Abstract
Coordinated execution of DNA replication, checkpoint activation, and postreplicative chromatid cohesion is intimately related to the replication fork machinery. Human AND-1/chromosome transmission fidelity 4 is localized adjacent to replication foci and is required for efficient DNA synthesis. In S phase, AND-1 is phosphorylated in response to replication arrest in a manner dependent on checkpoint kinase, ataxia telangiectasia-mutated, ataxia telangiectasia-mutated and Rad3-related protein, and Cdc7 kinase but not on Chk1. Depletion of AND-1 increases DNA damage, delays progression of S phase, leads to accumulation of late S and/or G2 phase cells, and induces cell death in cancer cells. It also elevated UV-radioresistant DNA synthesis and caused premature recovery of replication after hydroxyurea arrest, indicating that lack of AND-1 compromises checkpoint activation. This may be partly due to the decreased levels of Chk1 protein in AND-1-depleted cells. Furthermore, AND-1 interacts with cohesin proteins Smc1, Smc3, and Rad21/Scc1, consistent with proposed roles of yeast counterparts of AND-1 in sister chromatid cohesion. Depletion of AND-1 leads to significant inhibition of homologous recombination repair of an I-SceI-driven double strand break. Based on these data, we propose that AND-1 coordinates multiple cellular events in S phase and G2 phase, such as DNA replication, checkpoint activation, sister chromatid cohesion, and DNA damage repair, thus playing a pivotal role in maintenance of genome integrity.
Replication fork is not only the site of DNA synthesis but also the center for coordinated execution of various chromosome transactions. The preparation for replication forks starts in the G1 phase, when the prereplicative complex composed of origin recognition and minichromosome maintenance assembles on the chromosome. At the G1-S boundary, Cdc45, GINS complex, and other factors join the prereplicative complex to generate a complex capable of initiating DNA replication. A series of phosphorylation events mediated by cyclin-dependent kinase and Cdc7 kinase play crucial roles in this process and facilitate the generation of active replication forks (1–6). Purification of the putative replisome complex in yeast indicated the presence of the checkpoint mediator Mrc1 and fork protection complex proteins Tof1 and Csm3 in the replication fork machinery (7), consistent with a previous report on the genome-wide analyses with chromatin immunoprecipitation analyses on chip (microarray) (8). Mcm10 is another factor present in the isolated complex, required for loading of replication protein A (RPA)2 and primase-DNA polymerase α onto the replisome complex (7, 9, 10).
Replication fork machinery can cope with various stresses, including shortage of the cellular nucleotide pool and replication fork blockages that interfere with its progression. Stalled replication forks activate checkpoint pathways, leading to cell cycle arrest, DNA repair, restart of DNA replication, or cell death in some cases (11–14). Single-stranded DNAs coated with RPA at the stalled replication forks are recognized by the ATR-ATR-interacting protein kinase complex and Rad17 for loading of the Rad9-Rad1-Hus1 checkpoint clamp (14–16). Factors present in the replisome complex are also known to be required for checkpoint activation. Claspin, Tim, and Tipin functionally and physically associate with sensor and effector kinases and serve as mediator/adaptors (17–23). Mcm7, a component of the replicative DNA helicase in eukaryotes, was reported to associate with the checkpoint clamp loader Rad17 (24) and to have a distinct function in checkpoint (24, 25). We recently reported that Cdc7 kinase, known to be required for DNA replication initiation, plays a role in activation of DNA replication checkpoint possibly through regulating Claspin phosphorylation (26). Thus, it appears that DNA replication and checkpoint activation functionally and physically interact with each other.
Another crucial cellular event for maintenance of genome stability is sister chromatid cohesion. The cohesin complex, a conserved apparatus required for sister chromatid cohesion, contains Smc1, Smc3, and Rad21/Scc1/Mcd1 proteins. The assembled cohesin complexes are loaded onto chromatin prior to DNA replication in G1 phase and link the sister chromosomes during S and G2 phase until mitosis when they separate (27, 28). The mitotic cohesion defects are not rescued by supplementing cohesin in G2 phase, and it has been suggested that establishment of sister chromatid cohesion is coupled with DNA replication (29, 30). Indeed, yeast mutants in some replisome components show defect in sister chromosome cohesion or undergo chromosome loss (31–33). Cdc7 kinase is also required for efficient mitotic chromosome cohesion (34, 35).
Human AND-1 is the putative homolog of budding yeast CTF4/Pob1/CHL15 and fission yeast Mcl1/Slr3. The budding yeast counterpart was identified as a replisome component described above (7), which travels along with the replication fork (29). CTF4 is nonessential for viability, but its interactions with primase, Rad2 (FEN1 family of nuclease), and Dna2 have implicated CTF4 in lagging strand synthesis and/or Okazaki fragment processing (36–39). Yeast CTF4 and Mcl1 are involved in chromosome cohesion (33, 40, 41) and genetically interact with a cohesin, Mcd1/Rad21 (40, 42). Recently, it was reported that human AND-1 protein interacts with human primase-DNA polymerase α and Mcm10 and is required for DNA synthesis (43).
Here we confirm that human AND-1 protein is required for DNA replication and efficient progression of S phase, and we further show that it facilitates replication checkpoint. Depletion of AND-1 causes accumulation of DNA damage and cell cycle arrest at late S to G2 phase, ultimately leading to cell death. Furthermore, we also show that human AND-1 physically interacts with cohesin proteins Smc1, Smc3, Rad21/Scc1, suggesting a possibility that AND-1 may physically and functionally link replisome and cohesin complexes in vivo. Recent studies indicate that sister chromatid cohesion is required for recombinational DNA repair (44–47). Thus, we examined the requirement of AND-1 for repair of artificially induced double-stranded DNA breaks and showed that AND-1 depletion leads to significant reduction of the double strand break repair. Possible roles of AND-1 in coordination of various chromosome transactions at a replication fork and in maintenance of genome integrity during S phase will be discussed.
EXPERIMENTAL PROCEDURES
Cell Culture and Cell Cycle Synchronization
HeLa, HeLa S3, and U2OS cells were cultured in Dulbecco's modified Eagle's medium with 10% fetal bovine serum. For synchronization, cells were arrested twice at the G1/S boundary by incubation in the presence of 2.5 mm thymidine for 14–16 h with a 9-h interval of growth without the drug. Cells were released into the cell cycle and harvested at the indicated time intervals. Protein samples were resolved on SDS-polyacrylamide gel and analyzed by immunoblot. To detect the mobility shift of AND-1 in HeLa cells, proteins were separated on 7% polyacrylamide gel (acrylamide/bisacrylamide = 89:1). Cell cycle was analyzed by fluorescence-activated cell sorter as described previously (48).
siRNAs and Transfection
Transfection of small interfering RNA (siRNA) duplexes was conducted by using Oligofectamine (Invitrogen) or X-tremeGENE (Roche Applied Science) transfection reagents as described previously (23). The sense strands of the four different siRNA for human AND-1 were as follows: AND-1 number 1, 5′-GAUGGUCAAGAAGGCAGCAdTdT-3′; AND-1 number 3, 5′-AGGAAAACAUGCCUGCCACdTdT-3′; AND-1 number 4; and 5′-GGUGUAGGUAACAGGACAUdAdT-3′; AND-1 number 5, 5′-CAGAGAUGACUUGUGUAUUGdTdA-3′. siRNA for TopBP1 was 5′-CAGUGGAGGUGGAGUUCGdTdT-3′. siRNAs for Tipin, Cdc7 (Cdc7-1 and Cdc7-D), and control siRNA were described previously (23, 48, 49).
Antibodies
Anti-AND-1Npep antibody was raised by immunizing rabbits with the N-terminal 20-amino acid residue polypeptide of human AND-1, and anti-AND-1C antibody was generated by immunizing rabbits with the glutathione S-transferase-tagged C-terminal domain (712–1129 amino acids) of the human AND-1. Both antibodies were affinity-purified using antigen-coupled protein A-Sepharose or HiTrap NHS-activated HP (GE Healthcare). Rabbit anti-γH2AX antibody was a gift from Katsuyuki Tamai (Medical and Biological Laboratories, Nagoya, Japan). Anti-Mcm4 antibody was reported previously (3). Antibodies from commercial sources were as follows: Chk1, Chk2, PCNA, lamin B, and Mcm7 (Santa Cruz Biotechnology); α-tubulin and FLAG M2 (Sigma); Chk1 S317P, Chk1 S345P, Chk2 T68P, Smc1 S957P, and Cdc25C S216P (Cell Signaling); Claspin, Smc1, Smc3, Rad21, and TopBP1 (Bethyl Laboratories); Cdc7 (MBL); RPA p34 (NeoMarkers); HP1α (Upstate).
