Gallate dioxygenase (DesB) from Sphingobium sp. SYK-6, which belongs to the type II extradiol dioxygenase family, was purified and crystallized in two different crystal forms, which were subjected to X-ray analysis.
Keywords: extradiol dioxygenases, gallate dioxygenase
Abstract
Gallate dioxygenase (DesB) from Sphingobium sp. SYK-6, which belongs to the type II extradiol dioxygenase family, was purified and crystallized using the hanging-drop vapour-diffusion method. Two crystal forms were obtained. The form I crystal belonged to space group C2, with unit-cell parameters a = 136.2, b = 53.6, c = 55.1 Å, β = 112.8°, and diffracted to 1.6 Å resolution. The form II crystal belonged to space group P21, with unit-cell parameters a = 56.2, b = 64.7, c = 116.1 Å, β = 95.1°, and diffracted to 1.9 Å resolution. A molecular-replacement calculation using LigAB as a search model yielded a satisfactory solution for both crystal forms.
1. Introduction
Lignin is an abundant biomass in nature and its decomposition is essential to the terrestrial carbon cycle. Although the utilization of lignin as biomass has been expected, few practical uses for lignin have been established because of its high resistance to both chemical and biological degradation. Lignin is a complicated polymer of monolignols (Davin & Lewis, 2005 ▶). The structure of lignin provides mechanical strength to cell walls, but causes recalcitrance to microbial degradation. In nature, white rot fungi initiate the degradation of lignin and bacteria in the soil further degrade the resultant low-molecular-weight lignin compounds (Martinez et al., 2005 ▶).
Sphingobium sp. SYK-6 (formerly Sphingomonas paucimobilis SYK-6), isolated from pulping waste liquor, can grow on various low-molecular-weight lignin compounds with two aromatic rings (Katayama et al., 1988 ▶), including β-aryl ether (Masai et al., 1991 ▶, 1993 ▶, 2003 ▶; Sato et al., 2009 ▶), biphenyl (Peng et al., 1998 ▶, 1999 ▶; Sonoki et al., 2000 ▶) and diarylpropane, as sole carbon and energy sources. Analysis of the metabolic pathway in Sphingobium sp. SYK-6 revealed that low-molecular-weight lignin compounds are metabolized to vanillate and syringate, which are converted to protocatechuate (PCA) and 3-O-methylgallate (3MGA), respectively (Masai et al., 2007 ▶). The catechol ring of PCA, 3MGA and gallate (a 3MGA metabolite produced by LigM) is cleaved by extradiol dioxygenases (Kasai et al., 2004 ▶, 2005 ▶; Masai et al., 2007 ▶). Since the ring-opening reactions of the catechol derivatives are the key steps in lignin degradation, the metabolic pathways of these aromatic compounds have been intensively studied.
In Sphingobium sp. SYK-6, three extradiol dioxygenases, PCA 4,5-dioxygenase (LigAB), gallate dioxygenase (DesB) and 3MGA 3,4-dioxygenase (DesZ), catalyze the ring-opening reactions of the catechol derivatives generated in the catabolism of vanillate and syringate (Kasai et al., 2004 ▶, 2005 ▶; Masai et al., 2007 ▶; Fig. 1 ▶ a). These enzymes belong to the type II extradiol dioxygenase family and contain a nonhaem ferrous ion in their active sites (Eltis & Bolin, 1996 ▶). Type II extradiol dioxygenases are evolutionarily distinct from type I extradiol dioxygenases, the catalytic mechanism and substrate specificities of which have been intensively studied (Sugiyama et al., 1995 ▶; Han et al., 1995 ▶; Senda et al., 1996 ▶; Dai et al., 2002 ▶; Sato et al., 2002 ▶; Kovaleva & Lipscomb, 2007 ▶, 2008 ▶). The crystal structure of LigAB, which is the only available crystal structure of the type II extradiol dioxygenases, showed convergent evolution of the active-site structure towards that of type I extradiol dioxygenases, suggesting that types I and II extradiol dioxygenases share the same catalytic mechanism (Sugimoto et al., 1999 ▶).
