Abstract
Year by year, approximately two million people die from tuberculosis, a disease caused by the bacterium Mycobacterium tuberculosis. There is a tremendous need for new anti-tuberculosis therapies (antituberculotica) and drugs to cope with the spread of tuberculosis. Despite many efforts to obtain a better understanding of M. tuberculosis' pathogenicity and its survival strategy in humans, many questions are still unresolved. Among other cellular processes in bacteria, pathogenicity is controlled by transcriptional regulation. Thus, various studies on M. tuberculosis concentrate on the analysis of transcriptional regulation in order to gain new insights on pathogenicity and other essential processes ensuring mycobacterial survival. We designed a bioinformatics pipeline for the reliable transfer of gene regulations between taxonomically closely related organisms that incorporates (i) a prediction of orthologous genes and (ii) the prediction of transcription factor binding sites. In total, 460 regulatory interactions were identified for M. tuberculosis using our comparative approach. Based on that, we designed a publicly available platform that aims to data integration, analysis, visualization and finally the reconstruction of mycobacterial transcriptional gene regulatory networks: MycoRegNet. It is a comprehensive database system and analysis platform that offers several methods for data exploration and the generation of novel hypotheses. MycoRegNet is publicly available at http://mycoregnet.cebitec.uni-bielefeld.de.
INTRODUCTION
Year by year, approximately two million people die worldwide from tuberculosis (1) and one-third of the world's total population suffer from this communicable disease (http://www.who.int) caused by the bacterium Mycobacterium tuberculosis. Tuberculosis is the leading cause of death to people living with HIV and claims on average 200 000 lives every year, most of them in Africa. Persons infected with tuberculosis will not directly develop the characteristic full-blown clinical picture, but in most cases the latent form, which can progress to an active condition after years. About 10–15 people can be infected by a person with active tuberculosis a year, if she or he is left untreated (http://www.who.int). Although there is effective treatment to cure patients with tuberculosis, and new strategies have been developed to stop its further dissemination, its containment is still a serious problem (2). The number of multi-resistant strains not responding to standard drug treatments is increasing constantly worldwide (3,4). Consequently, there is a tremendous need for new anti-tuberculosis therapies (antituberculotica) and drugs to cope with the spread of tuberculosis. Despite many efforts to obtain a better understanding of the pathogenicity of M. tuberculosis and its survival strategy in humans, many questions are still unresolved. The molecular mechanisms responsible for resisting the human immune system and their activation are not perceived sufficiently so far; most notably, its ability to remain within the human host for years in a clinically latent state (5). Among other cellular processes in bacteria, pathogenicity is controlled by transcriptional regulation. Thus, various studies on M. tuberculosis concentrate in the analysis of transcriptional regulation in order to gain new insights on pathogenicity and other essential processes ensuring mycobacterial survival. The identification and characterization of transcriptional regulation on a genome-wide level will enable a better understanding of drug metabolism in M. tuberculosis and facilitate the development of new antibiotics, which are urgently needed. At present, studies focus mainly on the analysis of single regulons, or distinct subunits of the complex transcriptional regulatory network of M. tuberculosis [see e.g. (3,5,6)]. Bioinformatics platforms for data storage and public access of transcriptional regulation exist for M. tuberculosis, similar to other organisms such as Escherichia coli [RegulonDB (7)] or Corynebacterium glutamicum [CoryneRegNet (8)]. MtbRegList (9) and MTBreg (http://www.doe-mbi.ucla.edu/services/mtbreg) offer information relevant to regulatory interactions in M. tuberculosis H37Rv (MT) accumulated from literature or attained from computational predictions. While MtbRegList contains predicted and characterized regulatory DNA motifs cross-referenced with transcription factors (TFs), MTBreg combines a collection of conditionally regulated proteins together with information about selected TFs. However, both systems are designed as data repositories and only provide nonsatisfying bioinformatics support necessary for transcriptional gene regulatory network visualization, analysis, and reconstruction. Recently, the TB database has become available. This integrated online platform for tuberculosis research combines the annotated genome and expression data with a suite of bioinformatic tools for data analysis (10). The scope of TB database is placed on investigating and providing expression data, while little support is given for the reconstruction of regulatory networks based on these findings. Hence, there is currently no online platform or database system available, which aims to an appropriate data handling and analysis of transcriptional regulation in M. tuberculosis on a genome-wide level.
Here, we introduce MycoRegNet, an online accessible, user-friendly platform dedicated to the biomedical researcher, who is interested in the regulation of gene expression in the human pathogen M. tuberculosis. MycoRegNet is online available at http://mycoregnet.cebitec.uni-bielefeld.de.
The first idea of our approach is based on the assumption that orthologous TFs tend to regulate the expression of orthologous target genes for taxonomically closely related species (11–13). Corynebacterium glutamicum and M. tuberculosis are taxonomically classified into the suborder Corynebacterineae of the Actinobacteria phylum and are thus taxonomically closely related (14). Hence, the industrially important amino acid producer C. glutamicum has been successfully applied as model organism, e.g. for investigating cell envelope synthesis of M. tuberculosis (15–17). We therefore started with the well-examined regulatory network of C. glutamicum ATCC 13032 (CG) (18), which is stored in the corynebacterial reference database CoryneRegNet (8). Our comparative genomics approach aims for a reliable transfer of known regulatory interactions from CG to MT. Instead of relying exclusively on the detection of orthologous genes, we consider further evidence by means of an integrated TF binding site (TFBS) prediction. The resulting data were subsequently stored in an online platform designed for the visualization and analysis of the deduced transcriptional regulatory network, which enables the execution of bioinformatics tools for further hypotheses generation: MycoRegNet. The remainder of this article is structured as follows: we first describe the workflow used for the transfer of C. glutamicum data to M. tuberculosis in detail. The design of MycoRegNet is briefly introduced afterwards. It aims to overcome typical data integration problems and to supply online visualization and hypotheses generation tools. In the last section, we illustrate and discuss these functionalities. We finally conclude that MycoRegNet is an appropriate reference database and platform for gene regulatory network analysis of M. tuberculosis.
MATERIALS AND METHODS
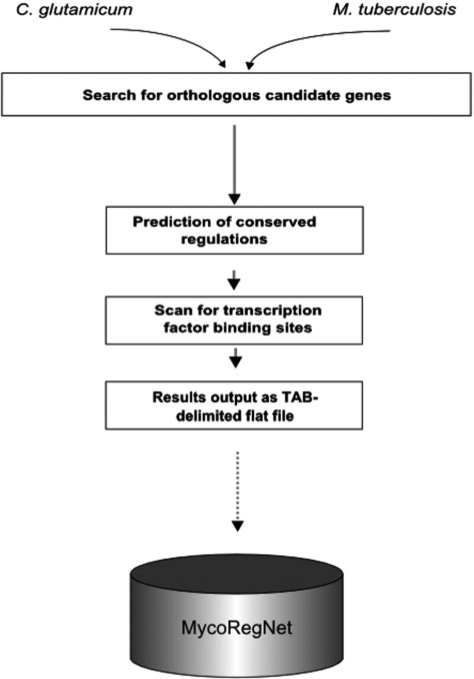
The network reconstruction pipeline mainly consists of the detection of (i) conserved genes between CG and MT and (ii) binding sites upstream the conserved genes in MT. Based on the corresponding results, a list of putative gene regulatory interactions in MT is generated and imported into the MycoRegNet database back-end (see Figure 1 for a graphical overview of the workflow).
Figure 1.
Diagram of the prediction pipeline. The diagram shows the main steps performed during transfer of gene regulations from C. glutamicum to M. tuberculosis. Starting with an orthology detection, the next step was a prediction of conserved regulations. Based on that, a TFBSs prediction provided further evidence. Finally, the results can be exported as TAB-delimited files and imported into the MycoRegNet data repository.
Detection of orthologous genes
Generally, the detection of orthologous genes is not straightforward, since analysis can be perturbed by factors like paralogs or sequence divergences in the genomes of interest. To reduce such effects, we searched for orthologous genes by performing bidirectional BLASTP (19) searches on the corresponding protein sequences. Therefore, we scanned the CG genome for sequence similarities with the MT genome and vice versa, performing BLASTP with an E-value cut-off of 10−4 in both directions. As a result, we obtained amino acid sequence pairs, so called bidirectional best hits (BBHs), representing the reciprocal best alignments of respective protein sequences. Thus, identified BBHs were considered to be putative orthologous proteins in CG and MT, which in turn indicates the respective genes to be regulated in both bacteria by orthologous TFs.
Transfer of regulatory interactions
Based on the previously identified BBHs, regulatory interactions characterized in CG were transferred to MT. We utilized the comprehensive data on transcriptional regulation in CG collected in the corynebacterial reference database CoryneRegNet (8), which contains 806 regulatory interactions of 72 TFs and 544 regulated target genes on CG (status: January 2009). For each regulatory interaction taken from CG, both the gene encoding the TF and the target gene were compared to the list of predicted orthologs in the MT genome. Only if both, the TF as well as its target gene, were identified as BBH, the regulatory interaction was transferred from CG to the orthologous counterparts in MT and was considered as a candidate transcriptional regulation in MT. Furthermore, we assume the regulatory role of the TF (activation or repression) to be conserved as well, including known autoregulations.
Further evidence through conserved TFBSs
In the last step of our regulatory network prediction pipeline, we add further evidence to the orthology-based approach introduced above by combining the preliminary results with the prediction of TFBSs. Therefore, all known binding sites of characterized TFs of CG with potential orthologs in MT were utilized to create appropriate motif profiles. TF binding motifs were modeled as so called position weight matrices (PWMs), the most widely used model for that purpose. However, we applied only PWMs of corynebacterial TFs deduced from more than 20 binding sites, i.e. of the TFs GlxR, RamB, AmtR, DtxR and LexA. To detect instances of the respective motifs in MT, we employed the TFBS matching tool PoSSuMsearch (20) and scanned 580-bp long, noncoding DNA sequences upstream all genes and operons, which have been detected as potential orthologs to target genes of the respective TF. The upstream sequences ranged from +20 bp relative to the transcription start. In our initial approach, we performed a restrictive search by setting the P-value threshold to 10−5. Due to the low number of detected binding sites in the first PoSSuMsearch runs, we decided to decrease the P-value threshold since the set P-value might be chosen too restrictive for our TFBS predictions. To determine a new P-value, we considered P-values of binding site matches of the PWM for GlxR upstream 26 target genes as marking value, where the binding of the GlxR ortholog Rv3676 in MT was experimentally verified (21–24). For P-value definition, we chose the binding site that match upstream one of these genes with the worst P-value. Thus, we finally set the P-value to <10−2 and defined for each target gene/operon in MT the TFBS match with the lowest P-value as prediction for the respective binding site.
