Abstract
Background
Hybridization events are relatively common in vascular plants. However, the frequency of these events is unevenly distributed across the plant phylogeny. Plant families in which individual species are pollinated by specific pollinator species are predicted to be less prone to hybridization than other families. However, exceptions may occur within these families, when pollinators shift host-plant species. Indeed, host shifts are expected to increase the rate of hybridization events. Pollinators of Ficus section Galoglychia are suspected to have changed host repeatedly, based on several cases of incongruence between plant phylogeny and taxonomy, and insect phylogeny and taxonomy. We tracked cyto-nuclear discordance across section Galoglychia as evidence for hybridization. To achieve a proper global view, we first clarified the monophyly of section Galoglychia as it had been questioned by recent phylogenetic studies. Moreover, we investigated if fig size could be a factor facilitating host shifts.
Results
Phylogenetic chloroplast and nuclear results demonstrated the monophyly of section Galoglychia. Within section Galoglychia, we detected several cases of statistically significant cyto-nuclear discordance. Discordances concern both terminal nodes of the phylogenetic trees and one deep node defining relationships between subsections. Because nuclear phylogeny is congruent with morphological taxonomy, discordances were caused by the chloroplast phylogeny. Introgressive hybridization was the most likely explanation for these discordances. We also detected that subsections pollinated by several wasp genera had smaller figs and were pollinated by smaller wasps than subsections pollinated by a single wasp genus.
Conclusion
As hypothesized, we discovered evidences of past hybridization in Ficus section Galoglychia. Further, introgression was only detected in subsections presenting incongruence between plant and pollinator phylogenies and taxonomy. This supports the hypothesis that host shift is the cause for plant-pollinator incongruence. Moreover, small fig size could facilitate host shifts. Eventually, this study demonstrates that non-coding chloroplast markers are valuable to resolve deep nodes in Ficus phylogeny.
Background
Frequent natural hybridization events in vascular plants were documented by early biologists (e.g. [1,2]). However, the prevalence of hybridization events is unevenly distributed across the plant phylogeny and seems to be concentrated within a small fraction of families and genera [3]. Grant [4] pointed out that the frequency of natural hybridizations varies with factors such as life history, breeding system, environmental disturbance, genetic predisposition, and eventually pollination syndrome. In this context, plant species pollinated by specific pollinator species are predicted to present almost no hybridization events. Indeed, in specific mutualisms as plant-pollinator associations, the associated species usually present co-adaptations involved in attraction, recognition and physical compatibility with the other species. Such adaptations can be viewed as pre-zygotic barriers limiting interspecific hybridization [5-7].
Host shift is one of the main ecological processes that can break patterns of strict cospeciation between tightly bound interacting species [8-11], as observed in some cases affecting postulated strict co-evolutionary pattern in specific pollination mutualisms [12]. When associated with a transitional use of two hosts by the pollinator, a host shift may allow plant hybridization and hence leave a signature of genetic introgression in the plant genome. Following such hybridization, recurrent backcrosses of offspring with one parent may also allow the introgression of a limited set of foreign alleles into a plant species [13]. Hence, plant lineages in which pollinators have jumped from one host to another are more likely to exhibit signs of introgression than other lineages.
Detecting past events of host shifts requires tracking incongruence between phylogenetic trees of interacting taxa [8,14,15]. However, pollinator species duplication in the absence of host speciation, followed by cospeciation and subsequent asymmetric extinction, a phenomenon we will call duplication/extinction hereafter, is another mechanism that can cause incongruence between host phylogeny and partner phylogeny [16]. Although different in essence, these phenomena lead to similar patterns of phylogenetic incongruence, and discriminating between them is often largely speculative [17]. Nevertheless, and contrary to host shift, duplication/extinction does not involve plant hybridization. Consequently, detecting introgressed genes in a plant lineage for which there is incongruence between plant and pollinator phylogenies would support the host shift hypothesis.
Fig trees (Ficus, Moraceae) and their pollinating wasps (Agaonidae, Chalcidoidae) constitute one of the well-known cases of species-specific pollination mutualism. In this system, the rule is that a wasp species generally pollinates a single host species [18,19]. Further Ficus and Agaonid wasp morphological taxonomy and molecular phylogenies are steadily improving, providing a unique set of information to infer past history of the association on a broad set of species. Based on phylogenetic data, several authors have suggested parallel diversification of host and associated pollinating wasps [5,20-23]. Moreover, fig tree sections or subsections appear to be generally monophyletic and pollinated by a single wasp genus or by a few genera. Nevertheless, exceptions to the rule of a single agaonid wasp species associated with a single Ficus species are quite frequent and rather well documented based on morphology [24,25] as well as on molecular methods [12,26-28]. Indeed, Rasplus et al. [29] estimated for Africa, on the basis of morphology, that 17% of fig species were pollinated by more than one wasp species and that 15% of fig pollinator wasps used two or more hosts. However, molecular data on Australian pollinators suggest that when two or more wasps pollinate the same host they seem to often be sister species [30].
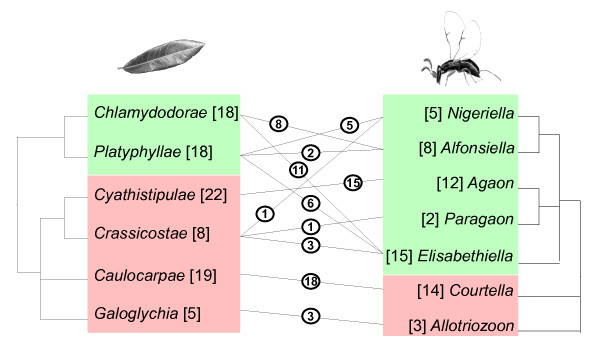
Within this general context, species of section Galoglychia show an unusual pattern of association with their pollinating wasps within genus Ficus. The 77 described species of section Galoglychia are distributed into six subsections [18,31], several of which are unambiguously defined. Seven morphologically unambiguous genera of fig wasps are known to pollinate section Galoglychia (Figure 1). Some genera are both subsection specific and the sole pollinators of the subsection. Other genera pollinate fig trees from different subsections and some subsections are pollinated by several wasp genera. Further some individual Ficus species may be pollinated by wasps belonging to different genera (e.g. Ficus natalensis pollinated by Alfonsiella and Elisabethiella) and one wasp species may pollinate several Ficus species (e.g. Elisabethiella stuckenbergii pollinates F. burkei, F. natalensis, F. petersii and F. lingua depauperata [32]).
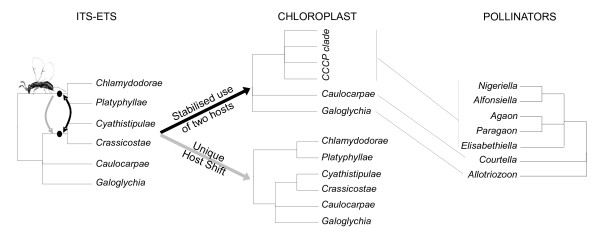
Figure 1.
Association between subsections of section Galoglychia in Ficus (left) and their genera of Agaonidae pollinating wasps (right). Numbers of taxa described for each wasp genus and each fig subsection are given within brackets. Linking lines inform on pollination associations between wasp genera and host subsections, with, for each link, the number of fig species known to be pollinated by a given wasp genus provided within circles. Colours highlight major clades to stress the discrepancy between plant (from [33]) and wasp (from [26]) phylogenies.
Recent molecular phylogenies supported, with some caveats, the monophyly of subsections within section Galoglychia [33] and that of wasp genera [26]. Hence the interactions between figs of section Galoglychia and their pollinating wasps present us with a series of instances of lack of strict specificity, and lack of congruence between fine wasp and Ficus taxonomy and phylogeny [18,34]. Comparing the phylogenies of trees and insects suggests additional lack of congruence, with respect to the order of branching of Ficus subsections and wasp genera (Figure 1). In the plant phylogeny [33], based on ITS and ETS nuclear markers, subsections Cyathistipulae and Crassicostae are grouped with subsections Galoglychia and Caulocarpae within a clade, while the remaining subsections are clustered together into a second clade. In the wasp pollinator phylogeny [26], based on 28S and ITS markers, subsection Galoglychia pollinators and Caulocarpae pollinators constitute the outgroups of a clade including the pollinators of the remaining subsections. Fig tree phylogeny is thus incongruent with pollinating wasp phylogeny with respect to the branching of subsections Cyathistipulae and Crassicostae versus Chlamydodorae and their pollinators. An additional finding of the molecular phylogeny of section Galoglychia is a suggestion that neotropical species of section Americana could be enclosed within section Galoglychia, although this result was only weakly supported by the various node validation methods [33]. If section Galoglychia is really paraphyletic to section Americana, pollinators of this last section could provide an example, pending phylogenetic and taxonomic re-evaluation, of discrepancy between the phylogenetic histories of Ficus and their pollinating wasps as the pollinators of section Americana and of section Galoglychia have been classified into different subfamilies (Agaoninae and Blastophaginae) [19].
Incongruence between pollinator and host phylogenies has been traditionally attributed to host shifts [12,35]. However, ancient pollinator species duplication events could also have lead to different wasp pollinator genera pollinating the same host figs. Subsequent cospeciation and asymmetric extinction could be responsible of the apparently haphazard host association exhibited by Alfonsiella and Elisabethiella pollinators [26,36].
The goal of this contribution is to study introgression events in the phylogeny of section Galoglychia. If host shifts and not duplications/extinctions are effectively responsible for plant-pollinator taxonomic and phylogenetic incongruence, then we expect to detect introgression events, especially in the plant lineages implied in this incongruence. We compared nuclear and chloroplast phylogenies to detect cases of genetic introgression between Ficus species, because the chloroplast genome of vascular plants possesses several features that facilitate its introgression into a new plant species after hybridization compared to the nuclear genome [13,37,38], as illustrated by the numerous cases of chloroplast introgression documented in the literature [37,39,40]. Our working hypothesis is that the current complex pattern of host association of genera Elisabethiella, Nigeriella and Alfonsiella result from host shifts. The prediction is that we may detect evidence for cytoplasm transfer between species of subsection Chlamydodorae, Platyphyllae and Crassicostae. We will also search for other potential cases of genetic introgression and especially for subsection Cyathistipulae for which available data suggests mismatch between the phylogenetic position of the subsection [33] and the phylogenetic position of its pollinators, namely genus Agaon [26]. To do so, we first developed specific chloroplast markers for Ficus and sequenced them in 58 individuals representing 38 species of section Galoglychia and 10 species of section Americana. We also sequenced ITS and ETS markers on these same individuals. Second, we searched for cases of discordance between chloroplast and nuclear phylogenies and tested their significance. Third, we showed that genetic introgression was the most likely mechanism to explain the cases of discordance. Besides, because section Galoglychia is the sole Ficus section subdivided into six subsections all of which are well defined (pending revision of the taxonomic position of some species assigned to subsection Platyphyllae, see discussion) and the sole section pollinated by six monophyletic agaonid wasp genera, this section offers a unique opportunity to investigate host shifts and their determinants. Within the section, known and suspected examples of pollinators using several hosts seem to involve host species presenting rather small figs, such as F. burkei, F. natalensis, F. petersii and F. lingua depauperata [32]. Therefore we investigated whether the fig subsections involved effectively presented small figs and whether their pollinator genera presented small body size.
