Abstract
Cystic fibrosis (CF) is a common recessive disorder caused by >1600 mutations in the CF transmembrane conductance regulator (CFTR) gene. About 13% of CFTR mutations are classified as “splicing mutations,” but for almost 40% of these, their role in affecting the pre-mRNA splicing of the gene is not yet defined. In this work, we describe a new splicing mutation detected in three unrelated Italian CF patients. By DNA analyses and mRNA studies, we identified the c.1002–1110_1113delTAAG mutation localized in intron 6b of the CFTR gene. At the mRNA level, this mutation creates an aberrant inclusion of a sequence of 101 nucleotides between exons 6b and 7. This sequence corresponds to a portion of intron 6b and resembles a cryptic exon because it is characterized by an upstream ag and a downstream gt sequence, which are most probably recognized as 5′- and 3′-splice sites by the spliceosome. Through functional analysis of this splicing defect, we show that this mutation abolishes the interaction of the splicing regulatory protein heterogeneous nuclear ribonucleoprotein A2/B1 with an intronic splicing regulatory element and creates a new recognition motif for the SRp75 splicing factor, causing activation of the cryptic exon. Our results show that the c.1002–1110_1113delTAAG mutation creates a new intronic splicing regulatory element in intron 6b of the CFTR gene exclusively recognized by SRp75.
Cystic fibrosis (CF2; MIM entry #219700) is the most frequent severe autosomic recessive disorder in the European population (1). Indeed, CF affects ∼1 in 2500 births, and ∼1 in 25 individuals are heterozygotes, with marked regional variations (2). CF is caused by mutations of the CF transmembrane conductance regulator (CFTR or ABCC7; MIM #602421) gene, which is also involved in a broad spectrum of phenotypes, including male infertility due to congenital bilateral absence of the vas deferens (3, 4), disseminated bronchiectasis (5, 6), and chronic pancreatitis (7, 8). The mutational spectrum of this disease is made up of >1600 different mutations, 98% of which consist of point mutations or microdeletions/insertions (Cystic Fibrosis Mutation Database). A significant fraction (∼13%) of CFTR mutations are classified as “splicing mutations,” but for almost 40% of these, their role in affecting the pre-mRNA splicing of the gene is not yet defined.
Splicing mutations act by directly disrupting splice sites or by creating new ones. Varied levels of alternative splicing have been detected for some of the splicing mutations (9–11). These differences are most likely due to different utilization of intragenic splicing elements by trans-acting splicing factors (12, 13). cis-Acting elements required for splicing in mammalian cells include short and poorly conserved consensus sequences, which specify the 5′-splice site, branch site, and 3′-splice site. The 3′-splice site is preceded by a polypyrimidine tract of variable length.
The branch point is typically located 18–40 nucleotides upstream of the polypyrimidine tract. These splicing signals are essential but do not contain sufficient information to specify correct splicing patterns, and they alone cannot be distinguished from numerous surrounding cryptic splice sites of similar strength (14). Their efficient recognition, as well as their regulated usage, requires sequences within exons and introns called enhancers and silencers. Pre-mRNA splicing requires the interaction of these critical cis-acting sequences with splicing regulating factors that form the spliceosome. The spliceosome is a highly complex RNA/protein machine consisting of five small nuclear RNAs and >100 proteins such as the SR (serine/arginine-rich) proteins and heterogeneous nuclear ribonucleoproteins (hnRNPs), allowing the fine-tuning of the splicing process (15–17).
In classical splicing defects, the cis-acting elements are mutated, and the effect on splicing can be predicted on the basis of genomic DNA sequence analysis. However, the effect of mutations in non-obvious regulatory elements is more difficult to evaluate directly from genome scanning methodologies. Patient CFTR mRNA analysis represents a good tool for the identification and definition of the pathogenic role of sequence variations not yet defined and splicing defects causing alternative products.
In this work, we describe three unrelated Italian patients affected by CF whose CFTR mRNA contains an insertion of 101 nucleotides between exons 6b and 7 corresponding to a portion of intron 6b. This sequence resembles a cryptic exon because it is characterized by an upstream ag and a downstream gt sequence, which are most probably recognized as 5′- and 3′-splice sites by the spliceosome. We assume that this anomalous transcript is caused by a mutation affecting an intronic splicing regulatory element that either abolishes or creates specific protein-binding domains, causing an alteration of the intronic splicing process. To elucidate the mechanisms involved in the aberrant splicing, we carried out molecular analysis of intron 6b as well as functional studies.
EXPERIMENTAL PROCEDURES
Patients
This study included three unrelated CF patients of Italian origin affected by a classical form of CF, with one undefined CF allele. Clinical features of affected individuals included pancreatic insufficiency, chronic lung disease with Pseudomonas aeruginosa infections, and a sweat test of >60 mmol/liter Cl−. These patients were selected after an extensive second level molecular screening (i.e. denaturing HPLC analysis and direct sequencing for searching for point mutations, breakpoint PCR, and multiple ligation-dependent probe amplification for detection of large deletions) carried out on 441 unrelated Italian CF patients. The CF genotype of the three patients was F508del/unknown, G542X/unknown, and exon 2 deletion (c.545811_164+2186del8108ins182)/unknown (18).
DNA and RNA Analyses
DNA extraction was carried out according to standard procedures (19). Informed consent was obtained in all cases before collecting patients' DNA. Screening for the most common CF mutations was carried out using INNO-LiPA CFTR 19, CFTR 17+Tn update, and CFTR Italian regional kits (Innogenetics). Denaturing HPLC and multiple ligation-dependent probe amplification analyses of all 27 CFTR exons were carried out to detect unknown mutations as well as large rearrangements.
