Abstract
Background
Symbiotic N2 fixation in legumes is constrained by many factors, including the paucity of suitable soil rhizobia To maximise growth of legume species therefore often requires the application of effective rhizobia as inoculants. But where native strains out-compete introduced rhizobia for nodule formation, it is important that the competitiveness of selected strains is tested in the field and glasshouse prior to their recommendation as commercial inoculants. However the methodology for strain identification inside nodules has often proved difficult and thus limited this field of research. In this study, the suitability of the antibiotic resistance technique (both intrinsic low-resistance fingerprinting and high-resistance marking) and the serological indirect ELISA method were assessed for their ability to detect selected Cyclopia rhizobia under glasshouse and field conditions. The four rhizobial strains that were used, namely PPRICI3, UCT40a, UCT44b and UCT61a, were isolated from wild Cyclopia species growing in the Western Cape fynbos of South Africa.
Results
The test strains formed two distinct groups with regard to their intrinsic resistance to the antibiotics streptomycin sulphate and spectinomycin dihydrochloride pentahydrate, making it impossible to use intrinsic antibiotic resistance to distinguish strains from within the same intrinsic resistance group. The use of strains marked with double antibiotic resistance was also investigated. A number of these strains lost their antibiotic marker tags after one plant passage; and some also lost their competitive ability. The indirect ELISA technique provided a more satisfactory method of identifying selected Cyclopia strains under both field and glasshouse conditions. The primary antibodies raised against strains UCT40a, UCT61a and UCT44b gave absorbance readings that were unambiguously negative (0.30 OD405), while those of strain PPRICI3 were ambiguous (0.50 OD405) with many false positive readings (1.0 A405). The indirect ELISA method showed a high level of analytical sensitivity in glasshouse experiments and there were no cross-reactions between the four test strains. The method was also suitable for detecting three of the four test strains in competition studies under field conditions, and can also be used to identify some strains under field conditions.
Conclusion
The antibiotic marker method was found unsuitable for identifying Cyclopia rhizobia in competition experiments in both glasshouse and field conditions. However, the indirect ELISA technique was found suitable for identifying these strains in glasshouse studies. The method was also appropriate for identifying strains UCT40a, UCT44b and UCT61a, but not strain PPRICI3, in field competition studies.
Background
Cyclopia Vent. (Fabaceae) is a shrubby perennial legume endemic to the Mediterranean heathland vegetation (fynbos) of the Western Cape of South Africa [1]. The shoots of several species of the genus have been harvested from the wild for centuries as a source of an herbal infusion known as honeybush tea. Due to its caffeine-free, flavonoid-rich, anti-oxidant properties, the demand for this tea has increased worldwide. To meet this demand requires the cultivation of Cyclopia as a commercial crop. Species of this genus exhibit indeterminate nodulation, and are therefore dependent on symbiotic N2 fixation for their N nutrition [2]. This suggests that manipulation of the symbiosis could lead to increased N nutrition, and hopefully greater tea yields in the low-nutrient environment of the Western Cape.
In Africa, symbiotic N2 fixation in legumes is constrained by many factors, including the paucity of suitable soil rhizobia, low concentrations of nutrients in the soil [3] and the quality of legume root exudates [4]. To maximise growth of the tea-producing Cyclopia species (which are adapted to highly acidic, low N and P environments) would require optimising soil conditions that enhance nodule formation and promote symbiotic N nutrition. This can be achieved via soil amelioration with exogenous nutrient inputs and/or the provision of sufficient quantities of an effective rhizobial symbiont as inoculant [5-7]. Although the initial stages of selecting high N2-fixing strains for inoculant production are usually conducted under controlled conditions in the glasshouse [8-10], subsequent testing is done under field conditions as biotic and abiotic factors can influence strain performance in the field, especially when in competition with indigenous native soil rhizobia. These native strains often out-compete introduced rhizobia for nodule formation in the host plant, leading to poor legume response to inoculation [11-13]. It is therefore important that the competitiveness of selected rhizobial strains is tested in the field prior to recommending their use as commercial inoculants.
The methodology of strain identification inside nodules has, however, often proved difficult, and thus limited this field of research. Three approaches that are routinely used, include 1) antibiotic resistance, 2) serological techniques, and more recently 3) genetic markers. Antibiotic resistance has traditionally been used as a marker in competition studies because the method is simple and requires no specialised equipment [14-19]. The intrinsic antibiotic resistance method can be used as a fingerprint to identify strains; just as mutants resistant to high antibiotic concentrations can be developed as markers for competition experiments.
Serological identification of rhizobial strains involves the use of antibodies raised against surface antigens of the test strain to detect the presence (or absence) of that strain in a suspension through agglutination, immunodiffusion, immunofluorescence or the enzyme-linked immunosorbent assay (ELISA). Because the antigenic properties of rhizobia are stable characteristics [24-26], the serological method is particularly useful in ecological studies as it does not modify the strain or alter its nodulation competitiveness. The immunofluorescence technique has also been successfully used to rapidly identify rhizobial strains [27-29], though this requires expensive equipment and large quantities of labelled antibody.
The ELISA technique is highly specific, reproducible, and commonly used to detect rhizobial strains directly from nodules. Additionally, the method is sensitive, can detect antigens in small nodules, uses small quantities of reagents, is relatively quick, and permits the rapid screening of large nodule samples. It can also detect double strain occupancy of nodules [30-34]. However, cross-reaction with native strains in field soils can lead to false positive results, thus limiting its application. A novel advance in strain detection has been the introduction of stable genetic markers [35-39]and DNA probes [40-43]into test rhizobial strains. However, the insertion of a foreign gene can increase the metabolic burden on the cell [44] and alter its competitive ability [45-47]. Furthermore, the release of such transgenic microbes into the environment is controversial [48-51]. The method also requires specialised knowledge and equipment and is therefore unsuitable for studies in developing countries with low-technology laboratories.
In this study, the suitability of the antibiotic resistance technique (both intrinsic low-resistance fingerprinting and high-resistance marking) and the serological indirect ELISA method were assessed for their ability to detect selected Cyclopia rhizobia under glasshouse and field conditions. Four rhizobial strains (PPRICI3, UCT40a, UCT44b and UCT61a) were used in this study. The strains were isolated from wild Cyclopia species growing in the Western Cape fynbos of South Africa. Strain PPRICI3 was isolated by the Plant Protection Research Institute (PPRI) within the Agricultural Research Council of South Africa, and is currently the recommended inoculant strain for Cyclopia cultivation, while strains UCT40a, UCT61a and UCT44b were isolated in our laboratory and selected for their high symbiotic performance under glasshouse conditions.
