Abstract
We developed and validated a new method to create automated 3D parametric surface models of the lateral ventricles in brain MRI scans, providing an efficient approach to monitor degenerative disease in clinical studies and drug trials. First, we used a set of parameterized surfaces to represent the ventricles in four subjects’ manually labeled brain MRI scans (atlases). We fluidly registered each atlas and mesh model to MRIs from 17 Alzheimer’s disease (AD) patients and 13 age-and gender-matched healthy elderly control subjects, and 18 asymptomatic ApoE4-carriers and 18 age- and gender-matched non-carriers. We examined genotyped healthy subjects with the goal of detecting subtle effects of a gene that confers heightened risk for Alzheimer’s disease. We averaged the meshes extracted for each 3D MR data set, and combined the automated segmentations with a radial mapping approach to localize ventricular shape differences in patients. Validation experiments comparing automated and expert manual segmentations showed that (1) the Hausdorff labeling error rapidly decreased, and (2) the power to detect disease- and gene-related alterations improved, as the number of atlases, N, was increased from 1 to 9. In surface-based statistical maps, we detected more widespread and intense anatomical deficits as we increased the number of atlases. We formulated a statistical stopping criterion to determine the optimal number of atlases to use. Healthy ApoE4-carriers and those with AD showed local ventricular abnormalities. This high-throughput method for morphometric studies further motivates the combination of genetic and neuroimaging strategies in predicting AD progression and treatment response.
Introduction
The ventricular system is a CSF-filled structure in the center of the brain, surrounded by gray and white matter structures whose volume and integrity are affected by neurodegenerative diseases (Powell et al., 1991). Ventricular changes reflect atrophy in surrounding structures, so ventricular measures and surface-based maps provide sensitive assessments of tissue reduction that correlate with cognitive deterioration in illnesses such as Alzheimer’s disease (AD) (Thompson et al., 2004; Carmichael et al., 2006), HIV/AIDS (Thompson et al., 2006) and schizophrenia (Styner et al., 2005; Narr et al., 2001, 2004), and offer a potential approach to evaluate disease progression in large-scale drug trials. In particular, published studies over the past 20 years have found that brain ventricle volume is higher in AD patients than in age-matched healthy controls, and ventricular volume has been proposed as a useful biomarker of disease progression (Thal et al., 2006; Carmichael et al., 2007a,b).
Despite years of efforts in automated surface parameterization, the concave shape, complex branching topology (Wang et al., 2005; Ferrarini et al., 2006) and extreme narrowness of the inferior and posterior horns have made it difficult for voxel-based classifiers to create accurate models of the ventricles. It is also difficult for surface parameterization approaches to impose a grid on the entire structure (Wang et al., 2005). In the engineering literature, automated ventricular segmentations often appear with the posterior or inferior horns omitted (Zhu and Jiang, 2004). This is a serious problem as the inferior horns are sensitive to the early degenerative changes in AD. Robust shape models are required to gauge these changes. For the vast 3D MR data sets now being collected (e.g., 339 subjects in Carmichael et al., 2006 and over 3000 scans in the Alzheimer's Disease Neuroimaging Initiative; http://www.loni.ucla.edu/ADNI/), manual segmentations would be impractical due to the time and expertise required; more automated methods are urgently needed.
One promising segmentation approach nonlinearly registers each brain MR volume to a target image volume in which the structures of interest have been labeled and parameterized; the labels are then projected onto the unlabeled data using the computed geometric transformations (Collins et al., 1994; Dawant et al., 1999; Fischl et al., 2002; Svarer et al., 2005; Heckemann et al., 2006). However, the chosen reference image volume influences the results of label propagation; different starting atlases create different segmentations for each subject. In Rohlfing et al. (2003, 2004) and Klein et al. (2005) multiple segmentations were combined, bringing the result closer to manual labels than if only one was used. "Targetless" image warping also combines multiple pairwise or groupwise registrations, reducing registration errors in groups of image volumes (Kochunov et al., 2005; Cootes et al., 2005; Zöllei et al., 2005; Babalola et al., 2006). A related effort in anatomical template construction uses multiple image registrations to create a mean anatomic template that is least distant from a group of anatomies in a strictly defined mathematical sense. Such mean templates may be defined using deformation vector field averaging (Collins et al., 1994; Christensen et al., 2006), by minimizing the mean strain tensors of deformation mappings to the template (Leporé et al., 2007), or by minimizing the total geodesic length of the deformations aligning each individual image volume to the template in the group of diffeomorphisms (Miller, 2004; Lorenzen et al., 2006). Each of these methods has been successful, but has limitations. Deformable atlases that rely on a single atlas template are limited by the unavoidable bias in selecting a specific individual scan as a template, with its unique contrast and geometry. This can make the results depend on the template used (as we show in this paper). Other methods have combined image volumes using multiple registrations. Although this minimizes bias, the resulting template commonly does not provide surface-based anatomical segmentations for the individuals mapped to it, or if it does, it suffers the limitation that a single registration deforms the segmentation in the atlas onto the individual. Surface-based segmentations may also provide additional benefit relative to voxel-based methods that examine image volumes and 3D deformations, as graphical models and maps, such as measures of radial thickness, may be derived from the surface geometry.
In this study, we validated a new automated technique for 3D segmentation that combines multiple registration-based surface models into one. We show that this increases the label propagation accuracy and the power to detect disease effects, without requiring any interactive human input other than the initial expert labeling of a small set of image volumes. Combining segmentations is not a new idea, but the approach has not been previously applied and validated for surface-based analyses of anatomy.
We first linearly registered MRI scans to a standard space. A small subgroup of image volumes was randomly chosen and manually traced. Lateral ventricular models were created in these image volumes and converted into parametric surface meshes (we will call these labeled image volumes “atlases”). We then fluidly registered each of these atlases and mesh models to all other subjects. A mesh averaging technique then combined the resulting fluidly propagated surface meshes for each image volume. To optimize the algorithm, we examined how the segmentation error (assessed using the symmetrized Hausdorff distance and 2-norm) and the disease detection power (assessed using cumulative distributions of surface statistics and the False Discovery Rate) depended on the number of atlases, leading to an empirical basis for choosing the number of templates in future studies that use multi-atlas segmentation.
We used this method to analyze morphometric differences in the lateral ventricles of subjects with AD, versus healthy elderly controls. The lateral ventricles of AD patients have been widely studied, and are known to display increased atrophy rates in surrounding tissues compared to aged matched controls, and are therefore an ideal system in which to validate our method.
Carriers of the apolipoprotein E epsilon-4 gene (ApoE4) are at heightened risk for early development of AD (Corder et al., 1993; Strittmatter et al., 1993). Structural imaging studies of AD patients have revealed morphological deficits in carriers of the ApoE4 allele versus non-carriers (Lehtovirta et al., 1995; Juottonen et al., 1998; Geroldi et al., 1999; 2000; Mori et al., 2002). In healthy elderly subjects, the ApoE4 allele is associated with diminished CNS glucose utilization (Reiman et al., 2004) and some studies have reported impairment in cognitive function (Deary et al., 2002). Inheritance of the ApoE4 allele is a risk factor for early development of AD, and, in epidemiological studies of several populations, between 10% and 18% of elderly healthy controls and 24–52% of AD patients have been found to carry at least one copy of the ApoE4 allele (Beyer et al., 1997). ApoE4-carriers with MCI (mild cognitive impairment) may respond especially well to cholinesterase treatment. In a study of 768 subjects with amnestic MCI, the ApoE4 carriers treated with donepezil showed a significant reduction in progression to AD at 3 years compared to carriers who received placebo, but the non-carriers did not (Petersen et al., 2005). There is, therefore, some interest in establishing whether neurodegeneration progresses differently in ApoE4-carriers than non-carriers and how atrophic changes interact with genotype. It is not yet known whether the ventricles of ApoE4 carriers differ from those of non-ApoE4 carriers. We thus validated our method further by looking at ventricular shape differences in a case that has not yet been studied, but where such changes are expected to exist.
Materials and methods
Fig. 1 shows the steps used to map multiple surface-based atlases into single average surface mesh via fluid registration. We detail our method and validation strategy in the following sections, and then present the steps used for ventricular shape modeling and statistical analysis. All t tests used here were two-tailed.
Fig. 1.
Mapping multiple surface-based atlases into new subjects’ scans via fluid registration. MRI scans of M different subjects (typically including both patients and controls, or a small sample representative of the individuals in the study) are first linearly registered to the ICBM standard space. N image volumes (subsequently called atlases) are randomly selected from the sample and the lateral ventricles are manually traced and converted into surface mesh models. M × N new ventricular models are then produced by fluid registration of each image volume to a different atlas. The N surface meshes per subject are integrated by simple mesh averaging for each individual subject.
