Abstract
Memory formation in the brain is thought to be depending upon long lasting plastic changes of synaptic contacts that require alterations on the transcriptional level. Here, we characterize LAPSER1, a putative cytokinetic tumor suppressor that binds directly to ProSAP2/Shank3 and the synaptic Rap-Gap protein SPAR1 as a novel postsynaptic density component. Postsynaptic LAPSER1 is in complex with all important members of the canonical Wnt pathway including β-catenin. Upon N-methyl-d-aspartate receptor-dependent activation, LAPSER1 and β-catenin comigrate from the postsynaptic density to the nucleus and induce the transcription and translation of known β-catenin target genes, including Tcfe2a and c-Myc. The nuclear export and cytoplasmic redistribution of β-catenin is tightly regulated by LAPSER1. We postulate a postsynaptic cross-talk between N-methyl-d-aspartate receptors and a LAPSER1-β-catenin complex that results in a self-regulated, synaptic activity-dependent expression of β-catenin target genes. This calls for a novel role of Tcfe2a and c-Myc in plastic changes of neural tissue.
Changes in gene expression and protein synthesis leading to alterations in spine and synaptic morphology are believed to be the neurobiological core elements of learning and memory (1–4). These complex neuronal processes require the transport of signals generated at the synapse to the nucleus and their conversion into the transcription of specific genes. This activity-dependent synapse-to-nucleus signaling is known to be Ca2+-dependent and mediated by nucleocytoplasmatic shuttling of transcription factors and/or their regulatory proteins (5–7).
Recent studies have demonstrated that newly identified PSD4 molecules like Abi-1 (8), AIDA-1 (9), and Jacob (10) take part in activity-dependent synapse-to-nucleus shuttling, thus being good candidates for the proposed synaptically activated second messengers and transcription factors involved in long lasting forms of synaptic plasticity (11). Fezzins are a family of PSD proteins (LAPSER1/LZTS2 (12), ProSAPiP1/LZTS3 (13), PSD-Zip70/LZTS1 (14, 15), and N4BP3 (16)) that are able to build homo- and heterooligomers and are defined by a conserved Fez1 domain with hitherto unknown function. LAPSER1 has been initially characterized as a putative tumor suppressor gene according to its ability to inhibit cell growth when overexpressed (12) and has been reported to be involved in cytokinesis colocalizing with centrosomes and interacting with γ-tubulin and p80 katanin (17). Thyssen et al. (18) showed that LAPSER1 containing various domains of a potential transcription factor can indeed enter the nuclear compartment and interacts with β-catenin. Moreover, it is able to influence nuclear localization and signaling of this key molecule of the Wnt/β-catenin pathway (19). Synaptic β-catenin as a well known part of cadherin-mediated cell-cell adhesions at synapses (20, 21) translocates from synapses to the nucleus even without a specific Wnt signal (22). These observations prompted us to investigate the putative role of LAPSER1 in the nervous system.
We found that LAPSER1 is highly expressed in brain and localizes to the postsynaptic compartment interacting with the PDZ domain of ProSAP2/Shank3, an important member of the ProSAP/Shank family of scaffolding proteins (23–26). Upon NMDA receptor activation, LAPSER1 alters its spatial localization and binding properties and shuttles with β-catenin rapidly to the nucleus. The translocation is followed by the expression of β-catenin target genes including c-Myc and TcfE2a.
EXPERIMENTAL PROCEDURES
Cloning of LAPSER1
According to a homology search we found the rat cDNA sequence of the Fezzin family member LAPSER1 in public data bases. In a PCR-based approach we cloned a full-length rat LAPSER1 cDNA using rat hippocampal and total brain cDNA libraries (Stratagene) as template. The correct cDNA was confirmed by DNA sequencing and inserted into different bacterial and eucaryotic expression vectors including a GFP (pEGFP; Clontech, Palo Alto, CA) and dsRed vector (pDsRed-Express; BD Bioscience, San Jose, CA). Similarly mutant constructs were generated by PCR or by the use of restriction enzymes. For the GFP-LAPSER1ΔATEI construct, the last 4 aa were deleted, the GFP-LAPSER1ΔC construct codes for aa 1–420.
Northern Blot Analysis
For Northern blot analysis a rat multiple tissue Northern blot (Clontech) was hybridized with a C-terminal LAPSER1 cDNA probe (bp 1157 to the end) according to the manufacturer's protocol (Clontech).
Antibodies
A partial cDNA of LAPSER1 (encoding aa 1–448) was subcloned in the bacterial expression vector pGEX-4T (Amersham Biosciences). A 68.5-kDa glutathione S-transferase-LAPSER1 fusion protein was expressed in Escherichia coli BL 21 and purified on glutathione-Sepharose 4B as recommended by the manufacturer (Amersham Biosciences). The fusion protein was used to generate LAPSER1 antibodies in rabbits and guinea pigs (Pineda, Berlin, Germany). The ProSAP1/Shank2 and ProSAP2/Shank3 antibodies have been described previously (8). GFP (Clontech), Bassoon (Stressgen), LAPSER1/LZTS2 (directed against the full-length protein; Novus), β-catenin (C terminus; Fitzgerald), N terminus (Upstate Biotechnology), SPAR1 (Santa Cruz), LRP (lipoprotein receptor-related proteins) 5/6 (Biozol), Axin2 (Biozol), APC (Biozol), Dvl-3 (Biozol), FZD3 (Acris), GSK-3β (Sigma), CK1ϵ (Acris), Myc (Roche Applied Science), and TcfE2a (Abnova) antibodies were purchased from commercial suppliers.
Western Blot Analysis, Isolation of Subcellular Protein Fractions, and Immunoprecipitation of Brain Lysates
Subcellular fractionation studies were performed as described previously (8). In brief, tissue from adult rats was homogenized in homogenization buffer (320 mm sucrose, 5 mm HEPES, pH 7.4) containing protease inhibitor mixture (Roche Applied Science); cell debris and nuclei were removed by 1,000 × g centrifugation. The supernatant was spun for 20 min by 12,000 × g, resulting in supernatant S2 and pellet P2 (crude membrane fraction). The S2 was centrifuged at 100,000 × g for 1 h, and the resulting supernatant was taken as cytoplasmatic section (S100). The P2 pellet was further fractionated by centrifugation in a sucrose step gradient (0.85, 1.0, and 1.2 m). For isolation of synaptic junctional proteins (PSD fraction), the synaptosomal fraction of the first gradient was diluted with 5 volumes of 1 mm Tris, pH 8.1. The suspension was kept on ice for 30 min and was centrifuged for 30 min at 33,000 × g. The pellet P3 was resuspended in 5 mm Tris, pH 8.1 (1.5 ml/g of wet tissue) and once again fractionated by centrifugation in a sucrose gradient. The 1.0/1.2 m interphase (synaptic junctions) was suspended in 320 mm sucrose, 0.5% Triton X-100, 5 mm Tris, pH 8.1 (120 ml/10 g wet tissue), kept on ice for 15 min, and centrifuged for 30 min at 33,000 × g (PSD fraction).
For immunoprecipitation the following primary antibodies were used: LAPSER1, ProSAP2/Shank3, SPAR1, and β-catenin (N terminus). The crude membrane fraction (P2) of an adult rat brain was solubilized with radioimmune precipitation assay buffer for 1 h at 4 °C. After centrifugation (20 min, 12,000 × g, 4 °C), the supernatant was added to the agarose and incubated overnight at 4 °C. The mixture was washed twice with radioimmune precipitation assay buffer and once with 20 mm Tris, pH 8.0. Immunoprecipitated proteins were separated by SDS-PAGE and blotted onto a polyvinylidene difluoride membrane. Immunoreactivity was visualized by using ProSAP2/Shank3, LAPSER1, SPAR1, β-catenin, and horseradish peroxidase-conjugated secondary antibodies and the SuperSignal detection system (Pierce).