Nasal epithelial cells were collected from two of the patients and from two non-CF control subjects using a Cytobrush. RNA was extracted using TRIzol LS (Invitrogen). cDNA synthesis was performed with a high capacity cDNA archive kit (Applied Biosystems) according to the manufacturer's instructions. The cDNA was amplified in six overlapping fragments as follows: an initial denaturation step at 95 °C for 12 min, followed by touchdown PCR, decreasing the annealing temperature by 0.5 °C every cycle for 14 cycles starting at 94 °C for 20 s, 62 °C for 40 s, and 72 °C for 60 s, followed by 35 cycles of regular PCR at 94 °C for 20 s, 55 °C for 40 s, and 72 °C for 60 s and a final extension step at 72 °C for 10 min.
A fragment of 400 bp of intron 6b including the mutation was amplified with IVS6b primers (forward, 5′-CTGGCAGAACCAGGATTCAAA-3′; and reverse, 5′-TTGGAGTCACTTGTGATCCTC-3′). Sequencing reactions of both amplified cDNA and amplified intron 6b were carried out according to the Sanger dideoxy chain termination method using a BigDye terminator cycle sequencing kit (Applied Biosystems). Sequence analyses were carried out with ABI PRISM 3100 (Applied Biosystems).
Minigene Constructs
The wild-type CFTR sequences consisting of an exon 6b-intron 6b-exon 7 cassette (∼4 kb long) were obtained by amplifying normal genomic DNA with exon 6b (forward, 5′-CACCATGGATCAGAGAGCTGGGAAGATC-3′) and exon 7 (reverse, 5′-CTGTATTTTGTTTATTGCTCC-3′) primers. The forward primer includes a Kozak translation initiation sequence with an ATG initiation codon. To enable directional cloning, the forward primer also contains a CACC sequence at its 5′-end. The PCR product was then cloned into the mammalian expression vector pcDNA3.1D/V5-His-TOPO (Invitrogen). By PCR-mediated site-directed mutagenesis, the mutant construct (i.e. lacking 4 nucleotides in intron 6b) was also created with primers IVS6b4bpdelfor (5′-GTTCTGCACTTGAGAATGAAGCTTTTCTGAATTATACA-3′), IVS6b4bpdelrev (5′-TGTATAATTCAGAAAAGGTTCATTCTCAAGTGCAGAAC-3′), IVS6bWTfor (5′-GTTCTGCACTTGAGAATGAAGCTTTTCTGAATTATACA-3′, and IVS6bWTrev (5′-TGTATAATTCAGAAAAGCTTCATTCTCAAGTGCAGAAC-3′). We also created several variants with single nucleotide changes at the deleted sequence. The oligonucleotides used for the PCR-mediated mutagenesis are available upon request.
Analysis of Minigene Expression
HeLa cells were grown in Dulbecco's modified Eagle's medium. Plasmid DNA was extracted and purified with a HiSpeed plasmid maxi kit (Qiagen). Cells were transfected with Lipofectamine 2000 (Invitrogen) with 3 μg of each reported plasmid (wild-type and mutants) and the control empty vector. RNA extraction was performed after 48 h using TRIzol LS, and reverse transcription (RT)-PCR was done as described above with exon 6b forward and exon 7 reverse primers. All PCR products were sequenced.
RNA Probe Preparation
Two synthetic RNA oligonucleotides, RNA-WT (5′-GCACUUGAGAAUGAGAAUAGCUUUUCUGAA-3′) and RNA-M (5′-GCACUUGAGAAUGAAGCUUUUCUGAAUUAU-3′), were either purchased from Eurofins MWG Operon (Ebersberg, Germany) or synthesized by in vitro transcription using T7 RNA polymerase and DNA oligonucleotide templates: WT, 5′-TAATACGACTCACTATAGGGCACTTGAGAATGAGAATAGCTTTTCTGAA-3′; and M, 5′-TAATACGACTCACTATAGGGCACTTGAGAATGAAGCTTTTCTGAA-3′.
HeLa Nuclear Extracts
HeLa nuclear extracts were obtained using a nuclear extract kit (Active Motif, Rixensart, Belgium) according to the manufacturer's instructions.
RNA Electrophoretic Mobility Shift Assay and UV Cross-linking of Protein and RNA
[α-32P]UTP-labeled RNA probes (1 μg) were synthesized by in vitro transcription for 60 min at 37 °C and treated with DNase for the same period of time. RNA probes were incubated with 5.2 mm HEPES-KOH (pH 7.9), 1 mm MgCl2, 0.8 mm magnesium acetate, 0.52 mm dithiothreitol, 3.8% glycerol, 0.75 mm ATP, 0.5 μg/μl heparin, and 30 μg of HeLa nuclear extract in a final volume of 20 μl for 20 min at room temperature. Following the addition of 5 μl of 50% (v/v) glycerol and tracking dye, the complexes were resolved on a 6% polyacrylamide gel (19:1 acrylamide/bisacrylamide ratio) in 75 mm Tris and 75 mm glycine at 100 V for 3–4 h at 4 °C. the gels were dried and exposed to X-Omat AR films for 3 h. The UV cross-linking assay was performed by incubation of the [α-32P]UTP-labeled RNA probes (1 × 106 cpm/incubation) with 20 μg of HeLa nuclear extract in a 20-μl final volume at 30 °C for 15 min. The final binding conditions were 20 mm HEPES (pH 7.9), 72 mm KCl, 1.5 mm MgCl2, 0.78 mm magnesium acetate, 0.52 mm dithiothreitol, 3.8% glycerol, 0.75 mm ATP, and heparin at a 5 μg/μl final concentration as a nonspecific competitor. The samples were then transferred to the wells of a human leukocyte antigen plate and irradiated with UV light on ice (0.8 kJ for 10 min). Unbound RNA was digested with 30 μg of RNase A by incubation for 37 °C for 30 min. The samples were run on 8–15% SDS-polyacrylamide gels, followed by autoradiography.
