Abstract
While chemokine and growth factor receptors are attractive and popular targets for cancer therapeutic intervention, structure-based targeting of the ligands themselves is generally not considered practical. New evidence indicates that a notable exception to this is macrophage migration inhibitory factor (MIF). MIF, an autocrine and paracrine acting cytokine/growth factor, plays a pivotal role in both the initiation and maintenance of neoplastic diseases. MIF possesses a non-physiologic enzymatic activity that is evolutionarily well conserved. Although small molecule antagonists of MIFs enzymatic active site have been reported to inhibit biological activities of MIF, universally high IC50s have limited their clinical appeal. Using a computational virtual screening strategy we have identified a unique small molecule inhibitor that serves as a suicide substrate for MIF resulting in the covalent modification of the catalytically active N-terminal proline. Our studies further reveal that this compound, 4-iodo-6-phenylpyrimidine (4-IPP), is ~ 5–10x times more potent in blocking MIF-dependent catalysis and lung adenocarcinoma cell migration and anchorage independent growth than the prototypical MIF inhibitor, ISO-1. Finally, using an in silico combinatorial optimization strategy, we have identified four unique congeners of 4-IPP that exhibit MIF inhibitory activity at concentrations 10–20x lower than that of parental 4-IPP.
Keywords: anchorage-independence, cancer, enzyme, inhibitor, migration, tautomerase
Introduction
Macrophage migration inhibitory factor (MIF) is a pleiotropic cytokine/growth factor that contributes to inflammatory, autoimmune and malignant disease pathologies. MIF exerts at least some of its biological actions through binding to its extracellular cognate receptor complex consisting of CD74 and CD44 (1, 2). Studies from this laboratory have revealed a requirement for extracellular MIF in the steady state activation of Rho GTPase family members leading to cell growth and migratory phenotypes (3, 4). These MIF-mediated effects are thought to be linked to CD74/CD44-initiated signaling given the fact that CD44-dependent signaling has been shown to promote Rho GTPase family member activation (5).
MIF possesses the unusual ability to catalyze the tautomerization of the non-physiological substrates D-dopachrome and L-dopachrome methyl ester into their corresponding indole derivatives (6). The N-terminal proline of MIF (Pro-1) appears to be a critical residue for enzymatic activity, as site-directed mutagenesis that substitutes a serine or glycine for this proline (P1G) is devoid of tautomerase activity (7). The physiologic role of MIFs catalytic activity remains in question, however, because of conflicting reports of loss or gain of biologic activity with various proline mutants (7, 8).
Of the MIF enzyme/biological antagonists reported, ISO-1 (S,R-3-(4-hydroxyphenyl)-4,5-dihydro-5-isoxazole acetic acid methyl ester), is the best characterized (9, 10). While other compounds have been shown to have improved biologic inhibitory activities against MIF (11, 12), ISO-1 is considered to be the prototypical MIF antagonist. We and others have reported that this compound is as effective in blocking MIF-dependent malignant phenotypes as MIF knockdown by siRNA (4, 13).
We now report on a unique virtual screening strategy that identified a novel suicide substrate for MIF. Our studies indicate that this compound is ~ 5–10x more potent than the best known MIF inhibitor in blocking MIF-dependent catalysis, migration and anchorage-independent growth.
Materials and Methods
Cells and reagents
Human A549 lung adenocarcinoma (ATCC, Manassas, VA) cells were grown in DMEM containing 10% FCS, 2 mM glutamine and 50 µg/ml gentamycin (Invitrogen, Carlsbad, CA). RNA interference transfections, migration assays and soft agar colony formation experiments were performed as previously described (4). Candidate MIF inhibitor compounds were purchased from Chemnavigator (San Diego, CA), 4-iodo-6-phenylpyrimidine was purchased from Specs (Wakefield, RI), 6-phenylpyrimidine was from Sigma-Aldrich (St. Louis, MO) and ISO-1 was a generous gift from Dr. Yousef Al-Abed (Feinstein Institute for Medical Research, Manhasset, NY).
