Abstract
The evaluation of Toxoplasma gondii isolates obtained from geographical environments other than Europe and North America revealed the existence of atypical strains that are not included in the three archetypal clonal lineages (lineages I, II, and III). GRA6 and GRA7 are polymorphic genes that have been used for the genotyping of Toxoplasma. The coding regions of GRA6 and GRA7 from 49 nonarchetypal strains were sequenced and compared with the sequences of type I, II, and III reference strains. Eighteen and 10 different amino acid sequences were found for GRA6 and GRA7, respectively. The polymorphisms found between the different sequences were analyzed, with the objective of defining peptides to be used for the serotyping of Toxoplasma infections. Two peptides specific for clonal lineages I and III (peptides GRA7I and GRA7III, respectively) were selected from the GRA7 locus. Three peptides specific for some atypical strains (peptides Am6, Af6, and Am7) were selected from both the GRA6 and the GRA7 loci. Serum samples from humans infected with Toxoplasma strains of known genotypes were serotyped with the selected peptides. Peptide GRA7III seems to be a good candidate for the serotyping of infections caused by type III strains. Peptide GRA7I had a very low sensitivity. Peptides Am6 and Af6 had low specificities, since they reacted with serum samples from patients infected with strains belonging to the three archetypal lineages. Although peptide Am7 was specific, it had low sensitivity.
The vast majority of Toxoplasma gondii isolates from human patients and domestic animals in Europe and North America belong to three archetypal clonal lineages, namely, types I, II, and III (1, 11, 30). However, nonarchetypal strains with atypical genotypes have recently been described in unusual hosts such as sea otters (10, 42, 43) and in tropical areas such as South America and Africa (2, 32, 39, 46, 54). Genotyping studies that distinguish different types of strains are important to gain knowledge of the biodiversity of the parasite in order to understand the molecular epidemiology of Toxoplasma and to highlight the correlation between the genotype of the parasite and the pathogenesis of human toxoplasmosis.
The dense granules (GRA) are parasitic organelles involved in cell invasion and in the intracellular survival of the parasite. GRA proteins are expressed by the three stages of T. gondii: the tachyzoite, bradyzoite (38), and sporozoite (55) stages. GRA6 is a GRA antigen of 32 kDa described for the first time by Lecordier et al. (38). In extracellular parasites, GRA6 exists in dense secretor granules mostly as soluble proteins. Like the other GRA proteins, GRA6 is involved in host cell invasion. GRA6 is a glycine-rich protein and behaves like an integral membrane protein within the parasitophorous vacuole (36, 41). GRA6 is considered a good marker of acute infection (27, 28, 52). However, the immune response to GRA6 is very heterogeneous (25).
GRA7 is a GRA antigen of 29 kDa (26, 31). Like GRA6, it is involved in host cell invasion. This protein is associated with the parasite membrane complex, the tubular elements of the intravacuolar network, and the parasitophorous vacuolar membrane. It migrates from the GRA to the parasitophorous vacuolar membrane through the intravacuolar network during host cell invasion (9). GRA7 is an antigen characteristic of the acute phase of the infection (27, 49, 51) and a target antigen in the intracerebral immune response during the chronic phase of infection (23, 45). These immunogenic properties make this antigen a good marker for serodiagnostic studies (5, 6, 7, 50).
GRA6 and GRA7 are polymorphic loci. The coding region of the GRA6 locus has been used as a marker for the genotyping of Toxoplasma (8, 15-20, 24, 33, 34, 40, 43, 47, 53). Sequencing of GRA6 detected a high degree of polymorphism (24, 57). Single nucleotide polymorphisms (SNP) in the region encoding GRA6 can be detected by methods based on PCR followed by restriction fragment length polymorphism (PCR-RFLP): digestion of the amplification products with a single endonuclease (MseI) can differentiate genotypes I, II, and III (24). Another method based on GRA6 polymorphisms is pyrosequencing (21). This technique allows the analysis of short DNA sequences and SNP. Two SNP located at positions 162 and 171 of the GRA6 gene allow the differentiation of types I, II, and III. GRA7 has been less explored as a marker for the genotyping of Toxoplasma. Preliminary results (I. Villena, unpublished data) showed that GRA7 allows the discrimination between genotypes I, II, and III and some atypical strains by PCR-RFLP.
Kong et al. (35) first proposed the use of the GRA6 and GRA7 proteins for serotyping. Basically, serotyping consists of a serological test with polymorphic peptides from Toxoplasma immunogens to detect strain-specific antibodies. Different peptides were proposed on the basis of the sequences of these antigens (35, 48, 53). However, the available peptides can differentiate only type II from non-type II infections. Infections due to nonarchetypal strains are misclassified as type II strains or type I or III strains, since GRA6 C-terminal peptides specific for type II and type I or III strains cross-react with serum samples from patients with infections caused by nonarchetypal strains (53).
We proposed to determine polymorphic regions of GRA6 and GRA7 with the objective of defining possible polymorphic peptides that could be used to distinguish type I from type III infections and infections due to atypical strains by serotyping. Some of the defined peptides were tested in order to evaluate their utility as serotyping markers.
MATERIALS AND METHODS
Human serum samples.
Thirty-one human serum samples related to 17 strains from the Toxoplasma bank of the Toxoplasma Biological Resource Center ToxoBS group were selected to evaluate the discriminative capacity of the selected peptides and, consequently, validate the serotyping approach (Table 1). The strains were previously typed by the analysis of five microsatellite markers (TUB2, TgM-A, W35, B17, B18) as described by Ajzenberg et al. (3); and the samples infected with these strains had the following genotypes or characteristics: (i) 4 serum samples corresponded to one type I infection, (ii) 8 serum samples corresponded to type II infections, (iii) 3 serum samples corresponded to type III infections, and (iv) 16 cases were associated with infections with 5 atypical strains (12 of which were caused by a single strain isolated during a Toxoplasma outbreak in Suriname) (12).
TABLE 1.
Serum samples from patients infected with Toxoplasma gondii strains
| Serum sample no. | Strain | Pathology | Geographical origin | Genotypea |
|---|---|---|---|---|
| 1 | LGE-01-1 | Congenital | France | II |
| 2 | LGE-00-2 | Congenital | France | II |
| 3 | LGE-2003-DUP | Congenital | France | II |
| 4 | LGE-2003-BAS | Congenital | France | II |
| 5 | LGE-00-1 | Congenital | France | II |
| 6 | LGE-2005-FRA | Congenital | France | II |
| 7 | LGE-2004-CAI | Congenital | France | II |
| 8 | LGE-07-LOR | Congenital | France | II |
| 9 | TOU-2004-FEU | Congenital | France | III |
| 10 | NED | Congenital | France | III |
| 11 | LGE-97-3 | Congenital | France | III |
| 12a | RH | Lymphadenopathy | France | I |
| 12b | RH | Lymphadenopathy | France | I |
| 12c | RH | Lymphadenopathy | France | I |
| 12d | RH | Lymphadenopathy | France | I |
| 13 | GUY-2004-TER | Congenital | Suriname | Atypical |
| 14 | GUY-2004-TER | Congenital | Suriname | Atypical |
| 15 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 16 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 17 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 18 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 19 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 20 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 21 | GUY-2004-TER | Multivisceral | Suriname | Atypical |
| 22 | GUY-2004-TER | Asymptomatic | Suriname | Atypical |
| 23 | GUY-2004-TER | Asymptomatic | Suriname | Atypical |
| 24 | GUY-2004-TER | Asymptomatic | Suriname | Atypical |
| 25 | GUY-2003-BAS | Multivisceral | French Guiana | Atypical |
| 26 | GUY-2004-AKO | Multivisceral | French Guiana | Atypical |
| 27 | GUY-2002-KOE | Multivisceral | French Guiana | Atypical |
| 28 | GUY-2002-MAT | Multivisceral | French Guiana | Atypical |
Genotype defined by the analysis of five microsatellite markers (TUB2, TgM-A, W35, B17, B18).
To establish cutoff values, 29 serum samples from France and 26 serum samples from Suriname and French Guiana negative for T. gondii were selected.
Strains.
