Abstract
Fe2+ is now shown to weaken binding between ferritin and mitochondrial aconitase messenger RNA noncoding regulatory structures ((iron-responsive element) (IRE)-RNAs) and the regulatory proteins (IRPs), which adds a direct role of iron to regulation that can complement the well known regulatory protein modification and degradative pathways related to iron-induced mRNA translation. We observe that the Kd value increases 17-fold in 5′-untranslated region IRE-RNA·repressor complexes; Fe2+, is studied in the absence of O2. Other metal ions, Mn2+ and Mg2+ have similar effects to Fe2+ but the required Mg2+ concentration is 100 times greater than for Fe2+ or Mn2+. Metal ions also weaken ethidium bromide binding to IRE-RNA with no effect on IRP fluorescence, using Mn2+ as an O2-resistant surrogate for Fe2+, indicating that metal ions bound IRE-RNA but not IRP. Fe2+ decreases IRP repressor complex stability of ferritin IRE-RNA 5–10 times compared with 2–5 times for mitochondrial aconitase IRE-RNA, over the same concentration range, suggesting that differences among IRE-RNA structures contribute to the differences in the iron responses observed in vivo. The results show the IRE-RNA·repressor complex literally responds to Fe2+, selectively for each IRE-mRNA.
Iron (e.g. ferrous sulfate, ferric citrate, and hemin) added to animal cells changes translation rates of messenger RNAs encoding proteins of iron traffic and oxidative metabolism (1–4). To cross cell membranes, iron ions are transported by membrane proteins such as DMT1 or carried on proteins such as transferrin. Inside the cells, iron is mainly in heme, FeS clusters, non-heme iron cofactors of proteins, and iron oxide minerals coated by protein nanocages (ferritins). Iron in transit is thought to be Fe2+ in labile “pools” accessible to small molecular weight chelators, and/or bound loosely by chaperones.
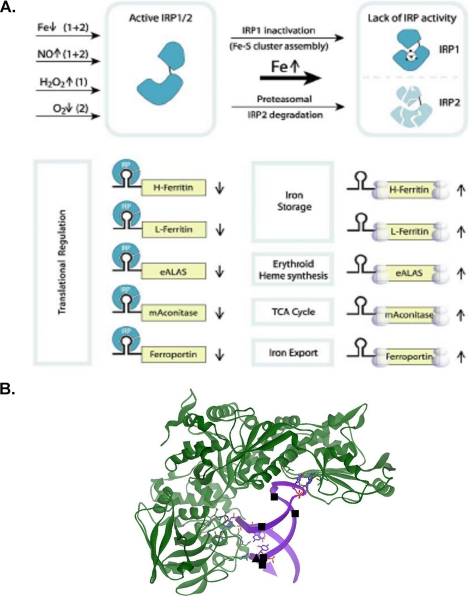
When iron concentrations in the cells increase, a group of mRNAs with three-dimensional, noncoding structures in the 5′-untranslated region (UTR)3 are derepressed (Fig. 1A), i.e. the fraction of the mRNAs in mRNA·repressor protein complexes, which inhibit ribosome binding, decreases and the fraction of the mRNAs in polyribosomes increases (5–7). The three-dimensional, noncoding mRNA structure, representing a family of related structures, is called the iron-responsive element, or IRE, and the repressors are called iron regulatory proteins (IRPs). Together they are one of the most extensively studied eukaryotic messenger RNA regulatory systems (1–4). In addition to large numbers of cell studies, structures of IRE-RNAs are known from solution NMR (8–12), and the RNA·protein complex from x-ray crystallography (13). Recent data indicate that demetallation of IRP1 and disruption of the [4Fe-4S] cluster that inhibits IRP1 binding to RNA will be enhanced by phosphorylation and low iron concentrations (1, 2, 14–16). Such results can explain the increased IRP1 binding to IRE-mRNAs and increased translational repression when iron concentrations are abnormally low. However, the mechanism to explain dissociation of IRE-RNA·IRP complexes, thereby allowing ribosome assembly and increased proteosomal degradation of IRPs (1, 2, 14, 15) (Fig. 1A), when high iron concentrations are abnormally high, is currently unknown.
FIGURE 1.
IRE-RNA·IRP complexes and a model for depression by excess iron. A, a representative model of iron-induced translation of 5′-UTR IRE-RNAs. This figure is modified from Ref. 7. B, IRE-RNA sites influenced by metal binding related to the crystal structure of the ferritin-IRE-RNA·IRP complex from Ref. 13. The figure was created by T. Tosha using Discovery Studio 1.6 and Protein Data Bank file 2IPY. ■, hydrated Mg2+, determined by solution NMR; ▴, Cu1+-1.10-phenanthroline, determined by RNA cleavage in O2.