Methods
Bacterial strains and growth media
The rhizobia used in this study included strains UCT40a, UCT44b, UCT61a and PPRI13, which were isolated from native Cyclopia species in the Western Cape of South Africa, using yeast-mannitol agar as growth medium. The choice of these four strains out of 39 bacterial isolates was based on their superior symbiotic performance. In general, some of the 39 bacterial isolates were faster in growth (appearing within two days of streaking and producing copious quantities of exopolysaccharide gum, e.g. UCT44b and UCT61a), while phenotypically similar strains only appeared 5 days after streaking.
Antibiotic Resistance
Intrinsic natural resistance to low antibiotic concentrations
The intrinsic resistance of the four Cyclopia strains to the antibiotics streptomycin sulphate (Sigma Chemical Co. Ltd.) and spectinomycin dihydrochloride pentahydrate (Fluka Biochemica Ltd.) was determined by streaking rhizobial culture onto yeast-mannitol agar (YMA52) plates containing incremental concentrations of streptomycin (0, 0.05, 0.1, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, 1.4, 1.6, 1.8, 2.0 and 5.0 μg ml-1) or spectinomycin (0, 0.2, 0.4, 0.6, 0.8, 1.0, 2.0, 5.0, 10.0 and 20.0 μg ml-1). The antibiotics were first sterilised by filtration through a 0.45 μm Millipore filter before addition to autoclaved YMA (cooled to < 50°C). Test strains were grown in yeast-mannitol broth (YMB52) at 20°C to 0.6 OD600, serially diluted to 10-6 and 0.1 ml streaked onto each plate. Plates were streaked in triplicates. Colony-forming units (CFU) per plate were counted after four days of growth. A strain was considered to have intrinsic resistance to an antibiotic if it attained 50% or more growth on antibiotic plates (colony-forming units, CFU, per plate) compared to antibiotic-free control plates.
Antibiotic marking
To develop spontaneous antibiotic-resistant mutants, streptomycin or spectinomycin was incorporated at 10 × the intrinsic resistance level of the test strain into YMA plates. Unmarked parent strains were each grown in YMB to 0.6 OD600 and 0.1 ml (107 – 108 cells), and streaked onto five replicate streptomycin-containing YMA plates. Mutants that appeared spontaneously within five days of growth were isolated, re-streaked onto YMA containing streptomycin, and stored at 0°C. For each test strain, three streptomycin-resistant mutants were randomly selected, grown in YMB broth to 0.6 OD600 and 0.1 ml streaked onto each of five replicate spectinomycin-marked plates. To develop a double marker, the spontaneous mutants were isolated and re-streaked onto plates containing both antibiotics. After growth, they were re-streaked on YMA slants containing the two antibiotics and incubated to grow at 20°C for storage at 0°C. For strain PPRICI3, only streptomycin-resistant mutants were obtained, as no doubly marked colonies appeared after 10 days of growth. For strain UCT40a, only two doubly-marked colonies were obtained.
Integrity test using plants in Leonard jars
Leonard jar assemblies supplied with N-free 1/4 strength Hoagland's nutrient solution [53] were used to assess the competitive ability of marked strains compared to their unmarked parents. Treatments included jars inoculated with the parent strains alone, the marked strains alone and 1:1 mixtures of parent and marked strains. Uninoculated jars served as negative controls. Jars were autoclaved prior to planting with pre-germinated seedlings of Cyclopia maculata raised from surface-sterilized seed. C maculata is a fast-growing species on which all parent strains are effective. Five replicate jars were used, each with one seedling. The glasshouse provided a 12-h day and night cycle, with a temperature range of 16 – 28°C.
Treatment strains were grown in YMB to 0.6 OD600, diluted to 0.2 OD600 and each jar inoculated with 1 ml of the appropriate strain. For the mixed treatments, the strains were mixed 1:1 before inoculation. Cell numbers were estimated as CFU ml-1 culture by streaking serial dilutions of the culture onto antibiotic-free YMA plates in triplicates and counting CFU after fours days of growth. Cell density across all strains ranged from 1 × 108 to 5 × 108 CFU ml-1 culture.
Plants were harvested at 16 weeks and each separated into shoots, roots and nodules. Nodules were counted and weighed, while shoots and roots were oven-dried at 60°C for dry matter determination. Rhizobia were isolated from the larger nodules (5 to 10 nodules per jar) as described by Vincent52. Each isolate was streaked onto three replicate plates containing the appropriate concentrations of the antibiotics streptomycin and spectinomycin for the test (Table 1). Three antibiotic-free plates were included for comparison. If a nodule isolate achieved more than 50% growth on antibiotic plates relative to growth on antibiotic-free plates, it was considered resistant to the antibiotic and therefore the marked strain occupying that nodule. The number of nodules occupied by the marked strain provided a measure of its competitive ability.
Table 1.
Levels of antibiotics used to develop resistant mutant strains of Cyclopia.
| Antibiotic | Concentration of antibiotics used (μg.ml-1) | |||
|---|---|---|---|---|
| PPRICI3 | UCT40a | UCT44b | UCT61a | |
| Streptomycin | 1 | 1 | 10 | 5 |
| Spectinomycin | 10 | 5 | 80 | 80 |
Nodule occupancy data were pooled for each test strain and analysed using a χ2 test against a null hypothesis of 50% expected nodule occupancy for equal competitive ability between marked and parent strains. The appropriateness of data pooling was assessed using heterogeneity χ2 tests [54]. For the marked strain treatments, the ability to grow on antibiotic plates provided a measure of the strains retention of antibiotic resistance after plant passage, while a reduction in growth indicated loss of the antibiotic marker.
Indirect ELISA technique
The indirect ELISA technique, modified from Kishinevsky and Maoz [55], was tested here for its ability to identify Cyclopia rhizobia under both glasshouse and field conditions. In the indirect ELISA method, the antigen is adsorbed, followed by the application of purified primary antibody and a single secondary antibody-conjugate. The antibody-conjugate (usually goat anti-rabbit conjugate) is commercially available and can be used in conjunction with a number of strain-specific antibody preparations. The method is simpler, but has lower analytical sensitivity than the direct method [55,56].
Production of strain-specific primary antibodies
The four test strains used in this study were grown in a defined broth medium containing 0.5 g K2HPO4, 0.2 g MgSO4.7H20, 0.1 g NaCl, 0.5 g KHPO4 and 10 g mannitol in 1 l distilled water53 and incubated at 20°C to obtain 0.4 OD600. To remove exopolysaccharides (produced in large quantities by strains UCT44b and UCT61a), the bacterial cells were washed three times by repeated centrifugation in phosphate-buffered saline (PBS) solution. The final sediment was suspended in 10 ml saline solution (150 mM NaCl) to a final concentration of > 109 CFU ml-1.