Culturing of Hippocampal Neurons
The preparation of hippocampal cultures was performed essentially as described by Dresbach et al. (27). Cell culture experiments of hippocampal primary neurons from rat (embryonic day 18) were performed as described previously (28). After preparation the hippocampal neurons were seeded on poly-l-lysine-coated (0,1 mg/ml; Sigma-Aldrich) glass coverslips. The cells were grown in Neurobasal medium (Invitrogen), complemented with B27 supplement (Invitrogen), 0.5 mm l-glutamine (Invitrogen), and 100 units/ml penicillin/streptomycin (Invitrogen) and maintained at 37 °C in 5% CO2.
All of the animal experiments were performed in compliance with the guidelines for the welfare of experimental animals issued by the Federal Government of Germany, the National Institutes of Health, and the Max Planck Society.
Neuronal Cell Culture and COS-7 Cell Transfections
LAPSER1 or a truncated 1form of LAPSER1 without the C terminus (GFP-LAPSER1-ΔC) were cloned into the GFP expression vector (pEGFP; Clontech) and dsRed vector (pDsRed-Express; BD Bioscience) using PCR strategies. Afterward the expression vectors were transfected into COS-7 cells or hippocampal neurons. Cotransfections with a dsRed-ProSAP2 construct (pdsRed2-ProSAP2-PDZ638–746), a ProSAP2/Shank3-Myc, or a GFP-SPAR1 (13) construct were performed. The cells were grown on glass coverslips, transfected using Polyfect (Qiagen) for COS-7 cells, and fixed with 4% paraformaldehyde 24 h after transfection. Cell cultures of hippocampal primary neurons were transfected as described previously (8).
Immunohistochemistry
Immunocytochemical staining was performed using 7-μm microtome sections from rat brains at different developmental stages, which were fixed by immersion in Bouin's fluid for 48 h, dehydrated, and embedded in paraplast. LAPSER1 was detected with the N-terminal rabbit anti LAPSER1 polyclonal antibody diluted 1:600 using the avidin-biotin-peroxidase complex (ABC) technique. Briefly, the sections were deparaffinized and incubated with primary antibodies in PBS containing 0.1% Triton X-100 for 24 h at room temperature, followed by incubation for 30 min at room temperature with the second antibody, biotin-labeled goat anti-rabbit IgG (Jackson ImmunoResearch, Soham, UK). The sections were then incubated for 30 min with a preformed complex of biotin-peroxidase-streptavidin (Jackson Immuno Research, Soham, UK), and peroxidase activity was revealed with 0.02% diaminobenzidine hydrochloride as chromogen.
For immunofluorescence experiments in cultured cells, the primary neurons were fixed with 4% paraformaldehyde, 1.5% sucrose, PBS at 4 °C for 20 min and processed for immunohistochemistry. After washing with PBS at room temperature, blocking was performed with 0.5% cold fish gelatin (Sigma) and 0.1% ovalbumin (Sigma), 1× PBS for 30 min at room temperature, and the cells were washed again with PBS, followed by the primary antibody at 4 °C overnight. After a washing step with PBS, incubation with the second antibody coupled to Alexa 488, Alexa 568, or Alexa 647 (1:1000) for 1 h room temperature followed. The cells were mounted in Mowiol with or without 4′,6-diamidino-2-phenylindole.
Yeast Two-hybrid Screening
Yeast two-hybrid screening was performed according to published methods. As bait, the cDNA coding for the Fez1 domain was fused to the GAL4 DNA-binding domain of vector pAS2–1 (Clontech). A rat brain cDNA library cloned into the pACT vector (GAL4 activation domain; Clontech) was screened. Eleven positive clones were identified; all partial cDNA clones coding for SPAR family members (30) were obtained and sequenced. For further analysis of the Fez1-SPAR1 interaction, the Fez1 domains of all Fezzins were isolated and cloned into the pAS2–1 vector. Moreover, the C terminus of SPAR1 was separated into three consecutive prey constructs that were all tested by a mini-mating approach.
Treatment of Hippocampal Cells
Hippocampal neurons were stimulated with either 100 μm NMDA; 30 μm glutamate; 50 μm AMPA; 200 ng/ml Wnt3a, 5a, or 7a; and 25 μm 5-amino-pyridine. NMDA or glutamate were added to the medium for 3 min, and the cells were incubated further in conditioned medium without NMDA/glutamate. Wnt proteins and 5-amino-pyridine were added continuously to the medium. Hanks' buffered salt solution (HBS) containing 110 mm NaCl, 5.4 mm KCl, 1.8 mm CaCl2, 0.8 mm MgCl2, 10 mm d-glucose, and 10 mm HEPES-NaOH, pH 7.4, was used for treatment with high potassium medium. For KCl stimulation, HBS medium was changed to high KCl-HBS (same as HBS except for 55 mm NaCl and 60 mm KCl).
mRNA Expression Study
10 μg of total RNA were isolated from control and NMDA-stimulated hippocampal cell culture neurons.
Microarray Analysis
The Oligo GEArray rat Wnt signaling pathway microarray ORN-043 was performed by SuperArray (Frederick, MD). All of the data sets were corrected using minimum value for background subtraction before being compared with each other.
Small Interference RNA Experiments
Knockdown of LAPSER1 was achieved by RNAi following published methods using the pSUPER vector (OligoEngine, Seattle, WA). Sequences for LAPSER1 RNAi constructs were chosen according to the Eurofins MWG Operon propositions. LAPSER1 target sequence AGCAGCAAC TGAAGGAATC and GCTGCAGC ACAACTATGTC oligonucleotides were ordered from Eurofins MWG Operon (Martinsried, Germany).
Statistical Analysis
The statistical analyses used in this study are indicated in each figure legend. To assess the amount of protein undergoing nuclear localization, the gray values of nuclei were measured after the indicated time points after NMDA stimulation using ImageJ 1.41o, and their means are compared with the mean gray values of nuclei of unstimulated cells.