Virtual Screening
SDF formatted files from the ACD (MDL) library were transformed using Genfra (Accelrys, San Diego, CA) obtaining a library of 343,802 structures suitable for the virtual screening software Ludi (Accelrys). Ludi considers hydrogen-bond donors and acceptors and aliphatic nature (parameters: 8 Å radius with no linkage, or rotatable bonds, 5000 fits, 300 lowest hits to be stored, aliphatic_aromatic and electrostatic_check options). The first three energy functions were used and a manual consensus score ranked the lists.
Enzymatic analyses
MIF enzymatic activity in cell lysates and liver tissue was carried out by lysing 1 × 107 A549 cells or ~ 1 gram pieces of liver in PBS containing 1 mM NaVO4, 2 mM NaF and a protease inhibitor cocktail (Roche Biochemical, Indianapolis, IN) using dounce-homogenization on ice. rMIF (16), cell lysate or liver lysate was added to a final volume of 700 µl PBS in plastic cuvettes. For rMIF inhibition assays, compounds were added to the rMIF/PBS solution and pre-incubated for 5 minutes. 4 mM L-3,4-dihydroxyphenylalanine methyl ester and 8 mM sodium periodate (Sigma-Aldrich) were combined in a 3:2 ratio to form L-dopachrome methyl ester. 300 µL of L-dopachrome methyl ester was then immediately added to the cuvettes, incubated for an additional 5 min to allow color change which was measured at OD475 nm.
Crystallization and structure determination
0.1 mM recombinant MIF protein (9) was mixed with a 10-fold molar excess of 4-IPP, incubated over night and concentrated to 1 mM protein with a Centricon 3000 M.W. centrifugal filter unit (Millipore). The complex was crystallized using the hanging drop vapour-diffusion method at 20 °C. Equal volumes of the MIF·inhibitor complex and reservoir solution (2M ammonium sulfate, 4% isopropanol, 0.1 M tris, pH 7.5) were mixed to form 2 µl drops. Crystals belonging to space group P212121 formed within 2 weeks. Data were collected in house using a Rigaku RAXIS-IV imaging plate area detector. The crystal was cryoprotected in the reservoir solution containing 25% glycerol. Diffraction data were collected at −180 °C, and HKL2000 (17) was used to integrate and scale the diffraction data to 1.8 Å. The structure was solved by molecular replacement with Phaser (18) using the structure of MIF in complex with Oxim-11 as search model (11). Refinement was performed using CNS and Refmac (19, 20) and manual manipulation of the model was done using MIFit 4.1. The structure is deposited with the RCSB Protein Data Bank and has been assigned the RCSB ID code RCSB045257 and PDB ID code 3B9S.
Mass Spectroscopy
MIF/4-IPP adducts were analyzed by liquid chromatography/mass spectrometry (LC-MS). Samples were run on a C18 (Waters) column attached to an HP1100 LC with auto sampler and Diode Array UV detector (Agilent Technologies, Santa Clara, CA) coupled to a Micromass Quattro LCZ (Waters Corp, Milford, MA) triple quadrupole mass spectrometer, using the electrospray ion source. The LC method used a reversed phase gradient, 10–60% acetonitrile and the mass spectrometer was set up in the electrospray positive-ion mode to scan over the range M/z 200 to 1000. LC peaks containing consecutive M/z scans were combined to produce mass spectra of the analyte, consisting of a series of multiply-charged ions of the intact protein. Spectra were further analyzed using MaxEnt1 feature of MassLynx (Waters Corp, Milford, MA) to determine mass of the proteins from the multiply-charged series of ions.
Results and Discussion
Virtual screening against Met-2 of MIF reveals a novel small molecule inhibitor of MIF
As opposed to prior virtual screens run against MIF that targeted the N-terminal proline, our computational screening strategy focused on targeting methionine at position A2 (A2 refers to monomer A, position 2 from the trimeric crystal structure of MIF). We chose this strategy because; 1) Met-2 resides at the base of the hydrophobic binding pocket while the N-terminal proline is situated on the side of the pocket (14); and, 2) prior studies have shown that disrupting this hydrophobic substrate-binding pocket by insertion of a single amino acid residue adjacent to Met-2 leads to a complete loss of enzymatic and biologic activity (9).