Results
Data set partitioning
Chloroplast marker development allowed producing fourteen pairs of primer specifically developed for Ficus (see Additional file 1). Five of them - atpB-rbcL, FcB, FcJ, FcL and trnL-trnF - were selected for this analysis based on the phylogenetic information provided by a subsample of the final data set used in this study. The final combined chloroplast matrix contained 58 accessions and 3,604 base pairs (see Additional file 2; TreeBASE accession number: SN4278). One hundred and twenty-two sites were variable and 52 (1.5%) were potentially parsimony informative. The model of sequence evolution varied across chloroplast partitions (Table 1). We used arithmetic mean -lnL of Bayesian posterior topologies to measure the ability of data partitioning to explain the entire data set. The analysis separating non-coding and coding regions (strategy S3) of the chloroplast data set returned a decisively better description of the data than the other two analyses according to Bayes factors (Table 2).
Table 1.
Data partitions, genome to which they are associated, total number of characters of each partition used in phylogenetic analyses and their estimated model of sequence evolution.
| Partition | Genome | Number of characters | Selected Model |
| P1: All chloroplast data combined | Chloropast | 3604 | GTR+I+Γ |
| P2: atpB-rbcL | Chloropast | 820 | HKY+Γ |
| P3: FcB | Chloropast | 779 | GTR |
| P4: FcJ | Chloropast | 760 | HKY |
| P5: FcL | Chloropast | 777 | GTR+Γ |
| P6: trnL-trnF | Chloropast | 468 | HKY |
| P7: Coding chloroplast DNA | Chloropast | 964 | HKY |
| P8: Non-coding chloroplast DNA | Chloropast | 2640 | GTR+Γ |
| P9: All nuclear data combined | Nuclear | 1320 | GTR+Γ |
| P10: ITS | Nuclear | 810 | GTR+Γ |
| P11: ETS | Nuclear | 510 | HKY+Γ |
Table 2.
Arithmetic mean -LnL for each partition strategy (Si) and 2ln Bayes factors results of comparisons between the strategies with the highest likelihood and alternative strategies.
| Partition strategy | Mean -lnL | 2ln Bayes factor |
| S1: P1 | 6158.25 | S1 vs S3: 214.3 |
| S2: P2+P3+P4+P5+P6 | 6030.59 | S2 vs S3: 11.38 |
| S3: P7+P8 | 6028.45 | |
| S4: P9 | 5584.63 | S4 vs S5: 21.06 |
| S5: P10+P11 | 5578.51 |
The combined ITS and ETS matrix contained 1,320 base pairs of which 348 were variable and 158 (12%) were potentially parsimony informative. The model of sequence evolution differed between the ITS and ETS partitions (Table 1). The strategy individualizing each of these partitions (S5) was decisively better than the strategy combining the entire nuclear data set (S4) (Table 2). In the following we will only present the results obtained using the best strategy for each data set.
Chloroplast phylogeny of section Galoglychia
The combined chloroplast matrix produced twelve most parsimonious trees of length 141. Consistency and Retention Indexes (CI = 0.88; RI = 0.93) suggested the presence of a single island of trees [41]. The strict consensus of all most parsimonious trees (SCMP) is shown in Figure 2A with bootstrap percentages (MP_BS). The two different runs during the BI analysis with partitioning strategy S3 reached model parameter convergence and generated a similar tree. This tree is shown in Figure 3 with the posterior probabilities (PP) indicated for each node. ML analysis produced one most likely tree (-lnL = 5960.04; not shown), slightly less well resolved than the Bayesian tree. Bootstrap percentages (ML_BS) are indicated over the Bayesian tree in Figure 3.
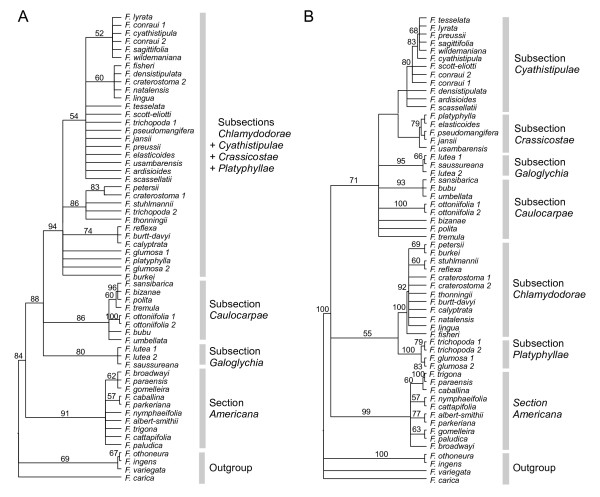
Figure 2.
Strict consensus trees of all most parsimonious trees. Analyses performed with (A) the chloroplast data set (length = 141 steps) and (B) the nuclear ITS+ETS data set (length = 521 steps). Values above branches indicate bootstrap supports.
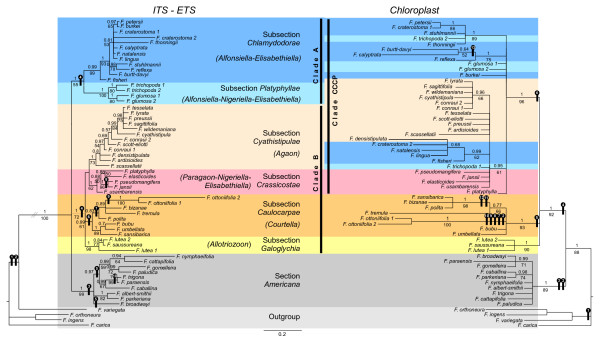
Figure 3.
ITS+ETS (left) and chloroplast (right) phylogenetic trees obtained with the Bayesian Inference (BI) analyses. Values above and under branches indicate posterior probability of BI and bootstrap support performed with the Maximum Likelihood analysis, respectively. Synapomorphic indels of each data set are mapped and numbered onto the corresponding tree (details about indels are given in Additional file 3). Genera of wasps known to pollinate a subsection are indicated between brackets.
The SCMP tree, the ML and the Bayesian trees were all similar in their overall topology. The monophyly of section Americana was strongly supported (91;89;1 for %MP_BS, %ML_BS and PP respectively; node supports are given in a similar order in the following) and formed a sister clade to section Galoglychia which was monophyletic (88;92;1). Subsections Galoglychia and Caulocarpae constituted monophyletic groups (80;90;1 and 86;93;1, respectively) and were basal within section Galoglychia. A third well supported clade was composed by species belonging to subsections Cyathistipulae, Crassicostae, Chlamydodorae and Platyphyllae (94;96;1; CCCP clade hereafter). However and contrary to the previous subsections, clade subsections did not form monophyletic groups within the CCCP. For example, the two accessions of F. trichopoda branched within distinct clades, as did the two accessions of F. craterostoma, and F. densistipulata (subsection Cyathistipulae) branched within a clade only composed of taxa of subsection Chlamydodorae.
One hundred and four distinct gaps were coded for the chloroplast data set. Five insertions and eight deletions were synapomorphic (Figure 3; see Additional file 3). A 10 bp insertion (chloroplast indel n°4) supported the monophyly of section Galoglychia since it was found neither in the outgroup nor in section Americana. A 7 bp deletion (n°6) supported the CCCP clade.
Nuclear phylogeny of section Galoglychia
The combined nuclear matrix produced 15,000 most parsimony informative trees of length 521 (RI = 0.84; CI = 0.75). The SCMP of all 15,000 trees is shown in Figure 2B. Figure 3 illustrates the unique Bayesian tree produced by the two different runs during the analysis and using the S5 partitioning strategy. Only one most likely tree (-lnL = 5150.78; not shown) was produced by the ML analysis. As with the chloroplast data, it differed from the Bayesian tree only in being slightly less resolved.
Here again, the SCMP, the ML and the Bayesian trees were all similar in their overall topology. A basal trichotomy divided the ingroup into three clades. The first one was the strongly supported section Americana (91;99;1). The second grouped subsection Chlamydodorae and part of subsection Platyphyllae (Clade A). Although it was not strongly supported by bootstraps (55;55), this clade was found in the 15,000 most parsimonious trees, and had a Bayesian posterior probability of one. The third clade grouped accessions from subsections Caulocarpae, Galoglychia, Cyathistipulae, Crassicostae and part of Platyphyllae (Clade B). It was not strongly supported by the bootstrap distribution (71;72) but it was present in every most parsimonious trees and is maximally supported by the BI analysis.
Sixty five indels were coded for the nuclear data set and 10 of them were synapomorphic (Figure 3; see Additional file 3). The ingroup is supported by one deletion and one insertion. Inside the ingroup, all accessions of section Americana presented a 3 bp deletion in their ETS sequence while all accessions of section Galoglychia shared a one base pair insertion in the ITS sequence.
Investigation of the cyto-nuclear discordance
While different phylogenetic reconstruction methods gave rise to similar topologies for a given data set, chloroplast and nuclear phylogenies appeared discordant for several internal and external nodes. Templeton tests applied to MP trees, and SH tests to ML trees always returned congruent results (Table 3). Both SCMP and ML trees explained significantly better the data set used to reconstruct these trees than the rival data set, demonstrating the overall discordance between nuclear and chloroplast topologies. We then tested the significance of local discordances involving F. trichopoda, F. craterostoma and F. densistipulata accessions. The original chloroplast topology in which both accessions of F. trichopoda are paraphyletic explained significantly better the chloroplast data than a modified topology in which these accessions were branched to form a monophyletic relationship. The same was true with the F. craterostoma chloroplast accessions. However, the original chloroplast topology was not significantly better than the topology in which F. densistipulata was branched with other Cyathistipulae taxa to make this subsection monophyletic. Alternatively, modifying the nuclear topology to branch one of the two F. trichopoda and one of the two F. craterostoma accessions with Chlamydodorae and with Cyathistipulae and Crassicostae accessions respectively, as in the chloroplast topology, produced significantly worse topologies than the unmodified nuclear topologies for the nuclear dataset. The same was true when F. densistipulata was branched with the Chlamydodorae species as in the chloroplast topology. For these last tests, it was possible to artificially reconstruct several alternative topologies. Table 3 gives results only for possibilities giving the highest P-value.