Subjects
3D T1-weighted MRI scans of 17 subjects with AD (7 male/10 female; age: 68.7 ± 6.8 [SD] years, including 9 ApoE4-carriers) and 13 age-matched healthy elderly controls (6 male/7 female; age: 71.3±3.3 [SD] years, including 3 ApoE4-carriers), and for 18 asymptomatic ApoE4-carriers (7 male/11 female; 69.8±6.2 [SD] years) and 18 age- and gender-matched non-carriers (67.5 ± 6.0 [SD] years) were analyzed. All image volumes were spatially normalized by a nine-parameter transformation (3 translations, 3 rotations, 3 scales) to the International Consortium for Brain Mapping (ICBM-53) average brain template (Collins et al., 1994). All registrations and labelings were performed in this space (resolution 1 × 1 × 1 mm and dimension 181 × 217 × 181 voxels). Patients were diagnosed with AD using criteria detailed in the Diagnostic and Statistical Manual of Mental Disorders (American Psychiatric Association, 2000), and had a typical clinical presentation. They also fulfilled the standards for probable AD according to criteria established by the National Institute of Neurological Disorders and Stroke and the Alzheimer's Disease and Related Disorders Association (McKhann et al., 1984). Their cognitive status was evaluated using the Mini-Mental State Exam (MMSE). Mean MMSE scores were, on average, 16.8 for the AD group (2 patients were severely demented (MMSE<10), 10 patients were moderately demented (9<MMSE<21) and 5 patients were mildly demented (MMSE>20)). The mean MMSE was 29.6 for the controls; the ApoE carriers did not differ in mean MMSE score from the non-carriers (29.1 versus 29.2; p = 0.61, t test). This AD cohort would be therefore regarded as having relatively advanced AD. We chose this AD sample, in preference to including only mildly affected patients, to validate the approach for detecting a disease effect before attempting studies of subtler effects in milder AD or MCI. For that reason, we did not explicitly select people with especially mild AD, although that may have more practical value for future applications.
We tried to maximize the number of subjects for the comparisons of AD versus Control and ApoE4 versus non-ApoE4 carriers, but we only had a limited number of subjects who had been genotyped. As such, the groups used for the comparisons partly overlap, so we could include some additional controls for the AD versus Control comparison. In particular, we did not want to use the non-ApoE4 carriers as a control group for both studies. If we had done so, we would have risked over-estimating the disease effect by comparing an AD group with a partially unknown genetic make-up (and likely carrying the ApoE4 gene in an unknown proportion) with a control group that lacked the ApoE4 risk gene altogether. Rather than compare the AD group with a set of controls who lacked the risk gene, we preferred to compare them with a control group who had not been genotyped, but who should carry the E4 gene with an incidence comparable to the general population.
MRI scanning
Image volumes were acquired on a 2-Tesla Bruker Medspec S200 whole-body scanner (Bruker Medical, Ettingen, Germany) at the Centre for Magnetic Resonance (University of Queensland, Australia). A linearly polarized birdcage head-coil (Bruker Medical) was used for signal reception. Three-dimensional T1-weighted images were acquired with an inversion recovery segmented 3D gradient echo sequence (known as magnetization prepared rapid gradient echo; MP-RAGE) to resolve anatomy at high resolution. Acquisition parameters were as follows: inversion time (TI)/repetition time (TR)/echo time (TE) = 850/1000/8.3 ms; flip angle=20°; 32 phase-encoding steps per segment; 23 cm field of view.
Ventricular delineation
To build the labeled atlases, the anterior, posterior and inferior horns of the lateral ventricles were manually traced separately in consecutive coronal brain sections by a single image volume analyst, following previously described criteria with established inter- and intrarater reliability (Narr et al., 2001). Digitized surface contours were displayed simultaneously in all three viewing planes to help identify neuroanatomic boundaries, using the software MultiTracer (Woods, 2003). The anterior, posterior and inferior horns were hand-traced separately. This procedure makes mesh averaging across subjects more accurate at the junctions between the ventricular horns, as these can be correctly matched when surfaces are compared and averaged together. The fluid registration was performed on the whole 3D volume at once.
Viscous fluid registration
We followed the approach proposed in Gramkow (1996) to deform each unlabeled image volume onto each target image volume on which the lateral ventricle had been manually traced. The deforming image volume was treated as embedded in a compressible viscous fluid with motion governed by the Navier– Stokes equation for conservation of momentum, simplified to a linear PDE, as pioneered by Christensen et al. (1996). This equation can be written as:
| (1) |
where µ and λ are viscosity constants, and F(x⃗, u⃗) is the force field that varies with the position x⃗ and drives the deformation u⃗, and with v⃗ (x⃗, t) the deformation velocity experienced by a particle at position x⃗:
| (2) |
The term ∇2v⃗ in the Navier–Stokes equation is the Laplacian operator, that enforces spatial regularity in the deformation velocity and ensures a smooth deformation of the fluid. ∇(∇T ⋅ v⃗) is the gradient of the divergence operator; it controls the expansion or contraction of the model.
A 3D convolution filter, derived in Bro-Nielsen and Gramkow (1996) and Gramkow (1996), was used to implement a Green's function solution of the fluid equation. The body force F(x,u), driving the images into register, is defined as the gradient of the summed squared differences in intensities between the deforming template T(x−u(x,t)) and target S(x):
| (3) |
The use of the summed squared intensity difference as the cost function for registration assumes that the intensities of spatially normalized ventricular CSF and periventricular brain tissue are consistent across the population of images being registered. Other cost functions could be used to drive the registration, such as mutual information or divergence measures (Chiang et al., 2007b). In addition, the high contrast of ventricular CSF versus adjacent tissues is also useful for accurate registration, as it gives the registration functional a high gradient (variational derivative) at the ventricular boundary. This high image contrast makes the registration-based label propagation more accurate. Transformations resulting from the fluid registration were applied to the manually traced ventricular boundary using tri-linear interpolation; this generated a propagated contour on the unlabeled image volumes.
Ventricular shape modeling and distance mapping
To identify regional differences in ventricular morphology, we used surface-based mesh modeling to match corresponding ventricular surface points across individuals (Thompson et al., 1996a,b, 2004). Sets of points representing the tissue boundaries from each region were resampled and made spatially uniform by stretching a regular parametric grid (100 × 150 surface points) over each surface. Grid-points from corresponding surfaces were then matched across subjects to obtain group-average parametric meshes. For each surface model, a medial curve was defined as the line traced out by the centroid of the ventricular boundary (illustrated in Fig. 2; Thompson et al., 2000, 2004; see Styner et al., 2005, for related work on M-reps). The medial curve was defined separately in each individual before averaging the surfaces. The operations of averaging surfaces and defining the medial curve from a surface are not commutative, in the sense that a medial curve derived from an average surface would not be the same as the average of the medial curves derived from each individual. Because we were interested in measuring radial ventricular expansion in each individual, we computed these measures in each subject with reference to their own medial curve, but plotted the resulting statistics on the average surface for the groups being compared.
Fig. 2.
A flow chart shows how distance maps are used to model ventricular shape. The three horns (anterior, posterior, and inferior) of the lateral ventricles are traced manually in consecutive coronal sections (1), and converted into 3D parametric surface meshes composed of spatially uniform triangular tiles (2). Distance maps are computed (3) that specify the 3D distance of each ventricular surface point to a medial reference curve [red curve; (2)]. The radial distance maps are averaged across subjects and displayed on the average surface across subjects, where the average surface is computed using a mesh averaging technique (4).
The local radial size was defined as the radial distance between a boundary point and its associated medial curve. This allows statistical comparisons of local surface contractions and expansions at equivalent surface locations between groups in 3D.
If ri(u,v) is the 3D position in stereotaxic space Ω of the point with parametric coordinates (u,v) on the ith mesh, then the average surface from N individual surface meshes is defined as:
| (4) |
The uniformly gridded meshes in each atlas are propagated through a fluid deformation to label the target image volume, which makes the mesh node density non-uniform. Because there is no known homology between points on the ventricles in different subjects, except possibly at the junctions of the different horns of the ventricles, we chose to re-grid the surface once again, to uniformly distribute the points over it. There is also a practical reason for doing this. If we did not re-grid the surface, the new surface would be meshed with a grid point density that depends on the shape of the individual atlas brain propagated onto it. Its nodes would be denser in regions where the ventricles are smaller in the target image volume than in the atlas, and sparser in regions where the ventricles are larger in the target image volume than in the atlas. If the surface is re-gridded, it removes this bias. In practice the regridding alters the mean geometry a little, and reduces the 2-norm measure of error (but not the Hausdorff error) between two labelings of a surface.
Since we computed the radial distance from the medial curve, our approach is only sensitive to the distance perpendicular to the medial curve, and will not be sensitive to the distance parallel to it.
Ventricular statistical maps and analysis
Surface contractions and expansions were statistically compared between groups at equivalent locations (Thompson et al., 2004. 2006). Local group differences in ventricular shape between patients and controls were detected using Student's t-tests at equivalent surface locations in 3D. The associated P values describing the uncorrected significance of group differences were plotted onto the average surface model, producing a color-coded map.
