SUMMARY
Primate lentiviruses including HIV-1 have evolved the capacity to transduce terminally differentiated, non-dividing myeloid cells and as a consequence, these viruses establish persistent infections of tissue macrophage and microglia in the host. In contrast, non-dividing myeloid cells are refractory to infection by gammaretroviruses such as MLV. Here we present evidence that a cellular restriction is the obstacle to transduction of macrophage by MLV. Neutralization of the restriction by Vpx, a primate lentiviral protein previously shown to protect primate lentiviruses from a macrophage restriction, rendered macrophage permissive to MLV infection. Packaging of Vpx within MLV virions was sufficient to confer a lentivirus phenotype for MLV. We further demonstrate that this restriction prevents transduction of quiescent monocytes by HIV-1. Monocyte- HeLa heterokaryons were resistant to HIV-1 infection while heterokaryons formed between monocytes and HeLa cells expressing Vpx were permissive to HIV-1 infection. Encapsidation of Vpx within HIV-1 virions conferred the ability to infect quiescent monocytes. Collectively our results indicate that the relative ability of lentiviruses and gammaretroviruses to transduce non-dividing, myeloid-cells is dependent upon their ability to neutralize a cellular restriction and that this restriction shapes the association of lentiviruses with myeloid cell reservoirs in the host.
INTRODUCTION
A fundamental characteristic that distinguishes lentiviruses from simple gammaretroviruses is their capacity to infect non-dividing cells (reviewed in Suzuki and Craigie, 2007; Yamashita and Emerman, 2006). Primate lentiviruses such as HIV-1 are able to transduce non-dividing cells (Bukrinsky et al., 1992; Lewis et al., 1992) and this underscores their ability to transduce terminally differentiated, non-dividing cells including macrophage, microglia and dendritic cells both in vitro and in vivo (Gartner et al., 1986; Ringler et al., 1989; Weinberg et al., 1991). This property of lentiviruses has been harnessed through lentivirus vectors which are used for the transduction of non-dividing cellular targets including microglia and neurons (Balcaitis et al., 2005; Naldini et al., 1996). In contrast, gammaretroviruses transduce cells in mitosis and non-dividing cells (G1/S/G2) are refractory to gammaretrovirus transduction (Bieniasz et al., 1995; Lewis et al., 1992; Lewis and Emerman, 1994; Roe et al., 1993). Furthermore, although lentiviruses have evolved the ability to infect terminally differentiated, non-proliferating cells, quiescent cells (G0) are refractory to lentivirus transduction. This is best exemplified by observations made with myeloid-lineage cells. Studies conducted with HIV-1 demonstrate that peripheral blood monocytes, which are the undifferentiated precursor to tissue macrophage, are highly refractory to infection (Collman et al., 1989; Di Marzio et al., 1998; Eisert et al., 2001; Naif et al., 1998; Neil et al., 2001; Rich et al., 1992; Sonza et al., 1996). Permissivity to HIV-1 infection is coordinated to the state of monocyte differentiation (Sonza et al., 1996; Triques and Stevenson, 2004). Similarly, transduction of CD4+ T lymphocytes is cell cycle dependent; G0 CD4+T lymphocytes are inefficiently transduced by HIV-1 but become highly permissive to infection after entry into cell cycle (Stevenson et al., 1990; Zack et al., 1990).
The mechanisms underscoring the differential ability of gammaretroviruses and lentiviruses to transduce non-dividing myeloid cells as well as the block to transduction of quiescent monocytes by lentiviruses are not well understood. Cell transduction by gammaretroviruses and lentiviruses requires synthesis of viral cDNA and translocation of viral cDNA to the nucleus in order for viral cDNA to integrate into cellular DNA. Synthesis of viral cDNA and transport of viral cDNA to the cell nucleus occurs within the context of a large (160s) ribonucleoprotein reverse transcription/ preintegration complex, which contains viral reverse transcriptase as well as the viral integrase that catalyzes formation of the integrated provirus (Bowerman et al., 1989). Therefore, transduction of a non-dividing cell requires translocation of this complex across the nuclear envelope in order for viral cDNA to contact chromatin. One possible explanation for the differential ability of lentiviruses and gammaretroviruses to transduce non-dividing cells is that reverse transcription complexes of lentiviruses harbor nucleophilic determinants that direct its nuclear translocation whereas reverse transcription complexes of gammaretroviruses lack these determinants. A number of viral factors (reviewed in (Suzuki and Craigie, 2007; Yamashita and Emerman, 2006) have been implicated in promoting nuclear translocation of the lentiviral reverse transcription complex including a triple stranded viral DNA intermediate referred to as the central DNA flap (Zennou et al., 2000). Viral proteins including integrase (Bouyac-Bertoia et al., 2001), the Vpr/Vpx accessory proteins (Fletcher et al., 1996; Heinzinger et al., 1994), mature matrix proteins (Bukrinsky et al., 1993) or capsid proteins from the incoming virion (Yamashita et al., 2007) have been suggested to play a role in non-dividing cell infection by HIV-1. There is also biochemical evidence that lentiviral reverse transcriptases, unlike oncoretroviral reverse transcriptases, synthesize cDNA in the presence of low dNTP concentrations that are equivalent to those found in macrophage and this has been suggested to account for the differential ability of lentiviruses and gammaretroviruses to transduce non-dividing macrophage (Diamond et al., 2004). However, there is no consensus as to which, if any, of these viral factors are ultimately responsible for the inability of gammaretroviruses and the ability of lentiviruses to transduce non-dividing myeloid cells.
A different set of factors have been proposed to regulate infection of quiescent monocytes by lentiviruses. G0 monocytes have low intracellular dNTP levels (O'Brien et al., 1994; Triques and Stevenson, 2004) and this has been proposed to limit the efficiency of viral cDNA synthesis in these quiescent cells. The cytidine deaminase Apobec 3, which is a target of the viral accessory protein Vif, has been shown to influence the permissivity of quiescent lymphocytes and monocytes to HIV-1 infection (Chiu et al., 2005; Ellery et al., 2007; Peng et al., 2007; Peng et al., 2006). Apobec 3G is sequestered in an enzymatically active low- molecular-mass (LMM) ribonucleoprotein complex or in an enzymatically inactive high-molecular-mass (HMM) complex. The LMM complex, which is the exclusive form in quiescent cells, has been shown to restrict infection of quiescent monocytes by HIV-1 (Chiu et al., 2005; Ellery et al., 2007; Peng et al., 2006).
A number of studies have suggested that the accessory proteins Vpr and Vpx of primate lentiviruses have evolved to specifically promote infection of non-dividing myeloid-lineage cells (Balliet et al., 1994; Connor et al., 1995; Fletcher et al., 1996; Goujon et al., 2008; Heinzinger et al., 1994; Sharova et al., 2008; Srivastava et al., 2008). By generating heterokaryons between cells in which Vpx was dispensable for infection and primary macrophages in which Vpx is required for SIV infection, we demonstrated that macrophages harbor a dominant restriction and that this restriction is specifically counteracted by the accessory protein Vpx (Sharova et al., 2008). In the current study, we demonstrate that this restriction is an obstacle to transduction of terminally differentiated, non-dividing cells by gammaretroviruses. Vpx-mediated neutralization of the restriction rendered macrophage permissive to MLV infection. Furthermore, we present evidence that the ability of lentiviruses to transduce quiescent monocytes is regulated by this same restriction and that neutralization of the restriction in monocytes confers susceptibility to lentivirus infection. Collectively, our results suggest that the relative ability of lentiviruses and gammaretroviruses to transduce non-dividing, myeloid cells is governed primarily by their ability to neutralize a restriction that is present within these cells.
RESULTS
A dominant restriction limits MLV infection of macrophage
The majority of studies that have examined obstacles to infection of non-dividing cells by gammaretroviruses have been conducted with artificially growth-arrested cell lines. Whether similar blocks exist in natural, non-dividing cells such as macrophage has not been fully examined. Intriguingly, it has been reported that a low level transduction of human macrophage with gammaretrovirus vectors occurs at high multiplicities of infection suggesting that the block to infection is saturable (Jarrosson-Wuilleme et al., 2006). In order to gain further insight into the mechanism underlying the block to macrophage transduction by MLV, we compared the extent of viral cDNA synthesis and the efficiency of viral transduction in primary macrophage. Transduction efficiency of HIV-1 and MLV in primary macrophage was assessed relative to transduction efficiencies in HeLa cells which are permissive to both HIV-1 and MLV transduction. Macrophage were transduced by HIV-1 at a level comparable to that observed in HeLa cells (Figure 1A). This was reflected by increasing levels of viral cDNA synthesis in macrophage transduced with increasing levels of input virus (Figure 1B). In contrast, transduction of macrophage by MLV was highly inefficient as evidenced by the frequency of GFP-positive cells (Figure 1A) and levels of viral cDNA synthesis (Figure 1B). Therefore, the primary block to transduction of macrophage by MLV appeared to be at the level of reverse transcription. In agreement with a previous study (Jarrosson-Wuilleme et al., 2006), we observed a low level of transduction (2-3% GFP positive) of primary macrophage by MLV. While artificially growth arrested HeLa cells are refractory to transduction by MLV (Lewis and Emerman, 1994; Roe et al., 1993), the block to infection of those cells by MLV was unrelated to the reverse transcription block in terminally differentiated macrophages (Figure 1C). Levels of MLV cDNA in aphidocolin-treated HeLa cells were comparable to those in untreated HeLa cells and nuclear localization of viral cDNA (as indicated by 2-LTR circles that are formed in the nucleus) were also comparable. However, integration of MLV cDNA was inefficient in aphidocolin-treated HeLa cells (Figure 1C). Therefore, the block that was observed in an artificially growth arrested cell line was distinct from the block that occurs in natural non-dividing targets of lentivirus infection.