Fifty-two strains of T. gondii were selected for the study of polymorphisms in GRA6 and GRA7 (Table 2). Strains RH, BEVERLEY, and NED were used as type I, II, and III reference strains, respectively. The remaining 49 T. gondii strains were selected for this study after a microsatellite analysis showed that they differed from the archetypal type I, II, and III strains. The microsatellite genotypes of these strains have been published previously (2, 53). Isolates TgCkCo24, GANGI, and ENVL-2002-MAC were considered variant of types I, II, and III, respectively, because of minor variations in their genotypes in comparison with those of the three archetypal clonal lineages. Isolates CRL-2004-MOT and IPB-2003-COG had different combinations of type I, II, or III alleles and were considered mixed type III-I and II-III genotypes, respectively. Isolates TgCkBr9, RMS-2003-DJO, and WIK exhibited the African no. 1 genotype and CCH-2004-NIA exhibited the African no. 2 genotype, which correspond to atypical genotypes previously described in immunocompromised African patients (4) The other isolates were atypical because of the presence of atypical alleles in their genotypes. Thirty-seven strains originated from South America or the Caribbean Islands, six originated from Africa, one originated from the Indian Ocean, five originated from Europe, and three originated from the United States.
TABLE 2.
GRA6 and GRA7 haplotypes for Toxoplasma gondii strains
| Strain | Host | Geographical origin | Genotypea | Haplotype
|
|
|---|---|---|---|---|---|
| GRA6 | GRA7 | ||||
| RH | Human | United States | I | 1 | 1 |
| TgCkCo24 | Chicken | Colombia | Variant of I | 1 | 1 |
| CRL-2004-MOT | Human | Africa (Cameroon) | III-I | 3 | 1 |
| CCH-2005-REN | Human | Caribbean (Guadeloupe) | Atypical | 7 | 1 |
| TgCkBr15 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 1 |
| TgCkBr5 | Chicken | Brazil (Sao Paulo) | Atypical | 11 | 1 |
| TgCkBr9 | Chicken | Brazil (Sao Paulo) | African no. 1 | 11 | 1 |
| TgCkBr23 | Chicken | Brazil (Sao Paulo) | Atypical | 11 | 1 |
| CCH-2004-NIA | Human | Africa (Senegal) | African no. 2 | 11 | 1 |
| PSP-2003-KOM | Human | Africa (Cameroon) | Atypical | 11 | 1 |
| RMS-2003-DJO | Human | Africa (Benin) | African no. 1 | 11 | 1 |
| WIK | Human | Africa | African no. 1 | 11 | 1 |
| GUY-2003-MEL | Human | French Guiana | Atypical | 13 | 1 |
| BEVERLEY | Rabbit | England | II | 2 | 2 |
| GANGI | Human | Africa | Variant of II | 2 | 2 |
| IPB-2003-COG | Human | Belgium | II-III | 3 | 2 |
| NED | Human | France | III | 3 | 3 |
| ENVL-2002-MAC | Monkey | Caribbean (Barbados) | Variant of III | 3 | 3 |
| TgCkBr21 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 3 |
| TgCkBr19 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 3 |
| TgCkBr10 | Chicken | Brazil (Sao Paulo) | Atypical | 2 | 4 |
| TgCkBr11 | Chicken | Brazil (Sao Paulo) | Atypical | 2 | 4 |
| TgCkBr25 | Chicken | Brazil (Sao Paulo) | Atypical | 2 | 4 |
| TgCkBr4 | Chicken | Brazil (Sao Paulo) | Atypical | 4 | 4 |
| RMS-1994-LEF | Human | France | Atypical | 4 | 4 |
| IPP-2002-BAT∴ | Human | French Guiana | Atypical | 7 | —b |
| P80∴ | Pig | United States | Atypical | 7 | — |
| TgCkBr1 | Chicken | Brazil (Sao Paulo) | Atypical | 7 | 4 |
| TgCkBr18 | Chicken | Brazil (Sao Paulo) | Atypical | 7 | 4 |
| MAR-2000-HOU | Human | Brazil | Atypical | 10 | 4 |
| TgCkBr24 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 4 |
| TgCkBr17 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 4 |
| TgCkBr16 | Chicken | Brazil (Sao Paulo) | Atypical | 18 | 4 |
| TgCkBr2 | Chicken | Brazil (Sao Paulo) | Atypical | 19 | 4 |
| TgCkBr3 | Chicken | Brazil (Sao Paulo) | Atypical | 16 | 5 |
| TgCkBr22 | Chicken | Brazil (Sao Paulo) | Atypical | 16 | 5 |
| GUY-2004-TER | Human | French Guiana | Atypical | 6 | 6 |
| TOU-2002-ALI | Human | Reunion Island | Atypical | 14 | 6 |
| TgCkBr6 | Chicken | Brazil (Sao Paulo) | Atypical | 17 | 6 |
| TgCkBr7 | Chicken | Brazil (Sao Paulo) | Atypical | 17 | 6 |
| TgCkAr23 | Chicken | Argentina | Atypical | 5 | 7 |
| IPP-2002-URB | Human | France | Atypical | 5 | 8 |
| GUY-2003-BAS | Human | French Guiana | Atypical | 6 | 9 |
| GUY-2004-LAB | Human | French Guiana | Atypical | 8 | 9 |
| GUY-2002-MAT | Human | French Guiana | Atypical | 8 | 9 |
| GUY-2001-DOS | Human | French Guiana | Atypical | 9 | 9 |
| GUY-2004-AKO | Human | French Guiana | Atypical | 10 | 9 |
| GUY-2002-KOE | Human | French Guiana | Atypical | 9 | 10 |
| TgCkBr14 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 11 |
| TgCkBr12 | Chicken | Brazil (Sao Paulo) | Atypical | 10 | 12 |
| P89 | Pig | United States | Atypical | 12 | 13 |
| GUY-2004-JAG | Jaguar | French Guiana | Atypical | 15 | 14 |
Genotype defined by the analysis of five microsatelites markers (TUB2, TgM-A, W35, B17, B18).
—, GRA7 was not sequenced for these strains.
DNA extraction.
DNA was extracted from the serum samples by using a QIAmp DNA minikit (Qiagen, Courtaboeuf, France), according to the manufacturer's protocol.
Sequencing.
The coding region of GRA6 gene was sequenced as described previously (53). The coding region of GRA7 was sequenced as described for the GRA6 gene, but with the following modifications: GRA7-specific primers 5′-ACCCTATATTGGGGCTTGCT-3′ (forward primer) and 5′-ACACTGTCCTCGAGCTCCTA-3′ (reverse primer) were used, and the annealing temperature was 63°C.
Sequence analysis.
The sequences were aligned by using Clustal W software. The nucleotide translation was performed with ExPASy (Expert Protein Analysis System) software (ExPASy Proteomics server). Haplotypes were defined with the program DnaSP (version 4.20.2) for Windows.
Antigen from whole-cell extracts.
Tachyzoites from the RH strain were obtained from the peritoneal cavities of mice 48 h after intraperitoneal infection. The tachyzoites were washed three times in phosphate-buffered saline (PBS), frozen at −80°C for 1 h 30 min, boiled at 100°C for 5 min, and sonicated three times (2 min per cycle) to induce tachyzoite lysis. The lysate was stored at −20°C.
Peptides.
Peptides were selected from the GRA6 and GRA7 antigens on the basis of the following criteria: the presence of strain-specific polymorphisms and the presence of possible immunogenic regions. Strain-specific polymorphisms were defined by GRA6 and GRA7 sequencing. Immunogenic epitopes were defined according to hydrophobicity (22), hydrophilicity (29), and antigenicity (56) characteristics (ExPASy Proteomics server; www.expasy.org/tools/protscale.html).
Six peptides were synthesized by Millegen, France. The amino acid sequences selected for the GRA7 peptides were synthesized as three repeats of the selected sequence in one peptide because the region selected had less than 20 amino acids (aa). The peptide control was described previously (53).
Enzyme-linked immunosorbent assay (ELISA) protocol with synthetic peptides.
Immobilizer amino plates (Nunc, Denmark) were coated with each peptide, which had been diluted to 10 μg/ml in 0.05 M carbonate/bicarbonate buffer (pH 9.6), by incubation overnight at 4°C. The wells were blocked with PBS-3% bovine serum albumin (BSA) for 1 h at 37°C in a moist atmosphere and were then washed three times with PBS-0.3% Tween 20. The sera were diluted to 1/50 in PBS-0.3% Tween 20-3% BSA and incubated for 2 h at 37°C in a moist atmosphere. The wells were washed three times with PBS-0.3% Tween 20. Anti-human immunoglobulin G alkaline phosphatase conjugate (Pierce) was diluted at 1/5,000 in PBS-3% BSA-0.3% Tween 20 and incubated for 1 h at 37°C in a moist atmosphere. The wells were washed three times with PBS-0.3% Tween 20 and developed with p-nitrophenyl phosphate for 15 min at 37°C. The absorbance was measured at 415 nm.
