Summary
Directed cellular movements are a universal feature of morphogenesis in multicellular organisms. Differential adhesion between the stationary and motile cells promotes these cellular movements to effect spatial patterning of cells. A prominent feature of Drosophila eye development is the 90° rotational movement of the multicellular ommatidial precursors within a matrix of stationary cells. We demonstrate that the cell adhesion molecules Echinoid (Ed) and Friend of Echinoid (Fred) act throughout ommatidial rotation to modulate the degree of ommatidial precursor movement. We propose that differential levels of Ed and Fred between stationary and rotating cells at the initiation of rotation create a permissive environment for cell movement, and that uniform levels in these two populations later contribute to stopping the movement. Based on genetic data, we propose that ed and fred impart a second, independent, `brake-like' contribution to this process via Egfr signaling. Ed and Fred are localized in largely distinct and dynamic patterns throughout rotation. However, ed and fred are required in only a subset of cells - photoreceptors R1, R7 and R6 - for normal rotation, cells that have only recently been linked to a role in planar cell polarity (PCP). This work also provides the first demonstration of a requirement for cone cells in the ommatidial rotation aspect of PCP. ed and fred also genetically interact with the PCP genes, but affect only the degree-of-rotation aspect of the PCP phenotype. Significantly, we demonstrate that at least one PCP protein, Stbm, is required in R7 to control the degree of ommatidial rotation.
Keywords: Drosophila eye, Egfr, Adhesion, Cell motility, Ommatidial rotation, Planar cell polarity, Tissue polarity
INTRODUCTION
Cell-cell adhesion is fundamental to metazoan development and to the growth and maintenance of adult tissues. In adult tissues, continuous regulation of cell adhesion underlies events such as spermatid development (Inagaki et al., 2006; Mueller et al., 2003; Ozaki-Kuroda et al., 2002), maintenance of apicobasal polarity (Nelson, 2003; Tsukita et al., 2001) and regeneration of tissues that require constant maintenance, such as the lining of the gut (Hermiston et al., 1996). Throughout metazoan development, cell adhesion plays key roles in events including maintenance of tissue integrity, boundary formation (Kim et al., 2000; Tepass et al., 2002), cell signaling (Jamora and Fuchs, 2002; Perez-Moreno et al., 2003; Sakisaka et al., 2007) and directed cellular movements (Hermiston et al., 1996; Pacquelet and Rorth, 2005). The precise and dynamic control of cell adhesion is also a critical regulator of tissue morphogenesis and patterning. For example, remodeling cell junctions within epithelia enables single cells and groups of cells to slide past their neighbors to reorganize tissues, such as during neural tube closure and convergent extension in vertebrates (Djiane et al., 2000; Formstone and Mason, 2005; Harrington et al., 2007) and ovary maturation and dorsal closure in Drosophila (Gorfinkiel and Arias, 2007; Lin et al., 2007; Niewiadomska et al., 1999). Loss-of-function mutations in cell adhesion molecules result in birth defects and disease states such as metastatic cancer (Matsushima et al., 2003; Naora and Montell, 2005; Pignatelli, 1998; Sozen et al., 2001; Suzuki et al., 1998; Suzuki et al., 2000). The coordinated regulation of cell adhesion also plays a key role in the rotational movement of subsets of cells that polarizes the Drosophila eye across its dorsal/ventral (D/V) midline, the equator, in an event known as ommatidial rotation. The mechanism by which changes in cell adhesion regulate this morphogenetic movement is poorly understood.
The 800 unit eyes, or ommatidia, of the Drosophila compound eye are polarized across the equator. This polarity is manifest as two chiral forms of `trapezoids' composed of the photosensitive membranes, or rhabdomeres, of seven of the eight photoreceptor cells. The apex of the trapezoid (R3) points north in the dorsal half and south in the ventral half of the eye (see Fig. 1) (reviewed by Wolff and Ready, 1993).
Fig. 1.
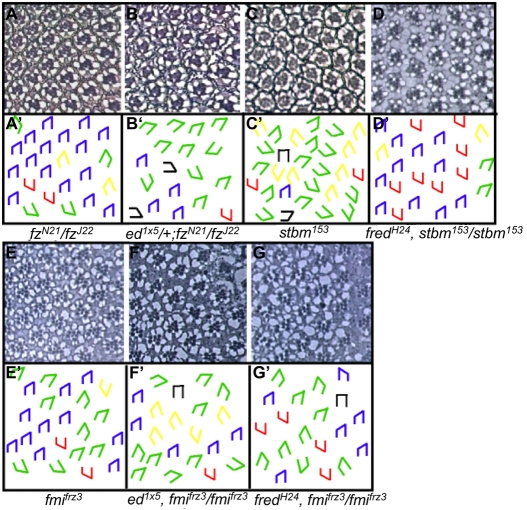
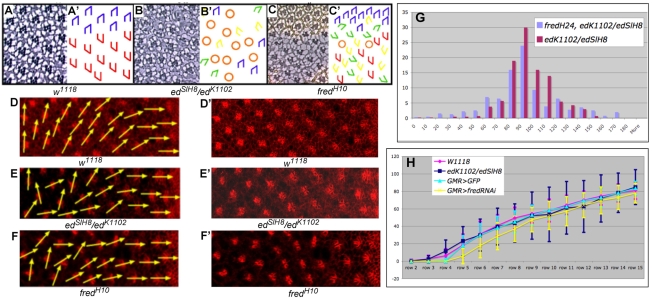
ed and fred mutant ommatidia misrotate. (A-C′) Tangential sections through adult Drosophila eyes (A-C) and corresponding schematics (A′-C′). (A) Wild type (w1118). Ommatidia come in two chiral forms, depicted in blue in the dorsal half and red in the ventral half of the eye. (B) Some edSlH8/edK1102 ommatidia under- or over-rotate (green and yellow trapezoids, respectively), and some contain an incorrect number of photoreceptors (orange circles). (C) Some ommatidia in fredH10 clones rotate correctly whereas others under- or over-rotate. (D-F′) The ed and fred orientation phenotypes result from aberrant ommatidial rotation. Anti-Arm staining (red) highlights the apical surface and outlines cell boundaries. Yellow vectors bisect R8 and run through the R3/R4 interface, highlighting the angle of orientation of each ommatidium. (D) Wild-type ommatidia follow a smooth progression of rotation. Ommatidial precursors in both edK1102/edSlH8 (E) and GMR>fredRNAi (F) knockdown eye discs misrotate. D′-F′ show D-F without arrows. (G) Reduction of fred activity enhances the ed mutant phenotype. Bar chart illustrating the percentage of ommatidia (y-axis) that are oriented at the angles indicated (x-axis) in edk1102/edSlH8 and edK1102, fredH24/edSlH8 eyes. (H) Graphical representation of data from D-F plotted as the mean angle of orientation (MAO) of ommatidia in each of four genotypes in rows 2-15. Error bars indicate the variance (s.d.). w1118 is the control for edK1102/edSlH8; GMR>GFP is the control for GMR>fredRNAi. The s.d. of ed and fred ommatidia is significantly different from that of the controls between rows 7-15. Trapezoid color for all schematics: blue and red, wild-type; green, under-rotated; yellow, over-rotated; black, fail to rotate; orange circles, incorrect number of photoreceptors.
Through a series of coordinated morphogenetic movements, the initially unpolarized retinal epithelium acquires polarity during third larval instar development. Groups of differentiating cells, the ommatidial precursors, rotate independently of their undifferentiated, stationary neighbors, the interommatidial cells (IOCs) (Fiehler and Wolff, 2007). These patterning events closely follow a moving front of differentiation, marked by the morphogenetic furrow (MF), which moves from posterior to anterior across the eye imaginal disc (Ready et al., 1976). Posterior to the furrow, the photoreceptors assemble into ommatidial units; ommatidial rotation begins coincident with assembly of the five-cell precluster, five rows posterior to the MF. Rotation continues as additional cells join the growing ommatidial unit. Ommatidia rotate 90° counterclockwise in the dorsal half of the eye and 90° clockwise in the ventral half. Rotation is complete by row 15 (Fiehler and Wolff, 2007).
Six core tissue polarity, or planar cell polarity (PCP), genes govern the establishment of this polarity: frizzled (fz), dishevelled (dsh), strabismus (stbm; Van Gogh - FlyBase), prickle (pk), diego (dgo) and flamingo (fmi; starry night - FlyBase) (Chae et al., 1999; Feiguin et al., 2001; Klein and Mlodzik, 2005; Klingensmith et al., 1994; Tree et al., 2002; Usui et al., 1999; Wolff and Rubin, 1998). Three phenotypes are evident when PCP signaling is disrupted, suggesting that three distinct events contribute to the establishment of polarity in the Drosophila eye: specification of the R3/R4 fates, direction of rotation (clockwise versus counterclockwise), and degree of rotation. Although mosaic analyses indicate a requirement for the PCP genes in specifying the R3 and R4 cell fates and additional work demonstrates a tight link between fate specification and the direction of rotation, the mechanisms that control the degree to which ommatidia rotate are poorly understood (Fanto and Mlodzik, 1999; Strutt et al., 2002; Wolff and Rubin, 1998). The identification of several proteins that affect only the degree of rotation, including the serine/threonine kinase Nemo (Nmo), E-cadherin (Shotgun - FlyBase), N-cadherin and members of the Egfr signaling pathway (Brown and Freeman, 2003; Choi and Benzer, 1994; Fiehler and Wolff, 2008; Gaengel and Mlodzik, 2003; Mirkovic and Mlodzik, 2006; Strutt and Strutt, 2003), suggests that a subset of genes might act to regulate this event.