Metal ion binding changes conformation and function of most RNA classes, e.g. rRNA (17), tRNA (18, 19), ribozymes (20–23), riboswitches (24, 25), possibly hammerhead mRNAs in mammals (26), and proteins. Although the effects of metal ion binding on eukaryotic mRNAs have not been extensively studied, Mg2+ is known to cause changes in conformation, shown by changes in radical cleavage sites of IRE-RNA with 1,10-phenanthrolene-iron and proton shifts in the one-dimensional NMR spectrum (12, 27). The Mg2+ effects are observed at low magnesium concentrations (0.1–0.5 mm) and low molar stoichiometries (1:1 and 2:1 = Mg:RNA).
We hypothesized that Fe2+ could directly change the binding of the IRE-mRNA to the iron regulatory protein for several reasons. First, other metal ions influence the IRE-RNA structure (12, 27). Second, in IRE-RNA/IRP cocrystals there are exposed RNA sites in the IRE-RNA/IRP complex that are accessible for interactions (13) (Fig. 1B). Third, regions in the IRE-RNA are hypersensitive to Fe2+-EDTA/ascorbate/H2O2, suggesting selective interactions with metals and/or solvent (28). We now report that Fe2+ weakens IRE-RNA/IRP binding, whereas Mg2+ requires 100 times the concentration and Mn2+ is comparable with Fe2+; the Fe2+ effect was diminished in mutant IRE-RNA and IRE-RNA selective in wild type sequences: ferritin IRE-RNA > mt-aconitase IRE-RNA.
EXPERIMENTAL PROCEDURES
Preparation of Binding Proteins and RNA
Isolation of recombinant rabbit IRP1 from yeast used previously described methods (29). RNA oligonucleotides for frog ferritin H, frog ferritin H ΔU6, and mt-aconitase (30, 29, and 29 nucleotides, respectively) were purchased from Genelink (Hawthorne, NY). The frog H (FerH) IRE-RNA was used as a model because of extensive structural information, e.g. “footprinting,” solution NMR, x-ray diffraction of cocrystals with IRP, and direct comparisons between natural, poly(A), 5′-UTR-IRE mRNA, with full-length transcripts and RNA aptamers (28, 30). After dissolving in 40 mm HEPES/KOH, pH 7.2, RNA was melted and annealed as described (31), by heating to 85 °C for 15 min with slow cooling to 25 °C. Melting and annealing decreased the Kd by ∼5-fold for the ferritin-IRE-RNA but had little effect on the mt-aconitase IRE-RNA.
Analysis of Fluorescence Data
Protein fluorescence intensities (332 nm (280 nm excitation)) were corrected for dilution, as needed, and for inner filter effects; maximum dilutions were <7%. Nonlinear least squares fitting of the data used KaleidaGraph software (version 2.1.3; Abelbeck Software). Ethidium bromide fluorescence was measured at 595 nm, with excitation at 510 nm. Scatchard analysis of EtBr binding used KaleidaGraph software.
Effect of Metals on RNA·Protein Complexes, RNA, and Protein
Solution
Solutions of Fe2+-O2, Mg2+, or Mn2+ were added to both RNA and protein solutions at the same concentrations, and the solutions were incubated separately for 15 min before adding to binding buffer that contained the same metal ion concentration as the RNA and protein solutions. RNA and protein mixtures were incubated in binding buffer (29), 40 mm HEPES/K+, pH 7.2, 100 mm KCl, 5% glycerol, and 2% 2-mercaptoethanol, for 15 min at 25 °C, before making fluorescence measurements. When Fe2+ was used, all incubations were anaerobic. The protein concentration was 0.1 μm and the RNA concentration varied from 0 to 1.0 μm; because of tight binding, 0.05 μm protein was also analyzed for ferritin RNA titrations to determine Kd. The order of addition to the binding buffer had no effect on the results. In the case of Fe2+, nitrogen-purged solutions of 0.1 m HCl used to dissolve FeSO4 and inhibit oxidation were diluted to 0.001 m H+ and further diluted 1:100 into the RNA or protein solutions.
Gels
Solutions of RNA and protein ± Fe2+-O2 were prepared as in the solution studies. Free and bound RNA was resolved by electrophoresis (EMSA) in 1% agarose gels followed by staining with ethidium bromide. The RNA concentrations were 0.1 μm and protein concentrations varied from 0 to 2.0 μm.
