Abstract
The coupling of environmental conditions to cell growth and division is integral to cell fitness. In Saccharomyces cerevisiae, the transcription factor Sfp1 couples nutrient status to cell growth rate by controlling the expression of ribosome biogenesis (Ribi) and ribosomal protein (RP) genes. Sfp1 is localized to the nucleus in rich nutrients, but upon nutrient limitation or target of rapamycin (TOR) pathway inhibition by rapamycin, Sfp1 rapidly exits the nucleus, leading to repression of the Ribi/RP regulons. Through systematic cell-based screens we found that many components of the secretory system influence Sfp1 localization. Notably, the essential Rab escort protein Mrs6 exhibited a nutrient-sensitive interaction with Sfp1. Overexpression of Mrs6 prevented nuclear localization of Sfp1 in rich nutrients, whereas loss of Mrs6 resulted in nuclear Sfp1 localization in poor nutrients. These effects were specific to Sfp1 and independent of the protein kinase C (PKC) pathway, suggesting that Mrs6 lies in a distinct branch of TOR and ribosome biogenesis regulation. Rapamycin-resistant alleles of MRS6 were defective in the cytoplasmic retention of Sfp1, the control of cell size, and in the repression of the Ribi/RP regulons. The Sfp1–Mrs6 interaction is a nexus for growth regulation that links the secretory system and TOR-dependent nutrient signaling to ribosome biogenesis.
Keywords: Ribosome biogenesis, secretion, target of rapamycin, Rab GTPase escort protein, Sfp1, Mrs6
Nutrient availability and environmental stress conditions dictate cellular growth rate, which in turn governs the onset of cell division (Jorgensen and Tyers 2004). The most significant draw on macromolecular synthesis is the biogenesis of new ribosomes. In yeast, the vast majority of total cellular transcriptional activity by RNA polymerase I (Pol I), Pol II, and Pol III is oriented toward production of ribosomal RNA (rRNA), ribosomal protein (RP), and tRNA (Warner 1999). An elaborate nutrient- and stress-sensing network allows cells to adapt rapidly to the ever-changing environment: In particular, two main nutrient-sensing conduits—the target of rapamycin (TOR) and protein kinase A (PKA) pathways—link nutrient status to critical cellular processes, including ribosome biogenesis, autophagy, and entry into G0 (Rohde et al. 2008; Zaman et al. 2008). TOR and PKA sense partially nonoverlapping nutrients and often converge on common targets (Jorgensen and Tyers 2004; Dechant and Peter 2008; Zaman et al. 2008).
The conserved TOR network relays amino acid concentrations, glucose, and perhaps other nutrient signals to the cellular machinery (De Virgilio and Loewith 2006; Wullschleger et al. 2006; Dechant and Peter 2008). The Tor kinases are most closely related to the phosphatidyl-inosital-3 kinase (PI3K) family of lipid kinases and exert their effects through two distinct complexes: TOR Complex 1 (TORC1) and TORC2, which control growth rate and cell polarization, respectively. In yeast, TORC1 is comprised of Kog1, Lst8, Tco89, and either the Tor1 or Tor2 kinase, whereas TORC2 is composed of Tor2 exclusively in association with Lst8, Avo1, Avo2, Avo3, Bit2, and Bit61 (Loewith et al. 2002; Wedaman et al. 2003). The TORC1 and TORC2 complexes are structurally and functionally conserved from yeast to humans (Wullschleger et al. 2006). Inhibition of TORC1 by the macrolide rapamycin mimics nutrient starvation and causes G1 arrest, inhibition of protein synthesis, glycogen accumulation, induction of autophagy, and entry into quiescence (Wullschleger et al. 2006). TORC1 is also intimately linked to various aspects of vesicular trafficking (Dechant and Peter 2008; Rohde et al. 2008).
TOR is a nexus for nutrient signaling. In metazoans, TOR is activated in response to growth factor signals via class I PI3K-mediated activation of the prosurvival kinase Akt/PKB, which inhibits the GTPase-activating proteing (GAP) complex Tsc1/2, leading to activation of the Rheb GTPases and stimulation of TOR (Wullschleger et al. 2006). Other signals also impinge on Tsc1/2, including hypoxia, ATP sufficiency through the LKB and AMP-activated kinase couple, and MAPK activity (Wullschleger et al. 2006). The Rag family of membrane-associated GTPases relays amino acid availability directly to mTORC1 by recruitment of TORC1 into close proximity with Rheb (Kim et al. 2008; Sancak et al. 2008). In both yeast and mammals, the class C Vps and EGO/GSE complexes also appear to activate TORC1 (Dubouloz et al. 2005; Nobukuni et al. 2005; Gao and Kaiser 2006; Zurita-Martinez et al. 2007). TOR, its activators, and effectors often localize to intracellular membranes, which may thus serve as platforms for TOR signaling complexes (Wullschleger et al. 2006; Yan et al. 2006; Aronova et al. 2007; Dechant and Peter 2008; Rohde et al. 2008).
TOR governs metabolism and growth through a host of downstream effectors. In yeast, TOR exerts control primarily at the level of gene expression, often through the cytoplasmic retention of stress- and nutrient-responsive transcription factors (Duvel et al. 2003; De Virgilio and Loewith 2006). In mammalian cells, TOR stimulates protein synthesis by phosphorylation of two critical targets: the translational activator S6 kinase (S6K) and the translational inhibitor 4E-BP1 (Wullschleger et al. 2006). Similarly, the yeast equivalent of S6K, the AGC kinase Sch9, directly mediates TORC1-dependent regulation of translation initiation, RP gene expression, and entry into G0 (Urban et al. 2007). TOR controls other aspects of ribosome biogenesis including the Pol I- and Pol III-dependent transcription of the rDNA and tRNA genes through phosphorylation of dedicated transcription factors (Mayer and Grummt 2006). In yeast, Tor1 itself may activate rDNA transcription in rich nutrient conditions by translocating to the nucleus and binding directly to promoter DNA (Li et al. 2006); however, in other studies, Tor1 has been localized to internal membrane structures but not the nucleus (Wedaman et al. 2003; Aronova et al. 2007; Sturgill et al. 2008).
Previously, to identify pathways that couple growth and division, we carried out a systematic screen for yeast deletion mutations that alter cell size (Jorgensen et al. 2002). Defects in ribosome biogenesis, as opposed to defects in protein translation per se, cause a pronounced small size, suggesting that a high rate of ribosome synthesis delays commitment to cell division (Jorgensen et al. 2002). Disruption of either Sch9 or the zinc finger transcription factor Sfp1 causes extremely small size and dramatic repression of genes required for the biogenesis of new ribosomes (Jorgensen et al. 2004a). Sch9 is specifically required for maximal expression of the RP regulon and is regulated in a nutrient-sensitive fashion by both phosphorylation and localization to the vacuolar membrane (Jorgensen et al. 2004a). Sfp1 controls a large cohort of >200 genes implicated in the complex pathways that assemble mature ribosomes, termed the ribosome biogenesis (Ribi) regulon; Sfp1 also directly or indirectly activates the RP regulon (Jorgensen et al. 2002). Sfp1 localization is highly responsive to nutrient conditions: In glucose medium, Sfp1 is located in the nucleus, but upon nutrient limitation or exposure to various stresses, Sfp1 relocalizes to the cytoplasm within minutes (Jorgensen et al. 2004a; Marion et al. 2004). In addition to Sch9 and Sfp1, RP gene expression depends on several other transcription factors and chromatin modifiers (Jorgensen et al. 2004b). Nutrient-responsive localization of Sfp1 depends on TOR and PKA signaling (Jorgensen et al. 2004a; Marion et al. 2004); however, the critical mechanism(s) that link Sfp1 localization to TOR and PKA activity are unknown.
To delineate how nutrients are linked to ribosome biogenesis and cell size, we undertook systematic screens for factors that alter Sfp1 localization in different nutrient conditions; these screens uncovered many components of the secretory system. We identify the Rab escort protein Mrs6 as the primary cytoplasmic retention factor for Sfp1 and demonstrate that the Sfp1–Mrs6 interaction links TOR signaling, the secretory network, and ribosome biogenesis in the control of cell growth.
Results
The endomembrane system influences nutrient-responsive localization of Sfp1
The subcellular localization of Sfp1 responds rapidly to nutrient status: Sfp1 partitions to the nucleus under optimal growth conditions and to a cytoplasmic compartment upon nutrient limitation or a variety of stresses (Jorgensen et al. 2004a; Marion et al. 2004). To identify the nutrient-dependent pathways that relay signals to Sfp1, we assayed the localization of Sfp1 in genome-wide loss- and gain-of-function screens. A collection of 800 tet-regulated essential genes was assayed for Sfp1GFP localization defects upon promoter repression (Mnaimneh et al. 2004). In a complementary approach, a set of 5280 galactose-regulated yeast ORFs was assayed for defects in Sfp1GFP localization upon promoter induction (Sopko et al. 2006). Each collection of strains was examined on an Evotec Opera automated high-resolution confocal microscopy platform in glucose medium, when Sfp1GFP is normally nuclear, and in glycerol medium, when Sfp1GFP is normally cytoplasmic (Fig. 1A). The distribution of Sfp1GFP between the nuclear and cytoplasmic compartments was quantified using a custom algorithm to determine nuclear and cytoplasmic areas of Sfp1GFP signal. The highest-ranking hits from each of the two screens are shown in Table 1; complete quantitative data for both screens is provided in Supplemental Table 1.
Figure 1.