Genetic evidence has revealed that the Egfr signaling pathway promotes rotation (Mirkovic and Mlodzik, 2006), even though ommatidia in Egfr pathway mutant eyes can over- or under-rotate, leading to a wide variance in the degree to which individual ommatidia rotate (Brown and Freeman, 2003; Gaengel and Mlodzik, 2003; Strutt and Strutt, 2003). Egfr pathway members signal through both the Mapk/Pnt transcriptional cascade and Canoe (Cno), the fly homolog of Afadin/AF-6 (Brown and Freeman, 2003; Gaengel and Mlodzik, 2003). Egfr pathway members also interact genetically with E-cadherin and N-cadherin during rotation (Mirkovic and Mlodzik, 2006). Whereas Egfr signaling and E-cadherin promote rotation, N-cadherin inhibits it (Mirkovic and Mlodzik, 2006). It remains unclear how rotating ommatidia coordinate changes in cell adhesion and cell signaling to initiate, advance and arrest rotation.
Here, we describe roles for two paralogous cell adhesion molecules (CAMs), Echinoid (Ed) and Friend of Echinoid (Fred), in controlling the degree of ommatidial rotation. Ed and Fred are large CAMs with extracellular immunoglobulin (Ig) C2 repeats and fibronectin type III domains (Bai et al., 2001; Chandra et al., 2003). Our work demonstrates that Ed and Fred are required at multiple steps during ommatidial rotation, using two functionally distinct mechanisms to either enable or slow rotation. We propose that Ed and Fred modulate adhesivity throughout ommatidial rotation, and, in addition, modify Egfr signaling during the second half of rotation. Initially, different levels of Ed and Fred in stationary and rotating cells create an environment permissive for rotation; later, equalizing levels between rotating and non-rotating populations of cells slows rotation. In addition, we demonstrate that ed and fred act in a subset of photoreceptor cells and in the cone cells, perhaps to regulate levels of Egfr signaling, ultimately inhibiting rotation. Notably, this requirement represents the first demonstration of a role for the cone cells in ommatidial rotation. We also demonstrate that ed and fred interact with the core PCP genes to control rotation. Finally, we identify a new and unexpected role for stbm in photoreceptor R7 in controlling the degree to which ommatidia rotate. This result raises the intriguing possibility that all the PCP genes function in distinct subsets of cells to control R3/R4 fate specification and the degree of rotation.
MATERIALS AND METHODS
Genetics
Fly lines: cno2/TM3 [U. Gaul (Boettner et al., 2003)], GMR-GAL4 (H. Chang, Perdue University), ro-GAL4 (J. Fischer, University of Texas), UAS-ed [J.-C. Hsu (Islam et al., 2003)], UAS-fredRNAi [H. Vaessin (Chandra et al., 2003)], y w eyFlp; edK1102FRT40A/BC, ed1x5FRT40A/CyO, edSlH8FRT40A/CyO, fredl(2)gH10FRT40A/CyO and fredl(2)gH24FRT40A/CyO (see de Belle et al., 1993), fredl(2)gH24, edK1102FRT40A/CyO, UAS-fred (S.A.S.). Remaining stocks were from the Bloomington Stock Center unless otherwise noted. Crosses were raised at 25°C.
Immunohistochemistry
Third instar eye imaginal discs were dissected, fixed and stained as described (Wolff, 2000). Tissue stained with anti-Cno antibody was fixed in PLP (Takahashi et al., 1998). Discs were incubated in primary antibody overnight at 4°C at the following concentrations: mouse anti-Armadillo, 1:10 (DSHB); rabbit anti-Ed, 1:1000 [A. Jarman (Rawlins et al., 2003)]; guinea pig anti-Fred, 1:1000; rabbit anti-Cno, 1:500 [D. Yamamoto (Takahashi et al., 1998)]. Alexa Fluor-conjugated secondary antibodies (Molecular Probes) were used at 1:300 and incubated in the dark for 2 hours at room temperature. Discs were mounted in 1:1 N-propylgallate:Vectashield and imaged on a Leica confocal microscope.
Phenotypic analyses
Adult eyes were fixed, embedded and sectioned as described (Wolff, 2000). Degree of rotation was determined using ImageJ software (NIH) to measure angles defined by vectors drawn (1) through the rhabdomeres of photoreceptors R1, R2 and R3, and (2) parallel to the equator. Only ommatidia with eight photoreceptor cells were scored. For all genotypes, 1000-1500 ommatidia (six to ten eyes) were scored. Statistical significance was determined using Student's t-test (mean angle of orientation) and F-test (variance).
Larval rotation phenotypes were scored in eye discs stained with anti-Armadillo to outline cells. ImageJ was used to measure the rotation angle between two vectors, one drawn between R3 and R4 and through R8, and the second drawn parallel to the equator. Angles of orientation were scored in rows 2 through 15 in 15 independent eye discs (i.e. one per larva) for each genotype.
Generation of mitotic clones
Mitotic clones were generated using the FLP/FRT technique (Xu and Rubin, 1993). For eyFLP clones, larvae were raised at 25°C. Mutant tissue is marked with w- in adult eyes and by the absence of GFP in larval discs.
Mosaic analysis
Mosaic analysis was performed in eyFLP and hsFLP clones. Phenotypes and photoreceptor genotypes were scored in adult eye sections and pupal eyes. Wild-type cells were marked with w+ in adult mosaic ommatidia and with GFP in pupal eyes. Only ommatidia with eight photoreceptor cells were scored.
RESULTS
ed and fred mutant ommatidia misrotate
ed was identified as a dominant suppressor of the ommatidial over-rotation phenotype caused by misexpression of nmo (Fiehler and Wolff, 2008). Although loss-of-function alleles of ed and nmo do not exhibit genetic interactions (data not shown), phenotypic analyses reveal key roles for ed and its paralog, fred, during ommatidial rotation. Wild-type ommatidia in adult eyes are oriented at almost precisely 90° (90.6°, s.d.=1.7). By contrast, many ed and fred mutant ommatidia were found to be oriented at either greater than, or less than, 90° (Fig. 1, Table 1). Whereas the mean angle of orientation (MAO) for both ed and fred loss-of-function alleles did not differ significantly from that of the wild type s.d., the variance, a quantifiable measurement of phenotype represented by the s.d., differed significantly from wild type (Table 1). Loss of fred function in clones of the hypomorphic allele fredH10 yielded a similar phenotype (s.d.=13.5, P=0, MAO=89.8°). ed and fred act cooperatively in R8 specification (Rawlins et al., 2003; Spencer and Cagan, 2003) and also cooperate to ensure that ommatidia orient at precisely 90°, as fredH24 dominantly enhanced the ommatidial orientation phenotype of edK1102/edSlH8 transheterozygotes (Fig. 1G, Table 1).
Table 1.