RESULTS
Ferrous Ions (-O2) Weaken Ferritin and mt-Aconitase IRE-RNA·IRP Repressor Complexes
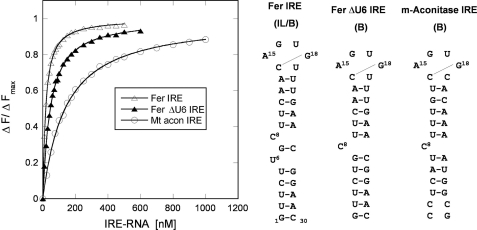
We selected two IRE-mRNAs to study, ferritin and mt-aconitase, because iron induces the synthesis of both proteins in whole animals and cells and because the iron responses are different: iron induces ferritin synthesis in the liver more than ∼150-fold and in mt-aconitase ∼4-fold (32). The two IRE-RNA structures are different (Fig. 2) as is the binding of IRP2 and IRP1 in the electrophoretic mobility shift assay (31). When the Kd for IRP1 binding to the two IRE-RNAs was determined by protein fluorescence quenching, the values differed 10-fold (Figs. 2 and 3).
FIGURE 2.
IRP1 preferentially binds to the ferritin (Fer H IRE-RNA) compared with mt-aconitase IRE-RNA, with nm affinity. IRE-RNA (30 nucleotides long) was melted and annealed prior to each titration and incubated with IRP1 protein in binding buffer, as previously described (29) (see “Experimental Procedures”). Concentrations were: RNA, 0–1.0 μm; protein, 0.1 μm. The same protein concentration was used for all three titrations for comparison purposes in the figure. However, for data fitting, a lower concentration of protein (0.05 μm) was used for ferritin RNA titrations in the absence of metal ions. RNA binding to protein was measured as decreased fluorescence intensity with excitation at 280 nm and emission at 332 nm (IRP1 protein) (at 25 °C). Data were analyzed as previously described (45, 46). The data points are the averages of three titrations and the solid lines are the fitted curves.
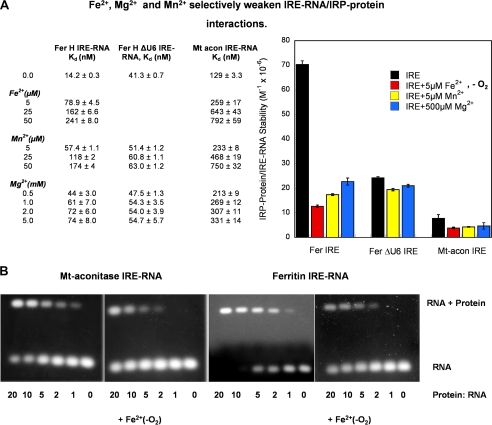
FIGURE 3.
Fe2+ selectively weakens IRE-RNA/IRP1 interactions. A, IRE-RNA binding to IRP was measured, ±Fe2+-O2, as IRP1 fluorescence quenching in solution (constant protein concentration) as described in the legend to Fig. 2 (see “Experimental Procedures”). The IRE-RNAs used were FerH IRE-RNA, FerH ΔU6 IRE-RNA, and mt-aconitase IRE-RNA. Data were analyzed as previously described (45, 46). The Kd values are the averages of three titrations. Relative stabilities (1/Kd) of the IRE-RNA·IRP1 complexes are used in the graph, right. B, IRE-RNA binding to IRP ± Fe2+-O2, was measured by electrophoretic mobility shift assay (constant RNA concentration). Anaerobic solutions of RNA and protein were prepared as in A; Fe2+ was 50 μm when present (see “Experimental Procedures”). A constant 0.1 μm RNA concentration was used with varying concentrations of protein; RNA was stained with ethidium bromide. Note the larger amount of IRP1 repressor required to bind (shift) an equivalent amount of mt-aconitase IRE-RNA compared with ferritin IRE-RNA ± Fe2+-O2. The data are representative of two independent experiments and the significance of the effect of metal ions on the Kd values is p < 0.01.
Fe2+, 5 μm, increased the Kd for ferritin IRE-RNA 6-fold and mt-aconitase RNA 2-fold (Fig. 3A), showing that Fe2+ weakens IRE-RNA/IRP1 binding and suggesting that excess cellular Fe2+ can derepress, i.e. facilitate formation of repressor-free, IRE-mRNA. Stabilities of IRE-RNA·IRP1 complexes were decreased 17- and 6-fold for ferritin and mt-aconitase IRE-RNA·IRP complexes, respectively, at 50 μm Fe2+. Thus, Fe2+, as well as the IRP1 repressor itself, recognize differences in IRE-RNA structure. The result is that Fe2+ weakens the RNA/protein repressor interaction differentially for the IRE-mRNAs that regulate ferritin and mt-aconitase synthesis.