Immobilization of RNA on Agarose Beads and Binding Assay (RNA Pulldown)
RNAs were covalently linked to adipic acid dihydrazide-agarose beads by modification of a published procedure (20). Ten micrograms of RNA were placed in 400 μl of reaction mixture containing 100 mm sodium acetate (pH 5.0) and 5 mm sodium m-periodate (Sigma), washed four times with 2 ml of 0.1 m sodium acetate (pH 5.0), and pelleted after each wash at 300 rpm for 3 min. After the final wash, 300 μl of 0.1 m sodium acetate (pH 5.0) were added to the beads, and the slurry was then mixed with the periodate-treated RNA and rotated for 16 h at 4 °C. The beads with the bound RNA was pelleted and washed three times with 1 ml of 2 m NaCl and three times with 1 ml of wash buffer (20 mm HEPES-KOH (pH 7.6), 6% (v/v) glycerol, 0.1 m KCl, 0.2 mm EDTA, and 0.5 mm dithiothreitol).
The beads containing immobilized RNA were incubated in 650 μl of wash buffer containing 600 μg of HeLa cell nuclear extract for 20 min at 30 °C. Beads were then pelleted by centrifugation at 1000 rpm for 3 min and washed four times with 1 ml of wash buffer. After the final centrifugation, the proteins bound to the immobilized RNA were eluted by addition of 20 μl of 4× protein sample buffer and boiled for 5 min.
Protein Analysis
Proteins were separated on 10% SDS-polyacrylamide gels and visualized by silver staining or electroblotted onto nitrocellulose membrane and probed with antibodies. Internal sequence analysis of the silver-stained band excised from the SDS-polyacrylamide gels was performed by the Nurex MALDI time-of-flight identification service (Sassari, Italy). Briefly, bands were excised from SDS gels and destained by washing twice with 50% acetonitrile in 5 mm ammonium bicarbonate and then dried for 30 min in 50% acetonitrile. Dried pieces of gel were subjected to protein digestion by trypsin. Mass spectrometric analysis were performed with a MALDI micro MX spectrometer (Micromass, Manchester, United Kingdom) equipped with a delayed extraction unit according to the tuning procedures suggested by the manufacturer. Sample was loaded onto the MALDI target using 2 ml of the tryptic digest mixed 1:1 with a solution of α-cyano-4-hydroxycinnamic acid (10 mg/ml in 40% acetonitrile and 0.1% (v/v) trifluoroacetic acid). A peak list was generated with ProteinLynx data preparation using the following parameters: external calibration with lock mass using mass 2465.1989 Da of adrenocorticotropic hormone, background subtract-type adaptive combining all scans, and performing deisotoping with a threshold of 5%. A peak list created as described above was used in Mascot using the Swiss-Prot Database (release 54.0, Jan 2008). Search settings allowed one missed cleavage with the trypsin enzyme selected, oxidation of methionine as a potential variable modification, a peptide tolerance of 100 ppm, and taxa human.
Antibodies
Anti-hnRNP A2/B1 monoclonal antibody was from Sigma. SRp75 was identified using a monoclonal antibody that recognizes a subset of non-small nuclear RNP splicing factors (SR proteins) such as SRp75, SRp55, SRp40, and SRp20 (Zymed Laboratories Inc., South San Francisco, CA).
Small Interfering RNA Transfections
Small Interfering RNA (siRNA) transfections were performed in HeLa cells using Lipofectamine RNAiMAX (Invitrogen). We used 1 μm ON-TARGETplus siRNA (Thermo Scientific).
HeLa cells were plated at 2 × 105 cells/well in 10-mm2 plates to achieve 40–50% confluence. The next day, 3 μl of Lipofectamine RNAiMAX were combined with 197 μl of Dulbecco's modified Eagle's medium (Sigma), and 100 μl of 1 μm siRNA was diluted in a final volume of 200 μl of Dulbecco's modified Eagle's medium. The two mixtures were combined and left for 20 min at room temperature. Finally, this mixture was added to the cells, which were maintained in 1.6 ml of Dulbecco's modified Eagle's medium. After 24 h, a second round of siRNA transfection was performed as described above. Eight hours later, the cells were transfected with 2 μg of minigenes and 3 μl of Lipofectamine 2000. On the 3rd day, HeLa cells were harvested and divided in two parts for protein and RNA extractions. RT-PCR of total RNA was performed as described for the transfection protocol described above. Nuclear protein extracts were obtained as described above and analyzed for SRp75 endogenous protein expression by immunoblotting using the antibody described above. Tubulin was used as a protein loading control. Each siRNA treatment was repeated at least three times.