ed and fred genetically interact with the Egfr and PCP signaling pathways
| Genotype | MAO | s.d. | P | N | n |
|---|---|---|---|---|---|
| w1118 | 90.6 | 1.85 | 8 | 1006 | |
| ed1 | 87.16 | 10.1 | 0 | 9 | 1383 |
| edSIH8/edK1102 | 90.56 | 19.6 | 0 | 6 | 417 |
| fredH10 clones | 89.82 | 13.5 | 0 | 10 | 420 |
| GMR>fredRNAi | 87.65 | 13.14 | 0 | 10 | 1039 |
| fredH24, edK1102/edSIH8 | 87.1 | 29.71 | 4×10−12 | 10 | 384 |
| PCP gene | |||||
| fzN21/fzJ22 | 88.38 | 6.54 | 10 | 1126 | |
| ed1×5/+; fzN21/fzJ22 | 85.5 | 13.56 | 1.0×10−126 | 10 | 1275 |
| fredH24/+; fzN21/fzJ22 | 88.49 | 7.69 | 4×10−4 | 10 | 1355 |
| dsh1/Y | 86.16 | 10.7 | 7 | 899 | |
| dsh1/Y; ed1×5/+ | 82.3 | 16.87 | 9×10−41 | 6 | 752 |
| dsh1/Y; fredH24/+ | 86.22 | 12.19 | 4×10−5 | 9 | 1025 |
| stbm153 | 76.27 | 22.66 | 11 | 1638 | |
| ed1×5/+; stbm153 | 68.29 | 24.95 | 0.15 | 10 | 1199 |
| fredH24/+; stbm153 | 83.01 | 15.4 | 1.0×10−45 | 6 | 842 |
| pksple | 88.34 | 4.69 | 10 | 1431 | |
| edK1102/+; pksple | 88.99 | 4.52 | 0.49 | 10 | 1505 |
| fredH24/+; pksple | 88.02 | 5.03 | 0.29 | 10 | 1430 |
| dgo380 | 88.61 | 11.83 | 8 | 755 | |
| ed1×5/+; dgo380 | 83.9 | 16.37 | 3×10−25 | 10 | 1235 |
| fredH24/+; dgo380 | 87.89 | 13.08 | 7×10−4 | 10 | 1142 |
| fmifrz3 | 88 | 10.24 | 6 | 612 | |
| ed1×5/+; fmifrz3 | 83.83 | 24.52 | 1.6×10−13 | 6 | 627 |
| fredH24/+; fmifrz3 | 85.17 | 15.49 | 1.2×10−11 | 9 | 1010 |
| Egfr pathway | |||||
| aosrlt | 77.84 | 40.42 | 6 | 645 | |
| ed1×5/+; aosrlt | 80.61 | 38.39 | 0.02 | 10 | 1055 |
| fredH10/+; aosrlt | 78.00 | 39.26 | 0.11 | 10 | 1040 |
| ed1 | 87.17 | 10.13 | 0 | 10 | 1383 |
| ed1, spis3547/ed1 | 89.88 | 5.71 | 1.3×10−76 | 10 | 1295 |
| cnomis1/cno2 | 94.30 | 26.28 | 6 | 728 | |
| ed1×5/+; cnomis1/cno2 | 88.39 | 33.48 | 4.9×10−25 | 10 | 867 |
| fredH10/+; cnomis1/cno2 | 95.99 | 20.65 | 1.8×10−20 | 10 | 1382 |
| EgfrElp | 92.08 | 12.12 | 5 | 466 | |
| fredh24/EgfrElp | 88.54 | 8.34 | 1×10−192 | 5 | 464 |
| pntΔ88/pnt1277 | 85.5 | 16.64 | 9 | 630 | |
| ed1×5/+; pntΔ88/pnt1277 | 83.53 | 20.84 | 2×10−6 | 6 | 611 |
| fredH10/+; pntΔ88/pnt1277 | 91.2 | 5.1 | 4×10−297 | 8 | 987 |
P-values are derived from F-test. F-test P-values are for a comparison between the s.d. of the genotype indicated and its respective baseline (i.e. the homozygous phenotype is the baseline for modified genotypes).
N, number of eyes scored; n, number of ommatidia scored.
The ed and fred orientation phenotypes could originate from defective rotation or defects in morphogenetic patterning events that occur during pupal life. To establish whether ed and fred function during ommatidial rotation, a row-by-row analysis of the degree to which individual ommatidia rotate was conducted between rows 2 and 15 in third instar eye imaginal discs lacking ed or fred function, and compared with age-matched, wild-type counterparts (w1118 or GMR>GFP, respectively; see Materials and methods; Fig. 1D-F,D′-F′). Rotation was measured in edSlH8/edK1102 and GMR>fredRNAi eye imaginal discs (fred alleles are lethal, necessitating the use of fredRNAi; the GMR>fredRNAi phenotype is identical to the fredH10 phenotype; Table 1). In edSlH8/edK1102 and GMR>fredRNAi, the variance in the degree of rotation (s.d.) was greater than in wild type (Fig. 1H). Importantly, the mutant s.d. did not become statistically distinct from the wild-type s.d. until row 7, approximately 2-3 rows after the initiation of rotation, the time at which the anterior and posterior cone cells join the ommatidial precursor (Fig. 1H) (Fiehler and Wolff, 2007). Importantly, the s.d. was still significantly greater in the mutants than in wild type at row 15, when rotation is complete in the wild type (w1118 s.d.=9.8, ed s.d.=20, P<1×10-7; GMR>GFP s.d.=8, GMR>fredRNAi s.d.=12, P<1×10-4). These results demonstrate a role for ed and fred in the cellular movements that drive ommatidial rotation and further indicate they are required for the post-initiation stages of rotation, not for the initiation of rotation. Although ed and fred might also participate in later patterning events that align ommatidia, their contributions to such events would be likely to play only a minor role in ommatidial rotation, as the MAO and s.d. for ed and fred ommatidia when rotation is complete (row 15) are essentially the same as in their adult counterparts.
Ed and Fred localize in dynamic and partially overlapping patterns in the eye imaginal disc
Ed localizes throughout the eye imaginal disc (Bai et al., 2001; Rawlins et al., 2003), but insight into the mechanisms by which Ed might regulate ommatidial rotation necessitate a cell-by-cell and row-by-row analysis of Ed localization. Immunolocalization of the C-terminal anti-Ed antibody revealed that high levels of Ed protein localize at the apical surface of all cells in the MF and through row 1 (the arc stage; Fig. 2A). In rows 3 and 4, Ed becomes reduced specifically within the cells that will soon begin to rotate (i.e. photoreceptors; Fig. 2B,C), making the photoreceptor clusters appear as holes within the imaginal disc epithelium, a staining pattern that persists until row 7 (Fig. 2H). Notably, cells with higher levels of Ed adhere more strongly to each other than to cells with lower levels of Ed (Spencer and Cagan, 2003; Wei et al., 2005). Ed levels were found to be high in photoreceptors R1, R6 and R7 when they are recruited into the growing ommatidium in rows 5/6 (Fig. 2D,E). Shortly following the recruitment of these photoreceptors and the consequent increase in Ed levels at the rotation interface, rotation slows (row 7).
Fig. 2.
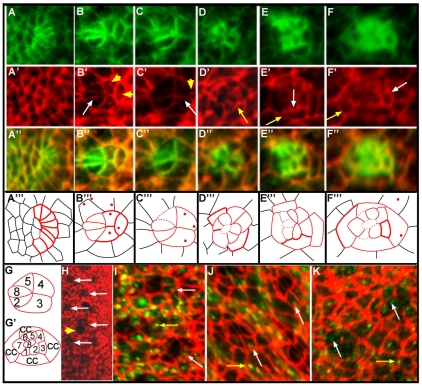
Ed localization is dynamic throughout rotation. (A-F‴) Anti-Arm (A-F, green) and anti-Ed (A′-F′, red) staining, and merge (A″-F″), in sequentially older ommatidial precursors in Drosophila third instar eye disc. Corresponding schematics (A‴-F‴) are shown with Ed localization in ommatidial precursors represented by solid red lines, Ed localization in cells outside the ommatidial precursors shown in black, and dashed red lines indicating cell boundaries where Ed is below detectable levels. The intensity of Ed staining correlates with the line weight. (A-A‴) In row 1, Ed is localized in all cells. (B-B‴) By row 3, Ed levels have diminished in R8, R2 and R5 (white arrow). Ed puncta are visible (yellow arrowheads). (C-C‴) Just prior to the start of rotation, Ed levels drop in the photoreceptor cells (see also H); Ed is visible at the R3/R4 (white arrow), R2/R3 and R4/R5 interfaces and in puncta (yellow arrowhead). (D-D‴) Ed levels increase in the photoreceptors as rotation progresses (yellow arrow). (E-E‴) In row 8, Ed remains high in the photoreceptor and cone cells (white arrow), and levels equalize between rotating and non-rotating cells (yellow arrow). (F-F‴) At the completion of rotation, Ed is enriched at the cone cell/interommatidial cell (IOC) (yellow arrow) and the cone cell/photoreceptor cell (white arrow) boundaries. (G,G′) Key to the schematics in A‴-F‴. cc, cone cell. (H) Low-magnification image of an eye disc stained with anti-Ed antibody. Just before rotation begins, ommatidia with low levels of Ed appear as `holes' in the staining pattern (white arrows). Mitotic cells, which also resemble `holes' (yellow arrowhead), are distinct. (I,J) Ed (red) vesicles colocalize with (I) Rab5-GFP (green) and (J) Rab7-GFP (green) positive puncta in both IOCs (puncta shown by yellow arrows) and photoreceptor cells (puncta shown by white arrows). (K) Vesicular Ed (red) does not colocalize with anti-Rab11 staining (green) in recycling endosomes in either IOCs (yellow arrows) or photoreceptors (white arrows).
When the cone cells are recruited, a dramatic shift was observed in the relative levels of Ed within the ommatidia and in the IOCs: Ed became prominent in two bands, one at the interface between the cone cells and the photoreceptors and a second at the interface between the cone cells and the IOCs (Fig. 2E,F). Notably, the recruitment of the cone cells and the resulting increase in Ed levels coincided with the second, slower 45° of rotation (Fiehler and Wolff, 2007). The distinct early and late patterns of Ed localization in rotating versus stationary cells suggest a model in which adhesion between rotating and non-rotating cells is reduced early to enable cells to slide past one another, and is subsequently increased during the slow phase of rotation to slow, and ultimately stop, rotation.
In addition to its membrane localization, Ed was also evident in intracellular vesicles throughout the eye disc (Fig. 2B,C) before, and at the initiation of, rotation. Photoreceptors R8, R2, R5, R3 and R4 contained large Ed puncta that frequently colocalized with either GFP-tagged Rab5 (early endosome marker, Fig. 2I) or Rab7 (late endosome/lysosome marker, Fig. 2J), but not with anti-Rab11 (which labels recycling endosomes, Fig. 2K), suggesting that Ed is endocytosed and degraded in these cells. By contrast, Ed was not vesicularized in photoreceptors R1, R6 and R7 (although Rab5 and Rab7 are prominent in these cells, Fig. 2I,J). The presence of Ed in endosomes prior to the onset of rotation suggests that some cells are actively reducing Ed levels and that rotation requires different levels of Ed in moving and stationary cells.