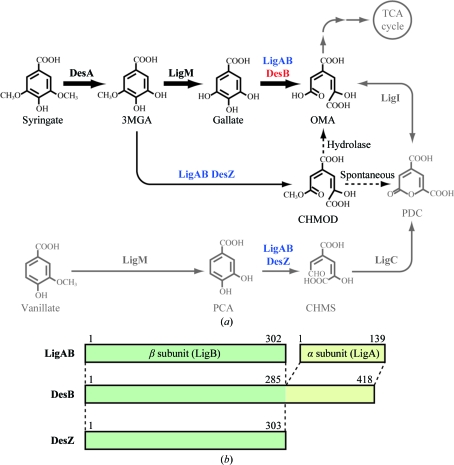
Figure 1.
(a) Degradation pathway of syringate and vanillate in Sphingobium sp. SYK-6. DesA, syringate O-demethylase; LigM, vanillate/3MGA O-demethylase; DesB, gallate dioxygenase; LigAB, PCA 4,5-dioxygenase; DesZ, 3MGA 3,4-dioxygenase; LigI, PDC hydrolase; LigC, 4-carboxy-2-hydroxymuconate-6-semialdehyde dehydrogenase; 3MGA, 3-O-methylgallate; OMA, 4-oxalomesaconate; PCA, protocatechuate; CHMOD, 4-carboxy-2-hydroxy-6-methoxy-6-oxohexa-2,4-dienoate; PDC, 2-pyrone-4,6-dicarboxylate; CHMS, 4-carboxy-2-hydroxymuconate-6-semialdehyde. (b) Schematic drawing of LigAB, DesB and DesZ from Sphingobium sp. SYK-6.
As shown in Fig. 1 ▶, Sphingobium sp. SYK-6 has a branched syringate-degradation pathway (Kasai et al., 2004 ▶, 2005 ▶). Syringate is mainly degraded via gallate and the pathway through CHMOD is a minor one (Kasai et al., 2005 ▶). However, the biological significance of the branched degradation pathway remains elusive. The differences in the substrate specificities and enzymatic activities of LigAB, DesB and DesZ are likely to play a critical role in controlling the metabolic flow in the branched syringate-degradation pathway.
Analysis of the primary structures of the three enzymes and the crystal structure of LigAB (Sugimoto et al., 1999 ▶) suggested that their different subunit compositions contribute to the different substrate specificities of LigAB, DesB and DesZ. LigAB has an α2β2 subunit composition (molecular weight 97.7 kDa), while DesB and DesZ are homodimeric enzymes with molecular weights of 93.8 and 73.1 kDa, respectively. In DesB, the N-terminal (residues 1–285) and the C-terminal (residues 286–418) regions exhibited 38 and 27% amino-acid identity to the sequences of the β and α subunits of LigAB, respectively (Fig. 1 ▶ b). DesZ, however, contains no region that corresponds to the α subunit of LigAB (Fig. 1 ▶ b). Since the active site of LigAB is composed of a deep cleft of the β subunit and a lid from the α subunit (Sugimoto et al., 1999 ▶), the difference in or total lack of the lid region in DesB or DesZ (Fig. 1 ▶ b) must have a significant effect on the substrate specificity of the enzymes. Therefore, the crystal structures of DesB and DesZ would provide novel insights into not only the origin of the substrate specificities of these type II extradiol dioxygenases but also the structural basis of the control of metabolic flow in the syringate-degradation pathway.
As the first step in this analysis, we analyzed the crystal structure of DesB (EC 1.13.11.–). Here, we report the purification, crystallization and preliminary X-ray diffraction studies of DesB.
2. Methods and results
2.1. Protein expression and purification
All purification steps were carried out at 277 K under aerobic conditions. Escherichia coli BL21 (DE3) cells were transformed with an expression plasmid containing the desB gene (GenBank AB190989; Kasai et al., 2005 ▶). Cells were grown in L-broth medium at 303 K. DesB expression was induced by IPTG when the culture reached an OD600 of 0.5–0.6. After 6 h expression, the cells were harvested, washed twice with 10% glycerol, 20 mM Tris–HCl pH 8.0 and resuspended in the same buffer. The cells were disrupted by sonication using a 500 W ultrasonic disintegrator (VCX500, Sonics). Cell debris and larger particles were removed by centrifugation.
