SUMMARY
RecQ DNA helicases act in conjunction with heterologous partner proteins to catalyze DNA metabolic activities that include recombination initiation and stalled replication fork processing. For the prototypical Escherichia coli RecQ protein, direct interaction with single-stranded DNA-binding (SSB) protein stimulates its DNA unwinding activity. Complex formation between RecQ and SSB is mediated by the RecQ winged-helix domain (WH), which binds the nine C-terminal-most residues of SSB, a highly conserved sequence known as the SSB-Ct element. Using nuclear magnetic resonance and mutational analyses, we identify the SSB-Ct binding pocket on E. coli RecQ. The binding site shares striking electrostatic similarity with the previously identified SSB-Ct binding site on E. coli Exonuclease I, although the SSB-binding domains in the two proteins are not otherwise related structurally. Substitutions that alter RecQ residues implicated in SSB-Ct binding impair RecQ binding to SSB and SSB/DNA nucleoprotein complexes. These substitutions also diminish SSB-stimulated DNA helicase activity in the variants, although additional biochemical changes in the RecQ variants indicate a role for the WH domain in helicase activity beyond SSB binding. Sequence changes in the SSB-Ct element are sufficient to abolish interaction with RecQ in the absence of DNA and to diminish RecQ binding and helicase activity on SSB/DNA substrates. These results support a model in which RecQ has evolved an SSB-Ct binding site on its winged-helix domain as an adaptation that aids its cellular functions on SSB/DNA nucleoprotein substrates.
Keywords: RecQ, helicase, single-stranded DNA-binding protein, genome maintenance, protein complex
INTRODUCTION
RecQ DNA helicases are evolutionarily conserved enzymes with diverse roles in genome maintenance 1; 2; 3; 4; 5. In bacteria, RecQ proteins aid in recombination, stalled replication fork processing, SOS signaling, and antigen variation pathways 6; 7; 8; 9; 10; 11; 12. In humans, individual mutation of three of the five RecQ helicase genes (BLM, WRN, and RECQ4) gives rise to Bloom’s, Werner’s, and Rothmund-Thompson syndromes, respectively 13; 14; 15. These conditions are marked by genome instability manifested as chromosome breakage and genomic rearrangements that are linked to cancer predisposition, underscoring the importance of the RecQ family of enzymes in maintaining the integrity of genomic DNA.
RecQ proteins associate with a large number of heterologous proteins, often functioning as components in multi-protein complexes 1; 5; 16; 17; 18. Direct interaction with single-stranded (ss) DNA-binding (SSB) protein (called Replication Protein A, or RPA, in eukaryotes) is a shared feature among bacterial and eukaryotic RecQ proteins. RecQ proteins known to associate with their cognate SSB proteins include E. coli RecQ 19, and human BLM 20, WRN 21, and RecQ1 22; 23. In addition, human RecQ5β is strongly stimulated by, and most likely associates with, RPA 24. Due to the conservation of RecQ interactions with SSB proteins, a physical and functional understanding of how these complexes assemble to act on DNA substrates is critical for appreciating the broad cellular roles of RecQ proteins.
In E. coli and nearly all other bacteria, SSB proteins function as homotetramers, each subunit of which comprises an N-terminal oligonucleotide/oligosaccharide binding (OB) fold linked to a structurally dynamic C-terminal tail 25; 26; 27; 28; 29 (Figure 1A). Whereas the OB fold is responsible for tetramerization and ssDNA binding, the C-terminal tail region ends with a highly conserved amphipathic peptide sequence (SSB-Ct) that associates with a diverse array of genome maintenance enzymes 30; 31 (Figure 1A). Interactions made with the SSB-Ct element are critical to proper genome maintenance and alterations in the SSB-Ct sequence can severely affect E. coli viability 32; 33; 34; 35; 36; 37. To date, every bacterial SSB-interacting protein examined binds to the SSB-Ct, including E. coli RecQ 19; 30, although structural data that indicate how such complexes are formed are only available for E. coli Exonuclease I 31. Additional studies are necessary to map the conserved structural features that define SSB-Ct binding sites.
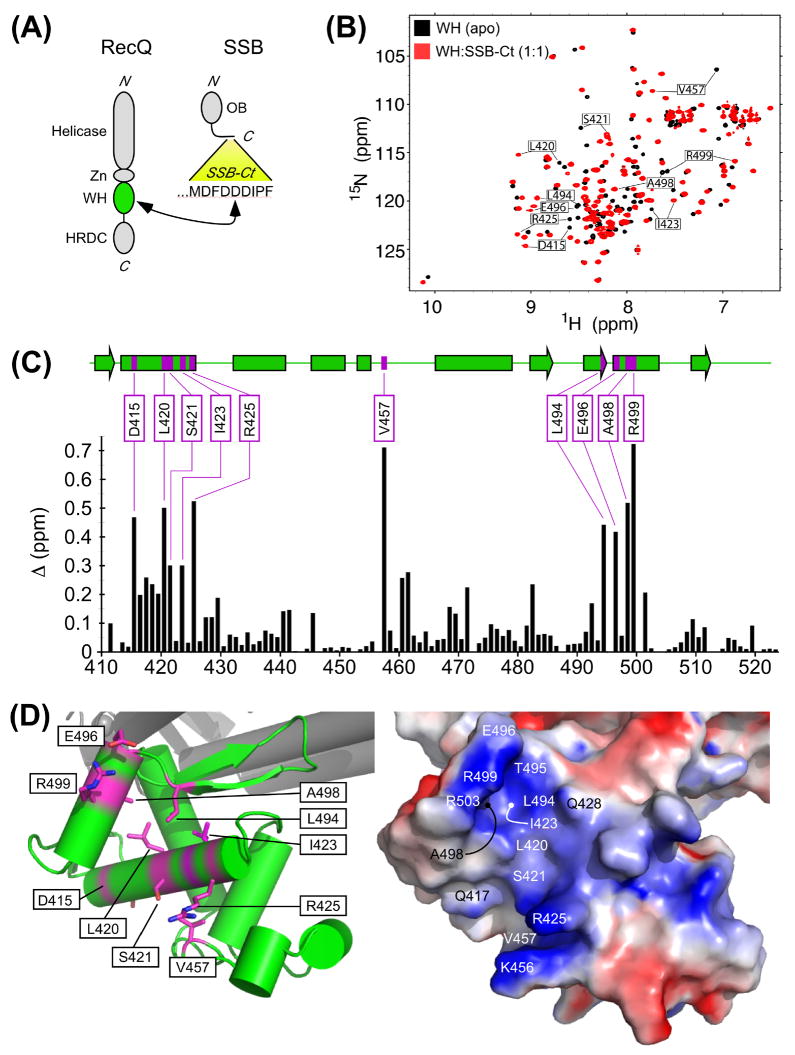
Figure 1. Structural basis of RecQ SSB-Ct binding.
(A) Schematic domain diagram of E. coli RecQ and SSB proteins. Domains in RecQ (Helicase, Zinc-binding (Zn), winged-helix (WH, green), and Helicase and RNase D C-terminal (HRDC)) and SSB (OB) are labeled. The sequence of the residues from SSB comprising the SSB-Ct element is given. An arrow indicates the elements of RecQ and SSB that are sufficient for complex formation19. (B) 15N HSQC spectral overlay of the RecQ-WH domain (black) and a 1:1 mixture of the RecQ-WH domain and SSB-Ct peptide (red). The largest 10 chemical shift differences between the spectra are labeled. (C) Histogram of the difference in amide proton and nitrogen resonance chemical shifts of the RecQ-WH with and without the addition of a 1:1 mixture of the SSB-Ct peptide. The largest differences (>0.3 ppm) are labeled and colored in magenta on secondary structural elements (helices as boxes, strands as arrows) of the RecQ-WH domain in the context of the crystal structure of the RecQ catalytic core 44. (D) Structure and electrostatics of the putative RecQ SSB-Ct binding site. (left) The ten RecQ-WH domain residues with the highest chemical shift differences upon SSB-Ct binding are labeled and colored in magenta on a ribbon diagram of the RecQ catalytic core structure. (right) Electrostatic representation of the surface of the RecQ catalytic core (red, blue and white for negative, positive and neutral, respectively) is shown in the same perspective as the ribbon diagram. Residues in addition to those identified by NMR that were altered in biochemical studies are labeled in the electrostatic diagram.
E. coli SSB is important for stimulating E. coli RecQ helicase activity and for aiding in joint activities between RecQ and other genome maintenance enzymes. SSB stimulates RecQ in at least two ways. First, SSB passively stimulates RecQ activity by binding and sequestering ssDNA generated by RecQ helicase activity and by preventing formation of non-productive ssDNA/RecQ complexes that inhibit the enzyme 38; 39. Second, SSB actively stimulates RecQ by recruiting RecQ to its DNA substrate and/or helping retain the enzyme on DNA through direct physical interaction 19. For active stimulation, interaction between SSB and RecQ is mediated by direct contacts made between the SSB-Ct sequence and the winged-helix domain of RecQ (RecQ-WH, Figure 1A) 19. Interestingly, the winged-helix domain of WRN is also important for contacting RPA 40; however, this interaction presumably occurs via a different mechanism, as RPA lacks the amphipathic SSB-Ct sequence that defines the bacterial SSB protein interaction site. Nonetheless, it is striking that for both E. coli RecQ and WRN, their respective winged-helix domains play important roles in complex formation with cognate SSB proteins. SSB is also a component in more complicated reactions that include recombination initiation with RecQ and RecA 41, DNA catenation and supercoiling, and converging replication fork resolution with RecQ and Topoisomerase III 6; 42; 43.
