Abstract
Systemic and local inflammation plays a prominent role in the pathogenesis of atherosclerotic coronary artery disease, but the relationship of whole blood gene expression changes with coronary disease remains unclear. We have investigated whether gene expression patterns in peripheral blood correlate with the severity of coronary disease and whether these patterns correlate with the extent of atherosclerosis in the vascular wall.
Patients were selected according to their coronary artery disease index (CADi), a validated angiographical measure of the extent of coronary atherosclerosis that correlates with outcome. RNA was extracted from blood of 120 patients with at least a stenosis greater than 50% (CADi≥23) and from 121 controls without evidence of coronary stenosis (CADi = 0).
160 individual genes were found to correlate with CADi (rho>0.2, P<0.003). Prominent differential expression was observed especially in genes involved in cell growth, apoptosis and inflammation. Using these 160 genes, a partial least squares multivariate regression model resulted in a highly predictive model (r2 = 0.776, P<0.0001). The expression pattern of these 160 genes in aortic tissue also predicted the severity of atherosclerosis in human aortas, showing that peripheral blood gene expression associated with coronary atherosclerosis mirrors gene expression changes in atherosclerotic arteries.
In conclusion, the simultaneous expression pattern of 160 genes in whole blood correlates with the severity of coronary artery disease and mirrors expression changes in the atherosclerotic vascular wall.
Introduction
Coronary artery disease, a multifactorial chronic disease, is the leading cause of death in Western countries. Despite considerable advances in the prevention and treatment of coronary artery disease and its complications, morbidity and mortality remains high. In half of patients with coronary artery disease, the first manifestation is death [1]. Consequently, substantial efforts are being put into the development of new strategies for accurate noninvasive diagnosis of coronary artery disease and the identification of novel treatment targets [2].
Systemic and local inflammation has been shown to play a prominent pathologic role in atherosclerotic coronary artery disease [3]. Adhesion of leukocytes to activated endothelial cells and their migration into the arterial wall are thought to initiate, propagate, and destabilize coronary plaques. All types of blood constituents appear to play a role in plaque formation, although the majority of inflammatory lesions in atherosclerotic vascular tissue consist of foam cell macrophages and activated T-cells [4]. Several studies have found distinct gene expression patterns in atherosclerotic arteries [5]–[8]. While other pathways are likely also important, a consistent feature has been differential expression of inflammatory genes and genes involved in cell cycle control [9]–[12].
Microarray analysis of peripheral blood cells is a practical approach to study gene expression changes that may reflect not only genetic predisposition but also presence and activity of disease, environmental modifier effects, and treatment responses [13]. Total peripheral leukocyte count correlates with the severity of coronary atherosclerosis and is a strong predictor of cardiovascular outcome [14], but little is known about the role of phenotypic changes in circulating blood cells of patients with coronary atherosclerosis. In a recent micro-array analysis, 526 genes were found to be differentially expressed in isolated mononuclear cells from 41 patients [15]. Gene expression patterns of 50 of these genes together with 56 genes selected from the literature were subsequently shown to be associated with the presence of coronary artery disease in two independent cohorts. The aim of the present study was 1) to identify distinct genomic markers in peripheral whole blood that correlate with the severity of coronary artery disease using micro-array analysis and 2) to investigate to what extent gene expression patterns in peripheral blood mirror those in atherosclerotic arteries.
Methods
Patient Selection and Characteristics
Patients and control subjects were recruited from individuals that had undergone catheterization in the Duke University Hospital Cardiac Catheterization Laboratory and participated in a proteomics study to discover candidate proteins that are differentially displayed in populations with and those without angiographic coronary artery disease [16]. After being approached and providing informed written consent, subjects had clinical and laboratory data collected. The investigation conforms to the principles outlined in the Declaration of Helsinki, and was approved by the Duke Institutional Review Board.
Patient selection, design and results from the main proteomics study have been reported previously [16]. Populations were initially defined in order to minimize differences in plasma proteins unrelated to the presence or absence of coronary artery disease. As a practical strategy, three different cohorts of subjects (cases and controls) were enrolled: 1) matched men (n = 106), who were matched for age and ethnic group, 2) unmatched men (n = 82), who did not fulfill the matching criteria and 3) unmatched women (n = 53). The severity of coronary artery disease was scored using the Duke Coronary Artery Disease Index (CAD-Index) [17], [18]. The CAD-index is a prognostic assessment of the extent of coronary artery disease, accounting for the number and severity of lesions and diseased vessels and involvement of left anterior descending and left main disease.
Inclusion criteria for the coronary artery disease patient population (cases) were: age between 40 and 65 and coronary artery stenosis of >50% in at least one major coronary artery. Inclusion criteria for the control population (controls) were: age between 40 and 65 for matched men cohort only, no angiographically detectable coronary artery stenosis on cardiac catheterization within the last two years, normal left ventricular ejection fraction and normal regional wall motion. Exclusion criteria for controls were typical signs of angina, or any history or evidence of myocardial ischemia on stress testing, myocardial infarction or unstable angina, any history of peripheral arterial or cerebrovascular disease, or significant vascular stenosis on noninvasive imaging or angiography. Exclusion criteria also included myocardial infarction within one month (for cases), diabetes, uncontrolled hypertension (systolic blood pressure >180 mmHg or diastolic blood pressure >100 mmHg) or with end-organ damage, renal insufficiency (creatinine >2.0 mg/dL or BUN>40 mg/dL), active malignancy, significant valvular heart disease, NYHA Class III or IV heart failure, cigarette smoking >2 packs per day, total cholesterol >300 mg/dL or triglyceride >400 mg/dL, anemia (hemoglobin <12.5 g/dL for females or <13.5 g/dL for males), and hypotension (systolic blood pressure <90 mmHg and diastolic blood pressure <50 mmHg).
Blood Sampling and Gene Expression Analysis
The blood samples (2.5 mL) were collected in PAXgene™ Blood RNA tubes and total RNA was isolated using the standardized RNA Kit (PreAnalytiX, Qiagen) [19]. RNA isolation started with a centrifugation step to pellet nucleic acids in the PAXgene Blood RNA Tube. The pellet was then washed, and Proteinase K added to digest proteins. Alcohol was added to adjust binding conditions, and the sample was applied to a PAXgene RNA spin column. During a brief centrifugation, RNA selectively bound to the PAXgene silica-gel membrane and eluted using an optimized buffer.
RNA was then quantified by absorbance at A260 nm and the purity was estimated by the ratio A260 nm/A280 nm. RNA integrity was confirmed by non-denaturing agarose gel electrophoresis. RNA was stored at −80°C until further analysis. The quality of 19 RNA samples was insufficient for microarray analysis due to degradation. The genomic studies were conducted in the Novartis Genomics Factory, Basel, Switzerland.
Genome-wide transcript profiling was assessed using human HGU133A oligonucleotide expression probe arrays (Affymetrix, Santa Clara, CA, U.S.A.), comprising 22,483 probe sets. The experiments were done according to the recommendations of the manufacturer [20]. Data was normalized using MAS5 (Affymetrix); the data is publicly available at the Gene Expression Omnibus (GEO) repository (accession number GSE12288, http:/www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE12288). As quality control, RT-PCR was performed on 8 selected genes in 2×20 subjects from the ‘matched men’ cohort.
Independent Evaluation of Predictive Gene Model in Human Aorta Tissue
To test whether the expression pattern in peripheral whole blood is representative for atherosclerosis in general, we have examined the capability of expression of genes derived from the peripheral blood cell study to predict the severity of atherosclerosis in human aortas. Gene expression data was generated using RNA extracted from a unique collection of freshly harvested human aortas with varying degrees of atherosclerosis (n = 67 donors). Donor identification, RNA extraction and micro-array methods (Affymetrix U95Av2) as well as gene expression signatures that differentiate between atherosclerotic disease states in human aortas have been reported previously [8]. As indicated in the original report, disease extent (normal, intermediate, severe) was scored by combining Sudan IV staining and raised lesion data. The “normal” or minimally diseased group showed no Sudan IV staining and contained no raised lesions, while the “intermediate” group showed more than 20% Sudan IV staining but contained no raised lesions. The “severe” group contained raised lesions covering more than 10% of the surface. We identified 20 normal, 25 intermediate and 22 severely diseased sections for this analysis.
Statistical Methods
Spearman rank correlation between CAD-index and gene expression was calculated (Partek Genomics Suite Version 6.3). An absolute correlation coefficient (rho) >0.2 was considered clinically relevant, corresponding to a p-value of 0.003 (n = 222). Among the 22,483 probe sets of the Affymetrix HGU133A chip, about 60 probe sets can be expected to have an absolute rho>0.2 by chance (false positives). Student's t test, parametric correlation and rank correlation according to Spearman were performed with the statistical software package S-Plus Version 6.
Projections to Latent Structures (PLS) analysis including Orthogonal Signal Correction (OSC) (SIMCA-P Version 10.0) was used to identify gene sets that discriminate between increasing CAD-indices or the three classes (normal, intermediate and severe) of atherosclerosis in the aorta samples. To reduce gene selection bias, models were subsequently repeatedly built based on data from two cohorts to predict CAD in the third cohort. In addition, extensive cross-validation by leave-one-out technique and validation by response permutation was applied to 7 groups of approximately 32 subjects to reduce bias in creating a predictive gene set.