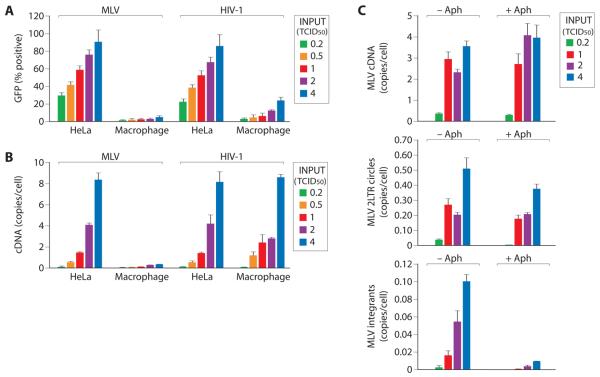
Figure 1. MLV infection of macrophage is blocked at, or prior, to reverse transcription of viral cDNA.
Terminally differentiated macrophage and HeLa cells were infected with MLV and HIV-1 variants expressing GFP at different levels of input virions. The frequency of GFP positive cells (A) and viral cDNA copies (B) were determined 48 hours post infection. (C) MLV infection of aphidicolin treated and untreated HeLa cells. Viral cDNA (upper two panels) and viral integrants (lower panel) were determined at different levels of input virus based on tissue culture infectious dose50 (TCID50) where 1 TCID50 is the amount of virus inoculum that yielded 50% transduction on HeLa cells. (error bars are s.d. of replicate samples from 3 independent experiments done on macrophage from different donors).
We have previously presented evidence that macrophage harbor a restriction that antagonizes HIV-1, HIV-2 and SIV at the level of reverse transcription and that the Vpx protein of HIV-2/SIVSM specifically overcomes this restriction (Sharova et al., 2008). In that study, the impact of the restriction on HIV-1 reverse transcription was revealed after the restriction was neutralized by Vpx and the efficiency of HIV-1 reverse transcription in macrophages was significantly enhanced (Sharova et al., 2008). Given that infection of macrophages by MLV was also being inhibited at the level of reverse transcription, we investigated whether the restriction that antagonizes lentivirus infection of macrophage may also be preventing infection of macrophage by MLV. We used a heterokaryon strategy that we previously adopted to demonstrate that Vpx countered a dominant restriction that was specifically expressed in macrophage (Sharova et al., 2008). Since HeLa cells are highly permissive to MLV infection, heterokaryons were generated between macrophage and HeLa cells and the susceptibility of the heterokaryons to MLV infection was assessed. When the fusogenic proteins of Newcastle Disease Virus (NDV) were expressed in HeLa cells, these cells readily fused with primary macrophage (Figure 2A). HeLa-macrophage heterokaryons (double-stained cells as indicated by the gate) were then sorted by FACS (Figure 2A, left panels). A FACS profile of sorted heterokaryons is shown (Figure 2A middle panel). Representative images of double-staining heterokaryons are shown (Figure 2A, right panels). Pre-sort images show one double-staining heterokaryon and two adjacent non-fused cells (DiO stained only) and one heterokaryon post-sort is shown. Because of the lipophilic nature of the dyes, fluorescence concentrates in lipid-rich regions of the cell. The block to MLV infection of macrophage was at the level of reverse transcription (Figure 1). Therefore, the ability of MLV to infect HeLa-macrophage heterokaryons was gauged by the relative levels of late MLV cDNA transcripts and 2-LTR circles which are formed only after completion of viral reverse transcription. While HeLa cells were permissive to MLV infection, macrophage and HeLa-macrophage heterokaryons were not permissive to MLV infection (Figure 2A). The restriction to lentivirus infection of macrophage is specifically counteracted by Vpx (Sharova et al., 2008). To determine whether the same restriction prevented MLV infection of macrophage, we examined the ability of Vpx to overcome the block to MLV infection of HeLa-macrophage heterokaryons. When Vpx was expressed in HeLa cells and those cells were allowed to fuse with macrophage, the resulting heterokaryons were rendered permissive to MLV infection (Figure 2B, right panels). In contrast, HeLa-macrophage heterokaryons not expressing Vpx remained refractory to MLV infection (Figure 2B). The expression of Vpx in HeLa cells did not increase their susceptibility to MLV infection (Figure 2C). Furthermore, the block imparted by aphidicolin treatment of HeLa cells was not released when Vpx was expressed in those cells (Figure 2C). Collectively, this data indicates that non-dividing macrophages harbor a dominant restriction that prevents MLV infection and Vpx overcomes the restriction. Furthermore, the block to MLV infection of non-dividing HeLa cells is distinct from that observed in macrophages and is not overcome by Vpx.
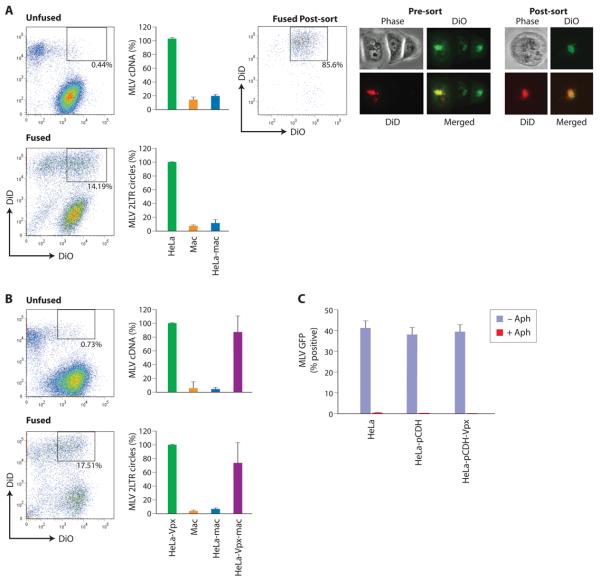
Figure 2. A restriction prevents transduction of macrophage by MLV.
Heterokaryons were formed between primary macrophage and between HeLa cells expressing fusogenic HN and F proteins of Newcastle Disease Virus (NDV). HeLa cells were stained with DiO (green) and macrophage were stained with DiD (red). Double-stained heterokaryons were sorted by FACS as indicated by the gate (A). FACS profile of heterokaryons post-sorting (fused post sort) is shown (middle panel) as are representative double staining heterokaryons pre-sort and post-sort (right panels). Because of the lipophilic nature of DiO and DiD, fluorescence concentrates in lipid-rich regions in the center of the cell rather than being evenly distributed throughout the cell. Susceptibility of HeLa-macrophage (HeLa-mac) heterokaryons to MLV infection was compared with infection levels in HeLa and in macrophage. Infection was gauged from the levels of late MLV cDNAs and 2-LTR circle cDNAs. Values were expressed relative to those obtained for HeLa cells (error bars are s.d. from 3 independent experiments). (B) Susceptibility of HeLa-macrophage heterokaryons to MLV infection was examined after expression of Vpx in HeLa cells. Double stained cells were sorted by FACS as indicated by the gate. MLV infection in HeLa-macrophage heterokaryons (HeLa-mac) and heterokaryons formed between macrophage and Vpx-expressing HeLa cells (HeLa-Vpx-mac) were gauged as outlined in A (error bars are s.d. of 3 independent experiments). (C) MLV infection of aphidicolin treated (+Aph) and untreated (−Aph) HeLa cells transfected with a Vpx expression vector (pCDH-Vpx) or an empty vector (pCDH).
Neutralization of the macrophage restriction confers permissivity to MLV infection
In HeLa-macrophage heterokaryons, neutralization of the restriction by Vpx rendered those heterokaryons permissive to MLV infection. Therefore, we examined whether neutralization of the restriction by Vpx would be sufficient to render macrophage permissive to MLV. We first examined whether introduction of Vpx into macrophage by wild-type SIV infection would render those macrophage susceptible to subsequent transduction by MLV. MLV transduction was gauged by levels of cDNA as detected by real-time PCR.). Infection of primary macrophage with increasing levels of wild-type SIV (PBj) led to a dose-dependent increase in the level of MLV transduction based on MLV cDNA synthesis (Figure 3A). Pre-infection of macrophage with a wild-type SIV but not a Vpx-deleted SIV also resulted in an increased ability of MLV to transduce macrophage as evidenced by MLV cDNA synthesis (Figure 3B,C) and expression of GFP from the MLV genome (Figure 3D). We have previously demonstrated that the restriction to infection of macrophage by lentiviruses can be overcome by Vpx from SIVPBj and HIV-2 but not Vpr of HIV-1 (Sharova et al., 2008). While Vpx alleles from SIVPBj and SIVmac239 enhanced infection of macrophages by MLV, no significant effect was observed with SIVagm Vpr (Figure 3C). Therefore, Vpx but no Vpr alleles of primate lentivirues appear to neutralize a restriction to lentivirus and gammaretrovirus infection of macrophage. Vpx also appeared to neutralize the restriction in cells that expressed Vpx since MLV transduction occurred predominantly in macrophage that had also been transduced by SIV (GFP expression, Figure 3E). We did not observe dsRed/GFP double positive cells in macrophages infected only with SIV (Figure 3E). Therefore, the presence of double positive cells was not simply due to bleeding of the GFP signal into the dsRed channel. The ability of wild-type SIV but not a Vpx-deleted SIV to render macrophage permissive to MLV transduction indicated that a restriction regulates the permissivity of primary macrophage to MLV transduction and that Vpx overcomes this restriction.
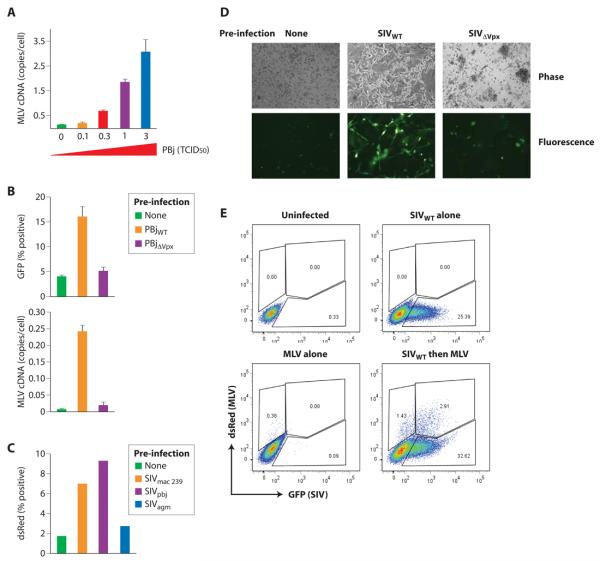
Figure 3. Vpx permits transduction of macrophage by MLV in trans.