Genome-wide screens for regulators of Sfp1 localization converge on the secretory system. (A) Schematic of cell-based screen for aberrant Sfp1 localization in strains depleted for one of 800 essential genes expressed from the tet promoter or overexpression of 5280 genes from the GAL1 promoter. (B) Elevated PKA activity alters Sfp1 distribution. An sfp1∷SFP1GFP strain bearing a GAL1-TPK1 plasmid was grown in galactose medium for 2 h prior to shift to glycerol medium for 40 min and visualization of Sfp1GFP. (C) Increased nuclear export alters Sfp1 distribution. An sfp1∷SFP1GFP strain bearing a GAL1-MSN5 plasmid was grown in galactose medium for 2 h prior to shift to glycerol medium for 40 min and visualization of Sfp1GFP. (D) Perturbation of secretory pathway function causes aberrant Sfp1 localization in rich medium. An sfp1∷SFP1GFP tet-YPT1 strain was grown in 10 μg/mL doxycycline in glucose medium for 16 h and shifted to glycerol medium for 40 min prior to visualization of Sfp1GFP. (E) Rapid inhibition of secretory pathway flux with Brefeldin A (BFA) causes cytoplasmic relocalization of Sfp1. An sfp1∷SFP1YFP erg6Δ strain was treated with 100 ng/mL BFA for the indicated times prior to visualization of SFP1YFP. (F) Strains defective in ER/Golgi trafficking exhibit aberrant Sfp1 localization. Subcellular distribution of Sfp1YFP was visualized in sfp1∷SFP1YFP sly1-1 or sfp1∷SFP1YFP ypt6169ts strains grown in synthetic complete media at 23°C prior to and following shift for 2 h to 37°C. (G) A log-phase culture of an sfp1∷SFP1YFP ypt1-1 strain was visualized at a permissive temperature of 30°C.
Table 1.
List of genes recovered in Sfp1 localization screens
(C) Cytoplasm; (ER) endoplasmic reticulum; (N) nucleus; (PM) plasma membrane
aFrom Saccharomyces Genome Database http://www.yeastgenome.org.
bRatio of Sfp1GFP signal area in given strain versus wild type.
cFraction of N signal detected as a function of total cell number.
The screening approach was validated by the isolation of components of the Ras/PKA signaling pathway, which we and others have shown previously to regulate Sfp1 localization (Jorgensen et al. 2004a; Marion et al. 2004). The catalytic subunits of PKA are encoded by partially redundant genes TPK1, TPK2, and TPK3 (Zaman et al. 2008), and deregulation of a single TPK gene suffices to reduce rapamycin-induced Sfp1 nuclear depletion (Marion et al. 2004). Aberrant nuclear retention of Sfp1 occurred in poor nutrient conditions in strains that overexpressed TPK1, TPK2, or TPK3 (Fig. 1B; Table 1). In contrast, another factor identified in the overexpression screen, the karyopherin MSN5/KAP142, caused the inappropriate nuclear export of Sfp1 in glucose medium (Fig. 1C). An msn5Δ strain did not exhibit constitutive Sfp1 nuclear localization (data not shown), implying functional compensation by other exportins. SFP1 itself also scored in the overexpression screen, presumably because competition for endogenous receptors lead to mislocalization of the Sfp1GFP reporter in all compartments.
Strikingly, the majority of genes that caused anomalous Sfp1 localization when depleted or overexpressed encoded constituents of the secretory pathway (Zerial and McBride 2001). The largest class of factors to affect Sfp1 localization participate in endoplasmic reticulum(ER)-to-Golgi vesicle transport and included the essential Rab GTPase Ypt1, the SNARE components Sec17 and Sed5, and the COPII vesicle-associated proteins Erv29 and Sec23 (Fig. 1D; Table 1). A second large class of secretory components recovered in the screens have roles in different aspects of Golgi vesicle trafficking, including the Rab GTPase Sec4; its cognate GAP Msb4 and its effector Sro7, which mediate transport from the Golgi to the plasma membrane; the SNAREs Sec20 and Use1, which mediate retrograde transport from the Golgi to the ER; and a variety of other essential and nonessential Golgi-associated factors such as Tlg1 and Avl9, which mediate endocytic and exocytic vesicle fusion events, respectively. A third class of factors recovered affect secretion through structural or metabolic mechanisms, including the glycosylphosphatidylinositol (GPI) anchor transamidase subunits Gpi16 and Gab1, the dolichol phosphate mannose synthase Dpm1, the ER chaperonin Rot1, the palmitoyl transferase Akr1, and the ergosterol saturase Erg5. These genetic results were supported by the observation that the early secretory pathway inhibitor Brefeldin A, which blocks ARF GTPase activities (Peyroche et al. 1999), also caused localization of Sfp1GFP to the cytoplasm (Fig. 1E).
Strains defective in the essential gene SLY1, which encodes a component of the ER–Golgi vesicle trafficking system, and YPT6, which encodes a Rab GTPase required for Golgi vesicle fusion, have been shown previously to be defective in rRNA and RP mRNA transcription (Mizuta and Warner 1994; Li and Warner 1996); however, these genes were not represented in our filtered tet screen data. To extend our results, we tested the effects of sly1-1 and ypt6169ts alleles on Sfp1 localization. Sfp1YFP redistributed to the cytoplasm upon shift of a sly1-1 strain to the nonpermissive temperature, but not upon shift of a ypt6169ts strain (Fig. 1F). Finally, we further examined the role of the Rab GTPase Ypt1, which is required for ER–Golgi trafficking and was isolated in our tet repression screen. Sfp1YFP localization was perturbed in a cold-sensitive ypt1-1 strain even at the permissive temperature (Fig. 1G). Notably, the SLY1-20 allele is a dominant suppressor of ypt1-1 (Dascher et al. 1991); moreover, many of the secretion genes recovered in our screen may either directly or indirectly affect the activity of Ypt1. Taken together, these results demonstrate that activity of the secretory system is intimately linked to Sfp1 localization.
Nutrients and stress alter association of Sfp1 and the Rab escort protein Mrs6
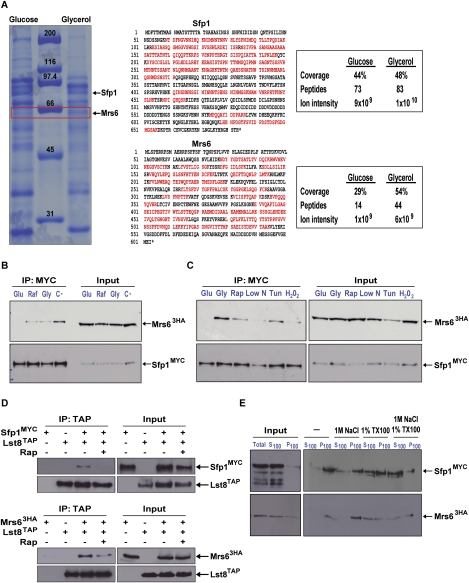
Given the pronounced effect of secretory pathway perturbation on Sfp1 localization, we sought to identify direct physical connections between Sfp1 and secretory proteins. SFP1Flag was transiently expressed from the GAL1 promoter, after which cultures were shifted briefly to either glucose or glycerol medium, Sfp1Flag complexes immunopurified, and associated proteins identified by mass spectrometry. An ∼66-kDa species that was preferentially bound to Sfp1 isolated from glycerol-treated cells was identified as Mrs6; this interaction has also been reported in two high-throughput studies, and recently verified (Ho et al. 2002; Tarassov et al. 2008; Lampiainen et al. 2009). The high-peptide sequence coverage of Sfp1 (48%) and Mrs6 (54%) and an estimation of total ion current for each protein species suggested that the Sfp1–Mrs6 complex was near stoichiometric in glycerol medium and that the interaction was reduced severalfold in glucose medium (Fig. 2A). Mrs6 is an essential escort protein that is required for the prenylation and membrane delivery of the Rab GTPases, including Ypt1, Sec4, Ypt6, Vps21, and other family members that control vesicle trafficking at different stages in the secretory system (Fujimura et al. 1994; Bialek-Wyrzykowska et al. 2000; Zerial and McBride 2001). Notably, depletion of Ypt1 and Sec4 markedly influenced Sfp1 localization (Fig. 1D; Table 1); we did not, however, recover MRS6 in our screen because the tet-MRS6 and GAL1-MRS6 strains in our collections were defective. Intriguingly, MRS6 is a multicopy suppressor of the iraΔ1 defect that results in hyperactivation of the Ras/PKA pathway (Fujimura et al. 1994).
Figure 2.
Nutrient-sensitive interaction between Sfp1 and the Rab escort protein Mrs6. (A) Identification of an Mrs6–Sfp1 complex. Sfp1FLAG complexes were immunoprecipitated from glucose or glycerol medium and resolved by SDS-PAGE, and differentially recovered species were identified by mass spectrometry. All peptides identified under each condition are indicated in red. (B) The Sfp1–Mrs6 complex is responsive to carbon source. An sfp1∷SFP1MYC13 mrs6∷MRS63HA strain was grown to log phase in glucose medium and incubated for 30 min in the indicated medium. Sfp1MYC13 was immunoprecipitated and the presence of Mrs63HA was assessed by immunoblot. (C) The Sfp1–Mrs6 complex is responsive to stress conditions. An sfp1∷SFP1MYC13 mrs6∷MRS63HA strain was grown to log phase in glucose medium, exposed to the indicated stresses, and processed as in B. (D) Interactions between Sfp1, Mrs6, and the TORC1 complex. sfp1∷SFP1MYC13 lst8∷LST8TAP or mrs6∷MRS63HA lst8∷LST8TAP strains were grown in rich glucose medium and treated for 30 min with 200 ng/mL rapamycin, and Lst8TAP immune complexes were analyzed for either Sfp1MYC13 (top panel) or Mrs63HA (bottom panel) by immunoblot. (E) Sfp1 and Mrs6 associate with an insoluble membrane fraction. (Left panel) A 100,000g P100 fraction was resuspended in lysis buffer and either untreated or incubated with 1 M NaCl and/or 1% Triton X-100, then repelleted at 100,000g and the presence of SFP1MYC13 and MRS63HA in the supernatant (S100) and the pellet (P100) was assessed by immunoblot.