Taken together, the outcome of the above introduced workflow is a list of transcriptional regulations for MT where (i) the TF is conserved, and (ii) the target gene is conserved between CG and MT as well, and additionally (iii) a binding site is predicted, if the target gene/operon is controlled by one of the five TFs where a TFBS search was performed for. Hence, the resulting predictions present most likely regulatory interactions in MT due to the taxonomically close relation between CG and MT. This is the data we aim to integrate into the MycoRegNet platform together with validated knowledge we have from (5,25–34).
Data integration with the MycoRegNet platform
Based on our experiences with CoryneRegNet, we designed MycoRegNet in a very similar way: as an ontology-based data warehouse for mycobacterial TFs and regulatory networks. We set it up as a sister project of CoryneRegNet to store, analyze and visualize the regulatory interactions in M. tuberculosis that are derived from the above introduced prediction pipeline. MycoRegNet is composed of two main parts: (i) A web front-end running on an Apache HTTP web server that manages user-database interactions as well as the execution of further online bioinformatics computations. (ii) The back-end consists of data preprocessing tools and a MySQL database that stores all data corresponding to the deduced and ontologically restructured mycobacterial gene regulatory interactions. This process comprises the integration of transcriptional regulations, the complete genome sequence of MT along with the genome annotation as stored in the GenBank database (NCBI) (35), operon predictions available from the Virtual Institute of Microbial Stress and Survival (VIMSS) (36), precalculated PWMs and other preprocessed data necessary for subsequent online TFBS detections, and stimulons derived from literature (25–31). The import and conversion software is implemented in Java, while the web pages generated at front-end level are developed in PHP. An embedded Java applet realizes the visualization of gene regulatory networks from the included data. A SOAP-based web Service (37) client/ server system implemented by means of NuSOAP enables a bidirectional interconnection with GenDB (38) and EMMA (39). The server is open access and provides well-structured data access via the SOAP interface to any other bioinformatics client. GenDB is an open source system for the annotation of prokaryotic genomes, while EMMA is a web-based application for the storage and analysis of transcriptomics data from microarrays. By means of the clients for GenDB and EMMA, data integrated in MycoRegNet is supplemented with up-to-date information on the genome annotation of MT (GenDB) and gene expression data preanalyzed with EMMA. To give one example, the Web Service client for GenDB facilitates the mapping of all genes controlled by a certain regulator to KEGG pathways (40) in order to provide an overview on the general nature of a TF of interest. Furthermore, the automatic annotation pipeline of GenDB can be used to regularly update gene function assignments.
RESULTS AND DISCUSSION
Here, we first summarize the database content. Subsequently, we present and discuss the benefits of MycoRegNet from the end-user perspective. We first describe the web interface with special attention to the TFBS prediction feature and the network visualization and analysis capability. We briefly describe the Web Service access afterwards and finally demonstrate the platforms' visualization functionality by means of an application example.
The database content
By using the above described transfer pipeline for regulatory interactions, we identified 1012 of 3991 proteins from MT as putative orthologs to proteins from CG. Based on the respective set of genes coding for the orthologous proteins, we detected 226 of 806 regulatory interactions from CG as likely conserved in MT (Table 1). Our initial findings reveal 24 partial conserved regulons affecting processes of the carbohydrate metabolism, cellular program, macroelement and metal homeostasis, SOS and stress response, specific biosynthesis as well as processes governed by sigma factors. By setting the P-value threshold to 102, we could put further evidence to 129 target genes 40 by predicting binding sites upstream the respective target genes/operons regulated by the TF orthologs of GlxR, RamB, AmtR, DtxR and LexA in MT (Table 2). All in all, we obtained a set of regulatory interactions which is based on good evidence. The database content comprises 618 regulatory interactions for 515 target genes regulated by 26 TFs. Several gene expression experiments are also directly stored within MycoRegNet's database back-end (data not shown). We also integrated genome annotation data of M. tuberculosis CDC1551 for future investigations concerning transcriptional regulation in another ecotype of M. tuberculosis.
Table 1.
Putative gene regulations of CG in MT
| TF | Target genes | |
|---|---|---|
| Carbohydrate metabolism | ||
| Rv0465c | Carbohydrate metabolism | |
| Rv0211 (pckA), Rv0247c (-), Rv0363c (fba), Rv0408 (pta) | ||
| Rv0409 (ackA), Rv0465c (-), Rv0467 (icl), Rv0896 (gltA2) | ||
| Rv0904c (accD3), Rv0951 (sucC), Rv0952 (sucD), Rv1475c (acn) | ||
| Rv1837c (glcB), Rv1862 (adhA), Rv2193 (ctaE), Rv2241 (aceE) | ||
| Rv2332 (mez), Rv2967c (pca), Rv3318 (sdhA) | ||
| Cell division and septation | ||
| Rv1009 (rpfB) | ||
| Specific biosynthesis pathways | ||
| Rv0884c (serC), Rv1010 (ksgA), Rv1011 (ispE), Rv1379 (pyrR) | ||
| Rv1380 (pyrB), Rv1381 (pyrC) | ||
| Rv0792c | Carbohydrate metabolism | |
| Rv0753c (mmsA) | ||
| Rv1719 | Carbohydrate metabolism | |
| Rv0554 (bpoC), Rv1074c (fadA3), Rv1719 (-), Rv2503c (scoB), Rv2504c (scoA) | ||
| Rv3676 | Carbohydrate metabolism | |
| Rv0211 (pckA), Rv0247c (-), Rv0400c (fadE7), Rv0465c (-) | ||
| Rv0467 (icl), Rv0896 (gltA2), Rv0904c (accD3), Rv0951 (sucC) | ||
| Rv0952 (sucD), Rv1098c (fumC), Rv1130 (-), Rv1161 (narG) | ||
| Rv1162 (narH), Rv1163 (narJ), Rv1436 (gap), Rv1437 (pgk) | ||
| Rv1438 (tpi), Rv1475c (acn), Rv1837c (glcB), Rv1854c (ndh) | ||
| Rv1862 (adhA), Rv1872c (lldD2), Rv2029c (pfkB), Rv2193 (ctaE) | ||
| Rv2194 (qcrC), Rv2195 (qcrA), Rv2196 (qcrB), Rv2200c (ctaC) | ||
| Rv2524c (fas), Rv2967c (pca), Rv3010c (pfkA), Rv3043c (ctaD) | ||
| Rv3279c (birA), Rv3280 (accD5), Rv3318 (sdhA), Rv3548c (-), Rv3676 (-) | ||
| Cell division and septation | ||
| Rv1009 (rpfB), Rv2145c (wag31), Rv2201 (asnB) | ||
| Macroelement and metal homeostasis | ||
| Rv0820 (phoT), Rv0928 (pstS3), Rv0929 (pstC2), Rv0930 (pstA1), Rv2220 (glnA1) | ||
| Rv2832c (ugpC), Rv2833c (ugpB), Rv2834c (ugpE), Rv2835c (ugpA), Rv2918c (glnD) | ||
| Rv2919c (glnB), Rv2920c (amt), Rv3859c (gltB) | ||
| SOS and stress response | ||
| Rv0867c (rpfA), Rv3048c (nrdF2), Rv3217c(-), Rv3219 (whiB1), Rv3681c (whiB4) | ||
| Specific biosynthesis pathways | ||
| Rv0884c (serC), Rv1010 (ksgA), Rv1011 (ispE), Rv1092c (coaA) | ||
| Rv3001c (ilvC) | ||
| Rv3002c (ilvN), Rv3003c (ilvB1) | ||
| Cellular Program | ||
| RelA | Sigma factor module | |
| Rv1221 (sigE), Rv2710 (sigB), Rv3221A (-), Rv3911 (sigM) | ||
| SOS and stress response | ||
| Rv2720 (lexA) | ||
| Macroelement and metal homeostasis | ||
| Rv0485 | Macroelement and metal homeostasis | |
| Rv0132c (fgd2) | ||
| PhoP | Macroelement and metal homeostasis | |
| Rv0545c (pitA), Rv0757 (phoP), Rv0758 (phoR), Rv0820 (phoT) | ||
| Rv0928 (pstS3), Rv0929 (pstC2), Rv0930 (pstA1), Rv1095 (phoH2) | ||
| Rv2832c (ugpC), Rv2833c (ugpB), Rv2834c (ugpE), Rv2835c (ugpA) | ||
| Rv0827c | Macroelement and metal homeostasis | |
| Rv0827c (-) | ||
| Rv1994c | Macroelement and metal homeostasis | |
| Rv1994c (-) | ||
| IdeR | Carbohydrate metabolism | |
| Rv0247c (-), Rv3318 (sdhA) | ||
| Macroelement and metal homeostasis | ||
| Rv0827c (-), Rv0844c (narL), Rv1285 (cysD), Rv1286 (cysN) | ||