DesB was purified from the obtained supernatant fraction by two chromatographic steps. The supernatant solution was applied onto a POROS PI column (Applied Biosystems) pre-equilibrated with 10% glycerol, 20 mM Tris–HCl pH 8.0. DesB was eluted with a linear NaCl gradient (0–0.5 M). The fractions containing enzyme activity were combined and dialyzed against 10% glycerol, 20 mM Tris–HCl pH 8.0. The dialyzed protein solution was applied onto a POROS HQ column (Applied Biosystems) which was pre-equilibrated with 10% glycerol, 20 mM Tris–HCl pH 8.0. DesB was eluted with a linear NaCl gradient (0–0.5 M). The purity of the sample was assessed by SDS–PAGE, showing that DesB was purified to near-homogeneity. The purified DesB was concentrated to 20 mg ml−1 by ultrafiltration using a Centricon YM-10 device (Millipore). During ultrafiltration, the buffer was exchanged to Milli-Q water.
2.2. Crystallization
Crystallization was carried out using the hanging-drop vapour-diffusion method at 277 K. A hanging drop was prepared by mixing 2 µl protein solution and 2 µl reservoir solution. Each hanging drop was placed over 1 ml reservoir solution. Initial crystallization trials were performed using Crystal Screen and Crystal Screen 2 (Hampton Research; Jancarik & Kim, 1991 ▶). The most promising condition was 30%(w/v) polyethylene glycol (PEG) 8000, 0.2 M sodium acetate trihydrate, 0.1 M sodium cacodylate pH 6.5 (Crystal Screen condition No. 28). In order to improve the crystal quality, the pH value (6.50–7.75) and the concentrations of the protein (14–30 mg ml−1), PEG 8000 (20–30%) and sodium acetate trihydrate (0.10–0.28 M) were systematically optimized. As a result, the highest quality crystals were obtained in 10–14 d at 277 K using 15 mg ml−1 protein solution and a reservoir solution containing 25%(w/v) PEG 8000, 0.1 M sodium acetate trihydrate, 0.1 M HEPES–NaOH pH 7.75 (Fig. 2 ▶).
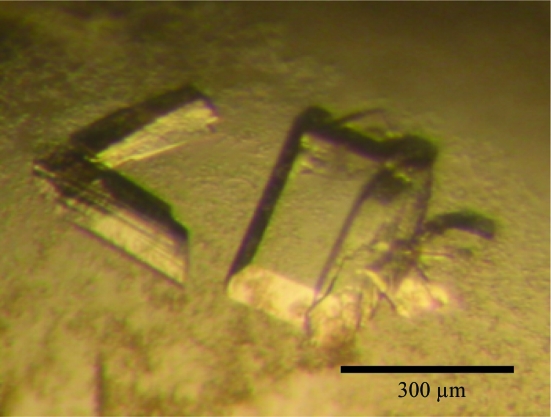
Figure 2.
Crystals of DesB. The largest dimension of the crystal is approximately 300 µm.
2.3. Data collection
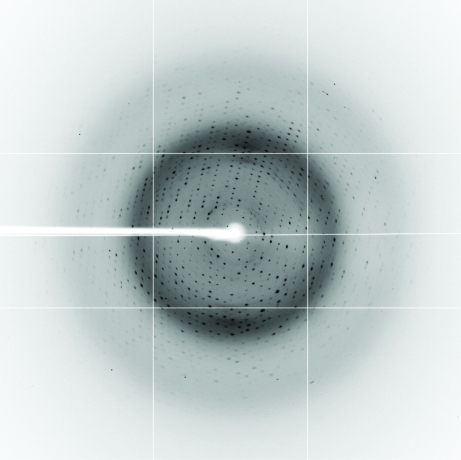
Diffraction data for the best crystal were collected using an ADSC Quantum 315r detector on beamline BL-5A of the Photon Factory (PF; KEK, Tsukuba, Japan; Fig. 3 ▶). After the crystal had been soaked in the reservoir solution for approximately 30 s, it was mounted on a cryo-loop (Hampton Research) and flash-cooled in an N2 stream. Crystal annealing was not performed. The diffraction data were processed and scaled using the XDS program suite (Kabsch, 1993 ▶). The crystal belonged to space group C2 (form I).
Figure 3.
X-ray diffraction pattern of the DesB crystal. The diffraction data were collected on BL-5A of PF using an ADSC Quantum 315r CCD detector.