To confirm nutrient regulation of the Mrs6–Sfp1 interaction, endogenous levels of Sfp1 and Mrs6 were tested for association under a variety of nutrient and stress conditions that inhibit ribosome biogenesis: carbon source limitation, rapamycin, nitrogen source limitation, and H2O2. The Sfp1–Mrs6 interaction was increased under each of the nutrient and stress conditions tested (Fig. 2B,C). We note that rapid harvest of cell cultures was needed to avoid transient starvation and artifactual association of Sfp1 and Mrs6 in glucose medium. As the Sfp1–Mrs6 interaction was sensitive to rapamycin, we determined whether Sfp1 and Mrs6 might interact with components of TORC1 by coimmunoprecipitation from yeast lysates. We detected specific interactions between Sfp1, Mrs6, and the TORC1 components Tco89, Lst8, and Tor1; association of the TORC1 complex with Sfp1 exhibited partial sensitivity to rapamycin (Fig. 2D; Supplemental Fig. 1).
As Mrs6 is associated peripherally with ER/Golgi membranes (Miaczynska et al. 1997), we examined the distribution of Sfp1 in crude cytosolic and membrane fractions isolated from rapamycin-treated cells (Fig. 2E). Approximately 10% of Sfp1 was detected in a 100,000g membrane pellet (P100) fraction, which is comparable with the membrane partitioning reported for the TOR effectors Gln3 and Tap42 (Yan et al. 2006; Puria et al. 2008). Treatment with NaCl and/or detergent released both Sfp1 and Mrs6 from the P100 fraction to approximately the same extent. The physical interactions of Mrs6 with TORC1 subunits and the similar membrane association of both Sfp1 and Mrs6 suggest that TOR signaling impinges on the Sfp1–Mrs6 complex at an endomembrane compartment. We were unable to discern regulated colocalization of Sfp1 and Mrs6 by fluorescence microscopy because Mrs6 was diffusely localized across the cytoplasm and nucleoplasm; intriguingly, however, a fraction of Mrs6 appeared to transiently partition into the nucleus upon shift from glucose to glycerol medium (data not shown).
Mrs6 is necessary and sufficient to control Sfp1 localization and cell size
We next tested whether the Sfp1–Mrs6 interaction might dictate the localization of Sfp1 by examining the distribution of Sfp1YFP upon artificial alteration of Mrs6 levels. Overexpression of MRS6 from the GAL1 promoter not only increased the cytoplasmic fraction of Sfp1 under all conditions, but also completely prevented nuclear accumulation of Sfp1 in glucose medium (Fig. 3A). This effect was evident within the time frame required to induce galactose gene expression (Jorgensen et al. 2002), consistent with a direct effect of Mrs6 on Sfp1 (Supplemental Fig. 2). Conversely, depletion of MRS6 prevented the relocalization of Sfp1 to the cytoplasm in glycerol medium (Fig. 3B). Because modulating the levels of Mrs6 was able to override the nutrient-dependent control of Sfp1 localization, the nutrient signaling pathways that govern the Sfp1–Mrs6 interaction may be limiting in vivo.
Figure 3.
Mrs6 controls Sfp1 localization. (A) MRS6 overexpression prevents nuclear accumulation of Sfp1 in glucose medium. An sfp1∷SFP1YFP strain expressing control vector or GAL1-MRS6Flag was grown in synthetic raffinose medium, prior to induction with galactose for 120 min. Sfp1YFP was visualized prior to (left) and after (right) shift into glucose medium. (B) Increased nuclear accumulation of Sfp1 upon Mrs6 depletion in glycerol medium. An sfp1∷SFP1YFP mrs6∷tet-MRS6 strain was grown in synthetic glucose medium that contained either 0 or 10 μg/mL doxycycline and shifted into glycerol medium prior to visualization of Sfp1YFP. (C) Mrs6 overexpression causes a reduction in cell size. Size distributions were acquired for log-phase cultures expressing either control vector (black) or GAL1-MRS6 (red) that had been induced for 6 h with galactose. (D) Mrs6 depletion causes an increase in cell size. Size distributions were acquired for log-phase cultures of an mrs6∷tet-MRS6 strain grown in glycerol medium, in the absence (black) or presence (red) of 10 μg/mL doxycycline. As a reference, a wild-type culture was grown in glucose medium without doxycycline (blue). (E) An sfp1Δ mutation bypasses the G2/M delay caused by MRS6 depletion. DNA content was determined for mrs6∷tet-MRS6 (left panel) or sfp1Δ mrs6∷tet-MRS6 (right panel) strains grown in glucose medium in the presence or absence of 10 μg/mL doxycycline. (F) Deletion of SFP1 is epistatic to depletion of MRS6 for cell size. Cell size distributions were acquired for mrs6∷tet-MRS6 and sfp1Δ mrs6∷tet-MRS6 strains grown in glucose medium in the absence (black) or presence (red) of 10 μg/mL doxycycline. (G) The Rab GTPase Ypt1 competes with Sfp1 for Mrs6. An in vivo PCA assay was used to monitor interactions between the indicated protein fusions as judged by growth in the presence of the DHFR inhibitor methotrexate. (H) An sfp1∷SFP1MYC13 mrs6∷MRS63HA strain bearing GAL1-YPT1, GAL1-SEC4, GAL1-RHO1 plasmids or empty vector was grown to early log phase in synthetic raffinose medium and induced with 2% galactose for 2 h prior to harvesting. Sfp1MYC13 immune complexes were analyzed for the presence of Mrs63HA and the indicated Flag-tagged proteins by immunoblot.
Importantly, from the perspective of size homeostasis, the alterations in Sfp1 distribution caused by manipulation of Mrs6 were accompanied by corresponding effects on cell size. Overexpression of MRS6 caused a decrease in cell size compared with wild type (Fig. 3C), whereas MRS6 depletion caused an increase in cell size on glycerol medium (Fig. 3D). These MRS6 effects are specific: Simple perturbation of secretory function would normally cause a delay in cell cycle progression and concomitant increase in cell size (Jorgensen et al. 2002); i.e., the opposite phenotype to that observed for overexpression of MRS6. Repression of the tet-MRS6 allele severely compromised cell growth and caused a G2/M cell cycle delay (Fig. 3E); this G2/M delay is reminiscent of that caused by overexpression of SFP1 (Jorgensen et al. 2004a). To determine if the tet-MRS6 depletion phenotype was due to Sfp1 hyperactivity, we analyzed the phenotype of a tet-MRS6 mrs6Δ sfp1Δ strain. As expected, an sfp1Δ mutation caused cells to accumulate in G1 phase (Jorgensen et al. 2002). The G2/M delay was suppressed by deletion of SFP1 (Fig. 3E). Depletion of MRS6 did not alter the mode size of a wild-type strain grown on glucose medium, but did cause a partial bimodal size distribution, perhaps indicative of misregulation in nutrient signaling (Fig. 3F, left panel). Unlike in a glycerol culture of a wild-type strain, depletion of MRS6 caused only a modest increase in the size of an sfp1Δ strain grown in glucose medium (Fig. 3, cf. D and F, right panel). These results suggest that Mrs6 acts upstream of Sfp1 in cell size control. The essential function of MRS6 was not bypassed by sfp1Δ because MRS6 is required for the prenylation of essential Rab GTPases (data not shown).
Given that Mrs6 is a necessary and sufficient factor for retention of Sfp1 in the cytoplasm, we sought to determine if other Mrs6 interaction partners might compete with Sfp1 in vivo. We monitored Sfp1 and Mrs6 interactions using a sensitive in vivo protein complementation assay (PCA) used previously to detect the Sfp1–Mrs6 interaction in a high-throughput study (Tarassov et al. 2008). As indicated by the ability of a PCA reporter strain to grow on methotrexate-containing medium, an SFP1-F[3] DHFR fusion protein interacted with an MRS6-F[1,2] fusion protein (Fig. 3G). Overexpression of the YPT1 Rab GTPase from the GAL1 promoter interfered with the interaction of a MRS6-F[3] fusion with a fusion to another of its cognate GTPases, SEC4-F[1,2]. Notably, overexpression of YPT1 also interfered with the growth of the SFP1-F[3]–MRS6-F[1,2] strain, suggesting that Sfp1 shares the same binding site on Mrs6 as the Rab GTPases. We validated the PCA competition studies by coimmunoprecipitation of Sfp1 and Mrs6 from strains that overexpressed various GTPases. The interaction between Sfp1 and Mrs6 was significantly compromised upon overexpression of Ypt1 but not Sec4 or Rho1 (Fig. 3H). All told, these results suggest that Mrs6 is the primary cytoplasmic anchor protein for Sfp1 in poor nutrient and stress conditions and that some Rab GTPases may compete with Sfp1 for binding to Mrs6.