Table 3.
Results of Templeton tests with most parsimonious (MP) trees and SH tests with maximum likelihood (ML) trees.
| Templeton test | SH test | |||
| Topology | Length | P-Value | -LnL | P-Value |
| Chloroplast Data | ||||
| Chloroplast | 143 | 5956.50 | ||
| Nuclear | 195 | < 0.001 | 6241.39 | < 0.001 |
| chloroplast - F. trichopoda monophyletic | 147 | 0.045 | 5981.30 | 0.021 |
| chloroplast - F. craterostoma monophyletic | 152 | 0.003 | 5999.10 | 0.001 |
| chloroplast - F. densistipulata within Cyathistipulae | 144 | 0.564 | 5961.60 | 0.280 |
| (Americana, (Galoglychia, Caulocarpae,(Cyathistipulae + Crassicostae + Chlamydodorae + Platyphyllae)) | 226 | 6061.80 | ||
| (Americana, (Galoglychia + Caulocarpae + Cyathistipulae + Crassicostae), (Chlamydodorae + Platyphyllae)) | 276 | 0.016 | 6340.26 | 0.005 |
| Nuclear Data | ||||
| Nuclear | 539 | 5170.68 | ||
| Chloroplast | 741 | < 0.001 | 6091.60 | < 0.001 |
| nuclear - F. trichopoda 2 within Chlamydodorae | 560 | < 0.001 | 5259.70 | 0.000 |
| nuclear - F. craterostoma 2 within Cyathistipulae and Crassicostae | 560 | < 0.001 | 5211.23 | 0.001 |
| nuclear - F. densistipulata within Chlamydodorae | 559 | < 0.001 | 5256.40 | < 0.001 |
| (Americana, (Galoglychia + Caulocarpae + Cyathistipulae + Crassicostae), (Chlamydodorae + Platyphyllae)) | 818 | 5407.92 | ||
| (Americana, (Galoglychia, Caulocarpae, (Cyathistipulae + Crassicostae + Chlamydodorae + Platyphyllae)) | 962 | 0.001 | 5890.20 | 0.002 |
P-values of a given line correspond to the test comparing the length (MP trees) or the likelihood (ML trees) of the topology of the corresponding line with the length/likelihood of the above topology written in bold characters. In the topology description, "," indicates that branching between subsections are left unresolved, and "+" indicates that the branching among and between subsections are left unresolved. Likelihoods were computed with PAUP.
Besides the terminal cases of discordance tested above, a cyto-nuclear discordance was generated by the branching of subsections Cyathistipulae and Crassicostae. To avoid interference due to cases of terminal discordance, we tested the significance of this deep discordance by comparing topologies differing only by the branching of these two subsections (Table 3). Therefore, branching was left unresolved (i.e. with a polytomy) inside subsections Galoglychia and Caulocarpae for the chloroplast topology, and inside the CCCP clade, Clade A and Clade B (see Figure 3). Both Templeton and SH tests showed that the position of subsections Cyathistipulae and Crassicostae obtained in trees reconstructed using one data set (nuclear or cytoplasmic) produced a significantly better topology than the rival topology for that set of data.
The Bayesian approach used to statistically evaluate discordances gave similar results (not shown). For each Bayesian analysis, we did not find any branching supporting a rival phylogenetic hypothesis in any tree of the 95% credible set sampled once stationarity was reached. The sole exception was constituted by F. densistipulata in the chloroplast data set analysis, which branched with other Cyatistipulae species in 17% of the trees of the 95% credible set.
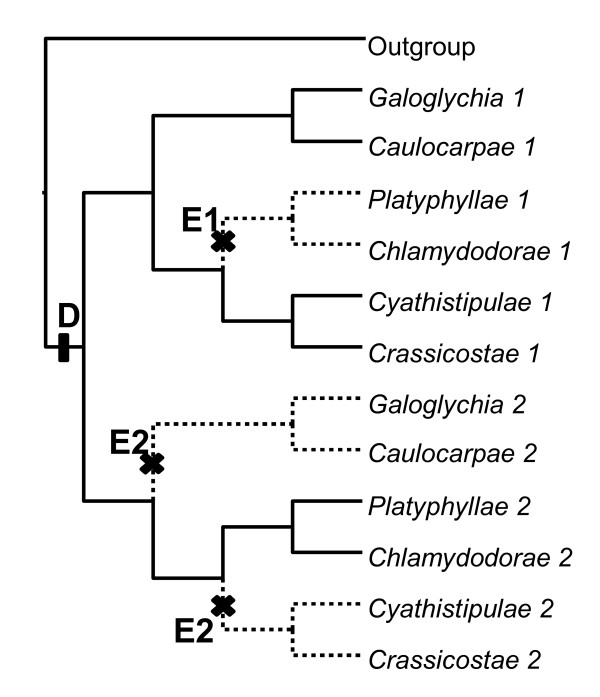
To check that apparent cyto-nuclear incongruence was not an artefact due to saturation by mutations, we plotted the estimated number of transitions and transversions against the TN93 [42] genetic distances (see Additional file 4). For all three analyses, transitions outnumber transversions indicating that substitutions are not saturated in the three types of data (chloroplast, ETS and ITS). Finally, considering ITS plus ETS as a single nuclear unit, a scenario based on gene duplication without genetic introgression and explaining the deep node discordance between nuclear and cytoplasmic phylogenies requires one duplication event at the base of section Galoglychia followed by three extinction events, each at the base of a group of subsections (Figure 4).
Figure 4.
The cyto-nuclear discordance explained by ITS and ETS duplication-extinction. The scenario requires a basal duplication (D) followed by three extinction events: one extinction of allele 1 (E1) and two extinctions of allele 2 (E2).
Fig and pollinator size
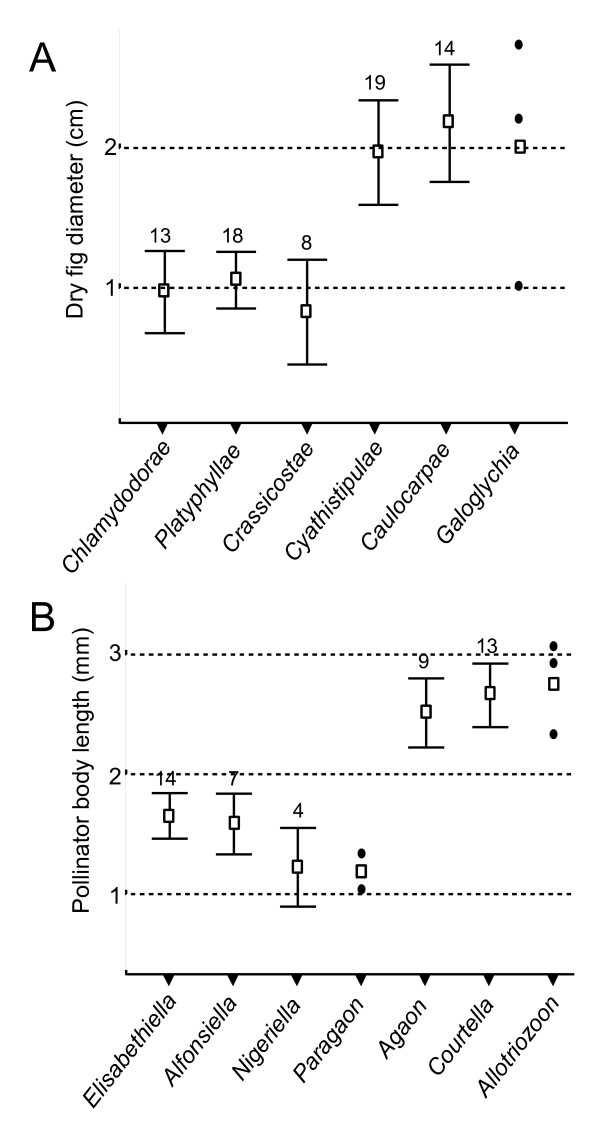
We investigated if fig and pollinator size were morphological traits associated at the subsection/genus level with plant-pollinator phylogenetic incongruence. Indeed, because of the as yet limited number of documented cases of incongruence, we limited our investigation to crude comparisons. Dry fig diameter of species belonging to section Galoglychia was distributed into two size categories (Figure 5A). Species belonging to subsections pollinated by a single pollinator genus (subsections Cyathistipulae, Caulocarpae and Galoglychia) presented significantly larger figs than species belonging to sections pollinated by at least two pollinator genera (t = 7.63; p < 10-3; not corrected for phylogenetic correlations). Similarly, body length of pollinating wasps was distributed into two size categories (Figure 5B): wasp genera that pollinated fig trees from a single subsection (i.e. Agaon, Paragaon, Courtella and Allotriozon) were significantly larger than genera pollinating several plant subsections (t = 7.43; p < 10-3; not corrected for phylogenetic correlations). Graphically, genus Paragaon presented small-sized wasps and therefore escapes this pattern. However, this genus includes only two species and for only one of them is the host known. Wasp size in this genus is thus little informative.
Figure 5.
Mean dry fig diameter and mean pollinator body length for each subsection of section Galoglychia and each pollinating wasp genus. Figures above whiskers give the number (n) of species included in the mean calculation. Whiskers give 95% credible interval (provided when n>3, otherwise single values are plotted).
Discussion
The monophyly of section Galoglychia
Rønsted et al. [33] did not demonstrate the monophyly of the African section Galoglychia. Based on ITS+ETS data, they suggested that section Galoglychia might be paraphyletic to the American section Americana, although this result was ambiguously supported. With our ITS+ETS data set, all three types of analysis (MP, ML and BI) also failed to resolve the placement of section Americana relative to section Galoglychia, although a 3 bp deletion in the ETS sequence characterises section Americana and a 1 bp insertion in the ITS sequence characterises section Galoglychia. However, the chloroplast phylogeny provides strong support for the monophyly of section Galoglychia with each of the three methods. Moreover, a 10 bp insertion and a 1 bp deletion in fragment FcL characterise species of section Americana and a 10 bp insertion in the same gene characterises species of section Galoglychia. Hence the phylogeny based on chloroplast data enables to resolve the trifurcation of the phylogeny based on ITS+ETS data and the result is further supported by insertion-deletion data which were not taken into account in the phylogenetic reconstructions. Eventually, our chloroplast data set provided less information on terminal nodes, but offered better resolution for deeper nodes. Because chloroplast genes overall evolve more slowly than nuclear genes do [43], they are less prone to homoplasy and are therefore usually recommended for deep phylogenies [44-46].