The optical density (OD) index was calculated by subtracting the OD of the peptide control from the OD of each peptide. The cutoff was set as the mean absorbance reading for the negative sera plus 2 standard deviations (SDs).
ELISA protocol with antigen from whole-cell extracts.
The ELISA protocol described above for the peptides was used for the ELISA protocol with antigen from whole-cell extracts, but the following changes were made. The antigen from the whole-cell extracts was diluted to 5 μg/ml. The wells were blocked with 1% gelatin solution in PBS. The serum samples and conjugate were diluted in PBS-0.3% Tween 20.
Nucleotide sequence accession numbers.
The nucleotide sequence data reported in this paper are available in the GenBank, EMBL, and DDBJ databases under accession numbers EF512265 (TgCkAr23), EF512264 (TgCkBr2), EF512263 (TgCkBr16), EF512262 (TgCkBr23), EF512261 (TgCkBr9), EF512260 (TgCkBr5), EF512259 (TgCkBr12), EF512258 (TgCkBr14), EF512257 (TgCkBr15), EF512256 (TgCkBr17), EF512255 (TgCkBr19), EF512254 (TgCkBr21), EF512253 (TgCkBr24), EF512252 (TgCkBr18), EF512251 (TgCkBr1), EF512250 (TgCkBr7), EF512249 (TgCkBr6), EF512248 (TgCkBr22), EF512247 (TgCkBr3), EF512246 (TgCkBr4), EF512245 (TgCkBr25), EF512244 (TgCkBr11), EF512243 (TgCkBr10), EF512242 (TOU-2002-ALI), EF512241 (GUY-2003-MEL), EF512240 (P89), EF512239 (WIK), EF512238 (RMS-2003-DJO), EF512237 (PSP-2003-KOM), EF512236 (CCH-2004-NIA), EF512235 (MAR-2000-HOU), EF512234 (GUY-2004-AKO), EF512233 (GUY-2002-KOE), EF512232 (GUY-2001-DOS), EF512231 (GUY-2002-MAT), EF512230 (GUY-2004-LAB), EF512229 (P80), EF512228 (IPP-2002-BAT), EF512227 (CCH-2005-REN), EF512226 (GUY-2004-TER), EF512225 (GUY-2003-BAS), EF512224 (IPP-2002-URB), EF512223 (IPB-2003-COG), EF512222 (ENVL-2002-MAC), EF512221 (CRL-2004-MOT), EF512220 (RMS-1994-LEF), EF512219 (GANGI), EF512218 (TgCkCo24), DQ187387 (GUY-2004-JAG), AF239286 (NED), AF239284 (BEVERLEY), and AF239283 (RH) for GRA6 and accession numbers EF639859 (RMS-2003-DJO), DQ473315 (GUY-2004-LAB), DQ459455 (NED), DQ459453 (MAR-2000-HOU), DQ459452 (RMS-1994-LEF), DQ459451 (GUY-2001-DOS), DQ459443 (RH), EU157183 (GUY-2004-JAG), EU157182 (TgCkBr2), EU157181 (TOU-2002-ALI), EU157180 (TgCkBr16), EU157179 (GUY-2003-MEL), EU157178 (P89), EU157177 (WIK), EU157176 (PSP-2003-KOM), EU157175 (CCH-2004-NIA), EU157174 (TgCkBr23), EU157173 (TgCkBr9), EU157172 (TgCkBr5), EU157171 (TgCkBr12), EU157170 (TgCkBr14), EU157169 (TgCkBr15), EU157168 (TgCkBr17), EU157167 (TgCkBr19), EU157166 (TgCkBr21), EU157165 (TgCkBr24), EU157164 (GUY-2004-AKO), EU157163 (GUY-2002-KOE), EU157162 (GUY-2002-MAT), EU157161 (CCH-2005-REN), EU157160 (TgCkBr18), EU157159 (TgCkBr1), EU157158 (GUY-2004-TER), EU157157 (GUY-2003-BAS), EU157156 (IPP-2002-URB), EU157155 (TgCkAr23), EU157154 (TgCkBr7), EU157153 (TgCkBr6), EU157152 (TgCkBr22), EU157151 (TgCkBr3), EU157150 (IPB-2003-COG), EU157149 (ENVL-2002-MAC), EU157148 (CRL-2004-MOT), EU157147 (TgCkBr4), EU157146 (GANGI), EU157145 (TgCkBr25), EU157144 (TgCkBr11), EU157143 (TgCkBr10), EU157142 (TgCkCo24), and EU157141 (BEVERLEY) for GRA7.
RESULTS
GRA6 amino acid sequences.
For the 19 haplotypes defined for GRA6, 18 different amino acid sequences were found for the 52 strains (Tables 2 and 3). The most polymorphic region of the GRA6 protein sequence was the C-terminal region. From this region (aa 198 to aa 230), 15 different peptides that allowed the differentiation of Toxoplasma strains could be selected. Peptides corresponding to haplotypes 1 and 3 (type I and type III strains, respectively) differed from each other by 3 aa substitutions (at positions 198, 213, and 219) and were clearly different from the peptide corresponding to haplotype 2. This haplotype is characterized by deletions from aa 204 to 208 and at position 213 and differed from haplotypes 1 and 3 by 4 and 5 aa substitutions from aa 219 to 230, respectively. The remaining 12 peptides were found in most of the nonarchetypal strains circulating in South America or Africa. They differed from the type I and III peptides by 1 to 3 aa substitutions. A peptide corresponding to haplotypes 8 and 9 (G-GYRGR-G-DRRA-ER-V-Y; hyphens indicate nonpolymorphic amino acids throughout the text) was found in 4 of 10 strains from French Guiana. A peptide corresponding to haplotypes 10 and 18 (G-GYRGR-G-DRRA-EH-V-Y) was found in 9 of 23 Brazilian strains. These two peptides differed from each other by a single amino acid (arginine instead of histidine at position 224) and from the type I and III peptides by 2 to 4 aa substitutions. A peptide corresponding to haplotype 11 (G-GYRGR-G-DGRA-ER-V-Y) was found in four of the six African strains and three strains from Brazil. Peptides R-GYGGR-G-DRRA-EH-V-Y for haplotype 16, G-GYRGR-G-DRRA-ES-V-Y for haplotype 17, G-GYGGR-DRRP-ER-E-Y for haplotype 19, and G-GDGGR-G-DRRP-ER-E-Y for haplotype 15 were exclusive to South American strains. The regions of the GRA6 protein between positions 47 and 50, positions 92 and 105, and positions 180 and 190 also showed polymorphisms that could be used to differentiate some atypical strains. However, in these regions, most of the polymorphisms present in atypical strains were shared by clonal lineages I, II, and III.
TABLE 3.