Mrs6 specifically regulates Sfp1 in parallel to the PKA and PKC pathways
The negative regulation of Sfp1 by Mrs6 is consistent with the ability of MRS6 overexpression to partially bypass the growth defects caused by hyperactivation of the PKA pathway (Fujimura et al. 1994). Although Sfp1 can be phosphorylated by PKA in vitro (Budovskaya et al. 2005) and driven into the nucleus by TPK overexpression (Fig. 1B; Table 1), overexpression of MRS6 was sufficient to inhibit the nuclear accumulation of Sfp1 in a RAS2Val19 strain in which the PKA pathway is hyperactive (Fig. 4A). Mutation of the two PKA consensus sites in Mrs6 (S9A and S16A) did not prevent nuclear retention of Sfp1 upon TPK1 overexpression in poor nutrient conditions (data not shown). These findings, in conjunction with the original isolation of MRS6 as a multicopy suppressor of high PKA activity, argue that MRS6 regulates Sfp1 in a pathway that is either downstream from or parallel to Ras/PKA, most likely through TOR signaling.
Figure 4.
Specificity of Sfp1 regulation by Mrs6. (A) Hyperactivation of the Ras/PKA pathway does not block MRS6-induced Sfp1 relocalization. Sfp1CFP localization was visualized in wild-type and RAS2VAL19 cells carrying either GAL1-MRS6 or an empty vector. Cells were grown to log phase in synthetic raffinose medium, induced for 120 min with galactose, and imaged prior to and following glucose addition. (B) The PKC pathway is not required for nutrient- and stress-induced Sfp1 relocalization. Sfp1YFP localization was visualized in a sfp1∷SFP1YFP pkc1Δ strain under the indicated conditions. (C) MRS6 does not influence TOR effectors that mediate the NCR response. Localization of Sfp1GFP, Gap1GFP, and Gln3GFP was determined in strains carrying either GAL1-MRS6 or an empty vector grown in raffinose medium following galactose induction for 120 min. (D) MRS6 does not influence the TOR and stress-responsive transcription factor Msn2. Localization of Msn2GFP and Sfp1GFP was determined in strains carrying either GAL1-MRS6 or an empty vector grown in galactose medium.
It has been demonstrated previously that the repression of RP, tRNA, and rDNA transcription in response to secretory pathway mutations or stress depends on PKC1, which encodes the yeast ortholog of protein kinase C (Nierras and Warner 1999; Li et al. 2000). This response is a consequence of the cell wall stress caused by secretory defects, and likely impinges on the Rap1 transcription factor, which is a primary determinant of RP gene expression (Li et al. 2000). To determine if secretory pathway defects might indirectly affect Sfp1 localization through a PKC-dependent signal arising from cell wall stress, we examined Sfp1 localization under different conditions in a pkc1Δ strain. Disruption of the PKC pathway did not affect the cytoplasmic relocalization of Sfp1 caused by a shift to a poor carbon source, inhibition of TOR signaling by rapamycin, or inhibition of glycoprotein synthesis by tunicamycin (Fig. 4B). The PKC-selective inhibitor staurosporine also did not affect Sfp1 relocalization in response to overexpression of Mrs6 (Supplemental Fig. 3); in addition, disruption of the actin cytoskeleton by latrunculin B did not affect Sfp1 relocalization (data not shown). These results indicate that the localization of Sfp1, and consequently the downstream Ribi regulon, is coupled to secretory function by a different mechanism than that which affects RP, tRNA, and rDNA transcription.
To discern whether Mrs6 is a specific regulator for Sfp1, we tested the influence of Mrs6 on other downstream effectors of TOR. In the nitrogen catabolite repression (NCR) response, the general amino acid permease Gap1 is sorted to the plasma membrane in a TOR-dependent manner (De Virgilio and Loewith 2006; Gao and Kaiser 2006). In addition, TOR regulates GAP1 expression by maintaining the transcriptional activator Gln3 in a cytoplasmic complex with its repressor Ure2 (Beck and Hall 1999; Cardenas et al. 1999). Overexpression of MRS6 had no effect on Gap1 localization (Fig. 4C, left panel) or Gln3 nuclear localization (Fig. 4C, right panel). TORC1 also promotes the phosphorylation and nuclear exclusion of Msn2/Msn4 (Beck and Hall 1999). The localization of Msn2/Msn4 was also unaffected by overexpression of MRS6 (Fig. 4D), nor was the localization of Rtg1/Rtg3 affected in the mitochondrial retrograde signaling response (data not shown). These results indicate that Mrs6 specifically regulates Sfp1 via TOR and that Mrs6 operates in parallel to the PKC pathway that controls other aspects of ribosome biogenesis.
Rapamycin-resistant alleles of MRS6 are defective for Sfp1 relocalization
Given that the essential function of MRS6 is independent of SFP1 and that Mrs6 negatively regulates Sfp1, we reasoned that it might be possible to isolate alleles of MRS6 that confer resistance to rapamycin without compromising cell viability. A collection of MRS6 alleles was generated by PCR mutagenesis followed by selection in the presence of a high concentration of rapamycin. Three such alleles, designated MRS6-R1, MRS6-R2, and MRS6-R3, were defective in the ability to relocalize Sfp1 to the cytoplasm in response to rapamycin (Fig. 5A; data not shown). The response of MRS6-R strains to nutrient limitation was more complex as Sfp1 partially relocalized to punctate cytoplasmic structures in glycerol medium (Fig. 5A). Sequence analysis indicated that the R1, R2, and R3 alleles encoded the amino acid alterations Ser335Pro, Trp67Arg/Ser125Phe, and Gly122Ser, respectively (Fig. 5B). Interestingly, the temperature-sensitive allele mrs6-2 encodes the identical Ser335Pro substitution, as well as a Gly227Val substitution in Domain II, which mediates essential interactions with the geranylgeranyltransferase II (GGTase II) that catalyzes Rab prenylation (Bialek-Wyrzykowska et al. 2000; Alory and Balch 2003). The Mrs6-2 protein is defective in prenylation activity and vesicle polarization/vacuole morphology, and accumulates in G2/M phase (Bialek-Wyrzykowska et al. 2000). In contrast, the MRS6-R strains had no obvious growth defect (Fig. 5C) and had a normal cell cycle distribution (data not shown). Moreover, the MRS6-R1 allele partly abrogated the reduction in cell size caused by acute rapamycin treatment (Fig. 5D); this observation buttresses the notion that Mrs6 is a principle conduit for size regulation by the TOR pathway. Significantly, the MRS6-R missense mutations all lie within or adjacent to the Domain I region of Mrs6, which corresponds to the Rab-binding platform (Alory and Balch 2003); this result suggests that Sfp1 engages the same interface on Mrs6 as the Rab GTPases. To further explore this interface, we created additional site-directed MRS6 mutations. Substitution of an Ala residue at position 335 resulted in a similar but milder phenotype than the MRS6-R1 allele, whereas substitution of a Glu residue caused a severe G2/M delay, heterogeneous colony growth on glucose medium, and irreversible sensitivity to rapamycin exposure (Supplemental Fig. 4). The inability of the Ser335Glu mutant to recover from rapamycin-induced arrest was accompanied by persistent macroautophagy (Supplemental Fig. 5), similar to the rapamycin recovery defect caused by defects in the EGO complex (Dubouloz et al. 2005). The above findings demonstrate that the Sfp1 anchor function of Mrs6 mediates TORC1-dependent growth and size control, and that this function is separable from the essential role of Mrs6 in secretion.
Figure. 5.
Isolation and characterization of rapamycin-resistant alleles of MRS6. (A) Nuclear retention of Sfp1 upon rapamycin treatment in the presence of rapamycin-resistant MRS6-R alleles. Localization of Sfp1GFP in synthetic glucose medium, either untreated or treated with rapamycin (200 ng/mL), and synthetic glycerol medium was assessed in the indicated strains. (B) Location of amino acid substitutions in MRS6-R alleles. Conserved structural Domain I and Domain II of Rab escort proteins and Rab GDI proteins are indicated in green; conserved regions of Rab escort proteins are indicated in purple (Alory and Balch 2003). (C) Growth of wild-type and mutant alleles of MRS6. The indicated strains and, as a control, a rapamycin-resistant TOR1-1 strain were streaked onto glucose medium containing 20 ng/mL rapamycin and imaged after 4 d. (D) Rapamycin-resistant alleles of MRS6 fail to adjust cell size in response to rapamycin. Cell size profiles of indicated strains grown in synthetic glucose medium either with or without rapamycin (200 ng/mL) and, as a control, synthetic glycerol medium.