In this study, we have used NMR to identify the SSB-Ct binding site on the RecQ-WH domain. This site, which is on the opposite face from the proposed DNA binding site of the domain 44, forms a pocket that includes non-polar and electropositive elements that resemble those found in the Exonuclease I SSB-Ct binding site 31. Using the NMR data and additional structural information from the E. coli RecQ catalytic core X-ray crystal structure 44 as guides, we constructed a panel of RecQ variant proteins with amino acid substitutions that were predicted to impair association with SSB. This panel was used to determine the extent to which alteration of particular residues influences RecQ binding to SSB, and RecQ DNA binding and unwinding in the presence and absence of SSB. Additional experiments using SSB variant proteins with altered SSB-Ct elements assessed the effect of altering SSB in the same reactions. Each of the RecQ variant proteins had deficiencies in SSB binding, with several arginine and glutamine substitutions having the greatest impact. RecQ proteins bearing one or two amino acid changes that diminish SSB binding exhibited modest nucleic acid binding defects, but more pronounced defects in binding to SSB-coated DNA. These RecQ variants retained the ability to be stimulated by SSB, albeit to a lesser extent than wild type proteins. Nonstandard SSB-Ct elements also mildly inhibited the helicase activity of wild type RecQ protein on a partial duplex DNA substrate. We postulate that SSB-Ct binding to the site identified in this study influences RecQ-mediated DNA unwinding of cellular nucleoprotein substrates.
RESULTS
NMR studies identify RecQ-WH domain residues involved in SSB-Ct binding
Our earlier results indicated that E. coli RecQ and SSB form a complex that is mediated by stoichiometric binding of the RecQ-WH domain to the SSB-Ct element 19, although the precise SSB-Ct binding site on the RecQ-WH domain was not defined. A structural approach was therefore taken to map the SSB-Ct binding site on RecQ. We initially attempted to map the site by soaking the SSB-Ct peptide into crystals of the E. coli RecQ catalytic core domain (which includes the RecQ-WH domain) but electron density for the peptide was never observed in these experiments (data not shown). Since the isolated RecQ-WH domain binds the SSB-Ct peptide with the same specificity and nearly the same affinity as the full-length RecQ protein 19, we used NMR to define the SSB-Ct binding site on this small (116-residue) domain.
15N- and 15N-,13C-labeled RecQ-WH domain samples provided well-dispersed NMR spectra that allowed determination of over 90% of the backbone resonance assignments from the domain as described in Materials and Methods (Supplementary Figure 1 and Supplementary Table 1). To map the SSB-Ct binding site, we titrated the SSB-Ct peptide into the RecQ-WH NMR samples and measured changes in the chemical shift positions of the domain’s backbone amides in 2D 1H-15N HSQC experiments. We anticipated that amides in residues directly involved in SSB-Ct binding would undergo significant chemical shift changes relative to those that are not involved in binding. Consistent with this hypothesis, the addition of the SSB-Ct peptide dramatically changed the chemical shifts of several HN resonances from the RecQ-WH domain (Figure 1B). Nineteen HN chemical shifts were substantially altered (≥ 0.2 ppm) by SSB-Ct addition: D415, A416, Q417, I418, A419, L420, S421, I423, R425, V457, M460, G461, S471, V482, L494, E496, A498, R499, and V501. Of these residues, the largest chemical shift changes (≥ 0.3 ppm) occurred for D415, L420, S421, I423, R425, V457, L494, E496, A498, and R499 (Figures 1B & 1C). For many of these latter residues, the corresponding peak intensities disappear at intermediate titration points (1:0.5 RecQ-WH:SSB-Ct) due to exchange between the free and bound conformations occurring during the NMR experiment. Therefore, detailed analysis of the 3D 1H-15N and 1H-13C NOESY-HSQC spectra (1:1 RecQ-WH:SSB-Ct) was required to assign the new HN chemical shifts for the bound form of the RecQ-WH domain. Eleven amide residues (Y410, G412, N443, H452, K456, V472, A487, T495, R503, G504, and V512) could not be unambiguously identified after 1:1 molar addition of SSB-Ct peptide to the RecQ-WH domain.
Each of the residues that undergoes a ≥ 0.3 ppm chemical shift upon binding to the SSB-Ct peptide maps to a single cleft on the E. coli RecQ-WH domain (Figure 1D). The cleft has electrostatic characteristics that include a hydrophobic pocket (L420, I423, L494, and A498) flanked by positively charged or polar elements (R499 and R503, with additional residues Q417, R425, and K456 near the pocket) (Figure 1D). As is described in the Discussion, these electrostatic features are strikingly similar to those found in the E. coli Exonuclease I SSB-Ct binding site, which is the only protein for which the SSB-Ct binding site has previously been defined 31. This cleft is positioned on the opposite face of the RecQ-WH domain relative to the proposed dsDNA-binding surface of the domain 44. Interestingly, this cleft also forms a crystal contact in RecQ catalytic core domain crystals 44, which could explain why SSB-Ct peptide soak experiments failed to produce the complex.
In addition to residues that comprise the putative SSB-Ct binding cleft on the E. coli RecQ-WH domain, HN chemical shift perturbation data revealed several other regions that undergo more minor chemical shift changes (0.1 – 0.2 ppm) (Figure 1C). Although some of these smaller shift changes correspond to residues adjacent to the putative SSB-Ct binding site (N427, Q428, R429, L492), several others do not (D411, R440, G441, Q445, H468, W469, Q509, and V512). These chemical shift changes are dispersed across loop and helical regions of the RecQ-WH domain. Therefore, it is likely that the observed minor chemical shift perturbations reflect small structural changes that are imparted upon SSB-Ct binding.
Panel of RecQ variant proteins
Based on the above structural studies, we hypothesized that the RecQ-WH domain cleft that was sensitive to the addition of the SSB-Ct peptide forms its SSB-Ct binding site. To test this idea, we created over-expression plasmids to produce a panel of single-site alanine RecQ variant proteins that individually alter several of the residues with large (≥ 0.3 ppm) SSB-Ct-dependent chemical shift changes: D415A, S421A, R425A, V457A, and R499A (Figure 1). Examination of the crystal structure of the RecQ catalytic core 44 revealed additional neighboring residues that form part of the putative SSB-Ct-binding cleft for which expression vectors were also created: Q417A, Q428A, K456A, T495A, and R503A. Based on the importance of basic residues in the SSB-Ct interface of Exonuclease I 31, charge-reversal variants were also made for R425, R499, and R503 (changing each to glutamate) along with each permutation of double and triple mutation for each arginine residue. Finally, a control variant was made in which an arginine distal from the putative SSB-Ct binding site was altered to alanine (R448A). Four of the RecQ variants (T495A, R425E/R499E, R499E/R503E, and R425E/R499E/R503E) could not be purified to homogeneity due to apparent proteolysis of the overexpressed proteins, and were not pursued further.
To determine whether any of the RecQ variants were misfolded, we measured the ssDNA-dependent ATP hydrolysis rates of each protein. Wild type RecQ hydrolyzed ATP with a rate of 1210 ± 140 min−1, which is in agreement with previously published measurements 51; 52. Of the fourteen variant RecQ proteins studied, five were indistinguishable from wild type RecQ (Q428A, R448A, V457A, R499A, and R499E) and the remaining nine only modestly reduced in ATP hydrolysis rates (ranging from 1.3-fold to 1.6-fold reductions) (Table 1). With the very modest effects on ATPase activity, these studies indicate that each of the RecQ variants is properly folded.
Table 1.