The localization pattern of Fred, as detected with an antibody raised against a peptide in the Fred intracellular domain (S.A.S., unpublished), differs from that of Ed. Like Ed, Fred protein was abundant in the MF. In contrast to Ed, however, early in rotation Fred was enriched in the photoreceptors relative to the surrounding IOCs (Fig. 3A,B). The localization of Fred in R3 and R4 was dynamic during the first half of rotation. At the initiation of rotation (rows 4-5), Fred was localized in a double-horseshoe pattern (UU), outlining photoreceptors R3 and R4, except where they abut R2 and R5 (Fig. 3B-B‴). One row, or 1.5 hours, later, Fred became restricted to the lateral edge of the R4 cell and the R3/R4 boundary (Fig. 3C-C‴,D-D‴). Fred levels remained high in R1, R6 and at the R7/R8 interface as they were recruited into the photoreceptor cluster in row 6 (Fig. 3C-C‴). By row 7, Fred was no longer detectable at the R3/R4 boundary but remained at the lateral edge of R4, the R7/R8 interface and in R1 and R6 (Fig. 3E-E‴). Following recruitment of the cone cells, the Fred pattern recapitulated the Ed pattern in that bands of Fred were evident at the interfaces between both the cone cells and the photoreceptors and the cone cells and IOCs (Fig. 3F-F‴). The localization pattern at the cone cell/photoreceptor and cone cell/IOC boundaries suggests that adhesion increases between these subsets of cells during the second, slower phase of rotation, perhaps serving as a brake for rotating cells.
Fig. 3.
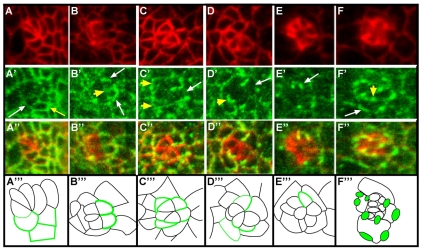
Fred localization is dynamic throughout rotation. (A-F‴) Anti-Arm (A-F, red) and anti-Fred (A′-F′, green) staining, and (A″-F″) merge, in increasingly older ommatidial precursors in Drosophila third instar eye disc. (A‴-F‴) Corresponding schematics, in which Fred localization is represented by green lines and line weight correlates with the intensity of Fred staining. (A-A‴) In row 3, Fred levels are enriched in R3 (white arrow), R4 (not evident in this image), and in the mystery cells (1-2 cells that lie between R3 and R4 in the developing ommatidial cluster but are ultimately expelled from the maturing ommatidial precursor; yellow arrow). (B-B‴) Just prior to the initiation of rotation, Fred localizes to the lateral edges of R3 and R4 (white arrows) and the R3/R4 boundary (yellow arrowhead). (C-C‴) In row 6, Fred begins to disappear from R3 (white arrow), but remains high in R4 and at the R3/R4 boundary. The newly added R1 and R6 contain high levels of Fred (yellow arrowheads). (D-D‴) In row 7, Fred disappears from R3 but is still high in R4 and at the R3/R4 boundary (white arrow). A bright band of Fred highlights the interface between R7 and R8 (yellow arrowhead), and Fred can still be seen faintly in R1 and R6. (E-E‴) By row 9, Fred is no longer present at the R3/R4 boundary, outlining only the periphery of R4 (white arrow). (F-F‴) At the end of rotation, Fred is enriched at the interfaces between the cone cells and the IOCs (white arrow) and also at the boundaries between the photoreceptors and the cone cells (yellow arrowhead).
Misexpression of ed and fred generates an under-rotation phenotype
The dynamic localization of Ed and Fred in rotating and stationary cells suggests that expression of ed and fred must be tightly regulated in time and space to achieve normal rotation. To test this hypothesis, cell-specific drivers were used to manipulate Ed and Fred levels in the photoreceptors and IOCs to either artificially equalize levels between rotating and non-rotating cells or to force high levels of expression in cells in which Ed and Fred are not normally elevated.
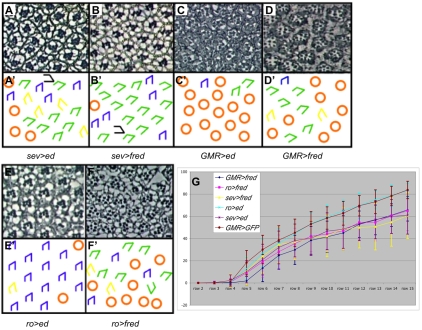
Ommatidial precursors rotated more slowly when driving UAS>ed or UAS>fred with the following drivers: sev>Gal4 (R3, R4, R1, R6, R7 and the cone cells); ro>Gal4 (R8, R2 and R5); and GMR>Gal4 (all cells posterior to the MF). Despite the distinct Ed and Fred localization patterns, the consequence of misexpression was similar for ed and fred. When either ed or fred was driven by the sev promoter, ommatidia under-rotated, on average, and exhibited a significant variance (Table 1, Fig. 4A,B). A similar phenotype resulted from misexpression of UAS>fred driven by ro>Gal4, although driving UAS>ed with ro>Gal4 did not cause a rotation phenotype (Fig. 4E,F), probably because the mechanism in place to clear Ed from the R8/R2/R5 membranes remains functional (see Fig. S1 in the supplementary material). Note that some ro>ed ommatidia had the expected missing photoreceptor phenotype owing to an effect on Egfr signaling (Rawlins et al., 2003; Spencer and Cagan, 2003). In the genotypes that under-rotate, aberrant rotation was evident from the start of rotation, or between rows 4 and 5 (Fig. 4G). By row 15, when rotation is complete in the wild type, ommatidia in these misexpression backgrounds had only rotated ∼60° (Fig. 4G). Overall, these data indicate that excess ed and fred activity early in rotation and forced equalization of levels in rotating versus non-rotating cells interfere with rotation, suggesting that dissimilar Ed and Fred levels in rotating and non-rotating cells are vital for the progression of rotation.
Fig. 4.
Misexpression of ed or fred results in under-rotation. (A-F′) Sections through adult Drosophila eyes (A-F) and corresponding schematics (A′-F′) of ed and fred misexpression lines. (A,B) sev>ed and sev>fred ommatidia frequently under-rotate; very few ommatidia are missing photoreceptors. (C) GMR>ed tissue is severely disrupted, precluding analysis of angles of orientation. (D) Most ommatidia in GMR>fred adult eyes under-rotate. Additional defects (spacing, morphology) might be due to disrupted Egfr signaling. (E) Most ommatidia rotate 90° in ro>ed eyes; some ommatidia are missing photoreceptors. (F) By contrast, many ommatidia under-rotate in ro>fred eyes. (G) Graph of larval rotation, or MAO, for ommatidia in rows 2-15. y-axis, degree of rotation. Ommatidia in misexpression lines are under-rotated in rows 4-15 compared with controls (GMR>GFP). Error bars indicate s.d.
ed and fred are required in a subset of cells for ommatidial rotation
Ed and Fred are dynamically localized in multiple cell types in the eye disc during rotation (Figs 2, 3). Since ed and fred have pleiotropic effects, the localization patterns of the proteins do not definitively identify those cells that require ed and fred for normal rotation, particularly as Ed and Fred regulate the reiterative Egfr signaling necessary for photoreceptor recruitment at this time (Freeman, 1997; Spencer and Cagan, 2003; Spencer et al., 1998). To identify the single photoreceptor or subsets of photoreceptor cells in which Ed and/or Fred function to regulate ommatidial rotation, we conducted a mosaic analysis. The FLP/FRT system (Xu and Rubin, 1993) was used to generate clones of either ed1x5 or fredH10 mutant tissue. The degree of rotation of mosaic ommatidia, i.e. those with a mixture of genetically mutant and genetically wild-type photoreceptors, was then assessed to evaluate the function of Ed or Fred in specific photoreceptors/groups of photoreceptors. Mosaic ommatidia mutant for ed (or fred) in a given photoreceptor were compared with mosaic ommatidia wild-type for ed in that photoreceptor; the genotypes of the remaining photoreceptors were not factored in. Parallel analyses were conducted for three groups of photoreceptors, R3/R4, R2/R5 and R1/R6/R7, and significant changes in the variance between each paired data set were identified. This analysis demonstrated that both ed and fred are required in R1, R6 and R7 for correct rotation: mosaic ommatidia genotypically wild-type for ed or fred in R1, R6 and R7 were more likely to have a smaller variance than mosaic ommatidia in which R1, R6 and R7 are genotypically mutant (Fig. 5A; see Table S1 in the supplementary material). Although a requirement for fred in R3 and R4 seemed likely given the prominent expression of Fred in R3 and R4 at a critical period of rotation, the mosaic analysis did not identify a requirement for either ed or fred in R3, R4, R8, R2 or R5 (Fig. 5B,C; see Table S1 in the supplementary material).
Fig. 5.
