Abstract
Arrhythmogenic cardiomyopathies are a heterogeneous group of pathological conditions that give rise to myocardial dysfunction with an increased risk for atrial or ventricular arrhythmias. Inherited defects in cardiomyocyte proteins in the sarcomeric contractile apparatus, the cytoskeleton and desmosomal cell-cell contact junctions are becoming recognized increasingly as major causes of sudden cardiac death in the general population. Animal models have been developed for the systematic dissection of the genetic pathways involved in the pathogenesis of arrhythmogenic cardiomyopathies. This review presents an overview of current animal models for arrhythmogenic right ventricular cardiomyopathy (ARVC), hypertrophic cardiomyopathy (HCM) and dilated cardiomyopathy (DCM) associated with cardiac arrhythmias and sudden cardiac death.
Introduction
Inherited arrhythmogenic cardiomyopathies are the most common cause of sudden cardiac death (SCD) in patients under 40 years of age. According to a recent expert consensus panel, cardiomyopathies are defined as ‘a heterogeneous group of diseases of the myocardium associated with mechanical and/or electrical dysfunction that usually exhibit inappropriate ventricular hypertrophy or dilatation, and are due to a variety of causes that frequently are genetic’ (Maron et al., 2006). Advances in molecular genetics led to the development of a new classification of cardiomyopathies and the recognition that a subset of inherited cardiomyopathies is associated with lethal ventricular tachyarrhythmias (Maron et al., 2006). The most prototypical of these diseases, arrhythmogenic right ventricular cardiomyopathy/dysplasia (ARVC/D), commonly presents clinically with ventricular tachyarrhythmias and tends to affect the right ventricle (RV) more than the left ventricle (LV) (MacRae et al., 2006). ARVC/D is typically caused by inherited mutations in desmosomal proteins (Moric-Janiszewska and Markiewicz-Loskot, 2007). Hypertrophic cardiomyopathy (HCM) is characterized by a hypertrophied, non-dilated LV in the absence of other systemic or cardiac disease. HCM is most commonly caused by a variety of mutations in genes encoding proteins that are found in the cardiac sarcomere. Familial forms of dilated cardiomyopathy (DCM) are characterized by ventricular enlargement and systolic dysfunction in the absence of LV hypertrophy. DCM is associated frequently with supraventricular arrhythmias and SCD (Boussy et al., 2008), and can be caused by mutations in sarcomeric, cytoskeletal or nuclear envelope proteins. Other forms of cardiomyopathy, including left ventricular noncompaction (LVNC) and restrictive cardiomyopathy (RCM) are less common and are not discussed in this review.
Animal models of arrhythmogenic cardiomyopathies have contributed to a better understanding of the pathophysiological processes leading to both cardiomyopathy and an increased susceptibility to cardiac arrhythmias (Awad et al., 2008). A variety of animals have been used to examine the pathological effects of mutations at both the whole organism/organ level and the cellular and molecular level (Berul, 2003). Initial animal models typically involved the serendipitous discovery of an inherited form of cardiomyopathy, followed by a forward genetic approach to identify the defect underlying the disease phenotype (Brink and Lochner, 1967; Kittleson et al., 1999). With the advent of genetically modified mouse models, inherited mutations identified in humans with cardiomyopathy are now modeled in transgenic or knock-in mice that may or may not recapitulate the clinical features of the disease (Berul, 2003). It is important to note that transgenic mouse models may have limitations from unintended alterations in gene expression besides the targeted gene. Likewise, single gene knockout models may introduce compensatory changes in other related structural genes, which may confound phenotypic results. However, animal models of arrhythmogenic cardiomyopathy continue to provide valuable insights into our understanding of the arrhythmogenic substrate, and the confluence of both cardiomyopathic and arrhythmogenic phenotypes. In this review, we will evaluate the current state of animal models of cardiomyopathies that are associated with cardiac arrhythmias, and will briefly discuss the new mechanistic insights obtained from these studies.
Models of arrhythmogenic right ventricular cardiomyopathy/dysplasia (ARVC/D)
ARVC/D is a heritable cardiomyopathy that primarily affects the RV (Anderson, 2006), and is the most common cause of SCD in competitive athletes in Italy (Maron et al., 2006). The typical manifestation of the disease involves the progressive destruction of desmosomal junctions causing apoptosis and eventual fibrotic replacement, predominantly of the RV (MacRae et al., 2006). Autosomal-dominant ARVC/D has been mapped to 12 chromosomal loci, and mutations have been identified in at least 12 genes thus far (Table 1). Two autosomal-recessive forms have been described in conjunction with woolly hair and kerataoderma (i.e. Naxos disease), and with Carvajal syndrome. Most forms of ARVC/D are caused by inherited mutations in desmosomal proteins that anchor the intermediate filaments to the desmosomes.
Table 1.
Animal models of ARVC/D
| Protein | Gene | Mutation | Model type | Animal | Phenotype | Reference |
|---|---|---|---|---|---|---|
| Ryanodine receptor type 2 | RYR2 | – | Spontaneous | Canine (Boxer) | Sustained VT, syncope, SCD, RV failure, reduced expression of wild-type RyR2 | Basso et al., 2004; Meurs et al., 2006 |
| Ryr2 | R176Q | KI | Mouse | VT induced by isoproterenol or programmed stimulation, reduced RV end-diastolic volume | Kannankeril et al., 2006 | |
| Laminin receptor | Lamr1 | – | Retroposon-related deletion | Mouse | QRS prolongation, RV fibrosis | Asano et al., 2004 |
| LBD3 (Cypher/ZASP) | Lbd3 | – | KO | Mouse | Biventricular dilation, fragmented myocyte Z-lines, muscle weakness | Zhou et al., 2001 |
| Lbd3 | – | CS KO | Mouse | Early cardiac death, LVNC dilated cardiomyopathy | Zheng et al., 2009 | |
| Desmoplakin | Dsp | V30M, Q90R, R2834H, W233X | CS Tg | Mouse | RV hypertrophy, systolic dysfunction, cardiomyocyte apoptosis, fibrosis, lipid accumulation | Yang et al., 2006 |
| Dsp | – | CS KO, KO | Mouse | Polymorphic VT, LV dilation, myocyte fibrosis, reduced fractional shortening | Garcia-Gras et al., 2006 | |
| Plakophilin 2 | Pkp2 | – | KO | Mouse | Embryonic lethality, RV wall thinning, hemopericardium | Grossman et al., 2004 |
| Desmocollin 2 | dsc2 | – | EMK | Zebra fish | Myocardial contractility dysfunction, reduced desmosomal plaques, loss of desmosome midlines | Heuser et al., 2006 |
| Junction plakoglobin | Jup | – | KO | Mouse | Spontaneous VT, increased RV volume, reduced RV function with age and exercise training | Kirchhof et al., 2006 |
| jup | – | EMK | Zebra fish | Aberrant valvulogenesis, decreased heart size, bradycardia, cardiac edema | Martin et al., 2009 |
CS, cardiac-specific; EMK, embryonic morpholino knockdown; KI, knock-in; KO, knockout; LV, left ventricle; LVNC, left ventricular noncompaction; RV, right ventricle; SCD, sudden cardiac death; Tg, transgenic; TI, tamoxifen-inducible; VT, ventricular tachycardia.