DNA Constructions and Transient Transfection
Full-length human AND-1 cDNAs were subcloned into pME18S-FLAG vector. HeLa S3 cells were transfected with DNA using Lipofectamine 2000 (Invitrogen) and were harvested at 48 h, followed by immunoprecipitation and immunoblot analysis.
Immunofluorescence Staining
Preparation of cells for immunofluorescence staining was performed as described previously (23, 50) with some modification. Briefly, for pre-extraction procedure, cells grown on coverslips were washed twice with PHEM buffer (60 mm PIPES, 25 mm HEPES-KOH, pH 6.9, 10 mm EGTA, and 2 mm MgCl2) and lysed with 0.5% Triton X-100. The cells were then fixed with methanol and immediately dried. For post-extraction procedure, PBS-washed cells were fixed with paraformaldehyde/PBS, washed with PBS, and lysed with 0.2% Triton X-100. For immunostaining, the cells on coverslips were blocked with 3% bovine serum albumin and then incubated with primary antibodies in AB buffer (0.1 m PIPES-KOH, pH 7.2, 1 mm MgSO4, 1 mm EGTA, 1.83% l-lysine, 1% bovine serum albumin, and 0.1% NaN3) for 1 h at 37 °C. After washing with PBS, cells were incubated with secondary antibodies (Cy3-conjugated anti-mouse antibody (Jackson ImmunoResearch), Alexa 488-conjugated anti-rabbit antibody (Molecular Probes), and 4,6-diamidino-2-phenylindole (DAPI, 1 μg per ml)) for 45 min at 37 °C. The coverslips were then washed with PBS three times and mounted with PBS containing 90% glycerol, 2.5% 1,4-diazabicyclo(2.2.2)octane (Sigma). Terminal deoxynucleotidyltransferase-mediated dUTP nick-end labeling (TUNEL) assays were performed using in situ cell death detection kit (Roche Applied Science) as instructed by the provider (51).
Bright Light and Fluorescence Microscopy
Cell morphology was observed by phase-contrast microscopy (IX50, Olympus), and images were captured with a digital camera (Camedia, Olympus). For counting cell death or mitotic index, cells were stained with Hoechst 33342, and morphologies of nuclei and mitotic chromosome were observed under a fluorescence microscope (Axiophot, Carl Zeiss) equipped with a Hamamatsu ORCA-ER CCD camera. At least 1,000 cells in 3–5 fields were counted for each transfectant. In Fig. 1, B and C, and supplemental Fig. S2B, grayscale images were pseudocolored and merged using software Aquacosmos (Hamamatsu Photonics).
FIGURE 1.
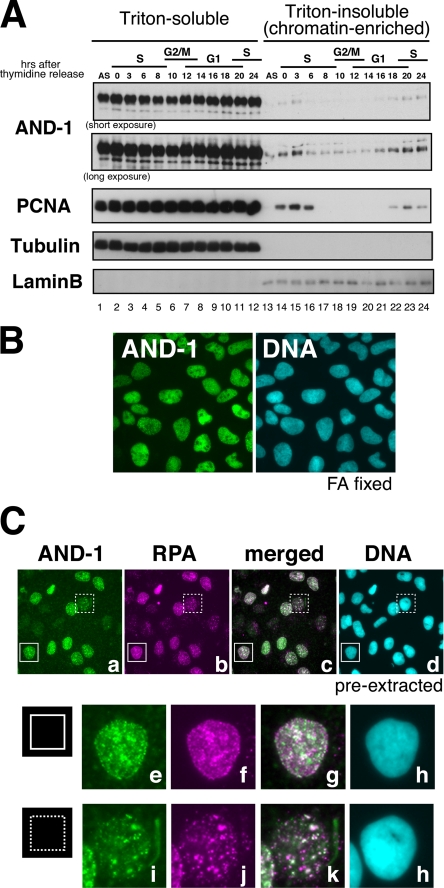
Expression and cellular localization of human AND-1 protein during cell cycle. A, HeLa cells were released from double thymidine block for the indicated time (hours) and fractionated into Triton-soluble and -insoluble fractions. Proteins were analyzed by Western blotting using the antibodies indicated. The affinity-purified anti-AND-1Npep antibody was used. S, S phase; G2/M, G2 phase and mitosis; G1, G1 phase. B, indirect immunofluorescence microscopy. U2OS cells fixed with paraformaldehyde (FA) were stained with anti-AND-1C antibody. DNA was counterstained with DAPI. C, U2OS cells were pre-extracted with detergent and immunostained with anti-AND-1C, anti-RPA, and DAPI (top panels). Enlarged images of two areas (indicated by solid line or dotted line square) in top panels are shown below.
Cell Fractionation
Cells were fractionated into Triton X-100-soluble and -insoluble fractions by the procedure described previously (48). The whole cell extracts were prepared by resuspending the cells in SDS-PAGE sample buffer. Fractionation of S phase cells into five fractions (Fig. 3C) was performed as follows: HeLa cells were extracted with CSK buffer containing 0.1% Triton X-100, centrifuged at 700 × g for 5 min, and separated into supernatants (S1) and pellets, which were further washed with CSK buffer without detergent (wash-out fraction). After digestion of the pellets with DNase I (1,000 units/ml) at 25 °C for 30 min, the soluble fraction was separated as chromatin fraction by centrifugation. Finally, pellets were extracted with CSK buffer containing 0.5 m NaCl, 0.1% Triton X-100 and fractionated into high salt-soluble nuclear fraction (S2) and nuclear scaffold fraction.
FIGURE 3.
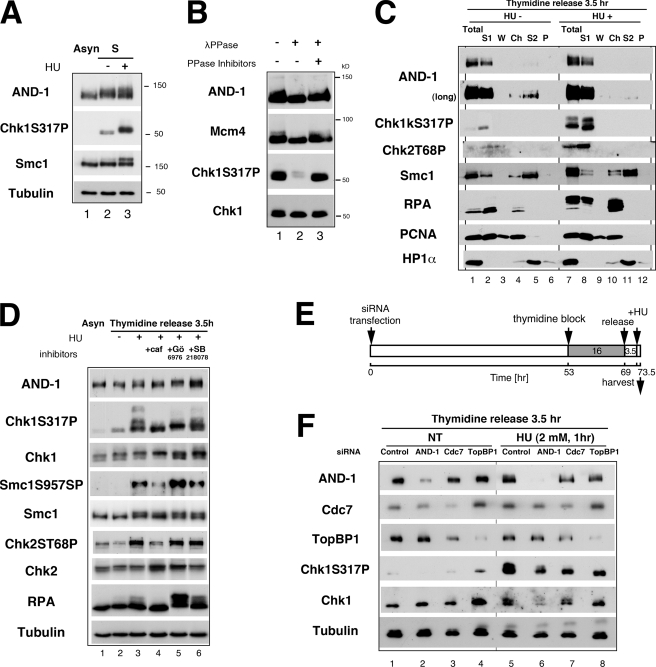
Mobility shift of AND-1 after replication stress depends on ATR/ATM and Cdc7 but not on Chk1 kinase. A, whole cell extracts were prepared from asynchronously (Asyn) growing (lane 1) or S phase-synchronized cells (3.5 h release from single thymidine block; lanes 2 and 3). In lane 3, cells were treated with 5 mm HU for 1 h before harvest. Indicated proteins were detected by immunoblot analysis. B, 30 μg of the whole cell extracts from the HU-treated cells (lane 3 of A) was subjected to phosphatase (λPPase) assay as described under “Experimental Procedures.” After treatment, samples were separated on SDS-PAGE, and proteins were detected by immunoblot with the indicated antibodies. C, HeLa cells, synchronized in S phase by release for 3.5 h from double thymidine block, were harvested untreated (HU−) or after additional incubation with 5 mm HU for 2 h (HU+). Cells were sequentially fractionated into Triton-soluble fractions (S1), washed fractions after extraction (W), supernatants after DNase I digestion (Ch), 0.5 m NaCl-soluble fractions (S2), and insoluble fractions (P). S1 fraction (20 μg of protein) and other fractions from equivalent amounts of cells were analyzed by immunoblot using the indicated antibodies. Total, whole cell extracts from the same numbers of the cells. D, effects of kinase inhibitors on AND-1 protein and other replication/checkpoint proteins. Asynchronous or S phase-synchronized (lanes 2–6; 3.5 h release from single thymidine block) HeLa cells were incubated with HU (2 mm) for 1 h. In lanes 4–6, the following reagents were added at 1.5 h before HU treatment; 5 mm caffeine (caf); 1 μm Gö6976; 5 μm SB218078. Twenty μg of whole cell extracts were resolved on SDS-PAGE, and proteins were detected with indicated antibodies. E, time schedule of the experiment in F as described under “Experimental Procedures.” F, HeLa cells were transfected with siRNA as shown, synchronized at S phase by 3.5 h release from thymidine block, and treated with HU. Whole cell extracts (20 μg of protein) were resolved on SDS-PAGE, and proteins were detected with indicated antibodies. Anti-AND-1C (in A, B, D, and F) or affinity-purified anti-AND-1NPep antibody (in C) and AND-1 number 3 siRNA (in F) were used.