Further data collection using crystals grown under similar conditions revealed another crystal form. A crystal grown with reservoir solution consisting of 25%(w/v) PEG 8000, 0.27 M sodium acetate trihydrate, 0.1 M HEPES–NaOH pH 7.25 belonged to space group P21 (form II). After the crystal had been soaked in the reservoir solution for a few days, it was flash-cooled in an N2 stream. The diffraction data were collected using an ADSC Quantum 4r detector on beamline BL-17A of the PF (KEK, Tsukuba, Japan). Data-collection statistics for the form I and from II crystals are given in Table 1 ▶.
Table 1. Data-collection statistics.
Values in parentheses are for the outermost resolution shell.
| Form I | Form II | |
|---|---|---|
| Beamline | BL-5A (PF) | BL-17A (PF) |
| Crystal-to-detector distance (mm) | 220.0 | 154.2 |
| Oscillation angle (°) | 1.0 | 1.0 |
| Exposure time (s) | 5 | 10 |
| Wavelength (Å) | 0.9790 | 1.0000 |
| Space group | C2 | P21 |
| Unit-cell parameters (Å, °) | a = 136.2, b = 53.6, c = 55.1, β = 112.8 | a = 56.2, b = 64.7, c = 116.1, β = 95.1 |
| Resolution limits (Å) | 17.0–1.60 (1.68–1.60) | 17.0–1.90 (2.00–1.90) |
| Observations | 218578 (24531) | 322721 (44622) |
| Unique reflections | 48034 (6404) | 63363 (8868) |
| Completeness (%) | 98.8 (97.0) | 96.6 (95.4) |
| Average I/σ(I) | 28.4 (5.3) | 20.0 (5.6) |
| Redundancy | 4.6 (3.8) | 5.1 (5.0) |
| Rmerge | 0.033 (0.264) | 0.067 (0.355) |
| Mosaicity (°) | 0.51 | 0.27 |
Assuming the presence of one subunit of DesB in the asymmetric unit, the Matthews coefficient (V M; Matthews, 1968 ▶) for the form I crystal was calculated to be 1.98 Å3 Da−1, corresponding to a solvent content of 37.8%. In the form II crystal, assuming the presence of one DesB dimer in the asymmetric unit, V M was 2.24 Å3 Da−1, corresponding to a solvent content of 45.2%.
The crystal structure was determined by the molecular-replacement method using the program MOLREP (Vagin & Teplyakov, 1997 ▶) in the CCP4 program suite (Collaborative Computational Project, Number 4, 1994 ▶) with a single α–β subunit pair from LigAB (PDB code 1bou; Sugimoto et al., 1999 ▶) as a search model. The molecular-replacement calculations gave satisfactory solutions for both crystal forms. No serious clashes between molecules were found in the crystal packing for both solutions. As expected from the V M values, the asymmetric units of the form I and form II crystals contained one and two subunits of DesB, respectively.
Model building for crystal form I was performed with the program Coot (Emsley & Cowtan, 2004 ▶). The molecular model of DesB demonstrated that the β and α subunits of LigAB correspond to the N- and C-terminal regions of DesB, respectively. Electron density for ferrous iron was clearly observed in the active site. Crystallographic refinement of the form I crystal is in progress.
Acknowledgments
This study was supported in part by Grants-in-Aid for Scientific Research from the Ministry of Education, Sports, Science, Culture and Technology (No. 17770096) and from the New Energy and Trial Technology Development Organization (NEDO) of Japan.
References
- Collaborative Computational Project, Number 4 (1994). Acta Cryst. D50, 760–763.
- Dai, S., Vaillancourt, F. H., Maaroufi, H., Drouin, N. M., Neau, D. B., Snieckus, V., Bolin, J. T. & Eltis, L. D. (2002). Nature Struct. Biol.9, 934–939. [DOI] [PubMed]
- Davin, L. B. & Lewis, N. G. (2005). Curr. Opin. Biotechnol.16, 407–415. [DOI] [PubMed]
- Eltis, L. D. & Bolin, J. T. (1996). J. Bacteriol.178, 5930–5937. [DOI] [PMC free article] [PubMed]
- Emsley, P. & Cowtan, K. (2004). Acta Cryst. D60, 2126–2132. [DOI] [PubMed]
- Jancarik, J. & Kim, S.-H. (1991). J. Appl. Cryst.24, 409–411.
- Han, S., Eltis, L. D., Timmis, K. N., Muchmore, S. W. & Bolin, J. T. (1995). Science, 270, 976–980. [DOI] [PubMed]
- Kabsch, W. (1993). J. Appl. Cryst.26, 795–800.