Morphologically, sections Galoglychia and Americana share the presence of two bracts subtending the figs as opposed to generally three in genus Ficus. Section Galoglychia is characterised by all the bracts of the ostiole (the passage leading into the fig) turned inward, the orifice of the ostiole forming a bilabiate slit. In section Americana, the bracts of the ostiole are imbricate so that generally three bracts are visible as closing the ostiole. The monophyly of section Galoglychia, as recognized by botanists since the major revision of African figs ([47]; recognized as subgenus Bibracteatae at that time), is hence most probably correct.
The infrasectional classification of section Galoglychia
The nuclear phylogeny proposes the division of section Galoglychia into two clades that do not include the same subsections as the two clades evidenced by the chloroplast analyses. Although the two nuclear clades are weakly supported (as previously observed by Rønsted et al. [33], several features suggest that this phylogeny is reliable. First, the alternative topology suggested by the chloroplast data set was demonstrated to be significantly less likely than the nuclear topology. Second, although 35% of our taxa were different from those used by Rønsted et al., we found a similar topology with similar values of bootstrap supports. It is thus unlikely that the sampling composition (which covers half of the species of section Galoglychia) biased the phylogenetic reconstruction. Third, the position of most species remains consistent with classical taxonomy [18]. A pattern seems to appear in the taxonomic modifications revealed by molecular phylogenies in Ficus, suggesting that they do not result from identification/manipulation errors or from limiting phylogenetic information. Indeed, within subsection Platyphyllae, F. platyphylla and F. jansii both present clusters of small figs at the axils of the leaves or just below, a trait they share with a number of species of subsection Crassicostae, with which they are grouped in the phylogenetic tree. Conversely, F. glumosa and F. stuhlmannii only present two figs at the axils of the leaves and are grouped with species from subsection Chlamydodorae, which all have figs in pairs at the axils of the leaves. Preliminary data on some other species of subsection Platyphyllae seem to support this splitting of the species into two different units, mainly according to this trait. Fourth, a potential source of error using ITS and ETS could be the amplification of paralogous copies of ITS and ETS leading to an incorrect phylogenetic reconstruction [48-50]. However, here again, the monophyly of each subsection is hardly compatible with the amplification of paralogous copies of ITS and ETS that generally evolve independently [51]. Four events of extinction following the duplication would have to be hypothesised to obtain an observed nuclear topology corresponding to the chloroplast topology (Figure 4). Although this number of duplication-extinctions can reasonably occur given the considered time lapse [39], these four events would have had to occur in the ancestor of each of the four extant subsections and not within the subsections, a pattern which is unlikely. Last, the separation of section Galoglychia into two nuclear clades is supported by ecological and biogeographical arguments [33]. All these lines of evidence lead us to conclude that subsections Galoglychia, Cyathistipulae, Crassicostae, Caulocarpae and part of Platyphyllae on one hand, and subsection Chlamydodorae and part of Platyphyllae on the other hand, form two monophyletic clusters.
A discordant chloroplast topology
Our investigation of the overall discordance between the chloroplast and the nuclear topology revealed two types of discordances occurring at two taxonomic levels. The first type concerns phylogenetic relationships between subsections: subsections Cyathistipulae and Crassicostae branch with subsections Chlamydodorae and Platyphyllae in the chloroplast topology, and with subsections Galoglychia and Caulocarpae in the nuclear topology. As highlighted above, the nuclear topology is consistent with morphological and biogeographical data. We can thus confidently assume that it is the chloroplast and not the nuclear topology that is discordant with the species tree. The second type of discordance concerns the monophyly of subsections: Chlamydodorae and Platyphyllae are paraphyletic in the chloroplast phylogeny (at least for F. craterostoma and F. trichopoda), and monophyletic in the nuclear phylogeny. The monophyly of subsections found in the nuclear tree is consistent with the morphological traits that led botanists to recognize these subsections for a century [47], meaning that here again, the nuclear but not the chloroplast phylogeny is more likely to reflect the species tree.
Examples of cyto-nuclear discordance are rather common in plants [52-57] and a limited number of causes leading to such discordances has been identified. Discordance between cytoplasmic and nuclear phylogenetic trees may either be artefactual or evidence distinct evolutionary histories followed by cytoplasmic and nuclear genomes [39]. Long Branch Attraction (LBA hereafter; [58]) is a source of artefact that may generate cyto-nuclear discordance [59,60]. Several arguments allow us to exclude LBA artefacts for our data set. First, the branches leading to discordance are not the longest branches of the tree, while some longer branches do not have unexpected positions in the tree. Further, the molecular data sets are not saturated which means that our markers did not evolve rapidly enough for phylogenetic signals to be lost. Finally, the discordance was the same whatever the phylogenetic reconstruction method employed. Although methodological concordance does not rule out LBA [61], parsimony has a stronger bias towards grouping long branches together than ML and BI which take into account unequal rates or branch lengths [62,63]. It is therefore unlikely that the observed cases of discordance resulted from LBA.
The cyto-nuclear discordance has therefore to be explained in terms of difference between species and chloroplast history. Two mechanisms may be involved: incomplete chloroplast lineage sorting and introgression. Differentiating between hypotheses of introgression and lineage sorting is difficult because both processes can generate very similar phylogenetic patterns [64]. Ancestral polymorphism is expected to be completely sorted out with a high probability approximately 4Ne generations (Ne = effective population size) after the separation [65]. The branch supporting the CCCP clade (BL = 0.1774) is longer than the branch supporting clades in which reciprocal monophyly has been reached, like the Caulocarpae clade (BL = 0.1699), the Galoglychia clade (BL = 0.1269) and section Americana (BL = 0.1756). Hence, this branch is expected to be long enough to have allowed chloroplast genes to reach reciprocal monophyly as well and hence to complete allele sorting. However, the sorting depends on Ne and not on branch length directly. Therefore, any variation in Ne over time and across species would alter conclusions based on relative branch length comparison. Methods based on coalescent simulations to calculate the probability of observing lineage sorting are also based on Ne remaining constant over time and among species [66] and both hypotheses are not realistic when comparing savannah and forest lineages over longer periods of time. Moreover, they require estimating ancestral population sizes that can only be obtained by intensive intraspecific sampling and this is not compatible with the phylogenetic approach used here. Eventually, these methods require generation time to be known and do not allow this parameter to vary across the phylogeny. Ficus lineages included in our phylogeny are highly different in term of height and life forms, with some lineages represented mainly by canopy emergent trees (e.g. subsection Caulocarpae) and other mainly by small shrubs and trees in open habitats (e.g., subsection Chlamydodorae). Because the generation time is known to vary with life form and height in trees [67], substantial variation in the generation time across the Galoglychia phylogeny would generate strong uncertainty for results based on coalescent methods. In conclusion, because of too many uncertainties in the inputs required for more subtle models, we assess that the method based on relative branch lengths, although not optimal, is appropriate to discuss lineage sorting with the current data set. Although we are not able to fully exclude it, the incomplete lineage sorting hypothesis thus does not appear a likely explanation for the observed cyto-nuclear discordance.
Host shift mediated introgression as the cause of phylogenetic discordance
Introgressive hybridization, i.e. the introduction of an allele inside the genome of a foreign taxon after hybridization [13], could explain the observed pattern. Introgressive hybridization in Ficus necessitates that a pollinating wasp fertilizes two hosts. Hybridization may result from sporadic visitation of an alternative host by a wasp, and may in that case not require successful development of wasp offspring. However, it may also result from transient or even stabilised utilisation of two hosts by a pollinating wasp species, and could result in subsequent speciation of the wasps on the new host. Thus, pollinator taxonomy and phylogeny may be informative to analyse whether introgressive hybridization could account for discrepancies between markers in the Ficus phylogeny.
Subsections Cyathistipulae and Crassicostae present discordant positions between the chloroplast and nuclear phylogenies. Subsection Cyathistipulae is strictly associated with genus Agaon. The phylogenetic analysis concludes to nuclear monophyly and does not reject cytoplasmic monophyly. On the other hand subsection Crassicostae is pollinated by what could be its own genus of pollinators, Paragaon (two species known, and host association only known for one species) but also more frequently by Elisabethiella and in one case by Nigeriella, i.e. genera also associated with subsections Chlamidodorae and Platyphyllae. Hence this part of the cyto-nuclear mismatch could be associated with repeated pollinator host shifts. Erasmus et al. [26] published a phylogeny of wasps pollinating Ficus species of section Galoglychia that may shed light on the origin of genera Agaon (the exclusive pollinators of subsection Cyathistipulae) and Paragaon (pollinating some species of Crassicostae). Based on a combined ITS/28S data set, they found the following topology: (((Agaon &Paragaon), (Alfonsiella &Nigeriella), Elisabethiella), Courtella, Allotriozoon). Hence, the cytoplasmic CCCP clade is exclusively pollinated by the Agaon, Paragaon, Alfonsiella, Nigeriella and Elisabethiella clade. One may therefore suggest that a first host shift of a pollinator from an ancestor of the Chlamydodorae to an ancestor of the Cyathistipulae and Crassicostae may have given rise to the Agaon-Paragaon lineage (detailed in Figure 6). This would be the first and major event bringing about the cyto-nuclear discordance. Then we need to hypothesize a second wasp transfer between these two large clades, from Chlamydodorae to Crassicostae. In this scenario, the pollinators bring with them the cytoplasm of their original host. A simple way to achieve such an a priori unexpected cytoplasm transfer is the stabilised occurrence of a species of fig pollinating wasps using two host species. Such a situation allows backcrosses of hybrids with both parental species. Indeed field observations report several cases of two Ficus species being pollinated by the same wasp species [26]. For instance Elisabethiella stuckenbergii is known to pollinate F. burkei, F. natalensis natalensis, F. lingua depauperata and F. petersii [32], a feature which could potentially allow cytoplasm transfer in any direction independently of which of these Ficus species, if any, was the original host of the wasp.
Figure 6.
Comparative phylogeny and the host shift scenario. If we accept current phylogenies, then a pollinating wasp of an ancestor of Chlamydodorae and Platyphyllae must have shifted to an ancestor of Cyathistipulae and Crassicostae, an event required to explain the discrepancy between ITS+ETS (left) and pollinator (right) phylogenies. However, under the hypothesis of a wasp suddenly changing host (grey arrows), there is no cytoplasm transfer and the Ficus lineage resulting from the hybridization necessary bears a chloroplast related to the Caulocarpae and Galoglychia lineages (middle bottom), a feature which is not observed in the cytoplasm phylogeny. On the contrary, if a wasp uses two hosts for a prolonged period of time (black arrows), repeated backcrosses may allow the introduction of a chloroplast originating in the Chlamydodorae plus Platyphyllae lineage inside the ancestor of Cyathistipulae and Crassicostae (middle top), as observed in the data set. The set of phylogenetic trees therefore support the role of an intermediate stage in which a pollinating wasp has consistently used two host species.