When we analyzed IRE-RNA/IRP1 interactions by fluorescence quenching, the titration used a constant protein concentration (0.1 μm) for the ferritin IRE-RNA, additional data were collected with 0.05 μm protein; RNA varied from 0 to 1.0 μm. To evaluate the possible effect of using a constant RNA concentration and titrating with variable protein concentrations, and to obtain data from electrophoretic mobility shift gel electrophoresis as in earlier studies of the IRE-RNA·protein complexes (31, 33, 34), we examined the effect of Fe2+ on RNA/protein interactions, in the absence of oxygen, by electrophoretic mobility shift assay. A constant 0.1 μm RNA was used with variable protein concentrations (0 to 2.0 μm) ± 50 μm Fe2+-O2. Gels were prepared and analyzed in the absence of air to stabilize Fe2+. The fraction of RNA complexed to the protein was diminished by the presence of Fe2+ (Fig. 3B). Differences between ferritin and mt-aconitase RNAs in the gels are observed most readily by comparing the unshifted RNA bands (see Fig. 3). Changing the order of addition of RNA and protein to the reaction mixture had no effect. The direct effects of Fe2+ observed by the stability of the RNA·protein complex (Fig. 3) complement the indirect effects of cellular iron, as previously observed, on the RNA/protein binding that is mediated by protein degradation and, for IRP1, changes in [4Fe-4S] assembly (1, 2, 15).
Magnesium and Manganese Weaken Ferritin and mt-Aconitase IRE-RNA·IRP Repressor Complexes
Mn2+ is chemically similar to Fe2+ but can be studied in air (22). Mn2+ also weakens IRE-RNA/IRP1 interactions, with differential effects for the two IRE-RNAs as well. For example, for 50 μm ferritin IRE-RNA, IRP1 binding was weakened 8-fold by Mn2+ and 17-fold by Fe2+. For the mt-aconitase IRE-RNA·IRP1 complex, the effects of Fe2+ and Mn2+ were identical (Fig. 3A), and smaller than for the ferritin IRE-RNA·IRP complex.
Mg2+ was studied because of the direct metal interactions observed with ferritin IRE-RNA by NMR spectroscopy (11) and because Mg2+ influences the structure/function of many classes of RNA (e.g. Refs. 17 and 26). Only when Mg2+ concentrations are 100 times higher than Mn2+ or Fe2+ was comparable weakening of the IRE-RNA·IRP complexes observed (Fig. 3A). The metal selectivity and RNA selectivity for altering IRE-RNA·IRP stability raises the possibility that metal ions played a selective role in the evolution of IRE-RNA·IRP complexes or the possibility that metal ions besides iron contribute to regulation of in vivo 5′-UTR IRE-RNA/IRP interactions.
Divalent Metal Ions Bound to IRE-RNA but Not IRP1
There is no evidence that metal ions bind to IRP1, except for the FeS cluster added to the apoIRP1 that yields c-aconitase. Furthermore, there are no obvious, additional, predicted metal binding sequences in the primary sequence of IRP1 (14). On the other hand there is evidence that metals bind to IRE-RNA from solution NMR and metal nuclease cleavage (27, 28, 35). To obtain more direct evidence that the effect of metal ions on the stability of the IRE-RNA·protein complexes reflects metal ions bound to IRE-RNA and not to the IRP repressor, we examined the effect of metal ions on the intrinsic fluorescence of IRP and the fluorescence of ethidium bromide bound to IRE-RNA. We used Mn2+ as a surrogate for Fe2+ to facilitate experiments in air, because the effects on stability of the RNA·protein complex are similar to Fe2+ (Fig. 3).
Mn2+ does not bind to IRP1, based on the absence of changes in intrinsic fluorescence (Fig. 4) when manganese is added; Mg2+ does not bind either, using the same analytical approach (Fig. 4). By contrast, addition of IRE-RNA to IRP1 decreases the intrinsic fluorescence of IRP1 (Fig. 2).
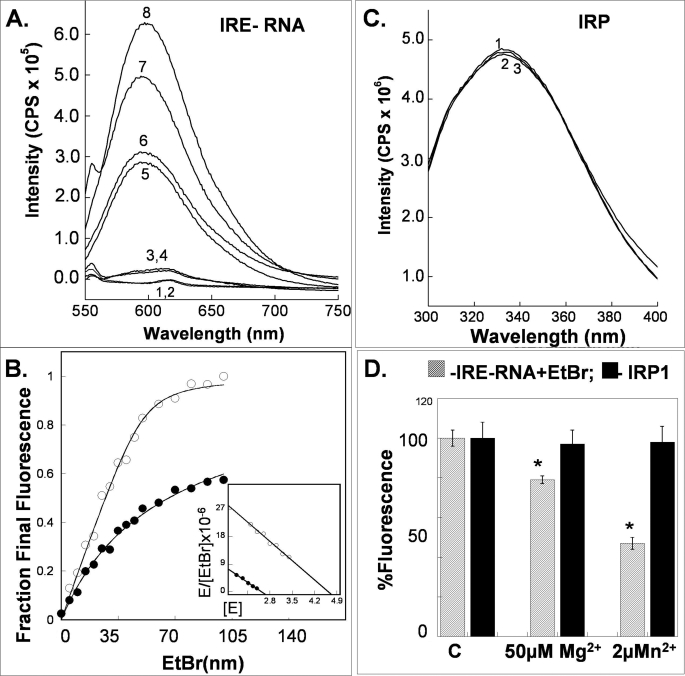
FIGURE 4.
