor cells in the Drosophila retina. Development. 2001;128:1183–91. doi: 10.1242/dev.128.7.1183. [DOI] [PubMed] [Google Scholar]
- ZHANG Q, ZHANG L, WANG B, OU CY, CHIEN CT, JIANG J. A Hedgehog-induced BTB protein modulates Hedgehog signaling by degrading Ci/Gli transcription factor. Dev Cell. 2006;10:719–29. doi: 10.1016/j.devcel.2006.05.004. [DOI] [PubMed] [Google Scholar]
- ZIPURSKY SL, RUBIN GM. Determination of neuronal cell fate: Lessons from the R7 neuron of Drosophila. Annu Rev Neurosci. 1994;17:373–97. doi: 10.1146/annurev.ne.17.030194.002105. [DOI] [PubMed] [Google Scholar]