Biochemical parameters of RecQ variant proteins
| RecQ variant | ATP Hydrolysis (min−1) | Apparent relative DNA binding affinity | Apparent relative SSB/DNA binding affinity | [RecQ] for half-maximal unwinding (nM) | [RecQ] for half-maximal unwinding with SSB (nM) | Fold stimulation of helicase activity with SSB |
|---|---|---|---|---|---|---|
| RecQ | 1210 ± 140 | +++++ | +++++ | 2.9 +/− 0.1 | 0.3 +/− 0.1 | 9.7 |
| R448A | 1210 ± 170 | +++++ | +++++ | 3.5 +/− 0.4 | 1.5 +/− 0.1 | 2.3 |
| D415A | 740 ± 130 | +++ | ++ | 13.9 +/− 0.7 | 6.8 +/− 1.5 | 2.0 |
| Q417A | 910 ± 70 | +++ | ++ | 6.4 +/− 1.8 | 2.0 +/− 0.2 | 3.2 |
| S421A | 940 ±100 | + | + | 9.5 +/− 1.1 | 4.7 +/− 1.6 | 2.0 |
| R425A | 730 ± 60 | +++ | ++ | 4.5 +/− 0.6 | 2.3 +/− 0.5 | 2.0 |
| R425E | 810 ± 100 | +++ | +++ | 8.1 +/− 0.3 | 5.4 +/− 0.6 | 1.5 |
| Q428A | 1310 ± 110 | +++++ | ++++ | 8.1 +/− 1.2 | 2.0 +/− 0.6 | 4.0 |
| K456A | 840 ± 60 | +++ | ++ | 7.4 +/− 3.0 | 2.3 +/− 0.3 | 3.2 |
| V457A | 1050 ± 80 | +++ | +++ | 3.0 +/− 0.4 | 1.0 +/− 0.1 | 3.0 |
| R499A | 1250 ± 140 | +++ | ++ | 4.9 +/− 0.8 | 3.5 +/− 0.1 | 1.4 |
| R499E | 1100 ± 110 | ++ | ++ | 8.7 +/− 1.2 | 4.3 +/− 0.3 | 2.0 |
| R503A | 770 ± 60 | ++ | + | 9.8 +/− 2.7 | 6.0 +/− 0.8 | 1.6 |
| R503E | 770 ± 90 | ++ | + | 6.8 +/− 1.1 | 4.1 +/− 0.6 | 1.7 |
| R425E/R503E | 960 ± 80 | +++ | ++ | 21.6 +/− 4.6 | 13.2 +/− 2.1 | 1.6 |
Amino acid substitutions in the putative SSB-Ct binding site impair SSB binding
Ammonium sulfate co-precipitation experiments were used to determine whether alterations in the RecQ variant proteins affect SSB binding. This method was developed originally to characterize SSB interactions with Exonuclease I and has been adapted to analyze SSB binding to other partners as well, including RecQ 6; 19; 37. The assay relies on the fact that SSB is precipitated in low concentrations of ammonium sulfate (150 g/L) whereas RecQ remains soluble in this condition. However, when SSB is mixed with RecQ prior to the addition of ammonium sulfate, RecQ that is in complex with SSB is co-precipitated and the amount of RecQ found in the insoluble fraction provides a relative measure of SSB binding.
Consistent with identification of the SSB-Ct binding site by NMR, each of the RecQ variants with altered residues in the putative SSB-Ct binding site showed deficiencies in SSB binding compared to wild type RecQ in the co-precipitation assay (Figure 2A). These binding defects could be divided into two groups: those with modest defects, which co-precipitated 20–50% as efficiently as wild-type RecQ (D415A, Q417A, S421A, K456A, and V457A), and those with severe defects, which co-precipitated 0–20% as efficiently as wild-type RecQ (R425A, R425E, Q428A, R499A, R499E, R503A, R503E, and R425E/R503E) (Figure 2A). In all cases, the solubility of the RecQ variant proteins in the absence of SSB was unaltered relative to wild type RecQ (data not shown). Notably, all but one of the substitutions that resulted in the largest deficiencies in SSB binding were those made in arginine residues at the periphery of the putative SSB-Ct binding pocket (Figure 2B). These residues could provide positive charges to bind the highly electronegative SSB-Ct element. Additionally, the R448A control RecQ variant co-precipitated nearly as efficiently as wild-type RecQ (~80%). These data support identification of the SSB-Ct binding site on RecQ by NMR.
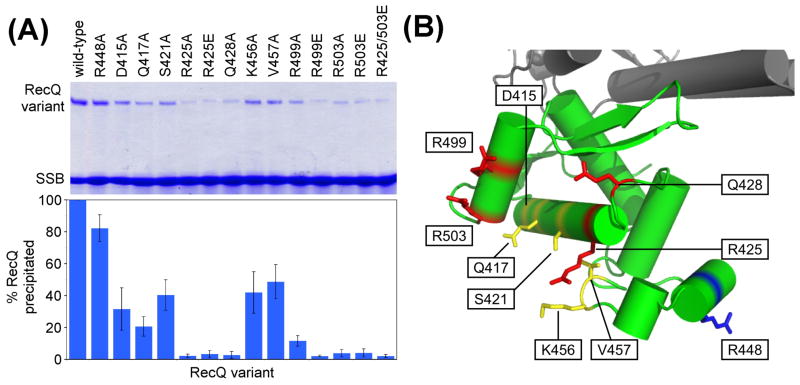
Figure 2. RecQ variant SSB binding to SSB.
(A) Ammonium sulfate co-precipitation pellets of RecQ variant and SSB mixtures were resolved by PAGE (top). Quantitation of the intensity of the RecQ variant band relative to wild-type protein is shown below the gel. Values are the mean of three measurements and one standard deviation as the error. (B) Alanine-substituted residues with modest (yellow) and severe (red) defects in SSB co-precipitation are shown on a ribbon diagram of the RecQ catalytic core structure.
RecQ variants have reduced DNA and SSB/DNA binding affinity
Previous electrophoretic mobility shift assays (EMSAs) have shown that RecQ can bind to a partial-duplex DNA structure (30-basepair duplex with a 70-base 3′ ssDNA overhang) and to a nucleoprotein complex formed between SSB and the DNA 19. However, substitution of an SSB that lacks the SSB-Ct element occludes RecQ binding, indicating that RecQ recognizes both DNA and SSB in the nucleoprotein complex 19. The same assay was used to test whether the RecQ variant proteins have impaired DNA or SSB/DNA binding.
Most of the RecQ protein variants had modestly weakened DNA binding affinities relative to wild type RecQ (Figure 3 (left panels) and Table 1). Given the presence of multiple bands that likely correspond to different stoichiometries in the protein/DNA complexes, we could not accurately estimate dissociation constants for the protein variants. However, the EMSA experiments provided qualitative evidence that the substitutions in the panel of proteins alter DNA binding. Most of the variants are only modestly defective in DNA binding (e.g. D415A, which required ~2-fold higher concentrations to shift the DNA mobility) but the S421A protein, which formed very low-mobility DNA complexes only at very high protein concentrations (50–100 nM), was strongly defective in DNA binding. The control variant (R448A) bound the DNA as well as wild type RecQ. The modest DNA binding defects of most of the RecQ variants might arise from substitution-induced changes in a DNA binding site distal from the SSB-Ct binding site although a direct role for the site in DNA binding can not be ruled out. Accordingly, studies with E. coli RecQ, WRN, and BLM proteins indicate that the winged-helix domain in each protein plays a direct role in DNA binding 45; 46; 47; 48.
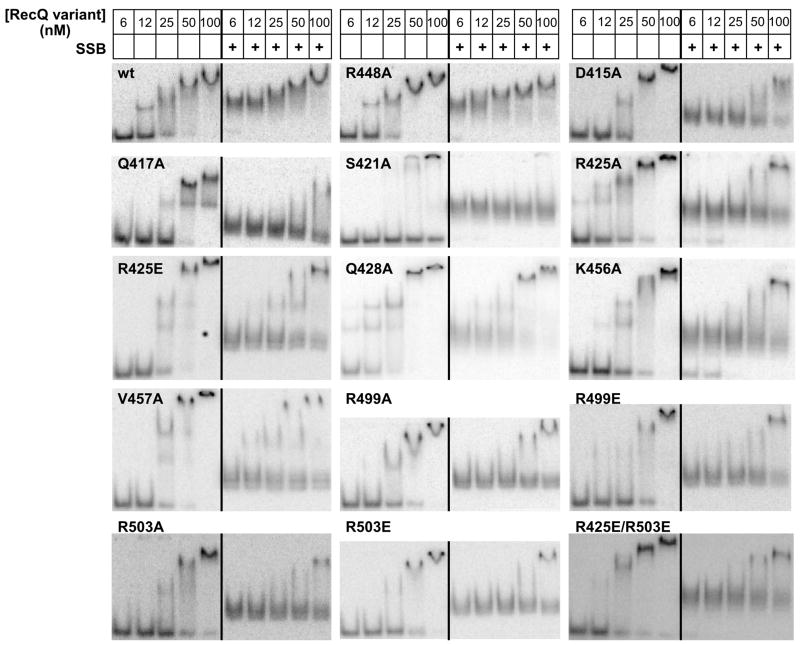
Figure 3. RecQ variants DNA and SSB/DNA binding.
Electrophoretic mobility shift analysis of RecQ variant proteins binding to partial duplex DNA is shown in the absence (left, each panel) and presence (right, each panel) of SSB.