Mrs6 is specifically required for regulation of the Ribi regulon by TORC1
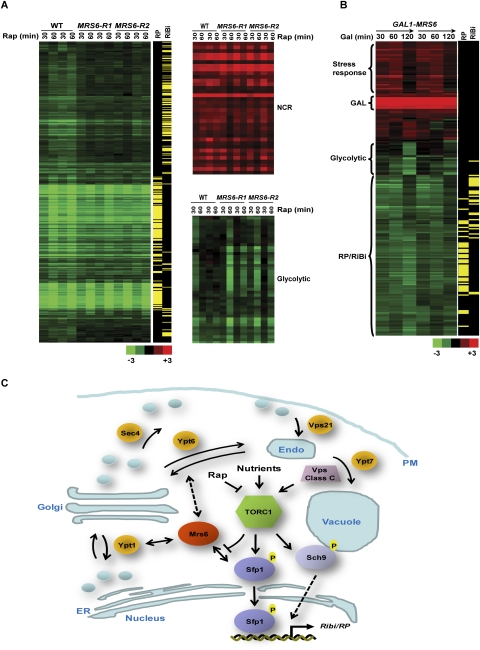
Rapamycin treatment results in a robust transcriptional signature caused by loss of TORC1 activity (Cardenas et al. 1999; Hardwick et al. 1999; Komeili et al. 2000; Shamji et al. 2000). To determine the specific role of Mrs6 in TORC1-dependent gene expression, we examined the transcriptional profiles of wild-type, MRS6-R1, and MRS6-R2 strains in the absence and presence of rapamycin. Untreated wild-type and the MRS6-R1/2 strains displayed no difference in transcriptional profile. As expected, rapamycin caused a massive transcriptional signature in wild-type cells (Fig. 6A; Supplemental Table 2). G0-specific Pol II genes, Msn2/4-dependent stress response genes, autophagy genes, NCR genes, tricarboxylic acid cycle genes, and Rtg1/3 and Hap2/3/4/5 target genes were all induced, while the glycolytic, RP, and Ribi regulons were repressed (Cardenas et al. 1999; Hardwick et al. 1999; Komeili et al. 2000; Shamji et al. 2000). The global expression profiles of rapamycin-treated MRS6-R1 and MRS6-R2 strains were similar to wild type, with the striking exception that repression of the Ribi regulon was markedly attenuated (Fig. 6A). Notably, repression of the RP regulon by rapamycin was only marginally affected in the MRS6-R strains; the residual repression may be a downstream consequence of repression of activators of RP genes that form part of the Ribi regulon (Jorgensen et al. 2002). The MRS6-R alleles also a caused delay in the repression of glycolysis genes (Fig. 6A); this may also be a downstream consequence of Ribi repression (Jorgensen et al. 2002), or may reflect a more direct role for Sfp1, Mrs6, and/or TORC1 in glycolytic gene regulation (Hardwick et al. 1999; Cipollina et al. 2008). The MRS6-R alleles had only a minor effect on activation of the NCR regulon by rapamycin (Fig. 6A). To validate the effects of the MRS6-R alleles on TOR-regulated gene expression, we also determined the effects of MRS6 overexpression on TOR-associated transcriptional profiles. Overexpression of MRS6 from the GAL1 promoter caused rapid repression of the Ribi regulon and delayed repression of the RP and glycolytic regulons (Fig. 6B). A transient induction of stress response genes was also observed, perhaps as a consequence of secretory system perturbation caused by excess Mrs6 (Fig. 6B). In summary, the transcriptional phenotypes conferred by MRS6-R alleles and by MRS6 overexpression demonstrate that Mrs6 serves to control Sfp1 transcriptional functions, consistent with the observed effects on Sfp1 localization and cell size.
Figure 6.
MRS6 specifically controls the Ribi regulon. (A) MRS6 is required for rapamycin-induced repression of the Ribi regulon. Expression profiles were determined by microarray analysis for mrs6Δ strains bearing MRS6, MRS6-R1, and MRS6-R2 alleles on a low-copy plasmid treated for 30 and 60 min with 100 ng/mL rapamycin. Values represent the average expression change relative to untreated cells of duplicate samples. Genes exhibiting at least a twofold change in expression are displayed in the clustergram. Genes were classified into the RP and Ribi regulons as described previously (Jorgensen et al. 2004a). Effects on NCR and glycolytic gene clusters are shown to the right. (B) Overexpression of MRS6 causes repression of the Ribi, RP, and glycolytic regulons. A strain bearing a GAL1-MRS6 plasmid was induced in galactose medium for 30, 60, and 120 min and processed as in A. Duplicate experiments are shown. (C) TORC1 regulation of the Sfp1–Mrs6 interaction. Competition between Sfp1 and Rab GTPases for Mrs6 may couple activity of the secretory system to ribosome biogenesis. TORC1 localizes to various elements of the endomembrane system. Membranous structures are shown in light blue, Rab GTPases are shown in orange, TORC1 activators and effectors are shown in steel blue. Not all Rab GTPases are shown; other membrane-associated small GTPases that activate TOR are also not shown. (ER) Endoplasmic reticulum; (PM) plasma membrane; (Endo) endosome; (Rap) rapamycin.
Discussion
Our systematic genetic screens recovered many critical secretory factors as determinants of Sfp1 localization. The identification of Mrs6 as a cytoplasmic tether for Sfp1 under poor nutrient conditions establishes a new mechanism that links the secretory system and ribosome biogenesis. Importantly, the effects of Mrs6 on Sfp1 are completely independent from the known PKC-dependent mechanism that indirectly monitors and transmits secretory defects to RP, tRNA, and rDNA transcription (Nierras and Warner 1999; Li et al. 2000). Mrs6 is a powerful cytoplasmic anchor that is both necessary and sufficient for retention of Sfp1 in the cytoplasm, regardless of nutrient status or signaling. Moreover, Mrs6 specifically controls Sfp1 and does not affect the localization of other known TOR effectors or the gene expression programs that are regulated by these effectors. The properties of rapamycin-resistant MRS6-R alleles demonstrate that the Sfp1 anchor function is separable from essential Mrs6 roles and that the Sfp1–Mrs6 interaction is dictated primarily by TOR activity. These findings elaborate the TOR paradigm in yeast and have potential ramifications for TOR signaling in metazoans.
Mrs6 integrates secretory flux, TOR signaling, and ribosome biogenesis
The regulation of the Sfp1–Mrs6 interaction by nutrient and stress conditions parallels known effects of these conditions on Ribi gene expression (Jorgensen et al. 2004a). Given that MRS6 overexpression is dominant over nutrient signaling pathways, it seems likely that Mrs6 itself is a limiting target for such signals and that the interaction equilibrium with Sfp1 may be readily shifted by changes in Mrs6 pools. This model is consistent with the rapid dynamics of Sfp1 relocalization in response to changes in nutrient status (Jorgensen et al. 2004a). Although the total cellular abundance of Mrs6 appears to greatly exceed that of Sfp1 (Ghaemmaghami et al. 2003; Lempiainen et al. 2009), the affinity of the interaction has undoubtedly been tuned by evolution to allow the nuclear distribution of Sfp1 to be controlled by both TORC1 signaling and direct competition with Rab GTPases. The location of the MRS6-R amino acid substitutions in the Rab-binding domain suggests that the Rab GTPases and Sfp1 compete for closely juxtaposed binding sites on Mrs6.
Because the Rab GTPases are critical mediators of vesicle compartment identity and directional flow in the secretory system (Novick et al. 1981; Grosshans et al. 2006), competition with Sfp1 for Mrs6 could serve to directly couple secretory flux to the rate of ribosome biogenesis. Mrs6 is ideally positioned to sense vesicle flux by virtue of its essential interactions with the Rab GTPases. Indeed, many of the secretory system genes isolated in our screens for altered Sfp1 localization directly or indirectly affect Rab GTPase activity, including GAPs, ER/Golgi trafficking complexes, GPI anchor/glycoprotein synthesis factors, and various SNARE components (Table 1). Perturbation of Rab GTPase recycling by secretory defects or stress would liberate Mrs6 to sequester Sfp1 and thereby attenuate ribosome biogenesis. Mrs6 may thus link the output of one form of growth—namely, secretion—to the input of another, ribosome biogenesis (Fig. 6C).
In addition to its potential role as a sensor of secretory system flux, the Mrs6–Sfp1 complex is a direct target of TORC1 activity, as demonstrated by both the physical interactions between Mrs6, Sfp1, and TORC1 components and the specific effects of MRS6-R alleles on Sfp1 localization and the response to rapamycin. Recently, it has been shown that Sfp1 is phosphorylated in vivo and in vitro by Tor1; Sfp1 phosphorylation in vivo depends in part on Mrs6 and is necessary for efficient nuclear localization of Sfp1 in glucose medium (Lempiainen et al. 2009). In contrast to our results, however, Lempiainen et al. (2009) suggest that the Sfp1–Mrs6 interaction is not regulated by nutrients and that Mrs6 is a positive regulator of Sfp1 nuclear import. These discrepancies may arise from extraction conditions (Supplemental Fig. 6) and/or the different nature of the mrs6 and sfp1 alleles used in each study. Whether or not phosphorylation of Mrs6 and/or other associated factors by TORC1 influences Sfp1 interactions remains to be determined. Similarly, the mechanism whereby PKA activity controls the Sfp1–Mrs6 interaction and/or Sfp1 localization also remains to be resolved (Jorgensen et al. 2004a). Regardless of these details, because alterations in Mrs6 levels can override nutrient signals, Mrs6 represents a gatekeeper for Sfp1-dependent regulation of ribosome biogenesis.
GTPases and endomembrane platforms for TOR signaling
Substantial evidence indicates that TORC1 signaling complexes are assembled on internal membrane platforms, such that regulation occurs at the level of localization rather than kinase activation (Dechant and Peter 2008; Rohde et al. 2008). Internal membrane dynamics are governed by a cohort of small GTPases that direct myriad vesicle fusion events. In metazoans, the Rheb class of GTPases plays a dedicated proximal role in TOR activation and the Tsc1/Tsc2 GAP proteins that inhibit Rheb activity are subject to a host of positive and negative signals from upstream kinases (Wang and Proud 2009). In parallel, critical amino acid inputs into TOR are transduced by the Rag class of GTPases, which recruit TORC1 to a Rab7 GTPase-positive perinuclear compartment where the Rheb GTPases reside (Kim et al. 2008; Sancak et al. 2008). In yeast, there is no evidence that the Rheb equivalent activates TORC1 and there is no apparent Tsc1/2 ortholog; importantly, however, the Rag orthologs Gtr1 and Gtr2 are linked to amino acid sensing through sorting of the Gap1 amino acid permease, which is a TORC1-dependent process (Gao and Kaiser 2006). The identification of the Rab escort protein Mrs6 as a cofactor for TORC1-mediated control of Sfp1 provides a further connection between membrane-associated GTPases and TOR signaling.