The scenario presented above hinges on the relative phylogenetic position of the different genera pollinating section Galoglychia. A recently published pollinator phylogeny [68] proposed that pollinators of subsections Chlamydodorae, Platyphyllae, Cyathistipulae and Crassicostae are monophyletic, as in the phylogeny of Erasmus et al. [26], and thus does not contradict the scenario proposed above. This new phylogeny did not include Allotriozoon and Paragaon and did not fully resolve the position of the different genera.
The second type of discordance, concerning the monophyly of subsections, suggests that introgressive hybridization has occurred inside the CCCP clade, with at least two evidenced instances. Both cases concerned species from subsections that are pollinated by several wasp genera. All wasp phylogenies demonstrated the monophyly of wasp genera [26,68]. Taken together, these observations support the hypothesis that hosts shift could account for several wasp genera pollinating a single Ficus subsection. Erasmus et al. [26] and Jousselin et al. [36] recently suggested that original pollinator species duplication followed by more recent extinction of one or other of the lineages in host figs, rather than host shifts, could explain this unusual association pattern. Beyond evidence for cytoplasmic gene transfers in our phylogenies, there are two reasons that do not favour this hypothesis. First we would need to apply this line of reasoning to subsection Crassicostae which is pollinated by three wasp genera (Paragaon, Elisabethiella, Nigeriella). As Elisabethiella and Nigeriella also pollinate subsection Chlamydodorae, these genera should then have differentiated into well defined genera very early in section Galoglychia history and only much later would one genus have become extinct in some species or even subspecies and survived in other species or even subspecies. The second reason is that Erasmus et al. [26] provided direct evidence of ongoing local pollinator exchange between Ficus species. Indeed Alfonsiella binghami samples from Nelspruit (South Africa), formed a single genetic group whether collected on F. petersii or on F. sthulmannii comparatively to samples of A. binghami also collected on F. sthulmannii but in Lekgalameetse (South Africa) and in Amani (Tanzania). Finally, recovering a phylogenetic signal suggesting some amount of parallel cladogenesis is not sufficient to infer that parallel cladogenesis is relevant within these taxonomic groups. Indeed, if host shifts preferentially occur between closely related hosts [69], then the limited amount of evidence suggesting host-pollinator co-cladogenesis in section Chlamydodorae [36] could easily result from host shifts. Hence the evidence in favour of the occurrence of host shifts within the pollinators of part of section Galoglychia is overwhelming.
Fig size: a factor facilitating host shifts?
Although we statistically demonstrated only three instances of cyto-nuclear discordance, it is remarkable that these signs of introgressive hybridization were only found in subsections that are implied in the incongruence between plant and insect phylogeny and taxonomy. If these results strongly supports the hypothesis that host shift is the cause of this incongruence, it also stresses the fact that host shifts have occurred preferentially in some particular lineages of section Galoglychia. Hence, host shift could have been facilitated by particular phenotypic traits in wasps or host plants rather than resulting from purely stochastic processes. When we compare the six subsections of section Galoglychia, a striking pattern emerges. All subsections that share pollinator genera and present cytoplasmic introgression also present, on average, small figs and are pollinated by small wasps. Conversely all subsections that have their own chloroplast lineage (or for which cytoplasm monophyly cannot be ruled out) and are pollinated by their own genus of agaonid wasps produce large figs and are pollinated by large wasps. Among pollinators of section Galoglychia, large wasp species seem therefore to be more host-specific than small wasp species, a pattern of association that could correlate with a simpler and shorter ostiole passages into small figs of section Galoglychia. Indeed, in Gabon, Michaloud [70] showed that Courtella sp. pollinating Ficus ottoniifolia (subsection Caulocarpae) looses its wings and antennae on entering the tight ostiole of the fig, while Alfonsiella fimbriata pollinating F. natalensis leprieuri (section Chlamydodorae) only looses its wings and retains its antennae, going through an ostiole that Michaloud describes as a simple slit.
Conclusion
As predicted by the hypothesis that host shifts are responsible for the lack of congruence between insect and plant phylogeny and taxonomy in section Galoglychia, we found evidence for several cases of chloroplast genetic introgression events. The alternative to host shifts would be duplication of insect taxa on a host followed by later extinction. Separating these two hypotheses is extremely difficult [69,71]. However, in our case, because two genetically independent markers, nuclear DNA carried by insects and chloroplasts, tell the same story, the hypothesis of host shifts becomes much more likely. Knowing which lineages are involved in host shifts within a Ficus section allowed us to infer that a facilitating factor for host shift could be small fig size. Detecting identical or analogous patterns to those described here in other sections of the genus will constitute the real test of their generality. Finally, while chloroplast DNA presents limited variability in Ficus [72], our initial screening of markers allowed increasing the number of polymorphic sites. We were eventually able to resolve deep nodes in the Ficus phylogeny for which ITS and ETS nuclear markers were not sufficiently informative. Non-coding chloroplast markers could thus be very helpful to resolve basal branching inside the Ficus phylogeny; a necessary step for an in depth analysis of the history of the coevolution between figs and their pollinating wasps.
Methods
Taxon sampling
Forty four specimens belonging to 38 species of Ficus out of 77 from the section Galoglychia were analysed in addition to 10 species from section Americana. Ficus carica, F. ingens, F. variegata and F. orthoneura were chosen as outgroups because they belong to distinct sections originating from Asia. This sampling encompasses all six subsections of section Galoglychia sensu Berg [31] and confirmed by Rønsted et al. [33]. For six species, two specimens from different locations were collected and used for validation. Most of the material consisted of dried leaves collected during field work mainly by FK, from herbaria and from living collections. Forty-one ITS and 33 ETS accessions were retrieved from GenBank (following previously published papers [5,22,33]). We chose to acquire our own set of nuclear sequences for species for which available sequences came from individuals of unknown geographic origin and belonging to species with large pan-african distribution ranges, or for species recognized to include several subspecies or morphotypes. This procedure allowed us to be confident that we were comparing chloroplast and nuclear genes of the same taxa. A list of taxa with origin and GenBank accession numbers is provided in Additional file 2.
Chloroplast DNA laboratory work
Although chloroplast markers were used in early Ficus phylogenetic studies [20], their low level of polymorphism and the associate difficulty to obtain resolved phylogenies down to genus level excluded them from subsequent studies [72]. We therefore developed our own chloroplast primer pairs. We focused on non-coding to select informative sequences. The detailed protocols used during molecular laboratory works are provided in Additional file 5. We designed primer pairs in Ficus using the Oligo4 software. The Staden Package was used to compile contiguous sequences of each accession and all polymorphic sites were checked against the original electrophoregrams. They were re-examined and adjusted manually using BioEdit [73]. To identify the position of Ficus sequences, they were blasted with the completely sequenced chloroplast genome, phylogenetically closest to Ficus, namely that of Morus indica [74]. Pairwise P-distance and average value with standard error estimated by 500 bootstraps were computed using the MEGA software version 3.1 [75].
Using universal primers developed by Grivet et al. [76] in the large single copy (LSC) region of chloroplast genome, we tested 25 amplifying DNA fragments smaller than 3 Kb on a sub-sample of 9 species (see Additional file 6) representing genus Ficus diversity with 6 sections and 3 species within section Galoglychia. We also tested two previously described sequences, atpB-rbcL [77] and trnL (UAA)3'exon-trnF(GAA) [78]. Based on the resulting sequences and their polymorphism, we selected 10 primer pairs and defined new primer pairs to amplify potentially informative sequences shorter than 1 kb (see Additional file 1). For the 3 fragments longer than 1 kb, we designed 2 to 3 primer pairs focusing on the most informative sequences with 6 to 15 substitutions. These 14 amplified sequences were distributed throughout the LSC chloroplast genome. We examined the amount of phylogenetic information conveyed by each marker on 33 Ficus species including 19 from sections Galoglychia and Americana. Finally, we eventually selected five markers on the basis of the phylogenetic information they conveyed: atpB-rbcL, FcB, FcL, FcJ and trnL-trnF.
ITS and ETS laboratory work
Based on 100 ITS sequences previously published [5,22,33], we designed new primer pairs specifically defined in Ficus using the Oligo4 software. The primers ITSF1 (ACAAGGTTTCCGTAGGTGA) and ITSF4 (GTATAGTTATTCGCCTCCT) were defined as forward and reverse primers in 18S and 26S ribosomal RNA genes, respectively. An additional reverse primer ITSF5 (CGGAGGTTACGCTGGGGTC) was defined in the zone at the junction between ITS2 and 26S conferring a high specificity in Ficus species. Indeed, ITS amplification using the ITSF1-ITSF5 primer pair was selective and successful for some samples producing a single, clearly defined, DNA fragment. For the remaining samples, we used the ITSF1-ITSF4 primer pair that amplified several bands since they were defined on conserved sequences. For samples which produced several amplified DNA fragments, we selected the one presenting the expected size based on previous published ITS sequences [22,72] by isolating it from the agarose gel following a long electrophoresis migration. We then performed a second PCR on this isolated DNA fragment for sequencing. ETS amplification was performed using the ETS-Hel-1/18S-ETS primer pair [79].
Alignment and phylogenetic analyses
Sequence alignment was performed using the program Clustal X [80] with manual adjustment and excluding ambiguous alignment positions (0.2% and 3.5% of the original chloroplast and nuclear data sets, respectively). All analyses with the chloroplast genes were performed with the five markers combined into a single matrix. This concatenation is not expected to modify the inference of the chloroplast history since this organelle is not known to recombine in wild Angiosperms [81]. For nuclear analyses, we followed previous studies that demonstrated that ITS and ETS sequences could be combined into a single matrix in Ficus [5,22]. In maximum parsimony (MP) analyses, all characters were equally weighted and treated as unordered. Heuristic searches using PAUP* 4b10 [82] were started with 1000 stepwise random addition sequence replicates, holding 10 trees at each step, followed by tree bisection-reconnection branch (TBR) swapping, saving maximally 100 most parsimonious trees. All shortest trees retained in memory were used as starting trees for a second round of searching using TBR branch swapping until all trees were found or a pre-set maximum of 15,000 trees were found. We eventually computed the strict consensus of the most parsimonious trees (SCMP). Relative levels of homoplasy in both nuclear and chloroplast data sets were assessed from all characters using the consistency index (CI) and the retention index (RI) as implemented in PAUP. Bootstrap support was assessed using 1000 replicates each consisting of 10 random addition sequence replicates with TBR swapping and no limits on the number of trees saved.
Bayesian inferences (BI) were performed with several partitioning strategies for each data matrix. Three strategies were tested with the chloroplast matrix: all chloroplast data combined (P1 in Table 1), one partition each for the five markers (P2-P6) and one partition each for the coding and non-coding DNA (P7-P8). Two strategies were tested with the nuclear matrix: all chloroplast data combined (P9) and one partition each for ITS and ETS (P10-P11). The appropriate model of sequence evolution for each partition was determined using the likelihood ratio test (LRT) implemented with MrModeltest [83].