Results
Patient Demographics
Demographic data, medical history and medication of the study population are summarized in table 1. A history of hypertension was significantly more common in the cases. Aspirin, statins, and blood pressure lowering agents were more frequently taken by the cases. All controls had no angiographically significant coronary artery disease (CAD-Index = 0). Within the cases, however, there was a wide distribution, with 81% of cases having a CAD-Index between 25 and 63. Although most cases (93%) had at least two-vessel disease or severe single-vessel disease, the distribution of cases is skewed towards the lower end of CAD-Index.
Table 1. Demographics and baseline characteristics.
| Matched Men | Unmatched Men | Unmatched Women | |||||||
| Controls | Cases | P | Controls | Cases | P | Controls | Cases | P | |
| n = 53 | n = 53 | n = 38 | n = 44 | n = 29 | n = 24 | ||||
| Age at time of study (mean±SD) | 52±7 | 53±6 | 0.77 | 51±8 | 58±7 | <.001 | 52±7 | 54±8 | 0.27 |
| Age at time of study (median, 25th–75th) | 52 (49–57) | 52 (48–57) | 50 (46–58) | 55 (54–63) | 56 (47–56) | 54 (50–60) | |||
| Age at last catheterization (mean±SD) | 51±6 | 51±7 | 0.51 | 49±8 | 56±7 | <.001 | 50±7 | 53±7 | 0.23 |
| Age at last catheterization (median, 25th–75th) | 50 (47–55) | 51 (47–56) | 48 (44–57) | 54 (52–62) | 50 (45–54) | 53 (49–57) | |||
| Ethnicity (Caucasian/African-American/Asian/Hispanic/Native Am) | 50/3/0/0/0 | 50/3/0/0/0 | 23/13/1/1/0 | 39/4/0/0/1 | 21/5/0/1/2 | 18/4/1/0/1 | |||
| Smoking, n (%) | 26 (49) | 29 (55) | 0.41 | 20 (53) | 34 (77) | <.001 | 9 (31) | 13 (54) | 0.02 |
| Diabetes, n (%) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | |||
| Hypertension, n (%) | 16 (30) | 27 (51) | 0.001 | 15 (39) | 27 (61) | <.001 | 13 (45) | 12 (50) | 0.30 |
| Myocardial infarction, n (%) | 0 (0) | 29 (55) | 2 (5) | 21 (48) | <.001 | 1 (3) | 10 (14) | <.001 | |
| PCI, n (%) | 0 (0) | 15 (28) | 0 (0) | 13 (30) | 0 (0) | 4 (20) | |||
| CABG, n (%) | 0 (0) | 15 (28) | 0 (0) | 15 (34) | 0 (0) | 7 (29) | |||
| Peripheral vascular disease, n (%) | 0 (0) | 2 (4) | 0 (0) | 5 (11) | 0 (0) | 3 (13) | |||
| Cerebrovascular disease, n (%) | 0 (0) | 1 (2) | 0 (0) | 4 (9) | 0 (0) | 3 (13) | |||
| Congestive heart failure, n (%) | 0 (0) | 6 (11) | 0 (0) | 4 (9) | 0 (0) | 2 (8) | |||
| Body Weight (kg) (mean±SD) | 97±19 | 94±16 | 0.40 | 101±19 | 94±30 | 0.27 | 89±27 | 84±24 | 0.43 |
| Systolic blood pressure (mmHg) (mean±SD) | 139±19 | 134±21 | 0.19 | 145±17 | 142±26 | 0.59 | 145±23 | 138±23 | 0.32 |
| Diastolic blood pressure (mmHg) (mean±SD) | 80±11 | 76±17 | 0.12 | 82±10 | 82±15 | 0.97 | 77±12 | 70±14 | 0.06 |
| LV ejection fraction (%) (mean±SD) | 64±7 | 56±11 | <.001 | 64±8 | 57±12 | 0.004 | 66±6 | 57±12 | 0.001 |
| Medication | |||||||||
| Aspirin, n (%) | 18 (34) | 47 (89) | <.001 | 10 (26) | 42 (95) | <.001 | 6 (21) | 21 (88) | <.001 |
| ACE inhibitor, n (%) | 5 (9) | 40 (75) | <.001 | 7 (18) | 26 (59) | <.001 | 4 (14) | 13 (54) | <.001 |
| ARB, n (%) | 3 (6) | 0 (0) | 0.07 | 2 (5) | 5 (11) | 0.03 | 0 (0) | 5 (21) | |
| Beta blocker, n (%) | 13 (25) | 43 (81) | <.001 | 6 (16) | 37 (84) | <.001 | 6 (21) | 20 (83) | <.001 |
| Calcium blocker, n (%) | 4 (8) | 10 (19) | 0.002 | 5 (13) | 9 (20) | 0.07 | 3 (10) | 4 (17) | 0.19 |
| Statin, n (%) | 10 (19) | 38 (72) | <.001 | 4 (11) | 29 (66) | <.001 | 5 (17) | 15 (63) | <.001 |
| Fibrate, n (%) | 2 (4) | 4 (8) | 0.15 | 0 (0) | 8 (18) | 0 (0) | 1 (4) | ||
| Clinical Laboratory parameters | |||||||||
| Total Cholesterol (mg/dL) (mean±SD) | 196±29 | 167±32 | <0.01 | 195±39 | 177±39 | 0.06 | 204±39 | 181±52 | 0.12 |
| Triglycerides (mg/dL) (mean±SD) | 183±120 | 142±71 | 0.05 | 155±96 | 181±142 | 0.40 | 146±67 | 161±76 | 0.50 |
| LDL Cholesterol (mg/dL) (mean±SD) | 117±22 | 100±31 | <0.01 | 117±35 | 102±40 | 0.10 | 119±31 | 100±40 | 0.10 |
| HDL Cholesterol (mg/dL) (mean±SD) | 44±11 | 39±9 | 0.03 | 47±12 | 42±9 | 0.09 | 56±21 | 49±16 | 0.24 |
| HbA1c (%) (mean±SD) | 5.4±0.4 | 5.5±0.5 | 0.30 | 5.8±1.1 | 5.7±0.8 | 0.92 | 5.6±0.6 | 5.6±0.6 | 0.81 |
| Creatinine (mg/dL) (%) (mean±SD) | 1.0±0.1 | 1.1±0.1 | 0.11 | 1.1±0.1 | 1.1±0.2 | 0.89 | 0.8±0.2 | 1.0±0.5 | 0.12 |
| Hematocrit (%) (mean±SD) | 44±5 | 43±3 | 0.15 | 43±2 | 43±3 | 0.89 | 41±3 | 40±2 | 0.03 |
| White blood cell count (109/L) (mean±SD) | 5.6±1.3 | 6.2±2.1 | 0.06 | 5.9±1.7 | 6.4±1.9 | 0.27 | 6.7±2.3 | 6.9±2.1 | 0.51 |
Clinical laboratory parameters were available for all subjects (table 1). Hematocrit and white blood cell counts were not significantly different. Total cholesterol and LDL-cholesterol levels were significantly lower in the coronary artery disease group, probably reflecting a higher use of statins.
Prediction of Coronary Disease Using Risk Factors and Biochemical Markers
Traditional risk factors, including body weight, smoking, and systolic and diastolic blood pressure did not correlate significantly with the extent of coronary disease in a rank correlation analysis. Total cholesterol (but not LDL-cholesterol) level was found to be inversely related with the CAD-index (rho = −0.41, P<0.0001), which may in part reflect the higher use of statins and better blood lipid control in cases. In addition, other parameters were found to positively (potassium, blood urea nitrogen, phosphorus and osmolarity) or negatively (calcium and HDL-cholesterol) correlate with CAD-index (rho>0.2). Of note, important clinical markers such as LDL-C (rho = 0.02), CRP (rho = −0.12) and homocysteine (rho = 0.02) exhibited a poor correlation with CAD-index, which could result from treatment of affected individuals. In a multivariate correlation analysis, the combination of risk factors and biochemical markers only poorly predicted the extent of coronary artery disease (r2 = 0.228).
Gene Expression
Gene expression data from 222 out of 241 subjects were available for this analysis (110/121 cases and 112/120 controls); RNA from the remaining 19 subjects did not pass quality control due to degradation.
In a univariate analysis, 160 genes were found to correlate with CAD-Index with an absolute rank correlation coefficient (rho) >0.2 (P<0.003). All probesets correlating with CAD-Index are listed in table 2. Most of these genes are known to be involved in hematopoietic cell differentiation, cell growth or growth arrest, apoptosis, cell adhesion, matrix modulation and inflammatory and immune response, processes known to modulate atherosclerosis.