The desmosome is a structural junction that links cardiomyocytes, and that transmits the tensile force of contraction from cell to cell (Fig. 1) (MacRae et al., 2006). The desmosome is comprised of intracellular and extracellular proteins, including desmoplakin, desmocollin, desmoglein, plakophilin 2 and junction plakoglobin (Marcus and Towbin, 2006). These proteins bind to cardiomyocyte intermediate filaments and N-cadherins in a complex array that tethers cardiomyocytes together through intermediate filaments (Stokes, 2007). Mutations in several of these proteins have been shown to destabilize the desmosome junction, leading to physical separation of cardiomyocytes and apoptosis. Cellular remodeling of the ventricle includes fibrosis and often adipogenesis, which incompletely replaces primarily the RV myocardium. ARVC/D usually presents clinically with monomorphic ventricular tachycardias. The hypothesized mechanism of arrhythmogenesis in ARVC/D is conduction block and formation of macro re-entry circuits that are caused by fibrotic myocardium resulting from desmosome dehiscence (Wolf and Berul, 2008). It is still unknown why the RV is targeted uniquely in this inherited disease.
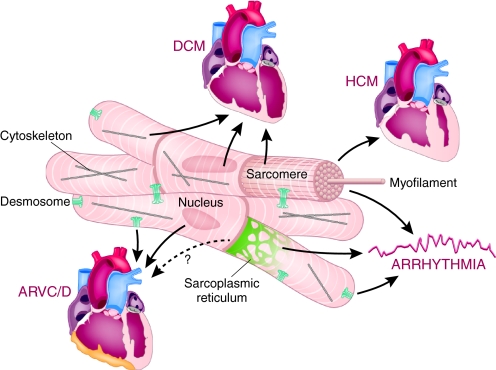
Fig. 1.
Inherited defects associated with various types of cardiomyopathy. A variety of genetic alterations in proteins that are involved in the cardiomyocyte cytoarchitecture manifest as cardiac arrhythmias and cardiomyopathy. HCM is caused typically by mutations in genes encoding proteins of the sarcomere, which is the fundamental contractile unit of the heart. DCM may be caused by mutations in some of the same sarcomeric genes, or in genes encoding cytoskeletal, nuclear envelope or transcription coactivator proteins. ARVC/D is caused typically by mutations in genes encoding desmosomal proteins, which comprise a key intercellular adhesive structure. Discrete changes in these structural and contractile elements of the cardiomyocyte have predictable effects on the phenotype of both cardiomyopathy and arrhythmias, as demonstrated in animal models.
Several animal models of ARVC/D caused by desmosomal proteins have been shown to recapitulate important aspects of the human disease phenotype (Table 1). Mutations in the gene encoding desmoplakin (DSP) are a classic example of desmosomal dysfunction leading to the ARVC/D type 8 (ARVC/D-8) phenotype. Using the human squamous carcinoma line SCC9 as a cellular model of desmosome formation, it was demonstrated that the N-terminal mutants V30M and Q90R failed to localize to the plasma membrane, whereas the C-terminal R2834H mutation did not affect the function of the N-terminus (Yang et al., 2006). Cardiac-specific overexpression of V30M or Q90R mutant desmoplakin in transgenic mice resulted in embryonic lethality owing to profound ventricular dilation. By contrast, cardiac-specific overexpression of the C-terminal mutant R2834H resulted in viable mice that developed ventricular enlargement and biventricular cardiomyopathy, which were detected on echocardiography (Yang et al., 2006). Histological examination revealed cardiomyocyte apoptosis, fibrosis and lipid accumulation, which are also found typically in patients with ARVC/D-8. Cardiac-restricted deletion of Dsp, encoding desmoplakin, impaired cardiac morphogenesis and caused embryonic lethality in homozygous knockout mice (Garcia-Gras et al., 2006). Heterozygous Dsp-deficient mice, however, exhibited excess adipocytes and fibrosis, increased apoptosis, defective cardiac contractility and ventricular arrhythmias, recapitulating the human ARVC/D phenotype.
Plakophilin 2 (or PKP2), a member of the armadillo family of proteins, is another integral member of the desmosome, where it binds to desmoplakin, desmocollin, desmoglein and plakoglobin (Hatzfeld, 2007). PKP2 is the only member of the plakophilin family to be expressed in the heart, and serves to tether desmosomal proteins (Rohr, 2007). Pkp2-deficient mouse embryos show abnormal heart morphogenesis and stability, followed by blood leakage, cardiac rupture and death on around embryonic day E11.5 (Grossmann et al., 2004). An evaluation of the embryonic hearts revealed evidence for myocardial wall thinning and aneurysm formation, which are also observed in patients with ARVC/D-9.
Junction plakoglobin (JUP) is another essential desmosomal protein which, when absent, causes an ARVC/D-12 phenotype in mice. Haploinsufficient Jup+/− mice developed RV dilation, reduced RV function and spontaneous ventricular ectopy (Kirchhof et al., 2006). Using a perfused heart preparation, an increased incidence of ventricular tachycardias of RV origin and prolonged RV conduction times were demonstrated. However, based on histology and electron microscopy, there was no evidence of changes in myocardial structure or desmosomal appearance, which suggests that Jup+/− mice only partially mimic the clinical ARVC/D phenotype. At present, there is no mouse model of desmocollin 2 (DSC2) mutations, which are identified in patients with ARVC/D-11.
In addition to mouse models, embryonic morpholino knockdown has been used in zebrafish to evaluate the effects of gene inhibition on embryogenesis. A morpholino is a synthetic antisense oligonucleotide with a high affinity for RNA, which acts as a blocker of translation and/or mRNA splicing (Chen and Ekker, 2004). Morpholino-induced knockdown of jup or dsc2 expression in zebrafish resulted in significant effects on cardiogenesis (Heuser et al., 2006; Martin et al., 2009). Zebrafish with reduced jup expression (morphants) developed small hearts, cardiac edema and valvular dysfunction leading to intrachamber reflux (Martin et al., 2009). Morphants exhibited a reduced number of desmosome and adheren junctions in the intercalated discs, consistent with the clinical ARVC/D-12 phenotype. However, morphants also showed abnormal valvulogenesis, and it is unclear whether this has clinical relevance. To model the physiological effects of a premature stop codon mutation, Heuser et al. (Heuser et al., 2006) performed morpholino knockdown of dsc2 in zebrafish. Morphants exhibited hearts with edema, bradycardia, reduced desmosomal areas, loss of desmosomal midlines and reduced contractility, as measured by fractional shortening. Rescue of the morphant phenotype by co-injection of wild-type human DSC2 mRNA implicated desmocollin 2 as a protein that is crucial for desmosomal function (Heuser et al., 2006). Whereas zebrafish morpholino models may offer insights into cardiac development and desmosomal organization in the absence of specific ARVC/D-associated genes, they do have important limitations because the morphology of the fish heart is very different from the human heart. Moreover, morpholino studies are limited typically to the embryonic stage and to knockdown models, whereas the aforementioned knock-in animal models offer a more accurate representation of the pathophysiology of human disease.