RESULTS
LAPSER1, a Fezzin Family Member That Localizes at PSDs
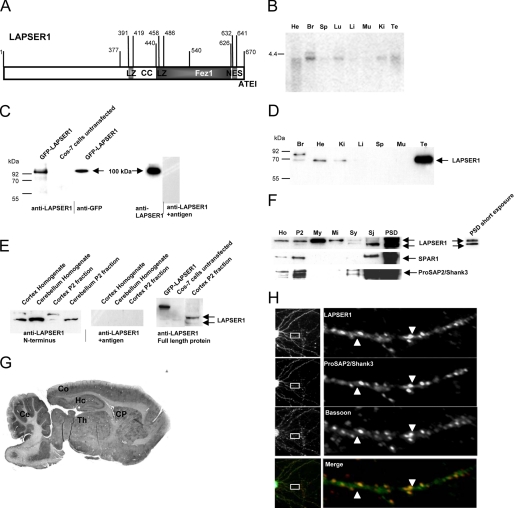
LAPSER1, a 670-aa-long leucine-rich protein is characterized by a coiled-coil domain, two leucine zippers, short proline- and serine-rich stretches, and a C-terminal ∼190-aa-long Fez1 (F37/esophageal cancer-related gene coding leucine zipper motif 1) domain (15) Moreover, it codes for a nuclear export signal as well as for a C-terminal PDZ domain-binding motif, ATEI (Fig. 1A; accession number DQ176638). Analysis of the expression profile of LAPSER1 revealed that on a multiple tissue Northern blot a 4.2-kb band could be detected in heart, lung, kidney, testis, and a stronger signal in brain (Fig. 1B). Western blot analysis with a polyclonal antibody directed specifically against LAPSER1 as shown for example by GFP-LAPSER1 transfection experiments into COS-7 cells and preabsorption of the antibody by the antigen (Fig. 1C) identified the protein at ∼70 kDa in the above mentioned tissues (Fig. 1D). In testis, LAPSER1 was most densely expressed. Interestingly, in brain the antibody detects an additional stronger band at a higher molecular mass of ∼85 kDa. The appearance of a second band in brain tissue was further verified using a commercially available antibody directed toward the full-length LAPSER1 protein. Further investigations using rat brain homogenates or P2 fractions from cortex and cerebellum revealed that the band of higher molecular mass could not be detected in the cerebellum (Fig. 1E). Subcellular fractionation from whole brain aimed to purify protein components of the PSD showed a strong enrichment of the relative amount of LAPSER1 immunoreactivity in the PSD fraction as compared with other fractions. This enrichment was as significant as the enrichment of the known PSD proteins SPAR1 and ProSAP2/Shank3. Within the PSD, the relative concentration of the 70-kDa band appears to be higher, compared with the 85-kDa band (Fig. 1F). In rat brain, LAPSER1 protein expression was found to be highest in the cerebral cortex, cerebellum, caudate putamen, thalamic nuclei, and hippocampus (Fig. 1G) and substantially increased from the early postnatal brain to adulthood as assessed by immunohistochemical staining of sagittal brain sections. Staining results not only showed that the proteins could be detected in cell cytoplasm but showed a punctate staining pattern in the neuropil of cerebellar, hippocampal, and cortical brain areas in the adult brain (data not shown). Furthermore, the enrichment of LAPSER1 immunoreactivity in PSD fractions was supported by the localization of LAPSER1 immunoreactivity at the PSD of excitatory synapses in 21-day-old hippocampal primary cultures. An analysis of the expression of LAPSER1 revealed that the molecule is localized in dendrites and spines, and the counterstaining with ProSAP2/Shank3 antibodies confirmed a match of both proteins at PSDs, whereas counterstaining with antibodies directed against the presynaptic marker protein Bassoon resulted in a staining of both antigens juxtaposed to each other (Fig. 1H).
FIGURE 1.
LAPSER1, a member of the Fezzin family of proteins, is mainly expressed in brain and localizes at postsynaptic densities. A, primary structural analysis of LAPSER1 revealed a central coiled-coil domain (CC) with two internal leucine zippers (LZ). The C-terminal part of the protein is characterized by a Fez1 domain (13) and a canonical Type I PDZ-binding motif (ATEI). Furthermore, between aa 632 and 641 a nuclear export signal is localized (accession number DQ176638). B, Northern blot analysis revealed an expression of LAPSER1 mRNA at ∼4.4 kb in brain (Br), lung (Lu), kidney (Ki), heart (He), and testis (Te). In the other tissues investigated (muscle (Mu), liver (Li), and spleen (Sp)) no LAPSER1 mRNA expression could be detected. C, the characterization of the self-generated polyclonal antibody against LAPSER1 showed that the antibody recognizes a GFP-LAPSER1 fusion protein in transfected COS-7 cells at a size of ∼100 kDa. This band was also detected by a GFP antibody. Preincubation of the antibody with the respected antigen abolished the signal. D and E, in rat brain homogenate the antiserum detected a double band in brain at ∼70 and 85 kDa, whereas in heart (He), kidney (Ki), and testis (Te), only the lower band is seen. Interestingly, Western blot experiments with homogenate or P2 fractions of different brain regions showed that the double band can be found in cortex and hippocampus (not shown) but not in the cerebellum. The GFP-LAPSER1 fusion protein as well as this double band was also detected by a commercially available LAPSER1 antibody directed against the full-length LAPSER1 protein. F, LAPSER1 immunoreactivity was slightly enriched in the insoluble protein fraction (P2). Further subcellular fractionation revealed a tight association with the synaptic cytoskeleton (PSD). The lower band seemed to be especially associated with the cytoskeleton of synaptic junctions because this band is mostly enriched toward the PSD fraction. Antibodies directed against the known PSD proteins SPAR1 and ProSAP2/Shank3 were used as quality controls for the fractions. Ho, homogenate; P2, membrane fraction; My, myelin fraction; Mi, mitochondrial fraction; Sy and Sj, synaptosomal fractions. G and H, in rat brain LAPSER1 protein could be readily detected in neurons of the hippocampal area (Hc), in the cortex (Co), in thalamic nuclei (Th), in the caudate putamen (CP), and in the cerebellum (Ce). LAPSER1 specifically localized to synaptic spines and PSDs in 3-week-old hippocampal neurons in culture. The LAPSER1 antigen was found in spines and PSDs colocalizing with ProSAP2/Shank3 used as postsynaptic marker. Bassoon staining defined the presynaptic cytomatrix.
LAPSER1 Binds to ProSAP2/Shank3 and Interacts with SPAR Family Members via the Fez1 Domain
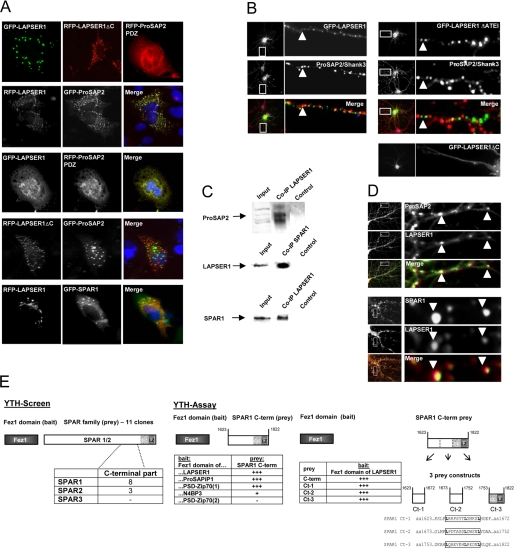
Previous work has shown that other Fezzins are able to interact with the PDZ domain of molecules of the ProSAP/Shank family via the C terminus and with the spine-associated Rap-Gap protein SPAR1 (29). To analyze whether LAPSER1 also interacts with the ProSAP2/Shank3 PDZ domain and SPAR1, we employed cotransfection experiments in COS-7 cells, immunoprecipitation and colocalization experiments. Full-length GFP- or RFP-LAPSER1 fusion proteins as well as a truncated form of LAPSER1 missing the C terminus (GFP-LAPSER1ΔC) could be detected as small clusters when transfected alone. After being cotransfected with ProSAP2/Shank3 or SPAR1, both proteins colocalized within these clusters. Interestingly, these clusters were lost when GFP-LAPSER1 is cotransfected with the ProSAP2/Shank3 PDZ domain as an RFP fusion protein. It seems that under these conditions LAPSER1 is recruited to the evenly distributed PDZ domain within the cell cytoplasm. The GFP-LAPSER1ΔC fusion protein missing the interaction domain with ProSAP2/Shank3 did not colocalize when cotransfected (Fig. 2A). Transfection of hippocampal neurons with the GFP-LAPSER1 fusion protein resulted in a clear targeting to postsynaptic sites. In contrast, the deletion of the last 4 aa in LAPSER1 abolished the synaptic localization, and further truncation led to an even distribution in the soma and dendritic compartment (Fig. 2B). Coimmunoprecipitation experiments showed that LAPSER1, SPAR1, and ProSAP2/Shank3 were coclustered within one postsynaptic complex. Using specific LAPSER1 antibodies in coimmunoprecipitation experiments of rat brain lysate revealed that ProSAP2/Shank3 and SPAR1 are readily detectable within the precipitate (Fig. 2C). Moreover, all three postsynaptic molecules colocalized at PSDs in hippocampal cells as demonstrated by immunostaining (Fig. 2D). To further characterize the functional role of the highly conserved Fez1 domain, we performed a yeast two-hybrid screen using the Fez1 domain as bait. This screen yielded several independent clones (n = 11) that were all coding for partial C-terminal sequences of members of the SPAR family of proteins (30). Eight clones were coding for SPAR1, three were coding for SPAR2, and none were coding for SPAR3. To determine whether SPAR binding is a universal feature of Fez1 domains, we subsequently cloned all Fez1 domains of the four Fezzin members into the bait vector and tested them in a yeast two-hybrid assay using the shortest SPAR1 prey clone. This filter lift assay revealed that the C-terminal Fez1 domains of the four fezzins interacted with SPAR; only the N-terminal Fez1 domain of PSD-Zip70 that showed only minor homology with the others was not a SPAR-interacting region (Fig. 2E). To further analyze the exact Fez1-interacting protein region within the SPAR1 C terminus, we subcloned this protein stretch into three approximately 80-aa-long prey proteins. Surprisingly, all three subclones were positive after mini-mating, plating, and filter lift assay. Finally, an alignment of these partial C-terminal SPAR proteins revealed repetitive leucine-rich regions in all of these proteins that could well be responsible for the very effective binding and interaction of all three C-terminal SPAR1 clones.