We tested every partitioning strategy with BI with the program MrBayes 3.1.2 [84]. Model parameters were always taken as unlinked and the rate multipliers set as variable across partitions. Initial runs were conducted starting with random neighbour joining trees to check the number of simultaneous Markov Chain Monte Carlo (MCMC) chains necessary not to get caught on local optima. A Dirichlet distribution was assumed for the rate matrix and base frequency and every tree topology was assumed to be equally probable. The MCMC process was set so that two simultaneous independent analyses starting from different random trees with four chains (three heated) ran simultaneously over 4,000,000 generations, and every 200th tree saved into a file. Variation in likelihood scores was examined graphically to determine apparent stationarity for each independent run resulting in a burnin of 10% trees. MCMC convergence was also explored by examining the Potential Scale Reduction Factor (PSRF) convergence diagnostics for all parameters in the model. Finally, we determined posterior probabilities of the phylogenies and associated branches based on the stationary trees pooled from the two runs for each analysis.
The results for each partitioning strategy were compared to the strategy with the best arithmetic mean of the likelihoods (-lnL; sampled from the posterior) using Bayes factors (see [85] for details). Bayes factors were approximated by the ratio of the harmonic means of the likelihoods of the two strategies being tested [86]. Arithmetic and harmonic means were calculated using the sump command of MrBayes. In this study, we used the traditional cutoff criterion of 2ln Bayes factor of ≥ 10 as very strong evidence against the compared strategy [87].
Maximum likelihood (ML) analyses were performed with RaxML [88] which offers the possibility to partition data. For each data set, we therefore ran analyses using the best partitioning strategy and model of sequence evolution calculated with MrModeltest. We replicated 200 heuristic searches using a randomized maximum parsimony tree as starting tree. The confidence of branching was assessed using 1000 non-parametric bootstrap resamplings generated as heuristic searches. The information from the 1000 bootstrapped trees was drawn on the best-scoring ML tree from the 200 runs.
Gap characters representing mainly deletions were studied separately from the nucleotide matrices. Gaps were coded for each data set using SeqState [89] with the Simple Indel Coding approach (SIC) [90]. This method scores all gaps, regardless of length, as separate presence/absence characters. Sequences with gaps that extend beyond both the 5'- and 3'-termini of the gap being coded, as well as sequences with gaps that extend beyond one terminus and to the other terminus, are scored as missing data for that character. We used preferentially the SIC approach because it has been demonstrated to outperform the classical method consisting in coding gaps as 5th states for each and every nucleotide position regardless of length [91]. We eventually mapped gaps onto the topologies resulting from the analyses of the respective data sets. Because of limited space along branches, we mapped only synapomorphic gaps.
Analyses of cyto-nuclear discordances
Chloroplast and nuclear phylogenies appeared discordant for several internal and external nodes. The nested pattern of the discordances prevented us from testing their individual significance by using the classical method of removing all discordant taxa and then adding them individually in comparative tests (see [39] for an example). Instead, we used Templeton tests [92] with trees modified from the SCMPs, and Shimodaira-Hasegawa tests (SH; [93]) with trees modified from the ML trees. SH tests were implemented in PAUP, using the REEL approximation with 10,000 bootstrap replicates. Because PAUP cannot account for partitions, models of sequence evolution specified in SH tests were those calculated with all data combined for each data set. Basically, for two phylogenetic hypotheses to be discordant, a topology A must explain significantly better a data set A than a topology B does it, and a topology B must explain significantly better a data set B than a topology A does it. Both conditions need to be full-filled to be able to assess the significance of a given discordance. For example, the global discordance between nuclear and chloroplast topology was tested by comparing the length of the SCMP of the chloroplast trees with the length of the nuclear SCMP, based on the chloroplast data set. The test was then repeated based on the nuclear data set. Similarly, we tested several local discordances individually by comparing a SCMP topology with the same SCMP topology in which only one discordant accession was moved manually in the position as found in the rival topology; each time duplicating the test with both data sets. Last, we tested discordances with a Bayesian approach [85]. This was achieved by first building 95% credible sets of unique trees (sampled at stationarity) with the sumt command in MrBayes for each the chloroplast and nuclear analysis. Then, we used the SumTrees [94] software to check for the presence of alternative phylogenetic hypotheses in the 95% credible set. If they were absent, these hypotheses could be rejected statistically.
Because saturation in substitutions can lead to incorrect phylogenetic inferences [95], we plotted the estimated number of transitions and transversions against the genetic distance using DAMBE V 4.5.32 [96]. In an unsaturated data set, transitions and transversions are both expected to increase linearly with the genetic distance, with a steeper slope for transitions than for transversions. Analyses were conducted with chloroplast, ITS and ETS sequence matrixes separately.
We investigated the hypothesis of amplifying paralogous copies of nuclear genes by assessing the minimum number of gene duplications and extinctions that are required to generate the observed discordant pattern with the help of the GeneTree 1.3.0 software [97,98]. This procedure was only done for the nuclear genes because gene duplication has not been documented for plastid loci [99]. Because ITS and ETS belong to the same unit of transcription, we did not separate them for this analysis.
Fig and pollinator size
Data on fig and pollinator size were retrieved from Berg & Wiebes [18]. As the data were collected by the same observers, the values were comparable. Dry fig diameter was averaged for each subsection of section Galoglychia. We used diameter of dry figs because it was the sole fig-size measure available for all species. We also used body length for each pollinator wasp species. Body length included head, thorax and gaster but excluded the ovipositor. Simple mean comparison tests were performed to compare fig/wasp size between subsections/genera involved in strict association with their partners and subsections/genera involved in multiple associations.
Authors' contributions
JPR, FK and BK jointly designed and coordinated the study. FK carried out the sampling and BK, CG and SS the laboratory analyses. JPR analysed the sequence data sets and wrote the first draft of the manuscript. FK and BK finalized the manuscript. All authors read and approved the final manuscript.
Supplementary Material
Description of the new chloroplast primer pairs specifically defined in Ficus. This table provides basic information about primers like name, sequence and position in Morus indica genome.
List of samples included in the final analyses with origin and GenBank accession numbers. This table provides species name, origin, voucher number and GenBank accession number for samples.
Insertions and deletions in chloroplast and nuclear markers. This data provides information about position and length for indels.
Substitution pattern of the chloroplast markers, ITS and ETS genes. The number of transitions and transversions is plotted against the TN93 distance.
Protocols for molecular laboratory works. A text providing details for extraction, amplification and sequencing procedures.
List of species used for the development of non coding chloroplast DNA markers for Ficus. A table giving voucher numbers and origins.
Acknowledgments
Acknowledgements
We are indebted to Champlain Djieto-Lordon, of the Faculty of Science at Yaoundé for help during field collections in Cameroun. We also thank E. Renoult and M. Charpentier for constructive comments on earlier version of the manuscript, and N. Alvarez for technical advices. We also thank N. Rønsted who kindly provided nine samples from her studies. JPR was supported by a grant from the French Ministry of Research. The work was supported by the ANR grant NICEFig.
Contributor Information
Julien P Renoult, Email: julien.renoult@cefe.cnrs.fr.
Finn Kjellberg, Email: finn.kjellberg@cefe.cnrs.fr.
Cinderella Grout, Email: groutc@yahoo.fr.
Sylvain Santoni, Email: Sylvain.Santoni@supagro.inra.fr.
Bouchaïb Khadari, Email: khadari@supagro.inra.fr.