| Rv2391 (nirA), Rv2392 (cysH), Rv2393 (-), Rv2895c (viuB) | ||
| Rv3044 (fecB), Rv3841 (bfrB) | ||
| Rv3160c | Macroelement and metal homeostasis | |
| Rv1848 (ureA), Rv1849 (ureB), Rv1850 (ureC), Rv1852 (ureG) | ||
| Rv2220 (glnA1), Rv2918c (glnD), Rv2919c (glnB), Rv2920c (amt) | ||
| Rv3664c (dppC), Rv3665c (dppB), Rv3666c (dppA), Rv3859c (gltB) | ||
| Rv3173c | Macroelement and metal homeostasis | |
| Rv0132c (fgd2), Rv0485 (-), Rv1079 (metB), Rv1133c (metE) | ||
| Rv1175c (fadH), Rv1285 (cysD), Rv1286 (cysN), Rv1294 (thrA) | ||
| Rv1296 (thrB), Rv1392 (metK), Rv2124c (metH), Rv2334 (cysK1) | ||
| Rv2391 (nirA), Rv2392 (cysH), Rv2393 (-), Rv3025c (iscS) | ||
| Rv3028c (fixB), Rv3029c (fixA), Rv3173c (-), Rv3340 (metC) | ||
| Rv3341 (metA) | ||
| Sigma factor module | ||
| SigB | Carbohydrate metabolism | |
| Rv0363c (fba), Rv1023 (eno), Rv1098c (fumC), Rv1436 (gap) | ||
| Rv1437 (pgk), Rv1438 (tpi), Rv3010c (pfkA) | ||
| SOS and stress response | ||
| Rv3132c (devS) | ||
| Specific biosynthesis pathways | ||
| Rv2210c (ilvE) | ||
| SigM | SOS and stress response | |
| Rv0384c (clpB), Rv1464 (csd), Rv1465, Rv1471 (trxB1) | ||
| Rv3418c (groES), Rv3913 (trxB2), Rv3914 (trxC) | ||
| SOS and stress response | ||
| HspR | SOS and stress response | |
| Rv0350 (dnaK), Rv0351 (grpE), Rv0352 (dnaJ1), Rv0353 (hspR) | ||
| Rv0384c (clpB), Rv0440 (groEL), Rv2745c | ||
| HrcA | SOS and stress response | |
| Rv0440 (groEL), Rv3418c (groES) | ||
| LexA | Cell division and septation | |
| Rv2748c (ftsK) | ||
| SOS and stress response | ||
| Rv1235 (lpqY), Rv1638 (uvrA), Rv1696 (recN), Rv2592c (ruvB) | ||
| Rv2593c (ruvA), Rv2594c (ruvC), Rv2720 (lexA), Rv2736c (recX) | ||
| Rv2737c (recA), Rv3370c (dnaE2), Rv3395c, Rv3585 (radA) | ||
| Rv2745c | SOS and stress response | |
| Rv0782 (ptrBb), Rv2460c (clpP2), Rv2461c (clpP1), Rv2725c (hflX) | ||
| Rv3596c (clpC1), Rv3715c (recR), Rv3716c | ||
| WhiB1 | SOS and stress response | |
| Rv3913 (trxB2), Rv3914 (trxC) | ||
| MtrA | SOS and stress response | |
| Rv0917 (betP), Rv3476c (kgtP) | ||
| CspA | Carbohydrate metabolism | |
| Rv1837c (glcB) | ||
| Specific biosynthesis pathways | ||
| PyrR | Specific biosynthesis pathways | |
| Rv1379 (pyrR), Rv1380 (pyrB), Rv1381 (pyrC), Rv2883c (pyrH) | ||
| ArgR | Specific biosynthesis pathways | |
| Rv1383 (carA), Rv1384 (carB), Rv1652 (argC), Rv1653 (argJ) | ||
| Rv1654 (argB), Rv1655 (argD), Rv1656 (argF), Rv1657 (argR) | ||
| Rv0488 | ||
Putative gene regulations of CG in MT, predicted in silico by using the introduced MycoRegNet pipeline
Table 2.
Detected binding sites upstream transferred target genes of CG in MT
| TF | Gene ID | Gene name | Operon | Binding motif |
|---|---|---|---|---|
| Rv0465c | Rv0211a | pckA | ATAACTACGCAGG | |
| Rv0249c | – | Rv0249c-Rv0248c-Rv0247ca | AGTAGTTCGCGAT | |
| Rv0363ca | fba | – | CGTACTTCTCAAA | |
| Rv0407 | pta | Rv0407-Rv0408a-Rv0409a | CGTGCTGTGCTCA | |
| Rv0465c | – | Rv0465ca-Rv0464c | CTAACTCTGCGAA | |
| Rv0467a | icl | – | CAAAATTTGCAAA | |
| Rv0884ca | serC | – | ATGGCATGGCCGA | |
| Rv0896a | gltA2 | – | TGAGCAGATCACT | |
| Rv0904ca | accD3 | – | ATTGCATGGCAAG | |
| Rv0951 | sucC | Rv0951a-Rv0952a | AGTGCTAAGCCGT | |
| Rv1009 | rpfB | Rv1009a-Rv1010a-Rv1011a | TCTACTTACCAAA | |
| Rv1379 | pyrR | Rv1379a-Rv1380a-Rv1381a-Rv1382-Rv1383-Rv1384-Rv1385 | AGTGCTACGCTGC | |
| Rv1475c | acn | Rv1475ca-Rv1474c | ACTGCTAGGCTGA | |
| Rv1837ca | glcB | – | TAGGCTGAGCAAT | |
| Rv1862a | adhA | – | TGTGCTGGGCTAA | |
| Rv2193 | ctaE | Rv2193a-Rv2194-Rv2195-Rv2196 | ACTACAAAGCGTC | |
| Rv2241 | aceE | Rv2241a-Rv2242 | CAAACAGCGCAAG | |
| Rv2332a | mez | – | TGCGCTCTGCGAA | |
| Rv2967ca | pca | – | CATGCAATGTCAA | |
| Rv3316 | sdhA | Rv3316-Rv3317-Rv3318a-Rv3319 | GTTGCATTGCCCC | |
| IdeR | Rv0249c | – | Rv0249c-Rv0248c-Rv0247ca | TTAGATGAGCGCACCCACG |
| Rv0827ca | – | – | CTATGGATCGCTGTACTAC | |
| Rv0844ca | narL | – | CGACGAGCAGCTAAACTCA | |
| Rv1285 | cysD | Rv1285a-Rv1286a | GAGGGCGAGGCACACGTCA | |
| Rv2391 | nirA | Rv2391a-Rv2392a-Rv2393a | TCAGGTGCGCGTCTCCCAG | |
| Rv2895ca | viuB | – | TAAGCGAAGCCGAACGCCA | |
| Rv3044a | fecB | – | GTAGACCAGGCTCCCCTTG | |
| Rv3316 | sdhA | Rv3316-Rv3317-Rv3318a-Rv3319 | CTAAGAAAAGCCAGCCTAA | |
| Rv3841a | bfrB | – | CTAGGAAAGCCTTTCCTGA | |
| LexA | Rv1235 | lpqY | Rv1235a-Rv1236-Rv1237-Rv1238 | TCGACTATCTATCCGA |
| Rv1638a | uvrA | – | TCGAATGTCAGCTCGC | |
| Rv1696a | recN | – | ||
| Rv2594c | ruvC | Rv2594ca-Rv2593ca-Rv2592ca | TCGAACGATTGTTCGG | |
| Rv2720a | lexA | – | TCGAACACATGTTTGA | |
| Rv2737c | recA | Rv2737ca-Rv2736ca | TCGAACAGGTGTTCGG | |
| Rv2748ca | ftsK | – | CCGACCAGGTGCTCGC | |
| Rv3370ca | dnaE2 | – | TCGAACAATTGTTCGA | |
| Rv3395c | – | Rv3395ca-Rv3394c | TCGAACATATTTTCGA | |
| Rv3160c | Rv1848 | ureA | Rv1848a-Rv1849-Rv1850a-Rv1851-Rv1852a-Rv1853 | GTGTCTACTGCGCGATGATCGAGAGCAT |
| Rv2220a | glnA1 | – | CAACACGGGGTTGACTGACGGGCAATAT | |
| Rv2920c | amt | Rv2920ca-Rv2919ca-Rv2918ca | AAGTTTTACGTTAATCCTGATGAAACAT | |
| Rv3666c | dppA | Rv3666ca-Rv3665c-aRv3664ca-Rv3663c-Rv3662c | GTGGTAGCTAACGGTCACCGGCGAGTGT | |
| Rv3859c | gltB | Rv3859ca-Rv3858c | CGCTTGACGGACAGCCTATCGACAAGAC | |
| Rv3676 | Rv0211a | pckA | TGTGAGCAGGCTTATA | |
| Rv0249c | – | Rv0249c-Rv0248c-Rv0247ca | TGTGATCTGTAACACC | |
| Rv0400ca | fadE7 | – | AGTGATGAGCACCCCG | |
| Rv0465c | – | Rv0465ca-Rv0464c | TTTGTCGAGGCTCACG | |
| Rv0467a | ivl | – | TGTTACAACGCTCACA | |
| Rv0820a | phoT | GGTGGTGATCCGCACC | ||
| Rv0867ca | rpfA | TGTGACATTACCCACA | ||
| Rv0884ca | serC | TGTGAGCTTGTTCACA | ||
| Rv0896a | gltA2 | – | GGCGTTGAACATCACC | |
| Rv0904ca | accD3 | – | CGTGAGTCGTATCACG | |
| Rv0928 | pstS3 | Rv0928a-Rv0929a-Rv0930a | ACTGAATTGAAACTCA | |
| Rv0951 | sucC | Rv0951a-Rv0952a | TGTGAGTTGGATCACG | |
| Rv1009 | rpfB | Rv1009a-Rv1010a-Rv1011a | GGTGGCGCTCATCACC | |
| Rv1092ca | coaA | TGCCACGTAGGTCACG | ||
| Rv1099c | fumC | Rv1099c-Rv1098ca-Rv1097c | ||
| Rv1130 | – | Rv1130a-Rv1131 | TGTGGATAAGTCCAGG | |
| Rv1161 | narG | Rv1161a-Rv1162a-Rv1163a-Rv1164-Rv1165-Rv1166 | TGCGTTGAACGGCACG | |
| Rv1436 | gap | Rv1436a-Rv1437a-Rv1438a | GGTTGTTTAGCCAACA | |
| Rv1475c | acn | Rv1475ca-Rv1474c | TGTAACTGCCGACATA | |
| Rv1837ca | glcB | – | AGGGATGCACTACACA | |
| Rv1854ca | ndh | – | TGTGGCTGATGACACA | |
| Rv1862a | adhA | – | CGTGGGGCGCCACACA | |
| Rv1872ca | lldD2 | – | GATGCCGTAGCGCACT | |
| Rv2029c | pfkB | Rv2029ca-Rv2028c-Rv2027c-Rv2026c | GGTGACGAGTCGCGCA | |
| Rv2145c | wag31 | CGTGACTGGCGTCCCA | ||
| Rv2193 | ctaE | Rv2193a-Rv2194a-Rv2195a-Rv2196a | GGTGGATAGGTTCACC | |
| Rv2200c | ctaC | Rv2200ca-Rv2199c | TGTGATACAGGAGGCG | |
| Rv2201 | asnB | GCTGTCGAAGACCACG | ||
| Rv2220a | glnA1 | TGTGACGGAAAAGACG | ||
| Rv2524ca | fas | – | CGTTACCCACGACACG | |
| Rv2835c | ugpA | Rv2835ca-Rv2834ca-Rv2833ca-Rv2832ca | GGTGATGCCGGGCACG | |
| Rv2920c | am | Rv2920ca-Rv2919ca-Rv2918ca | AGTGGACCAATTCCCC | |
| Rv2967ca | pca | – | CGTGGTGGTGGTCACC | |
| Rv3003ca | ilvB1 | Rv3003ca-Rv3002ca-Rv3001ca | TGTGGTGGCCACCCCA | |
| Rv3010ca | pfkA | – | GGTGATGGCGATGACC | |
| Rv3043c | ctaD | Rv3043ca-Rv3042c | AGTGGATCGCATCCCG | |
| Rv3048ca | nrdF2 | GGTGACTGGAAACGCA | ||
| Rv3217ca | – | TGTGGTGGCGGTCGCA | ||
| Rv3219a | whiB1 | AGTGAGATAGCCCACG | ||
| Rv3279c | birA | Rv3279ca-Rv3278c | TATCGGCTGCCGCACA | |
| Rv3280 | accD5 | Rv3280-aRv3281-Rv3282 | CGGGACGTCGACCACA | |
| Rv3316 | sdhA | Rv3316-Rv3317-Rv3318a-Rv3319 | CGAGACGTTTTCCACG | |
| Rv3549c | – | Rv3549c-Rv3548ca | GGTGATCGGCATTGCA | |
| Rv3676a | – | TGTCACCTACGACAGA | ||
| Rv3681ca | whiB4 | TGAGATACAGGTAACA | ||
| Rv3859c | gltB | Rv3859ca-Rv3858c | TGCTCCGGATTTCACA |
Detected binding sites of GlxR (ortholog in MT: Rv3676/Crp), RamB (ortholog in MT: Rv0465c), AmtR (ortholog in MT: Rv3160c), DtxR (ortholog in MT: IdeR/Rv3173c) and LexA (ortholog in MT: Rv2720/LexA) orthologs of CG in MT. Code:
aTransferred target gene of CG in MT.