Many other connections between membrane processes and TOR have been established recently. Synthetic lethal screens have revealed functional interactions between Tor1 and a number of vacuolar sorting steps, including the EGO/GSE and class C Vps complexes (Zurita-Martinez et al. 2007). Physical and functional connections between the vesicle trafficking system, the actin cytoskeleton, and TORC1/2 also have been reported (Aronova et al. 2007). Other TOR–membrane interactions in yeast include a role for vesicle trafficking in Gln3 relocalization (Puria et al. 2008), the requirement for the Golgi Ca2+/Mn2+ ATPase Pmr1 in TOR signaling (Devasahayam et al. 2006), the membrane association of inactive Tap42 complexes (Yan et al. 2006), and the association of Sch9 with the vacuole in rich nutrient conditions (Jorgensen et al. 2004a; Urban et al. 2007). In addition, a glucose-dependent interaction between the ER-resident dolichol mannose synthase Dpm1 and the phosphatidylinositol-4-phosphate phosphatase Sac1 appears to coordinate secretory capacity with nutrient conditions (Faulhammer et al. 2005).These observations all support the general notion that nutrient sensing occurs at the membrane–cytosol interface (De Virgilio and Loewith 2006; Rohde et al. 2008). The new functional and physical links between Sfp1, Mrs6, the TORC1 complex, and the secretory system elaborate the concept that TOR-dependent growth signals are coordinated across many different endomembrane platforms. How the activity of these spatially diverse TOR signaling complexes is coordinated to achieve balanced growth remains an outstanding systems-level problem.
Specificity of TOR effectors and coordination of growth control
In yeast, TORC1 coordinates cellular responses through two main effector trunks (Rohde et al. 2008). In one trunk, the transcriptional subprograms that enable nutrient and stress adaptation are governed by the Tap42-dependent localization of various transcription factors (Duvel et al. 2003). In a parallel trunk, ribosome production and protein translation are regulated by Sch9, Sfp1, and TOR-dependent phosphorylation of RNA Pol I-, Pol II-, and Pol III-associated factors (Jorgensen et al. 2004a; Li et al. 2006; Mayer and Grummt 2006; Urban et al. 2007). Sfp1 and Sch9 appear to specify at least partly nonoverlapping aspects of ribosome biogenesis, as indicated by the synthetic lethality of the sfp1Δ sch9Δ double mutant (Jorgensen et al. 2004a). Although the transcriptional programs controlled by each are clearly interdependent, Sfp1 preferentially activates the Ribi regulon, whereas Sch9 has a stronger effect on the RP regulon (Jorgensen et al. 2004a). Sfp1 and Sch9 also associate with distinct membrane environments—the secretory compartment in the case of Sfp1, and the vacuole in the case of Sch9. Finally, as we showed here, the different branches rely on different feedback mechanisms—the Sfp1 branch on Mrs6, and the secretory system and Sch9/RP gene branch on the PKC/cell wall stress pathway. These branches are not completely insulated, as shown by the suppression of mrs6 growth defects by increased dosage of the upstream PKC activator SLG1/WSC1 (Bialek-Wyrzykowska et al. 2000) and by potential feedback effects of Mrs6 on Sch9 phosphorylation (Lempiainen et al. 2009). The spatial and regulatory segregation of different aspects of ribosome biogenesis may enable more efficient anticipation of and/or adaptation to changing environmental conditions (Levy et al. 2007).
Conservation of growth control architecture
The overall conservation of the TOR system from yeast to humans attests to both its centrality and adaptability. The necessary coordination of membrane growth with macromolecular synthesis was undoubtedly a prime evolutionary impetus for the TOR signaling system. The TORC1 and TORC2 complexes themselves, coactivators such as the class C Vps complex and the Gtr1/Gtr2 Rag family GTPases, and effectors such as S6K and Sch9 all appear well conserved. Sfp1 itself shares a number of general attributes with the Myc transcription factor, including the regulation of ribosome biogenesis genes and overt effects on cell size and cell growth (Jorgensen et al. 2004a; Cook and Tyers 2007); however, only tentative connections have been made between Myc and TOR in metazoans (Ravitz et al. 2007; Teleman et al. 2008). Significantly, the trans-Golgi network component GOLPH3 has been identified recently as an oncogene in many human cancers, and as an activator of TOR signaling and a regulator of cell size (Scott et al. 2009). Finally, the human Rab escort proteins REP1/CHM and REP2/CHML share ∼50% sequence identity with Mrs6 (Alory and Balch 2003). Notably, REP1/CHM is the causative hereditary mutation in choroideremia, a disease of retinal degeneration and vision loss (Seabra et al. 1993). The identification of Mrs6 as a key regulator of Sfp1 in yeast raises the possibility that REP1/REP2 may link TOR signaling to the secretory system in higher organisms, with implications for human disease.
Materials and methods
Yeast strains, plasmids, and reagents
Yeast strains and plasmids used in this study are listed in Supplemental Tables 3 and 4. All strains are derivative of S288c. Standard molecular genetic techniques, growth conditions, and inhibitor concentrations were used (Jorgensen et al. 2004a). To avoid confounding nutrient depletion during cell harvest, all cultures were rapidly concentrated by low-speed centrifugation. Carbon starvation was in 0.02% glucose; for various other stresses, log-phase cultures were treated with 200 ng/mL rapamycin, 0.30 mM H2O2, or 2 μg/mL tunicamycin. Size distributions were acquired with a Beckman-Coulter Z2 Channelizer (Jorgensen et al. 2002). Protein fluorescence was visualized on a Nikon Eclipse E600FN microscope equipped with an Hamamatsu Orca II CCD camera. Random PCR mutagenesis was used to generate a collection of MRS6 alleles, which were selected for growth on 1 μg/mL rapamycin and then examined for failure to localize Sfp1GFP to the cytoplasm in 200 ng/mL rapamycin. Microarray analysis was performed on total RNA by the University Health Network Micorarray Center as described previously (Jorgensen et al. 2002). Average values of normalized log2 ratios of duplicate samples were calculated. All raw microarray data is provided in Supplemental Table 2.
Microscopic screen for regulators of SFP1 localization
The SGA technique (Tong et al. 2001) was used to mate an sfp1∷SFP1GFP reporter into a GAL1-ORF fusion strain collection (Sopko et al. 2006) and into a tet-controlled essential strain collection (Mnaimneh et al. 2004). Saturated liquid cultures were used to inoculate 100 μL of SC-glucose medium in Evotec 384-well flat-bottom μClear plates. After overnight growth to log phase, cells were imaged on an Evotec Opera automated spinning-disc confocal microplate imaging microscope (Perkin Elmer). Immediately following the first imaging pass, the plates were quickly centrifuged at low speed, and cells were washed once and resuspended in SC-glycerol medium, and then incubated for 40 min prior to reimaging. For tetracycline-repressible promoter strains, cells were grown for 16–18 h in the presence of 10 μg/mL doxycycline. Fluorescence signal upon excitation by a 488-nm laser light source was detected through a 525 ± 50-nm filter. Three fields of view were collected per well using a 12-bit Peltier-cooled CCD camera with a 60× water-immersible objective. Images were processed with the Acapella language, using the nuclei detection procedure script to quantitate signal area and intensity. Putative hits were identified on the basis of signal area and cell count, visually inspected, and individually confirmed by direct fluorescence microscopy. Cytoplasmic localization of Sfp1 in glucose medium was determined by calculating the total fluorescence signal over the entire area of the cytoplasm; this parameter was denoted as cell area. The area ratio in glucose was defined as the ratio between the cell area signal of the designated allele with respect to the wild-type cell area signal. Nuclear retention of Sfp1 in glycerol was determined by calculating the number of cells that had an area of fluorescence equivalent to the average nuclear area of fluorescence in a wild-type cell grown on glucose medium. The percent nuclear signal in glycerol was defined as the ratio between the number of distinct nuclear areas detected in the total cell population versus total cell count. False positives scored as mislocalized in glycerol arose from either residual glucose present in the microtiter plate well or misrecognition of punctuate cytoplasmic staining as nuclei. False positive small (whi) and large (lge) mutants were eliminated by concurrently small or large cell and nuclear area signals, respectively. Quantitative values for all genes in each screen are provided in Supplemental Table 1.
Protein analysis
Cell cultures were rapidly harvested, disrupted in lysis buffer (10 mM HEPES-KOH at pH 7.9, 50 mM KCl, 1.5 mM MgCl2, 1 mm EDTA, 0.5 mM DTT, protease inhibitor cocktail) and cleared by centrifugation at 15,000g. Immunoprecipitation and immunoblot analysis were carried out as described previously (Jorgensen et al. 2004a); some TORC1 coimmunoprecipitation experiments were carried out at a salt concentration of 600 mM. For subcellular fractionation, total cell lysate was fractionated at 100,000g for 45 min into soluble (S100) and membrane pellet (P100) fractions. The P100 fraction was resuspended in lysis buffer with or without 1 M NaCl and/or 1% Triton-X100, incubated for 30 min on ice, refractionated into soluble and insoluble fractions, and analyzed by immunoblot. For mass spectrometric analysis, a wild-type strain bearing a GAL1-SFP1FLAG plasmid was grown to early log phase in rich medium containing 2% w/v raffinose, induced with 2% w/v galactose for 2 h, and then resuspended and incubated for an additional 40 min in rich medium containing either 2% w/v glucose or 2% w/v glycerol prior to harvest. After lysis, protein complexes were captured on mouse anti-Flag M2 agarose affinity resin (Sigma), washed, and eluted with Flag peptide. Eluted proteins were precipitated with TCA, washed with acetone, and resolved by SDS-PAGE. Protein species were visualized by colloidal blue stain, excised, digested with trypsin, and analyzed by LC-MS/MS on a Thermo-Finnigan LTQ mass spectrometer essentially as described (Ho et al. 2002). In vivo protein interactions were detected using a PCA assay as described (Tarassov et al. 2008).