Immunoprecipitation
Cells were extracted with lysis buffer A (20 mm HEPES-NaOH, pH 7.4, 1 mm EDTA, 0.1 mm EGTA, 1 mm MgCl2, 150 mm NaCl, 1 mm Na3VO4, 20 mm NaF, 5% glycerol, 1% Nonidet P-40, 1 μg/ml pepstatin A, 1.5 μg/ml aprotinin, 1 μg/ml leupeptin, and 0.4 mm Pefabloc SC). After rotation for 30 min at 4 °C, the lysate was centrifuged for 20 min at 15,000 × g, and the supernatant was recovered. In Fig. 5A, nuclear extracts lysed with high salt buffer containing 0.35 m KCl and 0.1% Nonidet P-40 were used. Before immunoprecipitation, the extracts were precleared by incubation with protein A- and protein G-Sepharose and centrifugation for 3 min at 11,000 × g. The cleared extracts were immunoprecipitated with the antibodies indicated.
FIGURE 5.
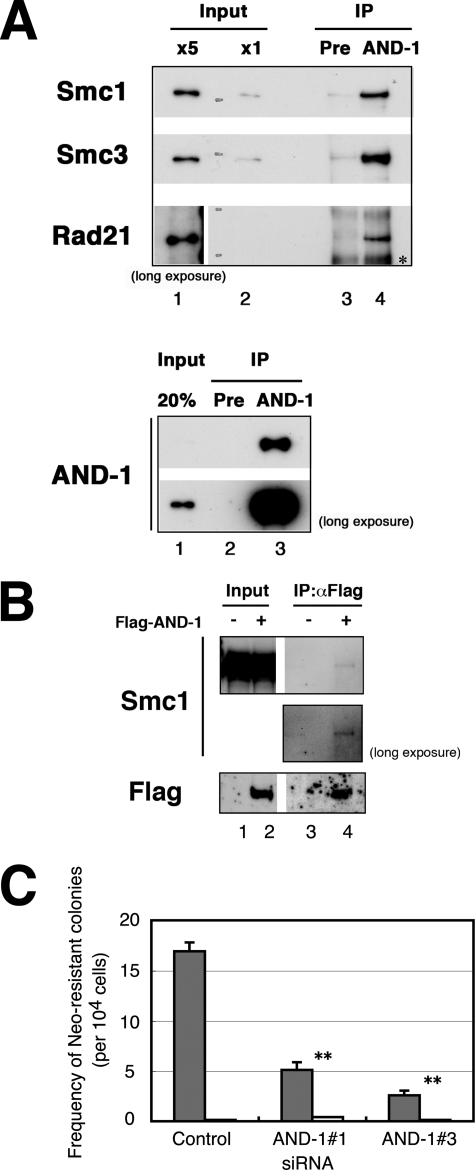
AND-1 interacts with cohesin complex and is required for recombinational repair. A, nuclear extracts (700 μg of protein in top panel; 30 μg of proteins in bottom panel) of HeLa S3 cells were immunoprecipitated (IP) with anti-AND-1C antibody or preimmune serum (Pre) and subjected to immunoblot analysis. In the top panel, input (x5 in lane 1 and x1 in lane 2) represents 0.1 or 0.02%, respectively, of the starting extracts used for immunoprecipitation in lanes 3 and 4. In the bottom panel, input represents 20% of the starting extracts used in lane 3. The asterisk indicates a nonspecific signal. B, HeLa cells were transfected with pME18S-FLAG-AND-1 (+) (lanes 2 and 4) or pME18S (−) (lanes 1 and 3) and harvested at 48 h after transfection. Cells were lysed and immunoprecipitated (IP) with anti-FLAG M2-agarose and analyzed for co-immunoprecipitation of Smc1 protein. Input (lanes 1 and 2) represents 10 μg of the starting extracts used for immunoprecipitation in lanes 3 and 4, respectively. In B, contrast-enhanced images of long exposed films are also shown. C, analysis of I-SceI-induced homologous recombinational repair using a neomycin-based reporter construct. The frequency of neomycin-resistant colonies was counted in control, AND-1 number 1, or AND-1 number 3 siRNA-transfected SW480sn3 cells. The error bars represent S.D. from three independent experiments, and p values are less than 0.01 for both AND-1 number 1 and AND-1 number 3 siRNAs compared with control. Values of cells transfected with I-SceI pCMV3his-I-SceI (gray bars) or empty vector (open bars) are shown. Similar defects in recombinational repair were observed also in cells transfected with AND-1 number 4 and number 5 siRNA (data not shown).
Treatment with Phosphatase and Kinase Inhibitors
For phosphatase assay in vitro, 30 μg of HeLa cell extracts were incubated with λ phosphatase (400 units) with or without phosphatase inhibitors (10 mm Na3VO4 and 50 mm NaF) at 30 °C for 30 min. To dephosphorylate Chk1 S317P, the extracts were treated at 10 °C for 15 min. For treatment with kinase inhibitors in vivo, HeLa cells were synchronized by thymidine block and released for 2 h and then were treated with the following reagents or mock-treated: caffeine (5 mm, Wako), Gö6976 and SB218078 (1 and 5 μm, respectively, Calbiochem). After incubation for 1.5 h, cells were further incubated in the presence of 2 mm hydroxyurea (HU) for 1 h and harvested.
Radioresistant DNA Synthesis (RDS) Assay
RDS after UV irradiation was assayed as reported previously (23, 52) with some modification as shown in Fig. 4, A, C, and D and legend.
FIGURE 4.
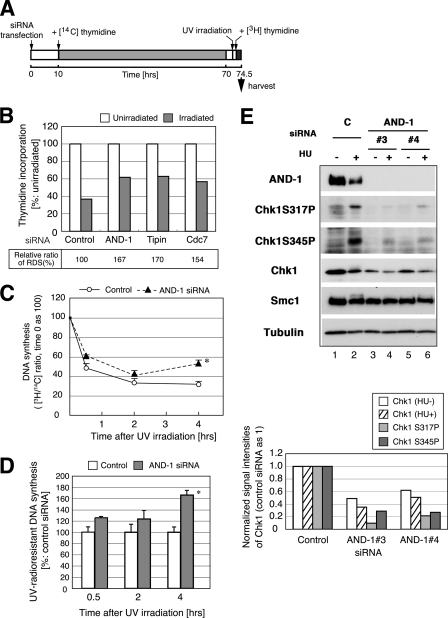
UV radio-resistant DNA synthesis (UV-RDS) in AND-1 knockdown U2OS cells. A, time schedule of the experiment. At 10 h after transfection with control or AND-1 number 3 siRNA, HeLa cells were transferred to the medium containing 20 nCi of [14C]thymidine per ml, followed by incubation for 60 h. After incubation with nonisotopic medium for 2 h, cells were irradiated with UVC (5 J/m2, UV) or mock-irradiated. After 1 h, 10 μCi of [3H]thymidine was added per ml of medium, and incubation was continued for 1.5 h. Cells were harvested, and incorporated radioactivity was measured by scintillation counter. B, ratio of 3H/14C incorporation of AND-1, Tipin, and Cdc7-depleted cells. The values of the unirradiated sample were taken as 100 in each siRNA-treated cells. At the bottom, RDS is shown as values relative to control siRNA, set as 100. We repeated the experiments three times using U2OS or HeLa cells, and the representative data are shown here. C and D, RDS was assessed in time course using the similar protocol described in A except for intensity of UV irradiation (3 J/m2) and for incorporation time of [3H]thymidine (30 min before indicated time). Ratio of 3H/14C incorporation at each time point was plotted. The efficiency of DNA synthesis of unirradiated cells (in C) or that of control siRNA-treated cells after irradiation (in D) was set as 100. Data with asterisks showed significant differences compared with control cells (p < 0.05). Open circles, control siRNA; filled triangles, AND-1 number 4 siRNA. E, knockdown of AND-1 leads to reduction of Chk1 protein level. HeLa cells were transfected with control (C) or AND-1 number 3 or number 4 siRNA and incubated with 2 mm HU for 1 h (+) or untreated (−) before harvest at 48 h after transfection. Twenty μg of protein was resolved on SDS-PAGE, and proteins were detected by the antibodies indicated. The anti-AND-1C antibody was used. In the bottom panel, the signal intensities of Chk1 (untreated, open column; HU-treated, shaded), Chk1 S317P (HU-treated, gray), and Chk1 S345P (HU-treated, dark gray) were normalized with those of α-tubulin. We repeated the experiments five times, and representative data are shown.