We next tested binding of the panel of RecQ variants to the same DNA pre-bound by SSB to examine the effects of the mutations on a nucleoprotein complex. Because our previous results indicated that RecQ recognizes both the DNA and the SSB components in SSB/DNA structures, variants that were defective in binding to either element should have weakened binding in this assay. Consistent with this prediction, each of the RecQ variant proteins (except for the R448A control) bound the SSB/DNA substrate with lower apparent affinity than the wild type protein (Figure 3 (right panels) and Table 1). The S421A variant, which had the weakest DNA binding, also bound the SSB/DNA substrate most poorly, requiring 100 nM protein to elicit a mobility shift. Several other RecQ variants predominantly formed smears in the SSB/DNA EMSA (D415A, Q417A, V457A, R499A, R499E, R503A, and R503E), which could be related to weakened complex formation. Nearly all of the RecQ variants formed distinctive low-mobility bands but required higher protein concentrations than wild type RecQ or the R448A control variant to do so. Notably, the effects of the RecQ-WH domain substitutions in SSB/DNA binding were less pronounced that those observed in the RecQ/SSB binding assay (Figure 2), which could be due to local concentration effects arising from RecQ and SSB binding to a common DNA substrate in the former assay. This possibility is discussed further below.
RecQ variants have reduced SSB-dependent helicase activity stimulation
Because SSB stimulation of RecQ DNA unwinding depends on direct physical interaction between the two proteins 19, we predicted that disrupting the SSB/RecQ interface would inhibit SSB-mediated stimulation of RecQ helicase activity. To test this hypothesis, the SSB-dependent helicase activity of each RecQ variant protein was measured utilizing the same partial-duplex DNA substrate that was used in the EMSA experiments. The 70-base 3′ ssDNA on the substrate provides a binding site for up to two tetramers of E. coli SSB 49; 50. Consistent with previously published data 19, wild type RecQ unwinds this substrate in a reaction that is stimulated ~10-fold by SSB (Table 1).
The variant RecQ proteins showed differential effects in unwinding the partial duplex DNA substrate. In each case, RecQ helicase activity was stimulated in the presence of SSB, albeit to a lesser extent than the ~10-fold increase observed with wild-type RecQ (Table 1). The degree of stimulation ranged from 4-fold in the case of the most-stimulated variant (Q428A) to 1.4-fold in the case of the least (R499A). Interestingly, each variant’s ability to bind SSB in solution did not correlate with SSB-stimulation of its helicase activity. This is perhaps best illustrated by the R448A control variant, which bound SSB and SSB/ssDNA with apparent wild type affinity but was only stimulated 2.3-fold by SSB. The S421A variant exhibits an intermediate ability to bind SSB in solution (~40% as well as wild type) but its helicase activity is only stimulated ~2-fold by SSB, presumably because its ability to bind the DNA substrate both in the presence and absence of SSB is greatly diminished (Figure 3). In contrast, the Q428A variant is an extremely poor SSB binding protein in solution (~3% of wild type levels) but displays the greatest degree of stimulation by SSB of any of the variants tested (Table 1). Therefore, because it appears that mutations in the RecQ-WH domain affect both SSB and DNA binding (and perhaps other functions of RecQ as well), the helicase results could not be correlated in a simple way to defects in SSB binding alone.
SSB variants with altered SSB-Ct sequences modify RecQ/SSB functions
To better evaluate the roles of the SSB/RecQ protein interface in their joint function, we employed several SSB variant proteins with altered SSB-Ct elements. These included: (1) SSB113, which changes the penultimate proline of the wild type SSB-Ct sequence (MDFDDDIPF) to serine; (2) SSB-mixed, which shuffles the order of the amino acids comprising the SSB-Ct sequence to DFMDDPFID; and (3) SSBΔC1, which removes the C-terminal-most phenylalanine residue. We have shown that RecQ is unable to bind SSB-Ct peptide equivalents of SSB113 and SSB-mixed sequences previously 19, and based on structural studies of Exonuclease I bound to the SSB-Ct peptide, we predicted that removing the C-terminal phenylalanine would disrupt binding as well 19; 31.
We first carried out ammonium sulfate co-precipitation and EMSA studies to determine the effects of the SSB mutations on complex formation with RecQ. Wild type RecQ does not co-precipitate with any of these SSB variant proteins (Figure 4A), confirming the importance of the SSB-Ct amino acid sequence for binding RecQ in the absence of DNA. RecQ was also diminished in its ability to bind DNA pre-coated with either the SSB-mixed or SSBΔC1 variant SSB proteins, whereas binding to SSB113-coated DNA was not significantly diminished relative to that observed with SSB-coated DNA (Figure 4B, compare with Figure 3). This difference may reflect the more modest alteration of SSB113 compared to the other SSB variant proteins. Nonetheless, these results support the notion that RecQ recognizes both DNA and SSB-Ct features in SSB/DNA substrates.
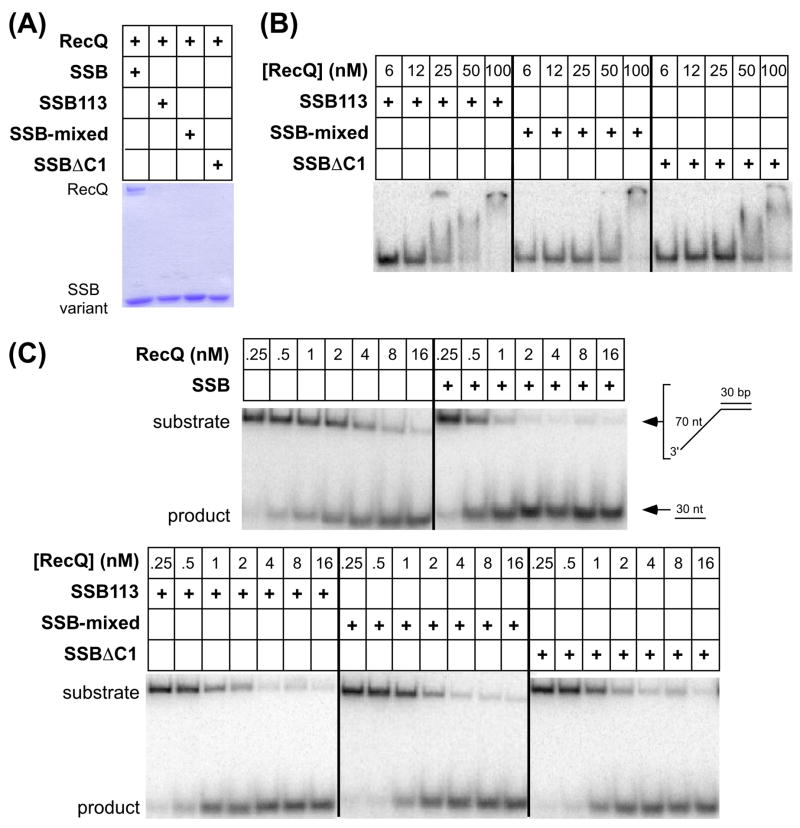
Figure 4. Effects of SSB-Ct sequence changes on RecQ binding.
(A) Ammonium sulfate co-precipitation pellets of RecQ and SSB variant mixtures were resolved by PAGE. (B) Electrophoretic mobility shift analysis of RecQ binding to partial duplex DNA prebound by SSB protein variants. (C) DNA unwinding by wild type RecQ in the absence and presence of SSB and SSB proteins variants.
When RecQ helicase activity was tested on DNA substrates pre-bound to the SSB variants, we observed that higher concentrations of RecQ protein were required to reach half-maximal unwinding when SSB-mixed or SSBΔC1 were included than was necessary for wild type SSB-bound DNA substrates, suggesting that the SSB variants modestly inhibit helicase activity (Figure 4C and Table 2). Taken together, these results suggest that the RecQ/SSB-Ct interaction is dependent upon an intact SSB-Ct element. It is interesting to note however that none of the SSB variants tested entirely blocked RecQ DNA binding and unwinding in the way that complete deletion of the SSB-Ct element was observed earlier 19. This could indicate that multiple elements of the SSB-Ct are important for associating with RecQ and subtle alterations of the SSB-Ct do not eliminate all of the contact points used to form the RecQ/SSB complex.
Table 2.
DNA helicase activity of RecQ with SSB variant proteins
| SSB Variant | [RecQ] for half-maximal unwinding (nM) | [RecQ] for half-maximal unwinding with SSB (nM) | Fold stimulation |
|---|---|---|---|
| SSB | 2.9 +/− 0.1 | 0.3 +/− 0.1 | 9.7 |
| SSB113 | 2.9 +/− 0.1 | 3.2 +/− 0.5 | 0.9 |
| SSB-mixed | 2.9 +/− 0.1 | 4.0 +/− 0.7 | 0.7 |
| SSBΔC1 | 2.9 +/− 0.1 | 4.7 +/− 0.8 | 0.6 |
DISCUSSION
More than a dozen proteins involved in DNA replication, repair, recombination, and replication restart in E. coli have been found to interact directly with SSB, establishing it as an important scaffold for a large number of genome maintenance processes 30. In all cases examined to date, interactions with SSB depend on the SSB-Ct element, an evolutionarily conserved amphipathic sequence of acidic and hydrophobic amino acids found at the C-terminus of SSB 30; 31; 32; 33; 35; 36; 53. Studies that offer structural and mechanistic insights into how the SSB-Ct element binds to heterologous proteins and the biochemical consequences of these associations are important for understanding the structural architecture that underlies several bacterial genome maintenance processes.