Metal ions alter fluorescence of IRE-RNA/EtBr complexes but not of apoIRP1. Experiments with hydrated manganous ions, models for ferrous iron because of the similarity of effects on IRE-RNA·IRP complexes (Fig. 3) and general chemical similarities, are facilitated because of the relative insensitivity of Mn2+ to air at neutral pH, contrasting with Fe2+. All experiments were in binding buffer (see “Experimental Procedures”). Protein fluorescence was measured as described in the legend to Fig. 2. EtBr-IRE-RNA fluorescence (excitation at 510 nm and emission at 525 nm) was measured in solutions with 0.1 μm ethidium bromide and 2.0 μm IRE-RNA. A, fluorescence spectra of EtBr-FerH IRE RNA complexes ± metal ions (50 μm Mn2+ and/or 50 μm Mg2+): 1) RNA; 2) RNA + Mn2+; 3) EtBr; 4) EtBr + Mn2+; 5) EtBr + RNA + Mn2+ + Mg2+; 6) EtBr + RNA + Mn2+; 7) EtBr + RNA + Mg2+; and 8) EtBr + RNA. B, stoichiometry of EtBr binding to IRE-RNA in the presence (●) and absence (○) of 2 μm Mn2+; C, fluorescence spectra from solutions of IRP1 ± 50 μm Mn2+ or 50 μm Mg2+: 1) IRP1; 2) IRP1 + Mn2+; and 3) IRP1 + Mg2+. D, bar graph comparing the effects of manganous ions on IRP1 and IRE-RNA. The data are the results of duplicate experiments and the error is the S.D. *, significantly different from controls, p < 0.01 for Mn2+ and <0.02 for Mg2+.
Mn2+ binds to ferritin IRE-RNA, in experiments using EtBr fluorescence as a reporter (Fig. 4A). For example, EtBr fluorescence decreases when Mn2+ was added to EtBr and RNA; the Ka between EtBr and RNA was 41.6 ± 2.3 × 106 liter/mol in the absence of Mn2+ and 18.0 ± 0.6 × 106 liter/mol in the presence of Mn2+; the decrease in Ka is significant (p < 0.01), and indicates metal binding to RNA. The stoichiometry of EtBr binding to IRE-RNA was 4.8 ± 0.3, and decreases to 2.6 ± 0.1 (Fig. 4) in the presence of Mn2+. Given the general similarities between Fe2+ and Mn2+ in destabilizing the IRE-RNA·IRP complex, undetectable Mn2+ binding to IRP1, and detectable Mn2+ binding to IRE-RNA and general similarities of Fe2+ and Mn2+ chemistry, the results indicate that Fe2+ destabilizes the IRE-RNA·IRP complex by binding to the RNA. In vivo when Fe2+ accumulates to concentrations that saturate normal Fe2+ chaperones and transporters, and fill ferritin, Fe2+ becomes available to destabilize IRE-RNA·IRP complexes and increase the synthesis of the encoded proteins.
Ferritin IRE-RNA Binds IRP1 Repressor More Stably Than the Ferritin ΔU6 Mutant or mt-Aconitase IRE-RNAs
Because ferritin IRE-RNA differs from mt-aconitase IRE-RNA by a U6 bulge in the helix, we analyzed the ferritin ΔU6 mutant IRE-RNA. Ferritin ΔU6 IRE-RNA should behave more like mt-aconitase than ferritin IRE-RNA, based on the predicted RNA secondary structure (Fig. 2) and the regions of contact in the IRE-RNA/IRP1 crystal structure (Fig. 1B). In fact, stability of the ferritin ΔU6 IRE-RNA·IRP1 complex is between that of the mt-aconitase IRE-RNA·IRP complex (3-fold higher) and the ferritin IRE-RNA·IRP complex (3-fold lower). Thus, the U6 bulge in ferritin RNA explains only part of the difference in IRP binding by ferritin and mt-aconitase IRE-RNAs. Mn2+ or Mg2+ had only small effects on the mutant IRE-RNA·IRP complex, in contrast to the complexes of wild type IRE·RNA with IRP (Fig. 3A), indicating that in the ferritin ΔU6 IRE-RNA·IRP1 complex the metal binding sites that influence stability or the RNA conformational change responsible for decreased IRP binding are blocked by deletion of U6. Because both ferritin and ferritin ΔU6 IRE-RNAs have identical base pairs but differences in IRP1 binding, because ferritin ΔU6 and mt-aconitase IRE-RNAs have similar secondary structures around the two IRP1 binding sites of the C8 bulge and CAGUG loop (Fig. 2), and because metal effects on the protein·RNA complex differ among ferritin, ferritin ΔU6, and mt-aconitase IRE-RNAs, the phylogenetically conserved differences among each IRE-mRNA clearly contribute to the inherent stability and metal responses of the IRE-RNA·IRP complexes.