There are fewer animal models of ARVC/D caused by mutations in genes encoding non-desmosomal proteins. To date, there are no models for ARVC/D-1, 3, 4 and 6, and the molecular basis of these syndromes remains to be elucidated at the whole animal level. Asano et al. (Asano et al., 2004) identified a mouse model of ARVC/D-5 while generating mice for the screening of anti-diabetic compounds. They found a line of mice with an intron-processed retroposon in the laminin receptor 1 (Lamr1) gene locus, which resulted in translation of a Lamr1 protein with a 13-amino acid deletion (Asano et al., 2004). Lamr1-mutant mice developed RV dysplasia and fibrosis, which had limited extension to the LV. The findings in Lamr1-mutant mice were confirmed in two lines of transgenic mice that expressed mutant Lamr1, either systemically or only in the heart (Asano et al., 2004). All strains of mutant Lamr1-transgenic mice were very susceptible to ARVC/D, suggesting that normal Lamr1 function is essential for normal cardiomyocyte adhesion. Although mutations in CYPHER/ZASP (also known as LBD3) were implicated previously in ARVC/D-7 (Moric-Janiszewska and Markiewicz-Loskot, 2007), it has been proposed more recently that these patients develop a different disease phenotype consisting of isolated noncompaction of the LV myocardium, associated with DCM. Models of cardiomyopathy that result from mutations in CYPHER/ZASP exist, including a CYPHER knockout (Zhou et al., 2001) and a conditional CYPHER knockout (Zheng et al., 2009), both of which develop severe biventricular DCM and early postnatal death. However, mutations in this sarcomeric Z-line protein are not associated primarily with arrhythmias, and thus CYPHER/ZASP is not within the scope of this review.
Mutations in the ryanodine receptor type 2 gene (RYR2) have been proposed to cause ARVC/D-2 (Dokuparti et al., 2005), although this association has been questioned subsequently by several groups (Wehrens, 2007). These patients typically have exercise- or emotion-induced ventricular tachycardia, often referred to as catecholaminergic polymorphic ventricular tachycardia, or CPVT. Most patients with CPVT have structurally normal hearts, although a subset might develop a mild cardiomyopathy with incomplete penetrance. Given its unique role in exercise-induced ventricular arrhythmias without significant evidence of cardiomyopathy, it is likely that the syndrome caused by RyR2 mutations represents a separate entity from ARVC/D (Tiso et al., 2001). One notable model of the contribution of RYR2 abnormalities in the ARVC/D phenotype is a spontaneously generated canine boxer (Basso et al., 2004). This model was found subsequently to have differential expression of the ryanodine receptor, with decreased RYR2 expression in the RV (Meurs et al., 2006), which may contribute to the ARVC/D phenotype. Further research is necessary to delineate the exact contribution of RYR2 to RV cardiomyopathy, and to determine whether there is a direct or indirect ultrastructural relationship between the desmosome and the sarcoplasmic reticulum.
Models of hypertrophic cardiomyopathy (HCM)
Familial hypertrophic cardiomyopathy (FHC) is an inherited disorder primarily involving the sarcomere, which results in the concentric hypertrophy of the ventricle (Marian, 2008). FHC is significant because it is the most common cause of SCD in the young. Many components of the sarcomere, when affected by genetic mutations, have been proven to lead to a HCM phenotype. Three sarcomeric genes have been implicated in over 70% of FHC cases in humans: MYH7, MYBPC3 and TNNT2, which encode for the myosin heavy chain, myosin-binding protein C and troponin T proteins, respectively (Keren et al., 2008). Additionally, genetic mutations in troponin I, and in the signaling proteins Gα1 and Rho have been implicated in FHC with arrhythmias. The precise mechanisms by which functional defects within the sarcomere could evoke arrhythmias remains incompletely understood, although experimental studies in animal models of HCM have pointed to abnormal ATP use and disturbances in Ca2+ cycling (Keren et al., 2008). Re-entrant arrhythmias could also arise in the presence of myocyte disarray and interstitial fibrosis, which are hallmarks of hypertrophic hearts. Table 2 lists the relevant FHC animal models for arrhythmogenic cardiomyopathy. For a more general discussion of animal models with HCM only, Morimoto offers an excellent review (Morimoto, 2008).
Table 2.
Animal models of arrhythmogenic mutations causing hypertrophic cardiomyopathy
| Protein | Gene | Mutation | Model type | Animal | Phenotype | Reference |
|---|---|---|---|---|---|---|
| Cardiac troponin T | Tnnt2 | – | KO | Mouse | Embryonic lethal, asystole, myo filament disarray | Nishii et al., 2008 |
| Tnnt2 | I79N | CS Tg | Mouse | Absence of hypertrophy or fibrosis, isoproterenol-induced ventricular ectopy, stress-induced non-sustained VT | Knollmann et al., 2003 | |
| Tnnt2 | F110I R278C |
CS Tg | Mouse | Isoproterenol- or pacing-induced VT in the F110I Tg, but not R278C Tg, mouse | Baudenbacher et al., 2008 | |
| Tnnt2 | Deletion of exon 16 | CS Tg | Rat | VT, SCD, ventricular dilation | Frey et al., 2000 | |
| α-Myosin heavy chain (α-MHC) | Myh6 | R403G | KI | Mouse | Pacing-induced VT, variable LV hypertrophy, severe myocyte disarray | Wolf et al., 2005 |
| Troponin I + α-MHC |
Myh6 Tnni3 |
R403G (Myh6) + G203S (Tnni3) | KI × Tg crossed line | Mouse | VT, ventricular dilation, severe heart failure | Tsoutsman et al., 2008 |
| β-Myosin heavy chain (β-MHC) | Myh7 | Q403 | Tg | Rabbit | VT, cardiac hypertrophy | Ripplinger et al., 2007 |
| Myosin binding protein C | Mybpc3 | – | KO | Mouse | Inducible VT, HCM | Berul et al., 2001 |
| Gα1 | Ras | – | TI CS Tg | Mouse | Atrial fibrillation, VT, conduction block, HCM | Ruan et al., 2007 |
CS, cardiac-specific; HCM, hypertrophic cardiomyopathy; KI, knock-in; KO, knockout; LV, left ventricle; SCD, sudden cardiac death; Tg, transgenic; TI, tamoxifen-inducible; VT, ventricular tachycardia.