The user interface
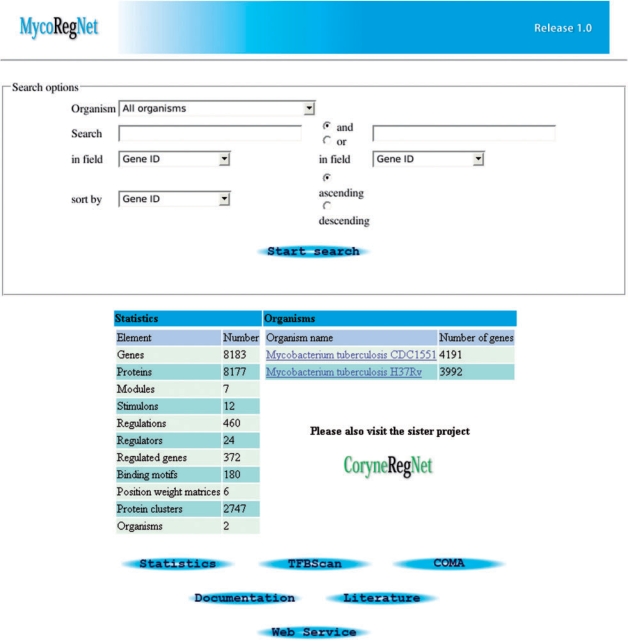
As for other online databases, MycoRegNet's web interface provides the three major capabilities: browsing the database content, searching by specifying filter criteria and basic visualization possibilities. Furthermore, the front-end offers the execution of computational features. At the main page (Figure 2), one has the option to search or to browse the database content. The user may browse the data repository by clicking on an ecotype name of interest and is provided with an overview on the selected organism. Alternatively, using one of the provided options within the search form, the database can be searched for specific gene/protein identifiers, gene/protein names, regulator types or functional modules. The search results are presented in tabular form, listing all relevant information for subsequent investigation. Furthermore, the following built-in features can be accessed from the main page, directly: TFBScan [for TFBS predictions; see below) and COMA (to check for contradictions within microarray gene expression studies, given the regulatory network stored in the database; refer to (8) for more details]. Detailed information on the results can be obtained via respective links at the result page. By selecting a particular gene, the corresponding gene details page is invoked. It presents a detailed overview of all available data attached to the gene of interest. Besides general information about the gene/protein (position in the genome, nucleotide sequence, etc.), it comprises a graphical representation of the genomic context, regulated target genes (if encoding a TF) including the TFBSs, etc., and stimulons that initiate a differential gene expression level. The integrated Web Service client for GenDB maintains the representation of up-to-date gene annotation data. General information (description, comments, an assigned function, etc.) is listed as well as the EC numbers for enzymes, and links to COG (41) and GO (42). Additionally, all target genes of a TF of interest, are linked to KEGG pathways and a list of regulated pathways is displayed.
Figure 2.
MycoRegNet main page. The main page includes a typical search mask, a statistical overview of the database content, an entry point to browse the integrated organisms, and links to more specific statistics, the system documentation and a tutorial on how to use the MycoRegNet Web Service.
TFBS prediction
With the integrated PoSSuMsearch software, MycoRegNet provides a statistically sound tool for the prediction of TFBSs based on PWMs, which have been precalculated during data import. To our knowledge, PoSSuMsearch is the only TFBSs profiling tool that offers exact P-value calculations and at the same time provides reasonable response times on genome-wide runs. There are three ways to access this feature through the MycoRegNet web site: (1) The TFBScan button at the main page offers the possibility to upload user-defined sequences in FASTA format. (2) At any gene details page the user can predict TFBSs in the upstream sequence of the selected gene. (3) If the gene of interest encodes a TF, the PWM learned from the known TFBSs of the TF may be used to scan for further TFBSs in the upstream sequences of all other mycobacterial genes. The predicted results may further be visualized as graphs. The interface is easy to use: one just has to choose a background model (nucleotide distribution) and a P-value threshold. For further details regarding the prediction of prokaryotic TFBSs by utilizing PoSSuMsearch, the reader is referred to (20,43).
Gene regulatory network visualization
As mentioned earlier, MycoRegNet also provides a network visualization toolkit: GraphVis. It is a Java applet, which graphically reconstructs regulatory networks as graphs based on selected genes and a user-defined graph depth cutoff. It traverses all regulatory interactions from the starting point until the graph depth cutoff has been reached. Finally, a Java applet window appears showing the regulatory network as graph, where nodes represent genes and edges regulatory interactions. GraphVis allows the user to zoom into the graph, apply different layout styles, remove selected elements or retrieve detailed information on selected genes. Furthermore, it is possible to extend the graph dynamically with more genes/regulations from the database and to display the operon grouping of presented genes. Visualized networks may also be graphically compared between two species or between a predicted and an evidenced network by utilizing special comparative graph layout algorithms. In addition, GraphVis features the projection of gene expression data onto the genes of a visualized network. The user can choose to apply gene expression data from the stimulon repository of MycoRegNet or from own tab-delimited text or MS-Excel files, which can be uploaded to GraphVis directly. It is also possible to use expression data extracted from EMMA by means of the integrated Web Service interconnection. According to the differential expression level of the genes, the concerned nodes are resized within the graph. Thus, the user can achieve a comprehensive overview of the transcriptional response of M. tuberculosis to a certain stimulus.
Well-structured data access by using Web Services
Although no real standard in bioinformatics, a growing number of platforms offer SOAP-based Web Service access to their data repositories [refer to some EBI resources (44) or to the BRENDA database (45), just to name two of them]. Many databases still provide flat files for exchange with other data processing systems. Thus, the developers of novel tools and platforms have to perform updates in regular time intervals and to adjust the downloaded data for their special purpose. On that account, gene regulatory data stored in MycoRegNet can also be accessed via the integrated Web Service server. The data can be integrated directly into corresponding projects without further time-consuming data processing. Detailed information on how to use the MycoRegNet Web Services is available from the main page via the Web Service button.
Application example—the regulatory network of the GlxR ortholog Rv3676 (CrpMT) in MT
Both GlxR (Cg0350) of C. glutamicum and CRPMT (Rv3676) of M. tuberculosis belong to the Crp-Fnr family of TFs (46) and have been characterized as cAMP sensing homologs of E. coli Crp (23,47,48). Crp-cAMP-dependent gene regulation is commonly involved in carbon catabolite repression and forms one of the possible connections between carbon metabolism and virulence (49,50).
In mycobacteria, cAMP signalling is the subject of intensive research, as it may be related to virulence of these strains (51,52). It is noteworthy that M. tuberculosis contains 16 putative adenylate cyclases, as well as 10 putative cyclic nucleotide binding proteins (53,54), hinting at a crucial and diverse role for cAMP signalling in mycobacteria.
GlxR of C. glutamicum has been in the focus of interest in the last years (48,55–59), and available data indicates GlxR as global regulator with about 150 target genes in functional diverse network modules, such as carbohydrate metabolism, aerobic and anaerobic respiration, fatty acid metabolism, aromatic compound degradation, glutamate uptake and nitrogen assimilation, the cellular stress response and resuscitation.
Previous studies suggested a similar vital role for CrpMT in M. tuberculosis. Published data implicate CrpMT in virulence, hypoxia and nutrient starvation (21,23,24,60). Deletion of CrpMT altered the expression of 16 genes, and caused an impaired growth phenotype in bone marrow-derived macrophages as well as in tuberculosis mouse models (24). Several suggestions for a putative CrpMT regulon have been made, although these studies relied solely on the detection of putative binding sites (23,24,60).