Antibodies were prepared against each test strain using adult New Zealand White rabbits. The rabbits were bled prior to inoculation to assess their pre-inoculation antibody levels. One rabbit was used for each test strain and was injected with the appropriate antigen according to the following protocol: Day 1: 0.5 ml intramuscular injections into each hind leg (with equal parts Freund's complete adjuvant mixed prior to injection); Day 14: 1 ml intravenous injection; Day 21: 1 ml intravenous injection; Day 28: 1 ml intravenous injection; Day 35: trial bleed to check antiserum titre; Day 37: bleed by cardiac puncture after 0.15 ml intravenous acetylpromazine (sedative) injection. Intravenous injections and trial bleeds were done via the marginal ear vein.
Collected blood was incubated for 1 h at 37°C to facilitate clotting and then held at 4°C overnight to extrude serum. The serum was removed, centrifuged to remove residual cells and stored at -20°C in 0.5 ml aliquots. Antiserum titres were tested using the long agglutination test of Vincent [52]. No precipitation reactions occurred with the pre-inoculation sera, but strong agglutinations occurred with the test antisera. Antisera agglutination titres were 1:600, 1:200, 1:400 and 1:500 for strains PPRICI3, UCT40a, UCT44b and UCT61a, respectively.
Antigen preparation from roots nodules
Cyclopia maculata seedlings were grown on nutrient-agar slants in individual sterile tubes. After three weeks of growth, the tubes were inoculated with test strains using three replicate tubes per strain and three uninoculated tubes as a negative control. The inoculated treatments included pure cultures of the four test strains and soil washes prepared from three field soils. The soils were sampled from three farms across the Western Cape: Waboomskraal near George (33° S, 22° W, CD), Kanetberg near Barrydale (33° S, 20° W, DD) and Reins Farms near Gourtismond (34° S, 21° W, BC). The three fields had no history of Cyclopia cultivation or the species. Nodules were harvested from the seedlings after sixteen weeks of growth. For each strain, three large nodules were harvested per replicate tube and 10 nodules per tube for each soil wash treatment. This gave a total of 9 nodules (all containing the same antigen) for each rhizobial strain and 30 nodules for each soil-wash treatment (with all 30 nodules probably containing different antigens). Antigens were extracted from the nodules by crushing individual nodules (mass ≈ 0.15 g) in 50 μl PBS and transferring 10 μl of the nodule macerate into 1 ml PBS (to give a low antigen concentration). The antigens were stored in 1.5 ml Eppendorf vials at 0°C and used within 48 h.
Testing the analytical sensitivity of antigen × antibody reactions
Checkerboard assays were carried out to determine the concentration effect of primary antibody (described above) and secondary antibody-conjugate (goat anti-rabbit antibody conjugated to alkaline-A-phosphatase, purchased from Sigma-Aldrich Chemical Co. Ltd.) on the sensitivity of antigen detection. The primary antibody concentration had no effect on absorbance readings, whereas a lower secondary antibody concentration of 1:4000 (diluted in 1% non-fat milk-PBS solution) significantly increased the analytical sensitivity of the test (data not shown).
Two sets of cross-reaction tests were carried out. The first used the antigens prepared from the four test strains (9 antigens per strain) and the second the soil-wash antigens (90 antigens prepared from three field soils). All possible primary antibody × antigen combinations were tested in duplicates. Wells of polysorp immunoplates (AEC-Amersham Co.) were coated with 100 μl of antigen and left at 5°C overnight. The plates were then washed three times with PBS (250 μl per well) and blocked with 200 μl 1% non-fat milk in PBS per well. After incubating at room temperature for two hours, 100 μl of the appropriate primary antibody (1:4000 diluted in 1% non-fat milk-PBS) was added to each well and the plates incubated for two hours at room temperature. After washing in PBS, 100 μl of secondary antibody was added to each well (1:4000 diluted in 1% non-fat milk-PBS) and the plates incubated at 37°C for one hour before washing (as before). Finally, a chromogenic enzyme substrate, p-nitrophenyl phosphate in 10% Tris-HCl buffer (Sigma-Aldrich chemical Co.), was added at the rate of 100 μl per well and the plates incubated in the dark and read when absorbance readings reached 1.0 OD405 for positive controls (approximately 30 min).
Results
Antibiotic Resistance
Intrinsic natural resistance of Cyclopia strains to low antibiotic concentrations
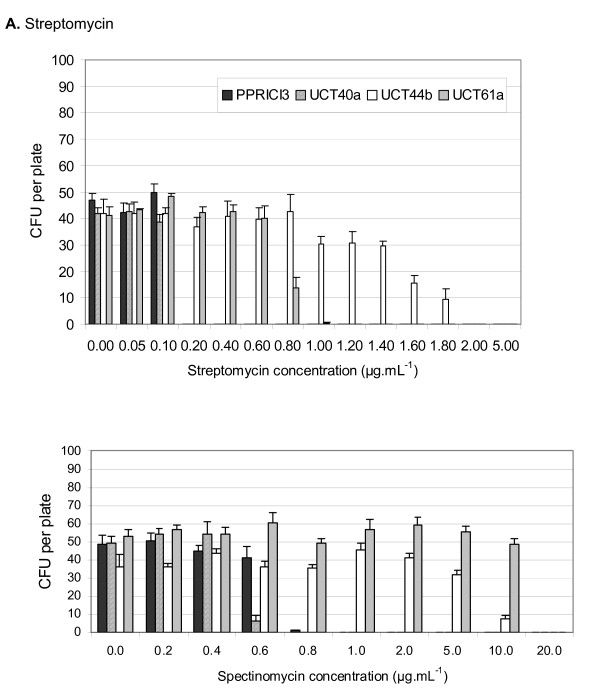
The Cyclopia strains fell into two distinct groups regarding their intrinsic antibiotic resistance, with strains UCT44b and UCT61a showing greater resistance than strains UCT40a and PPRICI3 to low concentrations of both streptomycin and spectinomycin (Figure 1). Strain UCT44b was tolerant to 1.4 – 1.6 μg ml-1 streptomycin and to 5.0 – 10 μg ml-1 spectinomycin. Strain UCT61a showed a slightly lower tolerance to streptomycin (about 0.6 – 0.8 μg ml-1) but exhibited a higher tolerance of spectinomycin (about 10.0 – 20.0 μg ml-1). Strains UCT40a and PPRICI3, on the other hand, were highly sensitive to low concentrations of the two antibiotics, with resistance to 0.1 – 0.2 μg ml-1 streptomycin and 0.4 – 0.8 μg ml-1 spectinomycin.