For stronger control over the likelihood of false rejections of the null hypotheses with multiple comparisons, we adopted the method of Storey (2002), which directly measures the positive false discovery rate (pFDR) under a given primary threshold. The pFDR method can be more powerful than the popular sequential P value FDR method (Benjamini and Hochberg, 1995; Genovese et al., 2002), as it estimates the probability that the null hypothesis is true, from the empirical distribution of observed P values (Manly et al., 2004). The false discovery rate (FDR) is defined as:
| (5) |
where V is the number of false discoveries, and R is the total number of discoveries. Briefly, pFDR is the false discovery rate conditioned on the event that positive findings, rejecting the null hypothesis, have occurred, and is given by
| (6) |
where π0 = Pr(H=0) is the probability that the null hypothesis is true, and γ is the rejection threshold for the individual hypothesis, which was set to 0.01 in our experiments. We refer readers to Storey and Tibshirani (2001) and Storey (2002) for details of the estimation procedures to obtain pFDR for statistical maps. The positive false discovery rate may also be defined as:
| (7) |
The pFDR measure is theoretically more suitable for representing the “rate at which discoveries are false” than the FDR measure when the primary rejection region is relatively small (Storey, 2002). When the rejection region is large, then γ is large, and the pFDR value is very close to the FDR value; they only differ, with pFDR>FDR, when the rejection region is small (in our data they are similar as 30% of the anterior and posterior horn surface has P<0.05 for N = 1 atlas, and 70% has P<0.05 for N = 4 atlases). The pFDR and FDR values for findings obtained with different numbers of atlases N are reported in Fig. 5.
Fig. 5.
The cumulative P value distribution is shown for the comparison of AD versus control groups, and the effect of varying N, the number of atlases, from 1 to 4, and the best subset of 4 atlases chosen from a set of 9. Also reported are the pFDR, FDR values and the percentage of significance (i.e., the proportion of surface points in the averaged maps with significance values less than 0.05). The pink dashed line represents the expected CDF under the null hypothesis. As can be seen, the statistical power is greater when more atlases are used. Notably, the effect sizes obtainable using the best 4 atlases are comparable to those obtained using 4 randomly selected atlases, but both of these situations give higher effect sizes than using fewer atlases. The proportion of the surface showing significant effects, at the voxel level, rises from a third to well over two-thirds when a greater number of atlases are used for segmentation. Interestingly, it was only marginally better to use the best 4 atlases (versus randomly picking 4), at least compared with the substantial improvement in effect size and Hausdorff error obtained by increasing the number of atlases from 1 to 4.
By convention, a statistical map with pFDR<0.05, i.e., a false discovery rate less than 5%, was considered to be significant. The pFDR was computed from the uncorrected P values in the significance maps using the method described in Storey (2002).
Validation study
We hand traced the left lateral ventricles from 8 AD patients and 10 controls as gold standard segmentations. The samples included the 9 subjects selected from the controls for atlas generation. These 9 manually labeled surfaces were not included when calculating the Hausdorff error (Fig. 3). For validation experiments, only 4 of them were used to make the ventricular maps (Fig. 4). Future studies will look at the effects of selecting atlases from a subset of data that is balanced from the control, ApoE4-carrier, and AD groups.
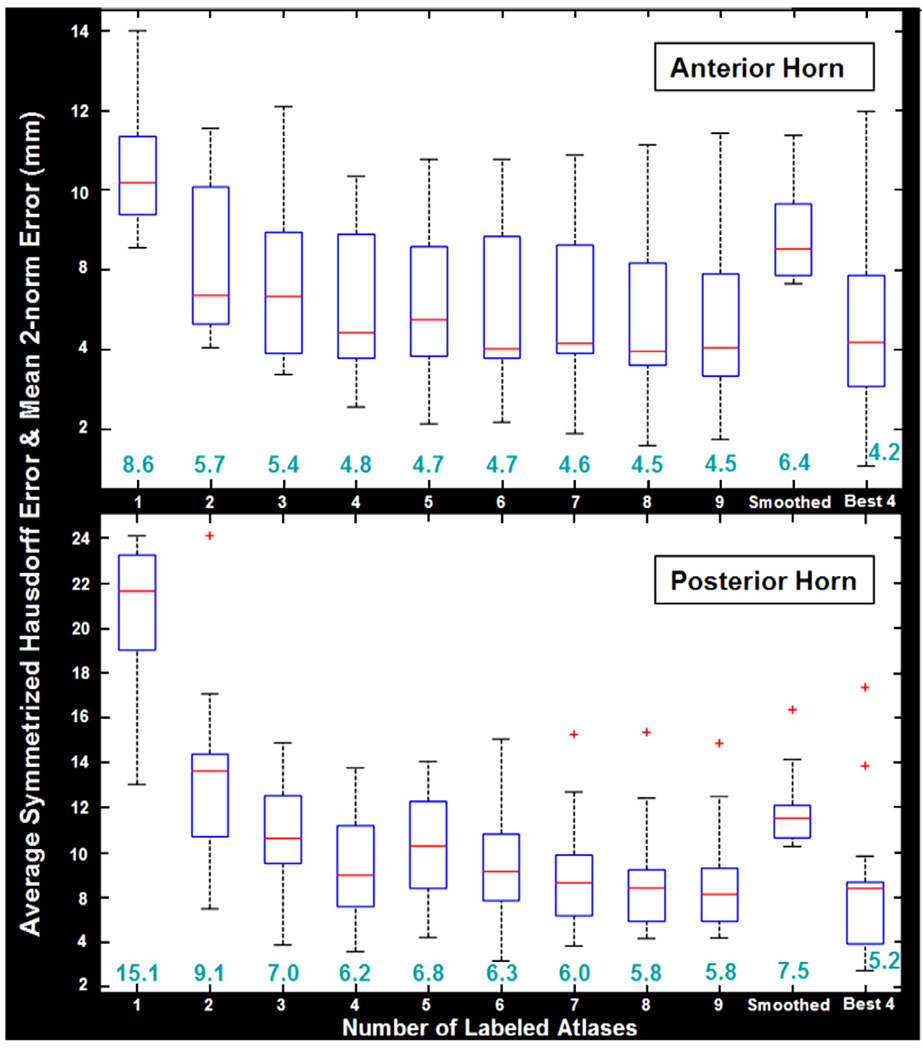
Fig. 3.
Plots of the average symmetrized Hausdorff error and mean 2-norm error for label propagation in the anterior and posterior horns of the ventricles. These plots are based on segmentations of 18 image volumes (8 AD patients, 10 controls), comparing automated segmentations with independent, manually derived left ventricular surface models. The horizontal axis indicates the results of using different numbers of labeled atlases (from 1 to 9) and the results obtained from surfaces generated from a single atlas after applying a 5 × 5 spatial smoothing filter. In the last column, the results are shown for the case where the “best 4” of the labeled atlases are used. The box plots are formed by drawing a line at the median and a box between the lower and upper quartiles. Any data observation that lies more than 1.5 times the interquartile range (IQR) lower than the first quartile or 1.5 times the IQR higher than the third quartile is considered an outlier, and whiskers indicate the most extreme values, on either side of the median, that are not outliers. Thus the box plot identifies the middle 50% of the data, the median, and the extreme points. As shown in the second to last column, applying some spatial smoothing to the surface generated from a single atlas does provide some benefit, but not as much as can be gained by generating multiple atlas surfaces. Segmentation results using a subset of the best 4 atlases are as good as using all 9, and marginally better than randomly picking 4. The mean 2-norm error is shown in green font, for each segmentation approach, and follows a similar pattern to the Hausdorff error.
Fig. 4.
Maps of the lateral surface of the left and right ventricular expansion using multiple atlases, (a) A measure of the radial distance is averaged across patients and, separately, across controls, and plotted on group average ventricular shapes using 4 atlases, (b) plots the ratio of the mean radial distance between patients and controls, revealing regions with 20% expansion radially (blue colors) using 4 atlases, (c) shows the significance of these differences at each surface point with different numbers of atlases (from 1 to 4) based on uncorrected P values.
To test our approach, surface models obtained automatically were compared to manual segmentations of the three horns of the lateral ventricles. The labeling error was measured using the symmetrized Hausdorff distance, as follows:
Let us first define the distance d(p,S′) between a point p belonging to a surface S and a surface S′ as:
| (8) |
where ‖·‖2 denotes the usual Euclidean norm. The Hausdorff distance is the maximum distance of a set to the nearest point in the other set (Daniel et al., 1990). More formally, the Hausdorff distance d(S,S′) between S and S′ is given by:
| (9) |
This distance is asymmetric, i.e., in general, d(S,S′)≠d(S′,S). The function d(S,S′) is called the forward Hausdorff distance from S to S′. It defines the point p ∈ S that is farthest from any point of S′, and measures the distance from p to its nearest neighbor in S′ using the given norm ‖·‖. Intuitively, if d(S,S′) = r, then each point of S must be within distance r of some point of S′, and there also is some point of S that is exactly distance r from the nearest point of S′. We also refer to d(S′,S) as the backward distance. It is then convenient to introduce the symmetrized Hausdorff distance ds(S, S′) as follows:
| (10) |
The Hausdorff distance accounts for overall discrepancies between two sets of surface points, and measures the discrepancy between two sets of points, without implying that there is any known correspondence between points on one surface and the other (i.e., it defines a distance between point sets). Even so, it does not provide any information about the accuracy of the surface as a whole. To assess this, we also computed the mean 2-norm between corresponding points of two surfaces, which is an alternative measure that assesses accuracy over the whole surface. This is defined as the distance between corresponding grid points, computed at each grid point and averaged over all points in the surface. This is naturally a symmetric measure, as it uses point pairs on each mesh. Each norm (Hausdorff and 2-norm) weights the errors a little differently, emphasizing outliers of different magnitudes to different degrees.
Results
Integrating multiple propagated labels into one
Fig. 2(4) shows a lateral ventricular surface extracted using different atlas image volumes. If the registrations were perfect and there were no digitization errors, all these image volumes would look identical. The right panel shows the average surface. The color bar represents the radial distance in mm.