(A) Vpx delivered to macrophage by wild-type SIV infection removes the block to synthesis of MLV cDNA in macrophage. Macrophage were initially infected with increasing titers of wild-type SIV and subsequently infected with MLV (4 TCID50) after 4 hours. Synthesis of MLV cDNA was assessed 48 hours after MLV infection. (B, C) Vpx but not Vpr is necessary for the ability of SIV to remove the block to macrophage transduction by MLV. Macrophage were infected by wild-type or Vpx-deleted SIV and subsequently infected by MLVGFP (4 TCID50) after 4 hours. In C, macrophages were infected with the indicated SIV infectious clones and then with MLV. The efficiency of MLV transduction was assessed after MLV infection. A representative field of macrophage transduced by MLV-GFP is shown in D. (E) Transduction of macrophage by MLV occurs primarily in SIV-infected macrophage. SIVGFP-infected macrophage were transduced with MLVdsRed and frequencies of coinfected cells was evaluated by FACS. FACS profiles of uninfected macrophage, MLV-transduced macrophage without prior SIV infection (MLV alone) or SIVWT without subsequent MLV infection (SIVWT alone) served as controls.
Packaging of Vpx within MLV virions confers a lentiviral phenotype
During lentivirus infection of macrophage, the restriction is neutralized by Vpx proteins that are encapsidated within the virus particle (Sharova et al., 2008). Therefore, we examined whether packaging of Vpx within MLV virions would be sufficient to confer upon MLV a lentiviral phenotype, i.e., the ability to transduce macrophage. The p6 domain of lentiviral gag proteins contains determinants for encapsidation of Vpr/Vpx proteins (Accola et al., 1999; Pancio and Ratner, 1998; Paxton et al., 1993; Wu et al., 1994). We fused the p6 domain of SIV gag to the C-terminus of the MLV gag protein (Figure 4A). Transfection of an MLV packaging cell line with plasmids expressing chimeric MLV gag-SIV p6 proteins, a Vpx expression vector and a VSV-G envelope-expression vector resulted in the production of VSV-G-pseudotyped, chimeric MLV virions containing Vpx. The presence of the VSV-G envelope bypassed the requirement for the presence of MLV receptor molecules on macrophages. Specific packaging of Vpx into MLV particles containing a chimeric gag-p6 domain was confirmed by Western blotting (Figure 4B). In contrast, MLV virions derived from a Vpx-expressing MLV packaging line containing wild-type MLV gag (lacking SIV p6) did not package Vpx proteins (Figure 4B).
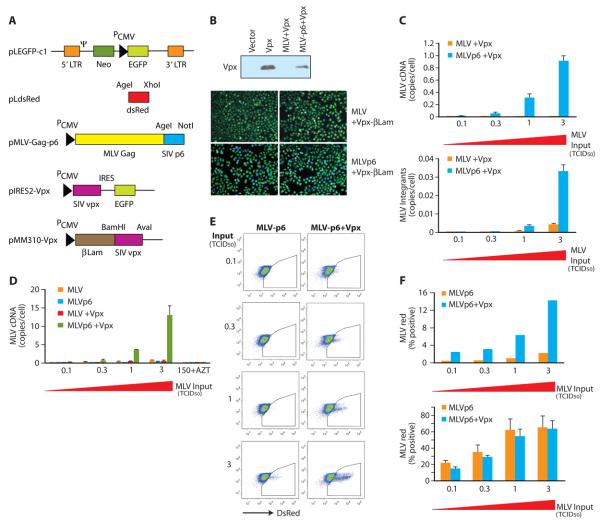
Figure 4. MLV virions encapsidating Vpx exhibit a lentiviral phenotype.
(A) A schematic of vectors used for expression of Vpx and chimeric MLV gag proteins containing the p6 domain of SIV gag which harbors the Vpx/Vpr packaging determinant. (B) Packaging of Vpx within MLV virions harboring an SIV gag p6 domain. Upper panel, packaging of Vpx within MLV virions containing or lacking an SIV p6 gag domain was examined by Western blotting with a Vpx-specific antibody. Lower panels, Vpx-β-lactamase fusion proteins were packaged in MLV variants containing or lacking the SIV gag p6 domain and β-lactamase activity was examined following infection of HeLa cells loaded with the β-lactamase substrate CCF2. (C) Packaging of Vpx within chimeric MLV virions containing SIV gag p6 (MLVp6) removes a block to reverse transcription in macrophage. Macrophage were infected with increasing concentrations of MLVp6 with or without encapsidated Vpx and viral cDNA synthesis (late cDNA, upper panel) and integration (lower panel) was assessed. (D-F) A p6 encapsidation signal and Vpx is required for MLV transduction of macrophage. MLV cDNA synthesis (D) was examined after infection of macrophage with MLV and MLVp6 variants with and without Vpx. Infections carried out in the presence of AZT verified de novo synthesis of MLV cDNA, (E). Packaging of Vpx permits transduction of primary macrophage by MLV. Macrophage were infected with increasing titers of chimeric MLV variants with and without Vpx as in C. Transduction was gauged by expression of dsRed from the MLV transgene. Frequencies of MLV transduction (dsRed expression) on macrophage (upper panel) and HeLa (lower panel) are indicated in panel F.
We next examined the functionality of the p6 domain within the chimeric MLV gag protein by its ability to package a Vpr-β-lactamase fusion protein within virions (Cavrois et al., 2002). Transfer of the β-lactamase-Vpr fusion protein into HeLa cells was then detected by enzymatic cleavage of CCF2 which is a fluorescent substrate of β-lactamase. Cleavage of CCF2 changes its fluorescence emission spectrum from green to blue which can be visualized by fluorescence microscopy. Infection of CCF2-loaded HeLa cells by chimeric MLV harboring a Vpr-β-lactamase fusion protein resulted in CCF2 cleavage as evidenced by the appearance of blue cells under fluorescence microscopy (Figure 4B). This was not the case for CCF2-loaded HeLa cells that had been infected with MLV harboring a wild-type gag protein (Figure 4B). Packaging of Vpx within MLV virions containing chimeric gag proteins markedly increased their ability to transduce primary macrophage both in terms of viral cDNA synthesis and integration (Figure 4C,D) and in terms of red fluorescent protein expression from the MLV genome (Figure 4E,F). The chimeric MLV variant containing the SIV gag p6 domain required Vpx for infection of macrophage since in the absence of Vpx, this chimeric MLV did not transduce macrophage (Figure 4D). Furthermore, MLV cDNA that was detected in these macrophages was synthesized de novo and was inhibited in the presence of AZT (Figure 4D). Transduction efficiencies of chimeric MLV particles containing Vpx (∼15% at high M.O.I.) approached those typically observed for lentivirus-based vectors (Figure 4F, upper panel). The transduction efficiency of MLV with or without packaged Vpx were similar when gauged on HeLa cells (Figure 4F, lower panel). Collectively, these results indicate that Vpx is sufficient to render primary macrophage permissive to MLV infection and that a restriction is the obstacle to MLV transduction of non-dividing macrophage.
The resistance of quiescent monocytes to lentivirus transduction is governed by a restriction
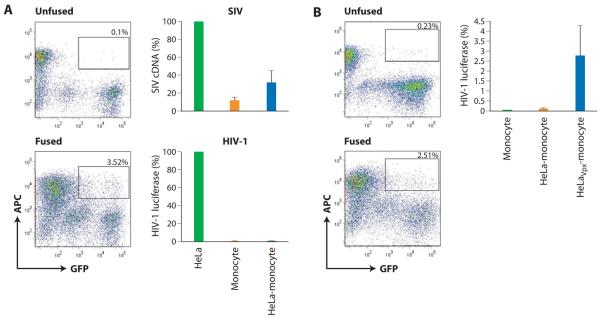
Circulating peripheral blood monocytes are highly refractory to lentivirus infection in vitro and infection is blocked at an early post-entry step (Collman et al., 1989; Naif et al., 1998; Neil et al., 2001; Rich et al., 1992; Sonza et al., 1996; Triques and Stevenson, 2004). Susceptibility to infection occurs only upon differentiation of monocytes to macrophage (Munk et al., 2002; Sonza et al., 1996; Triques and Stevenson, 2004). We first investigated whether the fusion of HeLa cells with monocytes would result in heterokaryons permissive to HIV-1 infection. To generate HeLa-monocyte heterokaryons, we exploited the fusogenic properties of Sendai Virus (hemmaglutinating virus of Japan; HVJ) envelope proteins. The susceptibility of those heterokaryons to HIV-1 and to SIV infection was then examined. SIV infection was gauged from the level of late cDNAs and HIV-1 infection was determined by luciferase activity expressed from the HIV-1 genome (values were expressed as percentages of those obtained with HeLa cells). As with unfused monocytes, HeLa-monocyte heterokaryons were highly refractory to transduction by HIV-1 (Figure 5A). Intriguingly, both primary monocytes and HeLa-monocyte heterokaryons were permissive to transduction by SIV (Figure 5A). To examine whether the ability of SIV to transduce primary monocytes was attributable to Vpx, we generated heterokaryons between monocytes and between HeLa cells that expressed the Vpx protein (Figure 5B). In this case, the permissivity of HeLa-monocyte heterokaryons to HIV-1 transduction was increased by Vpx (Figure 5B) whereas HeLa-monocyte heterokaryons not expressing Vpx remained refractory to HIV-1 transduction (Figure 5B). Since Vpx does not increase the efficiency of HIV-1 infection in HeLa cells, this result was not due to infection of unfused HeLa cells. Therefore, we conclude that heterokaryons formed between nonpermissive monocytes and permissive HeLa cells are nonpermissive due to the presence of a dominant restriction and that this restriction is overcome by Vpx. We titered the amount of Vpx needed to rescue SIVΔvpx infection in macrophage and observed that even a small amount of trans-packaged Vpx can counter the restriction present in macrophage (Figure S1). Vpx is packaged in molar amounts equivalent to gag proteins (Henderson et al., 1988). Assuming ∼2000 gag molecules per virion (Arthur et al., 1992), and assuming uniform Vpx: gag stoichiometry in each viral particle, Vpx packaged at ∼10% of wild-type levels still rescued a ΔVpx virus (Figure S1) suggesting that as few as 20 Vpx molecules can counteract the restriction.