Our initial virtual screen utilized the crystal structure of MIF (PDB - 1MIF) and the Available Chemicals Directory library, the results of which provided a ranking list of the top 100 compounds. Of the 76 commercially available compounds obtained from this list, only 41 were found to be soluble in aqueous solutions at 100 µM concentrations. Of these, 9 were found to be inhibitory at concentrations of 50 µM or less to the catalytic activity of purified, recombinant MIF (22.0% success rate; 9/41). When these compounds were tested against MIF-dependent catalytic activity in whole cell lysates, we found that only three compounds remained inhibitory at similar IC50s. Of these three remaining compounds only one was found to exhibit superior MIF catalytic inhibitory activity versus ISO-1: 4-iodo-6-phenypyrimidine (4-IPP) (Fig. 1A). 4-IPP and ISO-1 were tested for their relative inhibitory effects against the catalytic activity of recombinant human MIF. As shown in Fig. 1B, the IC50 of 4-IPP is 10 times lower than that of ISO-1.
FIGURE 1. 4-iodo-6-phenylpyrmidine inhibits MIF dopachrome tautomerase activity.
A. Structure comparison of 4-iodo-6-phenylpyrmidine and ISO-1. B. Indicated final concentrations of 4-IPP, 6-phenylpyrimidine or ISO-1 were pre-incubated with 50 nM MIF in PBS followed by addition of L-dopachrome methyl ester at a final concentration of 0.72 mM. Relative MIF tautomerase activity was assessed by measuring spectrophotometrically at OD475.
4-IPP is a suicide substrate for MIF
Because the kinetics observed with 4-IPP against MIF catalysis were reminiscent of an irreversible inhibitor (not shown), we next tested MIF/4-IPP admixtures to determine whether any shifts in the molecular weight of MIF were observable. As shown in Fig. 2A (top panel), liquid chromatography - mass spectroscopy (LC-MS) analysis revealed that MIF co-incubated with 4-IPP resulted in a mass increase of 154 units. Co-incubation of 4-IPP with a catalytically inactive mutant of MIF (point mutation of Pro-1 to Gly-1) (7), resulted in no covalent modification as assessed by mass spectroscopy (Fig. 2A, bottom panel).
FIGURE 2. 4-IPP covalently modifies MIF.
A. 5 µg wildtype (top panels) or P2G mutant (bottom panels) rMIF was incubated with either DMSO alone (top chromatograms) or 100 µM 4-IPP (bottom chromatograms) for 2 hours in a total volume of 50 µl PBS. Protein adducts were analyzed by LC-MS as described in Materials and Methods. B. Co-crystal structure of MIF:4-IPP with potential interacting residues surrounding the covalently attached phenylpyrimidine. C. Connelly surface of MIF colored by the Poisson Boltzman electrostatic potential with covalently bound 4-IPP in stick representation.
Co-crystallization of MIF and 4-IPP reveals 6-phenylpyrimidine covalently modified catalytic proline
Because 4-iodo-6-phenylpyrimidine is M.W. 282 and the iodine at position 4 of the pyrimidine ring can act as a leaving group, we suspected that the increase in mass of 154 units observed with wildtype rMIF/4-IPP (but not catalytically inactive mutants) was due to a covalent modification of rMIF by de-iodinated 4-IPP giving 6-phenylpyrimidine-modified MIF (M.W. 156 – with covalent attachment, M.W. 154). To test this, we performed co-crystalization of MIF/4-IPP admixtures. As shown in Fig. 2B, 4-IPP binds covalently to MIF at the nitrogen of the N-terminal proline by substitution of the iodo group at the 4-position of 4-IPP (Data collection details are found in Table S1, Supplemental Data). The phenyl substituent is oriented deep in the MIF catalytic pocket near Met-2 (Fig. 2C). This is similar to existing crystal structures of MIF with antagonists 7-hydroxy-2-oxo-chromene-3-carboxylic acid ethyl ester (15) and OXIM6 (11) (PDB entries 1GCZ and 2OOZ, respectively) although the phenyl substituent of bound 4-IPP is located slightly deeper into the binding pocket and in a different orientation. There is some apparent freedom of rotation of the phenyl ring as two different conformations exist in the tri-substituted trimeric MIF crystal structure. The major interaction partners for bound 4-IPP within 2.5 angstroms are MET A2,TYR A36, HIS A62, VAL A106, PHE A113, TYR B95, ASN B97.