By integrating multiple labels, random digitization errors from each hand-traced segmentation are significantly reduced. The resulting average model is also robust to inaccuracies in individual registrations that may occur when non-global minima of the intensity-based cost function are reached.
Comparison with manual labeling
Automatically propagated ventricular surface labels agreed closely with expertly labeled gold standard segmentations (Fig. 3) in tests on a randomly selected set of 8 patients and 10 controls.
Increasing the number of labeled atlases N resulted in an asymptotic decrease in both the average symmetric Hausdorff error and mean 2-norm between manually and automatically extracted models. The manually labeled surfaces were not included when calculating the Hausdorff error and mean 2-norm.
Using traditional volumetric measures calculated from the parametric meshes, the volume differences for the manually and automatically extracted ventricular surfaces (only anterior and posterior horn) did not reveal any significant difference between the controls and patients (P=0.97). In addition, we statistically assessed whether the automated or manual approaches gave systematically higher or lower volume values by comparing the mean volume difference between them with zero in each subject group (a one-sample t-test). We found that there was no bias in automated versus the manual approach in labeling the controls (P=0.56) or the patients (P=0.32). Even so, there was a significant difference for both the anterior horn (P = 8.3 × 10−8) and posterior horn (P = 0.04) between controls and patients when computing the labeling error by the Hausdorff distance (for anterior horn: controls—8.38 mm± 2.19 SD, patients—6.54 mm±1.80 SD; posterior horn: controls— 9.82 mm±4.49 SD, patients—8.37 mm±4.36 SD). This is because the registration is slightly more accurate in patients, as they have sharper image contrast at the ventricular boundaries and larger CSF volumes. Even so, there was no systematic bias in terms of the volumes being overestimated or underestimated in either patients or controls.
A box plot showing the average symmetrized Hausdorff error for the best subset of 4 atlases is shown in Fig. 3. Also, the cumulative P value distribution and the corresponding pFDR, FDR values and percentages of significance were reported in Fig. 5.
We also investigated whether the improvement in statistical power, with multiple atlases, may have resulted solely from the spatial smoothing inherent in averaging multiple surface meshes. As averaging is inherently a low-pass filtering technique, the Hausdorff error would be expected to decline slightly when averaging the meshes generated from several different atlases. As a result, we wanted to be sure that the benefit of averaging multiple atlases was greater than that obtainable by spatially filtering a single surface generated following fluid registration. If the extra power was purely attributable to a smoothing effect, then spatial smoothing of a single registered atlas surface would be a more efficient approach, as it would not require multiple atlas surfaces to be generated. To compare the averaging of meshes generated from multiple atlases with surfaces generated from a single atlas after applying a spatial smoothing filter, we plotted the results of neighborhood averaging over a 5 × 5 area, as a box plot, in Fig. 3. This smoothing operation uses the fact that there is already a mesh-like grid on the surface, and averages the coordinates of each point with those of its neighbors in a 5 × 5 neighborhood (an average of the available points was used in the case of points that were one or two grid positions from the boundary, where a full 5 × 5 neighborhood is not defined). As shown in the box plot, the symmetrized Hausdorff error (and mean 2-norm error) of a spatially smoothed single atlas is close to that obtainable by averaging the segmentation results using 2 atlases. When spatial smoothing is applied, some random digitization errors from each hand-traced segmentation are reduced, but errors from the inaccuracies in individual registrations still remain. Consequently, applying some spatial smoothing to the surface generated from a single atlas does provide some benefit, but not as much as can be gained by generating multiple atlas surfaces.
3D ventricular maps and statistical analysis
Statistical maps of mean radial distance comparing groups and maps of radial expansion for the AD patients versus controls and ApoE4-carriers versus non-carriers are shown in Figs. 4a,b. In Fig. 4b, the radial expansion of the ventricles is expressed as a percentage relative to the expected values in the corresponding group of controls. As expected, patients with AD and healthy ApoE4-carriers show localized volume enlargements in the posterior horn of the ventricles and in the mid-portion of the anterior horn (see also Carmichael et al., 2006). The group-averaged ventricular surface models were automatically generated using 4 randomly selected atlases from the control group. P value maps describing the significance of group differences were plotted onto the model surface. As shown in Fig. 4c, the total area of suprathreshold statistics (red) increased with N. The percentages of significance for different number of atlases are shown in Fig. 5. based on setting a significance level at the voxel level of 0.05.
Under the null hypothesis of no group difference, a plot of the cumulative distribution function (CDF) of the observed versus null P values is approximately diagonal. However, if significant changes are detected, the CDF increases rapidly from 0 over the small interval [0,α] and then levels off close to 1, since the CDF describes the probability distribution of the P values which are plotted in a sorted order and increase monotonically from 0 to 1. CDF plots could also be used to demonstrate the false discovery rate method for assigning overall significant values of group differences, as they show the proportion of suprathreshold voxels in a statistical map. Here we obtained P values for a set of 60,000 surface points covering the anterior and posterior horns. Inferior horn points were hard to delineate manually and were not included, due to concerns that a manual ‘gold standard’ for that region might be error-prone itself. Fig. 5 shows the cumulative P value distribution, and the effect of varying N from 1 to 4 with pFDR values. The pink dashed line represents the expected CDF under the null hypothesis. As shown, the statistical power increases with N.
As shown in Fig. 6, the P values are very sensitive to the individual chosen as an atlas, when a small number of atlases are used (e.g. a single subject). When using single atlas (top), the CDF lines vary greatly on all 4 possible sets, but they converge among 6 possible pairings when using the average of 2 atlases (bottom) from the set of 4 atlases.
Fig. 6.
The cumulative P value distribution of AD versus control based on segmentations obtained from various subsets of the set of 4 atlases used in Fig. 5. Top: 4 possible sets of “N = 1” CDF lines. This represents the effect sizes obtained when different single atlases are used as the basis for segmentation. Bottom: 6 possible sets of “N = 2” CDF lines. Note that in the case where a single atlas is chosen, the particular individual used affects the CDF and resulting P value, to some degree (top panel). This is undesirable in an empirical study, as the effects ought to be stable with respect to the arbitrary choices in the segmentation method. When 2 atlases are used, the resulting effect sizes and CDF plots are more stable with respect to the choice of atlases, which is to be expected given that some of the random errors associated with registration are canceled out regardless of which 2 atlases are used. Multi-atlas segmentation therefore not only increases the power to detect group differences, but the results are likely to be more reproducible.
To determine the optimal value of N, we performed 2-tailed t-tests to see by how much Hausdorff errors fell when adding an additional atlas, i.e., t-test (Error(N),Error(N−l)), for the anterior and posterior horns. In this study, values of N > 4 did not detectably increase the power (Figs. 7a,b). Values shown above the points represent the associated P values.
Fig. 7.
Two-tailed t-tests were performed to determine the optimal number of atlases N for (a) anterior and (b) posterior horns. For the current study, using more than 3 atlases did not yield a detectable increase in power. Values shown above the points represent the associated P values for the t-tests of the error reduction caused by adding another atlas.
Discussion
Morphometric alterations in subcortical structures are potential markers of a variety of neurodegenerative diseases and neuropsychiatric disorders, including Alzheimer's disease (Double et al., 1996; Jack et al., 2000), schizophrenia (Puri et al., 1999; Seidman et al., 1999; Frisoni et al., 2006) and other conditions (Halliday et al., 1996; Wolf et al., 2001). Volumes of brain structures are still commonly derived by expert manual labeling, a process that is both time-consuming and tedious. Furthermore, differences in interobserver delineation or drift over time may preclude the stable analysis of subjects over longer time-spans. In this paper, we combined a new automated anatomical surface modeling, based on multi-atlas fluid image alignment, with surface-based statistics as a method to visualize significant shape differences between two groups.
In our prior work (Carmichael et al., 2006), we developed a single-atlas-based surface segmentation approach, using a deformation model based on optical flow, a slightly simpler formulation than the fluid model used here. The fluid model guarantees that the resulting deformation mappings are diffeomorphic, i.e. formally guaranteed to remain one-to-one, invertible, and spatially smooth even if large image deformations are required. This one-to-one property arises from regularizing the template velocity rather than the deformation magnitude itself, and is not enforced with optical flow models (although it is a reasonable assumption for those models when deformations are small).
Compared with the ‘single-atlas’ approach, ‘multi-atlas’ fluid registration increased the statistical power in our surface-based morphometric study, and robustly detected differences in AD and ApoE4 carriers. In our sample, we did not detect overall cognitive differences between the ApoE4 and non-E4 carriers, at least on the MMSE, which is perhaps the most common and widely accepted measure of incipient cognitive decline used in studies of dementia. More sensitive neurocognitive tests may reveal associations between the ventricular differences reported here and cognition, or it may be that in the prodrome1 of disease, there is a cognitive buffer that leaves cognition intact until brain morphology has deteriorated beyond a certain point (see Stern, 2006).