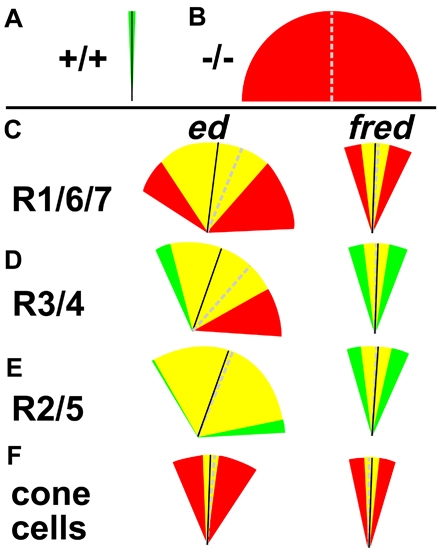
ed and fred are required in R1, R6, R7 and the cone cells for correct ommatidial rotation. (A) Schematic representation of wild-type MAO (black line; 90.6°) and wild-type variance (s.d., green wedge; 1.7). (B) Schematic representation of hypothetical mutant MAO (dashed gray line) and s.d. (red wedges). (C,F) ed and fred are required in photoreceptors R1, R6, R7 and the cone cells for rotation. (D,E) ed and fred are not required in R3/R4 or R2/R5 for rotation. The difference between the s.d. of the genetically wild-type and the genetically mutant populations is not significant. In C-F: black line, MAO when designated cells are genetically wild-type for ed or fred; dashed gray line, MAO when designated cells are genetically mutant for ed or fred; green wedges, genetically wild-type s.d; red wedges, genetically mutant s.d.; yellow wedges, overlap.
Wild-type ed or fred in R1, R6 and R7 does not completely rescue rotation. Furthermore, ommatidia with a full complement of genotypically wild-type photoreceptors can still misrotate. These observations suggest that ed and fred function in additional, non-photoreceptor cells to regulate ommatidial rotation. The most compelling candidates are the cone cells, as they express high levels of Ed and Fred until well after the completion of rotation. Mosaic ommatidia were therefore evaluated in mid-pupal (40 hours after puparium formation at 25°C) ed1x5 and fredH10 eyes, which revealed that mosaic ommatidia containing wild-type photoreceptors and between one and four mutant cone cells misrotate (for ed1x5 MAO=83°, s.d.=30, P<5×10-17; for fredH10 MAO=92°, s.d.=18, P<3×10-9; Fig. 5F; see Fig. S2 and Table S1 in the supplementary material), thus defining roles for Ed and Fred in the cone cells in ommatidial rotation. These results provide the first demonstration of a role for cone cells in ommatidial rotation.
ed and fred interact with Egfr signaling pathway members during rotation
The ed and fred ommatidial rotation phenotypes strongly resemble those observed in mutants of members of the Egfr signaling pathway. Furthermore, ed inhibits Egfr signaling (Bai et al., 2001; Rawlins et al., 2003; Spencer and Cagan, 2003). To determine whether Ed and/or Fred cooperate with Egfr signaling, we tested ed and fred for their ability to interact with Egfr pathway members and found that they dominantly modify the rotation phenotypes of pnt and cno. ed1x5 enhanced, whereas fredh24 strongly suppressed, the phenotype of both pntΔ88/pnt1277 and cnomis1/cno2 (Table 1, Fig. 6). Unlike in the wing, Cno localization was unchanged in clones of the null allele ed1x5 (data not shown). Although standard epistasis analysis cannot be employed to unambiguously order ed, fred and these Egfr signaling genes in a linear pathway (null alleles of these genes are lethal and the ommatidial rotation phenotypes are indistinguishable), our data indicate that in the simplest scenario, ed and fred function upstream of Ras85D, as ed and fred interact with both branches (cno and pnt) of the Egfr pathway. These results further suggest that ed and fred are likely to regulate rotation at least partly through control of Egfr signaling.
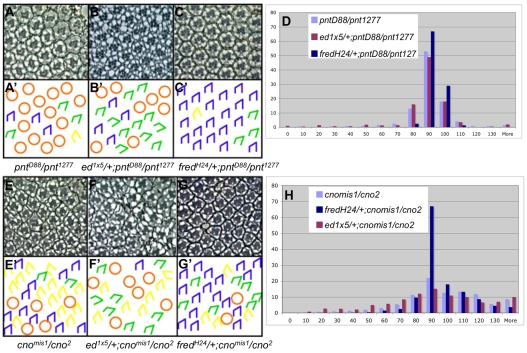
Fig. 6.
ed and fred interact genetically with pnt and cno. (A-C′) Sections through adult Drosophila eyes (A-C) and corresponding schematics (A′-C′). (A) pnt1277/pntΔ88 mutant eyes exhibit both over- and under-rotated ommatidia. (B) ed1x5/+, pntΔ88/pnt1277. Reducing ed activity enhances the pnt phenotype (i.e. the s.d. increases). (C) fredH24/+, pntΔ88/pnt1277. Reducing fred activity suppresses the pnt phenotype virtually to wild type. (D) Bar chart of angles of ommatidial orientation for pntΔ88/pnt1277, ed1x5/+, pntΔ88/pnt1277 and fredH24/+, pntΔ88/pnt1277. x-axis, MAO; y-axis, percentage. (E-G′) Sections (E-G) and corresponding schematics (E′-G′) for adult eyes of the indicated genotypes. (E) In cnomis1/cno2 mutant eyes, most ommatidia over-rotate. (F) ed1x5/+, cnomis1/cno2. Reducing ed activity enhances the cno phenotype. (G) fredH24/+; cnomis1/cno2. Reducing fred activity suppresses the cno phenotype. (H) Bar chart of angles of ommatidial orientation in cnomis1/cno2, ed1x5/+, cnomis1/cno2 and fredH24/+; cnomis1/cno2 adult eyes. x-axis, MAO; y-axis, percentage.
ed and fred interact with different subsets of planar cell polarity genes
The PCP genes control three events: specification of the R3 and R4 cell fates (Fanto et al., 1998; Fanto and Mlodzik, 1999; Wolff and Rubin, 1998), the direction of rotation with respect to the dorsal or ventral location of the ommatidium in the eye, and the degree of rotation. Of these three events, ed and fred regulate only the degree to which ommatidia rotate, suggesting that they might cooperate with the PCP genes in this event. Genetic assays designed to identify a possible link between ed and fred and PCP signaling revealed that ed and fred interact genetically with largely non-overlapping sets of the six core PCP genes as follows: ed dominantly enhances the mutant phenotypes of the genes that function in R3, i.e. fz, dsh, dgo and fmi; by contrast, fred dominantly interacts with genes that function in R4, as fred suppresses the hypomorphic stbm phenotype and enhances the fmi phenotype (Table 1; Fig. 7).
Fig. 7.
ed and fred interact genetically with different subsets of the PCP genes. (A-G′) Sections through adult Drosophila eyes (A-G) and corresponding schematics (A′-G′). Red trapezoids (this figure only) indicate dorsoventral inversions. (A) fzN21/fzJ22 mutant eyes exhibit both over- and under-rotated ommatidia. ed interacts specifically with the subset of PCP genes required in R3: fz, dgo and fmi. (B) ed1x5/+; fzN21/fzJ22. Reducing ed activity enhances the fz rotation phenotype without affecting the chirality phenotype. fred interacts with two PCP genes that are required in R4 for correct polarity: stbm and fmi. (C) stbm153 mutant eyes exhibit both over- and under-rotated ommatidia. (D) fredH24/+, stbm153/stbm153. Reducing fred activity strongly suppresses the stbm rotation phenotype. The fmifrz3 phenotype (E) is enhanced by both loss of ed function in ed1x5/+, fmifrz3/fmifrz3 (F) and loss of fred function in fredH24/+, fmifrz3/fmifrz (G).
These striking findings not only reveal a distinct association between ed and fred and the R3- and R4-specific PCP genes, respectively, but they are also unexpected in light of the mosaic analysis data, which identify roles for ed and fred in R1, R6 and R7, but not in R3 and R4 (Fig. 5). Furthermore, excess Ed and Fred protein in R3 and R4 at the beginning of rotation slows the process (sev>ed and sev>fred, Fig. 4). These findings raise the intriguing possibility that the PCP genes act in unique subsets of cells to control the three distinct events under their control. The localization patterns of the PCP proteins are consistent with such a model as several, including Stbm, localize not only at the R3/R4 interface, but also at the interfaces between the R7/R8, the R7/R1 and the R7/R6 photoreceptor cells. Furthermore, like nmo, ed and fred are clearly required in R7 to regulate ommatidial rotation. Although previous mosaic analyses have not uncovered a requirement for the PCP genes in any cells other than R3 and R4, those analyses measured the composite phenotype (R3/R4 fate, degree and direction of rotation). Consequently, a role for a subset of photoreceptors in one of these events could have been masked.