GRA6 polymorphic amino acids
| Haplotype(s) | Variable amino acid(s) at the following positionsa:
|
|||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 7 | 12 | 17 | 25 | 42 | 47 | 49 | 50 | 91 | 92 | 94 | 105 | 180 | 185 | 186 | 190 | 198 | 204-208 | 213 | 216-219 | 223 | 224 | 227 | 230 | |
| 1 (type I) | H | R | V | V | V | G | V | K | - | T | E | V | Q | D | G | D | G | GYGGR | A | DRRP | E | R | V | Y |
| 2 (type II) | Y | . | L | . | . | . | . | R | - | . | D | A | . | G | . | . | . | ----- | - | .... | G | S | E | F |
| 4 | Y | . | L | . | . | . | . | . | - | . | D | A | . | G | . | . | . | ..... | - | G.G. | G | S | E | F |
| 19 | Y | . | L | . | . | . | F | R | - | . | D | A | . | . | . | G | . | ..... | - | .... | . | . | E | . |
| 15 | Y | . | L | I | F | . | . | R | D | A | D | A | . | G | . | . | . | .D... | G | .... | . | . | E | . |
| 3 (type III) | . | . | L | . | . | D | . | . | - | . | D | . | P | . | . | . | R | ..... | G | ...A | . | . | . | . |
| 16 | . | . | L | . | . | . | . | . | - | . | D | . | P | . | . | . | R | ..... | G | ...A | . | H | . | . |
| 10, 18 | . | . | L | . | . | . | . | . | - | . | D | . | P | . | D | . | . | ..R.. | G | ...A | . | H | . | . |
| 17 | . | . | L | . | . | D | . | . | - | . | D | . | P | . | . | . | . | ..R.. | G | ...A | . | S | . | . |
| 5 | . | . | L | . | . | D | . | . | - | . | D | . | P | . | . | . | . | ..R.. | G | ...A | G | S | . | . |
| 6 | . | . | L | . | . | D | . | . | - | A | D | . | P | . | D | . | . | ..R.. | G | ...A | G | S | . | . |
| 7 | . | . | L | . | . | D | . | . | - | A | D | . | P | . | D | . | . | ----- | G | ...A | G | S | . | . |
| 14 | . | . | L | . | . | D | . | . | - | A | D | . | P | . | D | . | . | ..R.. | G | .... | G | S | . | . |
| 8 | . | H | L | . | . | . | . | . | - | . | D | . | P | . | D | . | . | ..R.. | G | ...A | . | . | . | . |
| 9 | . | . | L | . | . | . | . | . | - | . | D | . | P | . | D | . | . | ..R.. | G | ...A | . | . | . | . |
| 11 | . | . | L | . | . | . | . | Q | - | . | D | . | P | . | . | . | . | ..R.. | G | .G.A | . | . | . | . |
| 12 | . | . | L | . | . | . | . | Q | - | . | D | . | P | . | . | . | . | ..R.. | - | .G.A | . | . | . | . |
| 13 | . | . | L | . | . | . | . | . | - | . | D | . | P | . | D | . | . | ..R.. | G | .G.A | . | . | . | . |
Periods indicate that the amino acid is identical to that in haplotype 1, and hyphens indicate a deletion. Boldface indicates polymorphic amino acids for peptides Am6 (haplotype 7) and Af6 (haplotype 11).
On the basis of the high degree of polymorphism and the predicted immunogenicity (data not shown), two peptides were selected from the GRA6 C-terminal region (aa 198 to aa 230): one specific for South American strains (peptide Am6) and one specific for African strains (peptide Af6) (see Table 5). These two GRA6 peptides were selected from the most representative haplotypes found for strains from Africa and South America. Haplotype 11 was shared by four of six African strains, which justifies the selection of peptide Af6. Peptide Am6 was selected from haplotype 7. For the South American strains, the most representative haplotypes were haplotypes 7 and 10. However, the number of polymorphic amino acids between haplotypes 7 and 11 was higher than the number of polymorphic amino acids between haplotypes 10 and 11. Consequently, the better differentiation of atypical strains was expected with a peptide derived from haplotype 7 than with one derived from haplotype 10.
TABLE 5.
Amino acid sequences of GRA6 and GRA7 synthetic peptides
| Peptide | Locus | Amino acid positions | Strain type | Amino acid sequencea |
|---|---|---|---|---|
| GRA7I | GRA7 | 220-236 | I | LEQEVPESGEDGEDARQLEQEVPESGEDGEDARQLEQEVPESGEDGEDARQ |
| GRA7III | GRA7 | 220-236 | III | PEHEVPESGEDREDARQPEHEVPESGEDREDARQPEHEVPESGEDREDARQ |
| Am6 | GRA6 | 198-230 | Atypical (haplotype 7) | GGNEGRGEGGGEDDRRALHPGSVNVFDY |
| Af6 | GRA6 | 198-230 | Atypical (haplotype 11) | GGNEGRGYRGRGEGGGEDDGRALHPERVNVFDY |
| Am7 | GRA7 | 170-182 | Atypical (haplotypes 6, 9, and 10) | AGEEPLTTSQNVNAGEEPLTTSQNVNAGEEPLTTSQNVN |
Polymorphic amino acids are underlined.
GRA7 amino acid sequences.
Fourteen haplotypes with 10 different amino acid sequences were defined for GRA7 (Tables 2 and 4). The most polymorphic region in the GRA7 protein sequence comprised aa 170 to 182. Six different peptides from this region could be selected to differentiate strains. Peptides R-D-T-G-G and T-E-T-G-S were exclusive to strains that had the same GRA7 sequence as strains RH (haplotype 1) and BEVERLEY (haplotype 2), respectively. Most African strains (five of six) shared the peptide R-D-T-G-G characteristic of type I. Peptide T-D-S-G-S was shared by strains with the same GRA7 sequence as haplotypes 3, 4, 8, and 13. This peptide was found in 15 of the 23 Brazilian strains. Peptides A-E-T-S-N, A-D-T-S-N, and T-E-T-G-G were exclusive to nonarchetypal strains. Peptide A-E-T-S-N, characteristic of haplotype 6, 9, and 10 strains, was found in 10 strains. Except for strain TOU-2002-ALI, which was isolated on Reunion Island in the Indian Ocean, the remaining nine strains were atypical and were from South America. This peptide was found in 7 of 10 strains from French Guiana. It differed from peptide A-D-T-S-N, found in 2 Brazilian strains (strains TgCkBr3 and TgCkBr22), by a single amino acid. The regions between positions 106 and 120, 199 and 222, and 220 and 231 were less polymorphic. In these regions, most of the atypical strains shared the polymorphisms with type I, II, and III strains. The region between positions 106 and 120 allowed the differentiation of 12 of the 23 Brazilian strains. Only two different peptides (M-LT-L-Q and L-IK-P-H) could be selected from the region between aa 199 and 222. These peptides differed by 5 aa and allowed the differentiation of type I and II strains from type III strains, respectively. Peptide M-LT-L-Q was found in the 6 African strains and 9 of 10 strains from French Guiana. Three different peptides (L-Q-E-G, L-Q-K-G, and P-H-E-R) that allowed the differentiation of type I, II, and III strains were found between aa 220 and 231.
TABLE 4.
GRA7 polymorphic amino acids
| Haplotype(s) | Variable amino acid(s) at the following positionsa:
|
||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 8 | 38 | 106 | 115 | 120 | 160 | 161 | 167 | 170 | 172 | 176 | 178 | 182 | 185 | 194 | 197 | 199 | 201 | 202 | 220 | 222 | 229 | 231 | |
| 1 (type I) | S | I | R | N | H | L | V | E | R | D | T | G | G | L | A | L | M | L | T | L | Q | E | G |
| 14 | F | . | . | . | . | I | L | Q | T | E | . | . | . | . | . | . | . | . | . | . | . | . | . |
| 2 (type II) | F | . | . | . | . | I | L | Q | T | E | . | . | S | . | . | . | . | . | . | . | . | K | . |
| 3 (type III), 11 | F | . | G | H | N | I | L | Q | T | . | S | . | S | I | V | . | L | I | K | P | H | . | R |
| 4, 7, 12 | F | . | G | H | . | I | L | Q | T | . | S | . | S | I | V | . | L | I | K | P | H | . | R |
| 13 | . | . | G | H | N | I | L | Q | T | . | S | . | S | I | V | . | L | I | K | P | H | . | R |
| 8 | F | . | G | H | . | I | L | Q | T | . | S | . | S | I | V | F | L | I | K | P | H | . | R |
| 5 | F | . | . | . | . | I | L | Q | A | . | . | S | N | . | . | . | . | . | . | . | . | . | . |
| 9, 10 | F | T | . | . | . | I | L | Q | A | E | . | S | N | . | . | . | . | . | . | . | . | . | . |
| 6 | F | . | . | . | . | V | L | Q | A | E | . | S | N | . | . | . | . | . | . | . | . | . | . |
Periods indicate that the amino acid is identical to that in haplotype 1. Boldface indicates polymorphic amino acids for peptides GRA7I (haplotype 1), GRA7III (haplotypes 3 and 11), and Am7 (haplotypes 6, 9, and 10).
Three peptides were selected from GRA7. Two of them were from the C-terminal region, including the last five nonpolymorphic amino acids (aa 220 and 236), which had polymorphisms characteristic of type I (GRA7I) and type III (GRA7III) strains. Another peptide specific for the atypical strains belonging to haplotypes 6, 9, and 10 (peptide Am7) was selected from the region between aa 170 and 182 (Table 5). Both regions were predicted to be immunogenic (data not shown).
Serotyping.