Cardiac troponin T (TNNT2 or cTnT) encoded by the TNNT2 gene composes the sarcomeric thin filament, and is necessary for normal cardiac contractile function (Keren et al., 2008). TNNT2 is essential for normal heart development in the mouse embryo, and mice with a null allele at the TNNT2 locus undergo early embryonic death from impaired fibrillogenesis and uncoupling of troponin I and tropomyosin (Nishii et al., 2008). A transgenic rat model overexpressing human TNNT2 that was truncated at exon 16 was used to demonstrate systolic and diastolic dysfunction, as well as ventricular tachycardia (VT) and fibrillation, following exercise (Frey et al., 2000). By contrast, transgenic rats overexpressing wild-type TNNT2 were found to have myocardial hypertrophy and improved cardiac performance in the absence of cardiac arrhythmias (Frey et al., 2000). Thus, there are several robust mouse and rat models of TNNT2 mutations, which reproduce the predisposition to both HCM and/or ventricular arrhythmias when this filament is compromised.
Patients with some TNNT2 mutations, however, exhibit a much higher incidence of arrhythmias than would be expected based on the subtle degree or absence of cardiac hypertrophy, suggesting that these TNNT2 defects are somehow involved directly in the arrhythmia generation (Knollmann and Potter, 2001). To evaluate this hypothesis further, Knollmann et al. (Knollmann et al., 2003) created transgenic mice overexpressing the I79N-mutant Tnnt2. Similar to findings in patients carrying this mutation, Tnnt2-I79N mutant mice were significantly more likely to develop ventricular arrhythmias when compared with wild-type mice, or even with other troponin T mutant mice. This selected Tnnt2 mutation in mice markedly increased the sensitivity of cardiac myofilaments to Ca2+, thus providing insight into the enhanced vulnerability to ventricular arrhythmias.
Baudenbacher et al. (Baudenbacher et al., 2008) extended these findings by demonstrating that increased myofilament Ca2+ sensitivity is an independent risk factor for arrhythmia susceptibility. Transgenic mice with Ca2+-sensitizing mutations in Tnnt2 (I79N and F110I) or that overexpress slow skeletal troponin I (Tnni1 or ssTnI) had an increased propensity towards arrhythmias, whereas the non-sensitizing Tnnt2 mutation R278C did not (Baudenbacher et al., 2008; Fiset and Giles, 2008). Interestingly, the risk of developing VT was directly proportional to the severity of Ca2+ sensitization of the myofilaments. When the Ca2+-sensitizing drug EMD 57033 was administered to non-transgenic mice, they reproduced the susceptibility to VT following cardiac pacing that was found in mice with Ca2+-sensitizing mutations. Additionally, administration of the Ca2+-desensitizing agent blebbistatin, rescued transgenic and EMD 57033-treated mice from arrhythmias. Mice with enhanced Ca2+ sensitization had shorter effective refractory periods, more beat-to-beat variability of action potential durations, and increased dispersion of ventricular conduction velocities at increased heart rates, suggesting that Ca2+ sensitization creates a substrate for functional re-entry and arrhythmias (Baudenbacher et al., 2008). Thus, Ca2+ sensitization has emerged as an important mechanism for arrhythmogenesis in HCM.
Another essential element of the sarcomere that is involved in myocyte contraction is α-myosin heavy chain (α-MHC). By interdigitating and sliding past the troponin-tropomyosin complex, the myosin heavy chain is involved directly in transmitting and actuating force along the axis of the sarcomere (Keren et al., 2008). Genetic alterations in the myosin heavy chain have been shown to cause sarcomere disarray, resulting in ventricular arrhythmias and hypertrophy. A knock-in mouse that is heterozygous for the mutation R403G in α-MHC (also known as Myh6) (R403G+/−) was shown to be susceptible to VT following intracardiac programmed stimulation and exhibited a variable degree of LV hypertrophy. When controlling for genetic background by examining inbred strains in identical environments, no correlation was found between the amount of fibrosis and/or myocyte disarray and susceptibility to arrhythmias, supporting the view that hypertrophy and arrhythmogenesis may be two distinct phenotypic outcomes from a common pathological process, and that there may be a dose-independent effect of fibrosis upon arrhythmia risk (Wolf et al., 2005). Additionally, the α-MHC-R403G mouse line was crossed with a cardiac-specific transgenic line overexpressing G203S-mutant cardiac troponin I (Tnni3), implicating troponin I in the severity of arrhythmogenesis in HCM (Tsoutsman et al., 2008). The double mutant mouse line showed increased postnatal mortality reaching 100% by day 21, with severe conduction-system block, VT, ventricular dilation and cardiac fibrosis. The development of DCM in double mutant mice was unexpected since both single mutant lines were shown previously to be hypertrophic (Wolf et al., 2005; Tsoutsman et al., 2006).
Additionally, a transgenic rabbit model of the mutation R403Q in β-myosin heavy chain (β-MHC, also known as MYH7) was created to investigate arrhythmogenic vulnerability and myofibril disarray in HCM (Marian et al., 1999; Ripplinger et al., 2007). Transgenic rabbits showed an increased vulnerability to ventricular arrhythmias compared with wild-type rabbits. Cardiac magnetic resonance imaging (MRI) revealed LV transmural fiber rotation, suggesting that structural tissue remodeling might underlie increased arrhythmogenesis. Thus, the myosin heavy chain is essential in maintaining the structural integrity of the sarcomere, which upon breakdown manifests in HCM and sometimes DCM.
Other models for HCM include support structures to the sarcomere, such as myosin binding protein C (MYBPC3), and signaling molecules such as Gα1. One notable model of MYBPC3 is a transgenic model created by Berul (Berul et al., 2001). Homozygous transgenic mice expressing truncated Mybpc3 showed HCM, whereas heterozygous transgenic mice exhibited a mild HCM. Following intracardiac programmed stimulation, a significant increase in VT was noted in homozygous (64%) and heterozygous (20%) transgenic mice, compared with wild-type control mice (0%).
Signaling cascades such as the Ras–Raf–mitogen-activated protein kinases pathway have been shown to be involved in both cardiac remodeling and arrhythmogenesis (Ruan et al., 2007). Transgenic mice with a tamoxifen-inducible cardiac-specific overexpression of the Ras signaling protein exhibited early-onset ventricular hypertrophy, followed seven days later with ventricular arrhythmias. The onset of arrhythmias correlated temporally with cellular action potential prolongation, altered outward K+ currents and a loss of protein kinase A (PKA) phosphorylation of phospholamban, thus altering intracellular calcium handling. This effect, in turn, caused these mice to be more susceptible to ventricular arrhythmias.