References
- Losty JP. Evolution by means of hybridization. Dordrecht: Nijhoff; 1916. [Google Scholar]
- Raven PH. Systematics and plant population biology. Syst Biol. 1976;1:284–316. [Google Scholar]
- Ellstrand NC, Whitkus R, Rieseberg LH. Distribution of spontaneous plant hybrids. Proc Natl Acad Sci USA. 1996;93:5090–5093. doi: 10.1073/pnas.93.10.5090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grant V. Plant speciation. 2. New-York: Columbia University Press; 1981. [Google Scholar]
- Jousselin E, Rasplus JY, Kjellberg F. Convergence and coevolution in a mutualism: Evidence from a molecular phylogeny of Ficus. Evolution. 2003;57:1255–1269. doi: 10.1554/02-445. [DOI] [PubMed] [Google Scholar]
- Levin RA, Raguso RA, McDade LA. Fragrance chemistry and pollinator affinities in Nyctaginaceae. Phytochemistry. 2001;58:429–440. doi: 10.1016/S0031-9422(01)00257-6. [DOI] [PubMed] [Google Scholar]
- Pellmyr O. Yuccas, yucca moths, and coevolution: A review. Ann Missouri Bot Gard. 2003;90:35–55. doi: 10.2307/3298524. [DOI] [Google Scholar]
- Charleston MA, Robertson DL. Preferential host switching by primate lentiviruses can account for phylogenetic similarity with the primate phylogeny. Syst Biol. 2002;51:528–535. doi: 10.1080/10635150290069940. [DOI] [PubMed] [Google Scholar]
- Jackson AP. A reconciliation analysis of host switching in plant-fungal symbioses. Evolution. 2004;58:1909–1923. doi: 10.1111/j.0014-3820.2004.tb00479.x. [DOI] [PubMed] [Google Scholar]
- Menken SBJ. Pattern and process in the evolution of insect-plant associations: Yponomeuta as an example. Entomol Exp Appl. 1996;80:297–305. doi: 10.1007/BF00194779. [DOI] [Google Scholar]
- Schoonhoven LM, Jermy T, van Loon JJA. Insect-plant biology. From physiology to evolution. London: Chapman and Hall; 1998. [Google Scholar]
- Machado CA, Robbins N, Gilbert MTP, Herre EA. Critical review of host specificity and its coevolutionary implications in the fig/fig-wasp mutualism. Proc Natl Acad Sci USA. 2005;102:6558–6565. doi: 10.1073/pnas.0501840102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallet J. Hybridization as an invasion of the genome. Trends Ecol Evol. 2005;20:229–237. doi: 10.1016/j.tree.2005.02.010. [DOI] [PubMed] [Google Scholar]
- Huyse T, Volckaert FAM. Comparing host and parasite phylogenies: Gyrodactylus flatworms jumping from goby to goby. Syst Biol. 2005;54:710–718. doi: 10.1080/10635150500221036. [DOI] [PubMed] [Google Scholar]
- Ricklefs RE, Fallon SM, Bermingham E. Evolutionary relationships, cospeciation, and host switching in avian malaria parasites. Syst Biol. 2004;53:111–119. doi: 10.1080/10635150490264987. [DOI] [PubMed] [Google Scholar]
- Reed DL, Light JE, Allen JM, Kirchman JJ. Pair of lice lost or parasites regained: the evolutionary history of anthropoid primate lice. BMC Biol. 2007;5:7. doi: 10.1186/1741-7007-5-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson JA. Analysis of parasite host-switching: limitations on the use of phylogenies. Parasitology. 1999;119:S111–S123. [PubMed] [Google Scholar]
- Berg CC, Wiebes JT. African fig trees and fig wasps. Amsterdam: North Holland; 1992. [Google Scholar]
- Wiebes JT. Agaonidae (Hymenoptera-Chalcidoidea) and Ficus (Moraceae) - Fig wasps and their figs .14. (Conclusion - Old-World) Proc Koninklijke Nederl Akad Wetens (C) 1994;97:491–495. [Google Scholar]
- Herre EA, Machado CA, Bermingham E, Nason JD, Windsor DM, McCafferty SS, VanHouten W, Bachmann K. Molecular phylogenies of figs and their pollinator wasps. J Biogeog. 1996;23:521–530. doi: 10.1111/j.1365-2699.1996.tb00014.x. [DOI] [Google Scholar]
- Machado CA, Jousselin E, Kjellberg F, Compton SG, Herre EA. Phylogenetic relationships, historical biogeography and character evolution of fig-pollinating wasps. Proc R Soc Lond Series-B Biol Sci. 2001;268:685–694. doi: 10.1098/rspb.2000.1418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rønsted N, Weiblen GD, Cook JM, Salamin N, Machado CA, Savolainen V. 60 million years of co-divergence in the fig-wasp symbiosis. Proc R Soc Lond Series-B Biol Sci. 2005;272:2593–2599. doi: 10.1098/rspb.2005.3249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiblen GD. Phylogenetic relationships of fig wasps pollinating functionally dioecious Ficus based on mitochondrial DNA sequences and morphology. Syst Biol. 2001;50:243–267. [PubMed] [Google Scholar]
- Michaloud G, Michaloudpelletier S, Wiebes JT, Berg CC. The co-occurrence of two pollinating species of fig wasp and one species of fig. Proc Koninklijke Nederl Akad Wetens (C) 1985;88:93–119. [Google Scholar]
- Ramírez WB. Host specificity of fig wasps (Agaonidae) Evolution. 1970;24:680–691. doi: 10.2307/2406549. [DOI] [PubMed] [Google Scholar]
- Erasmus JC, van Noort S, Jousselin E, Greeff JM. Molecular phylogeny of fig wasp pollinators (Agaonidae, Hymenoptera) of Ficus section Galoglychia. Zool Scripta. 2007;36:61–78. doi: 10.1111/j.1463-6409.2007.00259.x. [DOI] [Google Scholar]
- Lopez-Vaamonde C, Rasplus JY, Weiblen GD, Cook JM. Molecular phylogenies of fig wasps: Partial cocladogenesis of pollinators and parasites. Mol Phylogenet Evol. 2001;21:55–71. doi: 10.1006/mpev.2001.0993. [DOI] [PubMed] [Google Scholar]
- Molbo D, Machado CA, Sevenster JG, Keller L, Herre EA. Cryptic species of fig-pollinating wasps: Implications for the evolution of the fig-wasp mutualism, sex allocation, and precision of adaptation. Proc Natl Acad Sci USA. 2003;100:5867–5872. doi: 10.1073/pnas.0930903100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rasplus JY. The one-to-one species specificity of the Ficus-Agaoninae mutualism: How casual? In: van der Maesen LJG, van der Burgt XM, van Medenbach de Rooy JM, editor. The biodiversity of African plants. Wageningen: Kluwer Academic Publisher; 1996. [Google Scholar]
- Lopez-Vaamonde C, Dixon DJ, Cook JM, Rasplus JY. Revision of the Australian species of Pleistodontes (Hymenoptera: Agaonidae) fig-pollinating wasps and their host-plant associations. Zool J Linn Soc. 2002;136:637–683. doi: 10.1046/j.1096-3642.2002.00040.x. [DOI] [Google Scholar]
- Berg CC. Subdivision of Ficus subg Urostigma sect Galoglychia (Moraceae) Proc Koninklijke Nederl Akad Wetens (C) 1986;89:121–127. [Google Scholar]
- Underhill JG. Phylogeny and codivergence in the fig-fig wasp mutualism: Sycoecine and Agaondid fig wasps (Chalcidoidea, Hymenoptera) associated with Ficus section Galoglychia (Moraceae) Cape Town: Faculty of Science; 2008. [Google Scholar]
- Rønsted N, Salvo G, Savolainen V. Biogeographical and phylogenetic origins of African fig species (Ficus section Galoglychia) Mol Phylogenet Evol. 2007;43:190–201. doi: 10.1016/j.ympev.2006.12.010. [DOI] [PubMed] [Google Scholar]
- Compton SG, Vannoort S. Southern African fig wasps (Hymenoptera, Chalcidoidea) - Resource utilization and host relationships. Proc Koninklijke Nederl Akad Wetens (C) 1992;95:423–435. [Google Scholar]
- Cook JM, Rasplus JY. Mutualists with attitude: Coevolving fig wasps and figs. Trends Ecol Evol. 2003;18:241–248. doi: 10.1016/S0169-5347(03)00062-4. [DOI] [Google Scholar]
- Jousselin E, van Noort S, Berry V, Rasplus JY, Rønsted N, Erasmus JC, Greeff JM. One fig to bind them all: Host conservatism in a fig wasp community unraveled by cospeciation analyses among pollinating and nonpollinating fig wasps. Evolution. 2008;62:1777–1797. doi: 10.1111/j.1558-5646.2008.00406.x. [DOI] [PubMed] [Google Scholar]
- Liston A, Kadereit JW. Chloroplast DNA evidence for introgression and long-distance dispersal in the desert annual Senecio flavus (Asteraceae) Pl Syst Evol. 1995;197:33–41. doi: 10.1007/BF00984630. [DOI] [Google Scholar]
- Rieseberg LH, Soltis DE. Phylogenetic consequences of cytoplasmic gene flow in plants. Evol Trends Pl. 1991;5:65–84. [Google Scholar]
- Niet T van der, Linder HP. Dealing with incongruence in the quest for the species tree: A case study from the orchid genus Satyrium. Mol Phylogenet Evol. 2008;47:154–174. doi: 10.1016/j.ympev.2007.12.008. [DOI] [PubMed] [Google Scholar]
- Van Droogenbroeck B, Kyndt T, Romeijn-Peeters E, Van Thuyne W, Goetghebeur P, Romero-Motochi JP, Gheysen G. Evidence of natural hybridization and introgression between Vasconcellea species (Caricaceae) from southern Ecuador revealed by chloroplast, mitochondrial and nuclear DNA markers. Ann Bot. 2006;97:793–805. doi: 10.1093/aob/mcl038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maddison DR. The discovery and importance of multiple islands of most-parsimonious trees. Syst Zool. 1991;40:315–328. doi: 10.2307/2992325. [DOI] [Google Scholar]
- Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial-DNA in humans and chimpanzees. Mol Biol Evol. 1993;10:512–526. doi: 10.1093/oxfordjournals.molbev.a040023. [DOI] [PubMed] [Google Scholar]
- Wolfe KH, Li WH, Sharp PM. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA. 1987;84:9054–9058. doi: 10.1073/pnas.84.24.9054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clegg MT, Gaut BS, Learn GH, Morton BR. Rates and patterns of chloroplast DNA evolution. Proc Natl Acad Sci USA. 1994;91:6795–6801. doi: 10.1073/pnas.91.15.6795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hilu KW, Borsch T, Muller K, Soltis DE, Soltis PS, Savolainen V, Chase MW, Powell MP, Alice LA, Evans R, et al. Angiosperm phylogeny based on matK sequence information. Am J Bot. 2003;90:1758–1776. doi: 10.3732/ajb.90.12.1758. [DOI] [PubMed] [Google Scholar]
- Schuettpelz E, Korall P, Pryer KM. Plastid atpA data provide improved support for deep relationships among ferns. Taxon. 2006;55:897–906. [Google Scholar]
- Mildbraed J, Burret M. Die Afrikanischen Arten der Gattung Ficus. Linn Bot Jarhb. 1911;46:163–269. [Google Scholar]
- Buckler ES, Ippolito A, Holtsford TP. The evolution of ribosomal DNA: Divergent paralogues and phylogenetic implications. Genetics. 1997;145:821–832. doi: 10.1093/genetics/145.3.821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartmann S, Nason JD, Bhattacharya D. Extensive ribosomal DNA genic variation in the columnar cactus Lophocereus. J Mol Evol. 2001;53:124–134. doi: 10.1007/s002390010200. [DOI] [PubMed] [Google Scholar]
- Ko KS, Jung HS. Three nonorthologous ITS1 types are present in a polypore fungus Trichaptum abietinum. Mol Phylogenet Evol. 2002;23:112–122. doi: 10.1016/S1055-7903(02)00009-X. [DOI] [PubMed] [Google Scholar]
- Alvarez I, Wendel JF. Ribosomal ITS sequences and plant phylogenetic inference. Mol Phylogenet Evol. 2003;29:417–434. doi: 10.1016/S1055-7903(03)00208-2. [DOI] [PubMed] [Google Scholar]
- Dorado O, Rieseberg LH, Arias DM. Chloroplast DNA introgression in southern california Sunflowers. Evolution. 1992;46:566–572. doi: 10.2307/2409874. [DOI] [PubMed] [Google Scholar]