Figure 1.
Intrinsic natural resistance of Cyclopia rhizobial strains to low concentrations of streptomycin sulphate (A) and spectinomycin dihydrochloride pentahydrate (B). Values are mean colony-forming units (CFU) per plate (n = 3 and error bars represent standard errors).
Nodulation and competitive ability of antibiotically-marked versus unmarked strains
The uninoculated control plants were not nodulated and thus showed significantly lower plant dry matter yield compared to the inoculated (nodulated) seedlings (P < 0.01, Table 2). The nodulation and N2-fixing ability of the mutants of strains PPRICI3, UCT44b and UCT61a were not altered by the antibiotic marker, as there were no significant differences in plant biomass, nodule mass or nodule number between strains (P < 0.05, Table 2). Marked strain UCT40aMkd3 produced no nodules, thus showing loss of symbiotic ability. Mutant strains UCT40aMkd1 and UCT40aMkd2 however showed no loss of their nodulation capacity compared to their parent strain (Table 2).
Table 2.
Nitrogen-fixing ability of marked rhizobial strains.
| Treatment | Total dry weight (mg) | Nodule biomass (mg) | Nodule number |
|---|---|---|---|
| Uninoculated | 0.06 ± 0.04 a | 0.00 ± 0.00 a | 0.0 ± 0.0 a |
| Inoculated | 0.72 ± 0.01 b | 33.33 ± 0.07 b | 19.6 ± 0.1 b |
| t(1,83) | 2.58 ** | 2.60 ** | 3.49 ** |
| PPRICI3 Parent | 0.87 ± 0.13 | 18.60 ± 0.64 | 14.8 ± 0.5 |
| PPRICI3Mkd1 | 0.70 ± 0.14 | 23.60 ± 0.78 | 13.2 ± 0.7 |
| PPRICI3Mkd2 | 0.68 ± 0.10 | 15.40 ± 0.48 | 11.2 ± 0.5 |
| PPRICI3Mkd3 | 1.26 ± 0.13 | 18.00 ± 0.62 | 12.6 ± 0.5 |
| F(3,16) | 2.06 ns | 0.51 ns | 0.17 ns |
| UCT40a Parent | 2.26 ± 0.19 a | 75.76 ± 1.36 a | 20.0 ± 0.7 a |
| UCT40aMkd1 | 1.83 ± 0.23 a | 74.70 ± 1.38 a | 24.3 ± 0.7 a |
| UCT40aMkd2 | 2.13 ± 0.20 a | 81.94 ± 1.20 a | 31.6 ± 0.7 a |
| UCT40aMkd3 | 0.12 ± 0.06 b | 0.00 ± 0.00 b | 0.0 ± 0.0 b |
| F(3,16) | 4.35 * | 10.30 ** | 8.13 ** |
| UCT44b Parent | 0.37 ± 0.13 | 31.25 ± 0.43 | 18.0 ± 0.4 |
| UCT44bMkd1 | 0.90 ± 0.12 | 56.00 ± 0.81 | 33.4 ± 0.8 |
| UCT44bMkd2 | 0.51 ± 0.09 | 23.20 ± 0.47 | 18.4 ± 0.5 |
| UCT44bMkd3 | 0.66 ± 0.12 | 25.60 ± 0.60 | 18.2 ± 0.6 |
| F(3,16) | 1.61 ns | 2.22 ns | 2.94 ns |
| UCT61a Parent | 0.84 ± 0.12 | 39.82 ± 0.93 | 25.4 ± 0.7 |
| UCT61aMkd1 | 0.54 ± 0.09 | 22.64 ± 0.44 | 16.0 ± 0.5 |
| UCT61aMkd2 | 0.61 ± 0.10 | 34.02 ± 0.73 | 21.6 ± 0.5 |
| UCT61aMkd3 | 1.07 ± 0.14 | 48.10 ± 1.04 | 32.0 ± 0.8 |
| F(3,16) | 2.79 ns | 1.63 ns | 1.79 ns |
Values are mean ± SE (n = 5) and different letters within a column indicate significant differences. ** Denotes a significant effect at P < 0.01, * = P < 0.05 and ns = no significant effect.
The antibiotically-marked strains showed varied abilities to compete with their parent strains for nodule occupancy (Table 3). The mutants of UCT44b and UCT61a showed significantly reduced competitive abilities, while those of PPRICI3 retained their competitiveness relative to the parent strain. Marked strain UCT40aMkd2 also retained its competitive ability, while mutant strain UCT40aMkd1 showed increased competitive ability compared to its unmarked parent (Table 3). The marked strains also varied in their retention of the antibiotic marker after plant passage (Table 4). Mutants of strain PPRICI3 retained their resistance marker, while those of UCT40a and UCT44b showed a slight reduction in the number resistant to antibiotics. Two of the three UCT61a mutants (i.e. UCT61aMkd1 and UCT61aMkd2) lost their antibiotic markers after plant passage (Table 4).
Table 3.
Competitiveness of antibiotically-marked strains compared to their unmarked parents.
| Treatment | Number of isolates tested | Number able to grow on YMA + antibiotics | % nodule occupancy by marked strain | Competitive ability of marked strain |
|---|---|---|---|---|
| UCT40a + UCT40aMkd1 | 40 | 30 | 75.0 * | I |
| UCT40a + UCT40aMkd2 | 28 | 14 | 50.0 | U |
| UCT44b + UCT44bMkd1 | 18 | 4 | 22.2 * | R |
| UCT44b + UCT44bMkd2 | 38 | 12 | 31.6 * | R |
| UCT44b + UCT44bMkd3 | 26 | 10 | 38.5 | U |
| UCT61a + UCT61aMkd1 | 50 | 0 | 0.0 * | R |
| UCT61a + UCT61aMkd2 | 52 | 0 | 0.0 * | R |
| UCT61a + UCT61aMkd3 | 60 | 0 | 0.0 * | R |
| PPRICI3 + PPRICI3Mkd1 | 35 | 21 | 60.0 | U |
| PPRICI3 + PPRICI3Mkd2 | 31 | 19 | 61.2 | U |
| PPRICI3 + PPRICI3Mkd3 | 31 | 10 | 32.3 | U |
* Denotes significant deviation from the expected frequency of 50% nodule occupancy using a χ2 test on pooled data, P < 0.05. Symbols indicate I = increased, U = unchanged and R = reduced competitive ability of the marked strain compared to its unmarked parent.
Table 4.