DNA Content and Incorporation of BrdUrd
Cells were incubated with BrdUrd and analyzed by fluorescence-activated cell sorter as described previously (48). The integral values of BrdUrd incorporation were determined from calculating the products of geometric means of BrdUrd incorporation and the numbers of BrdUrd-positive cells. The calculated values normalized by those of control siRNA-treated cells (supplemental Fig. S1G) or cells untreated (supplemental Fig. S3B) were set as 100.
Homologous Recombination Assay
The homologous recombination frequency was determined as described previously (53, 54). Briefly, the assay was conducted in SW480sn3 cells harboring a single integrated copy of a recombination substrate SCneo (55, 56). After transfection of cells with siRNAs (48 h), cells were further transfected with pCMV3nls-I-SceI or pME18S plasmid DNA. After 48 h, double strand break-introduced cells were either replated in selection media containing 1 mg of G418 per ml (2 × 105 cells per dish, triplicated) or nonselection media (500 cells per dish, triplicated; for control of colony-forming efficiency). After 2 weeks for selection, colonies were fixed and stained with Giemsa and counted. The recombination frequency was calculated as described previously (57).
RESULTS
Human AND-1 Protein Is Localized on Chromatin in Close Proximity to the Replication Foci during S Phase and Is Required for Efficient Cell Growth
Budding yeast CTF4 protein, the presumed functional homolog of human AND-1, has been reported as a component of replisome progressing complex (7). We have generated two anti-human AND-1 antibodies (see “Experimental Procedures”) and found that the expression of endogenous human AND-1 proteins was constant throughout the cell cycle (data not shown) as reported previously (43). In immunoblot analysis, we found that a portion of AND-1 protein was recovered in chromatin-enriched fraction in early to mid-S phase (Fig. 1A, lanes 14, 15, and 22–24) in a manner similar to that of PCNA. We then analyzed the cellular localization of AND-1 by immunostaining. AND-1 protein was detected in nuclei throughout the interphase in formaldehyde-fixed cells (Fig. 1B) (58). The signal intensities and overall patterns of detergent-resistant AND-1 staining in nuclei (Fig. 1C, panel a) varied dramatically in different cells and were mostly correlated with those of RPA (Fig. 1C, panels b and c), suggesting that the AND-1 may be associated with the sites of DNA replication spatiotemporally.
To explore the physiological functions of AND-1, it was depleted from cells by small interference RNA (siRNA). Four different siRNA oligonucleotide duplexes specific for human AND-1 were designed and introduced into U2OS or HeLa cells. Each transfected siRNA significantly reduced the level of AND-1 protein (Fig. 2A and data not shown). To examine the effects of knockdown of AND-1 on cell growth, cell numbers was monitored at 24, 48, and 72 h after transfection of HeLa cells. Treatment with AND-1 siRNA retarded cell growth (Fig. 2B and phase contrast images in supplemental Fig. S1B).
FIGURE 2.
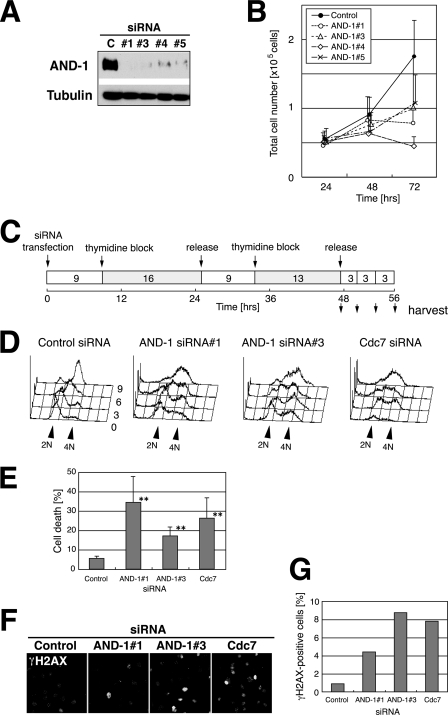
Knockdown of AND-1 delays S phase progression and induces DNA damage and cell death. A, U2OS cells were transfected with four different siRNAs against AND-1 (#1, #3, #4, or #5) or control (C) siRNA for 72 h. Twenty-five-μg proteins of whole cell extracts were analyzed by immunoblot using the antibodies indicated. The affinity-purified anti-AND-1NPep antibody was used. B, numbers of viable HeLa cells transfected with AND-1 siRNAs were counted at 24, 48, or 72 h after transfection. Black circle, control siRNA; open circle, AND-1 siRNA number 1; open triangle, AND-1 siRNA number 3; open diamond, AND-1 siRNA number 4; cross, AND-1 siRNA number 5. Error bars indicate S.D. of three independent transfection experiments. C, time schedule of the experiment in D as described under “Experimental Procedures.” D, DNA contents of siRNA-treated HeLa cells at various time points after double thymidine block and release. E, knockdown of AND-1 induces cell death. HeLa cells were transfected with the indicated siRNA, and the population of dead cells was counted at 77 h as described under “Experimental Procedures.” Error bars indicate S.D. Data with asterisks revealed significant differences compared with control siRNA-transfected cells (p < 0.01). F, DNA damage signals increased by knockdown of AND-1 protein. AND-1-depleted HeLa cells were stained with anti-γH2AX antibody at 72 h after transfection. G, populations of γH2AX-positive cells in F were counted and presented.
Depletion of AND-1 Inhibits DNA Replication and Delays S and G2 Phase Progression
The cell cycle profile of AND-1-depleted cells resulted in slight reduction in G1 phase population and accumulation of late S and/or G2 phase population in both HeLa (supplemental Fig. S1A) and U2OS cells (data not shown). Thus, we examined the effects of AND-1 knockdown on S phase progression and DNA synthesis in more detail. In control cells synchronously released from the G1/S boundary after transfection of siRNA (Fig. 2C), S phase was completed by 9 h after release as presented in the histograms of DNA content analyzed by flow cytometry (Fig. 2D). In AND-1 number 1 or AND-1 number 3 siRNA-treated cells, mid-S to late-S phase transition was retarded (compare the cell populations at 6 or 9 h after release), and DNA synthesis was not completed at 9 h after release. Similar delay in S phase progression was observed in Cdc7 siRNA-treated cells (Fig. 2D) as we reported previously (48). Delay in S phase progression was more directly shown by chasing the BrdUrd-incorporating S phase cells (supplemental Fig. S1, C–G). Thus, consistent with a previous report (43), we have concluded that depletion of human AND-1 compromises the rate of DNA synthesis and especially delays late S and G2 phase progression (see supplemental material).
In addition, cell death was prominently induced at 72 h after transfection in AND-1 siRNA-treated HeLa cells (Fig. 2E), and the extent of cell death was comparable with that observed in Cdc7 siRNA-treated cells (48, 49). Cell death was observed also in U2OS cells (data not shown). These results suggested activation of DNA damage responses by depletion of AND-1. Therefore, we examined the phosphorylated histone H2AX (γ-H2AX) signals, indicative of DNA damages, in AND-1-depleted cells. Immunostaining analysis showed that the population of γ-H2AX-positive cells increased in AND-1-depleted cells (Fig. 2, F and G), as observed in Cdc7-depleted cells (26). Smc1 protein is known to be phosphorylated by ATM/ATR checkpoint kinase in response to DNA damages (59–61). Similar to γ-H2AX, foci of phosphorylated Smc1 were strongly induced in AND-1-depleted cells (supplemental Fig. S2B). To verify the cell death observed was apoptotic, we stained the DNA breaks in situ by TUNEL methods (62) at 72 h after depletion of AND-1. Transfection of AND-1 number 1 or number 3 siRNA leads to an increase in the numbers of TUNEL-positive cells (data not shown). In conclusion, depletion of human AND-1 induced DNA damage and eventually caused apoptosis in HeLa cells.