Table 2. List of 160-gene model predictive of the extent of coronary artery disease.
| Symbol | U133A ID | 95Av2 ID | Name | Pathway | Rho |
| AIF1 | 207823_s_at | 37011_at 33641_g_at | allograft inflammatory factor 1 | Angiogenesis | 0.21 |
| 3640_at 37764_at | Inflammatory response | ||||
| MMP19° | 204575_s_at | matrix metalloproteinase 19 | Angiogenesis | 0.22 | |
| Response to metal ion | |||||
| Extracellular matrix modulation | |||||
| EPIM | 207346_at | epimorphin | Angiogenesis | 0.22 | |
| MMP24°° | 78047_s_at | Matrix metalloproteinase 24 (membrane-inserted) | Angiogenesis | 0.20 | |
| Response to metal ion | |||||
| Extracellular matrix modulation | |||||
| CRADD° | 209833_at | 822_s_at 1211_s_at | CASP2 and RIPK1 domain containing adaptor with death domain | Apoptosis | 0.25 |
| WDR13° | 222138_s_at | 727_at | WD repeat domain 13 | Apoptosis | 0.24 |
| PDE4D | 210837_s_at | 38526_at | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | Apoptosis | 0.22 |
| AK2°° | 212174_at | 40789_at 40788_at | adenylate kinase 2 | Apoptosis | 0.28 |
| FOLH1° | 217487_x_at | 1740_g_at 1739_at 1655_s_at | folate hydrolase (prostate-specific membrane antigen) 1 | Apoptosis | 0.22 |
| TGM5 | 207911_s_at | 33001_s_at | transglutaminase 5 | Apoptosis | 0.24 |
| P53AIP1° | 220403_s_at | p53-regulated apoptosis-inducing protein 1 | Apoptosis | 0.25 | |
| NALP1 | 211822_s_at | NACHT, leucine rich repeat and PYD (pyrin domain) containing 1 | Apoptosis | 0.26 | |
| Inflammatory response | |||||
| LGALS9° | 203236_s_at | 766_at 38091_at | galectin 9 | Cell adhesion | 0.25 |
| ICAM1° | 202637_s_at | 32640_at | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | Cell adhesion | 0.21 |
| PCDHGC3 | 205717_x_at | 657_at 35609_at 1691_at 1690_at 1169_at | protocadherin gamma subfamily C, 3 | Cell adhesion | 0.20 |
| GPLD1 | 206265_s_at | 934_at 1293_s_at | glycosylphosphatidylinositol specific phospholipase D1 | Cell adhesion | 0.21 |
| T/B cell proliferation | |||||
| CDH11° | 207173_x_at | 36976_at 2087_s_at | cadherin 11, type 2, OB-cadherin (osteoblast) | Cell adhesion | 0.24 |
| DSC3 | 206032_at | 32417_at | desmocollin 3 | Cell adhesion | 0.20 |
| Cytoskeleton | |||||
| LAMB3° | 209270_at | 36929_at | laminin, beta 3 | Cell adhesion | 0.25 |
| PKP4 | 214874_at | 33475_at | plakophilin 4 | Cell adhesion | 0.22 |
| Cytoskeleton | |||||
| FN1 | 214702_at | Fibronectin 1 | Cell adhesion | 0.21 | |
| IIp45° | 48659_at | IGFBP-2-Binding Protein, IIp45 (FLJ12438) | Cell adhesion | 0.22 | |
| PINK1°° | 209018_s_at | 35361_at | PTEN induced putative kinase 1 | Cell growth & growth arrest | 0.22 |
| Apoptosis | |||||
| FKBP8°° | 40850_at | 40850_at | FK506 binding protein 8, 38kDa | Cell growth & growth arrest | 0.25 |
| UBXD1°° | 220757_s_at | UBX domain-containing protein 1 | Cell growth & growth arrest | 0.21 | |
| RXRA° | 202426_s_at | 405_at 32800_at | retinoid X receptor, alpha | Cell growth & growth arrest | 0.24 |
| Apoptosis | |||||
| RIS1° | 213338_at | 35692_at | Ras-induced senescence 1 | Cell growth & growth arrest | 0.28 |
| NFYC° | 202215_s_at | 40466_at | nuclear transcription factor Y, gamma | Cell growth & growth arrest | 0.30 |
| CLN3 | 209275_s_at | 497_at | ceroid-lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | Cell growth & growth arrest | 0.27 |
| Apoptosis | |||||
| RARA | 211605_s_at | 1337_s_at | retinoic acid receptor, alpha | Cell growth & growth arrest | 0.26 |
| HCFC1 | 202473_x_at | 37910_at | host cell factor C1 (VP16-accessory protein) | Cell growth & growth arrest | 0.23 |
| PSG3 | 203399_x_at | 40857_f_at | pregnancy specific beta-1-glycoprotein 3 | Cell growth & growth arrest | 0.22 |
| STAU2° | 204226_at | 38341_at 32386_at | staufen, RNA binding protein, homolog 2 (Drosophila) | Cell growth & growth arrest | 0.26 |
| ELAVL2 | 208427_s_at | 36411_s_at 36410_f_at | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 2 (Hu antigen B) | Cell growth & growth arrest | 0.25 |
| TP53I11° | 214667_s_at | 36136_at | tumor protein p53 inducible protein 11 | Cell growth & growth arrest | 0.31 |
| NPR3 | 219789_at | 34519_at | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | Cell growth & growth arrest | 0.21 |
| Angiogenesis | |||||
| PTP4A1° | 200730_s_at | 843_at 33413_at | protein tyrosine phosphatase type IVA, member 1 | Cell growth & growth arrest | 0.27 |
| STC2 | 203439_s_at | 32043_at | stanniocalcin 2 | Cell growth & growth arrest | 0.24 |
| Response to metal ion | |||||
| SEMA3C | 203788_s_at | 377_g_at 376_at | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3C | Cell growth & growth arrest | 0.28 |
| Immune response | |||||
| CCNA1 | 205899_at | 1914_at | cyclin A1 | Cell growth & growth arrest | 0.20 |
| PTPRR° | 206084_at | 1658_g_at 1657_at | protein tyrosine phosphatase, receptor type, R | Cell growth & growth arrest | 0.23 |
| LHX2 | 206140_at | 40528_at | LIM homeobox 2 | Cell growth & growth arrest | 0.21 |
| T/B cell proliferation | |||||
| CPSF4 | 206688_s_at | 35743_at | cleavage and polyadenylation specific factor 4, 30kDa | Cell growth & growth arrest | 0.22 |
| Inflammatory response | |||||
| I–4° | 207377_at | 31735_at | type 1 protein phosphatase inhibitor | Cell growth & growth arrest | 0.21 |
| MEIS2 | 207480_s_at | 41388_at | Meis1, myeloid ecotropic viral integration site 1 homolog 2 (mouse) | Cell growth & growth arrest | 0.20 |
| NF2° | 211092_s_at | 38007_at 1894_f_at | neurofibromin 2 (bilateral acoustic neuroma) | Cell growth & growth arrest | 0.23 |
| Cytoskeleton | |||||
| BRRN1° | 212949_at | 41639_at | barren homolog (Drosophila) | Cell growth & growth arrest | 0.32 |
| CDC42 | 214230_at | 960_g_at 959_at 39736_at | cell division cycle 42 (GTP binding protein, 25kDa) | Cell growth & growth arrest | 0.23 |
| ZMYND10° | 216663_s_at | 32993_s_at | zinc finger, MYND domain containing 10 | Cell growth & growth arrest | 0.25 |
| CROC4 | 222301_at | 40483_at 40482_s_at | transcriptional activator of the c-fos promoter | Cell growth & growth arrest | 0.22 |
| PPP2R5B° | 635_s_at | 635_s_at | protein phosphatase 2, regulatory subunit B (B56), beta isoform | Cell growth & growth arrest | 0.20 |
| TRIM45° | 219923_at | tripartite motif-containing protein 45 | Cell growth & growth arrest | 0.23 | |
| TDRKH | 221053_s_at | tudor and KH domain containing | Cell growth & growth arrest | 0.26 | |
| PB1° | 221212_x_at | polybromo 1 | Cell growth & growth arrest | 0.25 | |
| NEIL1 | 219396_s_at | nei endonuclease VIII-like 1 | Cell growth & growth arrest | 0.31 | |
| PMS2L5° | 179_at | Postmeiotic segregation increased 2-like 5 | Cell growth & growth arrest | 0.24 | |
| BLOC1S1° | 202592_at | Biogenesis of lysosome-related organelles complex-1, subunit 1 | Cell growth & growth arrest | 0.23 | |
| BRF2° | 218955_at | subunit of RNA polymerase III transcription initiation factor, BRF1-like | Cell growth & growth arrest | 0.23 | |
| ASNA1°° | 202024_at | Arsenical pump-driving ATPase | Cell growth & growth arrest | 0.21 | |
| SIRT5°° | 221010_s_at | sirtuin (silent mating type information regulation 2 homolog) 5 | Cell growth & growth arrest | 0.21 | |
| HIST1H4G | 208551_at | histone 1, H4g | Cell growth & growth arrest | 0.27 | |
| SLD5 | 211767_at | SLD5 homolog | Cell growth & growth arrest | 0.27 | |
| MAN2A2° | 202032_s_at | 41766_at 38188_s_at | mannosidase, alpha, class 2A, member 2 | Cell-cell interaction | 0.28 |
| GJB3° | 215243_s_at | 41076_at | gap junction protein, beta 3, 31kDa (connexin 31) | Cell-cell interaction | 0.24 |
| PLXNA2 | 207290_at | 40395_at | plexin A2 | Cell-cell interaction | 0.21 |
| ADH1B | 209613_s_at | 35730_at | alcohol dehydrogenase IB (class I), beta polypeptide | Cellular metabolism | 0.22 |
| Immune response | |||||
| FGA | 205650_s_at | 38825_at | fibrinogen, A alpha polypeptide | Coagulation | 0.26 |
| Cell adhesion | |||||
| Inflammatory response | |||||
| SERPINB8° | 206034_at | 36312_at | serine (or cysteine) proteinase inhibitor, clade B (ovalbumin), member 8 | Coagulation | 0.23 |
| Inflammatory response | |||||
| TUBA3° | 212639_x_at | 32272_at | tubulin alpha 3 | Cytoskeleton | 0.28 |
| Apoptosis | |||||
| ADD1° | 214726_x_at | 32146_s_at 32145_at | adducin 1 (alpha) | Cytoskeleton | 0.21 |
| ARHGAP4 | 204425_at | 39649_at | Rho GTPase activating protein 4 | Cytoskeleton | 0.23 |
| LLGL1°° | 206123_at | 804_s_at 33200_at | lethal giant larvae homolog 1 (Drosophila) | Cytoskeleton | 0.22 |
| Cell growth & growth arrest | |||||
| KRT6A | 209125_at | 39016_r_at 39015_f_at | keratin 6A | Cytoskeleton | 0.28 |
| SMPX° | 219772_s_at | small muscle protein, X-linked | Cytoskeleton | 0.20 | |
| STXBP2°° | 209367_at | 38259_at | syntaxin binding protein 2 | Cytoskeleton - Exocytosis | 0.25 |