As part of our pipeline, the known regulatory interactions of GlxR collected in CoryneRegNet have been used to reconstruct the regulon of the orthologous TF CrpMT. Due to the apparent vital role of these regulators in their respective organisms, and the available data on putative target genes and characterized binding sites, we chose them as application case for our analysis. Employing our pipeline, regulatory interactions with 64 target genes could be transferred from C. glutamicum GlxR to M. tuberculosis CrpMT. Furthermore, we considered 26 genes with experimental evidence of regulation by CrpMT as potential target genes (21–24).
Based on experimentally verified binding sites of CrpMT (21–23) together with binding sites predicted by the TFBS search of our pipeline, we complemented the suggested regulon with the prediction of CrpMT binding sites in the upstream regions of putative target genes.
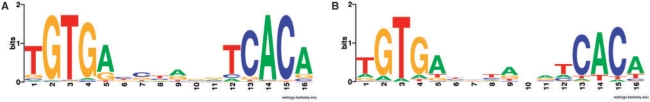
In contrast to the TFBS searched within our pipeline, we created an adopted and optimized PWM for CrpMT from experimentally verified and predicted binding sites, and applied it for TFBS search. To detect the novel binding sites, we set the P-value threshold to 10−5 and performed a restrictive search on sequences upstream genes/operons concering the whole genome of MT. Again, we used PoSSuMsearch for binding site prediction scanning 580-bp long upstream sequence, ranging from +20 bp relative to the transcription start site. Using Weblogo (61), we generated a sequence logo from the detected binding sites of CrpMT and from the appropriate binding sites of GlxR that were used for PWM creation (see Methods section). The resulting sequence logos are shown in Figure 3.
Figure 3.
Sequence logo of the predicted CrpMT binding sites (A) in comparison to the sequence logo of GlxR (B). The sequence logo models the binding site motif of CrpMT. It was deduced from the predicted binding sites in Table 3. The height of each letter within an individual stack represents the nucleotide's frequency relative to the particular motif position; thus, the degree of a nucleotide's conservation is indicated by the stack according to the respective position.
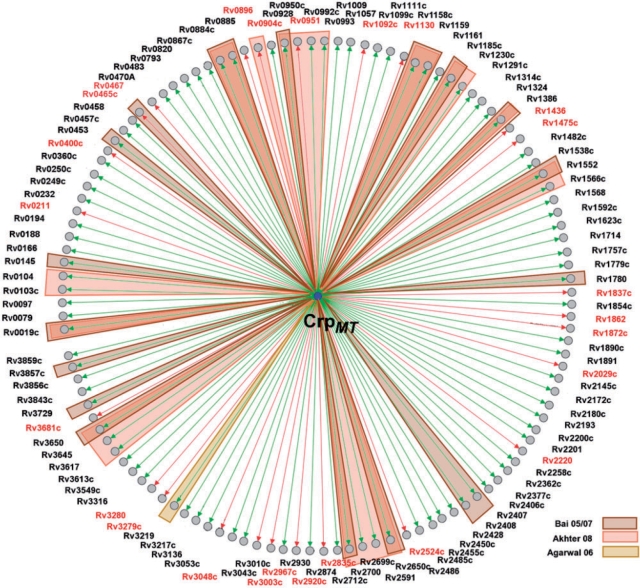
In total, we identified 207 putative target genes of CrpMT, organized in 121 transcription units (see Table 3 and Figure 4). Of this set, 17 genes belong to the mycobacterial core regulon (62) and 41 were reported as essential for M. tuberculosis (63,64). Furthermore, at least 17 genes of the suggested regulon are connected to antibiotic resistance and virulence of M. tuberculosis (65–69). Based on annotation information for M. tuberculosis (69), knowledge about orthologous C. glutamicum genes and operon structures, we attributed individual target genes to distinct functional modules.
Table 3.
Predicted CrpMT binding sites
| Gene ID | Gene | Motif positiond | Motif sequence | Operon |
|---|---|---|---|---|
| Carbohydarate metabolism | ||||
| Rv0211a | pckA | −166 | TGTGAGCAGGCTTATA | – |
| Rv0249c | sdhCD | −104 | TGTGATCTGTAACACC | Rv0249c-Rv0248c-Rv0247ca |
| Rv0249c | sdhCD | −410 | GGTGTCGGAGGTCACA | Rv0249c-Rv0248c-Rv0247ca |
| Rv0458 | adhA | −41 | TGTGAGCTGTATTACA | Rv0458-Rv0459 |
| Rv0465c | – | −167 | TTTGTCGAGGCTCACG | Rv0465ca–Rv0464cg |
| Rv0467a,g | icl | −341 | TGTTACAACGCTCACA | – |
| Rv0896a | gltA2 | −356 | GGCGTTGAACATCACC | – |
| Rv0951 | sucC | −173 | TGTGAGTTGGATCACG | Rv0951a,f–Rv0952a,f |
| Rv1099c | – | −515 | GCTGATGAATCCCACG | Rv1099c–Rv1098ca,f–Rv1097c |
| Rv1130 | prpD2 | −152 | TGTGGATAAGTCCAGG | Rv1130a–Rv1131 |
| Rv1436 | gap | −48 | GGTTGTTTAGCCAACA | Rv1436a,f–Rv1437a,f–Rv1438a,f |
| Rv1475c | acn | −462 | TGTAACTGCCGACATA | Rv1475ca,f–Rv1474c |
| Rv1552 | frdA | −284 | TGTGATCTAGGTCACGb | Rv1552–Rv1553–Rv1554–Rv1555 |
| Rv1837ca | glcB | −381 | AGGGATGCACTACACA | – |
| Rv1862a | adhA | −227 | CGTGGGGCGCCACACA | – |
| Rv1872ca | lldD2 | −200 | GATGCCGTAGCGCACT | – |
| Rv2029ca | pfkB | −410 | GGTGACGAGTCGCGCA | Rv2029c–Rv2028c–Rv2027c–Rv2026cf |
| Rv2967ca,f | pca | −389 | CGTGGTGGTGGTCACC | – |
| Rv3010ca | pfkA | −532 | GGTGATGGCGATGACC | – |
| Rv3316 | sdhC | −386 | CGAGACGTTTTCCACG | Rv3316–Rv3317–Rv3318a–Rv3319 |
| Rv3676a | CRP | −538 | TGTCACCTACGACAGA | – |
| Fatty acid metabolism | ||||
| Rv0097 | – | −526 | TGTCACGCCGGCCACG | Rv0097–Rv0098e–Rv0099c–Rv0100e–Rv0101 |
| Rv0166 | fadD5 | −84 | TGTGACCCAGACAACA | – |
| Rv0400ca,f | fadE7 | −5 | AGTGATGAGCACCCCG | – |
| Rv1185c | fadD21 | −168 | CGTGACGCCCCTCACG | – |
| Rv1714 | – | −405 | GGTGACGGCGGCCACA | Rv1714f–Rv1715f–Rv1716–Rv1717–Rv1718 |
| Rv2485cc | lipQ | −91 | TGTGATCCTCGACACA | – |
| Rv2486 | echA14 | −287 | TGTGTCGAGGATCACA | – |
| Rv2524ca,f | fas | −259 | CGTTACCCACGACACG | – |
| Rv2930c | fadD26 | −498 | TGTTAATCTCGTCACA | Rv2930–Rv2931–Rv2932f–Rv2933–Rv2934–Rv2935–Rv2936g–Rv2937g–Rv2938–Rv2939 |
| Rv3279c | birA | −38 | TATCGGCTGCCGCACA | Rv3279ca–Rv3278ce |
| Rv3280 | accD5 | −331 | CGGGACGTCGACCACA | Rv3280a–Rv3281e,f–Rv3282 |
| Rv3549c | – | −67 | GGTGATCGGCATTGCA | Rv3549c–Rv3548ca |
| Nitrogen assimilation | ||||
| Rv1538c | ansA | −187 | TGTGAGCACCACCACA | – |
| Rv2220a,f,g | glnA1 | −1 | TGTGACGGAAAAGACG | – |
| Rv2920c | amt | −2 | AGTGGACCAATTCCCC | Rv2920ca–Rv2919ca,g–Rv2918ca |
| Rv3859c | gltB | −398 | TGCTCCGGATTTCACA | Rv3859ca,f–Rv3858cf |
| PGRS | ||||
| Rv0453 | PPE11 | −269 | GGTGACCAAACTCACG | – |
| Rv1386 | PE15 | −133 | TGTGACCAAACTCACCb | Rv1386e–Rv1387c,e |
| Rv2408 | PE24 | −213 | GGTGATCGGCGTCACG | – |
| Rv2591 | P_PGRS44 | −38 | CGTGACATGTGTCACA | – |
| Rv3136c | PPE51 | −16 | AAGGAGCTGAGACACA | – |
| Rv3650 | PE33 | −83 | TGTGATGCACTTGACA | – |
| Respiration | ||||
| Rv1161 | narG | −512 | TGCGTTGAACGGCACG | Rv1161a–Rv1162a–Rv1163a–Rv1164–Rv1165–Rv1166f |
| Rv1623cc | cydA | −181 | CGTGGTGATCGGCACA | – |
| Rv1854ca | ndh | −109 | TGTGGCTGATGACACA | – |
| Rv2193 | ctaE | −517 | GGTGGATAGGTTCACC | Rv2193a,f–Rv2194a,f–Rv2195a,f–Rv2196a,f |
| Rv2200c | ctaC | −23 | TGTGATACAGGAGGCG | Rv2200ca,f–Rv2199c |
| Rv3043c | ctaD | −227 | AGTGGATCGCATCCCG | Rv3043ca,f–Rv3042cf |
| Other cellular processes | ||||
| Rv0019cg | fhaB | −69 | CGTGACTTTGCTGACGb | – |
| Rv0079 | – | −110 | GGTGACACAGCCCACA | Rv0079–Rv0080 |
| Rv0103c | ctpB | −159 | TGTGACGGGCGTCACA | – |
| Rv0104 | – | −1 | TGTGACGCCCGTCACA | – |
| Rv0145 | – | −59 | AGTGATGTGCCACACAb | Rv0145–Rv0146 |
| Rv0188c | – | −356 | AGAGAACAACGTCGCA | – |
| Rv0194 | – | −517 | TGTCATCTAGATCACG | – |
| Rv0232 | – | −53 | CGTGATGCAGCGCACA | Rv0232–Rv0233 |
| Rv0250ce | – | −37 | TGTGATCTGTAACACC | – |
| Rv0360c | – | −2 | CGTGACCAAGCGCACA | – |
| Rv0457c | – | −43 | TGTAATACAGCTCACA | – |
| Rv0470A | – | −212 | TGTGGTGGGAATCACA | – |
| Rv0483 | lprQ | −116 | TGTGTTTGGTATCACA | – |
| Rv0793 | – | −375 | TGTGATGGTGCGCACG | – |
| Rv0820a,g | phoT | −538 | GGTGGTGATCCGCACC | – |
| Rv0867ca | rpfA | −443 | TGTGACATTACCCACAb | – |
| Rv0884ca,f | serC | −91 | TGTGAGCTTGTTCACAb | – |