Phosphorylation of AND-1 Is Induced by Replication Arrest and Depends on ATM/ATR and Cdc7 but Not on Chk1 Kinase
Accumulating evidence has indicated that progression of cell cycle and activation of checkpoint are regulated by phosphorylation of numerous transducer and mediators. Therefore, we examined the phosphorylation status of human AND-1 protein during S phase and after replication stress. We detected retardation of the mobility of AND-1 protein on SDS-PAGE in S phase whole cell extracts (Fig. 3A, lane 2). Further retardation of the AND-1 protein band was observed after treatment of the S phase cells with HU (Fig. 3A, lane 3). Under the same conditions, Chk1 phosphorylation at Ser-317 and phosphorylation-induced mobility shift of Smc1 were also detected (Fig. 3A, lanes 2 and 3). Treatment of HU-treated cell extracts with phosphatase eliminated the mobility shift of AND-1 protein as well as that of Mcm4 and the Chk1 Ser-317 signal, indicating that the mobility shift is due to the phosphorylation by replication stress (Fig. 3B).
It is known that some proteins relocate to specific cellular sites in the presence of DNA damage or replication stress. Thus, we fractionated cells by the conventional method and analyzed the localization of AND-1 and checkpoint mediators after replication stress. Localization of PCNA, lamin B, or HP1α did not significantly change after HU treatment (Fig. 3C and data not shown). As reported previously, phosphorylated forms of checkpoint effector kinases Chk1 and Chk2 were detected only in the soluble fraction before and after replication stress (Fig. 3C, lanes 7 and 8) (63–65). In contrast, RPA and Smc1 were hyperphosphorylated in response to HU, and a significant portion of the proteins was detected in chromatin and insoluble fractions (Fig. 3C, lanes 10 and 11) (60, 66). AND-1 protein exhibited mobility shift on SDS-PAGE after HU treatment, as in Fig. 3A, but most of the proteins remained in soluble fractions, and an increase in chromatin and high salt-soluble fraction was not observed (Fig. 3C).
Mammalian ATM and ATR kinases are the key players in sensing the S phase checkpoint signal and phosphorylating effector kinases Chk1, Chk2, and other substrates, including histone H2AX (12, 67, 68). We examined whether phosphorylation of AND-1 under replication stress depends on these checkpoint kinases. As shown in Fig. 3C, HU treatment of S phase cells leads to phosphorylation of Chk1 Ser-317, Chk2 Thr-68, Smc1 Ser-957, and RPA (Fig. 3D, lane 3), and these phosphorylation events were clearly attenuated by the treatment with caffeine, an inhibitor of phosphatidylinositol 3-kinase, including ATM or ATR (Fig. 3D, lane 4). HU-induced mobility shift of AND-1 from S phase cells was also largely inhibited by caffeine treatment (Fig. 3D, lane 4). Curiously, when replication-arrested cells were treated with Chk1 kinase inhibitors, Gö6976 or SB218078, the mobility shift of AND-1 was further enhanced (Fig. 3D, compare lanes 5 and 6 with lane 3). The Chk1 inhibitors also stimulated the phosphorylation of Smc1 and RPA proteins, although they did not significantly affect phosphorylation of Chk1 and Chk2.
Recently, we reported that Cdc7 may be required for checkpoint kinase activation by replication stress in mammalian cells (26). Because phosphorylation of AND-1 was not completely inhibited by caffeine, we next examined whether Cdc7 kinase has any effects on AND-1 phosphorylation (Fig. 3E). The enhanced mobility shift of AND-1 observed in the presence of HU (Fig. 3F, lane 5) was compromised in Cdc7-depleted cells (Fig. 3F, lane 7). In contrast, depletion of TopBP1, the functional homolog of yeast Dpb11/Cut5 and known as a stimulator of ATR kinase (69, 70), only slightly reduced the HU-dependent mobility shift of AND-1 (Fig. 3F, lane 8). Thus, phosphorylation of AND-1 protein induced by replication arrest depends on ATM and/or ATR kinase as well as on Cdc7 but not on Chk1.
AND-1 Is Required for Efficient Execution of Intra-S Phase Checkpoint
Claspin, Tim, and Tipin proteins play roles in transmitting the checkpoint signals induced by stalled replication forks to downstream effector kinases and also in stable arrest of replication forks. In S phase, ultraviolet or ionizing radiation causes DNA damage or temporarily stalls DNA replication forks and results in inhibition of bulk DNA synthesis until the fork block is removed. We and other groups have previously shown that depletion of Tim or Tipin leads to inhibition of checkpoint activation (18–23) and to induced unscheduled radioresistant DNA synthesis (RDS) after UV irradiation (20, 21, 23). Therefore, we examined the role of AND-1, a possible fork component (7, 29), in intra-S phase checkpoint response. Cells were transfected with siRNA for AND-1, Tipin, or Cdc7, irradiated with UVC, and then were incubated with thymidine (Fig. 4A). AND-1-depleted cells, after irradiation, exhibited more DNA synthesis than did control cells (167 versus 100; Fig. 4B). The extent of the increase was similar to that shown by Tipin or Cdc7 depletion (Fig. 4B). During the time course, UV-RDS was elevated in AND-1-depleted cells at later time points, indicating that premature quenching of the checkpoint activation occurred by depletion of AND-1 (Fig. 4, C and D). We also detected knockdown of AND-1 also increased RDS after brief HU arrest in BrdUrd pulse-labeling methods with kinetics similar to that of UV-RDS (supplemental Fig. S3, A and B).
Mammalian Chk1 is an effector kinase in intra-S phase checkpoint, which is activated by ATR and/or ATM, and transmit the signals to downstream substrate such as Cdc25A (71, 72). Serines 317 and 345 of Chk1 are phosphorylated in response to DNA replication blocks and DNA damages (Fig. 5E, lane 2; also Fig. 4, A, C, D, and F) (73, 74), and these phosphorylations depend on ATR/ATM kinase activity (72). AND-1 depletion resulted in significant reduction of Ser-317 and Ser-345 phosphorylation of Chk1 (Fig. 4E, lanes 4 and 6; Fig. 3F, lane 6). This might be largely due to the decreased amount of Chk1 protein in AND-1-depleted cells (Fig. 4E, lanes 3–6; Fig. 3F, lane 6). To gain insight into regulation of the Chk1 protein level in AND-1-depleted cells, we examined the effect of the protein synthesis inhibitor, cycloheximide, or proteasome inhibitor, MG132. The stability of Chk1 protein did not significantly change in AND-1-depleted cells compared with that in control siRNA cells (supplemental Fig. S3C). Caspase-mediated degradation of Chk1 (75) was also not detected (data not shown).
AND-1 Associates with Cohesin Complexes and Is Required for Homologous Recombination-mediated Repair
The yeast CTF4/POB1 was originally isolated as DNA polymerase α-binding protein, and recently, human AND-1 was reported to bind to DNA polymerase α and Mcm10 through its central SepB homology domain (43). To shed light on the potential multifunctional roles of AND-1, we attempted to identify other AND-1-interacting proteins by immunoprecipitation using AND-1-specific antibodies. Among the cohesin and replication fork components, Smc1, Smc3, Rad21, and Mcm7 were specifically co-immunoprecipitated with endogenous AND-1 protein in asynchronous cell extracts (Fig. 5A and data not shown). The interaction of Smc1 with AND-1 was also confirmed by co-immunoprecipitation with ectopically expressed FLAG-tagged AND-1 protein (Fig. 5B). We next examined these interactions during the cell cycle, and in the presence or absence of replication stress or DNA damage. HeLa cells, synchronized at S, G2, M, or G1 phase, were treated with HU for replication arrest or with UV irradiation for DNA damage and were subjected to immunoprecipitation with anti-AND-1 antibody. Immunoblot analysis of the input for immunoprecipitation shows that Chk1 phosphorylation and mobility shift of Smc1 and Chk1 were detected in HU-treated cells (supplemental Fig. S4A). Smc1, Smc3, and Rad21 were detected throughout the cell cycle in the AND-1 immunoprecipitates, irrespective of DNA damage or replication arrest (supplemental Fig. S4B). In addition, immunoprecipitation from HU-treated cells indicated that phosphorylated and unmodified forms of Smc1 were equally co-immunoprecipitated by AND-1 antibody (supplemental Fig. S4B, lane 3).