| PLAUR°° | 211924_s_at | 189_s_at | plasminogen activator, urokinase receptor | Extracellular matrix modulation | 0.20 |
| Inflammatory response | |||||
| COL13A1° | 211809_x_at | collagen, type XIII, alpha 1 | Extracellular matrix modulation | 0.25 | |
| ABCC6 | 214033_at | Up-regulated gene 7 | Extracellular matrix modulation | 0.26 | |
| ITPK1°° | 210740_s_at | 35755_at | inositol 1,3,4-triphosphate 5/6 kinase | Hematopoietic cell differentation | 0.30 |
| FTL° | 212788_x_at | 35083_at | ferritin, light polypeptide | Hematopoietic cell differentation | 0.24 |
| Response to metal ion | |||||
| FANCC° | 205189_s_at | 35713_at 160034_s_at | Fanconi anemia, complementation group C | Hematopoietic cell differentation | 0.23 |
| Cell growth & growth arrest | |||||
| Inflammatory response | |||||
| Apoptosis | |||||
| SMAD5 | 205187_at | 39926_at 1952_s_at 1013_at | SMAD, mothers against DPP homolog 5 (Drosophila) | Hematopoietic cell differentation | 0.22 |
| NOTCH2° | 202445_s_at | 38083_at | Notch homolog 2 (Drosophila) | Hematopoietic cell differentation | 0.20 |
| Angiogenesis | |||||
| GATA1° | 210446_at | 36787_at | GATA binding protein 1 (globin transcription factor 1) | Hematopoietic cell differentation | 0.25 |
| IL9R° | 217212_s_at 208164_s_at | 938_at | interleukin 9 receptor | Hematopoietic cell differentation | 0.20 0.24 |
| CRSP2° | 202612_s_at | Cofactor required for Sp1 transcriptional activation subunit 2 | Hematopoietic cell differentation | 0.24 | |
| NOX4 | 219773_at | NADPH oxidase 4 | Hematopoietic cell differentation | 0.23 | |
| L3MBTL° | 206822_s_at | Lethal(3)malignant brain tumor-like protein | Hematopoietic cell differentation | 0.32 | |
| RNF24 | 210706_s_at | Ring finger 24 | Hematopoietic cell differentation | 0.28 | |
| KLF3°° | 219657_s_at | Kruppel-like factor 3 (basic) | Hematopoietic cell differentation | 0.21 | |
| MARCH2°° | 210075_at | 39910_at | membrane-associated ring finger (C3HC4) 2 | Immune response | 0.24 |
| GPSM3° | 214847_s_at | 39049_at | G-protein signalling modulator 3 (AGS3-like, C. elegans) | Immune response | 0.27 |
| IGSF4B | 213948_x_at | 39288_at | immunoglobulin superfamily, member 4B | Immune response | 0.25 |
| PRG3° | 220811_at | proteoglycan 3 | Immune response | 0.24 | |
| AGPAT1° | 32836_at | 32836_at | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | Inflammatory response | 0.29 |
| Lipid biosynthesis | |||||
| EPHB2° | 211165_x_at | 902_at 41678_at 2088_s_at | EPH receptor B2 | Inflammatory response | 0.22 |
| Cell-cell interaction | |||||
| PRKAR1B° | 212559_at | 1091_at | protein kinase, cAMP-dependent, regulatory, type I, beta | Inflammatory response | 0.25 |
| T/B cell proliferation | |||||
| PAFAH1B1° | 200815_s_at | 32569_at | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | Inflammatory response | 0.27 |
| NFX1 | 202585_s_at | 34667_at | nuclear transcription factor, X-box binding 1 | Inflammatory response | 0.26 |
| KCNMB1°° | 209948_at | 38298_at | potassium large conductance calcium-activated channel, subfamily M, beta member 1 | Ion channel | 0.22 |
| CHRNA5 | 206533_at | 36397_at | cholinergic receptor, nicotinic, alpha polypeptide 5 | Ion channel | 0.21 |
| Neurotransmission | |||||
| SAH° | 210377_at | 33279_s_at 33278_at | SA hypertension-associated homolog (rat) | Lipid metabolism | 0.30 |
| HLCS | 207833_s_at | 37764_at | holocarboxylase synthetase (biotin-[proprionyl-Coenzyme A-carboxylase (ATP-hydrolysing)] ligase) | Metabolic homeostasis | 0.21 |
| MAOA | 204388_s_at | 41772_at 41771_g_at 41770_at | monoamine oxidase A | Neurotransmission | 0.28 |
| GABRA6° | 207182_at | 34025_at | gamma-aminobutyric acid (GABA) A receptor, alpha 6 | Neurotransmission | 0.23 |
| CEPBA°° | 204039_at | 32550_r_at | CCAAT/enhancer binding protein (C/EBP), alpha | Progenitor cell differentiation | 0.35 |
| Cell growth & growth arrest | |||||
| SOX4° | 201416_at | 33131_at | SRY (sex determining region Y)-box 4 | Progenitor cell differentiation | 0.20 |
| ZNF305 | 206507_at | 37083_s_at 37082_at | zinc finger protein 305 | Progenitor cell differentiation | 0.21 |
| ZNFN1A2° | 220567_at | zinc finger protein, subfamily 1A, 2 | Progenitor cell differentiation | 0.23 | |
| ZNF3° | 219605_at | zinc finger protein 3 (A8–51) | Progenitor cell differentiation | 0.22 | |
| Response to metal ion | |||||
| Immune response | |||||
| CAPN5° | 205166_at | 38504_at | calpain 5 | Response to injury | 0.25 |
| Cell growth & growth arrest | |||||
| MTF1 | 205323_s_at | 38945_at | metal-regulatory transcription factor 1 | Response to metal ion | 0.25 |
| CA12° | 203963_at | 36454_at | carbonic anhydrase XII | Response to metal ion | 0.26 |
| NEDD4L° | 212445_s_at | 39356_at | neural precursor cell expressed, developmentally down-regulated 4-like | Response to metal ion | 0.22 |
| CABIN1° | 202624_s_at | 37652_at | calcineurin binding protein 1 | T/B cell proliferation | 0.21 |
| SH3BP2° | 209370_s_at | 1303_at | SH3-domain binding protein 2 | T/B cell proliferation | 0.23 |
| Immune response | |||||
| TNFRSF5 | 35150_at | 35150_at 35149_at | tumor necrosis factor receptor superfamily, member 5 | T/B cell proliferation | 0.22 |
| Inflammatory response | |||||
| Immune response | |||||
| PBX2°° | 211097_s_at | 38295_at | pre-B-cell leukemia transcription factor 2 | T/B cell proliferation | 0.25 |
| IL2 | 207849_at | interleukin 2 | T/B cell proliferation | 0.22 | |
| Cell growth & growth arrest | |||||
| Immune response | |||||
| HDAC5°° | 202455_at | histone deacetylase 5 | T/B cell proliferation | 0.28 | |
| Inflammatory response | |||||
| PIP° | 206509_at | 41094_at 325_s_at | prolactin-induced protein | T/B cell regulation | 0.23 |
| GAD2° | 216651_s_at | 32280_at 32279_at | glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) | T/B cell regulation | 0.21 |
| PNPLA2°° | 39854_r_at | 39854_r_at | patatin-like phospholipase domain containing 2 | Triglyceride homeostasis | 0.24 |
| MGLL° | 211026_s_at | 35792_at | monoglyceride lipase | Triglyceride homeostasis | 0.27 |
| MJD | 216657_at | 36819_at | Machado-Joseph disease (spinocerebellar ataxia 3, olivopontocerebellar ataxia 3, autosomal dominant, ataxin 3) | Ubiquitination | 0.2 |
| FBXO31 | 219784_at | F-box only protein 31 | Ubiquitylation | 0.26 | |
| GGA3° | 211815_s_at | 37959_at | golgi associated, gamma adaptin ear containing, ARF binding protein 3 | Ubiquitylation | 0.26 |
| N4BP1° | 48612_at | Nedd4 binding protein 1 | Ubiquitylation | 0.21 | |
| BC002942 | 31837_at | 31837_at | hypothetical protein BC002942 | 0.28 | |
| MGC21416° | 212340_at | 37891_a | hypothetical protein MGC21416 | 0.27 | |
| DKFZp586F1822 | 37891_at | DKFZp586F1822 | |||
| CDRT1 | 215999_at | 31781_at | CMT1A duplicated region transcript 1 | 0.23 | |
| KIAA0241° | 212475_at | 39761_at | KIAA0241 protein | 0.28 | |
| 13CDNA73° | 214319_at | 33276_at | hypothetical protein CG003 | 0.24 | |
| SH3TC1° | 219256_s_at | SH3 domain and tetratricopeptide repeats 1 | 0.28 | ||
| RER1° | 213114_at | RER1 homolog (S. cerevisiae) | 0.26 | ||
| DKFZp564M0616 | 215763_at | cDNA DKFZp564M0616 | 0.26 | ||
| KIAA0882° | 212960_at | KIAA0882 | 0.22 | ||
| FLJ11155 | 219750_at | hypothetical protein FLJ11155 | 0.23 | ||
| LOC51145 | 220752_at | erythrocyte transmembrane protein | 0.23 | ||
| SEC61A2 | 219499_at | Protein transport protein Sec61 alpha subunit 2 | 0.20 | ||
| C12orf4° | 218374_s_at | chromosome 12 open reading frame 4 | 0.24 | ||
| STYK1 | 221696_s_at | serine/threonine/tyrosine kinase 1 | 0.22 | ||
| FLJ20674 | 220137_at | hypothetical protein FLJ20674 | 0.24 | ||
| PRO1693 | 221137_at | PRO1693 | 0.22 | ||
| FLJ12058°° | 215971_at | FLJ12058 fis, clone HEMBB1002092 | 0.21 | ||
| FLJ10305° | 216501_at | hypothetical protein FLJ10305 | 0.25 | ||
| DKFZP434O047 | 208008_at | DKFZP434O047 protein | 0.25 | ||
| FLJ10970 | 219230_at | hypothetical protein FLJ10970 | 0.21 | ||
| FLJ22209 | 216450_x_at | cDNA: FLJ22209 fis, clone HRC01496 | 0.20 | ||
| FLJ11996 | 207487_at | hypothetical protein FLJ11996 | 0.24 | ||
| FLJ20241° | 207083_s_at | putative NFkB activating protein | 0.24 | ||
| FLJ14220° | 219310_at | FLJ14220, chromosome 20 open reading frame 39 | 0.21 | ||
| SYNGR1° | 213854_at | synaptogyrin 1 | 0.22 | ||
| FLJ11871° | 220915_s_at | FLJ11871, DKFZp686I0814 | 0.24 | ||
| LRCH4 | 204692_at | Leucine rich repeat neuronal 4 | 0.26 | ||
| PRO2533 | 220787_at | 0.23 | |||
| EST | 222308_x_at | 0.24 | |||
| EST | 222302_at | 0.22 | |||
| EST | 215906_at | 0.28 |
The 8 most predictive genes according to the VIP analysis are listed in bold; °° indicated the 19 genes and °or°° the 90 genes that were found to be differentially expressed in a multiway ANOVA.