FIGURE 2.
LAPSER1 interacts with ProSAP2/Shank3 and SPAR1. A, single transfections of GFP-LAPSER1 and RFP-LAPSER1-ΔC in COS-7 cells revealed that the fusion proteins were both found in small cytoplasmic clusters. The ProSAP2/Shank3 PDZ domain was evenly distributed in the cell. Cotransfection experiments of full-length RFP-LAPSER1 and GFP-ProSAP2/Shank3 in COS-7 cells resulted in a perfect overlay of both fusion proteins. This held also true for the cotransfection of GFP-LAPSER1 with the PDZ domain of ProSAP2/Shank3 as an RFP construct (RFP-ProSAP2-PDZ). Here, GFP-LAPSER1 seemed to be recruited to the ProSAP2/Shank3 PDZ domain localized in the perinuclear area. In contrast, the RFP-LAPSER1-ΔC fusion protein without the ProSAP2/Shank3 interaction motif did not cocluster with GFP-ProSAP2/Shank3. Full-length GFP-SPAR1 fusion proteins also perfectly colocalized with RFP-LAPSER1 when cotransfected in COS-7 cells. B, transfection of hippocampal neurons with the GFP-LAPSER1 fusion protein led to the postsynaptic localization of small fluorescent clusters colocalizing with ProSAP2/Shank3 (arrowheads). In contrast, the deletion of the very C-terminal ProSAP2/Shank3 interaction signal (GFP-LAPSER1ΔATEI) resulted in nonsynaptic localization of LAPSER1 clusters, and further deletion of the C-terminal coiled-coil domain (GFP-LAPSER1ΔC) completely prevented cluster formation. C and D, coimmunoprecipitations with rat brain homogenate showed that the protein-protein interactions investigated occur also in vivo. ProSAP2 as well as SPAR1 could be detected in the LAPSER1 precipitate, and vice versa LAPSER1 was present in the SPAR1 precipitate (5% of the homogenate is loaded as input). In hippocampal neurons endogenous LAPSER1 also colocalized with ProSAP2/Shank3 and SPAR1 at postsynaptic specializations of synaptic contacts (arrowheads). E, to determine the interaction partner of the Fez1 domain, we performed a yeast two-hybrid screen employing the Fez1 domain as bait. To our surprise all 11 positive clones were coding for members of the SPAR family of proteins (30). The shortest prey clone was only ∼200 aa long, indicating that the very C-terminal part contains the interaction motif. In a yeast two-hybrid assay we could show that the Fez1 domains of all family members are able to bind the SPAR1 C terminus, indicating that this binding specificity is a common feature of Fez1 domains. The Fez1 domain of N4BP3, and the not well defined second putative Fez1 domain of PSD-Zip70 did not bind SPAR1 as effective or do not bind at all. The separation of the SPAR1 C terminus into three subregions cloned as preys revealed that all subclones are interacting very well. Close analysis of the three subclones showed that all subclones are coding for a leucine-rich region that could be responsible for the interaction with the Fez1 domain.
LAPSER1 Binds to and Coclusters with β-Catenin and Colocalizes with Wnt Pathway Members Like the Casein Kinase1ϵ, LRP5/6, Dvl-3, Axin-1, and GSK-3β at Postsynaptic Clusters
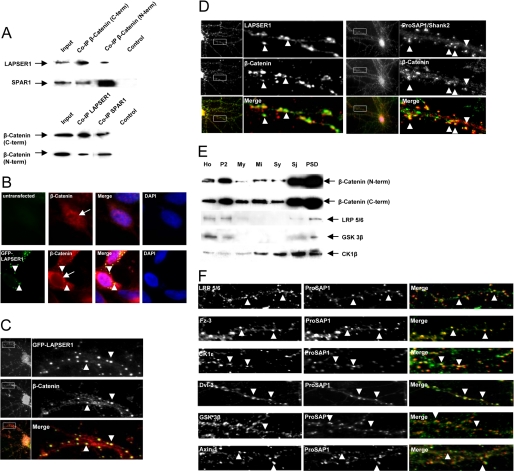
Because it was recently published that LAPSER1 directly interacts with β-catenin in tumor cells (18), we analyzed whether this protein complex could also be found at postsynaptic densities. To that end we performed coimmunoprecipitations with antibodies directed against the C- and N-terminal parts of β-catenin. LAPSER1 as well as SPAR1 could be detected in the precipitate, and vice versa precipitations with LAPSER1 and SPAR1 resulted in the staining of β-catenin on the Western blot (Fig. 3A). Overexpression of GFP-LAPSER1 in COS-7 cells and/or hippocampal neurons led to the recruitment of endogenous β-catenin into LAPSER1 positive clusters. Interestingly, in contrast to untransfected cells, the nuclei of transfected cells were free of any β-catenin staining (Fig. 3, B and C). The costaining of hippocampal neurons confirmed the colocalization of both endogenous proteins mainly at postsynaptic sites as also shown by the postsynaptic marker molecule ProSAP1/Shank2 (Fig. 3D). Because β-catenin is a key molecule of the canonical Wnt pathway being able to shuttle from the cytoplasm into the nucleus, we were interested whether other major components of this signaling cascade could be found within the PSD. Immunolabeling of PSD subfractions as well as staining of hippocampal neurons revealed the specific enrichment/cyolocalization of the LRP5/6 coreceptor, Fz-3 (frizzled 3 receptor) and CK1ϵ (casein kinase ϵ) at postsynaptic sites. Moreover, the Wnt scaffold protein Dvl-3 (dishevelled 3) and proteins of the β-catenin destruction complex (Axin-1 and GSK-3β) are specifically targeted to the postsynaptic compartment colocalizing with the PSD marker molecule (ProSAP1/Shank2) (Fig. 3, E and F).
FIGURE 3.