Acknowledgments
We are particularly grateful to Brett Larson for assistance with mass spectrometry and Harri Lempiainen and David Shore for critical discussions and sharing their unpublished data. We thank Marc Angeli, Zhen Lin, Vivian Nguyen, Danielle Dewar, Hille Tekotte and Ashton Breitkreutz for technical assistance; John Rohde, Angelika Amon, Paul Jorgensen, Liz Conibear, John Aitchison, Jeff Sharom, and Mike Cook for helpful comments; and Stephen Michnick, David Botstein, Brenda Andrews, and Jon Warner for providing reagents. This work was supported by grants to M.T. from the Canadian Institutes of Health Research (MOP-62830) and the Wellcome Trust. M.T. is a Research Professor of the Scottish Universities Life Sciences Alliance as supported by the Scottish Funding Council, and is also supported by a Royal Society Wolfson Research Merit Award.
Footnotes
Article is online at http://www.genesdev.org/cgi/doi/10.1101/gad.1804409.
Supplemental material is available at http://www.genesdev.org.
References
- Alory C, Balch WE. Molecular evolution of the Rab-escort-protein/guanine-nucleotide-dissociation-inhibitor superfamily. Mol Biol Cell. 2003;14:3857–3867. doi: 10.1091/mbc.E03-04-0227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aronova S, Wedaman K, Anderson S, Yates J, III, Powers T. Probing the membrane environment of the TOR kinases reveals functional interactions between TORC1, actin, and membrane trafficking in Saccharomyces cerevisiae. Mol Biol Cell. 2007;18:2779–2794. doi: 10.1091/mbc.E07-03-0274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beck T, Hall MN. The TOR signalling pathway controls nuclear localization of nutrient-regulated transcription factors. Nature. 1999;402:689–692. doi: 10.1038/45287. [DOI] [PubMed] [Google Scholar]
- Bialek-Wyrzykowska U, Bauer BE, Wagner W, Kohlwein SD, Schweyen RJ, Ragnini A. Low levels of Ypt protein prenylation cause vesicle polarization defects and thermosensitive growth that can be suppressed by genes involved in cell wall maintenance. Mol Microbiol. 2000;35:1295–1311. doi: 10.1046/j.1365-2958.2000.01782.x. [DOI] [PubMed] [Google Scholar]
- Budovskaya YV, Stephan JS, Deminoff SJ, Herman PK. An evolutionary proteomics approach identifies substrates of the cAMP-dependent protein kinase. Proc Natl Acad Sci. 2005;102:13933–13938. doi: 10.1073/pnas.0501046102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cardenas ME, Cutler NS, Lorenz MC, Di Como CJ, Heitman J. The TOR signaling cascade regulates gene expression in response to nutrients. Genes & Dev. 1999;13:3271–3279. doi: 10.1101/gad.13.24.3271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cipollina C, van den Brink J, Daran-Lapujade P, Pronk JT, Porro D, de Winde JH. Saccharomyces cerevisiae SFP1: At the crossroads of central metabolism and ribosome biogenesis. Microbiology. 2008;154:1686–1699. doi: 10.1099/mic.0.2008/017392-0. [DOI] [PubMed] [Google Scholar]
- Cook M, Tyers M. Size control goes global. Curr Opin Biotechnol. 2007;18:341–350. doi: 10.1016/j.copbio.2007.07.006. [DOI] [PubMed] [Google Scholar]
- Dascher C, Ossig R, Gallwitz D, Schmitt HD. Identification and structure of four yeast genes (SLY) that are able to suppress the functional loss of YPT1, a member of the RAS superfamily. Mol Cell Biol. 1991;11:872–885. doi: 10.1128/mcb.11.2.872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dechant R, Peter M. Nutrient signals driving cell growth. Curr Opin Cell Biol. 2008;20:678–687. doi: 10.1016/j.ceb.2008.09.009. [DOI] [PubMed] [Google Scholar]
- Devasahayam G, Ritz D, Helliwell SB, Burke DJ, Sturgill TW. Pmr1, a Golgi Ca2+/Mn2+-ATPase, is a regulator of the target of rapamycin (TOR) signaling pathway in yeast. Proc Natl Acad Sci. 2006;103:17840–17845. doi: 10.1073/pnas.0604303103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Virgilio C, Loewith R. Cell growth control: Little eukaryotes make big contributions. Oncogene. 2006;25:6392–6415. doi: 10.1038/sj.onc.1209884. [DOI] [PubMed] [Google Scholar]
- Dubouloz F, Deloche O, Wanke V, Cameroni E, De Virgilio C. The TOR and EGO protein complexes orchestrate microautophagy in yeast. Mol Cell. 2005;19:15–26. doi: 10.1016/j.molcel.2005.05.020. [DOI] [PubMed] [Google Scholar]
- Duvel K, Santhanam A, Garrett S, Schneper L, Broach JR. Multiple roles of Tap42 in mediating rapamycin-induced transcriptional changes in yeast. Mol Cell. 2003;11:1467–1478. doi: 10.1016/s1097-2765(03)00228-4. [DOI] [PubMed] [Google Scholar]
- Faulhammer F, Konrad G, Brankatschk B, Tahirovic S, Knodler A, Mayinger P. Cell growth-dependent coordination of lipid signaling and glycosylation is mediated by interactions between Sac1p and Dpm1p. J Cell Biol. 2005;168:185–191. doi: 10.1083/jcb.200407118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujimura K, Tanaka K, Nakano A, Toh-e A. The Saccharomyces cerevisiae MSI4 gene encodes the yeast counterpart of component A of Rab geranylgeranyltransferase. J Biol Chem. 1994;269:9205–9212. [PubMed] [Google Scholar]
- Gao M, Kaiser CA. A conserved GTPase-containing complex is required for intracellular sorting of the general amino-acid permease in yeast. Nat Cell Biol. 2006;8:657–667. doi: 10.1038/ncb1419. [DOI] [PubMed] [Google Scholar]
- Ghaemmaghami S, Huh WK, Bower K, Howson RW, Belle A, Dephoure N, O'Shea EK, Weissman JS. Global analysis of protein expression in yeast. Nature. 2003;425:737–741. doi: 10.1038/nature02046. [DOI] [PubMed] [Google Scholar]
- Grosshans BL, Ortiz D, Novick P. Rabs and their effectors: Achieving specificity in membrane traffic. Proc Natl Acad Sci. 2006;103:11821–11827. doi: 10.1073/pnas.0601617103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hardwick JS, Kuruvilla FG, Tong JK, Shamji AF, Schreiber SL. Rapamycin-modulated transcription defines the subset of nutrient-sensitive signaling pathways directly controlled by the Tor proteins. Proc Natl Acad Sci. 1999;96:14866–14870. doi: 10.1073/pnas.96.26.14866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho Y, Gruhler A, Heilbut A, Bader GD, Moore L, Adams SL, Millar A, Taylor P, Bennett K, Boutilier K, et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature. 2002;415:180–183. doi: 10.1038/415180a. [DOI] [PubMed] [Google Scholar]
- Jorgensen P, Tyers M. How cells coordinate growth and division. Curr Biol. 2004;14:R1014–R1027. doi: 10.1016/j.cub.2004.11.027. [DOI] [PubMed] [Google Scholar]
- Jorgensen P, Nishikawa JL, Breitkreutz BJ, Tyers M. Systematic identification of pathways that couple cell growth and division in yeast. Science. 2002;297:395–400. doi: 10.1126/science.1070850. [DOI] [PubMed] [Google Scholar]
- Jorgensen P, Rupes I, Sharom JR, Schneper L, Broach JR, Tyers M. A dynamic transcriptional network communicates growth potential to ribosome synthesis and critical cell size. Genes & Dev. 2004a;18:2491–2505. doi: 10.1101/gad.1228804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jorgensen P, Tyers M, Warner JR. Forging the factory: Ribosome synthesis and growth control in budding yeast. Cell growth: Control of cell size, Cold Spring Harbor Monograph Series 42. In: Hall MN, et al., editors. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, N.Y: 2004b. pp. 329–370. [Google Scholar]
- Kim E, Goraksha-Hicks P, Li L, Neufeld TP, Guan KL. Regulation of TORC1 by Rag GTPases in nutrient response. Nat Cell Biol. 2008;10:935–945. doi: 10.1038/ncb1753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeili A, Wedaman KP, O'Shea EK, Powers T. Mechanism of metabolic control. Target of rapamycin signaling links nitrogen quality to the activity of the Rtg1 and Rtg3 transcription factors. J Cell Biol. 2000;151:863–878. doi: 10.1083/jcb.151.4.863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lempiainen H, Uotila A, Urban J, Dohnal I, Ammerer G, Loewith R, Shore D. Sfp1 interaction with TORC1 and Mrs6 reveals feedback regulation on TOR signaling. Mol Cell. 2009;33:704–716. doi: 10.1016/j.molcel.2009.01.034. [DOI] [PubMed] [Google Scholar]
- Levy S, Ihmels J, Carmi M, Weinberger A, Friedlander G, Barkai N. Strategy of transcription regulation in the budding yeast. PLoS One. 2007;2:e250. doi: 10.1371/journal.pone.0000250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li B, Warner JR. Mutation of the Rab6 homologue of Saccharomyces cerevisiae, YPT6, inhibits both early Golgi function and ribosome biosynthesis. J Biol Chem. 1996;271:16813–16819. doi: 10.1074/jbc.271.28.16813. [DOI] [PubMed] [Google Scholar]