We therefore re-examined the requirement for stbm in PCP, but focused specifically on its role in ommatidial rotation. This analysis revealed a requirement for stbm function in photoreceptor R7 in regulating the degree of rotation. Remarkably, loss of stbm function in R7 could account for almost all of the degree-of-rotation errors in mosaic stbm15cn ommatidia: when R7 was genotypically wild-type for stbm, the variance in the degree of rotation, (s.d.=5) was very close to that of the wild type (s.d.=1.7); when R7 was genotypically mutant for stbm, the variance was significantly greater (s.d.=15.7) (see Fig. S3 and Table S2 in the supplementary material). These results provide the first demonstration (1) of a genetic requirement for any PCP gene outside the R3/R4 pair, and (2) that the PCP genes act in distinct subsets of cells to control the genetically separable aspects of the PCP phenotype. This novel result, in conjunction with the localization of Stbm at the tip of R7 and in the cone cells, provides an exciting new perspective as to how the PCP complex might regulate the degree to which ommatidia rotate.
The Ed, Fred and core PCP proteins localize to the R3/R4 boundary at approximately the same stage of development (Bastock et al., 2003; Strutt et al., 2002; Strutt, 2002). In addition, the Fred, Stbm and Fmi localization patterns during rotation bear a strong resemblance to one another (Rawls and Wolff, 2003). Although these observations raise the possibility that the PCP proteins might influence Ed and Fred localization, or vice versa, molecular epistasis analyses failed to uncover such a link, as Stbm and Fmi localization was unaffected in ed and fred mutant clones, and Ed and Fred were not mislocalized in clones of the PCP genes stbm and fmi (data not shown). Since protein localization does not appear to be the mechanism by which the PCP complex modulates Ed/Fred activity, an alternative possibility is that the core PCP genes act in a pathway parallel to ed and fred, indirectly regulating these two genes.
DISCUSSION
Here, we demonstrate that ed and fred have partially overlapping functions during the two phases of ommatidial rotation. We propose that different levels of Ed and Fred in rotating and non-rotating cells modulate the adhesivity of these cells, a prerequisite for rotation to occur. In the second phase, Ed and Fred are required in R1, R6, R7 and the cone cells, where they are likely to regulate the Egf receptor to contribute to the slowing of rotation.
There are two phases of rotation distinguishable by the rate at which the ommatidia rotate (Fiehler and Wolff, 2007). The initial phase (rows 4-7) is fast, with ommatidia rotating 10-15° per row, whereas rotation slows to 5-10° per row in the slow phase (rows 7-15). Our data demonstrate that Ed and Fred function during both phases and that they play unique roles in each phase (Fig. 8).
Fig. 8.
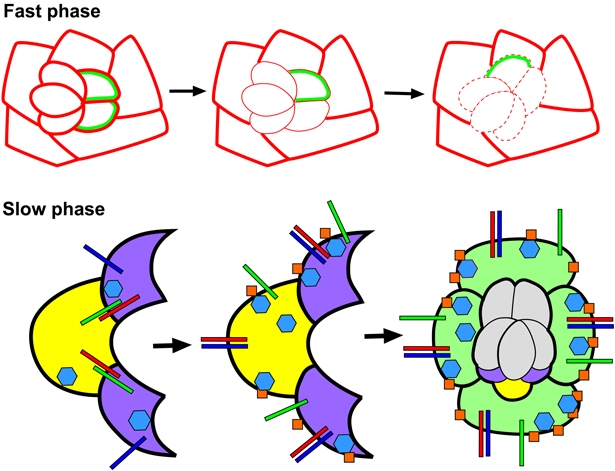
Ed and Fred contribute to both phases of rotation. (Above) Differential levels or expression domains of Ed and Fred in rotating and non-rotating cells create a permissive environment for the faster phase of rotation. Levels of Ed are equivalent in cells within nascent ommatidial preclusters and IOCs (solid red lines of equal weight in the left panel). Immediately before rotation, cells that will rotate actively reduce their levels/distribution of Ed and Fred (middle; reduced Ed levels, thin red line; reduced number of cells expressing Fred, green). A decrease in adhesion (dashed red line, right) between rotating and stationary cells enables rotation to proceed. (Below) Ed and Fred regulate Egfr signaling during the slow phase of ommatidial rotation. When photoreceptors R1, R6 (purple) and R7 (yellow) join the cluster, they contain high levels of both Ed (red bars) and Fred (green bars). During the fast phase, Ed/Fred binding reduces inhibition by Ed of the Egf receptor (blue bars, left). Robust Egfr signaling inhibits Cno (blue hexagons) activity and, consequently, few stable adherens junctions (AJs) form (orange squares). Concurrent with the slower phase of rotation, Ed levels increase in R1, R6 and R7. Ed associates with the Egf receptor, inhibiting Egfr signaling (middle). As a result, Cno activity increases and stable AJs form between moving and stationary cells, effectively applying a brake at the rotation interface (middle). Rotation-specific signaling events shift to a new rotation interface upon recruitment of the cone cells (light green) into the cluster. At the completion of rotation, levels of AJ proteins (Ed, Fred, Cno and Arm) are high, an indication that these two subsets of cells adhere strongly to one another.
In the first phase, we propose that the tight regulation of Ed and Fred levels between rotating and stationary cells creates an environment that is permissive to rotation. Immediately before rotation starts, Ed begins to be endocytosed in the ommatidial precluster cells. Concurrently, Ed levels fall dramatically in these cells while remaining high in the stationary IOCs, setting up an imbalance in Ed levels between these two populations of cells. We propose that the resulting differential adhesion between these two cell populations enables the rotating cells to slide past their stationary neighbors in accordance with Malcolm Steinberg's differential adhesion hypothesis (DAH) (Steinberg, 2007). The DAH suggests that cell populations maximize the strength of adhesive bonding between them and minimize the adhesive free energy, and use tension generated by adhesion between cells to drive events such as cell rearrangements during morphogenesis. Cells with equivalent levels of Ed (or Fred) adhere more tightly to one another and adhesion is reduced between cells with different levels of Ed (or Fred) (Spencer and Cagan, 2003), thereby enabling the two groups to slide past one another. In support of this hypothesis, artificially equalizing levels of Ed or Fred significantly slows rotation.
The data presented here are consistent with Ed and Fred playing two key roles in the slow phase by both directly and indirectly (through Egfr signaling) affecting the physical component of the process. We suggest that in both cases the outputs produce adhesive forces that slow/stop rotation. Ed and Fred are required in photoreceptors R1, R6 and R7 and the cone cells for normal ommatidial rotation. These cells do not become fully integrated into the ommatidial cluster until the second half of rotation. Furthermore, R1, R6 and R7 constitute the rotation interface until the cone cells are recruited, at which point the cone cells co-opt this position and role. Consequently, Ed and Fred are required in the right place (the subset of cells that lie at the rotation interface) and at the right time (the slower phase of rotation) to play a role in slowing rotation.
We propose that Ed and Fred activity in R1, R6, R7 and the cone cells regulates Egfr signaling in these cells to slow/stop rotation as follows. Egfr signaling promotes rotation via the Ras/Cno and Ras/Mapk/Pnt effectors (Brown and Freeman, 2003; Gaengel and Mlodzik, 2003), so its output must be dampened to slow rotation. Ed binds and inhibits the Egf receptor (Bai et al., 2001; Rawlins et al., 2003; Spencer and Cagan, 2003), whereas Fred binds Ed and interferes with this inhibition (S.A.S., unpublished). Therefore, cooperation between Ed and Fred precisely titrates Egfr activity in the cells in which Ed and Fred function. As R1, R6 and R7 are recruited into the ommatidial cluster, Ed levels are high in these cells, thereby decreasing Egfr signaling at their side of the rotation interface, thus impeding rotation. This inhibitory role switches to the cone cells when they are recruited, creating a new rotation interface.
Rotation may be slowed through Egfr signaling activity via its effector Cno/Afadin, an actin-binding adherens junction (AJ) protein (Mandai et al., 1997; Matsuo et al., 1997; Miyamoto et al., 1995; Ooshio et al., 2007). Afadin and its binding partners, nectins and α-actinin, build and stabilize dynamic AJs that undergo remodeling (Ooshio et al., 2007; Takahashi et al., 1999). The majority of cno mutant ommatidia over-rotate, indicating that Cno inhibits ommatidial rotation (Fig. 6E,H). Since Egfr signaling promotes and Cno inhibits rotation, Egfr signaling is likely to suppress Cno activity during rotation thereby blocking stable junction formation. In this scenario, high levels of Egfr would be required during the early phase of rotation to prevent Cno from promoting stable junctions between rotating and non-rotating cells. Consistent with this hypothesis, levels of Ed, an Egfr inhibitor, are very low in ommatidial cells both when rotation commences and during the fast phase of rotation.
Early in the second half of rotation, we propose that higher levels of Ed activity are necessary to repress Egfr signaling at the rotation interface, possibly increasing the amount of active Cno and consequently increasing the number of stable AJs between the moving and stationary cells. The more tightly the cells adhere to one another, the less permissive the environment is for movement, and the more difficult rotation becomes. Ed levels are high in the cells in which it would need to be high, i.e. R1, R6, R7 and the cone cells. Once rotation is complete, Ed and Fred are at high levels at the cell boundaries between the interommatidial and ommatidial cells, an indication that stable AJs now cement the fully rotated ommatidia in place.
ed and fred interact genetically with the R3 and R4 genes, respectively, modifying only the degree-of-rotation aspect of the PCP phenotype. Our genetic and molecular epistasis data suggest that ed and fred act in a pathway either downstream of, or parallel to, the PCP genes. First, localization of Ed and Fred does not require the PCP complex, nor do the PCP proteins require Ed and Fred for their localization. Second, mutations in ed and fred affect only one aspect of the PCP phenotype.