- Kellogg EA, Appels R, MasonGamer RJ. When genes tell different stories: The diploid genera of Triticeae (Gramineae) Syst Bot. 1996;21:321–347. doi: 10.2307/2419662. [DOI] [Google Scholar]
- Natcheva R, Cronberg N. Recombination and introgression of nuclear and chloroplast genomes between the peat mosses, Sphagnum capillifolium and Sphagnum quinquefarium. Mol Ecol. 2007;16:811–818. doi: 10.1111/j.1365-294X.2006.03163.x. [DOI] [PubMed] [Google Scholar]
- Okuyama Y, Fujii N, Wakabayashi M, Kawakita A, Ito M, Watanabe M, Murakami N, Kato M. Nonuniform concerted evolution and chloroplast capture: Heterogeneity of observed introgression patterns in three molecular data partition phylogenies of Asian Mitella (Saxifragaceae) Mol Biol Evol. 2005;22:285–296. doi: 10.1093/molbev/msi016. [DOI] [PubMed] [Google Scholar]
- Soltis DE, Kuzoff RK. Discordance between nuclear and chloroplast phylogenies in the Heuchera Group (Saxifragaceae) Evolution. 1995;49:727–742. doi: 10.2307/2410326. [DOI] [PubMed] [Google Scholar]
- Sutton BCS, Flanagan DJ, Gawley JR, Newton CH, Lester DT, Elkassaby YA. Inheritance of chloroplast and mitochondrial DNA in Picea and composition of hybrids from introgression zones. Theor Appl Genet. 1991;82:242–248. doi: 10.1007/BF00226220. [DOI] [PubMed] [Google Scholar]
- Felsenstein J. Cases in which parsimony or compatibility methods will be positively misleading. Syst Zool. 1978;27:401–410. doi: 10.2307/2412923. [DOI] [Google Scholar]
- Kennedy M, Holland BR, Gray RD, Spencer HG. Untangling long branches: Identifying conflicting phylogenetic signals using spectral analysis, neighbor-net, and consensus networks. Syst Biol. 2005;54:620–633. doi: 10.1080/106351591007462. [DOI] [PubMed] [Google Scholar]
- Sanderson MJ, Wojciechowski MF, Hu JM, Khan TS, Brady SG. Error, bias, and long-branch attraction in data for two chloroplast photosystem genes in seed plants. Mol Biol Evol. 2000;17:782–797. doi: 10.1093/oxfordjournals.molbev.a026357. [DOI] [PubMed] [Google Scholar]
- Bergsten J. A review of long-branch attraction. Cladistics. 2005;21:163–193. doi: 10.1111/j.1096-0031.2005.00059.x. [DOI] [PubMed] [Google Scholar]
- Pol D, Siddall ME. Biases in maximum likelihood and parsimony: A simulation approach to a 10-taxon case. Cladistics. 2001;17:266–281. doi: 10.1006/clad.2001.0172. [DOI] [PubMed] [Google Scholar]
- Swofford DL, Waddell PJ, Huelsenbeck JP, Foster PG, Lewis PO, Rogers JS. Bias in phylogenetic estimation and its relevance to the choice between parsimony and likelihood methods. Syst Biol. 2001;50:525–539. doi: 10.1080/106351501750435086. [DOI] [PubMed] [Google Scholar]
- Holder MT, Anderson JA, Holloway AK. Difficulties in detecting hybridization. Syst Biol. 2001;50:978–982. doi: 10.1080/106351501753462911. [DOI] [PubMed] [Google Scholar]
- Pamilo P, Nei M. Relationships between gene trees and species trees. Mol Biol Evol. 1988;5:568–583. doi: 10.1093/oxfordjournals.molbev.a040517. [DOI] [PubMed] [Google Scholar]
- Buckley TR, Cordeiro M, Marshall DC, Simon C. Differentiating between hypotheses of lineage sorting and introgression in New Zealand alpine cicadas (Maoricicada Dugdale) Syst Biol. 2006;55:411–425. doi: 10.1080/10635150600697283. [DOI] [PubMed] [Google Scholar]
- King DA. Allometry and life history of tropical trees. J Trop Ecol. 1996;12:25–44. doi: 10.1017/S0266467400009299. [DOI] [Google Scholar]
- Lopez-Vaamonde C, Wikström N, Kjer KM, Weiblen GD, Rasplus JY, Machado CA, Cook JM. Molecular dating and biogeography of fig-pollinating wasps. Mol Phylogenet Evol. 2009;52:715–726. doi: 10.1016/j.ympev.2009.05.028. [DOI] [PubMed] [Google Scholar]
- De Vienne DM, Giraud T, Shykoff JA. When can host shifts produce congruent host and parasite phylogenies? A simulation approach. J Evol Biol. 2007;20:1428–1438. doi: 10.1111/j.1420-9101.2007.01340.x. [DOI] [PubMed] [Google Scholar]
- Michaloud G. Figuiers tropicaux et pollinisation (Motion picture film) Distributed by Service du Film de Recherche Scientifique Paris, France. 1982.
- Light JE, Hafner MS. Codivergence in heteromyid rodents (Rodentia: Heteromyidae) and their sucking lice of the genus Fahrenholzia (Phthiraptera: Anoplura) Syst Biol. 2008;57:449–465. doi: 10.1080/10635150802169610. [DOI] [PubMed] [Google Scholar]
- Rønsted N, Yektaei-Karin E, Turk K, Clarkson JM, Chase MW. Species-level phylogenetics of large genera: prospects of studying co-evolution and polyploidy. In: Hodkinson T, Parnell J, Waldren S, editor. Reconstructing the tree of life: taxonomy and systematics of species rich taxa. Boca Raton, Florida: Systematic Association Series. CRC Press; 2007. [Google Scholar]
- Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Ac Symp Ser. 1999;41:95–98. [Google Scholar]
- Ravi V, Khurana JP, Tyagi AK, Khurana P. The chloroplast genome of mulberry: complete nucleotide sequence, gene organization and comparative analysis. Tree Genet Genom. 2007;3:49–59. doi: 10.1007/s11295-006-0051-3. [DOI] [Google Scholar]
- Kumar S, Tamura K, Nei M. MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinf. 2004;5:150–163. doi: 10.1093/bib/5.2.150. [DOI] [PubMed] [Google Scholar]
- Grivet D, Heinze B, Vendramin GG, Petit RJ. Genome walking with consensus primers: application to the large single copy region of chloroplast DNA. Mol Ecol Notes. 2001;1:345–349. doi: 10.1046/j.1471-8278.2001.00107.x. [DOI] [Google Scholar]
- Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH. Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA. 2005;102:8369–8374. doi: 10.1073/pnas.0503123102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taberlet P, Gielly L, Pautou G, Bouvet J. Universal primers for amplification of 3 noncoding regions of chloroplast DNA. Pl Mol Biol. 1991;17:1105–1109. doi: 10.1007/BF00037152. [DOI] [PubMed] [Google Scholar]
- Baldwin BG, Markos S. Phylogenetic utility of the external transcribed spacer (ETS) of 18S-26S rDNA: Congruence of ETS and ITS trees of Calycadenia (Compositae) Mol Phylogenet Evol. 1998;10:449–463. doi: 10.1006/mpev.1998.0545. [DOI] [PubMed] [Google Scholar]
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Ac Res. 1997;25:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trabelsi S, Gargouri-Bouzid R, Vedel F, Nato A, Lakhoua L, Drira N. Somatic hybrids between potato Solanum tuberosum and wild species Solanum vernei exhibit a recombination in the plastome. Plant Cell Tiss Org Cult. 2005;83:1–11. doi: 10.1007/s11240-005-3667-3. [DOI] [Google Scholar]
- Swofford DL. PAUP*: Phylogenetic Analysis Using Parsimony (* and other methods), Version 4.0b4. Sunderland, MA: Sinauer Associates; 2000. [Google Scholar]
- Nylander JAA. MrModeltest v2. Program distributed by the author. Uppsala University: Evolutionary Biology Centre; 2004. [Google Scholar]
- Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–755. doi: 10.1093/bioinformatics/17.8.754. [DOI] [PubMed] [Google Scholar]
- Brandley MC, Schmitz A, Reeder TW. Partitioned Bayesian analyses, partition choice, and the phylogenetic relationships of scincid lizards. Syst Biol. 2005;54:373–390. doi: 10.1080/10635150590946808. [DOI] [PubMed] [Google Scholar]
- Nylander JAA, Ronquist F, Huelsenbeck JP, Nieves-Aldrey JL. Bayesian phylogenetic analysis of combined data. Syst Biol. 2004;53:47–67. doi: 10.1080/10635150490264699. [DOI] [PubMed] [Google Scholar]
- Kass RE, Raftery AE. Bayes Factors. J Am Stat Assoc. 1995;90:773–795. doi: 10.2307/2291091. [DOI] [Google Scholar]
- Stamatakis A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 2006;22:2688–2690. doi: 10.1093/bioinformatics/btl446. [DOI] [PubMed] [Google Scholar]
- Müller K. SeqState - primer design and sequence statistics for phylogenetic DNA data sets. Appl Bioinf. 2005;4:65–69. doi: 10.2165/00822942-200504010-00008. [DOI] [PubMed] [Google Scholar]
- Simmons MP, Ochoterena H. Gaps as characters in sequence-based phylogenetic analyses. Syst Biol. 2000;49:369–381. doi: 10.1093/sysbio/49.2.369. [DOI] [PubMed] [Google Scholar]
- Simmons MP, Muller K, Norton AP. The relative performance of indel-coding methods in simulations. Mol Phylogenet Evol. 2007;44:724–740. doi: 10.1016/j.ympev.2007.04.001. [DOI] [PubMed] [Google Scholar]
- Templeton AR. Phylogenetic inference from restriction endonuclease cleavage site maps with particular reference to the evolution of humans and the apes. Evolution. 1983;37:221–244. doi: 10.2307/2408332. [DOI] [PubMed] [Google Scholar]
- Shimodaira H, Hasegawa M. Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol. 1999;16:1114–1116. [Google Scholar]
- SumTrees: Summarization of split support on phylogenetic trees. Version 1.0.2. Part of the DendroPy phylogenetic computation library version 2.1.3 http://packages.python.org/DendroPy/
- Swofford DL, Olsen G, Waddell PJ, Hillis DM. Phylogenetic inference in molecular systematics. Sunderland: Sinauer Associates Inc; 1996. pp. 407–514. [Google Scholar]
- Xia X, Xie Z. DAMBE: Software package for data analysis in molecular biology and evolution. J Hered. 2001;92:371–373. doi: 10.1093/jhered/92.4.371. [DOI] [PubMed] [Google Scholar]
- Page RDM. GeneTree: comparing gene and species phylogenies using reconciled trees. Bioinformatics. 1998;14:819–820. doi: 10.1093/bioinformatics/14.9.819. [DOI] [PubMed] [Google Scholar]
- Page RDM, Charleston MA. From gene to organismal phylogeny: Reconciled trees and the gene tree species tree problem. Mol Phylogenet Evol. 1997;7:231–240. doi: 10.1006/mpev.1996.0390. [DOI] [PubMed] [Google Scholar]
- Soltis DE, Soltis PS, Doyle JJ. Molecular systematics of plants II. Boston: Kluwer Academic Publishers; 1998. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of the new chloroplast primer pairs specifically defined in Ficus. This table provides basic information about primers like name, sequence and position in Morus indica genome.
List of samples included in the final analyses with origin and GenBank accession numbers. This table provides species name, origin, voucher number and GenBank accession number for samples.
Insertions and deletions in chloroplast and nuclear markers. This data provides information about position and length for indels.
Substitution pattern of the chloroplast markers, ITS and ETS genes. The number of transitions and transversions is plotted against the TN93 distance.
Protocols for molecular laboratory works. A text providing details for extraction, amplification and sequencing procedures.
List of species used for the development of non coding chloroplast DNA markers for Ficus. A table giving voucher numbers and origins.