Retention of the antibiotic resistance marker after plant passage.
| Marked Strain | Number of isolates tested | Number able to grow on YMA + antibiotics | % retention of antibiotic resistance |
|---|---|---|---|
| UCT40aMkd 1 | 25 | 23 | 92 |
| UCT40aMkd2 | 25 | 25 | 100 |
| UCT44bMkd 1 | 20 | 20 | 100 |
| UCT44bMkd 2 | 21 | 17 | 81 |
| UCT44bMkd 3 | 19 | 16 | 84 |
| UCT61aMkd 1 | 15 | 0 | 0 |
| UCT61aMkd 2 | 14 | 0 | 0 |
| UCT61aMkd 3 | 13 | 13 | 100 |
| PPRICI3Mkd 1 | 19 | 19 | 100 |
| PPRICI3Mkd 2 | 19 | 19 | 100 |
| PPRICI3Mkd3 | 20 | 20 | 100 |
Indirect ELISA assays
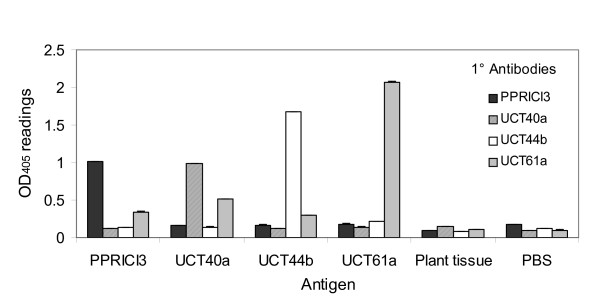
Results of the cross-reaction tests using pure antigens of PPRICI3, UCT40a, UCT44b and UCT61a (isolated from nodules of plants inoculated with these strains) are shown in Figure 2. Absorbance readings were clear and unambiguous; there were no cross-reactions, i.e. no false positive results for non-appropriate antigen × antibody combinations. In addition, non-specific adsorption using plant tissue or PBS substrate was low (≤ 0.15 OD405). There were some variations in the reactivity of the primary antibodies. For example, antibodies raised against strains UCT44b and UCT61a produced readings of ≥ 1.50 OD405, while strains PPRICI3 and UCT40a gave lower positive readings of about 1.0 OD405. The negative readings for all strains were ≤ 0.50 OD405, but were higher for strain UCT40a than the other three test strains.
Figure 2.
Cross-reaction tests of indirect ELISAs involving primary antibodies assayed against 4 test antigens, with plant tissue and PBS as controls. Nine antigens prepared for each test strain were assayed in duplicates. Error bars representing standard errors ranged from 0.001 – 0.006 OD405.
Cross-reaction tests using random antigens extracted from three field soils produced less defined readings with a number of distinct cross-reactions (Table 5). The primary antibodies raised against strains UCT40a and UCT61a gave absorbance readings that were unambiguously negative (≤ 0.30 OD405). Optical density readings were higher (≤ 0.50 OD405) for the antibody raised against strain UCT44b, but all readings were distinguishable as negative. The readings for the primary antibody raised against strain PPRICI3, on the other hand, were ambiguous (≥ 0.50 OD405) as the antibody produced many false positive readings (≥ 1.0 A405). The cross-reactions were more than 50% for each of the three field soils with the primary antibody of strain PPRICI3. Antigens isolated from the soil of Rein's Farms notably produced 90% false positive readings with the primary antibody raised against strain PPRICI3 in the indirect ELISA test (Table 5).
Table 5.
Cross-reaction tests of indirect ELISAs involving primary antibodies assayed against random antigens extracted from 3 different field soils.
| Antigen (field soil site) | 1° antibody | |||
|---|---|---|---|---|
| PPRICI3 | UCT40a | UCT44b | UCT61a | |
| Waboomskraal | 60 | 0 | 0 | 0 |
| Rein's Farms | 90 | 0 | 0 | 0 |
| Kanetberg | 55 | 0 | 3 | 0 |
Data are % antigens tested positive (≥ 1.0 OD405), n = 30, assayed in duplicates.
Discussion
Suitability of intrinsic antibiotic resistance for identification of Cyclopia rhizobia
The four Cyclopia strains fell into two distinct pairs with regard to their intrinsic natural resistance to the antibiotics streptomycin and spectinomycin. In the 0.0 – 0.1 μg ml-1 range, all four strains were resistant to streptomycin and could therefore not be distinguished by this technique. Over 0.2 μg ml-1, UCT40a and PPRICI3 were sensitive and did not grow, while UCT44b and UCT61a were resistant and could therefore be distinguished from the other two but not between themselves. However, from 1.2 – 1.8 μg ml-1 streptomycin, only strain UCT44b could grow and this strain could therefore be detected in a mixture with the other three strains. Test strain resistance to spectinomycin was similar in pattern to streptomycin, in that, all strains were resistant to the 0.0 – 0.6 μg ml-1 range, and were therefore not identifiable among them. However, between 1.0 and 10.0 μg ml-1 spectinomycin, only strains UCT44b and UCT61a could grow in the medium and could therefore be distinguished from any one of the other two in a mixture, but again not between themselves. Based on the distinct groupings obtained in this study, in hindsight, it would have been interesting to screen a broader population of isolates for their intrinsic antibiotic resistance.
It is interesting to note that the strains used could also be grouped with respect to colony characteristics such as colony morphology. Strains UCT40a and PPRICI3, which showed low resistance, both form small, discrete, opaque colonies with little exopolysaccharide gum production. Evidence from molecular analysis show that these two strains are in fact the same species [57]. Strains UCT44b and UCT61a, on the other hand, were found to be genetically different from each other and from strains PPRICI3 and UCT40a [57]. They form fast-growing colonies with large quantities of translucent exopolysaccharide gum. Our data on antibiotic resistance and colony morphology of the four test strains are consistent with the findings of other studies, which show that fast-growing "wet" colonies have higher antibiotic resistance than "dry" colonies [58,59].
Antibiotic markers as a tool for the detection of Cyclopia rhizobia
Analysis of root nodules for strain occupancy in the competition experiments conducted in Leonard jars revealed significant differences in the symbiotic ability and competitiveness of the antibiotic mutants relative to their unmarked parents. Marked strains from the intrinsically low resistance group (except strain UCT40aMkd3) performed well, retaining their symbiotic ability, competitive capacity, and their antibiotic-resistance marker tags. Strain UCT40aMkd1 even showed increased competitive ability compared to its parent strain. Marked strains of UCT44b and UCT61a, on the other hand, exhibited reduced competitive ability relative to their parent strains. This reduction in competitive ability was distinct for UCT61aMkd3, which showed zero nodule occupancy in competition with its parent strain. Strains UCT61aMkd1 and UCT61aMkd2 also lost their competitive ability, but this was most likely a reflection of the strains being unidentifiable through losing their antibiotic marker tag. Strain UCT44bMkd1 also showed some loss of its antibiotic resistance marker. The loss of symbiotic ability in strains with antibiotic tagging could suggest loss of their symbiotic plasmids. However because little is known about the rhizobia from native South African legumes, we also do not know anything about their plasmids and plasmid localization of symbiotic genes in these Cyclopia rhizobia. Whatever the case, this suggests genetic instability in the rhizobial strains isolated from Cyclopia species.