Ventricular measures also show a relatively high effect size in distinguishing disease from normality, even in groups where overall brain volumes provide poor discrimination. For example, in one longitudinal study (Thompson et al., 2004), measures of temporal horn volume expansion were around five times faster for AD patients than for controls (i.e., L: +18.1 ± 3.8% per year, R: +12.8 ± 4.7% per year in AD; and L: +3.7 ± 1.1% per year, R: +1.7 ± 1.1% per year in controls; group difference, P<0.0005). In the same study, total cerebral volumes were declining more rapidly in AD for both the left hemisphere (AD: 5.9 ± 2.5% per year; CTL: 1.0 ± 0.2%; group difference, P<0.023), and the right hemisphere (AD: 4.6±1.6% per year; CTL: 0.8±0.2%; group difference, P<0.0095), so group discrimination was slightly better for the ventricular volume measure. Others have also reported medium to large effect sizes for ventricular enlargement in cross-sectional studies of AD versus controls (e.g., Cohen’s d>0.5; Studholme et al., 2002).
Differences in the AD patients versus controls were greater than those of the ApoE4 carriers versus non-carriers, as the carriers of the risk gene had not developed AD at the time of the study. While the ApoE4 carrier versus non-carrier maps show differences primarily in anterior horn of the ventricles, the AD versus control maps also show differences in the posterior horn. This is reasonable as the occipital lobe is atrophied only in moderate to late AD, and is one of the last brain regions to be affected by Alzheimer pathology, so posterior horn differences found in AD may not be yet present in those at risk for AD.
As the current sample was small, we did not aim to differentiate subjects carrying a single ApoE4 risk allele from those who were homozygous (i.e., carrying two copies of E4). Individuals with two E4 copies are at greatly heightened risk for AD and may not be typical of those heterozygous for the risk allele (which is the more common situation). Future studies will focus on detecting interactions between E4 presence and therapeutic response. Also of interest are effects of the ApoE2 allele, which is rare but thought to be protective against future onset of AD. Other polymorphic alleles, such as BDNF (brain-derived neurotropic factor; Bueller et al., 2006) and other neurotrophins, may be relevant in determining ventricular morphology or its rate of change during development and disease.
Some of the shape differences recovered here, between AD and control groups, and between genetic subgroups, may reflect the widening of parts of the ventricle that were previously obscured or made invisible by partial volume effects. In normal subjects, the posterior horn is ‘pinched inwards’ at the calcar avis, near the depths of the calcarine fissure. When it opens up in AD, more of the posterior limit is visible. In healthy controls, the posterior horn is typically smaller and narrower, and partial volume effects are common (where some voxels contain both CSF and other tissues). This can lead to the incorrect assumption that the ventricle's posterior limit is growing rapidly backwards in AD, when in reality a part of the ventricles has opened up, that was previously obscured by partial volume effects has opened up. Changes in ventricular shape, detected here, are therefore likely to reflect both ventricular expansion and the appearance of parts of the ventricles previously obscured by partial volume effects.
Unfortunately, our deformation-based segmentation method is not optimal for mapping the inferior horns of the ventricles, since healthy subjects typically have very low volumes of ventricular CSF in the inferior horns, and their geometry appears narrow on coronally resliced MRI (typically a millimeter or less in width). Fortunately, the other ventricular horns do show very high effect sizes for disease and genetic effects, partly because the error in segmenting them is much less as a proportion of their overall volume and the percent difference in volume in disease is extremely large. In theory an inferior horn template could be propagated into new scans using a fluid registration approach, but it would be hard to propagate accurately onto controls in regions with very limited CSF, as large localized contractions (i.e., compressions of the template) would be needed. The fluid prior in the registration enforces a spatial smoothness in the deformation that limits very large localized compressions. In our tensor-based morphometry (TBM) studies, we have found that the smoothness of the deformation fields may limit the accuracy of the fluid registration for relatively small-scale structures (Hua et al., submitted for publication). For instance, surface-based methods detected hippocampal atrophy bilaterally in a group of 17 AD patients compared with 14 controls (Thompson et al., 2004), but a TBM study of a much larger sample (40 AD patients versus 40 controls) detected hippocampal atrophy in only one hemisphere (Hua et al., submitted for publication). The main reason for the poor effect size may be the relatively poor accuracy of fluid registration in labeling the hippocampus, compared with manual methods, and compared with fluid-based labeling of a large high-contrast structure such as the ventricles. Similarly, it would be difficult to measure cortical thickness using deformation based morphometry, as this would require very high frequency information to be recovered. This limitation may be overcome by future approaches using information-theoretic or log-tensor deformation priors that allow very large local contractions if they are shown to be expected statistically (Leow et al., 2007; Brun et al., 2007).
Our approach could be generalized, in principle, to create surface models of the hippocampus and the caudate. Recently, we used a single-atlas approach governed by optical flow image registration (rather than the fluid model used here) to segment the hippocampus in aging and AD (Carmichael et al., 2005). Hippocampal segmentation is generally regarded as more challenging than ventricular segmentation. The ventricles rely on high CSF/brain contrast to be extracted reliably and efficiently in very large numbers of subjects (Carmichael et al., 2006). The hippocampus, on the other hand, is relatively small and often does not present sufficient contrast or sharp borders, making automated or even manual segmentation less precise and less reliable and precise across raters and methods. Ultimately some hybrid combination of deformation-based methods (such as this one) and other computer vision and pattern recognition methods may outperform methods based on deformable atlases alone. In addition, although the hippocampus is more challenging to segment than the ventricles, the ventricles may have comparable or greater value as a biological marker of disease progression. In at least one study, inferior horn volume correlated more strongly with cognitive test scores than hippocampal volume did for AD patients (Jack et al., 2003). We recently found that an adaptive boosting algorithm (that learns discriminative features from a training set) combined with support vector machines (an approach for nonlinear regression and classification) provided high-quality hippocampal segmentations in an AD and control MRI database (Morra et al., submitted for publication, 2007). Ultimately, adaptive rather than a priori feature selection may be inevitable if the imaging pulse sequence or image contrast is varied, as is common in large multi-site studies.
In a follow-up study, we plan to cross-validate our AD and ApoE4 results with those of other methods to measure local differences in the ventricles, such as voxel-based morphometry (Ashburner and Friston, 1999), tensor-based morphometry (Hua et al., submitted for publication) and segmentation with a classifier based on adaptive boosting and support vector machines (Morra et al., submitted for publication, 2007).
Other groups have attempted fully automated subcortical segmentation (Fischl et al., 2002). Chupin et al. (2007) segmented brain structures using a Markov Random Field model based on a predefined set of features, invoking statistics on structure geometry and intensity in Bayesian prior distributions, rather than deforming a specific model under constraints. Our method is not Bayesian, unless the fluid model is regarded as a Gibbs prior. Bayesian approaches offer some advantages as they provide a case-by-case quantitative measure of confidence in the automated segmentations, that could be incorporated into the group statistical analysis.
Haller et al. (1997) and Hogan et al. (2000) used a fluid registration model to deform a single hippocampal surface model into new subjects, yielding a set of models that were analyzed for group differences in shape in Alzheimer's disease (Csernansky et al., 2005) and shape asymmetries (Wang et al., 2004). Zhou and Rajapaske (2005) proposed an automated method using fuzzy templates to segment different brain structures based on information extracted from a set of training image volumes. Ferrarini et al. (2006) used artificial neural networks to study ventricular shape variations in healthy elderly and AD subjects, generating a control average surface and a cloud of corresponding nodes across a data set. Heckemann et al. (2006) performed label propagation using free-form deformations and decision fusion to provide automated anatomical segmentations of the brain.
Level-set or active surface methods present an alternative to atlas-based approaches for deforming a pre-segmented anatomical model into new scans (Yushkevich et al., 2006). These methods typically use partial differential equations to evolve a deforming template or interface under image-derived forces, but they often require some interactive initialization of a template in every image volume, to allow accurate label propagation.
The optimal number of atlases depends on the distribution of registration errors for each structure, and whether there is any systematic bias between automated and manual segmentations. The accuracy and power gained may vary based on imaging protocol differences between the labeled and test image volumes. The incremental value of this method versus any standard approach depends on the statistical power required, the hypotheses, the study design, and the expected effect sizes. In addition, segmented models may be better combined using approaches that are robust to outliers, such as M-estimators or median filters on binarized labels. It may be most beneficial in drug trials or genetic studies where power is low and image acquisition is costly. This technique may also help neuroscientific efforts to find specific genes, risk factors or neuroprotective treatments that modify rates of brain degeneration.
Composition of the atlas group
For practical application of any type of atlas-based segmentation, it is beneficial to know whether it is better to propagate atlases that are matched by age or diagnosis to the individuals whose anatomy is being studied. Recent studies have examined a related problem, i.e., whether normalization of individual image volumes results in greater effect sizes in group study, or better labeling accuracy, when data are normalized to a group-specific template (defined using appropriate deformation and intensity metrics; Leporé et al., 2007), or optimized individual template (Chiang et al., 2007a). In multi-atlas setting, this concern is partially alleviated because the use of atlas templates from multiple subjects reduces the concern that the new anatomies to be labeled will differ severely from the labeled template. Fig. 8 shows the segmentation error for labeling the anterior and posterior ventricular horns of subjects aged 58 to 76, to examine whether labeling error depends on age. As expected, the labeling error, as quantified by the average symmetrized Hausdorff distance, is decreased as the number of templates propagated into the labeled brain is increased. In addition, there is no detectable age effect on accuracy. It might be hypothesized that anatomies of older subjects might be labeled more accurately than younger subjects, as age-related atrophy enlarges the CSF spaces, and this may alleviate any partial volume effects that make the structures hard to discern (e.g., at the posterior tip of the posterior horn). There is no obvious age effect even when only a single atlas is used (as shown by the upper trace in Fig. 8), and both the magnitude and cross-subject variability in labeling accuracy across subjects are greatly reduced as the number of atlases increases. Fig. 8 shows how well each brain (of different ages) is labeled, but the registration process is symmetrical so the same plot provides an index of how accurately each brain would label other brains if used as an atlas template. There is no apparent benefit of using older versus younger brains as templates, but one could imagine selecting the best N<k of K atlases based on their labeling error in a pilot study, and using this subset as a template set for a larger study. This principle for atlas selection and other meta-analytic improvements deserve further study.