Stimulation of DNA helicase activity by SSB was noted early in the characterization of RecQ 38. This and subsequent studies concluded that this stimulation resulted from SSB binding to and sequestering liberated ssDNA following RecQ helicase activity or from SSB preventing the formation of non-productive ssDNA/RecQ complexes, but more recent investigations have shown that direct contact between the two proteins also plays an important role 19; 38; 39. Here, we have combined structural and biochemical approaches to identify the SSB-Ct binding site on RecQ. Altering residues that comprise this site in the RecQ-WH domain results in greatly impaired binding to SSB and leads to more modest changes in SSB/DNA binding. This difference could be due to local concentration effects arising from SSB and RecQ binding to a common DNA substrate in the SSB/DNA experiments. Because it is involved in several distinct activities, alterations of the RecQ-WH domain affect biochemical functions of the RecQ variant proteins beyond just impairment of SSB binding. We have also shown that SSB variants with subtle changes in the SSB-Ct element generally inhibit RecQ substrate recognition and RecQ unwinding of SSB/ssDNA complexes.
Conservation of an SSB interaction site among genome maintenance enzymes
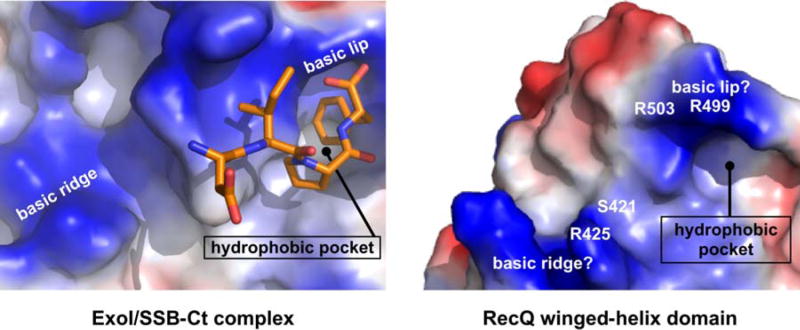
Our identification of the SSB-Ct binding site on RecQ allowed us to compare the RecQ site to that of Exonuclease I, which is the only other SSB-associated protein for which structural information on the complex is available 31. The Exonuclease I SSB-Ct binding site is defined by hydrophobic and basic chemical features that function to accommodate several determinants from the SSB-Ct peptide (Figure 5, left). These include a hydrophobic pocket on Exonuclease I that engulfs the side chain of the C-terminal phenylalanine of the SSB-Ct. Binding to the phenylalanine is further promoted by an arginine residue from Exonuclease I (the “basic lip” of the hydrophobic pocket), which complements the phenylalanine’s α-carboxyl oxygens via ionic interactions. Additional basic residues from Exonuclease I form a “basic ridge” that is thought to be important for binding the more N-terminal acidic residues of the SSB-Ct. Comparison of the SSB-Ct binding sites from Exonuclease I and RecQ reveals a number of structural features shared between the two sites, including a hydrophobic pocket lined by two potential “basic lip” residues (R499 and R503) and an adjoining “basic ridge” on RecQ (Figure 5). In all cases, alanine substitutions of residues in this region produce proteins with weakened SSB and SSB/DNA binding (Figures 2 and 3), consistent with this being the bona fide SSB-Ct binding site in RecQ. Remarkably, these common surface features in Exonuclease I and RecQ arise from unrelated protein folds, which could indicate that the two enzymes have evolved similar SSB-Ct binding sites independently. Because many of the other known SSB-associated proteins also lack structural homology with Exonuclease I and RecQ, these proteins appear to have independently evolved SSB binding sites as well. Future studies will be required to determine whether the features common to Exonuclease I and RecQ SSB-Ct binding sites are conserved among other SSB partner proteins. The similar SSB-Ct binding site structures in Exonuclease I and RecQ provide an electrostatic template that could prove useful for identifying SSB-Ct binding sites in these proteins.
Figure 5.
Comparison of the SSB-Ct binding site from Exonuclease I (left) 31, and RecQ (right). Surface representations of each molecule are colored by electrostatic features (red, blue and white for negative, positive and neutral, respectively). The SSB-Ct peptide is shown in orange on the Exonuclease I structure. The basic ridge, the arginine at the lip of the binding pocket, and the hydrophobic pocket are highlighted in both images.
The RecQ-WH domain as a node that integrates DNA and SSB binding
Winged-helix domains have been identified as common DNA-binding elements in many different proteins 47; 54; 55; 56; 57; 58; 59. In addition, a growing number of winged-helix domains with protein interaction roles have also been noted 47; 60; 61; 62. Based upon its structural similarity to the winged-helix domain of several dsDNA-binding proteins, a DNA binding role for the RecQ-WH domain has been predicted in addition to its role in SSB binding 19; 44. Several studies indicate that the winged-helix domains from E. coli RecQ and its eukaryotic homologs directly bind to DNA 45; 46; 47; 48, and recent work has shown that the winged helix domain of human RECQ1 plays an integral role in the DNA unwinding mechanism of the protein 72. Furthermore, experiments with the WRN winged-helix domain 47 and preliminary NMR experiments from our group using the RecQ-WH domain (data not shown) indicate that their respective winged-helix domain bind DNA using sites that are distinct from the SSB-Ct binding site described herein.
Identification of the SSB-Ct binding site in this study shows that SSB binds at a position in the RecQ-WH domain that is on the opposite face relative to the predicted dsDNA-binding surface, which highlights a potential integrative role for the domain. Related to this centrality, nearly every RecQ-WH variant tested exhibited modest DNA binding defects even in the absence of SSB (Figure 3). Likewise, single amino acid changes that are predicted to affect only SSB binding also alter RecQ helicase activity even in the absence of SSB (Table 1). These defects are distinct from the more dramatic defects in complex formation with SSB (Figure 2) but nonetheless could reflect larger roles for the RecQ-WH domain in RecQ activity. Given the compact size of the RecQ-WH domain, these observations could relate to changes on other surfaces on the domain from the amino acid substitutions or could reflect possible secondary roles for the SSB binding site in RecQ activity, such as DNA binding during helicase functions. Such interrelated effects could point to an integrative role for the RecQ-WH domain in coordinating joint DNA and SSB binding, as has been hypothesized for the WRN winged-helix domain 47. Interrelated binding could reflect an evolved coordination that is important for targeting the activity of RecQ within the cell.
MATERIALS AND METHODS
Proteins, synthetic peptides, and DNA substrates
Full-length E. coli RecQ and variant proteins
Hexahistidine-tagged E. coli RecQ and single- and double-site variant proteins were expressed, purified and quantitated as described previously 51. Alanine- and glutamic acid variants were generated by the QuikChange site-directed mutagenesis scheme (Stratagene). All plasmids were sequenced at the University of Wisconsin Biotechnology Center.
RecQ-WH domain
A T7-overexpression plasmid encoding the hexahistidine-tagged RecQ-WH domain (pET28-RecQ-WH, residues 408–523 of E. coli RecQ) has been described previously 19. E. coli BL21 (DE3) cells transformed with pET28-RecQ-WH were cultured at 30°C in M9 minimal media containing 150 μg/ml ampicillin, 15NH4Cl, and, in some cases, 13C-glucose, to late log phase (OD600 ~ 1.0). Protein expression was induced for 10 hours by the addition of isopropyl β-D-thiogalactopyranoside to 1 mM and the domain was purified as previously described 19. Purified RecQ-WH was dialyzed against 20 mM HEPES (pH 6.5), 0.1 M KCl, and concentrated to 0.3–0.8 mM. All NMR samples contained 5–10 % D2O.
SSB proteins
T7-overexpression plasmids for E. coli SSB, SSB113 and SSB-mixed were described previously 31. An overexpression plasmid encoding SSBΔC1 (E. coli SSB protein lacking its C-terminal-most residue) was made using the QuikChange site-directed mutagenesis method (Stratagene) and sequenced at the University of Wisconsin Biotechnology Center. E. coli SSB and SSB variant proteins were purified as described 63, with the following modifications: following ammonium sulfate fractionation, SSB was dialyzed into low-salt buffer (20 mM Tris (pH 8.0), 50 mM NaCl, 1 mM EDTA, 10% v/v glycerol) and purified chromatographically over Heparin Fastflow and Sephacryl S-100 size exclusion columns (GE Healthcare). Pure fractions were pooled, concentrated and dialyzed into storage buffer (20 mM Tris pH (pH 8.0), 0.5 M NaCl, 1 mM EDTA, 1 mM β-mercaptoethanol, 50% glycerol v/v) and stored at −20°C.
SSB-Ct peptide
A peptide comprising the E. coli SSB C-terminal sequence with an additional N-terminal Trp residue (SSB-Ct; Trp-Met-Asp-Phe-Asp-Asp-Asp-Ile-Pro-Phe) was synthesized and purified by the University of Wisconsin Peptide Synthesis Facility as described previously 19.