| Rv0885 | – | −133 | TGTGAACAAGCTCACA | Rv0885–Rv0886 |
| Rv0904ca | accD3 | −2 | CGTGAGTCGTATCACG | – |
| Rv0928 | pstS3 | −6 | ACTGAATTGAAACTCA | Rv0928a,g–Rv0929a,g–Rv0930a,g |
| Other cellular processes | ||||
| Rv0993 | galU | −8 | TGTGAACGATGTCACG | Rv0993f–Rv0994g–Rv0995g |
| Rv0950c | – | −153 | CGTGATCCAACTCACAb | – |
| Rv0992c | – | −109 | CGTGACATCGTTCACA | Rv0992–Rv0991–Rv0990 |
| Rv1009 | rpfB | −271 | GGTGGCGCTCATCACC | Rv1009a–Rv1010a,g–Rv1011a,f |
| Rv1057 | – | −248 | CGTGACCTAGGTAACA | – |
| Rv1092ca,f | coaA | −242 | TGCCACGTAGGTCACG | – |
| Rv1111ce | – | −411 | GGTGACATGAGTCACG | – |
| Rv1158c | – | −69 | TGTCACTTGAGTCACAb | Rv1158ce–Rv1157ce |
| Rv1159 | pimE | −77 | TGTGACTCAAGTGACA | – |
| Rv1230c | – | −79 | GGTGATCTAGTTCACGb | – |
| Rv1291c | – | −323 | TGTGATCGGCGCCACC | – |
| Rv1314c | – | −294 | GGTGATCCGGGCCACG | – |
| Rv1324e | – | −104 | TGTGATCTTGGTCATA | – |
| Rv1482c | – | −23 | TGTGACTCAGCACACT | – |
| Rv1566c | – | −235 | CGTGACTGAAATCACA | – |
| Rv1568 | bioA | −553 | TGTGATTTCAGTCACG | Rv1568–Rv1569g–Rv1570–Rv1571 |
| Rv1592cc | – | −215 | TGTGATAGGCGCCACG | – |
| Rv1757c | – | −351 | TGTGACGGCGGCCACG | – |
| Rv1779c | – | −89 | TGTGAACAACACCACA | – |
| Rv1780 | – | −147 | TGTGGTGTTGTTCACA | – |
| Rv1890c | – | −7 | TGTGTCGTGGCCCACA | – |
| Rv1891e,g | – | −63 | TGTGGGCCACGACACA | Rv1891–Rv1892–Rv1893e |
| Rv2145ca,f | wag31 | −463 | CGTGACTGGCGTCCCA | – |
| Rv2172ce | – | −2 | TGTGACCCTCAACACG | – |
| Rv2180c | – | −304 | TGTGTGGAACAACACA | – |
| Rv2201a,f | asnB | −336 | GCTGTCGAAGACCACG | – |
| Rv2258c | – | −459 | GGTGACGTCGACCACG | – |
| Rv2362c | recO | −224 | TGTGGGCTGGCTCACA | Rv2362c–Rv2361cf–Rv2360c |
| Rv2377c | mbtH | −268 | TGTGGTTCACCTCACT | – |
| Rv2406c | – | −34 | TGTGAACCAGCTCACC | – |
| Rv2407 | – | −242 | GGTGAGCTGGTTCACA | – |
| Rv2428c | ahpC | −93 | GGTGTGATATATCACC | – |
| Rv2450c | rpfE | −509 | TGTGGCGCAGGTCACC | – |
| Rv2450c | rpfE | −422 | CGTGATTCGGCTCACG | – |
| Rv2455c | – | −237 | AGTGACCAATACCACA | Rv2455c–Rv2454c–Rv2453c |
| Rv2650c | – | −305 | CGTGAGGAGCCTCACG | – |
| Rv2699c | – | −116 | TGTGATGTAAATCACA | – |
| Rv2700e,f | – | −138 | TGTGATTTACATCACA | – |
| Rv2712c | – | −296 | GGTGAGGTAGAGCACA | – |
| Rv2835c | ugpA | −513 | GGTGATGCCGGGCACG | Rv2835c–Rv2834ca–Rv2833ca,f–Rv2832ca,f |
| Other cellular processes | ||||
| Rv2874 | dipZ | −351 | TGTGGCGGAGTTCACA | – |
| Rv3003c | ilvB1 | −335 | TGTGGTGGCCACCCCA | Rv3003ca,f–Rv3002ca,f–Rv3001ca,f |
| Rv3048ca,f | nrdF2 | −2 | GGTGACTGGAAACGCA | – |
| Rv3053c | nrdH | −347 | GGTGATCTGCGACACG | Rv3053c–Rv3052c–Rv3051cf |
| Rv3217ca,e | – | −278 | TGTGGTGGCGGTCGCA | – |
| Rv3219 a,c | whiB1 | −176 | AGTGAGATAGCCCACGb | – |
| Rv3613cc | – | −458 | CGTGACGAATCCCCCA | – |
| Rv3617 | ephA | −315 | TGTGACCGGTGTCACT | Rv3617–Rv3618 |
| Rv3645 | – | −179 | TGTGAGCCGAATCACG | – |
| Rv3681ca | whiB4 | −106 | TGAGATACAGGTAACA | – |
| Rv3729 | – | −190 | TGTGACCACGGCCACG | – |
| Rv3843c | – | −505 | GGTGAGGTAAGTCACA | Rv3843ce–Rv3842cg |
| Rv3856c | – | −547 | TGTGGGCTTCGTCACA | – |
| Rv3857c | – | −341 | TGTGGGCTTCGTCACAb | – |
| Consensus | TGTGANNNNNNTCACA | |||
CrpMT binding sites detected by the TFBS search of the introduced pipeline and by the additional TFBS search with adopted and optimized PWMs. Bold letters indicate conserved pentamers of the motif. Codes:
aTransferred target gene from CG.
bExperimentally verified binding site by EMSA/CHiP/RT-PCR (21–23).
cGene showed altered expression in microarray studies of ΔRv3676 versus WT (24).
dMotif position relative to the translation start site.
eCore gene.
fEssential gene.
gGene involved in virulence processes
Figure 4.
Reconstructed network of the GlxR ortholog CrpMT. The network reconstruction of the CrpMT regulon is based on the 121 transcription units presented in Table 3. It was generated by the integrated network reconstruction tool GraphVis of MycoRegNet. Transcription units relying on binding site predictions/experimental verifications that were reported previously in (22–24,60) and correspond with our findings are colored according to the appropriate publication. Arrows and gene IDs (node labels) coloured in red indicate a repressive regulation of CrpMT, green arrows correspond to an activating regulation.
Similar to present knowledge on GlxR, results implicate CrpMT in the regulation of several functional modules such as carbohydrate metabolism (40 target genes), fatty acid metabolism (33 target genes), respiration (16 target genes) and nitrogen assimilation (7 target genes). Therefore, the position of the GlxR homolog CrpMT as global regulator in the transcriptional regulatory network seems to be conserved in M. tuberculosis. It is interesting to note that the suggested regulon comprises genes involved in essential functional modules, e.g. the citrate cycle, as well as genes involved in the synthesis of the cellular envelope which plays an important role in the virulence of M. tuberculosis. Together with the supposed regulation of further virulence−associated genes this might explain why a functional CrpMT is required for virulence in model systems (24).
CONCLUSIONS
With MycoRegNet, we have set up a system that allows researchers of the tuberculosis community to perform comprehensive analysis and visualizations of the gene regulatory network of MT. With its TFBS prediction it further provides easy access to a method that helps to generate new hypotheses in silico. As the sister project to CoryneRegNet, the MycoRegNet database content was generated through our comparative genomics pipeline, which provided us with reliable transfers of gene regulatory interactions from the reference organism C. glutamicum to M. tuberculosis. With MycoRegNet, the corresponding data are publicly available and can be accessed easily through the web interface, or in a well-structured manner by using the MycoRegNet Web Service to maintain the reconstruction, visualization, and validation of mycobacterial regulatory networks at different hierarchical levels. Taken together, MycoRegNet is a reference resource for the tuberculosis community to gain a better understanding of the complex coherences of transcriptional gene control. It has the potential to assist researchers at the development of new vaccines and drugs to treat and prevent tuberculosis. Although MycoRegNet has been initially designed for MT, it may also serve for other mycobacterial strains in future, such as the already integrated M. tuberculosis CDC1551.
FUNDING
GenoMik-Plus initiative of the German Federal Ministry of Education and Research (grant 0313805A); ERA-NET PathoGenoMics SPATELIS project (to J.K.); German Academic Exchange Service (to J.B. for his work at ICSI). Funding for open access charge: The work was financed in part by the GenoMik-Plus initiative of the German Federal Ministry of Education and Research, grant 0313805A.
Conflict of interest statement. None declared.