Figure 5. Transduction of primary monocytes by HIV-1 is blocked by a restriction.
Heterokaryons were formed between primary monocytes and HeLa cells using HVJ envelope fusion kit. (A) FACS analysis of HeLa-monocyte heterokaryons (left panels). HeLa cells expressed GFP and macrophages were stained with an APC-conjugated antibody to CD14. Double-stained cells were sorted as indicated by the gate. SIV infection was gauged from the levels of late cDNAs and HIV-1 infection was gauged from luciferase activity (right panels). Values were expressed relative to those obtained for HeLa cells (error bars are s.d. of 4 independent experiments.) (B) Vpx renders HeLa-monocyte heterokaryons permissive to HIV-1 infection. Heterokaryons were formed between primary monocytes and HeLa cells expressing Vpx as described in A). Susceptibility of HeLa-monocyte heterokaryons to HIV-1 infection was examined after expression of Vpx in HeLa cells. FACS analysis of HeLa-Vpx-monocyte heterokaryons is shown on the left panels. Double-stained cells were sorted as indicated by the gate. Infection of monocytes and infection of HeLa-monocyte heterokaryons with and without Vpx was gauged by luciferase activity (error bars are s.d. from 2 independent experiments).
Vpx renders primary monocytes permissive to HIV-1 transduction
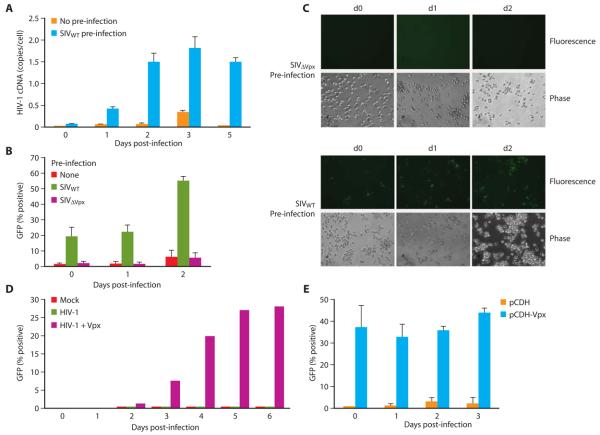
Since Vpx was sufficient to render HeLa-monocyte heterokaryons permissive to HIV-1 infection (Figure 5), we next examined whether Vpx was sufficient to render monocytes susceptible to HIV-1 transduction. Since monocytes were partially permissive to wild-type SIV transduction (Figure 5A), Vpx was introduced into monocytes by wild-type SIV infection and those monocytes were subsequently examined for permissivity to HIV-1. HIV-1 infection of monocytes was blocked at, or prior to, reverse transcription as evidenced by an absence of viral cDNA in HIV-1 infected monocytes (Figure 6A). SIV infection rendered monocytes highly permissive to subsequent HIV-1 infection as evidenced by the extent of HIV-1 cDNA synthesis (Figure 6A). In contrast, monocytes that had not been pre-infected with SIV remained refractory to HIV-1 (Figure 6A). Furthermore, monocytes infected with a wild-type SIV (SIVWT) but not a Vpx-deleted variant (SIVΔVpx) could be transduced by HIV-1 as evidenced by expression of GFP from the HIV-1 genome (Figure 6B, C). In some cases, the impact of SIV infection on monocyte permissivity was rapid and evident within one day of monocyte seeding (Figure 6B). We have previously demonstrated that packaging of Vpx within HIV-1 virions is sufficient to counteract a restriction that is expressed by differentiated macrophage (Sharova et al., 2008). Similarly, packaging of Vpx within HIV-1 virions (Figure 6D) or in a HIV-1 lentivirus vector (pCDH-Vpx, Figure 6E) markedly increased the efficiency of transduction in primary monocytes. We also examined whether the impact of the restriction was reversible. We speculated that following infection of macrophages by a Vpx-minus virus, we might be able to rescue the infection by subsequent introduction of Vpx. At various intervals following infection by a Vpx-minus virus (containing a GFP transgene) cells were super-infected by wild-type or ΔVpx SIV variants. The ability to rescue the initial SIVΔVpx infection was gauged by PCR using primers specific for GFP. We observed that SIVΔVpx GFP reverse transcription could be restored at least 5 hours later by a wild-type virus. Since this is in the time frame required for uncoating to occur, it suggests that the restriction might act subsequent to uncoating.
Figure 6. Vpx counteracts a monocyte restriction to HIV-1 infection in trans.
(A) Infection of monocytes by wild-type SIV removes a reverse transcription block to subsequent infection by HIV-1. SIVWT-infected monocytes were subsequently infected (4 hours later) by HIV-1 on the indicated intervals and levels of HIV-1 cDNA synthesis was gauged 48 hours post HIV-1 infection (B) Prior infection by wild-type but not Vpx-deleted SIV renders primary monocytes permissive to subsequent transduction by HIV-1. Monocytes were infected as in A. Transduction of HIV-1 (based on GFP expression) was assessed 72 hours post HIV-1 infection. Representative fields of primary monocytes following transduction by HIV-1-GFP are shown on C. (D) HIV-1 virions encapsidating Vpx efficiently transduce primary monocytes. Monocytes were infected with HIV-1 GFP variants in which Vpx was packaged. Levels of transduction (% GFP-positive monocytes) was determined at the indicated intervals post monocyte infection. (E) Transduction of monocytes with an HIV-1 lentivirus vector in which Vpx was or was not packaged. Monocytes were infected at the indicated intervals and GFP expression examined 72h post-infection (error bars are s.d. of replicate samples from 3 independent experiments done on macrophage from different donors).
Vpx affects monocytes permissivity independent of Apobec 3 or differentiation status
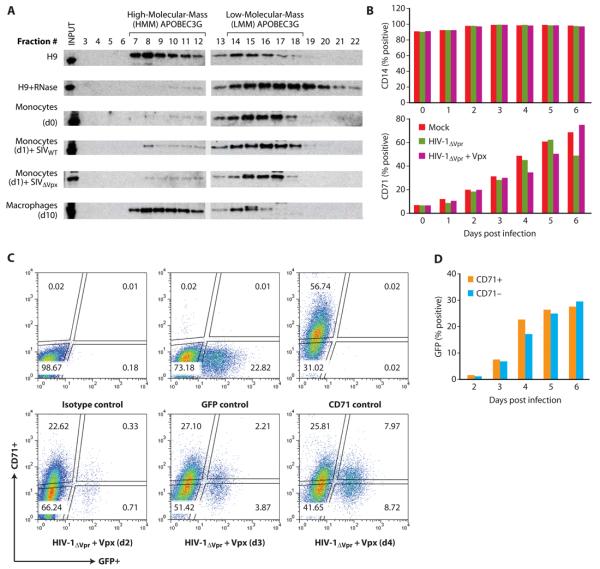
Previous studies have demonstrated that Apobec 3G exists predominantly in a low-molecular-mass (LMM) complex in quiescent monocytes and that this complex restricts infection of monocytes by HIV-1 (Chiu et al., 2005; Ellery et al., 2007; Peng et al., 2007). Differentiation of monocytes to macrophage redistributes Apobec 3G into a high-molecular-mass (HMM) complex and this correlates with the permissivity of macrophage to HIV-1 infection. To investigate the possibility that Vpx rendered monocytes permissive to infection by causing a shift in Apobec 3G from LMM to HMM complexes, we compared the distribution of Apobec 3G in uninfected monocytes and in monocytes infected with wild-type and Vpx-deleted SIV. As published previously (Chiu et al., 2005), Apobec 3G was sequestered primarily in a HMM complex in H9 cells and in differentiated (day 10) macrophage (Figure 7A). RNase-treatment of HMM complexes from H9 cells led to the formation of LMM Apobec 3G complexes (Figure 7A). In undifferentiated (day 0) monocytes, Apobec 3G was sequestered primarily in a LMM complex (Figure 7A). Infection of monocytes by wild-type or Vpx-deleted SIV did not noticeably alter distribution of Apobec 3G between LMM and HMM complexes (Figure 7A).
Figure 7. Vpx renders monocytes permissive to HIV-1 infection without inducing monocyte differentiation or Apobec 3G distribution.
(A) Distribution of Apobec 3G between LMM and HMM nucleoprotein complexes in undifferentiated (d0) monocytes, differentiated (d10) macrophage and SIV-infected monocytes. Distribution of Apobec 3G between H9 cell-derived HMM and LMM complexes before and after RNase treatment is shown for comparison. (B) Vpx does not affect differentiation status of monocytes in culture. Fresh monocytes were infected with HIV-1ΔVprGFP that had or had not packaged Vpx and the infection levels in monocyte/macrophage (CD14+) and differentiated monocyte (CD71+) subsets was determined at the indicated intervals post infection by FACS. (C) HIV-1 with encapsidated Vpx equally transduces undifferentiated (CD71−) and differentiated (CD71+) monocyte populations. Monocytes were infected with HIV-1 in which Vpx had been packaged (lower three panels) and the frequencies of infected (GFP positive) CD71+ macrophage and CD71− monocytes were determined by FACS. Upper three panels depict uninfected controls. The frequency of HIV-1 infection in CD71+ and CD71− cells at different intervals post- infection is shown in D. Supporting Information
A second possibility was that Vpx did not permit transduction of undifferentiated monocytes but rather, rapidly induced differentiation to macrophage that are permissive to HIV-1 infection. Typically, peripheral blood monocytes do not begin to express macrophage differentiation markers (CD71) until after 2-3 days in culture and infection of monocytes by HIV-1 either with or without Vpx did not have an effect on temporal expression of CD71 (Figure 7B). Nevertheless, it was possible that HIV-1 transduction was restricted to a small percentage of CD71+ macrophage in the culture. To examine this, frequencies of infected monocytes (CD71−) and macrophage (CD71+) were examined by FACS following infection with a GFP-expressing HIV-1 variant in which Vpx had been packaged. We compared GFP expression in CD71+ and CD71− cells at different intervals post-infection. As the frequency of GFP+ cells increased, there was no apparent bias to an increased frequency of CD71+/GFP+cells (Figure 7C). Indeed, the frequencies of infected CD71− monocytes at day 2, 3 and 4 post infection paralleled those for infected CD71+ cells (Figure 7C). In an independent experiment (Figure 7D), equivalent transduction of CD71+ and CD71− by HIV-1 over 6 days post infection was maintained. Collectively, these results indicate that Vpx directly renders undifferentiated monocytes permissive to HIV-1 transduction without inducing their differentiation.