DISCUSSION
The translation of 5′-UTR IRE-mRNAs increases when concentrations of environmental iron, such as ferrous salts, ferric chelates, and heme increase as the result of iron-induced changes in the IRE/IRP interactions and by degrading IRP (1–4). However, the chemical identity of the cellular iron signal targeted to IRE-RNA/IRP interactions is unknown, although insertion of a [4Fe-4S] cluster, after cluster synthesis/transport by ISC protein catalysts and chaperones (36), influences IRE-RNA/IRP1 interactions. When cellular iron concentrations are high, a larger fraction of each 5′-UTR IRE-mRNA is in the polyribosomes (5, 6, 37). Iron deficiency, by contrast stabilizes IRP concentrations and destabilizes the FeS cluster in IRP1, possibly mediated by IRP phosphorylation, and a smaller fraction of the 5′-UTR IRE-mRNAs are in polyribosomes (16). (A subset of the IRE-mRNA family, mainly involved in iron absorption and transport, regulates mRNA turnover with the mRNA stabilized by IRP binding. The 3′-UTR IRE-RNAs have specific structural features that place them outside the scope of this study.) IRE-RNA·IRP complexes occur in the cytoplasm, but whether the RNA·protein complex forms during IRE-mRNA processing in the nucleus, or during transport to the cytoplasm for mRNA storage or use remains unknown. The effect of Fe2+ on the stability of the 5′-UTR IRE-RNA·IRP complexes reported here indicates that the complex can sense increases in concentrations of aquated Fe2+ that result in weaker RNA/protein interactions.
Destabilization of the IRE-RNA·IRP complex by Fe2+ competes with the stabilization conferred by the very large number of bonds between the protein and the RNA complex in crystals (13); no IRE-RNA/IRP or IRE-RNA crystals have been obtained with metals to date (38). In the IRE-RNA·IRP complex, protein-RNA bonds are clustered at the RNA C8 bulge and the CAGUG terminal loop of the IRE-RNA structure (13). A number of RNA-protein bonds involve RNA sites that are hypersensitive to cleavage by Fe2+-EDTA/ascorbate/H2O2 (28), suggesting specific interactions with Fe2+-EDTA or solvent or both. The observed destabilization of the IRE-RNA·IRP complex is most likely explained by metal binding to exposed sites on the IRE-RNA (Fig. 1B), for several reasons. First, metal ions did not bind to the IRP1 in the current study. Second, there are no predicted metal binding sites in the IRP protein (14) beyond cysteine/[Fe-S] interactions. Third, in this study, metal ions decrease conformation-sensitive EtBr binding to RNA. Finally metal binding to IRE-RNA has been detected by NMR spectroscopy and metallonuclease cleavage (12, 27, 35).
Structural differences between ferritin and mt-aconitase IRE-RNAs are reflected in the stability differences of the RNA·protein complexes observed here, because the same IRP is present in both complexes and only the RNA is different. IRE-RNA structural differences then, explain, at least in part, the 30-fold difference in iron-induced synthesis in vivo for the two proteins and differences in IRP1 and IRP2 binding in vitro (31, 32). Because ferritin is an ancient IRE-RNA and mt-aconitase evolved more recently (39), the lower stability of the mt-aconitase IRE-RNA·protein complex and the smaller iron response may reflect the shorter time for evolutionary fine-tuning of the IRE-mRNA. However, the different iron responses of the two IRE-RNAs could also reflect the physiological function of each encoded protein. For example, large fluctuations in mt-aconitase synthesis could be deleterious to cell oxidative metabolism, which would explain the relatively small changes in synthesis induced by iron. By contrast large changes in ferritin synthesis allow cells to respond to large changes in iron and oxygen stress through ferritin mRNA translation (more sensitive to iron signals) as well as ferritin DNA transcription (more sensitive to oxidant signals) (40, 41).