REFERENCES
- 1.Frieden TR, Sterling TR, Munsiff SS, Watt CJ, Dye C. Tuberculosis. Lancet. 2003;362:887–899. doi: 10.1016/S0140-6736(03)14333-4. [DOI] [PubMed] [Google Scholar]
- 2.Raviglione MC, Smith IM. XDR tuberculosis–implications for global public health. N. Engl. J. Med. 2007;356:656–659. doi: 10.1056/NEJMp068273. [DOI] [PubMed] [Google Scholar]
- 3.Wilson M, DeRisi J, Kristensen HH, Imboden P, Rane S, Brown PO, Schoolnik GK. Exploring drug-induced alterations in gene expression in Mycobacterium tuberculosis by microarray hybridization. Proc. Natl Acad. Sci. USA. 1999;96:12833–12838. doi: 10.1073/pnas.96.22.12833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Singh JA, Upshur R, Padayatchi N. XDR-TB in South Africa: no time for denial or complacency. PLoS Med. 2007;4:e50. doi: 10.1371/journal.pmed.0040050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Park HD, Guinn KM, Harrell MI, Liao R, Voskuil MI, Tompa M, Schoolnik GK, Sherman DR. Rv3133c/dosR is a transcription factor that mediates the hypoxic response of Mycobacterium tuberculosis. Mol. Microbiol. 2003;48:833–843. doi: 10.1046/j.1365-2958.2003.03474.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Stewart GR, Snewin VA, Walzl G, Hussell T, Tormay P, O'Gaora P, Goyal M, Betts J, Brown IN, Young DB. Overexpression of heat-shock proteins reduces survival of Mycobacterium tuberculosis in the chronic phase of infection. Nat. Med. 2001;7:732–737. doi: 10.1038/89113. [DOI] [PubMed] [Google Scholar]
- 7.Gama-Castro S, Jimnez-Jacinto V, Peralta-Gil M, Santos-Zavaleta A, Pealoza-Spinola MI, Contreras-Moreira B, Segura-Salazar J, Muiz-Rascado L, Martnez-Flores I, Salgado H, et al. RegulonDB (version 6.0): gene regulation model of Escherichia coli K-12 beyond transcription, active (experimental) annotated promoters and Textpresso navigation. Nucleic Acids Res. 2008;36:D120–D124. doi: 10.1093/nar/gkm994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Baumbach J. CoryneRegNet 4.0 – a reference database for Corynebacterial gene regulatory networks. BMC Bioinformatics. 2007;8:429. doi: 10.1186/1471-2105-8-429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jacques PE, Gervais AL, Cantin M, Lucier JF, Dallaire G, Drouin G, Gaudreau L, Goulet J, Brzezinski R. MtbRegList, a database dedicated to the analysis of transcriptional regulation in Mycobacterium tuberculosis. Bioinformatics. 2005;21:2563–2565. doi: 10.1093/bioinformatics/bti321. [DOI] [PubMed] [Google Scholar]
- 10.Reddy TBK, Riley R, Wymore F, Montgomery P, DeCaprio D, Engels R, Gellesch M, Hubble J, Jen D, Jin H, et al. Tb database: an integrated platform for tuberculosis research. Nucleic Acids Res. 2009;37:D499–D508. doi: 10.1093/nar/gkn652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Babu MM, Teichmann SA. Evolution of transcription factors and the gene regulatory network in Escherichia coli. Nucleic Acids Res. 2003;31:1234–1244. doi: 10.1093/nar/gkg210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Babu MM, Luscombe NM, Aravind L, Gerstein M, Teichmann SA. Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol. 2004;14:283–291. doi: 10.1016/j.sbi.2004.05.004. [DOI] [PubMed] [Google Scholar]
- 13.Babu MM, Teichmann SA, Aravind L. Evolutionary dynamics of prokaryotic transcriptional regulatory networks. J. Mol. Biol. 2006;358:614–633. doi: 10.1016/j.jmb.2006.02.019. [DOI] [PubMed] [Google Scholar]
- 14.Stackebrandt E, Rainey FA, Ward-Rainey NL. Proposal for a new hierarchic classification system, Actionbacteria classis nov. Int. J. Sys. Bacteriol. 1997;47:479–491. [Google Scholar]
- 15.Eggeling L, Besra GS, Alderwick L. Structure and synthesis of the cell wall. In: Burkovski A, editor. Corynebacteria Genomics and Molecular Biology. Caister Academic Press; 2008. pp. 267–294. [Google Scholar]
- 16.Seidel M, Alderwick LJ, Sahm H, Besra GS, Eggeling L. Topology and mutational analysis of the single Emb arabinofuranosyltransferase of Corynebacterium glutamicum as a model of Emb proteins of Mycobacterium tuberculosis. Glycobiology. 2007;17:210–219. doi: 10.1093/glycob/cwl066. [DOI] [PubMed] [Google Scholar]
- 17.Portevin D, Sousa-D'Auria CD, Houssin C, Grimaldi C, Chami M, Daff M, Guilhot C. A polyketide synthase catalyzes the last condensation step of mycolic acid biosynthesis in mycobacteria and related organisms. Proc. Natl Acad. Sci. USA. 2004;101:314–319. doi: 10.1073/pnas.0305439101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Brinkrolf K, Brune I, Tauch A. The transcriptional regulatory network of the amino acid producer Corynebacterium glutamicum. J. Biotechnol. 2007;129:191–211. doi: 10.1016/j.jbiotec.2006.12.013. [DOI] [PubMed] [Google Scholar]
- 19.Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Beckstette M, Homann R, Giegerich R, Kurtz S. Fast index based algorithms and software for matching position specific scoring matrices. BMC Bioinformatics. 2006;7:389. doi: 10.1186/1471-2105-7-389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Agarwal N, Raghunand TR, Bishai WR. Regulation of the expression of whiB1 in Mycobacterium tuberculosis: role of cAMP receptor protein. Microbiology. 2006;152(Pt 9):2749–2756. doi: 10.1099/mic.0.28924-0. [DOI] [PubMed] [Google Scholar]
- 22.Bai G, Gazdik MA, Schaak DD, McDonough KA. The Mycobacterium bovis BCG cyclic AMP receptor-like protein is a functional DNA binding protein in vitro and in vivo, but its activity differs from that of its M. tuberculosis ortholog, Rv3676. Infect. Immun. 2007;75:5509–5517. doi: 10.1128/IAI.00658-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bai G, McCue LA, McDonough KA. Characterization of Mycobacterium tuberculosis Rv3676 (CRPMt), a cyclic amp receptor protein-like DNA binding protein. J. Bacteriol. 2005;187:7795–7804. doi: 10.1128/JB.187.22.7795-7804.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rickman L, Scott C, Hunt DM, Hutchinson T, Menndez MC, Whalan R, Hinds J, Colston MJ, Green J, Buxton RS. A member of the cAMP receptor protein family of transcription regulators in Mycobacterium tuberculosis is required for virulence in mice and controls transcription of the rpfA gene coding for a resuscitation promoting factor. Mol. Microbiol. 2005;56:1274–1286. doi: 10.1111/j.1365-2958.2005.04609.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sun R, Converse PJ, Ko C, Tyagi S, Morrison NE, Bishai WR. Mycobacterium tuberculosis ECF sigma factor sigC is required for lethality in mice and for the conditional expression of a defined gene set. Mol. Microbiol. 2004;52:25–38. doi: 10.1111/j.1365-2958.2003.03958.x. [DOI] [PubMed] [Google Scholar]
- 26.Manganelli R, Voskuil MI, Schoolnik GK, Smith I. The Mycobacterium tuberculosis ECF sigma factor sigmaE: role in global gene expression and survival in macrophages. Mol. Microbiol. 2001;41:423–437. doi: 10.1046/j.1365-2958.2001.02525.x. [DOI] [PubMed] [Google Scholar]
- 27.Manganelli R, Voskuil MI, Schoolnik GK, Dubnau E, Gomez M, Smith I. Role of the extracytoplasmic-function sigma factor sigma(H) in Mycobacterium tuberculosis global gene expression. Mol. Microbiol. 2002;45:365–374. doi: 10.1046/j.1365-2958.2002.03005.x. [DOI] [PubMed] [Google Scholar]
- 28.Davis EO, Dullaghan EM, Rand L. Definition of the mycobacterial SOS box and use to identify LexA-regulated genes in Mycobacterium tuberculosis. J. Bacteriol. 2002;184:3287–3295. doi: 10.1128/JB.184.12.3287-3295.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stewart GR, Wernisch L, Stabler R, Mangan JA, Hinds J, Laing KG, Young DB, Butcher PD. Dissection of the heat-shock response in Mycobacterium tuberculosis using mutants and microarrays. Microbiology. 2002;148(Pt 10):3129–3138. doi: 10.1099/00221287-148-10-3129. [DOI] [PubMed] [Google Scholar]
- 30.Parish T, Smith DA, Roberts G, Betts J, Stoker NG. The SenX3-RegX3 two-component regulatory system of Mycobacterium tuberculosis is required for virulence. Microbiology. 2003;149(Pt 6):1423–1435. doi: 10.1099/mic.0.26245-0. [DOI] [PubMed] [Google Scholar]
- 31.Betts JC, Lukey PT, Robb LC, McAdam RA, Duncan K. Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol. Microbiol. 2002;43:717–731. doi: 10.1046/j.1365-2958.2002.02779.x. [DOI] [PubMed] [Google Scholar]
- 32.Kendall SL, Movahedzadeh F, Rison SCG, Wernisch L, Parish T, Duncan K, Betts JC, Stoker NG. The Mycobacterium tuberculosis dosRS two-component system is induced by multiple stresses. Tuberculosis (Edinb) 2004;84:247–255. doi: 10.1016/j.tube.2003.12.007. [DOI] [PubMed] [Google Scholar]