Models of dilated cardiomyopathy (DCM)
DCM is characterized by decreased cardiac function and enlargement of the heart, and represents the most common form of non-ischemic cardiomyopathy. About 30% of DCM cases are caused by inherited mutations in a variety of proteins that are found in the cytoskeleton, sarcomere and nuclear envelope (Hershberger et al., 2009). There is a considerable body of evidence in animal models supporting the idea that cytoskeletal and sarcomeric mutations may manifest in DCM with a predisposition to arrhythmias. Many mutations in sarcomeric proteins, including the troponins, may manifest differentially in either HCM or DCM, and thus there is not always a clear genotype-phenotype correlation in animal models of DCM (Table 3).
Table 3.
Animal models of arrhythmogenic mutations causing dilated cardiomyopathy
| Protein | Affected gene | Mutation | Model type | Animal | Phenotype | Reference |
|---|---|---|---|---|---|---|
| Ena-VASP | Vasp | – | CS Tg | Mouse | Bradycardia, early lethality, DCM, myocyte hypertrophy | Eigenthaler et al., 2003 |
| Neuron-restrictive silencer factor (NRSF) | Rest | – | CS DN Tg | Mouse | Non-sustained VT, SCD, DCM | Kuwahara et al., 2003 |
| Vinculin | Vcl | – | CS KO | Mouse | Ventricular arrhythmias, SCD, DCM | Zemljic-Harpf et al., 2007 |
| Lamin-A/C | Lmna | – | KO | Mouse | Atrial fibrillation, AV conduction defects, sustained VT, DCM | Wolf et al., 2008 |
| Lmna | N195K | KI | Mouse | Second-degree AV block, marked diurnal HR variability, DCM | Mounkes et al., 2005 | |
| Phospholamban | Pln | R14del | KI | Mouse | VT, premature death, DCM, cardiac fibrosis | Haghighi et al., 2006 |
AV, atrioventricular; CS, cardiac-specific; DCM, dilated cardiomyopathy; DN, dominant-negative; HR, heart rate; KI, knock-in; KO, knockout; SCD, sudden cardiac death; Tg, transgenic; VT, ventricular tachycardia.
Genetic mutations in myosin have been shown to cause DCM and reduced fractional shortening in mice, but have not yet demonstrated increased susceptibility to arrhythmias (Schmitt et al., 2006). However, defects in modulators of myosin function have been shown to induce arrhythmias in mice with a DCM phenotype. For example, the mammalian enabled (Mena) and vasodilator-stimulated phosphoprotein (VASP) proteins are actin cytoskeletal signal modulators, which are required for normal cardiogenesis and function (Eigenthaler et al., 2003). Specifically, the Ena-VASP proteins are crucial for actin filament formation, which is involved in cell adhesion, cell motility and cell growth (Eigenthaler et al., 2003). Transgenic mice with cardiac-specific overexpression of the N-terminal Ena-VASP homology domain (VASP-EVH1) exhibited a severe DCM phenotype and showed an increased susceptibility to bradycardia and early postnatal lethality (Eigenthaler et al., 2003), which resulted from the displacement of endogenous VASP and Mena proteins from intercalated disks in cardiac myocytes. Thus, Ena-VASP, a modulator of myosin function, contributes to actin filament structure and function. When Ena and VASP are disrupted, mice show a DCM phenotype, which results from weakened cell-cell interactions and which may impair normal cardiac conduction and lead to arrhythmias.
Another modulator of myosin, the transcriptional regulator neuron-restrictive silencer factor (NRSF), was shown to be involved in arrhythmogenesis in cardiac-specific transgenic mice that overexpress dominant-negative Nrsf (Kuwahara et al., 2003). These mice developed a DCM phenotype and were susceptible to both VT and advanced second-degree heart block. Nrsf (also known as Rest) suppression was shown to upregulate the expression of ventricular If and ICa,T ion channels, which are usually restricted to pacemaker and Purkinje cells, suggesting that increased automaticity may underlie the predisposition to VT.
In cardiomyocytes, there is a tight interaction between the cytoskeleton and the nuclear envelope, which is forged by the nuclear lamina intermediate filament lamin (Wolf et al., 2008). Mutations in lamin have recently been shown to cause the DCM phenotype with arrhythmias. Heterozygous lamin knockout (Lmna+/−) mice were shown to develop an age-dependent atrioventricular nodal disease, atrial arrhythmias and VT (Wolf et al., 2008). Lmna+/− mice also developed DCM with decreased fractional shortening on echocardiography and abnormally shaped nuclei on histology. In another study, transgenic mice overexpressing N195K-mutant Lmna developed severe bradyarrhythmias and sinoatrial exit block in addition to the DCM phenotype (Mounkes et al., 2005). At the cellular level, increased fibrosis and abnormal desmin localization were demonstrated, suggesting that changes in cell-cell coupling in mutant hearts may underlie the arrhythmia propensity.
Other notable models of DCM associated with arrhythmias include a transgenic mouse with cardiac-specific overexpression of R14del-mutant phospholamban (Pln) (Haghighi et al., 2006), and a cardiac-specific vinculin (Vcl) knockout mouse (Zemljic-Harpf et al., 2007). Both models exhibited an increased incidence of early SCD, DCM and advanced heart failure. Thus, several animal models demonstrate that disruptions in cytoskeletal interactions with the sarcomere, nuclear envelope, and within other cytoskeletal elements, may manifest differentially in DCM with an increased incidence of both atrial and ventricular arrhythmias.
Current insights into mechanisms of arrhythmogenesis in animal models
The ‘final common pathway’ hypothesis by Towbin and Bowles was one of the first attempts to link structural protein mutations to the major presentations of heart failure: most notably dilated and hypertrophic cardiomyopathies (Bowles et al., 2000). According to this hypothesis, DCM arises primarily from alterations in cytoskeletal proteins, whereas ARVC/D is caused by structural weakening of the desmosome, which predisposes the RV to fibrosis and dilation (Stokes, 2007). Likewise, HCM is thought to arise from disruptions of sarcomere function, and from abnormal calcium handling and sensitization. Although this hypothesis was revolutionary in organizing the relationship between subcellular structures and cardiac function, the exact mechanisms underlying arrhythmogenesis in these diseases remained unexplained.
Current animal models have allowed us to begin to unlock the mechanisms underlying arrhythmogenic cardiomyopathies. These mechanisms may be divided into disorders of automaticity, triggered activity (such as delayed afterdepolarizations) and re-entry circuits (Wolf and Berul, 2008). In ARVC/D, RV fibrosis may contribute to conduction-system block or the separation of myocytes, which may promote re-entrant arrhythmias. Moreover, downregulation of connexin-43 may contribute to heterogeneity in repolarization and an increased likelihood of ventricular arrhythmias (Boukens et al., 2009). Likewise, reduced expression of connexin-43 in a Syrian cardiomyopathic hamster model of HCM was shown to lead to increased action potential dispersion and a propensity to develop VT at 20 weeks of age (Sato et al., 2008).