Using log-transformed data with signal intensities >80, only 19 probesets were found to be significantly differentially expressed in a multiway ANOVA (smoking, age, gender, cohort, race, CAD (i.e. case vs. control) and CAD-index as fixed factors or random effects, respectively) (table 2). However, when only taking the 20 controls with the least predicted CAD versus the 20 cases with the most predicted disease into account, a formal comparison yielded 90 out of the 160 probesets with statistically significant differential expression (p<0.05, no adjustment for multiple comparisons) (table 2). rt-PCR confirmed the Affymetrix results for 7 of the 8 genes tested in 20 cases and 20 controls (FKBP8, ITPK1, MARCH2, PNPLA2, TUBA3, UBXD1, FTL); the remaining gene (PINK1) did not show a significantly different expression on rt-PCR.
Correlation of Gene Expression Profile with Coronary Disease
All 160 genes with rho>0.2 were included in the PLS analysis, with CAD-Index as the only response variable. Polynomial regression analysis of the resulting t1-scores versus CAD-Index resulted in the prediction model including 95% confidence range of the regression and the 95% prediction interval with r2 = 0.764 (p<0.001) (figure 1). Predictive accuracy was found to be excellent in the overall population (RMSEE (root mean square error of estimation) = 0.323), but improved with increasing threshold of CAD (RMSEE = 0.249 for controls vs cases with CAD>40; RMSEE = 0.204 for controls vs cases with CAD>60 and RMSEE = 0.172 for controls vs cases with CAD>70).
Figure 1. Partial least squares plot of nominal CAD index versus predicted CAD index.
Result of the partial least squares analysis including all controls and all cases; n = 222 and 160 genes. Cases are represented as triangles and controls as circles. The CAD-index as predicted by the gene expression is plotted versus the nominal CAD-index as obtained from coronary angiography. Regression line of the predicted CAD index versus nominal CAD-Index is displayed by the full line including 95% confidence interval of the regression (dotted lines) and the 95% prediction interval (striped lines). Goodness of fit is indicated by r2 = 0.776 (p<0.001).
In order to test for robustness of the model, the PLS analysis was performed separately for each of the three cohorts, with the model repeatedly constructed using two cohorts (training sample) and tested in the third cohort (test sample). While the controls remain quite stable in the range of -2 standard deviations, the t1-scores of the cases were located mainly in the +2 standard deviation range and increase with increasing CAD-Index (figure 2). This relationship is clearly present in each cohort. Cross-validation of the model was also performed by dividing the data into 7 groups of on average 32 subjects and then developing a number of parallel models from reduced data with one of the groups deleted. The omitted group was then used as a test data set, and the differences between actual and predicted CAD-Indices were subsequently calculated for these data points. The reduced models validation demonstrated a Q2cum of 0.776, indicating an excellent predictive ability.
Figure 2. Partial least squares plot per cohort.
Results of the partial least squares regression analysis with 160 genes applied separately to each of the three cohorts (A) “Matched Men” (B) “Unmatched Men” and (C) ‘“Unmatched Women”. Models were each time constructed in two cohorts and then tested in the third cohort. Individual patients are ordered by their CAD-Index. Labels represent the individual CAD-Index. Controls (full line) have all CAD-Index 0, and the CAD-Index of cases (dotted line) increases from 23 up to 100. While the controls remain quite stable in the range of -2 standard deviations, the t1-scores increase with increasing CAD-Index (t1 indicates the t1 score vector result from the PLS analysis).
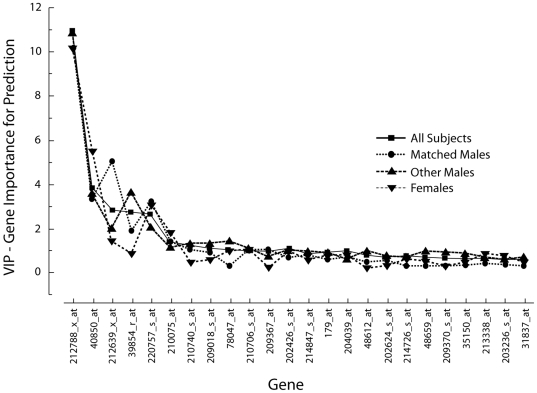
A Variable Importance in the Projection (VIP) of each gene for the separate PLS analyses of the three cohorts compared to the PLS analysis including all subjects was calculated. The VIP of the first 24 genes shows only little variation between the three cohorts suggesting a rather high stability of the prediction model (figure 3). A set of eight genes appears to have the highest impact on the model (FTL, FKBP8, TUBA3, PNPLA2, UBXD1, MARCH2, ITPK1, PINK1, in order of contribution; listed in bold in table 2). A PLS analysis only involving these eight highest ranking genes in the VIP analysis showed that the expressions profiles of these eight genes are also able to predict the CAD-Index (r2 = 0.752). Adding traditional risk factors and biochemical markers do not significantly improve this model (r2 = 0.782).
Figure 3. VIP.
Variable Importance in the Projection (VIP) for the separate PLS analyses of the three different cohorts compared to the PLS analysis including all subjects. Displayed are the 24 probesets with the highest VIP. The curve shows a steep decrease for the first 8 genes (listed in table 2); the contribution of further genes is comparable as suggested by almost linear curves.
Test of Predictability in Human Aorta Tissue Samples
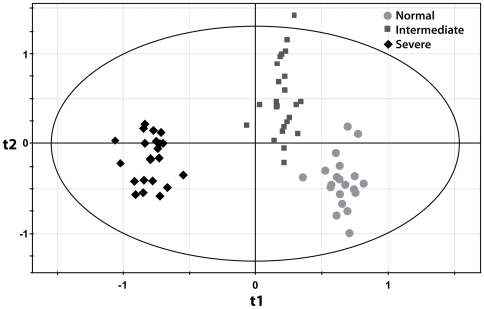
Since the genes whose expression contributes to prediction of CAD were studied within circulating leukocytes, we sought to define whether they actually reflect a molecular process that is ongoing within atherosclerotic arteries or not. Furthermore, as a test of reproducibility of the contribution of these 160 genes to predicting atherosclerotic disorders, we have investigated whether the in situ expression pattern of our 160 genes derived from peripheral blood could also adequately predict the severity of aorta atherosclerotic lesions. To achieve this goal, we have used gene expression data extracted from a large set of human aortas obtained from heart donors (n = 67), an independent human model of atherosclerosis. Excluding genes that are not present on the microarray used in the aorta expression study, the expression pattern of the remaining genes accurately separated the aorta samples according to the severity of atherosclerosis (figure 4). These results indicate that gene expression changes in peripheral blood are correlated with the extent of coronary atherosclerosis reflect similar pathophysiological changes in atherosclerotic arteries.