Fe2+, Mn2+, and 100× Mg2+ destabilize wild type IRE-RNA·IRP complexes, but the IRE-RNA mutation greatly reduced the metal ion effects; a single nucleotide deletion that eliminated the ferritin-selective IRE-RNA U6 bulge also greatly reduced the metal ion effects and weakened IRP binding. Thus, the genetic selective forces acting upon each IRE-mRNA appear to be extremely high. Moreover, the phylogenetic conservation of each IRE-RNA structure is extremely high (>90%) and includes variations among wild type IRE-RNA structures such as the initiator AUG in the mt-aconitase IRE-RNA, and base pairing flanking sequences around the ferritin-IRE RNAs of vertebrates (27, 28, 39, 42). The family of IRE-RNA structures with selective recognition by IRP 1 and 2 (29, 31, 43) yields a combinatorial array of RNA·protein complexes (44) with a range of physical stabilities that are tunable by Fe2+ and other cellular signals.
Acknowledgment
We are grateful to Dr. Takehiko Tosha for the graphics in Fig. 1B.
This work was supported, in whole or in part, by National Institutes of Health Grants DK20251 (to E. C. T.) and DK47281 (to W. E. W.). This work was also supported by National Science Foundation Grant MCB0814051 (to D. J. G. and M. K.), a Professional Staff Congress-City University of New York Faculty Award (to D. J. G.), and the Children's Hospital Oakland Research Institute Foundation (to E. C. T.).
- UTR
- untranslated region
- IRE
- iron-responsive element
- IRP
- iron regulatory protein repressor for IRE mRNA
- mt
- mitochondrial
- EtBr
- ethidium bromide.
REFERENCES
- 1.Wallander M. L., Leibold E. A., Eisenstein R. S. (2006) Biochim. Biophys. Acta 1763, 668–689 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rouault T. A. (2006) Nat. Chem. Biol. 2, 406–414 [DOI] [PubMed] [Google Scholar]
- 3.Leipuviene R., Theil E. C. (2007) Cell. Mol. Life Sci. 64, 2945–2955 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Muckenthaler M. U., Galy B., Hentze M. W. (2008) Annu. Rev. Nutr. 28, 197–213 [DOI] [PubMed] [Google Scholar]
- 5.Zähringer J., Baliga B. S., Munro H. N. (1976) Proc. Natl. Acad. Sci. U.S.A. 73, 857–861 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Melefors O., Goossen B., Johansson H. E., Stripecke R., Gray N. K., Hentze M. W. (1993) J. Biol. Chem. 268, 5974–5978 [PubMed] [Google Scholar]
- 7.Hentze M. W., Muckenthaler M. U., Andrews N. C. (2004) Cell 117, 285–297 [DOI] [PubMed] [Google Scholar]
- 8.Sierzputowska-Gracz H., McKenzie R. A., Theil E. C. (1995) Nucleic Acids Res. 23, 146–153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Laing L. G., Hall K. B. (1996) Biochemistry 35, 13586–13596 [DOI] [PubMed] [Google Scholar]
- 10.Addess K. J., Basilion J. P., Klausner R. D., Rouault T. A., Pardi A. (1997) J. Mol. Biol. 274, 72–83 [DOI] [PubMed] [Google Scholar]
- 11.Gdaniec Z., Sierzputowska-Gracz H., Theil E. C. (1998) Biochemistry 37, 1505–1512 [DOI] [PubMed] [Google Scholar]
- 12.Ke Y., Sierzputowska-Gracz H., Gdaniec Z., Theil E. C. (2000) Biochemistry 39, 6235–6242 [DOI] [PubMed] [Google Scholar]
- 13.Walden W. E., Selezneva A. I., Dupuy J., Volbeda A., Fontecilla-Camps J. C., Theil E. C., Volz K. (2006) Science 314, 1903–1908 [DOI] [PubMed] [Google Scholar]
- 14.Guo B., Yu Y., Leibold E. A. (1994) J. Biol. Chem. 269, 24252–24260 [PubMed] [Google Scholar]
- 15.Clarke S. L., Vasanthakumar A., Anderson S. A., Pondarré C., Koh C. M., Deck K. M., Pitula J. S., Epstein C. J., Fleming M. D., Eisenstein R. S. (2006) EMBO J. 25, 544–553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Deck K. M., Vasanthakumar A., Anderson S. A., Goforth J. B., Kennedy M. C., Antholine W. E., Eisenstein R. S. (2009) J. Biol. Chem. 284, 12701–12709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shenvi C. L., Dong K. C., Friedman E. M., Hanson J. A., Cate J. H. (2005) RNA 11, 1898–1908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wang B., Wilkinson K. A., Weeks K. M. (2008) Biochemistry 47, 3454–3461 [DOI] [PubMed] [Google Scholar]