Arrhythmias in patients with HCM or DCM may also be reentrant in nature, in particular in structurally remodeled hearts with myocyte disarray and fibrosis (Wolf and Berul, 2008). However, an increased susceptibility to arrhythmia has also been noted in patients and mice with HCM-linked mutations in which structural remodeling is absent. The increased expression of ion channels that are involved in pacemaking (If and ICa,T) in hypertrophic hearts might lead to increased automaticity (Wolf and Berul, 2008). Recent studies have also revealed that Ca2+ sensitization of myofilaments in HCM may promote afterdepolarizations and triggered arrhythmias (Baudenbacher et al., 2008). In fact, the degree of Ca2+ sensitization appears to be linked to the inducibility of ventricular arrhythmia, at least in mouse models (Baudenbacher et al., 2008). Although it is generally believed that desensitization of the myofilaments plays a role in the pathogenesis of DCM, the etiology of the arrhythmias associated with this condition remains poorly understood (Wolf and Berul, 2008). Finally, intracellular Ca2+ handling may be directly or indirectly impaired in inherited arrhythmogenic cardiomyopathies, suggesting that triggered activity as a result of afterdepolarizations might initiate arrhythmias.
Conclusions
Common inherited arrhythmias that are associated with cardiomyopathies arise from mutations in structures that span the entire cardiomyocyte, including the sarcomere, desmosome, cytoskeleton and the nuclear envelope. Although these regions seem disparate, the common unifying theme is that genetic defects leading to ineffective contraction cause ventricular remodeling, ultimately leading to cardiomyopathy. The resultant changes in ventricular fibrosis, hypertrophy and disordered calcium homeostasis create a substrate for arrhythmogenesis to occur. Further research is necessary to refine our understanding of the subtle ways in which these diseases are interrelated and to develop a common mechanism and nomenclature to describe these diseases. Current models show us that arrhythmogenicity and heart failure are separate entities in affected mutants, but that they may have a compounding effect as well. Thus, animal models will continue to serve as the key for unlocking the mechanism of how arrhythmogenic cardiomyopathies develop, and will continue to be useful in novel pharmacological therapies for treating the disease in humans.
Acknowledgments
M.D.M. is supported by the Molecular Medicine Scholars Training Grant (5T32HL066991). X.H.T.W. is a W.M. Keck Foundation Distinguished Young Scholar in Medical Research and is also supported by grants from the Muscular Dystrophy Association (69238); NIH/NHLBI grants R01-HL089598 and HL091947; a March of Dimes grant (MOD24172); the Hankamer Foundation; and the Alliance for Calmodulin Kinase Signaling in Heart Disease (08CVD01). Deposited in PMC for release after 12 months.
Footnotes
COMPETING INTERESTS
The authors declare no competing financial interests.
REFERENCES
- Anderson EL. (2006). Arrhythmogenic right ventricular dysplasia. Am Fam Physician 73, 1391–1398 [PubMed] [Google Scholar]
- Asano Y, Takashima S, Asakura M, Shintani Y, Liao Y, Minamino T, Asanuma H, Sanada S, Kim J, Ogai A, et al. (2004). Lamr1 functional retroposon causes right ventricular dysplasia in mice. Nat Genet. 36, 123–130 [DOI] [PubMed] [Google Scholar]
- Awad MM, Calkins H, Judge DP. (2008). Mechanisms of disease: molecular genetics of arrhythmogenic right ventricular dysplasia/cardiomyopathy. Nat Clin Pract Cardiovasc Med. 5, 258–267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Basso C, Fox PR, Meurs KM, Towbin JA, Spier AW, Calabrese F, Maron BJ, Thiene G. (2004). Arrhythmogenic right ventricular cardiomyopathy causing sudden cardiac death in boxer dogs: a new animal model of human disease. Circulation 109, 1180–1185 [DOI] [PubMed] [Google Scholar]
- Baudenbacher F, Schober T, Pinto JR, Sidorov VY, Hilliard F, Solaro RJ, Potter JD, Knollmann BC. (2008). Myofilament Ca2+ sensitization causes susceptibility to cardiac arrhythmia in mice. J Clin Invest. 118, 3893–3903 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berul CI. (2003). Electrophysiological phenotyping in genetically engineered mice. Physiol Genomics 13, 207–216 [DOI] [PubMed] [Google Scholar]
- Berul CI, McConnell BK, Wakimoto H, Moskowitz IP, Maguire CT, Semsarian C, Vargas MM, Gehrmann J, Seidman CE, Seidman JG. (2001). Ventricular arrhythmia vulnerability in cardiomyopathic mice with homozygous mutant Myosin-binding protein C gene. Circulation 104, 2734–2739 [DOI] [PubMed] [Google Scholar]
- Boukens BJ, Christoffels VM, Coronel R, Moorman AF. (2009). Developmental basis for electrophysiological heterogeneity in the ventricular and outflow tract myocardium as a substrate for life-threatening ventricular arrhythmias. Circ Res. 104, 19–31 [DOI] [PubMed] [Google Scholar]
- Boussy T, Paparella G, de Asmundis C, Sarkozy A, Chierchia GB, Brugada J, Brugada R, Brugada P. (2008). Genetic basis of ventricular arrhythmias. Cardiol Clin. 26, 335–353, v. [DOI] [PubMed] [Google Scholar]
- Bowles NE, Bowles KR, Towbin JA. (2000). The “final common pathway” hypothesis and inherited cardiovascular disease: the role of cytoskeletal proteins in dilated cardiomyopathy. Herz 25, 168–175 [DOI] [PubMed] [Google Scholar]
- Brink AJ, Lochner A. (1967). Work performance of the isolated perfused beating heart in the hereditary myocardiopathy of the Syrian hamster. Circ Res. 21, 391–401 [DOI] [PubMed] [Google Scholar]
- Chen E, Ekker SC. (2004). Zebrafish as a genomics research model. Curr Pharm Biotechnol. 5, 409–413 [DOI] [PubMed] [Google Scholar]
- Dokuparti MV, Pamuru PR, Thakkar B, Tanjore RR, Nallari P. (2005). Etiopathogenesis of arrhythmogenic right ventricular cardiomyopathy. J Hum Genet. 50, 375–381 [DOI] [PubMed] [Google Scholar]