RESULTS
Transcript Analysis of CF Patients
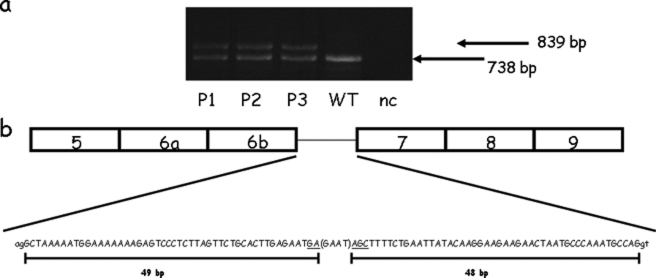
After an extensive second level molecular screening carried out at the DNA level on 441 unrelated Italian CF patients, we selected three patients with one undefined CF allele. To extend their molecular characterization, we performed an RNA study on the selected patients. The analysis showed a transcript with an insertion of 101 bp between exons 6b and 7 that resembles a cryptic exon (Fig. 1). Cryptic or pseudoexons have been shown to be activated by intron mutations that either create or reinforce splice sites or create a new branch site (21–25). The 101-bp inserted sequence is part of intron 6b (from nucleotides 1110 to 1113 upstream of exon 7) and is characterized by an upstream ag and a downstream gt sequence, which are most probably recognized as 5′- and 3′-splice sites by the spliceosome. Sequence analysis of this region in the DNA patients showed a 4-bp deletion most probably affecting an intronic splicing silencer. The deletion is located 49 bp downstream and 48 bp upstream of the 5′- and 3′-ends of the postulated cryptic exon, respectively. Inclusion of the cryptic exon out of the reading frame is predicted to lead to the insertion of nine amino acid residues followed by a stop codon. This means that the CFTR protein produced by this anomalous transcript should be truncated soon after exon 6b. According to recommendations for the description of sequence variants, the name of this new mutation is c.1002–1110_1113delTAAG (26).
FIGURE 1.
Splicing-processing defect in CFTR intron 6b does not affect canonical splicing regulatory elements. a, RT-PCR products obtained from RNA patients (P1–P3), the normal control (WT), and the negative control (nc) spanning exons 5–9 show an aberrant splicing event (101-bp insertion). b, sequence and localization of the CFTR cryptic exon in the patients' cDNA. The new sequence created by the GAAT deletion, located 49 bp downstream and 48 bp upstream of the 5′- and 3′-ends of the cryptic exon, respectively, is underlined.
Intronic 4-bp Deletion in the CFTR Gene Affects an Intronic Splicing Regulatory Element Causing Aberrant Splicing
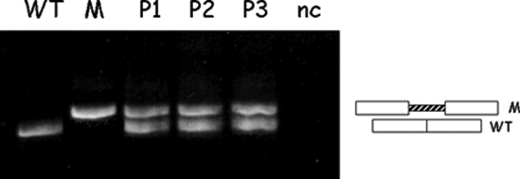
To demonstrate that the 4-bp deletion in CFTR intron 6b is the cause of aberrant splicing, we created two minigenes. We cloned the wild-type construct, spanning exon 6b, the entire intron 6b, and exon 7 of the CFTR gene (∼4 kb long), in a mammalian expression vector. By site-directed mutagenesis, we also created a mutant construct. HeLa cells were transfected with each construct (wild-type and mutant) and the control empty vector. We then performed RNA extraction and RT-PCR. The analysis showed that the wild-type minigene generated a transcript in which exons 6b and 7 were efficiently and accurately spliced, as expected. Conversely, the mutated minigene produced an abnormal transcript showing cryptic exon inclusion, as we found in the patients (Fig. 2).
FIGURE 2.
c.1002–1110_1113delTAAG mutation causes aberrant inclusion of the cryptic exon. Shown are the results from RT-PCR of RNA extracted from HeLa cells transfected with either the WT or mutated minigene. The WT minigene generates a transcript in which exons 6b and 7 are efficiently and accurately spliced. The mutated minigene (M) produces an abnormal transcript, showing cryptic exon inclusion, as we found in the patients (P1–P3). nc, HeLa cells transfected with the empty vector as a negative control.
4-bp Deletion in CFTR Intron 6b Creates a New Sequence with Enhancer Properties (Abolishes a Sequence with Silencer Properties)
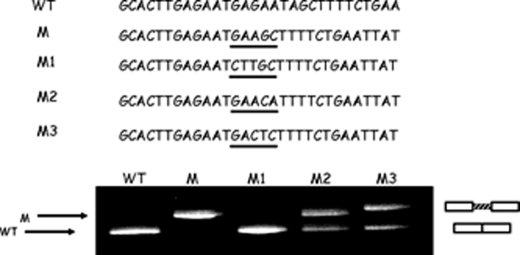
To understand the effects of the mutation on the efficiency of splicing, we performed three rounds of site-directed mutagenesis at the new sequence GAAGC (M), created after the GAAT deletion. Point mutations were introduced in the mutated expression vector (M) and evaluated by the functional splicing assay (Fig. 3). The first round of mutagenesis, M1 (consisting of the GAA → CTT substitution), caused a complete restoration of normal transcripts. Conversely, the two other rounds of mutagenesis, M2 and M3 (consisting of the GC → CA and AG → CT substitutions, respectively), caused only a partial restoration of normal transcripts. The presence of a GAA element in that position of intron 6b maybe plays a critical role in splicing efficiency. In fact, it has been shown previously that substitutions of adenines with uracils in GAA elements, as in M1, eliminate their enhancing ability (27). Probably, for this reason, substitutions in M2 and M3, by affecting GC nucleotides, have moderate effects on the splicing process. Hence, these results indicate that the c.1002–1110_1113delTAAG mutation most probably removes or weakens an element with silencer properties and creates a sequence with enhancer ability, resulting in a very high level of cryptic exon inclusion (Fig. 3).
FIGURE 3.
4-bp deletion in CFTR intron 6b creates a new sequence with enhancer properties. Shown is the RT-PCR analysis of HeLa cells transfected with minigenes carrying WT and mutant (M) constructs and mutants M1–M3. Multiple point mutations M1–M3 were introduced in the mutated expression vector and evaluated by functional splicing assay. M1 mutagenesis completely restored a correct splicing process; in contrast, M2 and M3 mutagenesis caused an alternative splicing process. Only the sequence created by the c.1002–1110_1113delTAAG mutation resulted in a very high level of cryptic exon inclusion.