DISCUSSION
Why non-dividing myeloid cells are permissive to lentivirus infection, yet nonpermissive to gammaretrovirus infection, is a long-standing question in the field. Our studies indicate that, a cellular restriction is the obstacle to transduction of terminally differentiated macrophage by MLV and that when the restriction is neutralized by the primate lentiviral Vpx protein, macrophage become permissive to MLV. Current models, based primarily on studies with artificially growth arrested fibroblast cell lines, suggest that the relative abilities of gammaretroviruses and lentiviruses to traverse the nuclear envelope dictates the differential abilities of these viruses to transduce non-dividing cells (reviewed in Yamashita and Emerman, 2006). However, we observed that MLV infection of artificially growth arrested HeLa cells was blocked at the level of integration and not viral cDNA synthesis or nuclear import of viral cDNA. This block was mechanistically distinct to the block we observed in natural non-dividing macrophages where MLV transduction was inhibited either prior to, or at the level of reverse transcription of viral cDNA. When the block to reverse transcription in macrophages was alleviated by Vpx, MLV integration and gene expression occurred. Therefore, the differential ability of lentiviruses and gammaretroviruses to transduce non-dividing macrophage is dictated by the degree to which they are sensitive to a restriction that acts prior to or at the level of reverse transcription.
Although our studies provide insight into mechanisms that restrict gammaretrovirus infection of non-dividing myeloid cells, there still remains the question as to how viral genomes access the nuclear compartment. Packaging of Vpx within MLV particles removed a block to reverse transcription and was sufficient to permit transduction of terminally differentiated macrophage. This indicates that if conditions for viral cDNA synthesis are met, subsequent events including synthesis, nuclear import and integration of viral cDNA and de novo gene expression occur in non-dividing macrophage following both HIV-1 and MLV infection. Therefore, presumably, the ability to traverse the nuclear envelope appears to be an intrinsic property of gammaretroviruses and lentiviruses rather than a characteristic that distinguishes these viruses. Models invoking a nuclear import role for Vpr/Vpx proteins have been supported by the fact that these proteins exhibit a nuclear localization (Di Marzio et al., 1995; Lu et al., 1993; Yao et al., 1995). While our data argues against the possibility that nuclear access is blocked during MLV infection of non-dividing macrophages, it is possible that the restriction is located in the nucleus and that Vpx must localize to the nucleus in order to counteract the restriction.
We previously demonstrated (Sharova et al., 2008) that infection of macrophage by HIV-1 is influenced by a restriction and that this restriction is sensitive to neutralization by Vpx but not SIVSM Vpr or HIV-1 Vpr. Here, we demonstrate that Vpx but not Vpr alleles of primate lentiviruses enhance infection of macrophages by MLV. All primate lentiviruses encode a Vpr protein. The vpx gene of the HIV-2 group, which includes HIV-2, SIVsmm and SIVmac, arose by duplication of the vpr gene within this group (Sharp et al., 1996; Tristem et al., 1992), that diverged from the other primate lentiviral groups around 200 years ago (Tristem et al., 1992). While Vpx represents a duplication, it does not share all the functional properties of Vpr. Vpr induces cell cycle arrest whereas Vpx does not (Fletcher et al., 1996). Conversely, the ability to neutralize a restriction in myeloid cells restriction is governed by Vpx but not Vpr. Presumably, this activity was manifest by the ancestral Vpr gene but, for unknown reasons, has been lost in the HIV-1 and SIVagm groups. It is possible that loss in the ability to counteract the myeloid cell restriction was compensated for by acquisition of partial resistance to the restriction as in the case of HIV-1.
Our studies further implicate a restriction as the obstacle to infection of quiescent monocytes by lentiviruses. It is likely that this same restriction antagonizes HIV-1 infection in monocytes and in macrophage. However, the degree to which HIV-1 is restricted in monocytes and macrophage differs considerably. In the absence of Vpx, HIV-1 still has the ability to transduce macrophage to some degree. Nevertheless, the efficiency with which HIV-1 transduces macrophage is greatly increased by Vpx. Therefore, while infection of macrophage by HIV-1 is antagonized by a restriction, this restriction is not sufficient to completely block transduction of these cells by HIV-1. In contrast, monocytes are totally refractory to HIV-1 infection in the absence of Vpx. Therefore, monocytes can be considered fully nonpermissive and macrophage semipermissive to HIV-1 transduction. The degree of permissivity might relate to the levels at which the restriction is expressed as monocytes differentiate to macrophage. A similar situation is seen with Apobec 3G in that some cell lines are semipermissive and that levels of Apobec 3G are insufficient to completely block virus transduction (Sheehy et al., 2002). In addition, because of low dNTP levels in monocytes, reverse transcription is inefficient in these cells (O'Brien et al., 1994). Since the restriction acts primarily at the reverse transcription step, it is possible that the restriction is able to more effectively perturb cDNA synthesis in an environment that is suboptimal for reverse transcription. Previous studies (Wolfrum et al., 2007) have also demonstrated that SIV is more efficient than HIV-1 in transducing primary monocytes and that deletion of Vpx prevented monocyte transduction by SIV. In that study, a HA-Vpx fusion protein, when packaged within HIV-1 virions, did not confer upon HIV-1 the ability to transduce monocytes. However, we have found that tags fused either to the N or C-terminus of Vpx abrogates its biological activity even though the fusion protein is competent for encapsidation. In this study, we demonstrate that, in its native form, Vpx is necessary and sufficient to permit transduction in macrophage by MLV and transduction of quiescent monocytes by HIV-1.
While the identity of the restriction that is counteracted by Vpx is as yet unidentified, it exhibits unique characteristics as compared to other known antiviral restrictions. Viral Vif and Vpu proteins that neutralize the antiviral restrictions Apobec 3G and tetherin/BST2 respectively, carry out their function in the virus producing cell (reviewed in (Malim and Emerman, 2008). Although some Vif is packaged within virions, there is no evidence that packaged Vif has a functional role in viral infection. By comparison, the ability of Vpx to neutralize the myeloid cell restriction appears to require that it is packaged within virions. Indeed, Vpx protein that was packaged into virions effected a durable removal of the block to subsequent infection by a restricted virus. This phenomenon is unprecedented in the arena of retroviral restrictions and suggests that the restriction has an extremely low turnover rate and takes a considerable time to recover after it has been neutralized by Vpx. Previous demonstrations that the biological activity of Vpx required association with damaged DNA binding protein 1 (DDB1) (Sharova et al., 2008, Srivastava et al., 2008) has parallels with viral proteins that circumvent the interferon response. For example, the v protein of paramyxovirus simian virus 5, requires DDB1 to promote targeted degradation of STAT1 (Andrejeva et al., 2002). However, of the various strategies used by viruses to impair the interferon response, all operate after initial infection and de novo expression of the viral defense protein. In this way, viruses prevent the infected cell from subsequently mounting an interferon response that would prevent surrounding cells from being infected by progeny virus. In the case of Vpx, the restriction must be neutralized by virion-associated Vpx in order for establishment of infection to occur. This indicates that the restriction is constitutively active in macrophage as opposed to being induced upon infection.
Our study underscores the powerful degree to which restrictions shape lentivirus biology. Primate lentiviruses exhibit tropism for macrophage lineage cells and reservoirs of tissue macrophage are evident in the gut, lung, lymph nodes and in the CNS (reviewed in (Alexaki et al., 2008; Carter and Ehrlich, 2008; Gonzalez-Scarano and Martin-Garcia, 2005). Tropism is dictated primarily by the expression of specific coreceptor molecules (mainly CCR5) on macrophage that permit virus binding and entry (reviewed in Gorry et al., 2005). Our study reveals a second level of tropism that is manifest post-entry and our findings would suggest that the ability of primate lentiviruses, and perhaps non-primate lentiviruses as well, to establish reservoirs in myeloid lineage cells is dependent upon their ability to counteract a myeloid cell-specific restriction. Given the potency with which the restriction antagonizes primate lentivirus infection, identification of the restriction itself, as well as pharmacologic agents which harness restrictions within macrophage, are important objectives.
EXPERIMENTAL PROCEDURES
Plasmids
The retroviral delivery vector pLEGFP-C1 contains MLV-derived retroviral elements along with a CMV promoter-driven EGFP gene (Clontech). Pseudotyping MLV and HIV-1 with VSV-G envelope involved cotransfection with a VSV-G expression plasmid. pMD-G (Naldini et al., 1996). pNL4-3.GFP contains the HIV-1 molecular clone NL4-3 with GFP in place of nef. pNL4-3.Luc plasmid contains luciferase reporter gene in place of envelope. The EGFP cassette in the expression vector pLEGFP-C1 was swapped with dsRed to obtain MLV with dsRed reporter expression (pLdsRed). The SIV clones were derived from SIV PBj (Fletcher et al., 1996). pMLV-Gagp6 was generated by replacing the RFP cassette in pMLV-Gag-RFP (Addgene plasmid 1814 obtained from Dr. W. Mothes (Sherer et al., 2003) ) with p6 amplified from SIVsm. The Vpx expression vector has been described previously (Sharova et al., 2008).