- Li Y, Moir RD, Sethy-Coraci IK, Warner JR, Willis IM. Repression of ribosome and tRNA synthesis in secretion-defective cells is signaled by a novel branch of the cell integrity pathway. Mol Cell Biol. 2000;20:3843–3851. doi: 10.1128/mcb.20.11.3843-3851.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Tsang CK, Watkins M, Bertram PG, Zheng XF. Nutrient regulates Tor1 nuclear localization and association with rDNA promoter. Nature. 2006;442:1058–1061. doi: 10.1038/nature05020. [DOI] [PubMed] [Google Scholar]
- Loewith R, Jacinto E, Wullschleger S, Lorberg A, Crespo JL, Bonenfant D, Oppliger W, Jenoe P, Hall MN. Two TOR complexes, only one of which is rapamycin sensitive, have distinct roles in cell growth control. Mol Cell. 2002;10:457–468. doi: 10.1016/s1097-2765(02)00636-6. [DOI] [PubMed] [Google Scholar]
- Marion RM, Regev A, Segal E, Barash Y, Koller D, Friedman N, O'Shea EK. Sfp1 is a stress- and nutrient-sensitive regulator of ribosomal protein gene expression. Proc Natl Acad Sci. 2004;101:14315–14322. doi: 10.1073/pnas.0405353101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayer C, Grummt I. Ribosome biogenesis and cell growth: mTOR coordinates transcription by all three classes of nuclear RNA polymerases. Oncogene. 2006;25:6384–6391. doi: 10.1038/sj.onc.1209883. [DOI] [PubMed] [Google Scholar]
- Miaczynska M, Lorenzetti S, Bialek U, Benito-Moreno RM, Schweyen RJ, Ragnini A. The yeast Rab escort protein binds intracellular membranes in vivo and in vitro. J Biol Chem. 1997;272:16972–16977. doi: 10.1074/jbc.272.27.16972. [DOI] [PubMed] [Google Scholar]
- Mizuta K, Warner JR. Continued functioning of the secretory pathway is essential for ribosome synthesis. Mol Cell Biol. 1994;14:2493–2502. doi: 10.1128/mcb.14.4.2493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mnaimneh S, Davierwala AP, Haynes J, Moffat J, Peng WT, Zhang W, Yang X, Pootoolal J, Chua G, Lopez A, et al. Exploration of essential gene functions via titratable promoter alleles. Cell. 2004;118:31–44. doi: 10.1016/j.cell.2004.06.013. [DOI] [PubMed] [Google Scholar]
- Nierras CR, Warner JR. Protein kinase C enables the regulatory circuit that connects membrane synthesis to ribosome synthesis in Saccharomyces cerevisiae. J Biol Chem. 1999;274:13235–13241. doi: 10.1074/jbc.274.19.13235. [DOI] [PubMed] [Google Scholar]
- Nobukuni T, Joaquin M, Roccio M, Dann SG, Kim SY, Gulati P, Byfield MP, Backer JM, Natt F, Bos JL, et al. Amino acids mediate mTOR/raptor signaling through activation of class 3 phosphatidylinositol 3OH-kinase. Proc Natl Acad Sci. 2005;102:14238–14243. doi: 10.1073/pnas.0506925102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Novick P, Ferro S, Schekman R. Order of events in the yeast secretory pathway. Cell. 1981;25:461–469. doi: 10.1016/0092-8674(81)90064-7. [DOI] [PubMed] [Google Scholar]
- Peyroche A, Antonny B, Robineau S, Acker J, Cherfils J, Jackson CL. Brefeldin A acts to stabilize an abortive ARF–GDP–Sec7 domain protein complex: Involvement of specific residues of the Sec7 domain. Mol Cell. 1999;3:275–285. doi: 10.1016/s1097-2765(00)80455-4. [DOI] [PubMed] [Google Scholar]
- Puria R, Zurita-Martinez SA, Cardenas ME. Nuclear translocation of Gln3 in response to nutrient signals requires Golgi-to-endosome trafficking in Saccharomyces cerevisiae. Proc Natl Acad Sci. 2008;105:7194–7199. doi: 10.1073/pnas.0801087105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravitz MJ, Chen L, Lynch M, Schmidt EV. c-myc repression of TSC2 contributes to control of translation initiation and Myc-induced transformation. Cancer Res. 2007;67:11209–11217. doi: 10.1158/0008-5472.CAN-06-4351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rohde JR, Bastidas R, Puria R, Cardenas ME. Nutritional control via Tor signaling in Saccharomyces cerevisiae. Curr Opin Microbiol. 2008;11:153–160. doi: 10.1016/j.mib.2008.02.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sancak Y, Peterson TR, Shaul YD, Lindquist RA, Thoreen CC, Bar-Peled L, Sabatini DM. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science. 2008;320:1496–1501. doi: 10.1126/science.1157535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott KL, Kabbarah O, Liang MC, Ivanova E, Anagnostou V, Wu J, Dhakal S, Wu M, Chen S, Feinberg T, et al. GOLPH3 modulates mTOR signalling and rapamycin sensitivity in cancer. Nature. 2009;459:1085–1090. doi: 10.1038/nature08109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seabra MC, Brown MS, Goldstein JL. Retinal degeneration in choroideremia: Deficiency of rab geranylgeranyl transferase. Science. 1993;259:377–381. doi: 10.1126/science.8380507. [DOI] [PubMed] [Google Scholar]
- Shamji AF, Kuruvilla FG, Schreiber SL. Partitioning the transcriptional program induced by rapamycin among the effectors of the Tor proteins. Curr Biol. 2000;10:1574–1581. doi: 10.1016/s0960-9822(00)00866-6. [DOI] [PubMed] [Google Scholar]
- Sopko R, Huang D, Preston N, Chua G, Papp B, Kafadar K, Snyder M, Oliver SG, Cyert M, Hughes TR, et al. Mapping pathways and phenotypes by systematic gene overexpression. Mol Cell. 2006;21:319–330. doi: 10.1016/j.molcel.2005.12.011. [DOI] [PubMed] [Google Scholar]
- Sturgill TW, Cohen A, Diefenbacher M, Trautwein M, Martin DE, Hall MN. TOR1 and TOR2 have distinct locations in live cells. Eukaryot Cell. 2008;7:1819–1830. doi: 10.1128/EC.00088-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tarassov K, Messier V, Landry CR, Radinovic S, Serna Molina MM, Shames I, Malitskaya Y, Vogel J, Bussey H, Michnick SW. An in vivo map of the yeast protein interactome. Science. 2008;320:1465–1470. doi: 10.1126/science.1153878. [DOI] [PubMed] [Google Scholar]
- Teleman AA, Hietakangas V, Sayadian AC, Cohen SM. Nutritional control of protein biosynthetic capacity by insulin via Myc in Drosophila. Cell Metab. 2008;7:21–32. doi: 10.1016/j.cmet.2007.11.010. [DOI] [PubMed] [Google Scholar]
- Tong AH, Evangelista M, Parsons AB, Xu H, Bader GD, Page N, Robinson M, Raghibizadeh S, Hogue CW, Bussey H, et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science. 2001;294:2364–2368. doi: 10.1126/science.1065810. [DOI] [PubMed] [Google Scholar]
- Urban J, Soulard A, Huber A, Lippman S, Mukhopadhyay D, Deloche O, Wanke V, Anrather D, Ammerer G, Riezman H, et al. Sch9 is a major target of TORC1 in Saccharomyces cerevisiae. Mol Cell. 2007;26:663–674. doi: 10.1016/j.molcel.2007.04.020. [DOI] [PubMed] [Google Scholar]
- Wang X, Proud CG. Nutrient control of TORC1, a cell-cycle regulator. Trends Cell Biol. 2009;19:260–267. doi: 10.1016/j.tcb.2009.03.005. [DOI] [PubMed] [Google Scholar]
- Warner JR. The economics of ribosome biosynthesis in yeast. Trends Biochem Sci. 1999;24:437–440. doi: 10.1016/s0968-0004(99)01460-7. [DOI] [PubMed] [Google Scholar]
- Wedaman KP, Reinke A, Anderson S, Yates J, III, McCaffery JM, Powers T. Tor kinases are in distinct membrane-associated protein complexes in Saccharomyces cerevisiae. Mol Biol Cell. 2003;14:1204–1220. doi: 10.1091/mbc.E02-09-0609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wullschleger S, Loewith R, Hall MN. TOR signaling in growth and metabolism. Cell. 2006;124:471–484. doi: 10.1016/j.cell.2006.01.016. [DOI] [PubMed] [Google Scholar]
- Yan G, Shen X, Jiang Y. Rapamycin activates Tap42-associated phosphatases by abrogating their association with Tor complex 1. EMBO J. 2006;25:3546–3555. doi: 10.1038/sj.emboj.7601239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaman S, Lippman SI, Zhao X, Broach JR. How Saccharomyces responds to nutrients. Annu Rev Genet. 2008;42:27–81. doi: 10.1146/annurev.genet.41.110306.130206. [DOI] [PubMed] [Google Scholar]
- Zerial M, McBride H. Rab proteins as membrane organizers. Nat Rev Mol Cell Biol. 2001;2:107–117. doi: 10.1038/35052055. [DOI] [PubMed] [Google Scholar]
- Zurita-Martinez SA, Puria R, Pan X, Boeke JD, Cardenas ME. Efficient Tor signaling requires a functional class C Vps protein complex in Saccharomyces cerevisiae. Genetics. 2007;176:2139–2150. doi: 10.1534/genetics.107.072835. [DOI] [PMC free article] [PubMed] [Google Scholar]