Nucleic acid substrates
Oligonucleotides o30 (5′-CTAATGACGGTCCTAGACGAACCGAGCGTC-3′) and o100 (5′-GACGCTCGGTTCGTCTAGGACCGTCATTAGTATGTTGATATACATAGACCTTACCGCAGTGATTCGCTTGTCAGTCCATTGAAGCACAATTACCCACGC-3′) were synthesized and purified by Integrated DNA Technologies and have been described previously 19. o30 was phosphorylated by T4 polynucleotide kinase (New England Biolabs) with [γ-32P]ATP (GE Healthcare) and annealed to unlabeled o100 to form a partial duplex DNA (3′-OH) consisting of a 30-basepair duplex with a 70-base 3′-overhang. The substrate was resolved by native polyacrylamide gel electrophoresis (PAGE), electroeluted, and dialyzed against 20 mM Tris (pH 8.0), 50 mM NaCl.
Determination of NMR resonance assignments
Backbone 1H, 13C, 15N resonance assignments were carried out at 25°C using Varian Inova 800 and 600 MHz spectrometers equipped with cryogenic probes. The 13C and 15N chemical shifts were referenced indirectly to 2,2-dimethyl-2-silapentane-5-sulfonic acid as described previously 64. Sequence-specific backbone assignments were achieved using 2D 1H-15N HSQC, 3D HNCO, 3D HNCA, 3D HNCACB, 3D CBCA(CO)NH, and 3D HBHA(CO)NH experiments, as previously described 65; 66). In addition, 3D 1H-15N NOESY-HSQC and 1H-13C NOESY-HSQC (60 ms mixing time) spectra were acquired to confirm backbone resonance assignments. All spectra were processed and initially analyzed with NMRPipe 67 and Sparky (http://www.cgl.ucsf.edu/home/sparky) software packages, respectively. The automated assignment program, PINE (miranda.nmrfam.wisc.edu/PINE/) 68 was used to initially assign the backbone resonances. Near complete (~90%) backbone resonance assignments for the RecQ-WH domain were obtained and HN shifts are labeled on the 1H-15N HSQC spectrum (Supplemental Figure 1). Eight mM SSB-Ct peptide aliquots in 10 mM HEPES (pH 6.5) were titrated into the RecQ-WH domain NMR sample at room temperature and incubated for 15 minutes at each titration stage. Spectra were collected at 1:0, 1:0.1, 1:0.2, 1:0.5, 1:0.8, 1:1, 1:1.2, 1:1.5, 1:2, 1:2.5, and 1:5 RecQ-WH domain:SSB-Ct peptide molar ratios. Three dimensional 1H-15N NOESY-HSQC spectra were acquired and analyzed at 1:1 and 5:1 RecQ-WH domain:SSB-Ct peptide molar ratios. Normalized changes in chemical shift between the 1:0 and 1:1 spectra were calculated from the equation Δppm = [(Δ1H ppm)2 + (Δ15N ppm X αN)2]0.5, where Δppm is the difference in ppm between the chemical shifts of the RecQ-WH domain and RecQ-WH domain:SSB-Ct complex and 0.1 serves as the scaling factor (based on the approximately 10-fold lower gyromagnetic ratio of 15N relative to 1H) of the nitrogen chemical shift changes (αN) 69.
DNA-dependent ATPase assays
ATP hydrolysis was measured by a standard assay as previously described 70. RecQ variants (5 nM) were added to reactions containing dT35 (1 nM nucleotides) and an ATP regeneration system that allows for spectrophotometric observation of ATP hydrolysis by coupling this process to the oxidation of NADH 71. Steady-state rates of ATP hydrolysis were calculated and normalized to the concentration of RecQ in order to determine the rate of ATP hydrolyzed per RecQ molecule per minute. Rates are reported as the mean of three measurements and one standard deviation as the error.
Ammonium sulfate co-precipitation
Co-precipitation experiments were performed as described previously 19; 37, except that pellet fractions were suspended in 30 μl of loading buffer prior to SDS-PAGE on 4-15% polyacrylamide gradient gels (Bio-Rad). Assays were performed in triplicate, and relative binding was quantified based on setting the intensity of the wild type RecQ band to 100% using ImageQuant software (GE Healthcare). Values are reported as the mean of three measurements and one standard deviation as the error.
Electrophoretic mobility shift assays
3′-OH DNA substrate radiolabeled with 32P was diluted to 1 nM, added to reaction buffer (final concentrations: 20 mM Tris (pH 8.0), 50 mM NaCl, 1 mM β-mercaptoethanol, 1 mM MgCl2, 0.1 g/liter bovine serum albumin, 5.5% (v/v) glycerol) and incubated with 20 nM SSB tetramers or reaction buffer at room temperature for five minutes. RecQ (or a RecQ variant protein) was added to a final concentration of 0–1000 nM, incubated for five minutes at room temperature, and resolved by 6% native PAGE. Gels were run at 90V for 3 hours at 4°C, dried, and imaged using a Molecular Dynamics Storm 820 phosphorimager. Assays were conducted multiple times with representative images shown. Qualitative assessments of the relative strength of RecQ DNA and SSB/DNA binding are given in Table 1.
Helicase assays
Twenty nM SSB (or SSB variant) and 1 mM ATP were added to 1 nM 3′-OH DNA substrate in reaction buffer described above. Reactions were started by the addition of RecQ protein (0–16 nM, as indicated) and incubated at room temperature for 20 minutes. Reactions were stopped by addition of stop solution (10 μg Proteinase K, 0.25% SDS, 30 mM EDTA, and 8.6 nM unlabeled o30) and incubated at 37°C for 30 minutes. Reaction products were resolved by 10% native PAGE (140V for 1.5 hours at 4°C), dried, and imaged using a Molecular Dynamics Storm 820 phosphorimager. Reactions were performed in triplicate and quantified using ImageQuant 5.1 software. Half-maximal unwinding values were determined using CurveExpert software. Values are reported as the mean of three measurements and one standard deviation as the error.
Supplementary Material
Acknowledgments
This work was supported by a grant to JLK from the NIH (GM068061). We thank the NMRFAM staff on the University of Wisconsin for technical assistance, Duo Lu for providing several SSB variant constructs and members of the Keck laboratory for assistance in revising this manuscript. RDS is a Cremer Scholar. We also thank Alessandro Vindigni and Opher Gileadi for communicating results prior to publication. This study made use of the National Magnetic Resonance Facility at Madison, which is supported by NIH grants P41RR02301 (BRTP/NCRR) and P41GM66326 (NIGMS). Additional equipment was purchased with funds from the University of Wisconsin, the NIH (RR02781, RR08438), the NSF (DMB-8415048, OIA-9977486, BIR-9214394), and the USDA.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Bachrati CZ, Hickson ID. RecQ helicases: suppressors of tumorigenesis and premature aging. Biochem J. 2003;374:577–606. doi: 10.1042/BJ20030491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bennett RJ, Keck JL. Structure and function of RecQ DNA helicases. Crit Rev Biochem Mol Biol. 2004;39:79–97. doi: 10.1080/10409230490460756. [DOI] [PubMed] [Google Scholar]
- 3.Cobb JA, Bjergbaek L. RecQ helicases: lessons from model organisms. Nucleic Acids Res. 2006;34:4106–14. doi: 10.1093/nar/gkl557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Seki M, Tada S, Enomoto T. Function of recQ family helicase in genome stability. Subcell Biochem. 2006;40:49–73. doi: 10.1007/978-1-4020-4896-8_5. [DOI] [PubMed] [Google Scholar]
- 5.Sharma S, Doherty KM, Brosh RM., Jr Mechanisms of RecQ helicases in pathways of DNA metabolism and maintenance of genomic stability. Biochem J. 2006;398:319–37. doi: 10.1042/BJ20060450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Suski C, Marians KJ. Resolution of converging replication forks by RecQ and topoisomerase III. Mol Cell. 2008;30:779–89. doi: 10.1016/j.molcel.2008.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mehr IJ, Seifert HS. Differential roles of homologous recombination pathways in Neisseria gonorrhoeae pilin antigenic variation, DNA transformation and DNA repair. Mol Microbiol. 1998;30:697–710. doi: 10.1046/j.1365-2958.1998.01089.x. [DOI] [PubMed] [Google Scholar]