Thirty-one serum samples from 28 patients infected with 17 different type I, II, and III and atypical strains were used to study the usefulness of the peptides described in Table 5 for the serotyping of T. gondii. Cutoff values were defined for each peptide according to the geographical origin of the strains. Patients who were infected with type I, II, and III strains and who originated from France were considered positive for GRA7I, GRA7III, Am6, Af6, Am7, and the antigen from the whole-cell extract when the OD indexes were equal to or greater than the cutoffs established for Europe (0.114, 0.126, 0.180, 0.096, 0.188, and 0.315, respectively). Patients who were infected with atypical strains and who originated from Suriname and French Guiana were considered positive for GRA7I, GRA7III, Am6, Af6, Am7, and the antigen from the whole-cell extract when the OD indexes were equal to or greater than the cutoffs established for South America (0.124, 0.120, 0.252, 0.079, 0.270, and 0.306, respectively). Cutoff values are indicated by the horizontal lines in Fig. 1 to 6.
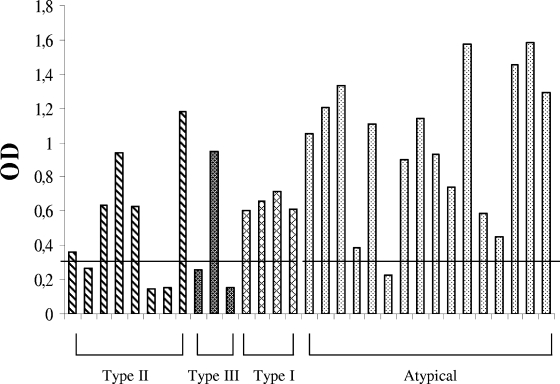
FIG. 1.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against peptide GRA7I. Horizontal lines represent the cutoff values (the cutoff values are different for sera for Europe and South America).
FIG. 6.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against the antigen from the whole-cell extract. Horizontal lines represent the cutoff values.
Peptide GRA7I did not recognize any serum sample from the patient with lymphadenopathy due to an accidental infection in the laboratory with strain RH (type I). However, this peptide reacted weakly with one serum sample from a patient infected with a type II strain (serum sample 6) and with one serum sample from a patient infected with an atypical strain (serum sample 23) (Fig. 1). Patient 23 was infected with an atypical strain (strain GUY-2004-TER) belonging to haplotype 6 for GRA7 which had the same polymorphisms in the GRA7 C-terminal region as type I strains. Peptide GRA7III was recognized by two of three serum samples from patients infected with type III strains. Cross-reactivity was observed between this peptide and the serum sample from one patient infected with a type II strain (serum sample 6) and with serum samples from three patients infected with atypical strains (serum samples 21, 23, and 27) (Fig. 2).
FIG. 2.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against peptide GRA7III. Horizontal lines represent the cutoff values.
The sequence of peptide Am6 differed from the sequence in atypical strains responsible for the infections studied by 5 and 7 aa, while the sequence of peptide Af6 differed by 1 to 3 aa. Peptides Am6 and Af6 reacted with serum samples from 8 and 11 of 16 patients infected with atypical strains, respectively. These two peptides also reacted with some serum samples from patients infected with archetypal strains (Fig. 3 and 4). The sequence of peptide Am6 differed by 9 aa from the sequence in type I strains, by 4 aa from the sequence in type II strains, and by 8 aa from the sequence in type III strains. The sequence of peptide Af6 differed by 4 aa from the sequence in type I strains, by 12 aa from the sequence in type II strains, and by 3 aa from the sequence in type III strains.
FIG. 3.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against peptide Am6. Horizontal lines represent the cutoff values (the cutoff values are different for sera from Europe and South America).
FIG. 4.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against peptide Af6. Horizontal lines represent the cutoff values.
Peptide Am7 had a lower sensitivity, being recognized by only three serum samples from 16 patients infected with atypical strains. However, this peptide did not react with any of the serum samples from patients infected with type I, II, or III strains (Fig. 5).
FIG. 5.
Pattern of reactivity of serum samples from patients infected with type I, II, and III and atypical strains when they were tested against peptide Am7. Horizontal lines represent the cutoff values (the cutoff values are different for sera from Europe and South America).
Not all serum samples had OD indexes greater than the respective cutoff when they were tested against the antigen from the whole-cell extract (Fig. 6). Six of 31 of the serum samples studied had OD indexes bellow the cutoff. However, some of those serum samples were positive when they were tested against the peptides studied. Serum samples 2, 7, and 9 were positive when they were tested against peptides Am6 and Af6, and serum samples 6 and 11 were positive when they were tested against peptide GRA7III.
DISCUSSION
The coding region for GRA6 has already been described as being highly polymorphic (24). Fazaeli et al. (24) identified nine different alleles in 30 strains by sequencing. Another GRA6 allele was found in type x, a genotype described in Californian sea otters (10, 42, 43). More recently, Zakimi et al. (57) found nine new sequences among 29 isolates from Japanese pigs. Most studies with GRA6 as a genetic marker have been conducted by PCR-RFLP (8, 14-20, 24, 43, 57). The GRA6 PCR-RFLP method is able to differentiate only three different groups among the strains and misses all atypical alleles. For example, in a study with sea otters, Miller et al. (43) could not distinguish type x by GRA6 PCR-RFLP because the SNP specific for type II was shared by type x strains. Similarly, Zakimi et al. (57) found by GRA6 PCR-RFLP only the three classical genotypes in pigs from Japan. The findings of our sequence analysis are in agreement with those of previous studies and confirm the high degree of polymorphism of the GRA6 gene. Six GRA6 sequences (haplotypes 1, 2, 3, 4, 5, and 11) found in our strains have also been described by Fazaeli et al. (24). The GRA6 sequence from strain GUY-2006-JAG is identical to the GRA6 sequence of the type x strain found in 60% of sea otters from California (13, 43). The nine new sequences found in pigs from Japan (57) were not found in our strains. Few studies that have genotyped GRA7 have been conducted (I. Villena et al., unpublished data).
The polymorphism of the GRA6 (35, 44, 48) and the GRA7 (35) antigens was used to develop serotyping methods. The objective of serotyping is to use an allele-specific peptide that would allow the identification of the strain type. The peptides described in two of these studies (35, 44) distinguish only type II infections from non-type II infections. They can be interesting in the context of a clonal population structure, like that observed in Europe and United States, but new data show that the population structure of Toxoplasma is much more complex and that strains circulating in other parts of the world diverge from the archetypal types (2, 32, 39, 46). According to the findings of the study performed by Lecordier et al. (37), only the N-terminal hydrophilic region of GRA6 is recognized by Toxoplasma-positive human sera. ELISA of serum samples for determination of their reactivity with the GRA6 C terminus showed a sensitivity of 10%, while ELISA of serum samples for determination of their reactivity with the GRA6 N terminus showed a sensitivity of 96%. On the basis of those findings, the authors suggested that a major B-cell epitope(s) is carried by the N-terminal peptide portion of the GRA6 antigen. However, the C-terminal region presents more polymorphisms than the N-terminal region. The results obtained by Kong et al. (35) showed that the C terminus is also probably immunogenic, since the peptides selected from that region reacted extremely well with human sera. The immunogenic epitopes predicted according to hydrophobicity (22), hydrophilicity (29), and antigenicity (56) characteristics with the ExPASy Proteomics server (www.expasy.org/tools/protscale.html) (data not shown) support this idea, since the C-terminal region between aa 198 and 226 was selected as an immunogenic epitope for GRA6.
The GRA6 peptides described by Kong et al. (35) were selected from this region and comprised the last four polymorphic sites (223, 224, 227, and 230). If we consider only the peptide sequences chosen in the study of Kong et al. (35), six different peptides can be described in our sample. Two peptides could differentiate type II (polymorphic amino acids GS-E-F) from type I and III (ER-V-Y), as described by Kong et al. (35) and Sousa et al. (53). In our study, we found four other peptides that differed from the peptide characterizing types I and III. Three peptides differed by a single amino acid (underlined) at position 224 (EH-V-Y and ES-V-Y instead of ER-V-Y) or at position 227 (ER-E-Y instead of ER-V-Y). The last peptide (GS-V-Y) is a mixture of amino acid sequences that characterize type II strains and type I and III strains. If we extend the selected peptides to aa 198 to 230, 3 aa substitutions are found between type I and type III strains; however, the large number of identical amino acids can increase the cross-reaction between these two peptides. Similarly, most nonarchetypal strains found in South America and Africa are characterized by peptides that differ from those in type I or III by a maximum of three nonconsecutive amino acids with a large number of identical amino acids, which may limit the use of these peptides for serotyping.