Figure 4. Partial least squares discriminant analysis in atherosclerotic aortas.
Result of the partial least squares discriminant analysis (t1/t2 score plot) including all aorta samples; n = 67. Dots represent normal aortas, squares represent intermediate atherosclerosis and diamonds indicate severe aorta atherosclerosis. Using expression data in aorta samples, the PLS analysis using the 160 peripheral blood genes adequately separates normal aortas from intermediate and severe atherosclerotic aortas (the ellipse indicates Hotelling's T2 95% confidence region; t1 and t2 indicate the t1 and t2 score vector results from the PLS-DA analysis).
Discussion
In this large-scale expression analysis of peripheral whole blood cells, we have found 160 genes whose expression correlates with the severity of angiographically documented coronary artery atherosclerosis. Taking into account that the CAD-Index is a semi-quantitative estimate of the extent of coronary atherosclerotic disease, which implies variation across subjects even with the same degree of disease, the prediction based on expression pattern of these genes is robust. Our findings are also robust as assessed by internal validation and consistency across three distinct subgroups. Importantly, the in situ expression pattern of the 160 genes derived from the peripheral blood analysis was also predictive of the severity of atherosclerosis in human aorta tissue. This provides validation of the association of this set of genes with atherosclerosis and support for the concept that peripheral blood gene expression reflects pathophysiology in the vascular wall. Taken together, the molecular signature in peripheral blood for varying degrees of coronary artery disease is remarkably consistent with that seen in the atherosclerotic arterial wall, providing valuable new information of the pathways and their genes that are involved in the atherosclerotic process.
Peripheral blood is easily accessible and routinely used for diagnostic laboratory analysis and thus is a good resource for additional tests that might define extent of coronary artery disease. Several inflammatory markers, including high sensitivity C-reactive protein (CRP) are associated with cardiovascular risk, independently from traditional risk factors [21]. Nevertheless, there is debate as to the additional prognostic value of these tests beyond traditional risk factors [22]. Other non-invasive analyses, such as coronary multislice CT can identify the extent of coronary artery disease, but such tests require specialized equipment and involve use of intravenous contrast and radiation. A simple blood test that predicts the extent of coronary artery disease could provide an additional useful tool for screening for coronary artery disease in at-risk populations. A similar approach has been successfully used for detection of cardiac allograft rejection and the response to immunosuppressive therapy [23].
Most of the differentially expressed genes in the present study are involved in bone marrow cell differentiation, cell growth or growth arrest, apoptosis, cell adhesion and matrix modulation, and inflammatory and immune response, processes known to modulate atherosclerosis. Since blood samples were taken in stable patients, our finding that circulating blood cells differentially express many pro-inflammatory genes supports the paradigm that inflammation is an important process in patients with coronary artery disease. Expression patterns of the same genes were found to correlate with the extent of atherosclerosis in human aortas as well, indicating that gene expression patterns in peripheral blood cells associated with coronary artery disease to some extent mirror gene expression changes in the atherosclerotic vessel wall. Indeed, many of the genes shared by our predictive models modulate monocyte or macrophage function, including MAN2A [24], RXRA [25], LGALS9 [26], PSG3 [27], CEPBA [28], ARGHAP4 [29], MADH5 [30], AIF1 [31], ELAVL2 [32], STXBP2 [33], KCNMB1 [34], PDE4D [35], EPHB2 [36], GGA3 [37], PLAUR [38], NPR3 [39] and TNFRSF5 (CD40) [40]. Interestingly, four of these genes (KCNMB1, NEDD4L, ADD1 and NPR3) have been implied in genetic susceptibility for hypertension [41]–[44], while two genes have been associated with genetic susceptibility for stroke (PDE4D) [45] or myocardial infarction (ADD1) [46]. The present results also appear to support a role for ferritin light chain (FTL) in atherosclerosis [47]. Ferritin is the major intracellular iron storage protein that plays a major role in the reaction to oxidative stress. Using a proteomic approach, You et al. found that the levels of ferritin light chain protein were significantly increased in atherosclerotic coronary arteries [48]. Ferritin light chain is also upregulated in circulating leukocytes of patients with juvenile rheumatoid arthritis, sickle cell disease, autoimmune renal disease or multiple sclerosis, indicating that altered FTL gene expression in peripheral cells of CAD patients might in at least in part reflect a general pro-inflammatory state that leads to degenerative changes [49]–[52].
We intentionally did not separate peripheral blood cells or leukocyte subtypes. There is currently little pathophysiological evidence that the study of leukocyte subgroups would add to our predictive model and the isolation process could, in itself, affect the gene expression pattern. Using whole blood cells not only allows aggregate RNA expression analysis per patient without the need to pool rare subtypes, but is also more practical from a clinical perspective. Leukocyte levels in all groups were very similar, although it cannot be excluded that the percentage of specific subtypes differ between groups, and hence that different numbers of subtypes are responsible for the observed effect. Peripheral whole blood might also include differential expression signatures from reticulocytes, platelets or rare hematopoietic progenitors.
In a recent paper, Wingrove et al reported 526 differentially expressed genes (>1.3-fold expression) from a genome-wide microarray analysis of peripheral blood mononuclear cells of 27 cases with angiographically documented CAD and 14 controls [15]. The authors found that 14 genes, out of a a set of 106 genes including the 50 most significant genes from the microarray analysis and 56 genes selected from the literature, were associated with the presence of CAD and the severity of CAD in two independent cohorts. The overlap between our study and the Wingrove study at the individual gene level appears to be very limited. This might be in part due to the considerably different design of our study. Not only did we prefer a correlation-based approach, the Wingrove study also used a much smaller subset of patients for unbiased microarray-based gene discovery, and added 56 literature-based genes for the subsequent analysis in their two cohorts. As a result of our correlation analysis, we also did not exclude genes with differential expression below 1.3-fold; since atherosclerosis is a chronic disease, small changes in gene expression might accrue over time and result in a clinically relevant phenotype. Moreover, in contrast with our study, a substantial proportion of microarray samples in the Wingrove analysis were taken from patients presenting with an acute coronary syndrome, which might have significantly influenced expression levels. Another reason for the discrepancies between the two studies might be the different types of microarray used and different types of cells studied. In our study, we analyzed RNA from whole blood in all patients, in contrast with isolated mononuclear cells used in the discovery phase of the Wingrove study. An Ingenuity Pathway Analysis (IPA, Ingenuity Systems, Redwood City, Ca; USA) comparing the 366 genes with p<0.05 (from the 526 probesets) and our 160 genes with rho>0.2 shows that similar biological functions were hit, despite the different microarrays and different matrices used (data not shown). In any case, the discrepancies between both studies suggest that these results need to be validated in larger and more diverse populations.
Of the 160 genes we found to be correlated with the extent of CAD, only 19 were significantly differentially expressed between all cases and controls, while gene expression was significantly different for 90 genes when comparing 20 patients with the least predicted CAD-index to 20 patients with the highest predicted CAD-index. Most of our cases only have mild to moderate disease, with only a minority having extensive disease. Thus, in part as a result of our proteomics-driven patient selection, there is likely to be a very gradual transition from controls to cases, with the distrubution of cases being skewed towards the lower end of CAD-index. We therefore assumed that the difference between controls and cases was not likely to be very large, hence our preference for a correlation-based analysis. Furthermore, since the average age of the controls was 52 years, it is highly likely that some degree of coronary atherosclerosis is present in these subjects. Interestingly, patients with normal angiograms but with microvascular dysfunction may also demonstrate peripheral monocyte activation, although not to the extent seen in patients with angiographically documented coronary artery disease [53]. Our findings that the present model also accurately predicts the severity of coronary artery disease in female patients, in whom advanced coronary artery disease is less likely at the age of 50, is reassuring. It is notable that CRP and LDL did not predict disease in our population. However, while these are excellent markers for future cardiovascular events [54], their ability to predict the severity of angiographically documented CAD is known to be low [55]–[58]. We even observed an inverse correlation between LDL-cholesterol levels and CAD-index. This might be at least in part due to differences in treatments, especially in statin use. Statins might indeed blunt gene expression differences in vascular cells and circulating monocytes to certain extent, which might have influenced our findings [59], [60].
In conclusion, the combined predictive value of differentially expressed genes in peripheral blood correlates with the extent of coronary atherosclerosis. Importantly, the expression pattern of the same genes is also correlated with the extent of disease in atherosclerotic aortas. While these findings need prospective validation in further populations, our findings also suggest that gene expression profiles might represent a novel and promising non-invasive test to assess the presence and extent of coronary artery disease. Although the extent of angiographic disease is a strong predictor of clinical outcome, further studies in larger and unselected populations will also be needed to examine the potential role of gene expression patterns in predicting outcome and to address potential confounding factors.
Acknowledgments
Dr Sinnaeve is a Clinical Investigator of the Fund for Scientific Research – Flanders.
Footnotes
Competing Interests: Drs Grass, Chibout and Vonderscher are full-time employees of Novartis.
Funding: The study was funded by Novartis. Drs Grass, Chibout and Vonderscher are full-time employees of Novartis, and participated in the study design and data collection and analysis, but had no influence in the decision to publish.