- 8.Sechman EV, Kline KA, Seifert HS. Loss of both Holliday junction processing pathways is synthetically lethal in the presence of gonococcal pilin antigenic variation. Mol Microbiol. 2006;61:185–93. doi: 10.1111/j.1365-2958.2006.05213.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Stohl EA, Seifert HS. Neisseria gonorrhoeae DNA recombination and repair enzymes protect against oxidative damage caused by hydrogen peroxide. J Bacteriol. 2006;188:7645–51. doi: 10.1128/JB.00801-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hishida T, Han YW, Shibata T, Kubota Y, Ishino Y, Iwasaki H, Shinagawa H. Role of the Escherichia coli RecQ DNA helicase in SOS signaling and genome stabilization at stalled replication forks. Genes Dev. 2004;18:1886–97. doi: 10.1101/gad.1223804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Courcelle J, Hanawalt PC. RecQ and RecJ process blocked replication forks prior to the resumption of replication in UV-irradiated Escherichia coli. Mol Gen Genet. 1999;262:543–51. doi: 10.1007/s004380051116. [DOI] [PubMed] [Google Scholar]
- 12.Courcelle J, Hanawalt PC. Participation of recombination proteins in rescue of arrested replication forks in UV-irradiated Escherichia coli need not involve recombination. Proc Natl Acad Sci U S A. 2001;98:8196–202. doi: 10.1073/pnas.121008898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ellis NA, Groden J, Ye TZ, Straughen J, Lennon DJ, Ciocci S, Proytcheva M, German J. The Bloom’s syndrome gene product is homologous to RecQ helicases. Cell. 1995;83:655–66. doi: 10.1016/0092-8674(95)90105-1. [DOI] [PubMed] [Google Scholar]
- 14.Yu CE, Oshima J, Fu YH, Wijsman EM, Hisama F, Alisch R, Matthews S, Nakura J, Miki T, Ouais S, Martin GM, Mulligan J, Schellenberg GD. Positional cloning of the Werner’s syndrome gene. Science. 1996;272:258–62. doi: 10.1126/science.272.5259.258. [DOI] [PubMed] [Google Scholar]
- 15.Kitao S, Shimamoto A, Goto M, Miller RW, Smithson WA, Lindor NM, Furuichi Y. Mutations in RECQL4 cause a subset of cases of Rothmund-Thomson syndrome. Nat Genet. 1999;22:82–4. doi: 10.1038/8788. [DOI] [PubMed] [Google Scholar]
- 16.Bohr VA, Cooper M, Orren D, Machwe A, Piotrowski J, Sommers J, Karmakar P, Brosh R. Werner syndrome protein: biochemical properties and functional interactions. Exp Gerontol. 2000;35:695–702. doi: 10.1016/s0531-5565(00)00145-5. [DOI] [PubMed] [Google Scholar]
- 17.Mankouri HW, Hickson ID. The RecQ helicase-topoisomerase III-Rmi1 complex: a DNA structure-specific ‘dissolvasome’? Trends Biochem Sci. 2007;32:538–46. doi: 10.1016/j.tibs.2007.09.009. [DOI] [PubMed] [Google Scholar]
- 18.Brosh RM, Jr, Bohr VA. Roles of the Werner syndrome protein in pathways required for maintenance of genome stability. Exp Gerontol. 2002;37:491–506. doi: 10.1016/s0531-5565(01)00227-3. [DOI] [PubMed] [Google Scholar]
- 19.Shereda RD, Bernstein DA, Keck JL. A central role for SSB in Escherichia coli RecQ DNA helicase function. J Biol Chem. 2007;282:19247–58. doi: 10.1074/jbc.M608011200. [DOI] [PubMed] [Google Scholar]
- 20.Brosh RM, Jr, Li JL, Kenny MK, Karow JK, Cooper MP, Kureekattil RP, Hickson ID, Bohr VA. Replication protein A physically interacts with the Bloom’s syndrome protein and stimulates its helicase activity. J Biol Chem. 2000;275:23500–8. doi: 10.1074/jbc.M001557200. [DOI] [PubMed] [Google Scholar]
- 21.Constantinou A, Tarsounas M, Karow JK, Brosh RM, Bohr VA, Hickson ID, West SC. Werner’s syndrome protein (WRN) migrates Holliday junctions and co-localizes with RPA upon replication arrest. EMBO Rep. 2000;1:80–4. doi: 10.1093/embo-reports/kvd004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cui S, Klima R, Ochem A, Arosio D, Falaschi A, Vindigni A. Characterization of the DNA-unwinding activity of human RECQ1, a helicase specifically stimulated by human replication protein A. J Biol Chem. 2003;278:1424–32. doi: 10.1074/jbc.M209407200. [DOI] [PubMed] [Google Scholar]
- 23.Cui S, Arosio D, Doherty KM, Brosh RM, Jr, Falaschi A, Vindigni A. Analysis of the unwinding activity of the dimeric RECQ1 helicase in the presence of human replication protein A. Nucleic Acids Res. 2004;32:2158–70. doi: 10.1093/nar/gkh540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Garcia PL, Liu Y, Jiricny J, West SC, Janscak P. Human RECQ5beta, a protein with DNA helicase and strand-annealing activities in a single polypeptide. EMBO J. 2004;23:2882–91. doi: 10.1038/sj.emboj.7600301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Weiner JH, Bertsch LL, Kornberg A. The deoxyribonucleic acid unwinding protein of Escherichia coli. Properties and functions in replication. J Biol Chem. 1975;250:1972–80. [PubMed] [Google Scholar]
- 26.Lohman TM, Ferrari ME. Escherichia coli single-stranded DNA-binding protein: multiple DNA-binding modes and cooperativities. Annu Rev Biochem. 1994;63:527–70. doi: 10.1146/annurev.bi.63.070194.002523. [DOI] [PubMed] [Google Scholar]
- 27.Raghunathan S, Kozlov AG, Lohman TM, Waksman G. Structure of the DNA binding domain of E. coli SSB bound to ssDNA. Nat Struct Biol. 2000;7:648–52. doi: 10.1038/77943. [DOI] [PubMed] [Google Scholar]
- 28.Savvides SN, Raghunathan S, Futterer K, Kozlov AG, Lohman TM, Waksman G. The C-terminal domain of full-length E. coli SSB is disordered even when bound to DNA. Protein Sci. 2004;13:1942–7. doi: 10.1110/ps.04661904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Williams KR, Spicer EK, LoPresti MB, Guggenheimer RA, Chase JW. Limited proteolysis studies on the Escherichia coli single-stranded DNA binding protein. Evidence for a functionally homologous domain in both the Escherichia coli and T4 DNA binding proteins. J Biol Chem. 1983;258:3346–55. [PubMed] [Google Scholar]
- 30.Shereda RD, Kozlov AG, Lohman TM, Cox MM, Keck JL. SSB as an organizer/mobilizer of genome maintenance complexes. Critical Reviews in Biochemistry and Molecular Biology. 2008 doi: 10.1080/10409230802341296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lu D, Keck JL. Structural basis of Escherichia coli single-stranded DNA-binding protein stimulation of exonuclease I. Proc Natl Acad Sci U S A. 2008;105:9169–74. doi: 10.1073/pnas.0800741105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wang TC, Smith KC. Effects of the ssb-1 and ssb-113 mutations on survival and DNA repair in UV-irradiated delta uvrB strains of Escherichia coli K-12. J Bacteriol. 1982;151:186–92. doi: 10.1128/jb.151.1.186-192.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chase JW, L’Italien JJ, Murphy JB, Spicer EK, Williams KR. Characterization of the Escherichia coli SSB-113 mutant single-stranded DNA-binding protein. Cloning of the gene, DNA and protein sequence analysis, high pressure liquid chromatography peptide mapping, and DNA-binding studies. J Biol Chem. 1984;259:805–14. [PubMed] [Google Scholar]
- 34.Curth U, Genschel J, Urbanke C, Greipel J. In vitro and in vivo function of the C-terminus of Escherichia coli single-stranded DNA binding protein. Nucleic Acids Res. 1996;24:2706–11. doi: 10.1093/nar/24.14.2706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kelman Z, Yuzhakov A, Andjelkovic J, O’Donnell M. Devoted to the lagging strand-the χ subunit of DNA polymerase III holoenzyme contacts SSB to promote processive elongation and sliding clamp assembly. EMBO J. 1998;17:2436–2449. doi: 10.1093/emboj/17.8.2436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yuzhakov A, Kelman Z, O’Donnell M. Trading places on DNA--a three-point switch underlies primer handoff from primase to the replicative DNA polymerase. Cell. 1999;96:153–63. doi: 10.1016/s0092-8674(00)80968-x. [DOI] [PubMed] [Google Scholar]
- 37.Genschel J, Curth U, Urbanke C. Interaction of E. coli single-stranded DNA binding protein (SSB) with exonuclease I. The carboxy-terminus of SSB is the recognition site for the nuclease. Biol Chem. 2000;381:183–92. doi: 10.1515/BC.2000.025. [DOI] [PubMed] [Google Scholar]
- 38.Umezu K, Nakayama H. RecQ DNA helicase of Escherichia coli. Characterization of the helix-unwinding activity with emphasis on the effect of single-stranded DNA-binding protein. J Mol Biol. 1993;230:1145–50. doi: 10.1006/jmbi.1993.1231. [DOI] [PubMed] [Google Scholar]