Cells and Viruses
Human monocytes were obtained from healthy donors by countercurrent centrifugal elutriation (Gendelman et al., 1988). 293T and HeLa cells were maintained in DMEM containing 10% FBS. Pseudotyped MLV (MLV-G) stocks were obtained by transfecting retroviral-packaging 293A cells with pLEGFP-c1 and pMD-G. Virus particles in culture supernatants were harvested after 24 and 48 hours, passed through 0.45 mm filter and concentrated by ultra-centrifugation. Vpx was packaged in MLV by co-transfecting 293A cells with pMD-G, pLdsRed , MLV-Gagp6 and Vpx expression vectors. Control virus was made with the same plasmids excluding MLVGagp6. Similarly, VSV-G pseudotyped HIV-1 (HIV-G) was prepared by transfecting 293T cells with pNL43-GFP and pMD-G. The viruses were titered by transducing HeLa or TZM-bl cells with increasing virus inputs followed by flow cytometry analysis of GFP-positive cells. One tissue culture infectious dose50 (TCID50) is the amount of transfected culture supernatant that generated ∼50% GFP-positive HeLa cells after 48 hours post-infection. Pseudotyped SIVsm viruses were obtained by transfecting 293T cells with a PBj1.9 molecular clone with (WT) or without Vpx (X2) (Fletcher et al., 1996) along with pMD-G. All virus stocks were treated with DNaseI (Worthington, NJ, USA) to remove residual transfection DNA. In all experiments, the SIVsmm-PBj strain has been used unless specified otherwise.
Infection Assays
HeLa cells as well as macrophage were infected with increasing virus inputs (TCID50) of HIV1-G and MLV-G. After 4 hours, cells were washed with fresh medium and incubated at 37°C for the remainder of the experiment. Pre-infection studies were performed by first infecting macrophages with pseudotyped SIVWT or SIVΔVpx variants and 4 hours later, the cells were infected with MLV-G (4 TCID50) for another 4 hours before washing cells with fresh medium. After 42-72 hours, the numbers of GFP/dsRed cells were quantitated by flow cytometry.
Analysis of Viral Infection by qPCR
Infected cells were washed with PBS before harvesting samples for DNA analysis. Total DNA was extracted from infected cells by a DNeasy kit (Qiagen). Quantitative analysis of MLV cDNA intermediates is as described (Bruce et al., 2005). PCR primers and probes for MLV include primers OJWB45 and OJWB48 for late MLV transcripts; OJWB45 and OJWB46 for 2LTR cDNA; MLV prb for cDNA detection (Bruce et al., 2005). PCR conditions for amplification of SIV and HIV-1 cDNAs are as described previously (Sharova et al., 2008). Copy number estimates of cDNA and 2LTR circles were determined on a ABI Prism 7500 fast machine. Integrants were quantitated by Alu-LTR real time PCR as described by Brussel & Sonigo (Brussel and Sonigo, 2003). Briefly, first PCR was done for 12 cycles using Alu primers and LTR-specific primer, tagged with lambda sequence. The PCR product was then diluted 10-fold and was used as template for a quantitative nested PCR using lambda primer and a LTR specific reverse primer. The number of cell equivalents in DNA lysates from HeLa cells, monocytes, macrophages and heterokaryons was determined by PCR using CCR5-specific primers (Hatzakis et al., 2000). The real-time PCR analysis from each sample was carried out in duplicate wells and most of the values shown in the figures are average of independent experiments using macrophage from at least 3 different donors.
APOBEC3G analysis
H9 cells, monocytes or macrophage were washed twice with PBS and incubated with lysis buffer containing 50 mM HEPES pH 7.4, 125 mM NaCl, 0.2% NP-40 and EDTA-free protease inhibitor cocktail (Roche). Cell lysates were clarified by centrifugation at 14000 rpm for 30 min at 4°C (Microfuge 22R, Beckman Coulter). Cleared cell lysates were quantitated (Protein assay kit, Bio-Rad) and analyzed by Fast Performance Liquid Chromatography (FPLC). For RNase treatment of HMM complexes from H9 cells, cell lysates was incubated with 50 μg/ml RNase A (Roche, DNase-free) at room temperature for 1 hour before analysis by FPLC. FPLC was run on an ÄKTAtm FPLC using a Superose 6 10/300 GL gel filtration column (GE healthcare). The running buffer contained 50 mM HEPES pH 7.4, 125 mM NaCl, 0.1% NP-40, 1mM DTT and 10% Glycerol. Fraction size was set at 1 ml. 20 μl of each fraction was boiled with Laemmli's SDS-sample buffer (6x reducing, BOSTON Bioproducts) and loaded onto a 10% SDS-PAGE. Proteins were transferred to nitrocellulose membranes and blotted with rabbit anti-Apobec 3G antibody (Courtesy of Dr. Tariq Rana) using a TROPIX CDP-star system (PerkinElmer).
FACS and macrophage immunophenotyping
Expression of CD14, CD71, or GFP/dsRed in monocytes/macrophage was monitored by flow cytometry. Cells were collected day 0 to day 6 post-infection and washed twice with buffer (PBS containing 0.1% fetal bovine serum and 2 mM EDTA). The washed cells were incubated with an antibody mixture containing PE conjugated anti-human CD14 (BD Biosciences) and APC conjugated anti-human CD71 (BD Biosciences) for 40 min. Cells were rinsed twice with washing buffer and fixed with 1% paraformaldehyde. Fixed cells were analyzed by cell flow cytometry analysis using a FACSCalibur System (BD Biosciences) and analyzed with Flowjo software (Tree Star, Inc). The percentage of infected CD71− monocytes and CD71+ macrophage were determined from the percentages of GFP+/CD71− or GFP+/CD71+ cells, respectively.
Cell fusion
Hela-macrophage fusion was achieved using paramyxovirus hemagglutinin-neuraminidase (HN) protein and fusion (F) proteins as described (Sharova et al., 2008). Briefly, HeLa cells were transfected with pCAGGS-HN and pCAGGS-F expression vectors encoding HN and F proteins of Newcastle disease virus (NDV). Sixteen hours post-transfection, HeLa cells were stained with 1.7 μM DiO and mixed with macrophages stained with 0.85 μM DiD (Molecular Probes) in a ratio of 1:2 and plated in 100 mm dishes. After overnight incubation, cells were infected with MLV for 40 h. Cell sorting was performed with a FACSAria flow cytometer using the FACSDiva software (Becton Dickinson). Double-stained cells were sorted and total DNA was isolated using a DNeasy Blood and Tissue Kit (Quiagen) and analyzed by real time PCR assay for late MLV cDNA and 2LTR circles. Hela-monocyte fusion was achieved using a GenomeONE-CFEX HVJ envelope fusion kit (Cosmo Bio Co., Ltd.). Manufacturer's instructions for fusion in suspension were followed. Briefly, GFP-expressing HeLa were mixed with monocytes (ratio 1:6) and incubated in the presence of HVJ-E suspension (1.25 ul/1× 106 cells) on ice for 5 min and subsequently at 37°C for 15 min. Cells were plated in 100 mm dishes and infected with HIV-1NL4-3 Luc or SIVWT for 40 hours. Prior to cell sorting, cells were stained with an APC-conjugated antibody to CD14 (BD Biosciences). Heterokaryons were sorted based on GFP and APC double staining. HIV-1NL4-3 Luc infection was measured by quantifying luciferase activity and SIVWT infection was analyzed by real time PCR assay for late cDNA and 2LTR circles.
Supplementary Material
Figure S1: Dose-dependent ability of packaged Vpx to counteract a macrophage restrictionPseudotyped SIVΔVpx was produced in 293T cells by co-transfecting ΔVpx PBj1.9 proviral DNA with pMD-G and increasing amount of a Vpx expression vector. (A) The amount of packaged Vpx in the purified virions was determined by western blotting with a Vpx antibody and normalization with p27 by densitometry. The integrated density was measured by Scion Image software. (B) The infectivity of these viruses in primary macrophage was determined from quantitation of SIV 2LTR cDNA 24 and 48 h post-infection. PBjWT was used as positive control for this experiment.
Figures S2: Reversibility of restrictionMacrophage were first infected with SIVΔvpx-GFP. After 0, 1, 3, 5, 24 and 48 h, these cells were super-infected with PBjΔVpx or PBjWT and 24 hours post SIVΔvpx-GFP infection, infection was gauged from quantitation of late viral cDNA using specific primers for the GFP transgene. SIVΔvpx-GFP infection of macrophage in the presence of AZT was used as a control to gauge de novo viral cDNA synthesis.