Nectins and afadins have been implicated in numerous human diseases and developmental defects, including breast cancer, metastasis and cleft palate. Defective cell adhesion and cell signaling also underlie these problems. Given the interspecies conservation of AJ genes, similar mechanisms might control ommatidial rotation and contribute to these human diseases.
Supplementary material
Supplementary material for this article is available at http://dev.biologists.org/cgi/content/full/136/19/3323/DC1
Supplementary Material
We thank A. Rawls, S. Arur and J. Skeath for critical comments on the manuscript; R. Fiehler and B. Grillo-Hill for helpful discussions; A. Jarman and K. Takahashi for antibodies; and J.-C. Hsu, H. Vaessin, U. Gaul and the Bloomington Stock Center for fly stocks. This investigation was supported by NIH grant R01 EY13136 to T.W. Deposited in PMC for release after 12 months.
References
- Bai, J., Chiu, W., Wang, J., Tzeng, T., Perrimon, N. and Hsu, J. (2001). The cell adhesion molecule Echinoid defines a new pathway that antagonizes the Drosophila EGF receptor signaling pathway. Development 128, 591-601. [DOI] [PubMed] [Google Scholar]
- Bastock, R., Strutt, H. and Strutt, D. (2003). Strabismus is asymmetrically localised and binds to Prickle and Dishevelled during Drosophila planar polarity patterning. Development 130, 3007-3014. [DOI] [PubMed] [Google Scholar]
- Boettner, B., Harjes, P., Ishimaru, S., Heke, M., Fan, H. Q., Qin, Y., Van Aelst, L. and Gaul, U. (2003). The AF-6 homolog canoe acts as a Rap1 effector during dorsal closure of the Drosophila embryo. Genetics 165, 159-169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown, K. E. and Freeman, M. (2003). Egfr signalling defines a protective function for ommatidial orientation in the Drosophila eye. Development 130, 5401-5412. [DOI] [PubMed] [Google Scholar]
- Chae, J., Kim, M. J., Goo, J. H., Collier, S., Gubb, D., Charlton, J., Adler, P. N. and Park, W. J. (1999). The Drosophila tissue polarity gene starry night encodes a member of the protocadherin family. Development 126, 5421-5429. [DOI] [PubMed] [Google Scholar]
- Chandra, S., Ahmed, A. and Vaessin, H. (2003). The Drosophila IgC2 domain protein Friend-of-Echinoid, a paralogue of Echinoid, limits the number of sensory organ precursors in the wing disc and interacts with the Notch signaling pathway. Dev. Biol. 256, 302-316. [DOI] [PubMed] [Google Scholar]
- Choi, K. W. and Benzer, S. (1994). Rotation of photoreceptor clusters in the developing Drosophila eye requires the nemo gene. Cell 78, 125-136. [DOI] [PubMed] [Google Scholar]
- de Belle, J. S., Sokolowski, M. B. and Hilliker, A. J. (1993). Genetic analysis of the foraging microregion of Drosophila melanogaster. Genome 36, 94-101. [DOI] [PubMed] [Google Scholar]
- Djiane, A., Riou, J., Umbhauer, M., Boucaut, J. and Shi, D. (2000). Role of frizzled 7 in the regulation of convergent extension movements during gastrulation in Xenopus laevis. Development 127, 3091-3100. [DOI] [PubMed] [Google Scholar]
- Fanto, M. and Mlodzik, M. (1999). Asymmetric Notch activation specifies photoreceptors R3 and R4 and planar polarity in the Drosophila eye. Nature 397, 523-526. [DOI] [PubMed] [Google Scholar]
- Fanto, M., Mayes, C. A. and Mlodzik, M. (1998). Linking cell-fate specification to planar polarity: determination of the R3/R4 photoreceptors is a prerequisite for the interpretation of the Frizzled mediated polarity signal. Mech. Dev. 74, 51-58. [DOI] [PubMed] [Google Scholar]
- Feiguin, F., Hannus, M., Mlodzik, M. and Eaton, S. (2001). The ankyrin repeat protein Diego mediates Frizzled-dependent planar polarization. Dev. Cell 1, 93-101. [DOI] [PubMed] [Google Scholar]
- Fiehler, R. W. and Wolff, T. (2007). Drosophila Myosin II, Zipper, is essential for ommatidial rotation. Dev. Biol. 310, 348-362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiehler, R. W. and Wolff, T. (2008). Nemo is required in a subset of photoreceptors to regulate the speed of ommatidial rotation. Dev. Biol. 313, 533-544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Formstone, C. J. and Mason, I. (2005). Combinatorial activity of Flamingo proteins directs convergence and extension within the early zebrafish embryo via the planar cell polarity pathway. Dev. Biol. 282, 320-335. [DOI] [PubMed] [Google Scholar]
- Freeman, M. (1997). Cell determination strategies in the Drosophila eye. Development 124, 261-270. [DOI] [PubMed] [Google Scholar]
- Gaengel, K. and Mlodzik, M. (2003). Egfr signaling regulates ommatidial rotation and cell motility in the Drosophila eye via MAPK/Pnt signaling and the Ras effector Canoe/AF6. Development 130, 5413-5423. [DOI] [PubMed] [Google Scholar]
- Gorfinkiel, N. and Arias, A. M. (2007). Requirements for adherens junction components in the interaction between epithelial tissues during dorsal closure in Drosophila. J. Cell Sci. 120, 3289-3298. [DOI] [PubMed] [Google Scholar]
- Harrington, M. J., Hong, E., Fasanmi, O. and Brewster, R. (2007). Cadherin-mediated adhesion regulates posterior body formation. BMC Dev. Biol. 7, 130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hermiston, M. L., Wong, M. H. and Gordon, J. I. (1996). Forced expression of E-cadherin in the mouse intestinal epithelium slows cell migration and provides evidence for nonautonomous regulation of cell fate in a self-renewing system. Genes Dev. 10, 985-996. [DOI] [PubMed] [Google Scholar]
- Inagaki, M., Irie, K., Ishizaki, H., Tanaka-Okamoto, M., Miyoshi, J. and Takai, Y. (2006). Role of cell adhesion molecule nectin-3 in spermatid development. Genes Cells 11, 1125-1132. [DOI] [PubMed] [Google Scholar]
- Islam, R., Weis, S., Wei-Hsin, C., Hortsch, M. and Hsu, J.-C. (2003). Neuroglian activates Echinoid to antagonize the Drosophila EGF receptor signaling pathway. Development 130, 2051-2059. [DOI] [PubMed] [Google Scholar]
- Jamora, C. and Fuchs, E. (2002). Intercellular adhesion, signalling and the cytoskeleton. Nat. Cell Biol. 4, E101-E108. [DOI] [PubMed] [Google Scholar]
- Kim, S. H., Jen, W. C., De Robertis, E. M. and Kintner, C. (2000). The protocadherin PAPC establishes segmental boundaries during somitogenesis in Xenopus embryos. Curr. Biol. 10, 821-830. [DOI] [PubMed] [Google Scholar]
- Klein, T. J. and Mlodzik, M. (2005). Planar cell polarization: an emerging model points in the right direction. Annu. Rev. Cell Dev. Biol. 21, 155-176. [DOI] [PubMed] [Google Scholar]
- Klingensmith, J., Nusse, R. and Perrimon, N. (1994). The Drosophila segment polarity gene dishevelled encodes a novel protein required for response to the wingless signal. Genes Dev. 8, 118-130. [DOI] [PubMed] [Google Scholar]
- Lin, H. P., Chen, H. M., Wei, S. Y., Chen, L. Y., Chang, L. H., Sun, Y. J., Huang, S. Y. and Hsu, J. C. (2007). Cell adhesion molecule Echinoid associates with unconventional myosin VI/Jaguar motor to regulate cell morphology during dorsal closure in Drosophila. Dev. Biol. 311, 423-433. [DOI] [PubMed] [Google Scholar]
- Mandai, K., Nakanishi, H., Satoh, A., Obaishi, H., Wada, M., Nishioka, H., Itoh, M., Mizoguchi, A., Aoki, T., Fujimoto, T. et al. (1997). Afadin: A novel actin filament-binding protein with one PDZ domain localized at cadherin-based cell-to-cell adherens junction. J. Cell Biol. 139, 517-528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuo, T., Takahashi, K., Kondo, S., Kaibuchi, K. and Yamamoto, D. (1997). Regulation of cone cell formation by Canoe and Ras in the developing Drosophila eye. Development 124, 2671-2680. [DOI] [PubMed] [Google Scholar]
- Matsushima, H., Utani, A., Endo, H., Matsuura, H., Kakuta, M., Nakamura, Y., Matsuyoshi, N., Matsui, C., Nakanishi, H., Takai, Y. et al. (2003). The expression of nectin-1alpha in normal human skin and various skin tumours. Br. J. Dermatol. 148, 755-762. [DOI] [PubMed] [Google Scholar]