References
- 1.Rosamond WD, Folsom AR, Chambless LE, Wang CH. Coronary heart disease trends in four United States communities. The Atherosclerosis Risk in Communities (ARIC) study 1987–1996. IntJ Epidemiol. 2001;30(Suppl 1):S17. doi: 10.1093/ije/30.suppl_1.s17. [DOI] [PubMed] [Google Scholar]
- 2.Gibbons GH, Liew CC, Goodarzi MO, Rotter JI, Hsueh WA, et al. Genetic markers: progress and potential for cardiovascular disease. Circulation. 2004;109:IV47. doi: 10.1161/01.CIR.0000133440.86427.26. [DOI] [PubMed] [Google Scholar]
- 3.Libby P. Inflammation in atherosclerosis. Nature. 2002;420:868. doi: 10.1038/nature01323. [DOI] [PubMed] [Google Scholar]
- 4.Eriksson EE, Xie X, Werr J, Thoren P, Lindbom L. Direct viewing of atherosclerosis in vivo: plaque invasion by leukocytes is initiated by the endothelial selectins. FASEB J. 2001;15:1149. doi: 10.1096/fj.00-0537com. [DOI] [PubMed] [Google Scholar]
- 5.Hiltunen MO, Tuomisto TT, Niemi M, Brasen JH, Rissanen TT, et al. Changes in gene expression in atherosclerotic plaques analyzed using DNA array. Atherosclerosis. 2002;165:23. doi: 10.1016/s0021-9150(02)00187-9. [DOI] [PubMed] [Google Scholar]
- 6.Nanni L, Romualdi C, Maseri A, Lanfranchi G. Differential gene expression profiling in genetic and multifactorial cardiovascular diseases. J Mol Cell Cardiol. 2006;41:934–948. doi: 10.1016/j.yjmcc.2006.08.009. [DOI] [PubMed] [Google Scholar]
- 7.Randi AM, Biguzzi E, Falciani F, Merlini P, Blakemore S, et al. Identification of differentially expressed genes in coronary atherosclerotic plaques from patients with stable or unstable angina by cDNA array analysis. J ThrombHaemost. 2003;1:829. doi: 10.1046/j.1538-7836.2003.00113.x. [DOI] [PubMed] [Google Scholar]
- 8.Seo D, Wang T, Dressman H, Herderick EE, Iversen ES, et al. Gene Expression Phenotypes of Atherosclerosis. Arterioscler Thromb Vasc Biol. 2004;24:1922–1927. doi: 10.1161/01.ATV.0000141358.65242.1f. [DOI] [PubMed] [Google Scholar]
- 9.Cagnin S, Biscuola M, Patuzzo C, Trabetti E, Pasquali A, et al. Reconstruction and functional analysis of altered molecular pathways in human atherosclerotic arteries. BMC Genomics. 2009;10:13. doi: 10.1186/1471-2164-10-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ijas P, Nuotio K, Saksi J, Soinne L, Saimanen E, et al. Microarray analysis reveals overexpression of CD163 and HO-1 in symptomatic carotid plaques. Arterioscler Thromb Vasc Biol. 2007;27:154–160. doi: 10.1161/01.ATV.0000251991.64617.e7. [DOI] [PubMed] [Google Scholar]
- 11.Mohler ER, 3rd, Sarov-Blat L, Shi Y, Hamamdzic D, Zalewski A, et al. Site-specific atherogenic gene expression correlates with subsequent variable lesion development in coronary and peripheral vasculature. Arterioscler Thromb Vasc Biol. 2008;28:850–855. doi: 10.1161/ATVBAHA.107.154534. [DOI] [PubMed] [Google Scholar]
- 12.Sluimer JC, Kisters N, Cleutjens KB, Volger OL, Horrevoets AJ, et al. Dead or alive: gene expression profiles of advanced atherosclerotic plaques from autopsy and surgery. Physiol Genomics. 2007;30:335–341. doi: 10.1152/physiolgenomics.00076.2007. [DOI] [PubMed] [Google Scholar]
- 13.Aziz H, Zaas A, Ginsburg GS. Peripheral blood gene expression profiling for cardiovascular disease assessment. Genomic Med. 2007;1:105–112. doi: 10.1007/s11568-008-9017-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Grimm RH, Jr, Neaton JD, Ludwig W. Prognostic importance of the white blood cell count for coronary, cancer, and all-cause mortality. JAMA. 1985;254:1932. [PubMed] [Google Scholar]
- 15.Wingrove JA, Daniels SE, Sehnert AJ, Tingley W, Elashoff MR, et al. Correlation of Peripheral-Blood Gene Expression With the Extent of Coronary Artery Stenosis. Circ Cardiovasc Genet. 2008;1:31–38. doi: 10.1161/CIRCGENETICS.108.782730. [DOI] [PubMed] [Google Scholar]
- 16.Donahue MP, Rose K, Hochstrasser D, Vonderscher J, Grass P, et al. Discovery of proteins related to coronary artery disease using industrial-scale proteomics analysis of pooled plasma. Am Heart J. 2006;152:478–485. doi: 10.1016/j.ahj.2006.03.007. [DOI] [PubMed] [Google Scholar]
- 17.Felker GM, Shaw LK, O'Connor CM. A standardized definition of ischemic cardiomyopathy for use in clinical research. J Am CollCardiol. 2002;39:210. doi: 10.1016/s0735-1097(01)01738-7. [DOI] [PubMed] [Google Scholar]
- 18.Mark DB, Nelson CL, Califf RM, Harrell FE, Jr, Lee KL, et al. Continuing evolution of therapy for coronary artery disease. Initial results from the era of coronary angioplasty. Circulation. 1994;89:2015. doi: 10.1161/01.cir.89.5.2015. [DOI] [PubMed] [Google Scholar]
- 19.Wang J, Robinson JF, Khan HM, Carter DE, McKinney J, et al. Optimizing RNA extraction yield from whole blood for microarray gene expression analysis. ClinBiochem. 2004;37:741. doi: 10.1016/j.clinbiochem.2004.03.013. [DOI] [PubMed] [Google Scholar]
- 20.Lockhart DJ, Dong H, Byrne MC, Follettie MT, Gallo MV, et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. NatBiotechnol. 1996;14:1675. doi: 10.1038/nbt1296-1675. [DOI] [PubMed] [Google Scholar]
- 21.Rosenson RS, Koenig W. Utility of inflammatory markers in the management of coronary artery disease. Am J Cardiol. 2003;92:10i. doi: 10.1016/s0002-9149(03)00504-6. [DOI] [PubMed] [Google Scholar]
- 22.Danesh J, Wheeler JG, Hirschfield GM, Eda S, Eiriksdottir G, et al. C-reactive protein and other circulating markers of inflammation in the prediction of coronary heart disease. NEnglJ Med. 2004;350:1387. doi: 10.1056/NEJMoa032804. [DOI] [PubMed] [Google Scholar]
- 23.Horwitz PA, Tsai EJ, Putt ME, Gilmore JM, Lepore JJ, et al. Detection of cardiac allograft rejection and response to immunosuppressive therapy with peripheral blood gene expression. Circulation. 2004;110:3815. doi: 10.1161/01.CIR.0000150539.72783.BF. [DOI] [PubMed] [Google Scholar]
- 24.Misago M, Liao YF, Kudo S, Eto S, Mattei MG, et al. Molecular cloning and expression of cDNAs encoding human alpha-mannosidase II and a previously unrecognized alpha-mannosidase IIx isozyme. ProcNatlAcadSciUSA. 1995;92:11766. doi: 10.1073/pnas.92.25.11766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Fritsche J, Stonehouse TJ, Katz DR, Andreesen R, Kreutz M. Expression of retinoid receptors during human monocyte differentiation in vitro. BiochemBiophysResCommun. 2000;270:17. doi: 10.1006/bbrc.2000.2371. [DOI] [PubMed] [Google Scholar]
- 26.Abedin MJ, Kashio Y, Seki M, Nakamura K, Hirashima M. Potential roles of galectins in myeloid differentiation into three different lineages. J LeukocBiol. 2003;73:650. doi: 10.1189/jlb.0402163. [DOI] [PubMed] [Google Scholar]
- 27.Snyder SK, Wessner DH, Wessells JL, Waterhouse RM, Wahl LM, et al. Pregnancy-specific glycoproteins function as immunomodulators by inducing secretion of IL-10, IL-6 and TGF-beta1 by human monocytes. Am J ReprodImmunol. 2001;45:205. doi: 10.1111/j.8755-8920.2001.450403.x. [DOI] [PubMed] [Google Scholar]
- 28.Heath V, Suh HC, Holman M, Renn K, Gooya JM, et al. C/EBPalpha deficiency results in hyperproliferation of hematopoietic progenitor cells and disrupts macrophage development in vitro and in vivo. Blood. 2004;104:1639. doi: 10.1182/blood-2003-11-3963. [DOI] [PubMed] [Google Scholar]
- 29.Tribioli C, Droetto S, Bione S, Cesareni G, Torrisi MR, et al. An X chromosome-linked gene encoding a protein with characteristics of a rhoGAP predominantly expressed in hematopoietic cells. ProcNatlAcadSciUSA. 1996;93:695. doi: 10.1073/pnas.93.2.695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Liu B, Mao N. Smad5: signaling roles in hematopoiesis and osteogenesis. IntJ BiochemCell Biol. 2004;36:766. doi: 10.1016/s1357-2725(03)00250-4. [DOI] [PubMed] [Google Scholar]