4-bp Deletion Creates New Binding with a trans-Acting Protein
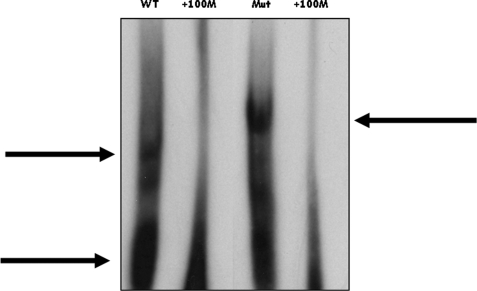
To identify the trans-acting factors possibly binding this intronic region, electrophoretic mobility shift assay was performed. We used in vitro transcribed RNAs including either the wild-type and mutant sequence. The RNAs were incubated with HeLa nuclear extract in the presence of the nonspecific competitor heparin. As shown in Fig. 4, the analysis showed two different shifted bands. The nature of the protein(s) that bind(s) to the mutant sequence was then investigated.
FIGURE 4.
Reactivity in electrophoretic mobility shift assay of HeLa nuclear extracts with WT and mutant labeled RNA oligonucleotides. Different complexes were formed in the presence of the WT or mutant (Mut) RNA (upper arrows). Competition analysis was done by adding unlabeled RNA oligonucleotides at a 100-fold molar excess (+100M). The lower arrow indicates free nucleic acids.
Identification of Nuclear Proteins That Bind to Wild-type and Mutant CFTR Intron 6b Elements
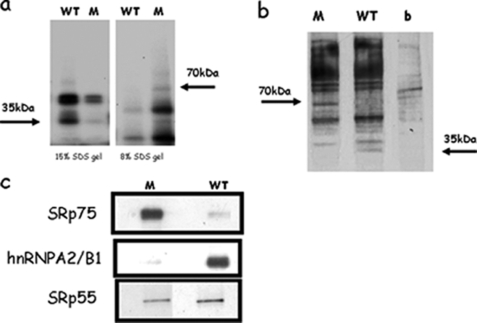
UV cross-linking experiments were done on wild-type and mutant CFTR intron 6b. The mutant and wild-type (WT) sequences were transcribed in vitro in the presence of [α-32P]UTP, and equal quantities of labeled transcripts were then used in a UV cross-linking assay with HeLa nuclear extracts. Among the numerous proteins that could be cross-linked to the labeled RNAs, an ∼35-kDa band could be observed only in the WT RNA, and an ∼75-kDa band could be observed only in the mutant RNA (Fig. 5a).
FIGURE 5.
Identification of nuclear proteins regulating the splicing process of intron 6b and involved in the aberrant splicing caused by the 4-bp deletion. a, UV cross-linking experiments carried out on WT and mutant CFTR intron 6b. WT and mutant (M) labeled RNAs cross-reacted with ∼35- and ∼75-kDa bands, respectively (arrows). Cross-reaction with the WT and mutant labeled probes was detected on 15 and 8% SDS-polyacrylamide gels, respectively. b, silver-stained 10% SDS-polyacrylamide gel of a pulldown analysis in which HeLa nuclear extracts were incubated with adipic dihydrazide beads derivatized with the target RNAs. The arrows indicate two bands of ∼35 kDa present in the WT lane and of ∼70 kDa present only in the M lane. b lane, beads not linked to the RNA probes as a negative control. c, binding analysis of hnRNP A2/B1 and SRp75 performed by Western blotting after pulldown assay. Binding of SRp55 showed no variations between WT and mutant RNAs.
To identify the trans-acting factors differentially binding to WT and mutant CFTR intron 6b sequences, we performed a pulldown analysis using two synthetic RNA oligonucleotides containing either the WT or mutant sequence. Agarose beads linked to the WT and mutant RNAs were separately incubated with HeLa nuclear extracts, and the associated proteins were analyzed by SDS-PAGE, followed by silver staining. Comparison of the binding patterns of the WT and mutant beads showed that the two patterns have many bands in common, but in the molecular mass range of 30–35 kDa, a protein was pulled down by WT RNA but not by the mutant one. Conversely, at 70–75 kDa, a protein complex was pulled down by mutant RNA but not by the WT one (Fig. 5b). Sequence analysis of the excised bands identified the binding proteins. The protein binding to the WT sequence is hnRNP A2/B1. The protein binding to the mutant sequence is SRp75. These results were confirmed by RNA pulldown experiments. WT and mutant synthetic RNA oligonucleotides were incubated with HeLa nuclear extracts, and the complexes were resolved by SDS-PAGE, followed by Western blotting using antibodies hnRNP A2/B1 and SRp75 (Fig. 5c). Binding of the well known specific factor SRp55 showed no variations between WT and mutant RNAs even though dedicated bioinformatics tools such as ESEfinder had suggested its possible involvement with mutant RNA (Fig. 5c).
hnRNP A2/B1 and SRp75 Overexpression in Transfection Assays Does Not Modify the Splicing Pattern
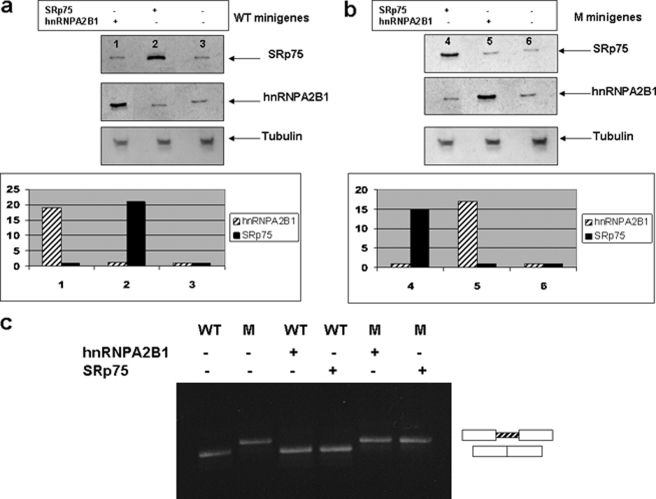
To evaluate the functional significance of the interactions of the splicing factors in the CFTR intron 6b splicing, both WT and mutant HeLa cells were transfected with plasmids coding for hnRNP A2/B1 and SRp75. Western blotting using the antibodies against the target proteins was carried out to test the overexpression achieved (Fig. 6, a and b).