Fig. 8.
Plots of labeling error for brains of subjects of different ages. To determine whether older subjects are labeled more accurately than younger ones, these plots show the average symmetrized Hausdorff error (in millimeters) for labeling each of a range of subjects of different ages. Data are presented for labelings using 1 to 9 different atlases. The labeling error for every subject falls when multiple atlases are used, and there is no obvious age effect on labeling accuracy. Conversely, these error metrics also measure the expected labeling error if each of these brains labeled here were used instead as an atlas template to label other brains. There is no obvious benefit to selecting subjects of a certain age as templates, but subsets of atlases could certainly be chosen, based on this type of data, to minimize the expected labeling error. The blue ticks on horizontal axis indicate the data points (8 AD patients, 10 controls) that actually occur in these line plots.
Acknowledgments
This work was funded by grants from the National Institute for Biomedical Imaging and Bioengineering, the National Center for Research Resources, and the National Institute on Aging (EB01651, RR019771, AG016570 to PT). Additional support was provided by NCRR Resource grant P41 RR13642 to AWT, and by the National Institute on Aging (AG021431 to JTB). The authors are grateful to Stephen Rose at the University of Queensland Center for Magnetic Resonance for his work acquiring the MRI scan data.
Footnotes
Prodrome: the prodrome is the early phase of a disease process before overt symptoms appear (from the Greek word for “forerunner”).
References
- Ashbumer J, Friston KJ. Spatial normalization. In: Toga AW, editor. Brain Warping, ch. 2. San Diego, USA: Academic Press; 1999. pp. 27–44. [Google Scholar]
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders. Ed 4. Washington, DC: American Psychiatric Association; 2000. (DSM-IV) [Google Scholar]
- Babalola KO, Cootes TF, Twining CJ, Petrovic V, Schestowitz R, Taylor CJ. Building 3D Statistical Shape Models using Groupwise Registration. Proceedings of the 12th annual meeting of the Organization for Human Brain Mapping; Florence, Italy. 2006. [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc, Ser. B Method. 1995;57:289–300. [Google Scholar]
- Beyer K, Lao JI, Fernandez-Novoa L, Alvarez XA, Sellers MA, Cacabelos R. APOE epsilon 4 allele frequency in Alzheimer’s disease and vascular dementia in the Spanish population. Ann. N. Y. Acad. Sci. 1997;826:452–455. doi: 10.1111/j.1749-6632.1997.tb48502.x. (1997 Sep 26) [DOI] [PubMed] [Google Scholar]
- Bro-Nielsen M, Gramkow C. Fast Fluid Registration of Medical Images. Proceedings of the 4th International Conference on Visualization in Biomedical Computing; September 22–25; Hamburg, Germany. 1996. pp. 267–276. [Google Scholar]
- Brun CA, Leporé N, Pennec X, Chou YY, Lopez OL, Aizenstein HJ, Becker JT, Toga AW, Thompson PM. Comparison of Standard and Riemannian Elasticity for Tensor-Based Morphometry in HIV/AIDS. 10th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI); Oct 29-Nov 2 (2007); Brisbane, Australia. 2007. [Google Scholar]
- Bueller JA, Aftab M, Sen S, Gomez-Hassan D, Burmeister M, Zubieta JK. BDNF Val66 Met allele is associated with reduced hippocampal volume in healthy subjects. Biol. Psychiatry. 2006;59(9):812–815. doi: 10.1016/j.biopsych.2005.09.022. [DOI] [PubMed] [Google Scholar]
- Carmichael OT, Aizenstein HA, Davis SW, Becker JT, Thompson PM, Meltzer CC, Liu Y. Atlas-based hippocampus segmentation in Alzheimer’s disease and mild cognitive impairment. NeuroImage. 2005;27(4):979–990. doi: 10.1016/j.neuroimage.2005.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carmichael OT, Thompson PM, Dutton RA, Lu A, Lee SE, Lee J, Kuller Y, Lopez LH, Aizenstein OL, Meltzer HJ, Liu CC, Yoga Y, Becker AW. Mapping ventricular changes related to dementia and mild cognitive impairment in a large community-based cohort. IEEE ISBI. 2006:315–318. [Google Scholar]
- Carmichael OT, Kuller LH, Lopez OL, Thompson PM, Lu A, Lee SE, Lee JY, Aizenstein HJ, Meltzer CC, Liu Y, Toga AW, Becker JT. Ventricular volume and dementia progression in the Cardiovascular Health Study. Neurobiol. Aging. 2007a;28(3):389–397. doi: 10.1016/j.neurobiolaging.2006.01.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carmichael OT, Kuller LH, Lopez OL, Thompson PM, Dutton RA, Lu A, Lee SE, Lee JY, Aizenstein HJ, Meltzer CC, Liu Y, Toga AW, Becker JT. Acceleration of cerebral ventricular expansion in the Cardiovascular Health Study. Neurobiol. Aging. 2007b;28(9):1316–1321. doi: 10.1016/j.neurobiolaging.2006.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiang MC, Dutton RA, Hayashi KM, Toga AW, Lopez OL, Aizenstein HJ, Becker JT, Thompson PM. 3D pattern of brain atrophy in HIV/AIDS visualized using tensor-based morphometry. NeuroImage. 2007a;34(1):44–60. doi: 10.1016/j.neuroimage.2006.08.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiang MC, Reiss AL, Lee AD, Bellugi U, Galaburda AM, Korenberg J, Mills DL, Toga AW, Thompson PM. 3D pattern of brain abnormalities in Williams syndrome visualized using tensor-based morphometry. NeuroImage. 2007b;36(4):1096–1109. doi: 10.1016/j.neuroimage.2007.04.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christensen GE, Rabbitt RD, Miller MI. Deformable templates using large deformation kinematics. IEEE Trans. Image Process. 1996;5:1435–1447. doi: 10.1109/83.536892. [DOI] [PubMed] [Google Scholar]
- Christensen GE, Johnson HJ, Vannier MW. Synthesizing average 3D anatomical shapes. NeuroImage. 2006;32(1):146–158. doi: 10.1016/j.neuroimage.2006.03.018. [DOI] [PubMed] [Google Scholar]
- Chupin M, Mukuna-Bantumbakulud R, Hasboun D, Bardinet E, Baillet S, Kinkingnehun S, Lemieux L, Dubois B, Garnero L. Anatomically constrained region deformation for the automated segmentation of the hippocampus and the amygdala: method and validation on controls and patients with Alzheimer’s disease. NeuroImage. 2007;34:996–1019. doi: 10.1016/j.neuroimage.2006.10.035. [DOI] [PubMed] [Google Scholar]
- Collins DL, Neelin P, Peters TM, Evans AC. Automatic 3D intersubject registration of MR volumetric data in standardized Talairach space. J. Comput. Assist. Tomogr. 1994;18(2):192–205. [PubMed] [Google Scholar]
- Cootes TF, Twining CJ, Petrovic V, Schestowitz R, Taylor CJ. Groupwise Construction of Appearance Models Using Piece-wise Affine Deformations. Vol. 2. Oxford, UK: BMVC; 2005. pp. 879–888. [Google Scholar]
- Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, Roses AD, Haines JL, Pericak-Vance MA. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science. 1993;261:921–923. doi: 10.1126/science.8346443. [DOI] [PubMed] [Google Scholar]
- Csernansky JG, Wang L, Swank J, Miller LP, Gado M, McKeel D, Miller MI, Morris JC. Preclinical detection of Alzheimer’s disease: hippocampal shape and volume predict dementia onset in the elderly. NeuroImage. 2005;25(3):783–792. doi: 10.1016/j.neuroimage.2004.12.036. [DOI] [PubMed] [Google Scholar]
- Daniel P, Huttenlocher P, Klara K. Computing the Minimum Hausdorff Distance for Point Sets Under Translation. Proc. of 6th Annual ACM Symp. on Comp. Geom. (SCG’90, Berkeley, CA); 1990. pp. 340–349. [Google Scholar]
- Dawant BM, Hartmann SL, Thirion JP, Vandermeulen D, Demaerel P. Automatic 3-D segmentation of internal structures of the head in MR images using a combination of similarity and free-form transformations: Part I. Methodology and validation on normal subjects. IEEE Trans. Med. Imag. 1999;18:909–916. doi: 10.1109/42.811271. [DOI] [PubMed] [Google Scholar]
- Deary IJ, Whiteman MC, Partie A, Starr JM, Hayward C, Wright AF, Carothers A, Whalley LJ. Cognitive change and the APOE epsilon 4 allele. Nature. 2002:418–932. doi: 10.1038/418932a. [DOI] [PubMed] [Google Scholar]