The most polymorphic region in the GRA7 sequences was located between residues 170 and 182. Peptide A-E-T-S-N, characteristic of haplotypes 6, 9, and 10, may be useful for the differentiation of strains from French Guiana. This region was not selected by Kong et al. (35). The peptides selected by Kong et al. (35) were located in the C-terminal region of GRA7 and comprised two polymorphic amino acids (aa 229 and 231). We propose that the size of this peptide be increased by including two other polymorphic amino acids (aa 220 and 222). In this region, only three different peptides that distinguish types I, II, and III can be selected. Atypical strains have the same polymorphism as type I and III strains. Peptides selected from this region may be good candidates for the differentiation of type I from type III strains (three different amino acids) as well as type II from type III strains (four different amino acids).
Five peptides were synthesized and tested in order to evaluate their usefulness in serotyping assays. Previous serotyping studies distinguished only serotype II (35, 44, 48). The peptides described by Sousa et al. (53) were derived from the archetypal strains and differentiated only strains of GRA6 type II from strains of GRA6 non-type II. Serotyping based on the peptides GRA6II and GRA6I/III showed a strong correlation between the GRA6 serotype and the microsatellite marker genotype for infections due to archetypal strains (53). It was therefore not possible to distinguish type I strains from type III strains and strains with atypical genotypes. The study reported here was a first attempt to select and test peptides capable of distinguishing type I strains from type III and atypical strains. These results, together with previous results obtained with GRA6II and GRA6I/III (53), suggest that, in the context of Europe and North America, it is possible to distinguish type II infections (reactivity against GRA6II only) from type III infections (reactivity against GRA6I/III and GRA7III). Type I infections seem to be more difficult to identify. One peptide derived from GRA6 with polymorphisms specific for GRA6 type I strains correctly recognized the serum samples from patients infected with type I strains (53). However, the type I peptide derived from GRA7 had sensitivity problems, since only one serum sample from patients infected with strains with the polymorphisms characteristic of this peptide recognized the peptides. However, more samples from natural type I infections need to be studied, since the patient with the type I infection was not infected by the natural oral route but was infected after an accidental needle inoculation. This may have induced a different immunological response. The cross-reactivity observed between peptides GRA7I and GRA7III with serum samples with other haplotypes was similarly described for peptides GRA6II and GRA6I/III (53). Peptides Am6 and Af6 had a high rate of reactivity, cross-reacting with serum samples from patients infected with strains that do not share the polymorphisms of these two peptides. The high rate of reactivity of these two peptides confirms that the C-terminal region of GRA6 is an immunogenic epitope, as predicted. The increase in the size of the peptide sequence seems to diminish the specificity of the peptide, and this is probably associated with an increase in the number of nonpolymorphic amino acids. Although peptide Am7 was specifically recognized by nonarchetypal strains, it had a low sensitivity. With strains from South America and Africa, the cross-reactivity with all the peptides that characterize archetypal strains and a lack of specificity of the atypical peptides designed here would not result in a valuable serotyping method.
Although serotyping is a promising method for strain typing, it presents sensitivity and specificity problems. The established cutoff values were defined by 2 SDs. The use of only 1 SD would diminish the specificity of the peptides, while 3 SDs give results similar to the chosen value (data not shown). The immune response against GRA6 and GRA7 is very heterogeneous since different patients infected with the same strain may present with different profiles of reactivity against the same peptides. GRA6 and GRA7 are antigens characteristic of recent infections (27, 28, 49, 51, 52). The use of single peptides derived from these two antigens may not allow serotyping of serum samples from patients with chronic infections. Besides, serotyping with peptides derived from only two antigens raises the same problems as genotyping of a single locus, leading to the misidentification of atypical strains. In order to achieve the correct differentiation between the different serotypes, the use of other polymorphic antigens must be explored. For each lineage, a large pool of polymorphic peptides from different antigens must be used to avoid the problems of serotyping of a single locus. Serotypes should be defined by the serological profiles against different peptides.
Our results indicate that other loci should be screened with the objective of selecting different peptides that, when they are combined with peptides that have already been defined, will be useful for the detection of a different humoral response in infected patients. The diversity of the immune response will be one of the main limits to the identification of clonal lineages by serotyping. The development of serotyping will be of special interest for epidemiological studies in order to determine if there is strain selection by the host, the geographical distributions of the different serotypes, and the hypothetical association between serotype and human disease.
Acknowledgments
We thank Marie-France Cesbron and Olivier Bastien for helpful discussions. The DNAs of strains from Brazilian chickens were kindly provided by T. Lehmann and J. P. Dubey. We thank Magalie Demar, Bernard Carme, and Rachida Boukhari for serum samples from French Guiana.
Footnotes
Published ahead of print on 3 June 2009.
REFERENCES
- 1.Ajzenberg, D., A. L. Bañuls, and M. L. Dardé. 2002. Microsatellite analysis of Toxoplasma gondii shows considerable polymorphism structured into two main clonal groups. Int. J. Parasitol. 3227-38. [DOI] [PubMed] [Google Scholar]
- 2.Ajzenberg, D., A. L. Bañuls, C. Su, A. Dumètre, M. Demar, B. Carme, and M. L. Dardé. 2004. Genetic diversity, clonality and sexuality in Toxoplasma gondii. Int. J. Parasitol. 341185-1196. [DOI] [PubMed] [Google Scholar]
- 3.Ajzenberg, D., A. Dumètre, and M. L. Dardé. 2005. Multiplex PCR for typing strains of Toxoplasma gondii. J. Clin. Microbiol. 431940-1943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ajzenberg, D., H. Yera, P. Marty, L. Paris, F. Dalle, J. Menotti, D. Aubert, J. Franck, M. H. Bessières, D. Quinio, P. Pelloux, L. Delhaes, N. Desbois, P. Thulliez, F. Robert-Gangneux, C. Kauffmann-Lacroix, S. Pujol, M. Rabodonirina, M. E. Bougnoux, B. Cuisenier, C. Duhamel, T. H. Duong, D. Filisetti, P. Flori, F. Gay-Andrieu, F. Pratlong, G. Nevez, A. Totet, C. Carme, H. Bonnabau, M. L. Dardé, and I. Villena. 2009. Genotype of 88 Toxoplasma gondii isolates associated with toxoplasmosis in immunocompromised patients, and correlation with clinical findings. J. Infect. Dis. 1991155-1167. [DOI] [PubMed] [Google Scholar]