LAPSER1 is part of a postsynaptic complex that contains β-catenin and other important Wnt pathway molecules. A, coimmunoprecipitation experiments with rat brain homogenate revealed that β-catenin interacts with LAPSER1 in brain. Two commercially available antibodies directed against the C- and N-terminal part of β-catenin were used for precipitation, and LAPSER1 could be detected in the precipitate. Moreover, SPAR1 was also found within the precipitate. Vice versa, experiments showed that after the precipitation with LAPSER1- or SPAR1 antibodies, β-catenin can be detected in the precipitate. B and C, overexpression of GFP-LAPSER1 in COS-7 cells or hippocampal neurons led to a recruitment of endogenous β-catenin (also shown in an untransfected COS-7 cell) into the GFP-LAPSER1 clusters in the cell cytoplasm and at postsynaptic sites (arrowheads). In contrast to untransfected cells, the overexpression of LAPSER1 led to the complete nuclear elimination of endogenous β-catenin that seemed now to be concentrated perinuclear (arrows). D, the endogenous complex of LAPSER1 and β-catenin could be seen at postsynaptic sites in cultured hippocampal neurons (21 days) after double-immunostaining (arrowheads). The mainly postsynaptic localization of β-catenin is verified with the postsynaptic marker ProSAP1/Shank2. E and F, the Wnt pathway members β-catenin, LRP5/6, GSK-3β, and CK1ϵ were highly enriched toward the PSD fraction of rat brain homogenate. Moreover, these proteins as well as the Wnt pathway components Fz-3 (frizzled receptor 3) and Dvl-3 (dishevelled 3) could be detected in hippocampal cells concentrated as dot-like structures that overlay with the labeling of the postsynaptic marker protein ProSAP1/Shank2 indicating the postsynaptic localization of the molecules (arrowheads).
LAPSER1 and β-Catenin Translocate to the Nucleus after Application of NMDA
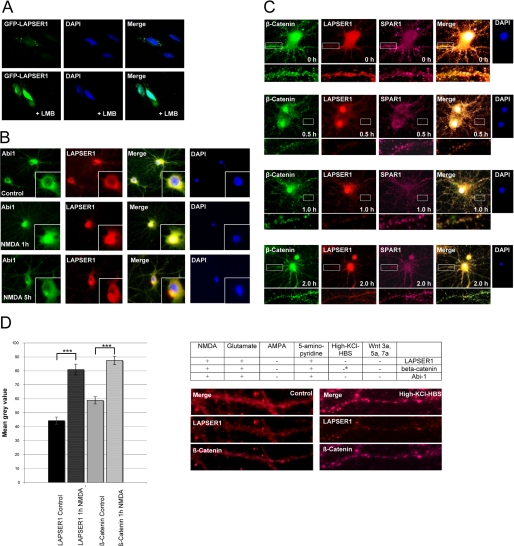
To determine the functional role of the LAPSER1-β-catenin interaction, we analyzed whether LAPSER1 could be found in different cellular subcompartments as shown for β-catenin. Application of the nuclear export inhibitor leptomycin B to GFP-LAPSER1 transfected COS-7 cells revealed that under this condition, GFP-LAPSER1 was mainly seen within the nuclear compartment (Fig. 4A). In accordance to previous observations that the abelson tyrosine kinase substrate Abi-1 is translocated from the PSD to the nucleus upon short application of NMDA (8), we observed that LAPSER1 was similarly directed to the nucleus and redistributed into dendrites and PSDs after 5 h (Fig. 4B). In the next step we analyzed the β-catenin-LAPSER1-SPAR1 complex with respect to nuclear shuttling. Our data showed that only β-catenin-LAPSER1 left the postsynaptic compartment after NMDA application, whereas SPAR1 stayed at synaptic sites. This translocation was only seen after application of NMDA, glutamate, or the release of endogenous glutamate by 5-aminopyridine. With the application of AMPA, some selected Wnt molecules or membrane depolarization by “high K+” had no effect on the distribution of neither the β-catenin-LAPSER1 complex nor Abi-1. However, we observed that high K+ levels significantly increased the local clustering and concentration of β-catenin at synaptic sites supporting previously published data by Murase et al. (31) (Fig. 4, C and D).
FIGURE 4.
LAPSER1 and β-catenin translocate from PSDs to the nucleus. A, GFP-LAPSER1 that was found in small clusters in COS-7 cells could be detected in the nucleus after application of the nuclear export inhibitor leptomycin B (LMB), indicating the shuttling of the molecule between both compartments. B, this phenomenon could also be seen in hippocampal neurons. After a short application (3 min) of NMDA and a change of medium, endogenous LAPSER1 was concentrated in the nuclear compartment after 1 h. Abi-1 that is known to also translocate into the nucleus was used as a positive control. The labeling of dendrites was markedly reduced. After 5 h Abi-1 as well as LASPER1 were redistributed into the dendritic arbors, spines, and PSDs, respectively. C and D, the nuclear accumulation after application of NMDA was also seen for β-catenin that shows a similar transport to the nucleus for at least 2 h. Nuclear β-catenin could be detected with the N- and C-terminal antibodies. SPAR1, however, which is part of the same PSD complex, did not change its synaptic localization under these experimental conditions. Statistical analysis revealed the significant accumulation of LAPSER1 and β-catenin in the nucleus after 1 h of NMDA application compared with controls. The nuclear translocation of LAPSER1 and β-catenin could be induced by NMDA, glutamate, and the release of endogenous glutamate by 5-aminopyridine, but not by AMPA, depolarization via high K+ levels, or application of different commercially available Wnt molecules. Depolarization by high K+ levels, however, resulted in an accumulation of β-catenin (indicated by a asterisk) at spines and PSDs as already published (31); LAPSER1 stainings were unaffected (t test; ***, p < 0.001). DAPI, 4′,6-diamidino-2-phenylindole.
LAPSER1 Is Important for the Nuclear Export of β-Catenin
In search of the functional role of LAPSER1 within the nuclear LAPSER1-β-catenin complex, we specifically down-regulated LAPSER1 expression by RNAi (Fig. 5A) and applied NMDA to neuronal cultures. Analysis of the RNAi-transfected cells and control cells that were transfected with the vector plasmid alone did not show any differences in the nuclear import of β-catenin after 1 h of post-NMDA application (Fig. 5B). After 4 h, however, β-catenin was still in the nuclear compartment in most of the LAPSER1 RNAi transfected neurons. This was significantly different from the control cells where β-catenin was mainly redistributed into the cytoplasm, dendrites, and spines of the neurons (Fig. 5C).
FIGURE 5.
LAPSER1 regulates the nuclear export of β-catenin after NMDA stimulation in neurons. A, LAPSER-RNAi transfected neurons were nearly completely depleted of endogenous LAPSER1 protein as revealed by immunostaining. B, after 1 h of NMDA application the nuclear import of β-catenin was not altered in cells transfected with either the LAPSER1 RNAi or empty vector construct (Ctrl). C, after 4 h, however, a significant percentage of LAPSER1 RNAi transfected neurons still displayed a prominent β-catenin localization within the nuclear compartment pointing toward the important role of LAPSER1 as a regulator of β-catenin shuttle between the neuronal cytoplasm and nucleus especially with respect to nuclear export. Statistical analysis was performed on the basis of 15 neurons at each time point from three independent experiments applying the Student's t test (*, p < 0.05). DAPI, 4′,6-diamidino-2-phenylindole.