- 19.Oshikane H., Sheppard K., Fukai S., Nakamura Y., Ishitani R., Numata T., Sherrer R. L., Feng L., Schmitt E., Panvert M., Blanquet S., Mechulam Y., Söll D., Nureki O. (2006) Science 312, 1950–1954 [DOI] [PubMed] [Google Scholar]
- 20.Rangan P., Masquida B., Westhof E., Woodson S. A. (2004) J. Mol. Biol. 339, 41–51 [DOI] [PubMed] [Google Scholar]
- 21.Vicens Q., Paukstelis P. J., Westhof E., Lambowitz A. M., Cech T. R. (2008) RNA 14, 2013–2029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Vogt M., Lahiri S., Hoogstraten C. G., Britt R. D., DeRose V. J. (2006) J. Am. Chem. Soc. 128, 16764–16770 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nelson J. A., Uhlenbeck O. C. (2008) RNA 14, 605–615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lipfert J., Das R., Chu V. B., Kudaravalli M., Boyd N., Herschlag D., Doniach S. (2007) J. Mol. Biol. 365, 1393–1406 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Coppins R. L., Hall K. B., Groisman E. A. (2007) Curr. Opin. Microbiol. 10, 176–181 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Martick M., Horan L. H., Noller H. F., Scott W. G. (2008) Nature 454, 899–902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang Y. H., Sczekan S. R., Theil E. C. (1990) Nucleic Acids Res. 18, 4463–4468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Harrell C. M., McKenzie A. R., Patino M. M., Walden W. E., Theil E. C. (1991) Proc. Natl. Acad. Sci. U.S.A. 88, 4166–4170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Erlitzki R., Long J. C., Theil E. C. (2002) J. Biol. Chem. 277, 42579–42587 [DOI] [PubMed] [Google Scholar]
- 30.Dix D. J., Lin P. N., McKenzie A. R., Walden W. E., Theil E. C. (1993) J. Mol. Biol. 231, 230–240 [DOI] [PubMed] [Google Scholar]
- 31.Ke Y., Wu J., Leibold E. A., Walden W. E., Theil E. C. (1998) J. Biol. Chem. 273, 23637–23640 [DOI] [PubMed] [Google Scholar]
- 32.Chen O. S., Schalinske K. L., Eisenstein R. S. (1997) J. Nutr. 127, 238–248 [DOI] [PubMed] [Google Scholar]
- 33.Butt J., Kim H. Y., Basilion J. P., Cohen S., Iwai K., Philpott C. C., Altschul S., Klausner R. D., Rouault T. A. (1996) Proc. Natl. Acad. Sci. U.S.A. 93, 4345–4349 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Phillips J. D., Guo B., Yu Y., Brown F. M., Leibold E. A. (1996) Biochemistry 35, 15704–15714 [DOI] [PubMed] [Google Scholar]
- 35.Thorp H. H., McKenzie R. A., Lin P. N., Walden W. E., Theil E. C. (1996) Inorg. Chem. 35, 2773–2779 [Google Scholar]
- 36.Lill R., Mühlenhoff U. (2008) Annu. Rev. Biochem. 77, 669–700 [DOI] [PubMed] [Google Scholar]
- 37.Dickey L. F., Wang Y. H., Shull G. E., Wortman I. A., 3rd, Theil E. C. (1988) J. Biol. Chem. 263, 3071–3074 [PubMed] [Google Scholar]
- 38.Selezneva A. I., Cavigiolio G., Theil E. C., Walden W. E., Volz K. (2006) Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 62, 249–252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Piccinelli P., Samuelsson T. (2007) RNA 13, 952–966 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hintze K. J., Theil E. C. (2005) Proc. Natl. Acad. Sci. U.S.A. 102, 15048–15052 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hintze K. J., Katoh Y., Igarashi K., Theil E. C. (2007) J. Biol. Chem. 282, 34365–34371 [DOI] [PubMed] [Google Scholar]
- 42.Kim H. Y., LaVaute T., Iwai K., Klausner R. D., Rouault T. A. (1996) J. Biol. Chem. 271, 24226–24230 [DOI] [PubMed] [Google Scholar]
- 43.Gunshin H., Allerson C. R., Polycarpou-Schwarz M., Rofts A., Rogers J. T., Kishi F., Hentze M. W., Rouault T. A., Andrews N. C., Hediger M. A. (2001) FEBS Lett. 509, 309–316 [DOI] [PubMed] [Google Scholar]
- 44.Theil E. C., Eisenstein R. S. (2000) J. Biol. Chem. 275, 40659–40662 [DOI] [PubMed] [Google Scholar]
- 45.Khan M. A., Miyoshi H., Gallie D. R., Goss D. J. (2008) J. Biol. Chem. 283, 1340–1349 [DOI] [PubMed] [Google Scholar]
- 46.Firpo M. A., Connelly M. B., Goss D. J., Dahlberg A. E. (1996) J. Biol. Chem. 271, 4693–4698 [DOI] [PubMed] [Google Scholar]