- Double K, Halliday G, Drill J, Harasty J, Cullen K, Brooks WS, Creasey H, Broe GA. Topography of brain atrophy during normal aging and Alzheimer’s disease. Neurobiol. Aging. 1996;17:513–521. doi: 10.1016/0197-4580(96)00005-x. [DOI] [PubMed] [Google Scholar]
- Ferrarini L, Palm WM, Olofsen H, Buchem MA, Reiber JHC, Admiraal-Behloul F. Shape differences of the brain ventricles in Alzheimer’s disease. NeuroImage. 2006;32(3):1060–1069. doi: 10.1016/j.neuroimage.2006.05.048. [DOI] [PubMed] [Google Scholar]
- Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C, van der Kouwe A, Killiany R, Kennedy D, Klaveness S, Montillo A, Makris N, Rosen N, Dale AM. Whole brain segmentation: automated labeling of neuroanatomical structures in the human brain. Neuron. 2002;33(3):341–355. doi: 10.1016/s0896-6273(02)00569-x. [DOI] [PubMed] [Google Scholar]
- Frisoni GB, Sabattoli F, Lee AD, Dutton RA, Toga AW, Thompson PM. In vivo neuropathology of the hippocampal formation in AD: a radial mapping MR-based study. NeuroImage. 2006;32(1):104–110. doi: 10.1016/j.neuroimage.2006.03.015. [DOI] [PubMed] [Google Scholar]
- Genovese CR, Lazar NA, Nichols T. Thresholding of statistical maps in functional neuroimaging using the false discovery rate. NeuroImage. 2002;15:870–878. doi: 10.1006/nimg.2001.1037. [DOI] [PubMed] [Google Scholar]
- Geroldi C, Pihlajamaki M, Laakso MP, DeCarli C, Beltramello A, Bianchetti A, Soininen H, Trabucchi M, Frisoni GB. APOE-epsilon4 is associated with less frontal and more medial temporal lobe atrophy in AD. Neurology. 1999;53:1825–1832. doi: 10.1212/wnl.53.8.1825. [DOI] [PubMed] [Google Scholar]
- Geroldi C, Laakso MP, DeCarli C, Beltramello A, Bianchetti A, Soininen H, Trabucchi M, Frisoni GB. Apolipoprotein E genotype and hippocampal asymmetry in Alzheimer’s disease: a volumetric MRI study. J. Neurol. Neurosurg. Psychiatry. 2000;68:93–96. doi: 10.1136/jnnp.68.1.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gramkow C. Registration of 2D and 3D medical images, Master’s thesis. Copenhagen, Denmark: Danish Technical University; 1996. [Google Scholar]
- Haller JW, Banerjee A, Christensen GE, Gado M, Joshi S, Miller MI, Sheline Y, Vannier MW, Csernansky JG. Three-dimensional hippocampal MR morphometry with high-dimensional transformation of a neuroanatomic atlas. Radiology. 1997;202(2):504–510. doi: 10.1148/radiology.202.2.9015081. [DOI] [PubMed] [Google Scholar]
- Halliday GM, McRitchie DA, Macdonald V, Double KL, Trent RJ, McCusker E. Regional specificity of brain atrophy in Huntington’s disease. Exp. Neurol. 1996;154:663–672. doi: 10.1006/exnr.1998.6919. [DOI] [PubMed] [Google Scholar]
- Heckemann RA, Hajnal J, Aljabar P, Rueckert D, Hammers A. Automatic anatomical brain MRI segmentation combining label propagation and decision fusion. NeuroImage. 2006;32:70–78. doi: 10.1016/j.neuroimage.2006.05.061. [DOI] [PubMed] [Google Scholar]
- Hogan RE, Mark KE, Wang L, Joshi S, Miller MI, Bucholz RD. Mesial temporal sclerosis and temporal lobe epilepsy: MR imaging deformation-based segmentation of the hippocampus in five patients. Radiology. 2000;216:291–297. doi: 10.1148/radiology.216.1.r00jl41291. [DOI] [PubMed] [Google Scholar]
- Hua X, Leow AD, Lee S, Klunder AD, Toga AW, Leporé N, Chou YY, Brun C, Chiang MC, Barysheva M, Jack CR, Bernstein MA, Britson PJ, Ward CP, Whitwell JL, Borowski B, Fleisher A, Fox NC, Boyes R, Barnes J, Harvey D, Kornak J, Schuff N, Boreta L, Alexander GE, Weiner MW, Thompson PM For the ADNI Study, submitted for publication. 3D characterization of brain atrophy in Alzheimer’s disease and mild cognitive impairment using tensor-based morphometry. NeuroImage. doi: 10.1016/j.neuroimage.2008.02.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jack CR, Jr, Petersen RC, Xu YC, O’Brien PC, Smith GE, Ivnik RJ, Boeve BF, Tangalos EG, Kokmen E. Rates of hippocampal atrophy correlate with change in clinical status in aging and AD. Neurology. 2000;55:484–489. doi: 10.1212/wnl.55.4.484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jack CR, Slomkowski M, Gracon S, Hoover TM, Felmlee JP, Stewart K, Xu Y, Shiung M, O’Brien PC, Cha R, Knopman D, Petersen RC. MRI as a biomarker of disease progression in a therapeutic trial of milameline for AD. Neurology. 2003;60(2):253–260. doi: 10.1212/01.wnl.0000042480.86872.03. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Juottonen K, Lehtovirta M, Helisalmi S, Riekkinen PJ, Sr, Soininen H. Major decrease in the volume of the entorhinal cortex in patients with Alzheimer’s disease carrying the apolipoprotein E epsilon 4 allele. J. Neurol. Neurosurg. Psychiatry. 1998;65:322–327. doi: 10.1136/jnnp.65.3.322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein A, Mensh B, Ghosh S, Tourville J, Hirsch J. Mindboggle: automated brain labeling with multiple atlas. BMC Medical Imaging. 2005;5:7. doi: 10.1186/1471-2342-5-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kochunov P, Lancaster J, Hardies J, Thompson PM, Woods RP, Cody JD, Hale DE, Laird A, Fox PT. Mapping structural differences of the corpus callosum in individuals with 18q deletions using targetless regional spatial normalization. Hum. Brain Mapp. 2005;24:325–331. doi: 10.1002/hbm.20090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehtovirta M, Laakso MP, Soininen H, Helisalmi S, Mannermaa A, Helkala EL, Partanen K, Ryynanen M, Vainio P, Hartikainen P. Volumes of hippocampus, amygdala and frontal lobe in Alzheimer patients with different apolipoprotein E genotypes. Neuroscience. 1995;67:65–72. doi: 10.1016/0306-4522(95)00014-a. [DOI] [PubMed] [Google Scholar]
- Leow AD, Chiang MC, Yanovsky I, Lee AD, Lu A, Klunder AD, Becker JT, Davis SW, Toga AW, Thompson PM. Statistical properties of Jacobian maps and inverse-consistent deformations in nonlinear image registration. IEEE Trans. Med. Imag.; May 2007; 2007. [DOI] [PubMed] [Google Scholar]
- Leporé N, Brun C, Pennec X, Chou Y, Lopez OL, Aizenstein HJ, Becker JT, Toga AW, Thompson PM. Mean Template for Tensor-Based Morphometry Using Deformation Tensors. 10th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI); Oct 29-Nov 2 (2007); Brisbane, Australia. 2007. [DOI] [PubMed] [Google Scholar]
- Lorenzen P, Prastawa M, Gerig G, Bullitt E, Joshi S. Multi-model image set registration and atlas formation. Med. Image Anal. 2006;10:440–451. doi: 10.1016/j.media.2005.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manly KF, Nettleton D, Hwang JT. Genomics, prior probability, and statistical tests of multiple hypotheses. Genome Res. 2004;14:997–1001. doi: 10.1101/gr.2156804. [DOI] [PubMed] [Google Scholar]
- McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadian EM. Clinical diagnosis of AD: reporter of the NINCDS-ARDRA Work Group under the auspices of the Health and Human Services Task Force on Alzheimer’s Disease. Neurology. 1984;34:934–944. doi: 10.1212/wnl.34.7.939. [DOI] [PubMed] [Google Scholar]
- Miller MI. Computational anatomy: shape, growth, and atrophy comparison via diffeomorphisms. NeuroImage. 2004;23(Suppl 1):S19–S33. doi: 10.1016/j.neuroimage.2004.07.021. [DOI] [PubMed] [Google Scholar]
- Mori E, Lee K, Yasuda M, Hashimoto M, Kazui H, Hirono N, Matsui M. Accelerated hippocampal atrophy in Alzheimer’s disease with apolipoprotein E e4 allele. Ann. Neurol. 2002;51:209–214. doi: 10.1002/ana.10093. [DOI] [PubMed] [Google Scholar]