ACKNOWLEDGEMENTS
We thank Natalia Sharova and Mark Sharkey for helpful suggestions on the experiments, Ann Dauphin for technical support, Bruce Blaise for FACS analysis and core facilities within the University of Massachusetts Center for AIDS Research (P30-AI42845) for reagents and cells. Viral clones were obtained from the AIDS Research and Reference Reagent Program, NIAID, NIH. This work was supported by NIH grants RR11589 and AI37475 to M. Stevenson.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- Accola MA, Bukovsky AA, Jones MS, Gottlinger HG. A conserved dileucine-containing motif in p6(gag) governs the particle association of Vpx and Vpr of simian immunodeficiency viruses SIV(mac) and SIV(agm) J Virol. 1999;73:9992–9999. doi: 10.1128/jvi.73.12.9992-9999.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexaki A, Liu Y, Wigdahl B. Cellular reservoirs of HIV-1 and their role in viral persistence. Curr HIV Res. 2008;6:388–400. doi: 10.2174/157016208785861195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrejeva J, Poole E, Young DF, Goodbourn S, Randall RE. The p127 subunit (DDB1) of the UV-DNA damage repair binding protein is essential for the targeted degradation of STAT1 by the V protein of the paramyxovirus simian virus 5. J Virol. 2002;76:11379–11386. doi: 10.1128/JVI.76.22.11379-11386.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arthur LO, Bess JW, Snowder RC, II, Benveniste RE, Mann DL, Chermann J-C, Henderson LE. Cellular proteins bound to immunodeficiency viruses: implications for pathogenesis and vaccines. Science. 1992;258:1935–1938. doi: 10.1126/science.1470916. [DOI] [PubMed] [Google Scholar]
- Balcaitis S, Weinstein JR, Li S, Chamberlain JS, Moller T. Lentiviral transduction of microglial cells. Glia. 2005;50:48–55. doi: 10.1002/glia.20146. [DOI] [PubMed] [Google Scholar]
- Balliet JW, Kolson DL, Eiger G, Kim FM, McGann KA, Srinivasan A, Collman R. Distinct effects in primary macrophages and lymphocytes of the human immunodeficiency virus type 1 accessory genes vpr, vpu, and nef: mutational analysis of a primary HIV-1 isolate. Virology. 1994;200:623–631. doi: 10.1006/viro.1994.1225. [DOI] [PubMed] [Google Scholar]
- Bieniasz PD, Weiss RA, McClure MO. Cell cycle dependence of foamy retrovirus infection. J Virol. 1995;69:7295–7299. doi: 10.1128/jvi.69.11.7295-7299.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bouyac-Bertoia M, Dvorin JD, Fouchier RA, Jenkins Y, Meyer BE, Wu LI, Emerman M, Malim MH. HIV-1 infection requires a functional integrase NLS. Mol Cell. 2001;7:1025–1035. doi: 10.1016/s1097-2765(01)00240-4. [DOI] [PubMed] [Google Scholar]
- Bowerman B, Brown PO, Bishop JM, Varmus HE. A nucleoprotein complex mediates the integration of retroviral DNA. Gen Develop. 1989;3:469–478. doi: 10.1101/gad.3.4.469. [DOI] [PubMed] [Google Scholar]
- Bruce JW, Bradley KA, Ahlquist P, Young JA. Isolation of cell lines that show novel, murine leukemia virus-specific blocks to early steps of retroviral replication. J Virol. 2005;79:12969–12978. doi: 10.1128/JVI.79.20.12969-12978.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brussel A, Sonigo P. Analysis of early human immunodeficiency virus type 1 DNA synthesis by use of a new sensitive assay for quantifying integrated provirus. J Virol. 2003;77:10119–10124. doi: 10.1128/JVI.77.18.10119-10124.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukrinsky M, Haggerty S, Dempsey MP, Sharova N, Adzhubei A, Spitz L, Lewis P, Goldfarb D, Emerman M, Stevenson M. A nuclear localization signal within HIV-1 matrix protein that governs infection of non-dividing cells. Nature. 1993;365:666–669. doi: 10.1038/365666a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bukrinsky MI, Sharova N, Dempsey MP, Stanwick TL, Bukrinskaya AG, Haggerty S, Stevenson M. Active nuclear import of human immunodeficiency virus type 1 preintegration complexes. Proc Natl Acad Sci USA. 1992;89:6580–6584. doi: 10.1073/pnas.89.14.6580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carter CA, Ehrlich LS. Cell biology of HIV-1 infection of macrophages. Annu Rev Microbiol. 2008;62:425–443. doi: 10.1146/annurev.micro.62.081307.162758. [DOI] [PubMed] [Google Scholar]
- Cavrois M, De Noronha C, Greene WC. A sensitive and specific enzyme-based assay detecting HIV-1 virion fusion in primary T lymphocytes. Nat Biotechnol. 2002;20:1151–1154. doi: 10.1038/nbt745. [DOI] [PubMed] [Google Scholar]
- Chiu YL, Soros VB, Kreisberg JF, Stopak K, Yonemoto W, Greene WC. Cellular APOBEC3G restricts HIV-1 infection in resting CD4+ T cells. Nature. 2005;435:108–114. doi: 10.1038/nature03493. [DOI] [PubMed] [Google Scholar]
- Collman R, Hassan NF, Walker R, Godfrey B, Cutilli J, Hastings JC, Friedman H, Douglas SD, Nathanson N. Infection of monocyte-derived macrophages with human immunodeficiency virus type 1 (HIV-1). Monocyte-tropic and lymphocyte-tropic strains of HIV-1 show distinctive patterns of replication in a panel of cell types. J Exp Med. 1989;170:1149–1163. doi: 10.1084/jem.170.4.1149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Connor RI, Chen BK, Choe S, Landau NR. Vpr is required for efficient replication of human immunodeficiency virus type-1 in mono-nuclear phagocytes. Virology. 1995;206:935–944. doi: 10.1006/viro.1995.1016. [DOI] [PubMed] [Google Scholar]
- Di Marzio P, Choe S, Ebright M, Knoblauch R, Landau NR. Mutational analysis of cell cycle arrest, nuclear localization and virion packaging of human immunodeficiency virus type 1 Vpr. J Virol. 1995;69:7909–7916. doi: 10.1128/jvi.69.12.7909-7916.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Di Marzio P, Tse J, Landau NR. Chemokine receptor regulation and HIV type 1 tropism in monocyte-macrophages. AIDS Res Hum Retroviruses. 1998;14:129–138. doi: 10.1089/aid.1998.14.129. [DOI] [PubMed] [Google Scholar]
- Diamond TL, Roshal M, Jamburuthugoda VK, Reynolds HM, Merriam AR, Lee KY, Balakrishnan M, Bambara RA, Planelles V, Dewhurst S, Kim B. Macrophage tropism of HIV-1 depends on efficient cellular dNTP utilization by reverse transcriptase. J Biol Chem. 2004;279:51545–51553. doi: 10.1074/jbc.M408573200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisert V, Kreutz M, Becker K, Konigs C, Alex U, Rubsamen-Waigmann H, Andreesen R, von Briesen H. Analysis of cellular factors influencing the replication of human immunodeficiency virus type I in human macrophages derived from blood of different healthy donors. Virology. 2001;286:31–44. doi: 10.1006/viro.2001.0940. [DOI] [PubMed] [Google Scholar]
- Ellery PJ, Tippett E, Chiu YL, Paukovics G, Cameron PU, Solomon A, Lewin SR, Gorry PR, Jaworowski A, Greene WC, Sonza S, Crowe SM. The CD16+ monocyte subset is more permissive to infection and preferentially harbors HIV-1 in vivo. J Immunol. 2007;178:6581–6589. doi: 10.4049/jimmunol.178.10.6581. [DOI] [PubMed] [Google Scholar]
- Fletcher TM, Brichacek B, Sharova N, Newman MA, Stivahtis G, Sharp PM, Emerman M, Hahn BH, Stevenson M. Nuclear import and cell cycle arrest functions of the HIV-1 Vpr protein are encoded by two separate genes in HIV-2/SIMSM. EMBO Journal. 1996;15:6155–6165. [PMC free article] [PubMed] [Google Scholar]
- Gartner S, Markovits P, Markovitz DM, Kaplan MH, Gallo RC, Popovic M. The role of mononuclear phagocytes in HTLV-III/LAV infection. Science. 1986;233:215–219. doi: 10.1126/science.3014648. [DOI] [PubMed] [Google Scholar]
- Gendelman HE, Orenstein JM, Martin MA, Ferruca C, Mitra R, Phipps T, Wahl LA, Lane HC, Fauci AS, Burke DS. Efficient isolation and propagation of human immunodeficiency virus on recombinant colony-stimulating factor 1-treated monocytes. J Exp Med. 1988;167:1428–1441. doi: 10.1084/jem.167.4.1428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez-Scarano F, Martin-Garcia J. The neuropathogenesis of AIDS. Nat Rev Immunol. 2005;5:69–81. doi: 10.1038/nri1527. [DOI] [PubMed] [Google Scholar]
- Gorry PR, Churchill M, Crowe SM, Cunningham AL, Gabuzda D. Pathogenesis of macrophage tropic HIV-1. Curr HIV Res. 2005;3:53–60. doi: 10.2174/1570162052772951. [DOI] [PubMed] [Google Scholar]
- Goujon C, Arfi V, Pertel T, Luban J, Lienard J, Rigal D, Darlix JL, Cimarelli A. Characterization of simian immunodeficiency virus SIVSM/human immunodeficiency virus type 2 Vpx function in human myeloid cells. J Virol. 2008;82:12335–12345. doi: 10.1128/JVI.01181-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatzakis A, Touloumi G, Karanicolas R, Karafoulidou A, Mandalaki T, Anastassopoulou C, Zhang L, Goedert JJ, Ho DD, Kostrikis LG. Effect of recent thymic emigrants on progression of HIV-1 disease. Lancet. 2000;355:599–604. doi: 10.1016/S0140-6736(99)10311-8. [DOI] [PubMed] [Google Scholar]
- Heinzinger N, Bukrinsky M, Haggerty S, Ragland A, Lee M-A, Kewalramani V, Gendelman H, Ratner L, Stevenson M, Emerman M. The Vpr protein of human immunodeficiency virus type 1 influences nuclear localization of viral nucleic acids in nondividing host cells. Proc Natl Acad Sci USA. 1994;91:7311–7315. doi: 10.1073/pnas.91.15.7311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson LE, Sowder RC, Copeland TD, Benveniste RE, Oroszlan S. Isolation and characterization of a novel protein (X-ORF product) from SIV and HIV-2. Science. 1988;241:199–201. doi: 10.1126/science.3388031. [DOI] [PubMed] [Google Scholar]