To explore physiological significance of interaction between AND-1 and the cohesin complex, we investigated the function of AND-1 in sister chromatid cohesion in mammalian cells. In yeasts, CTF4 or Mcl1 mutation exhibited defects in proper sister chromatid cohesion (33, 40, 41, 76). We first examined the effects of AND-1 depletion on sister chromatid cohesion in mitotic chromosome spread. However, the result was not definitive because of the variations in chromosome morphology that might have overridden the effects of knockdown (data not shown). Lately, several observations in different organisms indicate that proper establishment of sister chromatid cohesion is required for postreplicative repair of double-stranded DNA breaks (44–46, 77). Therefore, we examined the homologous recombination-dependent repair by utilizing a reporter assay in SW480sn3 cells, in which recombinational repair restores the neomycin-resistant gene and one can measure recombination-dependent repair efficiency by counting colony-forming units in the presence of neomycin (55, 56). Transfection of AND-1 number 1 or AND-1 number 3 siRNA resulted in reduction of appearance of neomycin-resistant recombinants by 70 or 85%, respectively, compared with the control siRNA cells (Fig. 5C). The repair efficiency is not due to the lower level of sister chromatids caused by reduced replicated molecules, because late S and G2 phase population increased in AND-1-depleted cells compared with control cells (supplemental Fig. S1A). Thus, these results clearly suggest a novel function of human AND-1 in homologous recombinational repair.
DISCUSSION
In this study, we developed a specific antibody against human AND-1 protein and characterized its cellular localization and functions during the cell cycle progression. Our results confirmed the previous report by Zhu et al. (43) that AND-1 is required for efficient DNA replication. Although the previous report showed that AND-1 depletion specifically affects S phase progression, our data indicate that loss of AND-1 delays the progression of late S through G2 phase (43). This may be because we examined the effect of AND-1 depletion on synchronized cell population, which has permitted us to more precisely identify the cell cycle stage affected.
S phase defects by depletion of human AND-1 may be due to incomplete formation of the replisome complex. Like yeast counterpart CTF4 (7, 29), human AND-1 is likely to be a component of the replisome complex, because its localization and staining pattern were similar to those of RPA (Fig. 1C). It was reported previously that AND-1 is essential for the stability of DNA polymerase α p180 (43). However, the level of DNA polymerase α was only slightly reduced by AND-1 siRNA in HeLa cells (supplemental Fig. S5) in our experimental condition, suggesting that S phase defect may not be simply due to the decreased level of DNA polymerase α protein. Alternatively, the defect of AND-1-depleted cells in recombinational repair (Fig. 5C) of naturally occurring DNA lesions during S phase may result in delay in completion of DNA replication. Increase of BrdUrd-negative cells at very late S and/or G2 phase (circles in supplemental Fig. S1D and supplemental Fig. S1A) could be due to accumulation of unrepaired damages at those stages. The accumulation of G2 cells may not be explained solely by normal DNA damage checkpoint-mediated G2 arrest because no obvious increase of phosphorylation of Chk1 (Fig. 3F and Fig. 4E), Chk2, or Cdc25C Ser-216 (data not shown) is detected.
Replication fork machinery is intimately involved in DNA replication checkpoint signaling induced by replication stress (67). Human fork factors, Tim, Tipin, and Claspin, have been proposed to function as checkpoint mediators as we and other groups previously reported (17–23). Here we show that another human replisome component, AND-1, is required for full activation of replication checkpoint. It is noteworthy that RDS in the absence of human AND-1 was elevated in UV-treated cells, to a level previously seen in Tipin or Cdc7 siRNA-treated cells (Fig. 4C) (20, 23). Although the mechanism of AND-1-mediated checkpoint is not yet clear, we observed the levels of some of the key proteins decreased, including Chk1 (Fig. 3F and Fig. 4E). We also note that Chk1 protein level was reduced by AND-1 depletion, but to a smaller extent in U2OS cells (data not shown), suggesting that genetic background may affect the cellular responses. Down-regulation of the Chk1 protein level was previously reported after depletion of HCLK-2, a clock protein known to be involved in intra-S phase checkpoint (57), and also in the treatment with a number of genotoxins (75, 78, 79), indicating that regulation of Chk1 stability may be a part of the mechanisms that modulate activation or quenching of S phase checkpoint signaling. In AND-1-depleted cells, the replication stress tends to increase the instability of Chk1 and attenuate its phosphorylation (Fig. 4E), which is consistent with higher RDS in the later stage (Fig. 4, C and D). Moreover, AND-1 is phosphorylated in response to replication arrest in an ATR- and Cdc7-dependent manner and may play unknown roles in checkpoint activation. We have recently identified a couple of phosphorylated segments in the N-terminal region of AND-1 protein in vivo, although its functional significance remains to be clarified (data not shown).
Several studies in yeasts have revealed genetic interactions between DNA replication and chromatid cohesion factors (80, 81) and have suggested alternative functions of some the replisome components in sister chromatid cohesion. Among them, CTF18, a paralog of RFC1, travels along the chromosome as a replisome component (29), and its depletion leads to severe defects in cohesion and chromosome segregation both in the mitotic and meiotic cell cycle (33, 41, 82, 83). In this study, we found that human AND-1 physically interacts with cohesin components Smc1, Smc3, and Rad21/Scc1 in the nuclear extracts even in the presence of high salt. The interactions were not affected by DNA damage, replication stress, nor the phosphorylation status of Smc1. Thus, these interactions may be involved in the establishment of sister chromatid cohesion that occurs in a replication-dependent manner during the normal S phase. Other replisome components are also known to bind to cohesin. Human CTF18 binds not only to RFC subcomponents but also to Smc1 and Scc1 (84). In early biochemical studies of the bovine cohesin complex, DNA polymerase ϵ was co-purified (85). Nematode Tim1 (homolog of the human replisome component Tim) binds to cohesin proteins and facilitates establishment of the meiotic chromatid cohesion (86). AND-1 and the cohesin complex may be a part of a putative macromolecular complex at the replication fork and may help to coordinate execution of DNA replication and sister chromatid cohesion.
Furthermore, we provide evidence that human AND-1 is also required for homologous recombinational repair. In neomycin-resistant recombination reporter assays, the frequency of recombination events was significantly impaired by AND-1 depletion (Fig. 5C) to the extent similar to that shown by depletion or chemical inhibition of Chk1, HCLK2, or Rad18 (54, 57, 87). In yeasts, Mcl1 was also implicated in DNA repair (39, 88), and CTF4 was recently reported to be involved in homologous recombination-based repair through intra- and inter-chromatid exchange (89).
A number of studies have indicated that yeast cohesin proteins are required for DNA damage repair in S and G2 phase, possibly due to cohesin-dependent tethering of recombination proteins, which facilitates homologous recombination-mediated repair (44–47, 77). However, it should be noted that depletion of cohesins leads to enhancement, but not to inhibition, of recombination repair induced in similar assays, presumably because intra-chromatid recombination and/or unequal sister chromatid recombination has been accelerated by abrogation of cohesin-dependent equal sister chromatid recombination pathway (90). In contrast, AND-1 seems to be required not only for equal sister chromatid recombination, which exclusively depends on cohesin proteins, but also for intra- or unequal sister chromatid recombination. AND-1 may facilitate the loading of repair machinery at the damage sites in a manner independent of cohesins.
Given the multiple functions played by AND-1, it is tempting to speculate that AND-1 may function as a structural scaffold to which replication factors, checkpoint mediators, cohesins, as well as recombination factors bind for coordinated transactions of replication, checkpoint signaling, sister chromatid cohesion, and damage repair. Formation of the multiprotein complex on AND-1 may not only facilitate efficient execution of these events but also stabilize the entire protein machinery.
Supplementary Material
Acknowledgments
We thank Chika Taniyama for conducting the experiment of Fig. 1A; Naoko Kakusho for excellent technical assistance; and Dr. Shunichi Takeda and Dr. Thomas Helleday for SW480sn3 cells and pCMV3nls-I-SceI. We thank all the members of our laboratory for helpful discussions.
This work was supported by grants-in-aid for scientific research “A” and scientific research on priority areas “Chromosome Cycle” (to H. M.), by scientific research “C” (to N. Y.) from the Ministry of Education, Science, Sports and Culture, and by Astellas Foundation for Research on Metabolic Disorders (to H. M.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental “Results” and Figs. S1–S5.
- RPA
- replication protein A
- ATR
- ataxia-telangiectasia-mutated (ATM) and Rad3-related
- HU
- hydroxyurea
- siRNA
- small interfering RNA
- TopBP1
- topoisomerase II-binding protein 1
- PCNA
- proliferating cell nuclear antigen
- PBS
- phosphate-buffered saline
- DAPI
- 4,6-diamidino-2-phenylindole
- BrdUrd
- bromodeoxyuridine
- RDS
- radioresistant synthesis
- PIPES
- 1,4-piperazinediethanesulfonic acid
- TUNEL
- terminal deoxynucleotidyltransferase-mediated dUTP nick-end labeling.