- Morra JH, Tu Z, Apostolova LG, Green A, Toga AW, Thompson PM. Comparison of Adaboost and support vector machines for detecting Alzheimer’s disease through automated hippocampal segmentation. submitted to IEEE Transactions on Medical Imaging; Sept. 7 2007; 2007b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narr KL, Thompson RM, Sharma T, Moussai J, Blanton R, Anvar B, Edris A, Krupp R, Rayman J, Khaledy M, Toga AW. Three-dimensional mapping of temporo-limbic regions and the lateral ventricles in Schizophrenia: gender effects. Biol. Psychiatry. 2001;50:84–97. doi: 10.1016/s0006-3223(00)01120-3. [DOI] [PubMed] [Google Scholar]
- Narr KL, Thompson PM, Szeszko P, Robinson D, Jang S, Woods RP, Kim S, Hayashi KM, Asunction D, Toga AW, Bilder RM. Regional specificity of hippocampal volume reductions in first-episode schizophrenia. NeuroImage. 2004;21:1563–1575. doi: 10.1016/j.neuroimage.2003.11.011. [DOI] [PubMed] [Google Scholar]
- Petersen RC, Thomas RG, Gruridman M, Bennett D, Doody R, Ferris S, Galasko D, Jin S, Kaye J, Levey A, Pfeiffer E, Sano M, van Dyck CH, Thal LJ for the Alzheimer’s Disease Cooperative Study Group. Vitamin E and donepezil for the treatment of mild cognitive impairment. N. Engl. J. Med. 2005;352(23):2379–2388. doi: 10.1056/NEJMoa050151. [DOI] [PubMed] [Google Scholar]
- Powell AL, Mezrichb RS, Wang JZ. Abnormal cerebral ventricular change in Alzheimer’s disease recorded with cardiac gated MRI: a preliminary report. Dement. Geriatr. Cogn. Disord. 1991;2:45–50. [Google Scholar]
- Puri BK, Richardson AJ, Oatridge A, Hajnal JV, Saeed N. Cerebral ventricular asymmetry in schizophrenia: a high resolution 3D magnetic resonance imaging study. Int. J. Psychophysiol. 1999;34:207–211. doi: 10.1016/s0167-8760(99)00078-1. [DOI] [PubMed] [Google Scholar]
- Reiman EM, Chen K, Alexander GE, Caselli RJ, Bandy D, Osborne D, Saunders AM, Hardy J. Functional brain abnormalities in young adults at genetic risk for late-onset Alzheimer’s dementia. Proc. Natl. Acad. Sci. U. S. A. 2004;101:284–289. doi: 10.1073/pnas.2635903100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rohlfing T, Russakoff DB, Maurer CR. Expectation maximization strategies for multi-atlas multi-label segmentation. IPMI. 2003:210–221. doi: 10.1007/978-3-540-45087-0_18. [DOI] [PubMed] [Google Scholar]
- Rohlfing T, Brandt R, Menzel R, Maurer CR., Jr Evaluation of atlas selection strategies for atlas-based image segmentation with application to confocal microscopy images of bee brains. NeuroImage. 2004;21:1428–1442. doi: 10.1016/j.neuroimage.2003.11.010. [DOI] [PubMed] [Google Scholar]
- Seidman LJ, Faraone SV, Goldstein JM, Goodman JM, Kremen WS, Toomey R, Tourville J, Kennedy D, Makris N, Caviness VS, Tsuang MT. Thalamic and amygdala-hippocampal volume reductions in first degree relatives of schizophrenic patients: an MRI-based morphometric analysis. Biol. Psychiatry. 1999;46:941–954. doi: 10.1016/s0006-3223(99)00075-x. [DOI] [PubMed] [Google Scholar]
- Stern Y. Cognitive reserve and Alzheimer disease. Alzheimer Dis. Assoc. Disord. 2006;20:112–117. doi: 10.1097/01.wad.0000213815.20177.19. [DOI] [PubMed] [Google Scholar]
- Storey JD, Tibshirani R. Technical Report 2001–28 . Estimating false discovery rates under dependence, with applications to DNA microarrays. Department of Statistics, Stanford University; 2001. [Google Scholar]
- Storey JD. A direct approach to false discovery rates. J. R. Stat. Soc, Ser. B Stat. Method. 2002;64:479–498. [Google Scholar]
- Strittmatter WJ, Saunders AM, Schmechel D, Pericak-Vance M, Enghild J, Salvesen GS, Roses AD. Apolipoprotein E: high-avidity binding to beta-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc. Natl. Acad. Sci. U. S. A. 1993;90:1977–1981. doi: 10.1073/pnas.90.5.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studholme C, Cardenas V, Schuff N, Krishnan M, Chui H, Weiner MW. The effects of cerebrovascular and Alzheimer’s disease on patterns of ventricular expansion rate derived from serial MRI. ISMRM 2002 [Google Scholar]
- Styner M, Lieberman JA, McClure RK, Weinberger DR, Jones DW, Gerig G. Morphometric analysis of lateral ventricles in schizophrenia and healthy controls regarding genetic and disease-specific factors. Proc. Natl. Acad. Sci. 2005;102(12):4872–4877. doi: 10.1073/pnas.0501117102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svarer C, Madsen K, Hasselbalch SG, Pingborg LH, Haugbol S, Frokjer VG, Holm S, Paulson OB, Lnudsen GM. MR-based automatic delineation of volumes of interests in human brain PET images using probability maps. NeuroImage. 2005;24:969–979. doi: 10.1016/j.neuroimage.2004.10.017. [DOI] [PubMed] [Google Scholar]
- Thal LJ, Kantarci K, Reiman EM, Klunk WE, Weiner MW, Zetterberg H, Galasko D. The role of biomarkers in clinical trials for Alzheimer’s disease. Alzheimer Dis. Assoc. Disord. 2006;20:6–15. doi: 10.1097/01.wad.0000191420.61260.a8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson PM, Schwartz C, Lin RT, Khan AA, Toga AW. Three-dimensional statistical analysis of sulcal variability in the human brain. J. Neurosci. 1996a;16:4261–4274. doi: 10.1523/JNEUROSCI.16-13-04261.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson PM, Schwartz C, Toga AW. High-resolution random mesh algorithms for creating a probabilistic 3D surface atlas of the human brain. NeuroImage. 1996b;3:19–34. doi: 10.1006/nimg.1996.0003. [DOI] [PubMed] [Google Scholar]
- Thompson PM, Mega MS, Narr KL, Sowell ER, Blanton RE, Toga AW. Brain image analysis and atlas construction, invited chapter. In: Fitzpatrick M, Sonka M, editors. SPIE Handbook on Medical Image Analysis. Society of Photo-Optical Instrumentation Engineers (SPIE) Press; 2000. pp. 1063–1131. [Google Scholar]
- Thompson PM, Hayashi KM, de Zubicaray G, Janke AL, Rose SE, Semple J, Hong MS, Herman D, Gravano D, Doddrell DM, Toga AW. Mapping hippocampal and ventricular change in Alzheimer’s disease. NeuroImage. 2004;22:1754–1766. doi: 10.1016/j.neuroimage.2004.03.040. [DOI] [PubMed] [Google Scholar]
- Thompson PM, Dutton RA, Hayashi KM, Lu A, Lee SE, Lee JY, Lopez OL, Aizenstein HJ, Toga AW, Becker JT. 3D mapping of ventricular and corpus callosum abnormalities in HIV/AIDS. NeuroImage. 2006;31(1):12–23. doi: 10.1016/j.neuroimage.2005.11.043. [DOI] [PubMed] [Google Scholar]
- Wang L, Trein J, Gado M, Hosakere M, Miller MI, Csernansky JG. Volume, area and thickness abnormalities of the anterior cingulate in schizophrenia subjects. Annu. Meet. Soc. Neurosci. 2004 [Google Scholar]
- Wang YL, Gu X, Hayashi KM, Chan TF, Thompson PM, Yau ST. Surface parameterization using Riemann surface structure. Proc. ICCV2005. 2005:1061–1066. doi: 10.1007/11566489_81. [DOI] [PubMed] [Google Scholar]
- Woods RP. Multitracer: a Java-based tool for anatomic delineation of grayscale volumetric images. NeuroImage. 2003;19:1829–1834. doi: 10.1016/s1053-8119(03)00243-x. [DOI] [PubMed] [Google Scholar]
- Wolf H, Grunwald M, Kruggel F, Riedel-Heller SG, Angerhofer S, Hojjatoleslami A, Hensel A, Arendt T, Gertz HJ. Hippocampal volume discriminates between normal cognition; questionable and mild dementia in the elderly. Neurobiol. Aging. 2001;22:177–186. doi: 10.1016/s0197-4580(00)00238-4. [DOI] [PubMed] [Google Scholar]
- Yushkevich PA, Piven J, Hazlett C, Smith H, Smith G, Ho R, Ho S, Gee J, Gerig G. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. NeuroImage. 2006;31:1116–1128. doi: 10.1016/j.neuroimage.2006.01.015. [DOI] [PubMed] [Google Scholar]
- Zhou J, Rajapakse JC. Segmentation of subcortical brain structures using fuzzy templates. NeuroImage. 2005;28(4):915–924. doi: 10.1016/j.neuroimage.2005.06.037. [DOI] [PubMed] [Google Scholar]
- Zhu L, Jiang T. Parameterization of 3D brain structures for statistical shape analysis. Proc. SPIE Med. Imag. 2004;5370:1254–1261. [Google Scholar]
- Zöllei L, Learned-Miller E, Grimson E, Wells W. Efficient Population Registration of 3D Data. Proc ICCV, Computer Vision for Biomedical Image Applications; Beijing, China. 2005. [Google Scholar]