FIGURE 6.
Overexpression of hnRNP A2/B1 and SRp75 in mutant and WT cells does not modify the slicing pattern. a, Western blot analysis of HeLa cells showing the overexpression achieved after co-transfection of the WT minigene and hnRNP A2/B1 (lane 1) or the WT minigene and SRp75 (lane 2). Lane 3, HeLa cells transfected with the WT minigene alone. b, Western blot analysis of HeLa cells showing the overexpression achieved after co-transfection of the mutated minigene and hnRNP A2/B1 (lane 4) or the mutated minigene and SRp75 (lane 5). Lane 6, HeLa cells transfected with the mutated minigene alone. The graphs in a and b show the level of expression of hnRNP A2/B1 and SRp75. The y axes represent –fold changes. Tubulin was used as a protein loading control. S.D. was not detectable. c, effect of overexpression of hnRNP A2/B1 and SRp75 on the splicing pattern of the WT and mutated (M) minigenes. The WT and mutated minigenes were transfected alone (−) or co-transfected (+) with hnRNP A2/B1 or SRp75. The resulting RT-PCR products were separated on 2% agarose gel.
Overexpression of both hnRNP A2/B1 and SRp75 in the WT cells did not show any variation in the splicing pattern. Moreover, overexpression of hnRNP A2/B1 in the mutant cells and of SRp75 did not modify the slicing pattern (Fig. 6c). These results indicate that the normal sequence in intron 6b contains a strong splicing regulating element. In addition, the new intronic motif created by the mutation shows a strong enhancer activity.
siRNA against SRp75 Affects CFTR Intron 6b Splicing
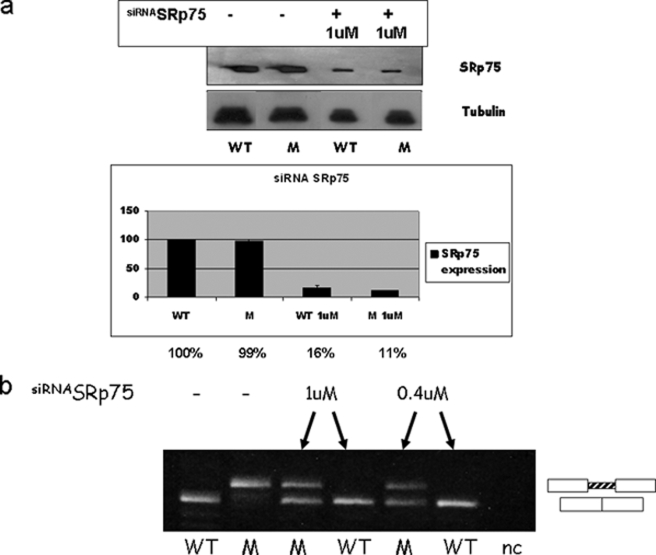
To evaluate the functional role of SRp75 in the in vivo splicing of WT and mutant CFTR intron 6b, we used siRNA against this splicing factor. We transiently transfected HeLa cells, and the silencing effects were analyzed by Western blotting using the antibody against the target protein. HeLa cells transfected with the corresponding siRNA showed a strong reduction in SRp75 levels (Fig. 7a). At the same time, the splicing pattern was evaluated by RT-PCR using total RNA. In the mutant cells, the siRNA-mediated knockdown of SRp75 resulted in partial restoration of CFTR intron 6b splicing. The correct splicing of intron 6b in the WT cells was not affected by siRNA-mediated depletion of SRp75 (Fig. 7b).
FIGURE 7.
In vivo depletion of SRp75 partially restores correct CFTR intron 6b splicing. a, Western blot analysis of HeLa cells transfected with the WT and mutated (M) minigenes treated with (+) and without (−) 1 μm SRp75 siRNA. The graph shows the expression SRp75 after siRNA treatment in WT and mutant cells compared with untreated cells. Tubulin was used as a protein loading control. b, effect of siRNA against SRp75 on the WT and mutated minigenes. Untreated (−) and siRNA-treated (+) HeLa cells were transfected with the mutated and WT minigenes. Two different amounts (1 and 0.4 μm) of siRNA were used. nc, negative control.
DISCUSSION
CF is a common recessive disorder caused by >1600 mutations in the CFTR gene. Despite the extensive mutation characterization work around the world, there is still a substantial fraction of patients with typical clinical diagnosis of CF but unknown mutations in the CFTR gene (28). About 13% of CFTR mutations are termed as affecting the pre-mRNA splicing, but for almost 40% of these, their role in affecting the pre-mRNA splicing is not yet defined. Genotype screening in the CFTR gene frequently identifies sequence variations whose association with the disease phenotype is unclear. In fact, the pathologic effect of an apparently benign polymorphism is difficult to assess. For this reason, patient CFTR mRNA analysis represents a good tool for the identification and definition of the pathogenic role of sequence variations not yet defined and splicing defects causing alternative products.