- 33.Kendall SL, Withers M, Soffair CN, Moreland NJ, Gurcha S, Sidders B, Frita R, Bokum AT, Besra GS, Lott JS, et al. A highly conserved transcriptional repressor controls a large regulon involved in lipid degradation in Mycobacterium smegmatis and Mycobacterium tuberculosis. Mol. Microbiol. 2007;65:684–699. doi: 10.1111/j.1365-2958.2007.05827.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Florczyk MA, McCue LA, Purkayastha A, Currenti E, Wolin MJ, McDonough KA. A family of acr-coregulated Mycobacterium tuberculosis genes shares a common DNA motif and requires Rv3133c (dosR or devR) for expression. Infect. Immun. 2003;71:5332–5343. doi: 10.1128/IAI.71.9.5332-5343.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2009;37:D26–D31. doi: 10.1093/nar/gkn723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Alm EJ, Huang KH, Price MN, Koche RP, Keller K, Dubchak IL, Arkin AP. The MicrobesOnline web site for comparative genomics. Genome Res. 2005;15:1015–1022. doi: 10.1101/gr.3844805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Curbera F, Duftler M, Khalaf R, Nagy W, Mukhi N, Weerawarana S. Unraveling the web services web: an introduction to SOAP, WSDL, and UDDI. IEEE Internet Comput. 2002;6:86–93. [Google Scholar]
- 38.Meyer F, Goesmann A, McHardy AC, Bartels D, Bekel T, Clausen J, Kalinowski J, Linke B, Rupp O, Giegerich R, et al. GenDB–an open source genome annotation system for prokaryote genomes. Nucleic Acids Res. 2003;31:2187–2195. doi: 10.1093/nar/gkg312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Dondrup M, Albaum S, Griebel T, Henckel K, Jünemann S, Kahlke T, Kleindt C, Küster H, Linke B, Mertens D, et al. EMMA 2 – a MAGE-compliant system for the collaborative analysis and integration of microarray data. BMC Bioinformatics. 2009;10:50. doi: 10.1186/1471-2105-10-50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kanehisa M, Goto S, Hattori M, Aoki-Kinoshita KF, Itoh M, Kawashima S, Katayama T, Araki M, Hirakawa M. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res. 2006;34:D354–D357. doi: 10.1093/nar/gkj102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV, Krylov DM, Mazumder R, Mekhedov SL, Nikolskaya AN, et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics. 2003;4:41. doi: 10.1186/1471-2105-4-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Baumbach J, Brinkrolf K, Wittkop T, Tauch A, Rahmann S. CoryneRegNet 2: an integrative bioinformatics approach for reconstruction and comparison of transcriptional regulatory networks in prokaryotes. J. Integr. Bioinformat. 2006;3:24. [Google Scholar]
- 44.Pillai S, Silventoinen V, Kallio K, Senger M, Sobhany S, Tate J, Velankar S, Golovin A, Henrick K, Rice P. Soap-based services provided by the European Bioinformatics Institute. Nucleic Acids Res. 2005;33:W25–W28. doi: 10.1093/nar/gki491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Barthelmes J, Ebeling C, Chang A, Schomburg I, Schomburg D. BRENDA, AMENDA and FRENDA: the enzyme information system in 2007. Nucleic Acids Res. 2007;35:D511–D514. doi: 10.1093/nar/gkl972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Körner H, Sofia HJ, Zumft WG. Phylogeny of the bacterial superfamily of Crp-Fnr transcription regulators: exploiting the metabolic spectrum by controlling alternative gene programs. FEMS Microbiol. Rev. 2003;27:559–592. doi: 10.1016/S0168-6445(03)00066-4. [DOI] [PubMed] [Google Scholar]
- 47.Akhter Y, Tundup S, Hasnain SE. Novel biochemical properties of a CRP/FNR family transcription factor from Mycobacterium tuberculosis. Int. J. Med. Microbiol. 2007;297:451–457. doi: 10.1016/j.ijmm.2007.04.009. [DOI] [PubMed] [Google Scholar]
- 48.Kim HJ, Kim TH, Kim Y, Lee H-S. Identification and characterization of GlxR, a gene involved in regulation of glyoxylate bypass in Corynebacterium glutamicum. J. Bacteriol. 2004;186:3453–3460. doi: 10.1128/JB.186.11.3453-3460.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Deutscher J. The mechanisms of carbon catabolite repression in bacteria. Curr. Opin. Microbiol. 2008;11:87–93. doi: 10.1016/j.mib.2008.02.007. [DOI] [PubMed] [Google Scholar]
- 50.Görke B, Stülke J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 2008;6:613–624. doi: 10.1038/nrmicro1932. [DOI] [PubMed] [Google Scholar]
- 51.Shenoy AR, Capuder M, Draskovic P, Lamba D, Visweswariah SS, Podobnik M. Structural and biochemical analysis of the Rv0805 cyclic nucleotide phosphodiesterase from Mycobacterium tuberculosis. J. Mol. Biol. 2007;365:211–225. doi: 10.1016/j.jmb.2006.10.005. [DOI] [PubMed] [Google Scholar]
- 52.Shenoy AR, Visweswariah SS. New messages from old messengers: cAMP and mycobacteria. Trends Microbiol. 2006;14:543–550. doi: 10.1016/j.tim.2006.10.005. [DOI] [PubMed] [Google Scholar]
- 53.Shenoy AR, Visweswariah SS. Mycobacterial adenylyl cyclases: biochemical diversity and structural plasticity. FEBS Lett. 2006;580:3344–3352. doi: 10.1016/j.febslet.2006.05.034. [DOI] [PubMed] [Google Scholar]
- 54.McCue LA, McDonough KA, Lawrence CE. Functional classification of cNMP-binding proteins and nucleotide cyclases with implications for novel regulatory pathways in Mycobacterium tuberculosis. Genome Res. 2000;10:204–219. doi: 10.1101/gr.10.2.204. [DOI] [PubMed] [Google Scholar]
- 55.Kohl TA, Baumbach J, Jungwirth B, Phler A, Tauch A. The GlxR regulon of the amino acid producer Corynebacterium glutamicum: in silico and in vitro detection of DNA binding sites of a global transcription regulator. J. Biotechnol. 2008;135:340–350. doi: 10.1016/j.jbiotec.2008.05.011. [DOI] [PubMed] [Google Scholar]
- 56.Jungwirth B, Emer D, Brune I, Hansmeier N, Pühler A, Eikmanns BJ, Tauch A. Triple transcriptional control of the resuscitation promoting factor 2 (rpf2) gene of Corynebacterium glutamicum by the regulators of acetate metabolism RamA and RamB and the cAMP-dependent regulator GlxR. FEMS Microbiol. Lett. 2008;281:190–197. doi: 10.1111/j.1574-6968.2008.01098.x. [DOI] [PubMed] [Google Scholar]
- 57.Han SO, Inui M, Yukawa H. Effect of carbon source availability and growth phase on expression of Corynebacterium glutamicum genes involved in the tricarboxylic acid cycle and glyoxylate bypass. Microbiology. 2008;154(Pt 10):3073–3083. doi: 10.1099/mic.0.2008/019828-0. [DOI] [PubMed] [Google Scholar]
- 58.Han SO, Inui M, Yukawa H. Expression of Corynebacterium glutamicum glycolytic genes varies with carbon source and growth phase. Microbiology. 2007;153(Pt 7):2190–2202. doi: 10.1099/mic.0.2006/004366-0. [DOI] [PubMed] [Google Scholar]
- 59.Letek M, Valbuena N, Ramos A, Ordez E, Gil JA, Mateos LM. Characterization and use of catabolite-repressed promoters from gluconate genes in Corynebacterium glutamicum. J. Bacteriol. 2006;188:409–423. doi: 10.1128/JB.188.2.409-423.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Akhter Y, Yellaboina S, Farhana A, Ranjan A, Ahmed N, Hasnain SE. Genome scale portrait of cAMP-receptor protein (CRP) regulons in mycobacteria points to their role in pathogenesis. Gene. 2008;407:148–158. doi: 10.1016/j.gene.2007.10.017. [DOI] [PubMed] [Google Scholar]
- 61.Crooks GE, Hon G, Chandonia J-M, Brenner SE. WebLogo: a sequence logo generator. Genome Res. 2004;14:1188–1190. doi: 10.1101/gr.849004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Marmiesse M, Brodin P, Buchrieser C, Gutierrez C, Simoes N, Vincent V, Glaser P, Cole ST, Brosch R. Macro-array and bioinformatic analyses reveal mycobacterial ‘core’ genes, variation in the ESAT-6 gene family and new phylogenetic markers for the Mycobacterium tuberculosis complex. Microbiology. 2004;150(Pt 2):483–496. doi: 10.1099/mic.0.26662-0. [DOI] [PubMed] [Google Scholar]
- 63.Sassetti CM, Boyd DH, Rubin EJ. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 2003;48:77–84. doi: 10.1046/j.1365-2958.2003.03425.x. [DOI] [PubMed] [Google Scholar]
- 64.Lamichhane G, Zignol M, Blades NJ, Geiman DE, Dougherty A, Grosset J, Broman KW, Bishai WR. A postgenomic method for predicting essential genes at subsaturation levels of mutagenesis: application to Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA. 2003;100:7213–7218. doi: 10.1073/pnas.1231432100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Rengarajan J, Bloom BR, Rubin EJ. Genome-wide requirements for Mycobacterium tuberculosis adaptation and survival in macrophages. Proc. Natl Acad. Sci. USA. 2005;102:8327–8332. doi: 10.1073/pnas.0503272102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Tullius MV, Harth G, Horwitz MA. Glutamine synthetase GlnA1 is essential for growth of Mycobacterium tuberculosis in human THP-1 macrophages and guinea pigs. Infect. Immun. 2003;71:3927–3936. doi: 10.1128/IAI.71.7.3927-3936.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.McAdam RA, Quan S, Smith DA, Bardarov S, Betts JC, Cook FC, Hooker EU, Lewis AP, Woollard P, Everett MJ, et al. Characterization of a Mycobacterium tuberculosis H37Rv transposon library reveals insertions in 351 ORFs and mutants with altered virulence. Microbiology. 2002;148(Pt 10):2975–2986. doi: 10.1099/00221287-148-10-2975. [DOI] [PubMed] [Google Scholar]
- 68.McKinney JD, Höner zu Bentrup K, Muñoz-Elías EJ, Miczak A, Chen B, Chan WT, Swenson D, Sacchettini JC, Jacobs WR, Russell DG. Persistence of Mycobacterium tuberculosis in macrophages and mice requires the glyoxylate shunt enzyme isocitrate lyase. Nature. 2000;406:735–738. doi: 10.1038/35021074. [DOI] [PubMed] [Google Scholar]
- 69.Cole ST. Learning from the genome sequence of Mycobacterium tuberculosis H37Rv. FEBS Lett. 1999;452:7–10. doi: 10.1016/s0014-5793(99)00536-0. [DOI] [PubMed] [Google Scholar]