β-Catenin Regulates Known Target Genes of the Canonical Wnt/β-Catenin Pathway Depending upon LAPSER1 Expression
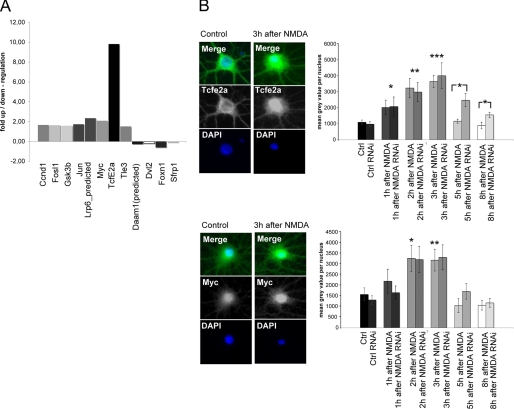
Because NMDA application was shown to induce the nuclear translocation of β-catenin in neurons, we were interested to explore putative target genes of this novel signaling pathway. To that end we performed a microarray analysis (Oligo GEArray Rat Wnt Signaling Pathway Microarray) comparing mRNA levels of NMDA-stimulated and unstimulated hippocampal neurons. 12 Wnt pathway-related genes were found to be regulated. A 10-fold up-regulation was seen for a known β-catenin target gene, the transcription factor TcfE2a that specifically interacts with β-catenin in the nucleus, and among others a 2-fold increase was observed for the transcription factor Myc (Fig. 6A). We analyzed both of these factors by immunohistochemical staining after NMDA application and investigated the possible role of LAPSER1 for this regulatory pathway. We found TcfE2a as well as the Myc protein significantly up-regulated in nuclei of NMDA-treated neurons that were either transfected with the empty vector or with the LAPSER1 RNAi construct. Interestingly, 5 and 8 h after NMDA treatment the TcfE2a and Myc protein levels went back to basic values in control neurons but stayed elevated in the LAPSER1 RNAi transfected neurons. At these time points the differences between the experimental groups were significant for TcfE2a (Fig. 6B).
FIGURE 6.
NMDA application leads to the enhanced transcription of several known β-catenin target genes including c-Myc and Tcfe2a. A, microarray analysis (Oligo GEArray Rat Wnt Signaling Pathway Microarray) comparing mRNA levels of NMDA stimulated and unstimulated hippocampal neurons revealed the specific up-/down-regulation of several Wnt pathway genes. The mRNA was prepared 1 h after NMDA treatment. The β-catenin nuclear receptor gene TcfE2a shows a 10-fold up-regulation, and Myc shows a 2-fold up-regulation. B, staining of hippocampal neurons transfected with LAPSER1 RNAi or the empty vector construct revealed 3 h after NMDA application a significant increase of TcfE2a and Myc protein levels in both experimental groups as determined by the measurement of staining intensity within the nuclear compartment of hippocampal neurons. Interestingly, after 5 and 8 h, the TcfE2a and Myc protein levels went back to control (Ctrl) values in neurons transfected with the empty RNAi vector construct but stayed elevated in the LAPSER1 RNAi transfected neurons. For TcfE2a the differences were significantly different at 5 and 8 h after NMDA application. Statistical analysis was performed on the basis of 20 neurons from three independent experiments applying the t test (*, p < 0.05; **, p < 0.01; ***, p < 0.001). DAPI, 4′,6-diamidino-2-phenylindole.
DISCUSSION
In this study, we provide first insights into the functional role of the Fezzin protein LAPSER1 (13) in brain. LAPSER1 is a ProSAP2/Shank3-interacting PSD protein adding a new member to the growing list of molecules that interact with the PDZ domain of ProSAP/Shank proteins (32). This interaction is essential for the postsynaptic targeting and adherence of LAPSER1. One LAPSER1 isoform seems to exist exclusively in brain. Western blot analysis of P2, Sy, and PSD fractions indicated two different bands at ∼70 and 85 kDa, respectively, that were recognized by an N-terminal as well as by an antibody directed against the full-length LAPSER1 protein. Although alternative splicing of the human LAPSER1 gene has been proposed to take place in testis and prostate (12), we could not detect any LAPSER1 splice variants by Northern blot analysis. Therefore, the two isoforms may derive from post-translational modification. Putative phosphorylation sites have been predicted for the amino acid sequences of LAPSER1 and homologues (12, 15). Because serine phosphorylation of PSD-Zip70 resulted in slightly larger protein bands on SDS-PAGE (15), it might well be that the LAPSER1 doublet in some brain areas represents phosphorylated and dephosphorylated isoforms with specific functions in different neuronal cell types and/or compartments.
Moreover, we were able to characterize the potential of the novel protein-protein interaction domain “Fez1” that is highly conserved among the Fezzin proteins. Fez1 domains are approximately 200 aa long and were found to specifically interact with the C termini of actin binding SPAR molecules (30) that localize at PSDs. It seems that this interaction is secured by redundant short interaction motifs because we found that even partial SPAR C-terminal sequences are efficiently bound to the LAPSER1 Fez1 domain. Because Fezzin family members are able to build large homo- and heteromeric scaffolds (13), one can envision that Fezzin protein scaffolds are functioning as multimodular “docking stations” for SPAR proteins at spines and PSDs.
Apart from its crucial role in the canonical Wnt pathway, β-catenin has already been shown to be present at synaptic sites interacting with the axodendritic adhesion molecule N-cadherin (21) and the actin-binding protein α-catenin, which links cadherin-coupled β-catenin to the cytoskeleton (33). Here, we provide evidence that all key molecules of the canonical Wnt/β-catenin pathway (19) including casein kinase 1ϵ, LRP5/6, Dvl-3, Axin-1, and GSK-3β are also expressed in neurons throughout the brain as revealed by in situ hybridization (data not shown) and are specifically localized at PSDs of excitatory synapses. LAPSER1 is connected to this signaling complex by the direct interaction with β-catenin. Synaptic β-catenin has also shown to be a binding partner of synaptic scaffolding molecule (S-SCAM), a MAGUK family member that is associated with the NMDA receptor cluster either indirectly via SAP90/PSD-95 or directly via the NMDAR subunit NR2A (34, 35). Wnt signaling is achieved through the binding of a receptor complex that includes Frizzled (Fz) receptors, low density LRP-5/6 and tyrosine kinase-related receptor RYK (36). This Wnt binding subsequently activates dishevelled (Dvl), a cytosolic protein bringing together signaling components for efficient transduction (37–39). Recent studies have revealed the diverse roles of Wnts in the CNS that include differentiation of synaptic specializations, synaptic protein organization, and regulation of gene expression in the developing as well as in the adult lesioned or unlesioned brain (39–43). Identification of neurons secreting Wnts and those containing the molecular components downstream of the Wnt receptor apparatus suggest that Wnts function both as anterograde and retrograde signals in the brain at synaptic junctions (40, 44–46). Here, we show that PSDs of the vertebrate brain are indeed equipped with the molecular machinery for the canonical Wnt pathway. These data underline the important role for the Wnt pathway with respect to neuronal cell fate, differentiation, synaptogenesis, and synaptic function (41, 44, 47–49). We were not able to identify the synaptic canonical Wnt/β-catenin pathway; however, we could show that NMDA receptor activation induces the nuclear translocation of a LAPSER1-β-catenin complex. Under these conditions the PSD protein SPAR1 that has been shown to be regulated and phosphorylated by Wnt-CKIϵ-Rap1 in Xenopus (50) stays at synaptic sites. In the case of β-catenin the nuclear translocation fits nicely to the data published by Takeichi and Abe (22), who showed that cleaved β-catenin itself might shuttle from synapses even without a specific Wnt signal. In contrast to their observations, we could detect nuclear β-catenin with the N- and C-terminal antibody, indicating that we identified nuclear β-catenin most likely as an uncleaved protein. The nuclear shuttle of LAPSER1-β-catenin could not be elicited by membrane depolarization alone or AMPA receptor activation but seems to be coupled to the NMDA receptor. As described above Wnts are normally needed to rescue β-catenin from the destruction complex. Therefore, we propose that neuronal cells have developed either a different mechanism to stabilize β-catenin at PSDs that might involve LAPSER1 or that Wnts might be coreleased from the presynaptic compartment. We screened for the neuronal expression of different Wnt molecules by in situ hybridization (data not shown) and found most of them broadly expressed in brain (2b, 3, 3a, 4, 5a, 5b, 7a, 8a, 9a, 10b, and 16), some were specific for the hippocampal area (2 and 7b), and some were not expressed (1, 6, and 10a). This list still opens up the possibility that, for example, a steady state release of Wnt proteins at synapses is important for the maintenance of Wnt signaling pathways. The commercially available Wnt proteins 3a, 5a, and 7a had no effect on synaptic β-catenin and did not influence NMDA induced β-catenin translocation. This might, however, be due to the fact that we did not apply the appropriate Wnt glycoprotein that influences the synapto-nuclear communication in neurons.