In this study, we have described a new splicing mutation detected in three unrelated Italian CF patients. The molecular characterization of these patients was carried out at either the DNA or RNA level. The splicing defect described here abolishes the interaction of hnRNP A2/B1 with an intronic splicing regulatory element and creates a new recognition motif for the SRp75 splicing factor, causing the activation of a cryptic exon. To our knowledge, this is the first time that the SR splicing protein SRp75 has been directly implicated in the creation of a cryptic exon in the CFTR transcript.
SR proteins (15) are typically involved in positive regulation of splicing, stimulating splicing by interacting with exonic or intronic splicing enhancer elements. In contrast, negative regulation is promoted most frequently by hnRNPs (29), which function by binding sequences known as exonic or intronic splicing silencers. Several hnRNPs have been identified as key splicing repressors. Among these, the hnRNP A1 protein has been extensively characterized. The first hnRNP A1-dependent exonic splicing silencer was identified in studies of human immunodeficiency virus Tat exon 2 repression (30, 31). This exonic splicing silencer was found to bind hnRNP A1, and mutations disrupting the exonic splicing silencer prevented hnRNP A1 binding and allowed enhanced exon 2 splicing.
Despite the abundance in the genome of potential pseudoexons (intronic sequences between 50 and 200 nucleotides in length with apparently possible 5′- and 3′-splice sites at either end), their inclusion does not seem to be a frequent event during the normal pre-mRNA processing. It was indeed noted that many of these non-spliced pseudoexons contain multiple defects in splicing-controlling regions despite their apparently good agreement with real exonic sequences (32). In accord with this, three putative splicing silencer motifs have been found to be enriched in these non-spliced pseudoexons compared with their surrounding intronic regions (33). This observation is in agreement with a study that showed that human DNA is particularly rich in sequences that can inhibit splicing (34).
Many intronic sequences seem to be close to becoming exons because most of the disease-associated pseudoexon inclusion events originate from a single activating mutation. This mutation generally involves the ex novo creation of a strong donor or acceptor site, followed by the subsequent selection of weaker opportunistic acceptor or donor site sequences. However, other mechanisms that cause the inclusion of pseudoexon-activating mutations may also involve the creation or deletion of splicing regulatory elements.
There are in fact at least two cases, the α-galactosidase A (35) and ATM (36) genes, in which the creation/deletion of splicing regulatory elements has been observed to occur. In the ATM gene, the inclusion of a 65-nucleotide long pseudoexon is the consequence of a 4-nucleotide deletion in ATM intron 20 that disrupts a high affinity U1 small nuclear ribonucleoprotein-binding site located within the pseudoexon sequence itself. This region is called the intronic splicing processing element, and its disruption results in the activation of an immediately upstream cryptic acceptor site and of a downstream cryptic donor site.
The splicing mutation affecting the CFTR gene described in this work brings to mind the splicing defect in the ATM gene described by Pagani et al. (36). In summary, our results indicate that the c.1002–1110_1113delTAAG mutation creates a new intronic splicing regulatory element in intron 6b of the CFTR gene specifically recognized by the splicing factor SRp75. This acts as enhancer that increases the pseudoexon inclusion during the splicing process.
We have established that the basic mechanism affecting splicing due to mutation is not always obvious and cannot be simply derived from in silico predictions. The creation of an intronic splicing enhancer may be a common event in human pathology and has to be taken into account to assess the phenotypic risk in CF patients and to devise new therapies.
Acknowledgments
We thank Dr. Rosaria Casciaro (Istituto G. Gaslini, Clinica Pediatrica, Centro Fibrosi Cistica, Genoa, Italy) and Prof. Carla Colombo (Centro Regionale di Riferimento, Fibrosi Cistica, Milan, Italy) for clinical evaluation.
This work was supported by an Italian Cystic Fibrosis Research Foundation grant (to M. C. R.), Project FFC24/2006, adopted by Silvana Ratti (Pesaro, Italy).
- CF
- cystic fibrosis
- CFTR
- CF transmembrane conductance regulator
- hnRNP
- heterogeneous nuclear ribonucleoprotein
- HPLC
- high pressure liquid chromatography
- RT
- reverse transcription
- MALDI
- matrix-assisted laser desorption ionization
- siRNA
- small interfering RNA
- WT
- wild-type.