Inhibition of MIF-dependent migration and anchorage-independence by 4-IPP
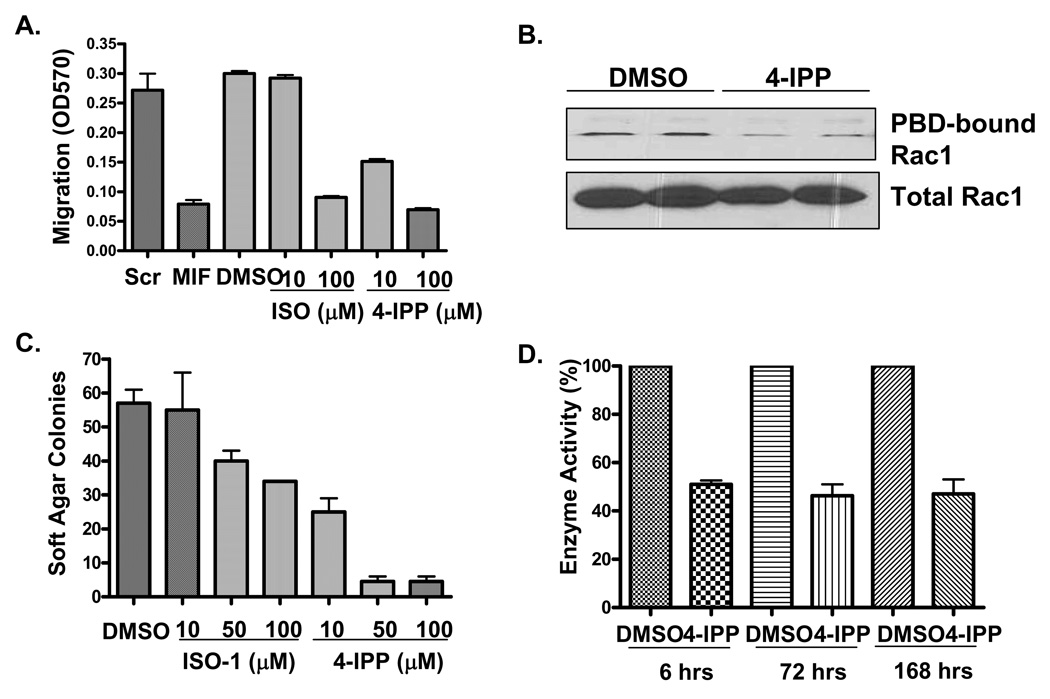
Studies from this (4) and other laboratories (13) reveal that ISO-1 effectively inhibits MIF-dependent cell migration and anchorage-independent growth in human cancer cell lines. As such, we tested 4-IPP against ISO-1 and determined the relative inhibitory effects on the migration of A549 lung adenocarcinoma cells on collagen substrates. Consistent with our recent findings, MIF siRNA strongly reduced the migration of A549 cells (4) (Fig. 3A). Similar to MIF siRNA, pre-treatment of A549 cells with either ISO-1 or 4-IPP at 100 µM reduced A549 motility by greater than 70%. In contrast, 10 µM (Fig. 3A) and 50 µM (not shown) ISO-1 had no effect on A549 migration while 10 µM 4-IPP attenuated migration by greater than 50% that of vehicle alone (Fig. 3A). In order to ensure that this effect was not restricted to A549 cells, H1299 and H23 NSCLC cell lines showed a similar effect with 4-IPP albeit more pronounced (Supplemental Fig. 1). Interestingly, both of these cell lines were largely resistant to ISO-1 at 100 µM while 4-IPP was maximally effective at this concentration. As MIF is thought to promote NSCLC motility in a Rac1-dependent fashion, 4-IPP was tested for its ability to block Rac1 binding to the PAK-binding domain of Pak (PBD) (4). As shown in Fig. 3B, 4-IPP reduced Rac1 binding to PBD similar to what we have observed with siRNA in the same cells.