- 5.Altcheh, J., N. S. Diaz, C. M. Pepe, V. Martin, M. Nigro, H. Freilij, and S. O. Angel. 2006. Kinetic analysis of the humoral immune response against 3 Toxoplasma gondii-recombinant proteins in infants with suspected congenital toxoplasmosis. Diagn. Microbiol. Infect. Dis. 56161-165. [DOI] [PubMed] [Google Scholar]
- 6.Aubert, D., G. T. Maine, I. Villena, J. C. Hunt, L. Howard, M. Sheu, S. Brojanac, L. E. Chovan, S. F. Nowlan, and J. M. Pinon. 2000. Recombinant antigens to detect Toxoplasma gondii-specific immunoglobulin G and immunoglobulin M in human sera by enzyme immunoassay. J. Clin. Microbiol. 381144-1150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Beghetto, E., A. Spadoni, L. Bruno, W. Buffolano, and N. Gargano. 2006. Chimeric antigens of Toxoplasma gondii: toward standardization of toxoplasmosis serodiagnosis using recombinant products. J. Clin. Microbiol. 442133-2140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Belfort-Neto, R., V. Nussenblatt, L. Rizzo, C. Mucciolo, C. Silveira, R. Nussenblatt, A. Khan, L. D. Sibley, and R. Belfort, Jr. 2007. High prevalence of unusual genotypes of Toxoplasma gondii infection in pork meat samples from Erechim, southern Brazil. An. Acad. Bras. Cienc. 79111-114. [DOI] [PubMed] [Google Scholar]
- 9.Bonhomme, A., G. T. Maine, A. Beorchia, H. Burlet, D. Aubert, I. Villena, J. Hunt, L. Chovan, L. Howard, S. Brojanac, M. Sheu, J. Tyner, M. Pluot, and J. M. Pinon. 1998. Quantitative immunolocalization of a P29 protein (GRA7), a new antigen of Toxoplasma gondii. J. Histochem. Cytochem. 461411-1422. [DOI] [PubMed] [Google Scholar]
- 10.Conrad, P. A., M. A. Miller, C. Kreuder, E. R. James, J. Mazet, H. Dabritz, D. A. Jessup, F. Gulland, and M. E. Grigg. 2005. Transmission of Toxoplasma: clues from the study of sea otters as sentinels of Toxoplasma gondii flow into the marine environment. Int. J. Parasitol. 351155-1168. [DOI] [PubMed] [Google Scholar]
- 11.Dardé, M. L., B. Bouteille, and M. Pestre-Alexandre. 1992. Isoenzyme analysis of 35 Toxoplasma gondii isolates and the biological and epidemiological implications. J. Parasitol. 78786-794. [PubMed] [Google Scholar]
- 12.Demar, M., D. Ajzenberg, D. Maubon, F. Djossou, D. Panchoe, W. Punwasi, N. Valery, C. Peneau, J. L. Daigre, C. Aznar, B. Cottrelle, L. Terzan, M. L. Dardé, and B. Carme. 2007. Fatal outbreak of human toxoplasmosis along the Maroni River: epidemiological, clinical and parasitological aspects. Clin. Infect. Dis. 4588-95. [DOI] [PubMed] [Google Scholar]
- 13.Demar, M., D. Ajzenberg, B. Serrurier, M. L. Dardé, and B. Carme. 2008. Atypical Toxoplasma gondii strain from a free-living jaguar (Panthera onca) in French Guiana. Am. J. Trop. Med. Hyg. 78195-197. [PubMed] [Google Scholar]
- 14.Djurkovic-Djakovic, O., I. Klun, A. Khan, A. Nikolic, S. Knezevic-Usaj, B. Bobic, and D. Sibley. 2006. A human origin type II strain of Toxoplasma gondii causing severe encephalitis in mice. Microbes Infect. 82206-2212. [DOI] [PubMed] [Google Scholar]
- 15.Dubey, J. P., L. Applewhaite, N. Sundar, G. V. Velmurugan, L. A. Bandini, O. C. Kwok, R. Hill, and C. Su. 2007. Molecular and biological characterization of Toxoplasma gondii isolates from free-range chickens from Guyana, South America, identified several unique and common parasite genotypes. Parasitology 181-7. [DOI] [PubMed] [Google Scholar]
- 16.Dubey, J. P., S. M. Gennari, N. Sundar, M. C. Vianna, L. M. Bandini, L. E. Yai, C. H. Kwok, and C. Su. 2007. Diverse and atypical genotypes identified in Toxoplasma gondii from dogs in São Paulo, Brazil. J. Parasitol. 9360-64. [DOI] [PubMed] [Google Scholar]
- 17.Dubey, J. P., J. A. Morales, N. Sundar, G. V. Velmurugan, C. R. Gonzalez-Barrientos, G. Hernandez-Mora, and C. Su. 2007. Isolation and genetic characterization of Toxoplasma gondii from striped dolphin (Stenella coeruleoalba) from Costa Rica. J. Parasitol. 93710-711. [DOI] [PubMed] [Google Scholar]
- 18.Dubey, J. P., R. P. Rajapakse, R. R. Wijesundera, N. Sundar, G. V. Velmurugan, O. C. Kwok, and C. Su. 2007. Prevalence of Toxoplasma gondii in dogs from Sri Lanka and genetic characterization of the parasite isolates. Vet. Parasitol. 146341-346. [DOI] [PubMed] [Google Scholar]
- 19.Dubey, J. P., C. Su, J. A. Cortés, N. Sundar, J. E. Gomez-Marin, L. J. Polo, L. Zambrano, L. E. Mora, F. Lora, J. Jimenez, O. C. H. Kwok, S. K. Shen, X. Zhang, A. Nieto, and P. Thulliez. 2006. Prevalence of Toxoplasma gondii in cats from Colombia, South America and genetic characterization of Toxoplasma gondii isolates. Vet. Parasitol. 14142-47. [DOI] [PubMed] [Google Scholar]
- 20.Dubey, J. P., C. Su, J. Oliveira, J. A. Morales, R. V. Bolanos, N. Sundar, O. C. H. Kwok, and S. K. Shen. 2006. Biological and genetic characterization of Toxoplasma gondii isolates in free-range chickens from Costa Rica, Central America. Vet. Parasitol. 13929-36. [DOI] [PubMed] [Google Scholar]
- 21.Edvinsson, B., M. L. Dardé, H. Pelloux, and B. Evengard. 2007. Rapid genotyping of Toxoplasma gondii by pyrosequencing. Clin. Microbiol. Infect. 13424-429. [DOI] [PubMed] [Google Scholar]
- 22.Eisenberg, D., E. Schwarz, M. Komaromy, and R. Wall. 1984. Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J. Mol. Biol. 179125-142. [DOI] [PubMed] [Google Scholar]
- 23.Fatoohi, A. F., G. J. Cozon, T. Greenland, J. Ferrandiz, J. Bienvenu, S. Picot, and F. Peyron. 2002. Cellular immune responses to recombinant antigens in pregnant women chronically infected with Toxoplasma gondii. Clin. Diagn. Lab. Immunol. 9704-707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fazaeli, A., P. E. Carter, M. L. Darde, and T. H. Pennington. 2000. Molecular typing of Toxoplasma gondii strains by GRA6 gene sequence analysis. Int. J. Parasitol. 30637-642. [DOI] [PubMed] [Google Scholar]
- 25.Ferrandiz, J., C. Mercier, M. Wallon, S. Picot, M. F. Cesbron-Delauw, and F. Peyron. 2004. Limited value of assays using detection of immunoglobulin G antibodies to the two recombinant dense granule antigens, GRA1 and GRA6 Nt of Toxoplasma gondii, for distinguishing between acute and chronic infections in pregnant women. Clin. Diagn. Lab. Immunol. 111016-1021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fischer, H. G., S. Stachelhaus, M. Sahm, H. E. Meyer, and G. Reichmann. 1998. GRA7, an excretory 29 kDa Toxoplasma gondii dense granule antigen released by infected host cells. Mol. Biochem. Parasitol. 91251-262. [DOI] [PubMed] [Google Scholar]
- 27.Gatkowska, J., E. Hiszcynska-Sawicka, J. Kur, L. Holec, and H. Dlugonska. 2006. Toxoplasma gondii: an evaluation of diagnostic value of recombinant antigens in a murine model. Exp. Parasitol. 114220-227. [DOI] [PubMed] [Google Scholar]
- 28.Golkar, M., K. Azadmanesh, G. Khalili, B. Khoshkholgh-Sima, J. Babie, C. Mercier, M. P. Brenier-Pinchart, H. Fricker-Hidalgo, H. Pelloux, and M. F. Cesbron-Delauw. 2008. Serodiagnosis of recently acquired Toxoplasma gondii infection in pregnant women using enzyme-linked immunosorbent assays with a recombinant dense granule GRA6 protein. Diagn. Microbiol. Infect. Dis. 6131-39. [DOI] [PubMed] [Google Scholar]
- 29.Hopp, T. P., and K. R. Woods. 1981. Prediction of protein antigenic determinants from amino acid sequences. Proc. Natl. Acad. Sci. USA 783824-3828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Howe, D. K., and L. D. Sibley. 1995. Toxoplasma gondii comprises three clonal lineages: correlation of parasite genotype with human disease. J. Infect. Dis. 1721561-1566. [DOI] [PubMed] [Google Scholar]