To understand how the LAPSER1-β-catenin nuclear localization could be involved in NMDA activity-dependent gene transcription, we screened for induced target genes of the Wnt pathway. Here, we found the nuclear binding molecule Tcf to be significantly induced. Tcf/Lef is a well characterized nuclear complex that translates the Wnt signal into the enhanced transcription of specific target genes after association with nuclear β-catenin (51). The strong induction of TcfE2a mRNA and protein might be a hint for a positive feedback mechanism that enhances LAPSER1-β-catenin induced alteration of gene transcription and might help to understand the proposed induction of gene expression in neurons with respect to learning and memory formation (11). We found that LAPSER1 seems to regulate the localization of β-catenin in neuronal cells by enhancing its nuclear export via a functional nuclear export sequence. Because a permanent increase of activated β-catenin within the nucleus is known to inhibit cell differentiation and apoptosis causing cancer in several tissues (52), these findings support the proposed function of LAPSER1 as a putative tumor suppressor gene (12, 17) also in neurons. Because the elevated expression of TcfE2a was significantly prolonged in LAPSER1-depleted neurons, first evidence was provided that the regulated nuclear export of β-catenin is one mechanism to terminate β-catenin-dependent gene transcription. This points toward a self-limiting function of the LAPSER1-β-catenin complex in neuronal cell nuclei after synaptic activation. The redistribution of the molecules into the synaptic compartment should therefore prepare the neuron for novel events of synaptic activation.
Interestingly, the transcription factor c-Myc was also among the induced target genes pointing toward an important role of this molecule in postmitotic neurons with respect to neuronal plasticity. This hypothesis is supported by the fact that after synaptic activation nuclear Abi-1 was found to be in complex with the Myc/Max dimer (8). What could be the role of activity induced Myc or Myc/Max complexes, respectively? Several thousand genes are potentially regulated by Myc/Max, and by now only the in vivo regulation of some specific genes involved in cell cycle regulation, transformation, tumorigenesis, and development are characterized (53). Knoepfler (54) recently proposed that elevated Myc transcription factors might act more general in a potential nontranscriptional function but by the regulation of chromatin structure in a global fashion. This concept is especially supported by the phenotype of N-Myc knock-out mice that show severe defects of neural progenitor proliferation and brain growth (55) and a striking alteration of the nuclear structure with extremely condensed chromatin (56). In turn elevated Myc levels might influence surprisingly large chromatin domains and enable gene transcription in activated neurons.
Acknowledgments
We gratefully acknowledge the professional technical assistance of M. Manz, N. Damm, and U. Pika-Hartlaub.
This work was supported by Deutsche Forschungsgemeinschaft Grants SFB497-B8 (to T. M. B.) and BO 1718;3-1.
- PSD
- postsynaptic density
- NMDA
- N-methyl-d-aspartate
- GFP
- green fluorescent protein
- RFP
- red fluorescent protein
- aa
- amino acid(s)
- PBS
- phosphate-buffered saline
- RNAi
- RNA interference
- AMPA
- α-amino-3-hydroxy-5-methyl-4-isoxazolpropionic acid.
REFERENCES
- 1.Kelleher R. J., 3rd, Govindarajan A., Jung H. Y., Kang H., Tonegawa S. (2004) Cell 116, 467–479 [DOI] [PubMed] [Google Scholar]
- 2.Kelleher R. J., 3rd, Govindarajan A., Tonegawa S. (2004) Neuron 44, 59–73 [DOI] [PubMed] [Google Scholar]
- 3.Costa-Mattioli M., Gobert D., Harding H., Herdy B., Azzi M., Bruno M., Bidinosti M., Ben Mamou C., Marcinkiewicz E., Yoshida M., Imataka H., Cuello A. C., Seidah N., Sossin W., Lacaille J. C., Ron D., Nader K., Sonenberg N. (2005) Nature 436, 1166–1173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wu H., Zhou Y., Xiong Z. Q. (2007) FEBS J. 274, 3218–3223 [DOI] [PubMed] [Google Scholar]
- 5.West A. E., Griffith E. C., Greenberg M. E. (2002) Nat. Rev. Neurosci. 3, 921–931 [DOI] [PubMed] [Google Scholar]
- 6.Deisseroth K., Mermelstein P. G., Xia H., Tsien R. W. (2003) Curr. Opin. Neurobiol. 13, 354–365 [DOI] [PubMed] [Google Scholar]
- 7.Flavell S. W., Cowan C. W., Kim T. K., Greer P. L., Lin Y., Paradis S., Griffith E. C., Hu L. S., Chen C., Greenberg M. E. (2006) Science 311, 1008–1012 [DOI] [PubMed] [Google Scholar]
- 8.Proepper C., Johannsen S., Liebau S., Dahl J., Vaida B., Bockmann J., Kreutz M. R., Gundelfinger E. D., Boeckers T. M. (2007) EMBO J. 26, 1397–1409 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Jordan B. A., Fernholz B. D., Khatri L., Ziff E. B. (2007) Nat. Neurosci. 10, 427–435 [DOI] [PubMed] [Google Scholar]
- 10.Dieterich D. C., Karpova A., Mikhaylova M., Zdobnova I., König I., Landwehr M., Kreutz M., Smalla K. H., Richter K., Landgraf P., Reissner C., Boeckers T. M., Zuschratter W., Spilker C., Seidenbecher C. I., Garner C. C., Gundelfinger E. D., Kreutz M. R. (2008) Biology 6, e34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Alberini C. M. (1999) J. Exp. Biol. 202, 2887–2891 [DOI] [PubMed] [Google Scholar]
- 12.Cabeza-Arvelaiz Y., Thompson T. C., Sepulveda J. L., Chinault A. C. (2001) Oncogene 20, 6707–6717 [DOI] [PubMed] [Google Scholar]
- 13.Wendholt D., Spilker C., Schmitt A., Dolnik A., Smalla K. H., Proepper C., Bockmann J., Sobue K., Gundelfinger E. D., Kreutz M. R., Boeckers T. M. (2006) J. Biol. Chem. 281, 13805–13816 [DOI] [PubMed] [Google Scholar]
- 14.Konno D., Ko J. A., Usui S., Hori K., Maruoka H., Inui M., Fujikado T., Tano Y., Suzuki T., Tohyama K., Sobue K. (2002) J. Cell Sci. 115, 4695–4706 [DOI] [PubMed] [Google Scholar]