REFERENCES
- 1.Welsh M. J., Tsui L. C., Boat T. F., Beaudet A. I. (1995) in Cystic Fibrosis: The Metabolic and Molecular Bases of Inherited Disease ( Scriver C. R., Beaudet A. I., Sly W. S., Valle D. eds) pp. 3799–3876, McGraw-Hill Book Co., New York [Google Scholar]
- 2.Estivill X., Bancells C., Ramos C. (1997) Hum. Mutat. 10, 135–154 [DOI] [PubMed] [Google Scholar]
- 3.Chillón M., Casals T., Mercier B., Brassas L., Lissens W., Silber S., Romey M. C., Ruiz-Romero J., Verlingue C., Claustres M. (1995) N. Engl. J. Med. 332, 1475–1480 [DOI] [PubMed] [Google Scholar]
- 4.Costes B., Girodon E., Ghanem N., Flori E., Jardin A., Soufir J. C., Goossens M. (1995) Eur. J. Hum. Genet. 3, 285–293 [DOI] [PubMed] [Google Scholar]
- 5.Pignatti P. F., Bombieri C., Marigo C., Benetazzo M., Luisetti M. (1995) Hum. Mol. Genet. 4, 635–639 [DOI] [PubMed] [Google Scholar]
- 6.Girodon E., Cazeneuve C., Lebargy F., Chinet T., Costes B., Ghanem N., Martin J., Lemay S., Scheid P., Housset B., Bignon J., Goossens M. (1997) Eur. J. Hum. Genet. 5, 149–155 [PubMed] [Google Scholar]
- 7.Cohn J. A., Friedman K. J., Noone P. G., Knowles M. R., Silverman L. M., Jowel P. S. (1998) N. Engl. J. Med. 339, 653–658 [DOI] [PubMed] [Google Scholar]
- 8.Sharer N., Schwarz M., Malone G., Howarth A., Painter J., Super M., Brazanga J. (1998) N. Engl. J. Med. 339, 645–652 [DOI] [PubMed] [Google Scholar]
- 9.Zielenski J., Patrizio P., Corey M., Handelin B., Markiewicz D., Asch R., Tsui L. C. (1995) Am. J. Hum. Genet. 57, 958–960 [PMC free article] [PubMed] [Google Scholar]
- 10.Chiba-Falek O., Kerem E., Shoshani T., Aviram M., Augarten A., Bentur L., Tal A., Tullis E., Rahat A., Kerem B. (1998) Genomics. 53, 276–283 [DOI] [PubMed] [Google Scholar]
- 11.Chiba-Falek O., Parad R. B., Kerem E., Kerem B. (1999) Am. J. Respir. Crit. Care Med. 159, 1998–2002 [DOI] [PubMed] [Google Scholar]
- 12.Nissim-Rafinia M., Chiba-Falek O., Sharon G., Boss A., Kerem B. (2000) Hum. Mol. Genet. 9, 1771–1778 [DOI] [PubMed] [Google Scholar]
- 13.Pagani F., Buratti E., Stuani C., Romano M., Zuccato E., Niksic M., Giglio L., Faraguna D., Baralle F. E. (2000) J. Biol. Chem. 275, 21041–21047 [DOI] [PubMed] [Google Scholar]
- 14.Cartegni L., Chew S. L., Krainer A. R. (2002) Nat. Rev. Genet. 3, 285–298 [DOI] [PubMed] [Google Scholar]
- 15.Graveley B. R. (2000) RNA 6, 1197–1211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Nilsen T. W. (2003) BioEssays 25, 1147–1149 [DOI] [PubMed] [Google Scholar]
- 17.Zhou Z., Licklider L. J., Gygi S. P., Reed R. (2002) Nature 419, 182–185 [DOI] [PubMed] [Google Scholar]
- 18.Faà V., Bettoli P. P., Demurtas M., Zanda M., Ferri V., Cao A., Rosatelli M. C. (2006) J. Mol. Diagn. 8, 499–503 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Miller S. A., Dykes D. D., Polesky H. F. (1988) Nucleic Acids Res. 16, 1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Caputi M., Mayeda A., Krainer A. R., Zahler A. M. (1999) EMBO J. 18, 4060–4067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Vervoort R., Gitzelmann R., Lissens W., Liebaers I. (1998) Hum. Genet. 103, 686–693 [DOI] [PubMed] [Google Scholar]
- 22.Chillón M., Dörk T., Casals T., Giménez J., Fonknechten N., Will K., Ramos D., Nunes V., Estivill X. (1995) Am. J. Hum. Genet. 56, 623–629 [PMC free article] [PubMed] [Google Scholar]
- 23.Wang M., Dotzlaw H., Fuqua S. A., Murphy L. C. (1997) Breast Cancer Res. Treat. 44, 145–151 [DOI] [PubMed] [Google Scholar]
- 24.Highsmith W. E., Burch L. H., Zhou Z., Olsen J. C., Boat T. E., Spock A., Gorvoy J. D., Quittel L., Friedman K. J., Silverman L. M.(1994) N. Engl. J. Med. 331, 974–980 [DOI] [PubMed] [Google Scholar]
- 25.Ars E., Serra E., García J., Kruyer H., Gaona A., Lázaro C., Estivill X. (2000) Hum. Mol. Genet. 9, 237–247 [DOI] [PubMed] [Google Scholar]
- 26.den Dunnen J. T., Antonarakis S. E. (2000) Hum. Mutat. 15, 7–12 [DOI] [PubMed] [Google Scholar]
- 27.Yeakley J. M., Morfin J. P., Rosenfeld M. G., Fu X. D. (1996) Proc. Natl. Acad. Sci. U.S.A. 93, 7582–7587 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Groman J. D., Meyer M. E., Wilmott R. W., Zeitlin P. L., Cutting G. R. (2002) N. Engl. J. Med. 347, 401–407 [DOI] [PubMed] [Google Scholar]
- 29.Dreyfuss G., Kim V. N., Kataoka N. (2002) Nat. Rev. Mol. Cell Biol. 3, 195–205 [DOI] [PubMed] [Google Scholar]
- 30.Amendt B. A., Si Z. H., Stoltzfus C. M. (1995) Mol. Cell. Biol. 11, 4606–4615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Staffa A., Cochrane A. (1995) Mol. Cell. Biol. 8, 4597–4605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sun H., Chasin L. A. (2000) Mol. Cell. Biol. 20, 6414–6425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fairbrother W. G., Chasin L. A. (2000) Mol. Cell. Biol. 20, 6816–6825 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sironi M., Menozzi G., Riva L., Cagliani R., Comi G. P., Bresolin N., Giorda R., Pozzoli U. (2004) Nucleic Acids Res. 32, 1783–1791 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ishii S., Nakao S., Minamikawa-Tachino R., Desnick R. J., Fan J. Q. (2002) Am. J. Hum. Genet. 70, 994–1002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Pagani F., Buratti E., Stuani C., Bendix R., Dörk T., Baralle F. E. (2002) Nat. Genet. 30, 426–429 [DOI] [PubMed] [Google Scholar]