Only marked strains of PPRICI3 could be confidently used in competition studies in the glasshouse, as they retained their symbiotic trait, their antibiotic markers and showed unchanged competitive abilities. The antibiotic markers did not therefore allow for a full comparative study across the four test strains. The method is also dubious for field studies as strains from the high resistance group may exist in the field environment, and thus prove intrinsically resistant to the marker level used for low-level fingerprinting (Table 1). As all four strains were isolated from the same region and from the same area proposed for Cyclopia cultivation (the fynbos in the Western Cape of South Africa), the presence of intrinsically high-resistance rhizobia in the field is probable and may present problems when identifying antibiotically-marked strains from the low resistance group in field competition experiments. In addition, concerns have been raised regarding the consequences of releasing antibiotic-resistant bacteria into field environments [60,61,49].
Indirect ELISA technique
The indirect ELISA technique is more suitable than the antibiotic resistance methods for identifying Cyclopia strains in nodules in glasshouse and field studies. There were no cross-reactions between the four test strains, showing that they are antigenically different (Figure 2). All four primary antibodies reacted strongly with their appropriate homologous strain, producing absorbance readings that were easily distinguished from heterologous strains, and thus made this technique ideal for strain identification in comparative glasshouse and field competition studies.
The antibodies raised against strains UCT40a and UCT61a did not cross-react with antigens from any of the three field soils and the antibody raised against strain UCT44b provided only one ambiguous positive result (0.69 OD405 with an antigen derived from the Kanetberg soil), but did not cross-react with antigens from the other field sites (Table 5). The antibody raised against strain PPRICI3, on the other hand, produced many false positive results, making the indirect ELISA method unsuitable for identifying this strain in field experiments. The reason for the high level of cross-reactions with the PPRICI3 antibody remains unclear. According to the polyphasic taxonomic investigations of Kock [53], strain PPRICI3 is genetically identical to strain UCT40a. However, because the two strains produced antibodies with different specificity levels, clearly indicates they differ in their surface antigen characteristics.
Conclusion
The antibiotic markers were found to be unsuitable for identifying Cyclopia rhizobia in competition experiments under both glasshouse and field conditions. In contrast, the indirect ELISA technique was very successful in identifying the four Cyclopia strains under glasshouse conditions, as well as identifying strains UCT40a, UCT44b and UCT61a (but not strain PPRICI3) in field studies.
Authors' contributions
AS conducted the studies as a PhD student in FD's laboratory, and prepared the draft paper. FD conceptualized the study, supervised all aspects of the work, and critically edited the paper. All authors read and approved the final manuscript.
Contributor Information
Amy C Spriggs, Email: amy.spriggs@sanbi.org.
Felix D Dakora, Email: dakorafd@tut.ac.za.
Acknowledgements
This research was supported with funds from the Dr. C. Fred Bentley Fellowship (International Development Research Centre, Canada) and B.P. Southern Africa Ltd to AC Spriggs, and with a grant from the National Research Foundation, Pretoria, to FDD.
References
- Arnold T, de Wet BC. Plants of Southern Africa. National Botanical Institute of South Africa; 1994. [Google Scholar]
- Spriggs AC, Stock WD, Dakora FD. Influence of mycorrhizal associations on foliar δ15 N values of legume and non-legume shrubs and trees in the fynbos of South Africa: Implications for estimating N2 fixation using the 15N natural abundance method. Plant and Soil. 2003;255:495–502. doi: 10.1023/A:1026044831178. [DOI] [Google Scholar]
- Dakora FD, Keya SO. Contribution of legume nitrogen fixation to sustainable agriculture in Sub-Saharan Africa. Soil Biology and Biochemistry. 1997;29:809–817. doi: 10.1016/S0038-0717(96)00225-8. [DOI] [Google Scholar]
- Dakora FD, Phillips DA. Root exudates as mediators of mineral acquisition in low nutrient environments. Plant and Soil. 2002;245:35–47. doi: 10.1023/A:1020809400075. [DOI] [Google Scholar]
- Peterson HL, Loynachan TE. The significance and application of Rhizobium in agriculture. International Review of Cytology. 1981;13:311–331. [Google Scholar]
- Brockwell J, Bottomley PJ. Recent advances in inoculant technology and prospects for the future. Soil Biology and Biochemistry. 1995;27:683–697. doi: 10.1016/0038-0717(95)98649-9. [DOI] [Google Scholar]
- Peoples MB, Ladha JK, Herridge DF. Enhancing legume N2 fixation through plant and soil management. Plant and Soil. 1995;174:83–101. doi: 10.1007/BF00032242. [DOI] [Google Scholar]
- Svenning MM, Junttila O, Solheim B. Symbiotic growth of indigenous white clover (Trifolium repens) with local Rhizobium leguminosarum biovar trifolii. Physiologia Plantarum. 1991;83:381–389. doi: 10.1111/j.1399-3054.1991.tb00109.x. [DOI] [Google Scholar]
- Howieson JG, Malden J, Yates RJ, O'Hara GW. Techniques for the selection and development of elite inoculant strains of Rhizobium leguminosarum in southern Australia. Symbiosis. 2000;28:33–48. [Google Scholar]
- Fening JO, Danso SKA. Variation in symbiotic effectiveness of cowpea bradyrhizobia indigenous to Ghanaian soils. Applied Soil Ecology. 2002;21:23–29. doi: 10.1016/S0929-1393(02)00042-2. [DOI] [Google Scholar]
- Carter JM, Tieman JS, Gibson AH. Competitiveness and persistence of strains of rhizobia for faba bean in acid and alkaline soils. Soil Biology and Biochemistry. 1995;27:617–623. doi: 10.1016/0038-0717(95)98640-A. [DOI] [Google Scholar]