- 15.Ishii H., Vecchione A., Murakumo Y., Baldassarre G., Numata S., Trapasso F., Alder H., Baffa R., Croce C. M. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 10374–10379 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Murillas R., Simms K. S., Hatakeyama S., Weissman A. M., Kuehn M. R. (2002) J. Biol. Chem. 277, 2897–2907 [DOI] [PubMed] [Google Scholar]
- 17.Sudo H., Maru Y. (2007) FASEB J. 21, 2086–2100 [DOI] [PubMed] [Google Scholar]
- 18.Thyssen G., Li T. H., Lehmann L., Zhuo M., Sharma M., Sun Z. (2006) Mol. Cell Biol. 26, 8857–8867 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Moon R. T., Kohn A. D., De Ferrari G. V., Kaykas A. (2004) Nat. Rev. Genet. 5, 691–701 [DOI] [PubMed] [Google Scholar]
- 20.Salinas P. C., Price S. R. (2005) Curr. Opin. Neurobiol. 15, 73–80 [DOI] [PubMed] [Google Scholar]
- 21.Takeichi M., Abe K. (2005) Trends Cell Biol. 15, 216–221 [DOI] [PubMed] [Google Scholar]
- 22.Abe K., Takeichi M. (2007) Neuron 53, 387–397 [DOI] [PubMed] [Google Scholar]
- 23.Boeckers T. M., Bockmann J., Kreutz M. R., Gundelfinger E. D. (2002) J. Neurochem. 81, 903–910 [DOI] [PubMed] [Google Scholar]
- 24.Boeckers T. M. (2006) Cell Tissue Res. 326, 409–422 [DOI] [PubMed] [Google Scholar]
- 25.Sheng M., Kim E. (2000) J. Cell Sci. 113, 1851–1856 [DOI] [PubMed] [Google Scholar]
- 26.Durand C. M., Betancur C., Boeckers T. M., Bockmann J., Chaste P., Fauchereau F., Nygren G., Rastam M., Gillberg I. C., Anckarsäter H., Sponheim E., Goubran-Botros H., Delorme R., Chabane N., Mouren-Simeoni M. C., de Mas P., Bieth E., Rogé B., Héron D., Burglen L., Gillberg C., Leboyer M., Bourgeron T. (2007) Nat. Genet. 39, 25–27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Dresbach T., Hempelmann A., Spilker C., tom Dieck S., Altrock W. D., Zuschratter W., Garner C. C., Gundelfinger E. D. (2003) Mol. Cell Neurosci. 23, 279–291 [DOI] [PubMed] [Google Scholar]
- 28.Seidenbecher C. I., Langnaese K., Sanmartí-Vila L., Boeckers T. M., Smalla K. H., Sabel B. A., Garner C. C., Gundelfinger E. D., Kreutz M. R. (1998) J. Biol. Chem. 273, 21324–21331 [DOI] [PubMed] [Google Scholar]
- 29.Pak D. T., Yang S., Rudolph-Correia S., Kim E., Sheng M. (2001) Neuron 2, 289–303 [DOI] [PubMed] [Google Scholar]
- 30.Spilker C., Acuña Sanhueza G. A., Böckers T. M., Kreutz M. R., Gundelfinger E. D. (2008) J. Neurochem. 104, 187–201 [DOI] [PubMed] [Google Scholar]
- 31.Murase S., Mosser E., Schuman E. M. (2002) Neuron 3, 91–105 [DOI] [PubMed] [Google Scholar]
- 32.Gundelfinger E. D., Boeckers T. M., Baron M. K., Bowie J. U. (2006) Trends Biochem. Sci. 31, 366–373 [DOI] [PubMed] [Google Scholar]
- 33.Drees F., Pokutta S., Yamada S., Nelson W. J., Weis W. I. (2005) Cell 123, 903–915 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nishimura W., Yao I., Iida J., Tanaka N., Hata Y. (2002) J. Neurosci. 22, 757–765 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Al-Hallaq R. A., Conrads T. P., Veenstra T. D., Wenthold R. J. (2007) J. Neurosci. 27, 8334–8343 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yoshikawa S., McKinnon R. D., Kokel M., Thomas J. B. (2003) Nature 422, 583–588 [DOI] [PubMed] [Google Scholar]
- 37.Tamai K., Semenov M., Kato Y., Spokony R., Liu C., Katsuyama Y., Hess F., Saint-Jeannet J. P., He X. (2000) Nature 407, 530–535 [DOI] [PubMed] [Google Scholar]
- 38.Wehrli M., Dougan S. T., Caldwell K., O'Keefe L., Schwartz S., Vaizel-Ohayon D., Schejter E., Tomlinson A., DiNardo S. (2000) Nature 407, 527–530 [DOI] [PubMed] [Google Scholar]
- 39.Ciani L., Salinas P. C. (2005) Nat. Rev. Neurosci. 6, 351–362 [DOI] [PubMed] [Google Scholar]
- 40.Packard M., Mathew D., Budnik V. (2003) Nat. Rev. Neurosci. 4, 113–120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lie D. C., Colamarino S. A., Song H. J., Désiré L., Mira H., Consiglio A., Lein E. S., Jessberger S., Lansford H., Dearie A. R., Gage F. H. (2005) Nature 437, 1370–1375 [DOI] [PubMed] [Google Scholar]
- 42.Caricasole A., Bakker A., Copani A., Nicoletti F., Gaviraghi G., Terstappen G. C. (2005) Biosci. Rep. 25, 309–327 [DOI] [PubMed] [Google Scholar]
- 43.De Ferrari G. V., Moon R. T. (2006) Oncogene 25, 7545–7553 [DOI] [PubMed] [Google Scholar]
- 44.Ataman B., Ashley J., Gorczyca M., Ramachandran P., Fouquet W., Sigrist S. J., Budnik V. (2008) Neuron 57, 705–718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tang S. J. (2007) Synapse 61, 866–868 [DOI] [PubMed] [Google Scholar]
- 46.Speese S. D., Budnik V. (2007) Trends Neurosci. 30, 268–275 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bryja V., Schulte G., Rawal N., Grahn A., Arenas E. (2007) J. Cell Sci. 120, 586–595 [DOI] [PubMed] [Google Scholar]
- 48.Chen J., Park C. S., Tang S. J. (2006) J. Biol. Chem. 281, 11910–11916 [DOI] [PubMed] [Google Scholar]
- 49.Hall A. C., Lucas F. R., Salinas P. C. (2000) Cell 100, 525–535 [DOI] [PubMed] [Google Scholar]
- 50.Tsai I. C., Amack J. D., Gao Z. H., Band V., Yost H. J., Virshup D. M. (2007) Dev. Cell 12, 335–347 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Logan C. Y., Nusse R. (2004) Annu. Rev. Cell Dev. Biol. 20, 781–810 [DOI] [PubMed] [Google Scholar]
- 52.Polakis P. (2007) Curr. Opin. Genet. Dev. 17, 45–51 [DOI] [PubMed] [Google Scholar]
- 53.Baudino T. A., Cleveland J. L. (2001) Mol. Cell Biol. 21, 691–702 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Knoepfler P. S. (2007) Cancer Res. 1, 5061–5063 [DOI] [PubMed] [Google Scholar]
- 55.Knoepfler P. S., Zhang X. Y., Cheng P. F., Gafken P. R., McMahon S. B., Eisenman R. N. (2006) EMBO J. 21, 2723–2734 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Knoepfler P. S., Cheng P. F., Eisenman R. N. (2002) Genes Dev. 15, 2699–2712 [DOI] [PMC free article] [PubMed] [Google Scholar]