- Jarrosson-Wuilleme L, Goujon C, Bernaud J, Rigal D, Darlix JL, Cimarelli A. Transduction of nondividing human macrophages with gammaretrovirus-derived vectors. J Virol. 2006;80:1152–1159. doi: 10.1128/JVI.80.3.1152-1159.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis P, Hensel M, Emerman M. Human immunodeficiency virus infection of cells arrested in the cell cycle. EMBO J. 1992;11:3053–3058. doi: 10.1002/j.1460-2075.1992.tb05376.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis PF, Emerman M. Passage through mitosis is required for oncoretroviruses but not for the human immunodeficiency virus. Journal of Virology. 1994;68:510–516. doi: 10.1128/jvi.68.1.510-516.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu YL, Spearman P, Ratner L. Human immunodeficiency virus type 1 viral protein R localization in infected cells and virions. J Virol. 1993;67:6542–6550. doi: 10.1128/jvi.67.11.6542-6550.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malim MH, Emerman M. HIV-1 accessory proteins--ensuring viral survival in a hostile environment. Cell Host Microbe. 2008;3:388–398. doi: 10.1016/j.chom.2008.04.008. [DOI] [PubMed] [Google Scholar]
- Munk C, Brandt SM, Lucero G, Landau NR. A dominant block to HIV-1 replication at reverse transcription in simian cells. Proc Natl Acad Sci U S A. 2002;99:13843–13848. doi: 10.1073/pnas.212400099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naif HM, Li S, Alali M, Sloane A, Wu L, Kelly M, Lynch G, Lloyd A, Cunningham AL. CCR5 expression correlates with susceptibility of maturing monocytes to human immunodeficiency virus type 1 infection. J Virol. 1998;72:830–836. doi: 10.1128/jvi.72.1.830-836.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naldini L, Blomer U, Gallay P, Ory D, Mulligan R, Gage FH, Verma IM, Trono D. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science. 1996;272:263–267. doi: 10.1126/science.272.5259.263. [DOI] [PubMed] [Google Scholar]
- Neil S, Martin F, Ikeda Y, Collins M. Postentry restriction to human immunodeficiency virus-based vector transduction in human monocytes. J Virol. 2001;75:5448–5456. doi: 10.1128/JVI.75.12.5448-5456.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Brien WA, Namazi A, Kalhor H, Mae S-H, Zack JA, Chen IS. Kinetics of human immunodeficiency virus type 1 reverse transcription in blood mononuclear phagocytes are slowed by limitations of nucleotide precursors. J Virol. 1994;68:1258–1263. doi: 10.1128/jvi.68.2.1258-1263.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pancio HA, Ratner L. Human immunodeficiency virus type 2 Vpx-Gag interaction. J Virol. 1998;72:5271–5275. doi: 10.1128/jvi.72.6.5271-5275.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paxton W, Connor RI, Landau NR. Incorporation of Vpr into human immunodeficiency virus type 1 virions: requirement for the p6 region of gag and mutational analysis. J Virol. 1993;67:7229–7237. doi: 10.1128/jvi.67.12.7229-7237.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng G, Greenwell-Wild T, Nares S, Jin W, Lei KJ, Rangel ZG, Munson PJ, Wahl SM. Myeloid differentiation and susceptibility to HIV-1 are linked to APOBEC3 expression. Blood. 2007;110:393–400. doi: 10.1182/blood-2006-10-051763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng G, Lei KJ, Jin W, Greenwell-Wild T, Wahl SM. Induction of APOBEC3 family proteins, a defensive maneuver underlying interferon-induced anti-HIV-1 activity. J Exp Med. 2006;203:41–46. doi: 10.1084/jem.20051512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rich EA, Chen IS, Zack JA, Leonard ML, O'Brien WA. Increased susceptibility of differentiated mononuclear phagocytes to productive infection with human immunodeficiency virus-1 (HIV-1) J Clin Invest. 1992;89:176–183. doi: 10.1172/JCI115559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ringler DJ, Wyand MS, Walsh DG, MacKey JJ, Chalifoux LV, Popovic M, Minassian AA, Sehgal PK, Daniel MD, Desrosiers RC, et al. Cellular localization of simian immunodeficiency virus in lymphoid tissues. I. Immunohistochemistry and electron microscopy. Am J Pathol. 1989;134:373–383. [PMC free article] [PubMed] [Google Scholar]
- Roe T, Reynolds TC, Yu G, Brown PO. Integration of murine leukemia virus DNA depends on mitosis. EMBO J. 1993;12:2099–2108. doi: 10.1002/j.1460-2075.1993.tb05858.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharova N, Wu Y, Zhu X, Stranska R, Kaushik R, Sharkey M, Stevenson MS. Primate lentiviral Vpx commandeers DDB1 to counteract a macrophage restriction. PLoS Pathog. 2008;4:4. doi: 10.1371/journal.ppat.1000057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp PM, Bailes E, Stevenson M, Emerman M, Hahn BH. Gene acquisition in HIV and SIV. Nature. 1996;383:586–587. doi: 10.1038/383586a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheehy AM, Gaddis NC, Choi JD, Malim MH. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature. 2002;418:646–650. doi: 10.1038/nature00939. [DOI] [PubMed] [Google Scholar]
- Sherer NM, Lehmann MJ, Jimenez-Soto LF, Ingmundson A, Horner SM, Cicchetti G, Allen PG, Pypaert M, Cunningham JM, Mothes W. Visualization of retroviral replication in living cells reveals budding into multivesicular bodies. Traffic. 2003;4:785–801. doi: 10.1034/j.1600-0854.2003.00135.x. [DOI] [PubMed] [Google Scholar]
- Sonza S, Maerz A, Deacon N, Meanger J, Mills J, Crowe S. Human immunodeficiency virus type 1 replication is blocked prior to reverse transcription and integration in freshly isolated peripheral blood monocytes. J Virol. 1996;70:3863–3869. doi: 10.1128/jvi.70.6.3863-3869.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Srivastava S, Swanson SK, Manel N, Florens L, Washburn MP, Skowronski J. Lentiviral Vpx accessory factor targets VprBP/DCAF1 substrate adaptor for cullin 4 E3 ubiquitin ligase to enable macrophage infection. PLoS Pathog. 2008;4:e1000059. doi: 10.1371/journal.ppat.1000059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevenson M, Stanwick TL, Dempsey MP, Lamonica CA. HIV-1 replication is controlled at the level of T cell activation and proviral integration. EMBO J. 1990;9:1551–1560. doi: 10.1002/j.1460-2075.1990.tb08274.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki Y, Craigie R. The road to chromatin - nuclear entry of retroviruses. Nat Rev Microbiol. 2007;5:187–196. doi: 10.1038/nrmicro1579. [DOI] [PubMed] [Google Scholar]
- Triques K, Stevenson M. Characterization of restrictions to human immunodeficiency virus type 1 infection of monocytes. J Virol. 2004;78:5523–5527. doi: 10.1128/JVI.78.10.5523-5527.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tristem M, Marshall C, Karpas A, Hill F. Evolution of the primate lentiviruses: evidence from vpx and vpr. EMBO J. 1992;11:3405–3412. doi: 10.1002/j.1460-2075.1992.tb05419.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinberg JB, Matthews TJ, Cullen BR, Malim MH. Productive human immunodeficiency virus type 1 (HIV-1) infection of nonproliferating human monocytes. J Exp Med. 1991;174:1477–1482. doi: 10.1084/jem.174.6.1477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfrum N, Muhlebach MD, Schule S, Kaiser JK, Kloke BP, Cichutek K, Schweizer M. Impact of viral accessory proteins of SIVsmmPBj on early steps of infection of quiescent cells. Virology. 2007;364:330–341. doi: 10.1016/j.virol.2007.03.008. [DOI] [PubMed] [Google Scholar]
- Wu X, Conway JA, Kim J, Kappes JC. Localization of the Vpx packaging signal within the C terminus of the human immunodeficiency virus type 2 gag precursor protein. J Virol. 1994;68:6161–6169. doi: 10.1128/jvi.68.10.6161-6169.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamashita M, Emerman M. Retroviral infection of non-dividing cells: old and new perspectives. Virology. 2006;344:88–93. doi: 10.1016/j.virol.2005.09.012. [DOI] [PubMed] [Google Scholar]
- Yamashita M, Perez O, Hope TJ, Emerman M. Evidence for direct involvement of the capsid protein in HIV infection of nondividing cells. PLoS Pathog. 2007;3:1502–1510. doi: 10.1371/journal.ppat.0030156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yao XJ, Subbramanian RA, Rougeau N, Boisvert F, Bergeron D, Cohen EA. Mutagenic analysis of human immunodeficiency virus type 1 Vpr: Role of a predicted N-terminal alpha-helical structure in Vpr nuclear localization and virion incorporation. J Virol. 1995;69:7032–7044. doi: 10.1128/jvi.69.11.7032-7044.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zack JA, Arrigo SJ, Weitsman SR, Go AS, Haislip A, Chen IS. HIV-1 entry into quiescent primary lymphocytes: molecular analysis reveals a labile, latent viral structure. Cell. 1990;61:213–222. doi: 10.1016/0092-8674(90)90802-l. [DOI] [PubMed] [Google Scholar]
- Zennou V, Petit C, Guetard D, Nerhbass U, Montagnier L, Charneau P. HIV-1 genome nuclear import is mediated by a central DNA flap. Cell. 2000;101:173–185. doi: 10.1016/S0092-8674(00)80828-4. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1: Dose-dependent ability of packaged Vpx to counteract a macrophage restrictionPseudotyped SIVΔVpx was produced in 293T cells by co-transfecting ΔVpx PBj1.9 proviral DNA with pMD-G and increasing amount of a Vpx expression vector. (A) The amount of packaged Vpx in the purified virions was determined by western blotting with a Vpx antibody and normalization with p27 by densitometry. The integrated density was measured by Scion Image software. (B) The infectivity of these viruses in primary macrophage was determined from quantitation of SIV 2LTR cDNA 24 and 48 h post-infection. PBjWT was used as positive control for this experiment.
Figures S2: Reversibility of restrictionMacrophage were first infected with SIVΔvpx-GFP. After 0, 1, 3, 5, 24 and 48 h, these cells were super-infected with PBjΔVpx or PBjWT and 24 hours post SIVΔvpx-GFP infection, infection was gauged from quantitation of late viral cDNA using specific primers for the GFP transgene. SIVΔvpx-GFP infection of macrophage in the presence of AZT was used as a control to gauge de novo viral cDNA synthesis.