REFERENCES
- 1.Zou L., Stillman B. (1998) Science 280, 593–596 [DOI] [PubMed] [Google Scholar]
- 2.Mimura S., Takisawa H. (1998) EMBO J. 17, 5699–5707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Masai H., Taniyama C., Ogino K., Matsui E., Kakusho N., Matsumoto S., Kim J. M., Ishii A., Tanaka T., Kobayashi T., Tamai K., Ohtani K., Arai K. (2006) J. Biol. Chem. 281, 39249–39261 [DOI] [PubMed] [Google Scholar]
- 4.Takayama Y., Kamimura Y., Okawa M., Muramatsu S., Sugino A., Araki H. (2003) Genes Dev. 17, 1153–1165 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Tanaka S., Umemori T., Hirai K., Muramatsu S., Kamimura Y., Araki H. (2007) Nature 445, 328–332 [DOI] [PubMed] [Google Scholar]
- 6.Yabuuchi H., Yamada Y., Uchida T., Sunathvanichkul T., Nakagawa T., Masukata H. (2006) EMBO J. 25, 4663–4674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gambus A., Jones R. C., Sanchez-Diaz A., Kanemaki M., van Deursen F., Edmondson R. D., Labib K. (2006) Nat. Cell Biol. 8, 358–366 [DOI] [PubMed] [Google Scholar]
- 8.Katou Y., Kanoh Y., Bando M., Noguchi H., Tanaka H., Ashikari T., Sugimoto K., Shirahige K. (2003) Nature 424, 1078–1083 [DOI] [PubMed] [Google Scholar]
- 9.Ricke R. M., Bielinsky A. K. (2004) Mol. Cell 16, 173–185 [DOI] [PubMed] [Google Scholar]
- 10.Pacek M., Tutter A. V., Kubota Y., Takisawa H., Walter J. C. (2006) Mol. Cell 21, 581–587 [DOI] [PubMed] [Google Scholar]
- 11.Nyberg K. A., Michelson R. J., Putnam C. W., Weinert T. A. (2002) Annu. Rev. Genet. 36, 617–656 [DOI] [PubMed] [Google Scholar]
- 12.Osborn A. J., Elledge S. J., Zou L. (2002) Trends Cell Biol. 12, 509–516 [DOI] [PubMed] [Google Scholar]
- 13.Sancar A., Lindsey-Boltz L. A., Unsal-Kaçmaz K., Linn S. (2004) Annu. Rev. Biochem. 73, 39–85 [DOI] [PubMed] [Google Scholar]
- 14.Bartek J., Lukas C., Lukas J. (2004) Nat. Rev. Mol. Cell Biol. 5, 792–804 [DOI] [PubMed] [Google Scholar]
- 15.Zou L., Elledge S. J. (2003) Science 300, 1542–1548 [DOI] [PubMed] [Google Scholar]
- 16.Zou L., Liu D., Elledge S. J. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 13827–13832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chini C. C., Chen J. (2003) J. Biol. Chem. 278, 30057–30062 [DOI] [PubMed] [Google Scholar]
- 18.Unsal-Kaçmaz K., Mullen T. E., Kaufmann W. K., Sancar A. (2005) Mol. Cell. Biol. 25, 3109–3116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Errico A., Costanzo V., Hunt T. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 14929–14934 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Unsal-Kaçmaz K., Chastain P. D., Qu P. P., Minoo P., Cordeiro-Stone M., Sancar A., Kaufmann W. K. (2007) Mol. Cell. Biol. 27, 3131–3142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gotter A. L., Suppa C., Emanuel B. S. (2007) J. Mol. Biol. 366, 36–52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chou D. M., Elledge S. J. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 18143–18147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yoshizawa-Sugata N., Masai H. (2007) J. Biol. Chem. 282, 2729–2740 [DOI] [PubMed] [Google Scholar]
- 24.Tsao C. C., Geisen C., Abraham R. T. (2004) EMBO J. 23, 4660–4669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cortez D., Glick G., Elledge S. J. (2004) Proc. Natl. Acad. Sci. U.S.A. 101, 10078–10083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kim J. M., Kakusho N., Yamada M., Kanoh Y., Takemoto N., Masai H. (2008) Oncogene 27, 3475–3482 [DOI] [PubMed] [Google Scholar]
- 27.Nasmyth K. (2001) Annu. Rev. Genet 35, 673–745 [DOI] [PubMed] [Google Scholar]
- 28.Haering C. H., Löwe J., Hochwagen A., Nasmyth K. (2002) Mol. Cell 9, 773–788 [DOI] [PubMed] [Google Scholar]
- 29.Lengronne A., McIntyre J., Katou Y., Kanoh Y., Hopfner K. P., Shirahige K., Uhlmann F. (2006) Mol. Cell 23, 787–799 [DOI] [PubMed] [Google Scholar]
- 30.Uhlmann F., Nasmyth K. (1998) Curr. Biol. 8, 1095–1101 [DOI] [PubMed] [Google Scholar]
- 31.Xu H., Boone C., Klein H. L. (2004) Mol. Cell. Biol. 24, 7082–7090 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Skibbens R. V., Corson L. B., Koshland D., Hieter P. (1999) Genes Dev. 13, 307–319 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hanna J. S., Kroll E. S., Lundblad V., Spencer F. A. (2001) Mol. Cell. Biol. 21, 3144–3158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Takeda T., Ogino K., Tatebayashi K., Ikeda H., Arai Ki, Masai H. (2001) Mol. Biol. Cell 12, 1257–1274 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bailis J. M., Bernard P., Antonelli R., Allshire R. C., Forsburg S. L. (2003) Nat. Cell Biol. 5, 1111–1116 [DOI] [PubMed] [Google Scholar]
- 36.Formosa T., Nittis T. (1999) Genetics 151, 1459–1470 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhou Y., Wang T. S. (2004) Mol. Cell. Biol. 24, 9568–9579 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wittmeyer J., Formosa T. (1997) Mol. Cell. Biol. 17, 4178–4190 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tsutsui Y., Morishita T., Natsume T., Yamashita K., Iwasaki H., Yamao F., Shinagawa H. (2005) Curr. Genet. 48, 34–43 [DOI] [PubMed] [Google Scholar]
- 40.Williams D. R., McIntosh J. R. (2002) Eukaryot. Cell 1, 758–773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Petronczki M., Chwalla B., Siomos M. F., Yokobayashi S., Helmhart W., Deutschbauer A. M., Davis R. W., Watanabe Y., Nasmyth K. (2004) J. Cell Sci. 117, 3547–3559 [DOI] [PubMed] [Google Scholar]
- 42.Tong A. H., Lesage G., Bader G. D., Ding H., Xu H., Xin X., Young J., Berriz G. F., Brost R. L., Chang M., Chen Y., Cheng X., Chua G., Friesen H., Goldberg D. S., Haynes J., Humphries C., He G., Hussein S., Ke L., Krogan N., Li Z., Levinson J. N., Lu H., Ménard P., Munyana C., Parsons A. B., Ryan O., Tonikian R., Roberts T., Sdicu A. M., Shapiro J., Sheikh B., Suter B., Wong S. L., Zhang L. V., Zhu H., Burd C. G., Munro S., Sander C., Rine J., Greenblatt J., Peter M., Bretscher A., Bell G., Roth F. P., Brown G. W., Andrews B., Bussey H., Boone C. (2004) Science 303, 808–813 [DOI] [PubMed] [Google Scholar]
- 43.Zhu W., Ukomadu C., Jha S., Senga T., Dhar S. K., Wohlschlegel J. A., Nutt L. K., Kornbluth S., Dutta A. (2007) Genes Dev. 21, 2288–2299 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sjögren C., Nasmyth K. (2001) Curr. Biol. 11, 991–995 [DOI] [PubMed] [Google Scholar]
- 45.Cortés-Ledesma F., Aguilera A. (2006) EMBO Rep. 7, 919–926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Schär P., Fäsi M., Jessberger R. (2004) Nucleic Acids Res. 32, 3921–3929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Ström L., Sjögren C. (2007) Curr. Opin. Cell Biol. 19, 344–349 [DOI] [PubMed] [Google Scholar]