FIGURE 3. 4-IPP inhibits NSCLC migration and anchorage independent growth.
A. 2 × 105 inhibitor pre-treated and siRNA transfected A549 human lung adenocarcinoma cells were added to the upper chamber of collagen coated transwell chambers. 20 hours later, cells in the upper chamber were removed by swabbing and the bottom of the membrane was stained, washed, solubilized with isopropanol and read spectrophotometrically at OD570. Results are representative of four independent experiments and were performed in duplicate. B. 4 × 105 A549 cells were treated in duplicate with 0.1 % DMSO or 50 µM 4-IPP in fresh media. 16 hours later, corresponding cell lysates were subjected to PAK binding domain (PBD) GST-fusion protein sepharose bead pull downs or were directly assessed for total Rac1 by immunoblotting. Washed PBD-GST pull downs were subjected to western blotting and active Rac1 was detected using anti-Rac1. C. 1 × 105 A549 human lung adenocarcinoma cells were plated in duplicate plates of soft agar containing vehicle (0.1% DMSO) or 10, 50 or 100 µM ISO-1 or 4-IPP. 20 days later, colonies were stained and enumerated. Data are representative of 2 independent experiments. D. Mice were injected intraperitoneally for 6 hours (n = 2) or once daily for 3 (n = 4) or 7 days (n = 3). Mice were weighed both pre- and post-treatment for the 3 and 7 day treatments before being sacrificed and livers resected. After homogenizing tissue, 500 µg total liver protein was assessed for L-dopachrome methyl ester tautomerase activity.
We next tested and compared the relative efficacies of ISO-1 and 4-IPP to inhibit lung adenocarcinoma anchorage-independent growth. As shown in Fig. 3C, both ISO-1 and 4-IPP inhibited A549 soft agar growth, but 4-IPP was between 5–10 fold more potent (similar to IC50 data from Fig. 1A) than ISO-1.
We then tested in vivo inhibition of tissue-associated MIF by 4-iodo-6-phenylpyrimidine. 1 mg (~ 50 mg/kg) injections of 4-iodo-6-phenylpyrimidine inhibited liver MIF enzyme tautomerase activity at 6 hours or for 3 and 7 days (daily injections of 1 mg 4-IPP) by greater than 50% (Fig. 3D). Importantly, we observed no weight loss or obvious toxicity with 1, 2 or 4 mg daily intraperitoneal injections (50, 100 and 200 mg/kg, respectively) of 4-IPP over the course of seven days (Fig. 3D and data not shown). While longer treatment regimens remain to be tested, acute toxicity of 4-IPP appears to be nominal.
In order to identify more potent analogs of our lead MIF antagonist, we performed an in silico combinatorial docking screen using the co-crystal structure of the MIF:4-IPP complex defining MIF as the receptor and the phenylpyrimidine rings of 4-IPP as the core. Four unique congeners of 4-IPP were predicted to have substantially increased binding potential within the MIF substrate binding pocket (Fig. 4A). We subsequently synthesized the four analogs and, as predicted, the IC50s of all four individual analogs were between ~ 10x – 20x more active (IC50s = A1 ~ 200 nM, A2 ~275 nM, A3 ~ 400 nM, A4 ~ 475 nM) than parental 4-IPP in blocking MIF-dependent catalysis (Fig. 4B). Finally, to determine if enhanced enzyme inhibitory activity correlates with enhanced biologic inhibitory activity we tested the most potent of the four analogs against 4-IPP in an in vitro migration assay. As shown in Fig. 4C, A1-4-IPP was substantially more potent in blocking migration than our lead parent compound, 4-IPP.
FIGURE 4. 4-IPP analog optimization improves MIF inhibitor efficacy.