- 39.Harmon FG, Kowalczykowski SC. Biochemical characterization of the DNA helicase activity of the escherichia coli RecQ helicase. J Biol Chem. 2001;276:232–43. doi: 10.1074/jbc.M006555200. [DOI] [PubMed] [Google Scholar]
- 40.Doherty KM, Sommers JA, Gray MD, Lee JW, von Kobbe C, Thoma NH, Kureekattil RP, Kenny MK, Brosh RM., Jr Physical and functional mapping of the replication protein a interaction domain of the werner and bloom syndrome helicases. J Biol Chem. 2005;280:29494–505. doi: 10.1074/jbc.M500653200. [DOI] [PubMed] [Google Scholar]
- 41.Harmon FG, Kowalczykowski SC. RecQ helicase, in concert with RecA and SSB proteins, initiates and disrupts DNA recombination. Genes Dev. 1998;12:1134–44. doi: 10.1101/gad.12.8.1134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Harmon FG, DiGate RJ, Kowalczykowski SC. RecQ helicase and topoisomerase III comprise a novel DNA strand passage function: a conserved mechanism for control of DNA recombination. Mol Cell. 1999;3:611–20. doi: 10.1016/s1097-2765(00)80354-8. [DOI] [PubMed] [Google Scholar]
- 43.Harmon FG, Brockman JP, Kowalczykowski SC. RecQ helicase stimulates both DNA catenation and changes in DNA topology by topoisomerase III. J Biol Chem. 2003;278:42668–78. doi: 10.1074/jbc.M302994200. [DOI] [PubMed] [Google Scholar]
- 44.Bernstein DA, Zittel MC, Keck JL. High-resolution structure of the E.coli RecQ helicase catalytic core. EMBO J. 2003;22:4910–21. doi: 10.1093/emboj/cdg500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.van Brabant AJ, Ye T, Sanz M, German IJ, Ellis NA, Holloman WK. Binding and melting of D-loops by the Bloom syndrome helicase. Biochemistry. 2000;39:14617–25. doi: 10.1021/bi0018640. [DOI] [PubMed] [Google Scholar]
- 46.von Kobbe C, Thoma NH, Czyzewski BK, Pavletich NP, Bohr VA. Werner syndrome protein contains three structure-specific DNA binding domains. J Biol Chem. 2003;278:52997–3006. doi: 10.1074/jbc.M308338200. [DOI] [PubMed] [Google Scholar]
- 47.Hu JS, Feng H, Zeng W, Lin GX, Xi XG. Solution structure of a multifunctional DNA- and protein-binding motif of human Werner syndrome protein. Proc Natl Acad Sci U S A. 2005;102:18379–84. doi: 10.1073/pnas.0509380102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Huber MD, Duquette ML, Shiels JC, Maizels N. A conserved G4 DNA binding domain in RecQ family helicases. J Mol Biol. 2006;358:1071–80. doi: 10.1016/j.jmb.2006.01.077. [DOI] [PubMed] [Google Scholar]
- 49.Bujalowski W, Lohman TM. Escherichia coli single-strand binding protein forms multiple, distinct complexes with single-stranded DNA. Biochemistry. 1986;25:7799–802. doi: 10.1021/bi00372a003. [DOI] [PubMed] [Google Scholar]
- 50.Lohman TM, Overman LB. Two binding modes in Escherichia coli single strand binding protein-single stranded DNA complexes. Modulation by NaCl concentration. J Biol Chem. 1985;260:3594–603. [PubMed] [Google Scholar]
- 51.Bernstein DA, Keck JL. Domain mapping of Escherichia coli RecQ defines the roles of conserved N- and C-terminal regions in the RecQ family. Nucleic Acids Res. 2003;31:2778–85. doi: 10.1093/nar/gkg376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Zittel MC, Keck JL. Coupling DNA-binding and ATP hydrolysis in Escherichia coli RecQ: role of a highly conserved aromatic-rich sequence. Nucleic Acids Res. 2005;33:6982–91. doi: 10.1093/nar/gki999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Meyer RR, Glassberg J, Scott JV, Kornberg A. A temperature-sensitive single-stranded DNA-binding protein from Escherichia coli. J Biol Chem. 1980;255:2897–901. [PubMed] [Google Scholar]
- 54.Schultz SC, Shields GC, Steitz TA. Crystal structure of a CAP-DNA complex: the DNA is bent by 90 degrees. Science. 1991;253:1001–7. doi: 10.1126/science.1653449. [DOI] [PubMed] [Google Scholar]
- 55.Lima CD, Wang JC, Mondragon A. Three-dimensional structure of the 67K N-terminal fragment of E. coli DNA topoisomerase I. Nature. 1994;367:138–46. doi: 10.1038/367138a0. [DOI] [PubMed] [Google Scholar]
- 56.Mondragon A, DiGate R. The structure of Escherichia coli DNA topoisomerase III. Structure. 1999;7:1373–83. doi: 10.1016/s0969-2126(00)80027-1. [DOI] [PubMed] [Google Scholar]
- 57.Gajiwala KS, Burley SK. Winged helix proteins. Curr Opin Struct Biol. 2000;10:110–6. doi: 10.1016/s0959-440x(99)00057-3. [DOI] [PubMed] [Google Scholar]
- 58.Rodriguez AC, Stock D. Crystal structure of reverse gyrase: insights into the positive supercoiling of DNA. EMBO J. 2002;21:418–26. doi: 10.1093/emboj/21.3.418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Jonsson H, Peng SL. Forkhead transcription factors in immunology. Cell Mol Life Sci. 2005;62:397–409. doi: 10.1007/s00018-004-4365-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Littlefield O, Nelson HC. A new use for the ‘wing’ of the ‘winged’ helix-turn-helix motif in the HSF-DNA cocrystal. Nat Struct Biol. 1999;6:464–70. doi: 10.1038/8269. [DOI] [PubMed] [Google Scholar]
- 61.Zheng N, Fraenkel E, Pabo CO, Pavletich NP. Structural basis of DNA recognition by the heterodimeric cell cycle transcription factor E2F-DP. Genes Dev. 1999;13:666–74. doi: 10.1101/gad.13.6.666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Mer G, Bochkarev A, Gupta R, Bochkareva E, Frappier L, Ingles CJ, Edwards AM, Chazin WJ. Structural basis for the recognition of DNA repair proteins UNG2, XPA, and RAD52 by replication factor RPA. Cell. 2000;103:449–56. doi: 10.1016/s0092-8674(00)00136-7. [DOI] [PubMed] [Google Scholar]
- 63.Lohman TM, Green JM, Beyer RS. Large-scale overproduction and rapid purification of the Escherichia coli ssb gene product. Expression of the ssb gene under lambda PL control. Biochemistry. 1986;25:21–5. doi: 10.1021/bi00349a004. [DOI] [PubMed] [Google Scholar]
- 64.Markley JL, Bax A, Arata Y, Hilbers CW, Kaptein R, Sykes BD, Wright PE, Wuthrich K. Recommendations for the presentation of NMR structures of proteins and nucleic acids. J Mol Biol. 1998;280:933–52. doi: 10.1006/jmbi.1998.1852. [DOI] [PubMed] [Google Scholar]
- 65.Reiter NJ, Lee D, Wang YX, Tonelli M, Bahrami A, Cornilescu CC, Butcher SE. Resonance assignments for the two N-terminal RNA recognition motifs (RRM) of the S. cerevisiae pre-mRNA processing protein Prp24. J Biomol NMR. 2006;36(Suppl 1):58. doi: 10.1007/s10858-006-9039-4. [DOI] [PubMed] [Google Scholar]
- 66.Sattler M, Scheucher J, Griesenger C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Progress in Nuclear Magnetic Resonance Spectroscopy. 1999;34:93–158. [Google Scholar]
- 67.Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR. 1995;6:277–93. doi: 10.1007/BF00197809. [DOI] [PubMed] [Google Scholar]
- 68.Eghbalnia HR, Bahrami A, Wang L, Assadi A, Markley JL. Probabilistic Identification of Spin Systems and their Assignments including Coil-Helix Inference as Output (PISTACHIO) J Biomol NMR. 2005;32:219–33. doi: 10.1007/s10858-005-7944-6. [DOI] [PubMed] [Google Scholar]
- 69.Schweimer K, Hoffmann S, Bauer F, Friedrich U, Kardinal C, Feller SM, Biesinger B, Sticht H. Structural investigation of the binding of a herpesviral protein to the SH3 domain of tyrosine kinase Lck. Biochemistry. 2002;41:5120–30. doi: 10.1021/bi015986j. [DOI] [PubMed] [Google Scholar]
- 70.Killoran MP, Keck JL. Three HRDC domains differentially modulate Deinococcus radiodurans RecQ DNA helicase biochemical activity. J Biol Chem. 2006;281:12849–57. doi: 10.1074/jbc.M600097200. [DOI] [PubMed] [Google Scholar]
- 71.Morrical SW, Lee J, Cox MM. Continuous association of Escherichia coli single-stranded DNA binding protein with stable complexes of recA protein and single-stranded DNA. Biochemistry. 1986;25:1482–94. doi: 10.1021/bi00355a003. [DOI] [PubMed] [Google Scholar]
- 72.Pike A, Shrestha B, Popuri V, Burgess-Brown N, Muzzolini L, Costantini S, Vindigni A, Gileadi O. Structure of the human RECQ1 helicase: identification of a putative strand separation pin. Proc Natl Acad Sci USA. 2008 doi: 10.1073/pnas.0806908106. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.