- Eigenthaler M, Engelhardt S, Schinke B, Kobsar A, Schmitteckert E, Gambaryan S, Engelhardt CM, Krenn V, Eliava M, Jarchau T, et al. (2003). Disruption of cardiac Ena-VASP protein localization in intercalated disks causes dilated cardiomyopathy. Am J Physiol Heart Circ Physiol. 285, H2471–H2481 [DOI] [PubMed] [Google Scholar]
- Fiset C, Giles WR. (2008). Cardiac troponin T mutations promote life-threatening arrhythmias. J Clin Invest. 118, 3845–3847 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frey N, Franz WM, Gloeckner K, Degenhardt M, Muller M, Muller O, Merz H, Katus HA. (2000). Transgenic rat hearts expressing a human cardiac troponin T deletion reveal diastolic dysfunction and ventricular arrhythmias. Cardiovasc Res. 47, 254–264 [DOI] [PubMed] [Google Scholar]
- Garcia-Gras E, Lombardi R, Giocondo MJ, Willerson JT, Schneider MD, Khoury DS, Marian AJ. (2006). Suppression of canonical Wnt/beta-catenin signaling by nuclear plakoglobin recapitulates phenotype of arrhythmogenic right ventricular cardiomyopathy. J Clin Invest. 116, 2012–2021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grossman A, Barenboim E, Azaria B, Sherer Y, Goldstein L. (2004). Arrhythmogenic right ventricular dysplasia: a possible cause of sudden incapacitation. Aviat Space Environ Med. 75, 697–699 [PubMed] [Google Scholar]
- Grossmann KS, Grund C, Huelsken J, Behrend M, Erdmann B, Franke WW, Birchmeier W. (2004). Requirement of plakophilin 2 for heart morphogenesis and cardiac junction formation. J Cell Biol. 167, 149–160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haghighi K, Kolokathis F, Gramolini AO, Waggoner JR, Pater L, Lynch RA, Fan GC, Tsiapras D, Parekh RR, Dorn GW, 2nd, et al. (2006). A mutation in the human phospholamban gene, deleting arginine 14, results in lethal, hereditary cardiomyopathy. Proc Natl Acad S ci USA 103, 1388–1393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatzfeld M. (2007). Plakophilins: multifunctional proteins or just regulators of desmosomal adhesion? Biochim Biophys Acta 1773, 69–77 [DOI] [PubMed] [Google Scholar]
- Hershberger RE, Lindenfeld J, Mestroni L, Seidman CE, Taylor MR, Towbin JA. (2009). Genetic evaluation of cardiomyopathy: a heart failure society of america practice guideline. J Card Fail. 15, 83–97 [DOI] [PubMed] [Google Scholar]
- Heuser A, Plovie ER, Ellinor PT, Grossmann KS, Shin JT, Wichter T, Basson CT, Lerman BB, Sasse-Klaassen S, Thierfelder L, et al. (2006). Mutant desmocollin-2 causes arrhythmogenic right ventricular cardiomyopathy. Am J Hum Genet. 79, 1081–1088 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kannankeril PJ, Mitchell BM, Goonasekera SA, Chelu MG, Zhang W, Sood S, Kearney DL, Danila CI, De Biasi M, Wehrens XH, et al. (2006). Mice with the R176Q cardiac ryanodine receptor mutation exhibit catecholamine-induced ventricular tachycardia and cardiomyopathy. Proc Natl Acad Sci USA 103, 12179–12184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keren A, Syrris P, McKenna WJ. (2008). Hypertrophic cardiomyopathy: the genetic determinants of clinical disease expression. Nat Clin Pract Cardiovasc Med. 5, 158–168 [DOI] [PubMed] [Google Scholar]
- Kirchhof P, Fabritz L, Zwiener M, Witt H, Schafers M, Zellerhoff S, Paul M, Athai T, Hiller KH, Baba HA, et al. (2006). Age- and training-dependent development of arrhythmogenic right ventricular cardiomyopathy in heterozygous plakoglobin-deficient mice. Circulation 114, 1799–1806 [DOI] [PubMed] [Google Scholar]
- Kittleson MD, Meurs KM, Munro MJ, Kittleson JA, Liu SK, Pion PD, Towbin JA. (1999). Familial hypertrophic cardiomyopathy in maine coon cats: an animal model of human disease. Circulation 99, 3172–3180 [DOI] [PubMed] [Google Scholar]
- Knollmann BC, Potter JD. (2001). Altered regulation of cardiac muscle contraction by troponin T mutations that cause familial hypertrophic cardiomyopathy. Trends Cardiovasc Med. 11, 206–212 [DOI] [PubMed] [Google Scholar]
- Knollmann BC, Kirchhof P, Sirenko SG, Degen H, Greene AE, Schober T, Mackow JC, Fabritz L, Potter JD, Morad M. (2003). Familial hypertrophic cardiomyopathy-linked mutant troponin T causes stress-induced ventricular tachycardia and Ca2+-dependent action potential remodeling. Circ Res. 92, 428–436 [DOI] [PubMed] [Google Scholar]
- Kuwahara K, Saito Y, Takano M, Arai Y, Yasuno S, Nakagawa Y, Takahashi N, Adachi Y, Takemura G, Horie M, et al. (2003). NRSF regulates the fetal cardiac gene program and maintains normal cardiac structure and function. EMBO J. 22, 6310–6321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacRae CA, Birchmeier W, Thierfelder L. (2006). Arrhythmogenic right ventricular cardiomyopathy: moving toward mechanism. J Clin Invest. 116, 1825–1828 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marcus F, Towbin JA. (2006). The mystery of arrhythmogenic right ventricular dysplasia/cardiomyopathy: from observation to mechanistic explanation. Circulation 114, 1794–1795 [DOI] [PubMed] [Google Scholar]
- Marian AJ. (2008). Genetic determinants of cardiac hypertrophy. Curr Opin Cardiol. 23, 199–205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marian AJ, Wu Y, Lim DS, McCluggage M, Youker K, Yu QT, Brugada R, DeMayo F, Quinones M, Roberts R. (1999). A transgenic rabbit model for human hypertrophic cardiomyopathy. J Clin Invest. 104, 1683–1692 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maron BJ, Towbin JA, Thiene G, Antzelevitch C, Corrado D, Arnett D, Moss AJ, Seidman CE, Young JB. (2006. Contemporary definitions and classification of the cardiomyopathies: an American Heart Association scientific statement from the Council on Clinical Cardiology, Heart Failure and Transplantation Committee; quality of care and outcomes research and functional genomics and translational biology interdisciplinary working groups; and council on epidemiology and prevention. Circulation 113, 1807–1816 [DOI] [PubMed] [Google Scholar]