A. Indicated final concentrations of 4-IPP analogs A1–A4 were pre-incubated with 50 nM MIF in PBS followed by addition of L-dopachrome methyl ester at a final concentration of 0.72 mM. Relative MIF tautomerase activity was assessed by measuring spectrophotometrically at OD475. B. A549 lung adenocarcinoma cells were pre-incubated with the indicated concentrations of either 4-IPP or A1-4-IPP for 16 hours. 2 × 105 cells were added to the upper chamber of collagen coated transwell chambers and processed 16 hours later. Results are +/− SD of the averages of duplicate samples.
Using a novel virtual screening strategy, we have discovered a family of suicide substrates of MIF that covalently modify the N-terminal, catalytically active proline of MIF. Our results indicate that 4-iodo-6-phenylpyrimidine is de-halogenated upon contact with enzymatically active, but not inactive, MIF and a subsequent covalent bond then forms between C-4 of pyrimidine and the N-terminal nitrogen of Proline at position 1. Importantly, this covalently modified MIF is rendered inactive in both its catalytic and biologic functions by the suicide substrate, 4-IPP. Our studies further indicate that 4-IPP treatment inhibits the migration and anchorage-independence of human lung adenocarcinoma cell lines in vitro and MIF-associated liver enzyme activity when administered in vivo.
Several small molecule inhibitors of MIF have been described since MIF’s enzymatic activity was first discovered (9, 11, 12, 15). Orita and colleagues first described virtual screening as a tool to identify novel small molecule antagonists of MIF (15). Using the catalytically active N-terminal proline as a target, the authors discovered that SBDD-identified MIF inhibitory compounds could be classified into 4 independent groups. Despite excellent IC50 activities against MIF catalysis of lead compounds identified from this study, no functional testing of these inhibitors has been reported to date.
Our group and others have recently reported that ISO-1 blocks the migration, invasion and anchorage-independent growth of human cancer cell lines (4, 13). As the MIF receptor CD74 has been found to be necessary for anti-tumor activities of ISO-1, it is likely that 4-IPP and it’s more active analogs similarly block MIF/receptor interactions albeit more efficiently than ISO-1.
In summary, we have identified a unique family of small compound inhibitors of MIF that act as suicide substrates to irreversibly inhibit MIF-dependent tumor cell invasive and anchorage-independent properties. While further studies are needed to elucidate the anti-tumor efficacy of this novel lead compound, we propose that 4-iodo-6-phenylpyrimidine represents a unique class of anti-MIF therapeutic antagonists.
Supplementary Material
Abbreviations
- NSCLC
non-small cell lung cancer
- 4-IPP
4-iodo-6-phenylpyrimidine
References
- 1.Leng L, Metz CN, Fang Y, et al. MIF Signal Transduction Initiated by Binding to CD74. J Exp Med. 2003;197:1467–1476. doi: 10.1084/jem.20030286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Shi X, Leng L, Wang T, et al. CD44 is the signaling component of the macrophage migration inhibitory factor-CD74 receptor complex. Immunity. 2006;25:595–606. doi: 10.1016/j.immuni.2006.08.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Swant JD, Rendon BE, Symons M, Mitchell RA. Rho GTPase-dependent signaling is required for macrophage migration inhibitory factor-mediated expression of cyclin D1. J Biol Chem. 2005;280:23066–23072. doi: 10.1074/jbc.M500636200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rendon BE, Roger T, Teneng I, et al. Regulation of human lung adenocarcinoma cell migration and invasion by macrophage migration inhibitory factor. J Biol Chem. 2007;282:29910–29918. doi: 10.1074/jbc.M704898200. [DOI] [PubMed] [Google Scholar]
- 5.Singleton PA, Dudek SM, Ma SF, Garcia JG. Transactivation of sphingosine 1-phosphate receptors is essential for vascular barrier regulation. Novel role for hyaluronan and CD44 receptor family. J Biol Chem. 2006;281:34381–34393. doi: 10.1074/jbc.M603680200. [DOI] [PubMed] [Google Scholar]