- Denton MW, Reeve WG, Howieson JC, Coventry DR. Competitive abilities of common field isolates and a comercial strain of Rhizobium leguminosarum bv. trifolii for clover nodule occupancy. Soil Biology and Biochemistry. 2003;35:1039–1048. doi: 10.1016/S0038-0717(03)00146-9. [DOI] [Google Scholar]
- Okogun JA, Sanginga N. Can introduced and indigenous rhizobial strains compete for nodule formation by promiscuous soybean in the moist savanna agroecological zone of Nigeria? Biology and Fertility of Soils. 2003;38:26–31. doi: 10.1007/s00374-003-0611-8. [DOI] [Google Scholar]
- Josey DP, Beynon JL, Johnston AWB, Beringer JE. Strain identification in Rhizobium using intrinsic antibiotic resistance. Journal of Applied Bacteriology. 1979;46:343–350. [Google Scholar]
- Jones DG, Bromfield ESP. Studies on double strain occupancy of nodules and the competitive ability of Rhizobium trifolii on red and white clover grown in soil and agar. Annals of Applied Biology. 1979;94:51–59. [Google Scholar]
- Schwinghamer EA, Dudman WF. In: Methods for Evaluating Biological Nitrogen Fixation. Bergersen FJ, editor. New York: John Wiley; 1980. Methods of identifying strains of diazotrophs; pp. 337–365. [Google Scholar]
- Stein M, Bromfield ESP, Dye M. An assessment of a method based on intrinsic antibiotic resistance for identifying Rhizobium strains. Annals of Applied Biology. 1982;101:261–267. doi: 10.1111/j.1744-7348.1982.tb00821.x. [DOI] [Google Scholar]
- Dakora FD. Use of intrinsic antibiotic resistance for characterisation and identification of rhizobia from nodules of Vigna unguiculata (L) Walp. and Phaseolus vulgaris (L) Acta Microbiologica Polonica. 1985;34:187–196. [PubMed] [Google Scholar]
- Ramírez ME, Israel DW, Wollum AG II. Using spontaneous antibiotic-resistant mutants to assess competitiveness of bradyrhizobial inoculants for nodulation of soybean. Canadian Journal of Microbiology. 1998;44:753–758. doi: 10.1139/cjm-44-8-753. [DOI] [Google Scholar]
- Zelazna-Kowalska I. Correlation between streptomycin resistance and infectiveness in Rhizobium trifolii. Plant and Soil. 1971;special:67–71. doi: 10.1007/BF02661839. [DOI] [PubMed] [Google Scholar]
- Turco RF, Moorman TB, Bezdicek DF. Effectiveness and competitiveness of spontaneous antibiotic-resistant mutants of Rhizobium leguminosarum and Rhizobium japonicum. Soil Biology and Biochemistry. 1986;18:259–262. doi: 10.1016/0038-0717(86)90059-3. [DOI] [Google Scholar]
- Lochner HH, Strijdom BW, Law IJ. Unaltered nodulation competitiveness of a strain of Bradyrhizobium sp. (Lotus) after a decade in soil. Applied and Environmental Microbiology. 1989;55:3000–3008. doi: 10.1128/aem.55.11.3000-3008.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lochner HH, Strijdom BW, Steyn PL. Limitations of colony morphology and antibiotic resistance in the identification of a Bradyrhizobium sp. (Lotus) in soil. Biology and Fertility of Soils. 1991;11:128–134. doi: 10.1007/BF00336377. [DOI] [Google Scholar]
- Brockwell J, Schwinghamer EA, Gault RR. Ecological studies of root-nodule bacteria introduced into field environments V: A critical examination of the stability of antigenic and streptomycin-reistant markers for identification of strains of Rhizobium trifolii. Soil Biology and Biochemistry. 1977;9:19–24. doi: 10.1016/0038-0717(77)90056-6. [DOI] [Google Scholar]
- Diatloff A. Ecological studies of root-nodule bacteria introduced into field environments 6: Antigenic and symbiotic stability in Lotononis rhizobia over a 12-year period. Soil Biology and Biochemistry. 1977;9:85–88. doi: 10.1016/0038-0717(77)90042-6. [DOI] [Google Scholar]
- Berger JA, May SN, Berger LR, Bohlool BB. Colorometric enzyme-limked immunosorbent assay for the idenitification of strains of Rhizobium in culture and in the nodules of lentils. Applied and Environmental Microbiology. 1979;37:642–646. doi: 10.1128/aem.37.3.642-646.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohlool BB, Schmidt EL. Immunofluorescent detection of Rhizobium japonicum in soils. Soil Science. 1970;10:229–236. doi: 10.1097/00010694-197010000-00002. [DOI] [Google Scholar]
- Kosslak RM, Bohlool BB, Dowdle S, Sadowsky MJ. Competition of Rhizobium japonicum strains in early stages of soybean nodulation. Applied and Environmental Microbiology. 1983;46:870–873. doi: 10.1128/aem.46.4.870-873.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Josephson KL, Bourque DP, Bliss FA, Pepper IL. Competitiveness of Kim 5 and Viking 1 bean rhizobia: Strain by cultivar interactions. Soil Biology and Biochemistry. 1991;23:249–253. doi: 10.1016/0038-0717(91)90060-W. [DOI] [Google Scholar]
- Fuhrmann J, Wollum AG II. Simplified Enzyme-linked Immunosorbent Assay for Routine Identification of Rhizobium Japonicum Antigens. Applied and Environmental Microbiology. 1985;49:1010–1013. doi: 10.1128/aem.49.4.1010-1013.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martensson AM, Gustafsson JG, Ljunggren HD. A modified, highly sensitive enzyme-linked immunosorbent assay (ELISA) for Rhizobium meliloti strain identification. Journal of General Microbiology. 1984;130:247–253. [Google Scholar]
- Smith GB, Wollum AG. Nodulation of Glycine max by six Bradyrhizobium japonicum strains with different competitive abilities. Applied Environmental Microbiology. 1989;55:1957–1962. doi: 10.1128/aem.55.8.1957-1962.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- George MLC, Robert FM. Competition among Rhizobium leguminosarum bv. phaseoli strains for nodulation of common bean. Canadian journal of Microbiology. 1992;38:157–160. doi: 10.1139/m92-026. [DOI] [PubMed] [Google Scholar]
- Brutti L, Rivero E, Basurco JCP, Nicolas M, Iriarte L, Abbiati N, Ljunggren H, Martensson A. Persistence of Bradyrhizobium japonicum in arable soils of Argentina. Applied Soil Ecology. 1998;10:87–94. doi: 10.1016/S0929-1393(98)00038-9. [DOI] [Google Scholar]