- Kasai, D., Masai, E., Miyauchi, K., Katayama, Y. & Fukuda, M. (2004). J. Bacteriol.186, 4951–4959. [DOI] [PMC free article] [PubMed]
- Kasai, D., Masai, E., Miyauchi, K., Katayama, Y. & Fukuda, M. (2005). J. Bacteriol.187, 5067–5074. [DOI] [PMC free article] [PubMed]
- Katayama, K., Nishikawa, S., Murayama, A., Yamasaki, M., Morohoshi, N. & Haraguchi, T. (1988). FEBS Lett.233, 129–133. [DOI] [PubMed]
- Kovaleva, E. G. & Lipscomb, J. D. (2007). Science, 316, 453–457. [DOI] [PMC free article] [PubMed]
- Kovaleva, E. G. & Lipscomb, J. D. (2008). Nature Chem. Biol.4, 186–193. [DOI] [PMC free article] [PubMed]
- Martinez, A. T., Speranza, M., Ruiz-Duenas, F. J., Ferreira, P., Camarero, S., Guillen, F., Martinez, M. J., Gutierrez, A. & del Rio, J. C. (2005). Int. Microbiol.8, 195–204. [PubMed]
- Masai, E., Ichimura, Y., Sato, K., Miyauchi, K., Katayama, Y. & Fukuda, M. (2003). J. Bacteriol.181, 1768–1775. [DOI] [PMC free article] [PubMed]
- Masai, E., Katayama, Y. & Fukuda, M. (2007). Biosci. Biotechnol. Biochem.71, 1–15. [DOI] [PubMed]
- Masai, E., Katayama, Y., Kawai, S., Nishikawa, M., Yamasaki, M. & Morohoshi, N. (1991). J. Bacteriol.173, 7950–7955. [DOI] [PMC free article] [PubMed]
- Masai, E., Katayama, Y., Kubota, S., Kawai, S., Yamasaki, M. & Morohoshi, N. (1993). FEBS Lett.323, 135–140. [DOI] [PubMed]
- Matthews, B. W. (1968). J. Mol. Biol.33, 491–497. [DOI] [PubMed]
- Peng, X., Egashira, T., Hanashiro, K., Masai, E., Nishikawa, S., Katayama, Y., Kimbara, K. & Fukuda, M. (1998). Appl. Environ. Microbiol.64, 2520–2527. [DOI] [PMC free article] [PubMed]
- Peng, X., Masai, E., Katayama, Y. & Fukuda, M. (1999). Appl. Environ. Microbiol.65, 2789–2793. [DOI] [PMC free article] [PubMed]
- Sato, N., Uragami, Y., Nishizaki, T., Takahashi, Y., Sazaki, G., Sugimoto, K., Nonaka, T., Masai, E., Fukuda, M. & Senda, T. (2002). J. Mol. Biol.321, 621–636. [DOI] [PubMed]
- Sato, Y., Moriuchi, H., Hishiyama, S., Otsuka, Y., Oshima, K., Kasai, D., Nakamura, M., Ohara, S., Katayama, Y., Fukuda, M. & Masai, E. (2009). Appl. Environ. Microbiol.75, 5195–5201. [DOI] [PMC free article] [PubMed]
- Senda, T., Sugiyama, K., Narita, H., Yamamoto, T., Kimbara, K., Fukuda, M., Sato, M., Yano, K. & Mitsui, Y. (1996). J. Mol. Biol.255, 735–752. [DOI] [PubMed]
- Sonoki, T., Obi, T., Kubota, S., Higashi, M., Masai, E. & Katayama, Y. (2000). Appl. Environ. Microbiol.66, 2125–2132. [DOI] [PMC free article] [PubMed]
- Sugimoto, K., Senda, T., Aoshima, H., Masai, E., Fukuda, M. & Mitsui, Y. (1999). Structure, 7, 953–965. [DOI] [PubMed]
- Sugiyama, K., Senda, T., Narita, H., Yamamoto, T., Kimbara, K., Fukuda, M., Yano, K. & Mitsui, M. (1995). Proc. Jpn Acad. Ser. B Phys. Biol. Sci.71, 33–35.
- Vagin, A. & Teplyakov, A. (1997). J. Appl. Cryst.30, 1022–1025.