- 31.Jacobs, D., J. F. Dubremetz, A. Loyens, F. Bosman, and E. Saman. 1998. Identification and heterologous expression of a new dense granule protein (GRA7) from Toxoplasma gondii. Mol. Biochem. Parasitol. 91237-249. [DOI] [PubMed] [Google Scholar]
- 32.Khan, A., B. Fux, C. Su, J. P. Dubey, M. L. Dardé, J. W. Ajioka, B. M. Rosenthal, and L. D. Sibley. 2007. Recent transcontinental sweep of Toxoplasma gondii driven by a single monomorphic chromosome. Proc. Natl. Acad. Sci. USA 10414872-14877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Khan, A., C. Jordan, C. Mucciolo, A. L. Vallochi, L. V. Rizzo, R. Belfort, Jr., R. W. Victor, C. Silveira, and L. D. Sibley. 2006. Genetic divergence of Toxoplasma gondii strains associated with ocular toxoplasmosis, Brazil. Emerg. Infect. Dis. 12942-949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Khan, A., C. Su, M. German, G. A. Storch, D. B. Clifford, and L. D. Sibley. 2005. Genotyping of Toxoplasma gondii strains from immunocompromised patients reveals high prevalence of type I strains. J. Clin. Microbiol. 435881-5887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kong, J. T., M. E. Grigg, L. Uyetake, S. Parmley, and J. C. Boothroyd. 2003. Serotyping of Toxoplasma gondii infections in humans using synthetic peptides. J. Infect. Dis. 1871484-1495. [DOI] [PubMed] [Google Scholar]
- 36.Labruyere, E., M. Lingnau, C. Mercier, and L. D. Sibley. 1999. Differential membrane targeting of the secretory proteins GRA4 and GRA6 within the parasitophorous vacuole formed by Toxoplasma gondii. Mol. Biochem. Parasitol. 102311-324. [DOI] [PubMed] [Google Scholar]
- 37.Lecordier, L., M. P. Fourmaux, C. Mercier, E. Dehecq, E. Masy, and M. F. Cesbron-Delauw. 2000. Enzyme-linked immunosorbent assays using the recombinant dense granule antigens GRA6 and GRA1 of Toxoplasma gondii for detection of immunoglobulin G antibodies. Clin. Diagn. Lab. Immunol. 7607-611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lecordier, L., I. Moleon-Borodowsky, J. F. Dubremetz, B. Tourvieille, C. Mercier, D. Deslée, A. Capron, and M. F. Cesbron-Delauw. 1995. Characterization of a dense granule antigen of Toxoplasma gondii (GRA6) associated to the network of the parasitophorous vacuole. Mol. Biochem. Parasitol. 7085-94. [DOI] [PubMed] [Google Scholar]
- 39.Lehmann, T., P. L. Marcet, D. H. Graham, E. R. Dahl, and J. P. Dubey. 2006. Globalization and the population structure of Toxoplasma gondii. Proc. Natl. Acad. Sci. USA 10311423-11428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lin, A., E. H. Shin, T. Y. Kim, J. H. Park, S. M. Guk, and J. Y. Chai. 2005. Genetic characteristics of the Korean isolate KI-1 of Toxoplasma gondii. Korean J. Parasitol. 4327-32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Mercier, C., K. D. Z. Adjogble, W. Däubener, and M. F. Cesbron-Delauw. 2005. Dense granules: are they key organelles to help understand the parasitophorous vacuole of all apicomplexa parasites? Int. J. Parasitol. 35829-849. [DOI] [PubMed] [Google Scholar]
- 42.Miller, M. A., P. Conrad, E. R. James, A. Packham, S. Toy-Choutka, M. J. Murray, D. Jessup, and M. Grigg. 2008. Transplacental toxoplasmosis in a wild southern sea otter (Enhydra lutris nereis). Vet. Parasitol. 15312-18. [DOI] [PubMed] [Google Scholar]
- 43.Miller, M. A., M. E. Grigg, C. Kreuder, E. R. James, A. C. Melli, P. R. Crosbie, D. A. Jessup, J. C. Boothroyd, D. Brownstein, and P. A. Conrad. 2004. An unusual genotype of Toxoplasma gondii is common in California sea otters (Enhydra lutris nereis) and is associated with mortality. Int. J. Parasitol. 34275-284. [DOI] [PubMed] [Google Scholar]
- 44.Morisset, S., F. Peyron, J. R. Lobry, J. Garweg, J. Ferrandiz, K. Musset, J. E. Gomez-Marin, A. Torre, M. Demar, B. Carme, C. Mercier, J. F. Garin, and M. F. Cesbron-Delauw. 2008. Serotyping of Toxoplasma gondii: striking homogeneous pattern between symptomatic and asymptomatic infections within Europe and South America. Microbes Infect. 10742-747. [DOI] [PubMed] [Google Scholar]
- 45.Neudeck, A., S. Stachelhaus, N. Nischik, B. Striepen, G. Reichmann, and H. G. Fischer. 2002. Expression variance, biochemical and immunological properties of Toxoplasma gondii dense granule protein GRA7. Microbes Infect. 4581-590. [DOI] [PubMed] [Google Scholar]
- 46.Pena, H. F. J., S. M. Gennari, J. P. Dubey, and C. Su. 2008. Population structure and mouse-virulence of Toxoplasma gondii in Brazil. Int. J. Parasitol. 38561-569. [DOI] [PubMed] [Google Scholar]
- 47.Petersen, E., B. Edvinsson, B. Lundgren, T. Benfield, and B. Evengard. 2006. Diagnosis of pulmonary infection with Toxoplasma gondii in immunocompromised HIV-positive patients by real-time PCR. Eur. J. Clin. Microbiol. Infect. Dis. 25401-404. [DOI] [PubMed] [Google Scholar]
- 48.Peyron, F., J. Lobry, K. Musset, J. Ferrandiz, J. E. Gomez-Marin, E. Peterson, V. Meroni, B. Rausher, C. Mercier, S. Picot, and M. F. Cesbron-Delauw. 2006. Serotyping of Toxoplasma gondii in chronically infected pregnant women: predominance of type II in the Old World and types I and III in the New World. Microbes Infect. 82333-2340. [DOI] [PubMed] [Google Scholar]
- 49.Pfrepper, K. I., G. Enders, M. Gohl, D. Krezal, H. Hlobil, D. Wassenberg, and E. Soutschek. 2005. Seroreactivity to and avidity for recombinant antigens in toxoplasmosis. Clin. Diagn. Lab. Immunol. 12977-982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Pietkiewicz, H., E. Hiszezynska-Sawicka, J. Kur, E. Peterson, H. V. Nielsen, M. Paul, M. Stankiewicz, and P. Myjak. 2007. Usefulness of Toxoplasma gondii recombinant antigens (GRA1, GRA7 and SAG1) in an immunoglobulin G avidity test for the serodiagnosis of toxoplasmosis. Parasitol. Res. 100333-337. [DOI] [PubMed] [Google Scholar]
- 51.Pietkiewicz, H., E. Hiszcynska-Sawicka, J. Kur, E. Peterson, H. V. Nielsen, M. Stankiewicz, I. Andrzejewska, and P. Myjak. 2004. Usefulness of Toxoplasma gondii-specific recombinant antigens in serodiagnosis of human toxoplasmosis. J. Clin. Microbiol. 421779-1781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Redlich, A., and W. A. Muller. 1998. Serodiagnosis of acute toxoplasmosis using a recombinant form of the dense granule antigen GRA6 in an enzyme-linked immunosorbent assay. Parasitol. Res. 84700-706. [DOI] [PubMed] [Google Scholar]
- 53.Sousa, S., D. Ajzenberg, M. Vilanova, J. Costa, and M. L. Dardé. 2008. Usefulness of GRA6 derived synthetic polymorphic peptides in Toxoplasma gondii serotyping of worldwide human samples by an immunoenzymatic assay. Clin. Vaccine Immunol. 151380-1386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Su, C., X. Zhang, and J. P. Dubey. 2006. Genotyping of Toxoplasma gondii by multilocus PCR-RFLP markers: a high resolution and simple method for identification of parasites. Int. J. Parasitol. 36841-848. [DOI] [PubMed] [Google Scholar]
- 55.Tilley, M., M. E. Fichera, M. E. Jerome, D. S. Roos, and M. W. White. 1997. Toxoplasma gondii sporozoites form a transient parasitophorous vacuole that is impermeable and contains only a subset of dense-granule proteins. Infect. Immun. 654598-4605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Welling, G. W., W. J. Weijer, R. van der Zee, and S. Welling-Wester. 1985. Prediction of sequential antigenic regions in proteins. FEBS Lett. 188215-218. [DOI] [PubMed] [Google Scholar]
- 57.Zakimi, S., H. Kyan, M. Oshiro, C. Sugimoto, X. Xuenan, and K. Fujisaki. 2006. Genetic characterization of GRA6 genes from Toxoplasma gondii from pigs in Okinawa, Japan. J. Vet. Med. Sci. 681105-1107. [DOI] [PubMed] [Google Scholar]