- 48.Yoshizawa-Sugata N., Ishii A., Taniyama C., Matsui E., Arai K., Masai H. (2005) J. Biol. Chem. 280, 13062–13070 [DOI] [PubMed] [Google Scholar]
- 49.Montagnoli A., Tenca P., Sola F., Carpani D., Brotherton D., Albanese C., Santocanale C. (2004) Cancer Res. 64, 7110–7116 [DOI] [PubMed] [Google Scholar]
- 50.Sugata N., Li S., Earnshaw W. C., Yen T. J., Yoda K., Masumoto H., Munekata E., Warburton P. E., Todokoro K. (2000) Hum. Mol. Genet. 9, 2919–2926 [DOI] [PubMed] [Google Scholar]
- 51.Kim J. M., Nakao K., Nakamura K., Saito I., Katsuki M., Arai K., Masai H. (2002) EMBO J. 21, 2168–2179 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Heffernan T. P., Simpson D. A., Frank A. R., Heinloth A. N., Paules R. S., Cordeiro-Stone M., Kaufmann W. K. (2002) Mol. Cell. Biol. 22, 8552–8561 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Saleh-Gohari N., Bryant H. E., Schultz N., Parker K. M., Cassel T. N., Helleday T. (2005) Mol. Cell. Biol. 25, 7158–7169 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Saberi A., Hochegger H., Szuts D., Lan L., Yasui A., Sale J. E., Taniguchi Y., Murakawa Y., Zeng W., Yokomori K., Helleday T., Teraoka H., Arakawa H., Buerstedde J. M., Takeda S. (2007) Mol. Cell. Biol. 27, 2562–2571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Mohindra A., Hays L. E., Phillips E. N., Preston B. D., Helleday T., Meuth M. (2002) Hum. Mol. Genet. 11, 2189–2200 [DOI] [PubMed] [Google Scholar]
- 56.Johnson R. D., Jasin M. (2001) Biochem. Soc. Trans. 29, 196–201 [DOI] [PubMed] [Google Scholar]
- 57.Collis S. J., Barber L. J., Clark A. J., Martin J. S., Ward J. D., Boulton S. J. (2007) Nat. Cell Biol. 9, 391–401 [DOI] [PubMed] [Google Scholar]
- 58.Köhler A., Schmidt-Zachmann M. S., Franke W. W. (1997) J. Cell Sci. 110, 1051–1062 [DOI] [PubMed] [Google Scholar]
- 59.Kim S. T., Xu B., Kastan M. B. (2002) Genes Dev. 16, 560–570 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Yazdi P. T., Wang Y., Zhao S., Patel N., Lee E. Y., Qin J. (2002) Genes Dev. 16, 571–582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Kitagawa R., Bakkenist C. J., McKinnon P. J., Kastan M. B. (2004) Genes Dev. 18, 1423–1438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Desjardins L. M., MacManus J. P. (1995) Exp. Cell Res. 216, 380–387 [DOI] [PubMed] [Google Scholar]
- 63.Smits V. A., Reaper P. M., Jackson S. P. (2006) Curr. Biol. 16, 150–159 [DOI] [PubMed] [Google Scholar]
- 64.Li J., Stern D. F. (2005) J. Biol. Chem. 280, 37948–37956 [DOI] [PubMed] [Google Scholar]
- 65.Niida H., Katsuno Y., Banerjee B., Hande M. P., Nakanishi M. (2007) Mol. Cell. Biol. 27, 2572–2581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Robison J. G., Elliott J., Dixon K., Oakley G. G. (2004) J. Biol. Chem. 279, 34802–34810 [DOI] [PubMed] [Google Scholar]
- 67.Abraham R. T. (2001) Genes Dev. 15, 2177–2196 [DOI] [PubMed] [Google Scholar]
- 68.Burma S., Chen B. P., Murphy M., Kurimasa A., Chen D. J. (2001) J. Biol. Chem. 276, 42462–42467 [DOI] [PubMed] [Google Scholar]
- 69.Kumagai A., Lee J., Yoo H. Y., Dunphy W. G. (2006) Cell 124, 943–955 [DOI] [PubMed] [Google Scholar]
- 70.Mäkiniemi M., Hillukkala T., Tuusa J., Reini K., Vaara M., Huang D., Pospiech H., Majuri I., Westerling T., Mäkelä T. P., Syväoja J. E. (2001) J. Biol. Chem. 276, 30399–30406 [DOI] [PubMed] [Google Scholar]
- 71.Sørensen C. S., Syljuåsen R. G., Falck J., Schroeder T., Rönnstrand L., Khanna K. K., Zhou B. B., Bartek J., Lukas J. (2003) Cancer Cell 3, 247–258 [DOI] [PubMed] [Google Scholar]
- 72.Liu Q., Guntuku S., Cui X. S., Matsuoka S., Cortez D., Tamai K., Luo G., Carattini-Rivera S., DeMayo F., Bradley A., Donehower L. A., Elledge S. J. (2000) Genes Dev. 14, 1448–1459 [PMC free article] [PubMed] [Google Scholar]
- 73.Zhao H., Piwnica-Worms H. (2001) Mol. Cell. Biol. 21, 4129–4139 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Gatei M., Sloper K., Sorensen C., Syljuäsen R., Falck J., Hobson K., Savage K., Lukas J., Zhou B. B., Bartek J., Khanna K. K. (2003) J. Biol. Chem. 278, 14806–14811 [DOI] [PubMed] [Google Scholar]
- 75.Matsuura K., Wakasugi M., Yamashita K., Matsunaga T. (2008) J. Biol. Chem. 283, 25485–25491 [DOI] [PubMed] [Google Scholar]
- 76.Mayer M. L., Pot I., Chang M., Xu H., Aneliunas V., Kwok T., Newitt R., Aebersold R., Boone C., Brown G. W., Hieter P. (2004) Mol. Biol. Cell 15, 1736–1745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Sonoda E., Matsusaka T., Morrison C., Vagnarelli P., Hoshi O., Ushiki T., Nojima K., Fukagawa T., Waizenegger I. C., Peters J. M., Earnshaw W. C., Takeda S. (2001) Dev. Cell 1, 759–770 [DOI] [PubMed] [Google Scholar]
- 78.Zhang Y. W., Otterness D. M., Chiang G. G., Xie W., Liu Y. C., Mercurio F., Abraham R. T. (2005) Mol. Cell 19, 607–618 [DOI] [PubMed] [Google Scholar]
- 79.Nomura M., Nomura N., Yamashita J. (2005) Biochem. Biophys. Res. Commun. 335, 900–905 [DOI] [PubMed] [Google Scholar]
- 80.Warren C. D., Eckley D. M., Lee M. S., Hanna J. S., Hughes A., Peyser B., Jie C., Irizarry R., Spencer F. A. (2004) Mol. Biol. Cell 15, 1724–1735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Pan X., Ye P., Yuan D. S., Wang X., Bader J. S., Boeke J. D. (2006) Cell 124, 1069–1081 [DOI] [PubMed] [Google Scholar]
- 82.Naiki T., Kondo T., Nakada D., Matsumoto K., Sugimoto K. (2001) Mol. Cell. Biol. 21, 5838–5845 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Mayer M. L., Gygi S. P., Aebersold R., Hieter P. (2001) Mol. Cell 7, 959–970 [DOI] [PubMed] [Google Scholar]
- 84.Bermudez V. P., Maniwa Y., Tappin I., Ozato K., Yokomori K., Hurwitz J. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 10237–10242 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Jessberger R., Riwar B., Baechtold H., Akhmedov A. T. (1996) EMBO J. 15, 4061–4068 [PMC free article] [PubMed] [Google Scholar]
- 86.Chan R. C., Chan A., Jeon M., Wu T. F., Pasqualone D., Rougvie A. E., Meyer B. J. (2003) Nature 423, 1002–1009 [DOI] [PubMed] [Google Scholar]
- 87.Sørensen C. S., Hansen L. T., Dziegielewski J., Syljuåsen R. G., Lundin C., Bartek J., Helleday T. (2005) Nat. Cell Biol. 7, 195–201 [DOI] [PubMed] [Google Scholar]
- 88.Bennett C. B., Lewis L. K., Karthikeyan G., Lobachev K. S., Jin Y. H., Sterling J. F., Snipe J. R., Resnick M. A. (2001) Nat. Genet. 29, 426–434 [DOI] [PubMed] [Google Scholar]
- 89.Ogiwara H., Ui A., Lai M. S., Enomoto T., Seki M. (2007) Biochem. Biophys. Res. Commun. 354, 222–226 [DOI] [PubMed] [Google Scholar]
- 90.Potts P. R., Porteus M. H., Yu H. (2006) EMBO J. 25, 3377–3388 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.