- 6.Rosengren E, Bucala R, Aman P, et al. The immunoregulatory mediator macrophage migration inhibitory factor (MIF) catalyzes a tautomerization reaction. Mol Med. 1996;2:143–149. [PMC free article] [PubMed] [Google Scholar]
- 7.Swope M, Sun HW, Blake PR, Lolis E. Direct link between cytokine activity and a catalytic site for macrophage migration inhibitory factor. EMBO J. 1998;17:3534–3541. doi: 10.1093/emboj/17.13.3534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hermanowski-Vosatka A, Mundt SS, Ayala JM, et al. Enzymatically inactive macrophage migration inhibitory factor inhibits monocyte chemotaxis and random migration. Biochemistry. 1999;38:12841–12849. doi: 10.1021/bi991352p. [DOI] [PubMed] [Google Scholar]
- 9.Lubetsky JB, Dios A, Han J, et al. The tautomerase active site of macrophage migration inhibitory factor is a potential target for discovery of novel anti-inflammatory agents. J Biol Chem. 2002;277:24976–24982. doi: 10.1074/jbc.M203220200. [DOI] [PubMed] [Google Scholar]
- 10.Al Abed Y, Dabideen D, Aljabari B, et al. ISO-1 binding to the tautomerase active site of MIF inhibits its pro-inflammatory activity and increases survival in severe sepsis. J Biol Chem. 2005;280:36541–36544. doi: 10.1074/jbc.C500243200. [DOI] [PubMed] [Google Scholar]
- 11.Crichlow GV, Cheng KF, Dabideen D, et al. Alternative chemical modifications reverse the binding orientation of a pharmacophore scaffold in the active site of macrophage migration inhibitory factor. J Biol Chem. 2007;282:23089–23095. doi: 10.1074/jbc.M701825200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dabideen DR, Cheng KF, Aljabari B, Miller EJ, Pavlov VA, Al Abed Y. Phenolic hydrazones are potent inhibitors of macrophage migration inhibitory factor proinflammatory activity and survival improving agents in sepsis. J Med Chem. 2007;50:1993–1997. doi: 10.1021/jm061477+. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Meyer-Siegler KL, Iczkowski KA, Leng L, Bucala R, Vera PL. Inhibition of macrophage migration inhibitory factor or its receptor (CD74) attenuates growth and invasion of DU-145 prostate cancer cells. J Immunol. 2006;177:8730–8739. doi: 10.4049/jimmunol.177.12.8730. [DOI] [PubMed] [Google Scholar]
- 14.Sun HW, Bernhagen J, Bucala R, Lolis E. Crystal structure at 2.6-A resolution of human macrophage migration inhibitory factor. Proc Natl Acad Sci. 1996;93:5191–5196. doi: 10.1073/pnas.93.11.5191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Orita M, Yamamoto S, Katayama N, et al. Coumarin and chromen-4-one analogues as tautomerase inhibitors of macrophage migration inhibitory factor: discovery and X-ray crystallography. J Med Chem. 2001;44:540–547. doi: 10.1021/jm000386o. [DOI] [PubMed] [Google Scholar]
- 16.Bernhagen J, Mitchell RA, Calandra T, Voelter W, Cerami A, Bucala R. Purification, bioactivity, and secondary structure analysis of mouse and human macrophage migration inhibitory factor (MIF) Biochemistry. 1994;33:14144–14155. doi: 10.1021/bi00251a025. [DOI] [PubMed] [Google Scholar]
- 17.Otwinowski Z, Minor W. Processing of X-ray Diffraction Data Collected in Oscillation Mode. In: Carter CW Jr, Sweet RM, editors. Methods in Enzymology, Macromolecular Crystallography. Volume 276. New York: Academic Press; 1997. pp. 307–326. [DOI] [PubMed] [Google Scholar]
- 18.Storoni LC, McCoy AJ, Read RJ. Likelihood-enhanced fast rotation functions. Acta Crystallogr D Biol Crystallogr. 2004;60:432–438. doi: 10.1107/S0907444903028956. [DOI] [PubMed] [Google Scholar]
- 19.Brunger AT, Adams PD, Clore GM, et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D Biol Crystallogr. 1998;54:905–921. doi: 10.1107/s0907444998003254. [DOI] [PubMed] [Google Scholar]
- 20.Murshudov GN, Vagin AA, Dodson EJ. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D Biol Crystallogr. 1997;53:240–255. doi: 10.1107/S0907444996012255. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.