- Jefferson RA. The Gus reporter-gene system. Nature. 1989;342:157–160. doi: 10.1038/342837a0. [DOI] [PubMed] [Google Scholar]
- Meighen EA. Molecular biology of bacterial bioluminescence. Microbiology Reviews. 1991;55:123–142. doi: 10.1128/mr.55.1.123-142.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streit W, Botero L, Werner D, Beck D. Competition for nodule occupancy on Phaseolus vulgaris by Rhizobium etli and Rhizobium tropici strains can be effectively monitored in an utisol during the early stages of growth using a constitutive GUS gene fusion. Soil Biology and Biochemistry. 1995;27:1075–1081. doi: 10.1016/0038-0717(95)00003-W. [DOI] [Google Scholar]
- Wilson KJ, Peoples MB, Jefferson RA. New techniques for studying compeition by rhizobia and for assessing nitrogen fixation in the field. Plant and Soil. 1995;174:241–253. doi: 10.1007/BF00032250. [DOI] [Google Scholar]
- Sessitsch A, Hardarson G, de Vos WM, Wilson KJ. Use of marker genes in competition studies of Rhizobium. Plant and Soil. 1998;204:35–45. doi: 10.1023/A:1004322809676. [DOI] [Google Scholar]
- Steffan RJ, Goksoyr J, Bej AK, Atlas RM. Recovery of DNA from soils and sediments. Applied Environmental Microbiology. 1988;54:2908–2915. doi: 10.1128/aem.54.12.2908-2915.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armann R, Springer W, Ludwig W, Gortz HD. Identification in situ and phylogeny of uncultured bacterial endosymbionts. Nature. 1991;351:92–96. doi: 10.1038/351161a0. [DOI] [PubMed] [Google Scholar]
- Krishnan BH, Pueppke SG. A nodC-lacZ gene fusion in Rhizobium fredii facilitates direct assessment of competition for nodulation of soybean. Canadian Journal of Microbiology. 1992;38:515–519. [Google Scholar]
- Bjourson AJ, Stone CE, Cooper JE. Combined subtraction hybridization and polymerase chain reaction amplification procedure for isolation of strain specific Rhizobium DNA sequences. Applied and Environmental Microbiology. 1992;58:2296–2301. doi: 10.1128/aem.58.7.2296-2301.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCormick D. Detection technology: the key to environmental biotechnology. Biotechnology. 1986;4:419–422. doi: 10.1038/nbt0586-419. [DOI] [Google Scholar]
- Pankhurst CE, MacDonald PE, Reeves JM. Enhanced nitrogen fixation and competitiveness for nodulation of Lotus pedunculatus by a plasmid-cured derivative of Rhizobium loti. Journal of General Microbiology. 1986;132:2321–2328. [Google Scholar]
- Law IJ, Strijdom BW. Negative effects of agrocin 84-encoding Agrobacterium plasmids on symbiotic properties of Rhizobium meliloti. Archives of Microbiology. 1989;152:463–467. doi: 10.1007/BF00446930. [DOI] [Google Scholar]
- Liu R, Tran VM, Schmidt EL. Nodulating competitiveness of a non-motile Tn7 mutant of Bradyrhizobium japonicum in non-sterile soil. Applied Environmental Microbiology. 1989;55:1895–1900. doi: 10.1128/aem.55.8.1895-1900.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veal DA, Stokes HW, Grant D. Genetic exchange in natural communities. Advanced Microbiology Ecology. 1992;12:383–430. [Google Scholar]
- van Veen JA, van Overbeek LS, van Elsas JD. Fate and activity of microorganisms introduced into soil. Microbiology Molecular Biology Reviews. 1997;61:121–135. doi: 10.1128/mmbr.61.2.121-135.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davidson J. Genetic exchange between bacteria and the environment. Plasmid. 1999;42:73–91. doi: 10.1006/plas.1999.1421. [DOI] [PubMed] [Google Scholar]
- Sessitsch A, Howieson JC, Perret X, Antoun H, Martinez-Romero E. Advances in Rhizobium Research. Critical Reviews in Plant Sciences. 2002;21:323–378. doi: 10.1080/0735-260291044278. [DOI] [Google Scholar]
- Vincent JM. A Manual for the Study of Root-Nodule Bacteria. IBP Handbook No. 15. England: Oxford; Blackwell scientific Publications; 1970. [Google Scholar]
- Hewitt EJ. Sand and Water Culture Methods Used in the Study of Plant Nutrition. Technical Communication No. 22. England: Farnham Royal; Commonwealth Agricultural Bureau; 1966. [Google Scholar]
- Zar JH. Biostatistical Analysis. 2. New Jersey: Prentice Hall; 1984. pp. 49–52. [Google Scholar]
- Kishinevsky B, Maoz A. ELISA identification of rhizobium strains by use of enzyme-labelled protein A. Current Microbiology. 1983;9:45–49. doi: 10.1007/BF01567132. [DOI] [Google Scholar]
- Evans J, Gregory A, Dobrowolski N, Morris SG, O'Connor GE, Wallace C. Nodulation of field-grown Pisum sativum and Vicia faba: Competitiveness of inoculant strains of Rhizobium leguminosarum bv. viciae determined by an indirect, competitive ELISA method. Soil Biology and Biochemistry. 1996;28:247–255. doi: 10.1016/0038-0717(95)00121-2. [DOI] [Google Scholar]
- Kock M. Ph.D Thesis. University of Pretoria, Pretoria, South Africa, Microbiology Department; 2003. Diveristy of root-nodulating bacteria associated with Cyclopia species. [Google Scholar]
- Sinclair MJ, Eaglesham ARJ. Intrinsic antibiotic resistance in relation to colony morphology in three populations of West African cowpea rhizobia. Soil Biology and Biochemistry. 1984;16:247–252. doi: 10.1016/0038-0717(84)90009-9. [DOI] [Google Scholar]
- Lucrecia M, Ramos G, Magalhaes FM, Boddey RM. Native and inoculated rhizobia isolated from field grown Phaseolus vulgaris: Effects of liming an acid soil on antibiotic resistance. Soil Biology and Biochemistry. 1987;19:179–185. doi: 10.1016/0038-0717(87)90078-2. [DOI] [Google Scholar]
- Davies J. Origins and evolution of antibiotic resistance. Microbiologia. 1996;12:9–16. [PubMed] [Google Scholar]
- Salyers AA, Shoemaker NB. Resistance gene transfer in anaerobes: New insights, new problems. Clinical Infectious Diseases. 1996;23:36–43. doi: 10.1093/clinids/23.supplement_1.s36. [DOI] [PubMed] [Google Scholar]
- Kishinevsky B, Bar-Joseph M. Rhizobium strain identification in Arachis hypogaea by enzyme-linked immunosorbent assay (ELISA) Canadian Journal of Microbiology. 1978;24:1537–1543. doi: 10.1139/m78-245. [DOI] [PubMed] [Google Scholar]