- Martin ED, Moriarty MA, Byrnes L, Grealy M. (2009). Plakoglobin has both structural and signalling roles in zebrafish development. Dev Biol. 327, 83–96 [DOI] [PubMed] [Google Scholar]
- Meurs KM, Lacombe VA, Dryburgh K, Fox PR, Reiser PR, Kittleson MD. (2006). Differential expression of the cardiac ryanodine receptor in normal and arrhythmogenic right ventricular cardiomyopathy canine hearts. Hum Ge net. 120, 111–118 [DOI] [PubMed] [Google Scholar]
- Moric-Janiszewska E, Markiewicz-Loskot G. (2007). Review on the genetics of arrhythmogenic right ventricular dysplasia. Europace 9, 259–266 [DOI] [PubMed] [Google Scholar]
- Morimoto S. (2008). Sarcomeric proteins and inherited cardiomyopathies. Cardiovasc Res. 77, 659–666 [DOI] [PubMed] [Google Scholar]
- Mounkes LC, Kozlov SV, Rottman JN, Stewart CL. (2005). Expression of an LMNA-N195K variant of A-type lamins results in cardiac conduction defects and death in mice. Hum Mol Genet. 14, 2167–2180 [DOI] [PubMed] [Google Scholar]
- Nishii K, Morimoto S, Minakami R, Miyano Y, Hashizume K, Ohta M, Zhan DY, Lu QW, Shibata Y. (2008). Targeted disruption of the cardiac troponin T gene causes sarcomere disassembly and defects in heartbeat within the early mouse embryo. Dev Biol. 322, 65–73 [DOI] [PubMed] [Google Scholar]
- Ripplinger CM, Li W, Hadley J, Chen J, Rothenberg F, Lombardi R, Wickline SA, Marian AJ, Efimov IR. (2007). Enhanced transmural fiber rotation and connexin 43 heterogeneity are associated with an increased upper limit of vulnerability in a transgenic rabbit model of human hypertrophic cardiomyopathy. Circ Res. 101, 1049–1057 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rohr S. (2007). Molecular crosstalk between mechanical and electrical junctions at the intercalated disc. Circ Res. 101, 637–639 [DOI] [PubMed] [Google Scholar]
- Ruan H, Mitchell S, Vainoriene M, Lou Q, Xie LH, Ren S, Goldhaber JI, Wang Y. (2007). Gi alpha 1-mediated cardiac electrophysiological remodeling and arrhythmia in hypertrophic cardiomyopathy. Circulation 116, 596–605 [DOI] [PubMed] [Google Scholar]
- Sato T, Ohkusa T, Honjo H, Suzuki S, Yoshida MA, Ishiguro YS, Nakagawa H, Yamazaki M, Yano M, Kodama I, et al. (2008). Altered expression of connexin43 contributes to the arrhythmogenic substrate during the development of heart failure in cardiomyopathic hamster. Am J Physiol Heart Circ Physiol. 294, H1164–H1173 [DOI] [PubMed] [Google Scholar]
- Schmitt JP, Debold EP, Ahmad F, Armstrong A, Frederico A, Conner DA, Mende U, Lohse MJ, Warshaw D, Seidman CE, et al. (2006). Cardiac myosin missense mutations cause dilated cardiomyopathy in mouse models and depress molecular motor function. Proc Natl Acad S ci USA 103, 14525–14530 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stokes DL. (2007). Desmosomes from a structural perspective. Curr Opin Cell Biol. 19, 565–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tiso N, Stephan DA, Nava A, Bagattin A, Devaney JM, Stanchi F, Larderet G, Brahmbhatt B, Brown K, Bauce B, et al. (2001). Identification of mutations in the cardiac ryanodine receptor gene in families affected with arrhythmogenic right ventricular cardiomyopathy type 2 (ARVD2). Hum Mol Genet. 10, 189–194 [DOI] [PubMed] [Google Scholar]
- Tsoutsman T, Chung J, Doolan A, Nguyen L, Williams IA, Tu E, Lam L, Bailey CG, Rasko JE, Allen DG, et al. (2006). Molecular insights from a novel cardiac troponin I mouse model of familial hypertrophic cardiomyopathy. J Mol Cell Cardiol. 41, 623–632 [DOI] [PubMed] [Google Scholar]
- Tsoutsman T, Kelly M, Ng DC, Tan JE, Tu E, Lam L, Bogoyevitch MA, Seidman CE, Seidman JG, Semsarian C. (2008). Severe heart failure and early mortality in a double-mutation mouse model of familial hypertrophic cardiomyopathy. Circulation 117, 1820–1831 [DOI] [PubMed] [Google Scholar]
- Wehrens XH. (2007). The molecular basis of catecholaminergic polymorphic ventricular tachycardia: what are the different hypotheses regarding mechanisms? Heart Rhythm 4, 794–797 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolf CM, Berul CI. (2008). Molecular mechanisms of inherited arrhythmias. Curr Genomics 9, 160–168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolf CM, Moskowitz IP, Arno S, Branco DM, Semsarian C, Bernstein SA, Peterson M, Maida M, Morley GE, Fishman G, et al. (2005). Somatic events modify hypertrophic cardiomyopathy pathology and link hypertrophy to arrhythmia. Proc Natl Acad Sci USA 102, 18123–18128 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolf CM, Wang L, Alcalai R, Pizard A, Burgon PG, Ahmad F, Sherwood M, Branco DM, Wakimoto H, Fishman GI, et al. (2008). Lamin A/C haploinsufficiency causes dilated cardiomyopathy and apoptosis-triggered cardiac conduction system disease. J Mol Cell Cardiol. 44, 293–303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Z, Bowles NE, Scherer SE, Taylor MD, Kearney DL, Ge S, Nadvoretskiy VV, DeFreitas G, Carabello B, Brandon LI, et al. (2006). Desmosomal dysfunction due to mutations in desmoplakin causes arrhythmogenic right ventricular dysplasia/cardiomyopathy. Circ Res. 99, 646–655 [DOI] [PubMed] [Google Scholar]
- Zemljic-Harpf AE, Miller JC, Henderson SA, Wright AT, Manso AM, Elsherif L, Dalton ND, Thor AK, Perkins GA, McCulloch AD, et al. (2007). Cardiac-myocyte-specific excision of the vinculin gene disrupts cellular junctions, causing sudden death or dilated cardiomyopathy. Mol Cell Biol. 27, 7522–7537 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng M, Cheng H, Li X, Zhang J, Cui L, Ouyang K, Han L, Zhao T, Gu Y, Dalton ND, et al. (2009). Cardiac-specific ablation of Cypher leads to a severe form of dilated cardiomyopathy with premature death. Hum Mol Genet. 18, 701–713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou Q, Chu PH, Huang C, Cheng CF, Martone ME, Knoll G, Shelton GD, Evans S, Chen J. (2001). Ablation of Cypher, a PDZ-LIM domain Z-line protein, causes a severe form of congenital myopathy. J Cell Biol. 155, 605–612 [DOI] [PMC free article] [PubMed] [Google Scholar]