- Mirkovic, I. and Mlodzik, M. (2006). Cooperative activities of Drosophila DE-cadherin and DN-cadherin regulate the cell motility process of ommatidial rotation. Development 133, 3283-3293. [DOI] [PubMed] [Google Scholar]
- Miyamoto, H., Nihonmatsu, I., Kondo, S., Ueda, R., Togashi, S., Hirata, K., Ikegami, Y. and Yamamoto, D. (1995). canoe encodes a novel protein containing a GLGF/DHR motif and functions with Notch and scabrous in common developmental pathways in Drosophila. Genes Dev. 9, 612-625. [DOI] [PubMed] [Google Scholar]
- Mueller, S., Rosenquist, T. A., Takai, Y., Bronson, R. A. and Wimmer, E. (2003). Loss of nectin-2 at Sertoli-spermatid junctions leads to male infertility and correlates with severe spermatozoan head and midpiece malformation, impaired binding to the zona pellucida, and oocyte penetration. Biol. Reprod. 69, 1330-1340. [DOI] [PubMed] [Google Scholar]
- Naora, H. and Montell, D. J. (2005). Ovarian cancer metastasis: integrating insights from disparate model organisms. Nat. Rev. Cancer 5, 355-366. [DOI] [PubMed] [Google Scholar]
- Nelson, W. J. (2003). Adaptation of core mechanisms to generate cell polarity. Nature 422, 766-774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niewiadomska, P., Godt, D. and Tepass, U. (1999). DE-Cadherin is required for intercellular motility during Drosophila oogenesis. J. Cell Biol. 144, 533-547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ooshio, T., Fujita, N., Yamada, A., Sato, T., Kitagawa, Y., Okamoto, R., Nakata, S., Miki, A., Irie, K. and Takai, Y. (2007). Cooperative roles of Par-3 and afadin in the formation of adherens and tight junctions. J. Cell Sci. 120, 2352-2365. [DOI] [PubMed] [Google Scholar]
- Ozaki-Kuroda, K., Nakanishi, H., Ohta, H., Tanaka, H., Kurihara, H., Mueller, S., Irie, K., Ikeda, W., Sakai, T., Wimmer, E. et al. (2002). Nectin couples cell-cell adhesion and the actin scaffold at heterotypic testicular junctions. Curr. Biol. 12, 1145-1150. [DOI] [PubMed] [Google Scholar]
- Pacquelet, A. and Rorth, P. (2005). Regulatory mechanisms required for DE-cadherin function in cell migration and other types of adhesion. J. Cell Biol. 170, 803-812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perez-Moreno, M., Jamora, C. and Fuchs, E. (2003). Sticky business: orchestrating cellular signals at adherens junctions. Cell 112, 535-548. [DOI] [PubMed] [Google Scholar]
- Pignatelli, M. (1998). Integrins, cadherins, and catenins: molecular cross-talk in cancer cells. J. Pathol. 186, 1-2. [DOI] [PubMed] [Google Scholar]
- Rawlins, E. L., White, N. M. and Jarman, A. P. (2003). Echinoid limits R8 photoreceptor specification by inhibiting inappropriate EGF receptor signalling within R8 equivalence groups. Development 130, 3715-3724. [DOI] [PubMed] [Google Scholar]
- Rawls, A. S. and Wolff, T. (2003). Strabismus requires Flamingo and Prickle function to regulate tissue polarity in the Drosophila eye. Development 130, 1877-1887. [DOI] [PubMed] [Google Scholar]
- Ready, D. F., Hanson, T. E. and Benzer, S. (1976). Development of the Drosophila retina, a neurocrystalline lattice. Dev. Biol. 53, 217-240. [DOI] [PubMed] [Google Scholar]
- Sakisaka, T., Ikeda, W., Ogita, H., Fujita, N. and Takai, Y. (2007). The roles of nectins in cell adhesions: cooperation with other cell adhesion molecules and growth factor receptors. Curr. Opin. Cell Biol. 19, 593-602. [DOI] [PubMed] [Google Scholar]
- Sozen, M. A., Suzuki, K., Tolarova, M. M., Bustos, T., Fernandez Iglesias, J. E. and Spritz, R. A. (2001). Mutation of PVRL1 is associated with sporadic, non-syndromic cleft lip/palate in northern Venezuela. Nat. Genet. 29, 141-142. [DOI] [PubMed] [Google Scholar]
- Spencer, S. A. and Cagan, R. L. (2003). Echinoid is essential for regulation of Egfr signaling and R8 formation during Drosophila eye development. Development 130, 3725-3733. [DOI] [PubMed] [Google Scholar]
- Spencer, S. A., Powell, P. A., Miller, D. T. and Cagan, R. L. (1998). Regulation of EGF receptor signaling establishes pattern across the developing Drosophila retina. Development 125, 4777-4790. [DOI] [PubMed] [Google Scholar]
- Steinberg, M. S. (2007). Differential adhesion in morphogenesis: a modern view. Curr. Opin. Genet. Dev. 17, 281-286. [DOI] [PubMed] [Google Scholar]
- Strutt, D., Johnson, R., Cooper, K. and Bray, S. (2002). Asymmetric localization of frizzled and the determination of notch-dependent cell fate in the Drosophila eye. Curr. Biol. 12, 813-824. [DOI] [PubMed] [Google Scholar]
- Strutt, D. I. (2002). The asymmetric subcellular localisation of components of the planar polarity pathway. Semin. Cell Dev. Biol. 13, 225-231. [DOI] [PubMed] [Google Scholar]
- Strutt, H. and Strutt, D. (2003). EGF signaling and ommatidial rotation in the Drosophila eye. Curr. Biol. 13, 1451-1457. [DOI] [PubMed] [Google Scholar]
- Suzuki, K., Bustos, T. and Spritz, R. A. (1998). Linkage disequilibrium mapping of the gene for Margarita Island ectodermal dysplasia (ED4) to 11q23. Am. J. Hum. Genet. 63, 1102-1107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki, K., Hu, D., Bustos, T., Zlotogora, J., Richieri-Costa, A., Helms, J. A. and Spritz, R. A. (2000). Mutations of PVRL1, encoding a cell-cell adhesion molecule/herpesvirus receptor, in cleft lip/palate-ectodermal dysplasia. Nat. Genet. 25, 427-430. [DOI] [PubMed] [Google Scholar]
- Takahashi, K., Matsuo, T., Katsube, T., Ueda, R. and Yamamoto, D. (1998). Direct binding between two PDZ domain proteins Canoe and ZO-1 and their roles in regulation of the jun N-terminal kinase pathway in Drosophila morphogenesis. Mech. Dev. 78, 97-111. [DOI] [PubMed] [Google Scholar]
- Takahashi, K., Nakanishi, H., Miyahara, M., Mandai, K., Satoh, K., Satoh, A., Nishioka, H., Aoki, J., Nomoto, A., Mizoguchi, A. et al. (1999). Nectin/PRR: an immunoglobulin-like cell adhesion molecule recruited to cadherin-based adherens junctions through interaction with Afadin, a PDZ domain-containing protein. J. Cell Biol. 145, 539-549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tepass, U., Godt, D. and Winklbauer, R. (2002). Cell sorting in animal development: signalling and adhesive mechanisms in the formation of tissue boundaries. Curr. Opin. Genet. Dev. 12, 572-582. [DOI] [PubMed] [Google Scholar]
- Tree, D. R., Shulman, J. M., Rousset, R., Scott, M. P., Gubb, D. and Axelrod, J. D. (2002). Prickle mediates feedback amplification to generate asymmetric planar cell polarity signaling. Cell 109, 371-381. [DOI] [PubMed] [Google Scholar]
- Tsukita, S., Furuse, M. and Itoh, M. (2001). Multifunctional strands in tight junctions. Nat. Rev. Mol. Cell Biol. 2, 285-293. [DOI] [PubMed] [Google Scholar]
- Usui, T., Shima, Y., Shimada, Y., Hirano, S., Burgess, R. W., Schwarz, T. L., Takeichi, M. and Uemura, T. (1999). Flamingo, a seven-pass transmembrane cadherin, regulates planar cell polarity under the control of Frizzled. Cell 98, 585-595. [DOI] [PubMed] [Google Scholar]
- Wei, S. Y., Escudero, L. M., Yu, F., Chang, L. H., Chen, L. Y., Ho, Y. H., Lin, C. M., Chou, C. S., Chia, W., Modolell, J. et al. (2005). Echinoid is a component of adherens junctions that cooperates with DE-Cadherin to mediate cell adhesion. Dev. Cell 8, 493-504. [DOI] [PubMed] [Google Scholar]
- Wolff, T. (2000). Histological techniques for the Drosophila eye. Parts I and II. In Drosophila Protocols (ed. M. A. W. Sullivan and R. S. Hawley), pp. 201-244. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
- Wolff, T. and Ready, D. F. (1993). Pattern formation in the Drosophila retina. In The Development of Drosophila melanogaster (ed. M. Bate and A. Martinez-Arias). Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press.
- Wolff, T. and Rubin, G. M. (1998). Strabismus, a novel gene that regulates tissue polarity and cell fate decisions in Drosophila. Development 125, 1149-1159. [DOI] [PubMed] [Google Scholar]
- Xu, T. and Rubin, G. M. (1993). Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223-1237. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.