- 31.Arvanitis DA, Flouris GA, Spandidos DA. Genomic rearrangements on VCAM1, SELE, APEG1and AIF1 loci in atherosclerosis. J Cell MolMed. 2005;9:153. doi: 10.1111/j.1582-4934.2005.tb00345.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.King PH, Fuller JJ, Nabors LB, Detloff PJ. Analysis of the 5′ end of the mouse Elavl1 (mHuA) gene reveals a transcriptional regulatory element and evidence for conserved genomic organization. Gene. 2000;242:125. doi: 10.1016/s0378-1119(99)00537-5. [DOI] [PubMed] [Google Scholar]
- 33.Logan MR, Odemuyiwa SO, Moqbel R. Understanding exocytosis in immune and inflammatory cells: the molecular basis of mediator secretion. J Allergy ClinImmunol. 2003;111:923. [PubMed] [Google Scholar]
- 34.Hanley PJ, Musset B, Renigunta V, Limberg SH, Dalpke AH, et al. Extracellular ATP induces oscillations of intracellular Ca2+ and membrane potential and promotes transcription of IL-6 in macrophages. ProcNatlAcadSciUSA. 2004;101:9479. doi: 10.1073/pnas.0400733101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Shepherd MC, Baillie GS, Stirling DI, Houslay MD. Remodelling of the PDE4 cAMP phosphodiesterase isoform profile upon monocyte-macrophage differentiation of human U937 cells. BrJ Pharmacol. 2004;142:339. doi: 10.1038/sj.bjp.0705770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yu G, Luo H, Wu Y, Wu J. Ephrin B2 induces T cell costimulation. J Immunol. 2003;171:106. doi: 10.4049/jimmunol.171.1.106. [DOI] [PubMed] [Google Scholar]
- 37.Kzhyshkowska J, Gratchev A, Martens JH, Pervushina O, Mamidi S, et al. Stabilin-1 localizes to endosomes and the trans-Golgi network in human macrophages and interacts with GGA adaptors. J LeukocBiol. 2004;76:1151. doi: 10.1189/jlb.0504300. [DOI] [PubMed] [Google Scholar]
- 38.Paysant J, Vasse M, Soria J, Lenormand B, Pourtau J, et al. Regulation of the uPAR/uPA system expressed on monocytes by the deactivating cytokines, IL-4, IL-10 and IL-13: consequences on cell adhesion to vitronectin and fibrinogen. BrJ Haematol. 1998;100:45. doi: 10.1046/j.1365-2141.1998.00528.x. [DOI] [PubMed] [Google Scholar]
- 39.Baldini PM, De Vito P, Martino A, Fraziano M, Grimaldi C, et al. Differential sensitivity of human monocytes and macrophages to ANP: a role of intracellular pH on reactive oxygen species production through the phospholipase involvement. J LeukocBiol. 2003;73:502. doi: 10.1189/jlb.0702377. [DOI] [PubMed] [Google Scholar]
- 40.Schonbeck U, Libby P. CD40 signaling and plaque instability. CircRes. 2001;89:1092. doi: 10.1161/hh2401.101272. [DOI] [PubMed] [Google Scholar]
- 41.Fernandez-Fernandez JM, Tomas M, Vazquez E, Orio P, Latorre R, et al. Gain-of-function mutation in the KCNMB1 potassium channel subunit is associated with low prevalence of diastolic hypertension. J ClinInvest. 2004;113:1032. doi: 10.1172/JCI20347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lanzani C, Citterio L, Jankaricova M, Sciarrone MT, Barlassina C, et al. Role of the adducin family genes in human essential hypertension. J Hypertens. 2005;23:543. doi: 10.1097/01.hjh.0000160210.48479.78. [DOI] [PubMed] [Google Scholar]
- 43.Pankow JS, Dunn DM, Hunt SC, Leppert MF, Miller MB, et al. Further evidence of a quantitative trait locus on chromosome 18 influencing postural change in systolic blood pressure: the Hypertension Genetic Epidemiology Network (HyperGEN) Study. Am J Hypertens. 2005;18:672. doi: 10.1016/j.amjhyper.2004.12.004. [DOI] [PubMed] [Google Scholar]
- 44.Pitzalis MV, Sarzani R, Dessi-Fulgheri P, Iacoviello M, Forleo C, et al. Allelic variants of natriuretic peptide receptor genes are associated with family history of hypertension and cardiovascular phenotype. J Hypertens. 2003;21:1491. doi: 10.1097/00004872-200308000-00012. [DOI] [PubMed] [Google Scholar]
- 45.Gretarsdottir S, Thorleifsson G, Reynisdottir ST, Manolescu A, Jonsdottir S, et al. The gene encoding phosphodiesterase 4D confers risk of ischemic stroke. NatGenet. 2003;35:131. doi: 10.1038/ng1245. [DOI] [PubMed] [Google Scholar]
- 46.Tobin MD, Braund PS, Burton PR, Thompson JR, Steeds R, et al. Genotypes and haplotypes predisposing to myocardial infarction: a multilocus case-control study. Eur Heart J. 2004;25:459. doi: 10.1016/j.ehj.2003.11.014. [DOI] [PubMed] [Google Scholar]
- 47.You SA, Wang Q. Ferritin in atherosclerosis. Clin Chim Acta. 2005;357:1–16. doi: 10.1016/j.cccn.2005.02.001. [DOI] [PubMed] [Google Scholar]
- 48.You SA, Archacki SR, Angheloiu G, Moravec CS, Rao S, et al. Proteomic approach to coronary atherosclerosis shows ferritin light chain as a significant marker: evidence consistent with iron hypothesis in atherosclerosis. Physiol Genomics. 2003;13:25. doi: 10.1152/physiolgenomics.00124.2002. [DOI] [PubMed] [Google Scholar]
- 49.Alcorta D, Preston G, Munger W, Sullivan P, Yang JJ, et al. Microarray studies of gene expression in circulating leukocytes in kidney diseases. ExpNephrol. 2002;10:139. doi: 10.1159/000049909. [DOI] [PubMed] [Google Scholar]
- 50.Bomprezzi R, Ringner M, Kim S, Bittner ML, Khan J, et al. Gene expression profile in multiple sclerosis patients and healthy controls: identifying pathways relevant to disease. HumMolGenet. 2003;12:2191. doi: 10.1093/hmg/ddg221. [DOI] [PubMed] [Google Scholar]
- 51.Jarvis JN, Dozmorov I, Jiang K, Frank MB, Szodoray P, et al. Novel approaches to gene expression analysis of active polyarticular juvenile rheumatoid arthritis. Arthritis ResTher. 2004;6:R15. doi: 10.1186/ar1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Jison ML, Munson PJ, Barb JJ, Suffredini AF, Talwar S, et al. Blood mononuclear cell gene expression profiles characterize the oxidant, hemolytic, and inflammatory stress of sickle cell disease. Blood. 2004;104:270. doi: 10.1182/blood-2003-08-2760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Lin CP, Lin WT, Leu HB, Wu TC, Chen JW. Differential mononuclear cell activity and endothelial inflammation in coronary artery disease and cardiac syndrome X. IntJ Cardiol. 2003;89:53. doi: 10.1016/s0167-5273(02)00428-x. [DOI] [PubMed] [Google Scholar]
- 54.Ridker PM, Rifai N, Rose L, Buring JE, Cook NR. Comparison of C-reactive protein and low-density lipoprotein cholesterol levels in the prediction of first cardiovascular events. N Engl J Med. 2002;347:1557–1565. doi: 10.1056/NEJMoa021993. [DOI] [PubMed] [Google Scholar]
- 55.Arroyo-Espliguero R, Avanzas P, Cosin-Sales J, Aldama G, Pizzi C, et al. C-reactive protein elevation and disease activity in patients with coronary artery disease. Eur Heart J. 2004;25:401–408. doi: 10.1016/j.ehj.2003.12.017. [DOI] [PubMed] [Google Scholar]
- 56.Hunt ME, O'Malley PG, Vernalis MN, Feuerstein IM, Taylor AJ. C-reactive protein is not associated with the presence or extent of calcified subclinical atherosclerosis. Am Heart J. 2001;141:206–210. doi: 10.1067/mhj.2001.112488. [DOI] [PubMed] [Google Scholar]
- 57.Schmidt SB, Wasserman AG, Muesing RA, Schlesselman SE, Larosa JC, et al. Lipoprotein and apolipoprotein levels in angiographically defined coronary atherosclerosis. Am J Cardiol. 1985;55:1459–1462. doi: 10.1016/0002-9149(85)90953-1. [DOI] [PubMed] [Google Scholar]
- 58.Zebrack JS, Muhlestein JB, Horne BD, Anderson JL. C-reactive protein and angiographic coronary artery disease: independent and additive predictors of risk in subjects with angina. J Am Coll Cardiol. 2002;39:632–637. doi: 10.1016/s0735-1097(01)01804-6. [DOI] [PubMed] [Google Scholar]
- 59.Morikawa S, Takabe W, Mataki C, Wada Y, Izumi A, et al. Global analysis of RNA expression profile in human vascular cells treated with statins. J Atheroscler Thromb. 2004;11:62–72. doi: 10.5551/jat.11.62. [DOI] [PubMed] [Google Scholar]
- 60.Schirmer SH, Fledderus JO, van der Laan AM, van der Pouw-Kraan TC, Moerland PD, et al. Suppression of inflammatory signaling in monocytes from patients with coronary artery disease. J Mol Cell Cardiol. 2009;46:177–185. doi: 10.1016/j.yjmcc.2008.10.029. [DOI] [PubMed] [Google Scholar]