Abstract
UV light induces phosphorylation of the α subunit of the eukaryotic initiation factor 2 (eIF2α) and inhibits global protein synthesis. Both eIF2 kinases, protein kinase-like endoplasmic reticulum kinase (PERK) and general control of nonderepressible protein kinase 2 (GCN2), have been shown to phosphorylate eIF2α in response to UV irradiation. However, the roles of PERK and GCN2 in UV-induced eIF2α phosphorylation are controversial. The one or more upstream signaling pathways that lead to the activation of PERK or GCN2 remain unknown. In this report we provide data showing that both PERK and GCN2 contribute to UV-induced eIF2α phosphorylation in human keratinocyte (HaCaT) and mouse embryonic fibroblast cells. Reduction of expression of PERK or GCN2 by small interfering RNA decreases phosphorylation of eIF2α after UV irradiation. These data also show that nitric-oxide synthase (NOS)-mediated oxidative stress plays a role in regulation of eIF2α phosphorylation upon UV irradiation. Treating the cells with the broad NOS inhibitor NG-methyl-l-arginine, the free radical scavenger N-acetyl-l-cysteine, or the NOS substrate l-arginine partially inhibits UV-induced eIF2α phosphorylation. The results presented above led us to propose that NOS mediates UV-induced eIF2α phosphorylation by activation of both PERK and GCN2 via oxidative stress and l-arginine starvation signaling pathways.
UV irradiation inhibits translation initiation through activation of kinases that phosphorylate the α-subunit of eukaryotic initiation factor 2 (eIF2α).2 Two eIF2α kinases, double strand RNA-dependent protein kinase-like ER kinase (PERK) and general control of amino acid biosynthesis kinase (GCN2), are known to phosphorylate the serine 51 of eIF2α in response to UV irradiation (1–4). However, the one or more upstream pathways that activate eIF2α kinase(s) upon UV irradiation are not known. In this report, we provide evidence that UV-induced nitric-oxide synthase (NOS) activation and nitric oxide (NO•) production regulate both PERK and GCN2 activation upon UVB irradiation.
Expression of inducible nitric-oxide synthase in a mouse macrophage cell line leads to the phosphorylation of eIF2α and inhibition of translation (5). In cultured neuronal and pancreatic cell lines, production of NO• and peroxynitrite (ONOO−) induces endoplasmic reticulum (ER) stress, which activates PERK and results in cell dysfunction and apoptosis (6–9). Cytokine-stimulated inducible nitric-oxide synthase activation in astrocytes depletes l-arginine and activates GCN2, which phosphorylates eIF2α (10). UV irradiation also activates NOS and elevates cellular NO• (11–13). However, the UV-induced NOS activation and NO• production have never been shown to be related to the activation of eIF2α kinase(s). Now we demonstrate that UV-induced activation of NOS mediates the activation of both PERK and GCN2, which coordinately regulate the phosphorylation of eIF2α.
EXPERIMENTAL PROCEDURES
Cell Culture
HaCaT cells (kindly provided by Dr. Hongtao Yu, Jackson State University) were grown in Dulbecco's minimal essential medium (Cellgro) supplemented with 10% fetal calf serum and penicillin/streptomycin. The cells were cultured at 37 °C with 5% CO2. Wild-type mouse embryonic fibroblasts (MEFwt), knock-out MEF cells (MEFPERK−/−) and GCN2 knock-out MEF (MEFGCN2−/−) cells (kindly provided by Dr. Randal Kaufman, University of Michigan Medical School, Ann Arbor, MI) were grown in 10% fetal calf serum-enriched Dulbecco's modified Eagle's medium (Cellgro). The cells were incubated at 37 °C and 10% CO2.
UVB Irradiation
UVB light was generated from two 15-watt UVB lamps (UVP Inc.). The intensity of UVB was measured by a UVB meter (UVP Inc.). Cells were treated with the UVB dose at 50 mJ/cm2. Medium was removed before the UVB treatment.
Chemical Treatments of Cultured Cells
NG-Methyl-l-arginine (LNMMA, Sigma) was used to inhibit NOS activity and N-acetyl-l-cysteine (LNAC, Sigma) was used to scavenge free radicals. The cells were pretreated with LNMMA (100 μm for HaCaT or 200 μm for MEFs) or LNAC (25 mm) for 1 h and then UVB-irradiated (50 mJ/cm2). Immediately after irradiation, the cells were cultured in the same medium with the inhibitors until further analysis. For l-Arg treatment, cells were supplemented with l-Arg (50 mm for HaCaT, 25 mm for MEFs, or as indicated) in the complete medium 1 h before UVB irradiation. Immediately after irradiation, the cells were cultured in fresh medium without l-Arg until further analysis.
Western Blot Analysis
After treatment, cells were lysed by Nonidet P-40 lysis buffer (2% Nonidet P-40, 80 mm NaCl, 100 mm Tris-HCl, 0.1% SDS) with proteinase inhibitor mixture (CompleteTM, Roche Molecular Biochemicals). When phosphorylated proteins were analyzed, the phosphatase inhibitor mixture (Sigma) was added into the lysis buffer according to the manufacturer's protocol. The cell lysate was incubated on ice for 5 min and then centrifuged at 14,000 rpm for 10 min at 4 °C. The protein concentration was measured by using a Bio-Rad Protein DC Assay kit (Bio-Rad Laboratories). Equal amounts of protein were subjected on SDS-PAGE and transferred to a nitrocellulose membrane at 100 V for 30 min using a Trans-Blot SD Semi-Dry Electrophoretic Cell (Bio-Rad Laboratories). For non-phosphorylated proteins, the membranes were blocked in 5% milk in phospho-buffered saline (PBS) with 0.02% Tween 20 (PBST) for 30 min. For phosphorylated proteins, the membranes were blocked with 5% bovine serum albumin instead of milk. The membranes were incubated with primary antibodies at 4 °C overnight.
Assay for Total eIF2α and Phosphorylated eIF2α
Cells were treated with chemicals and irradiated with UVB as indicated. Immediately after UV irradiation, the cells were cultured in fresh medium. At 4 h post irradiation, the cells were lysed by the Nonidet P-40 lysis buffer with proteinase inhibitor mixture (CompleteTM, Roche Molecular Biochemicals) and phosphatase inhibitor mixture (Sigma). The cell lysate was incubated on ice for 5 min and then centrifuged at 14,000 rpm for 10 min at 4 °C. The protein concentration was measured by using the Bio-Rad Protein DC Assay kit (Bio-Rad Laboratories). Equal amounts of proteins were resolved by SDS-PAGE and electroblotted to a nitrocellulose membrane. The total amount of eIF2α and phosphorylated eIF2α were probed with antibodies against eIF2α (Cell Signaling) and phosphorylated eIF2α (Cell Signaling) using Western blot analysis.
For analysis of the ratio of phosphorylated eIF2α/total eIF2α after UVB irradiation, COS-1 cells were transiently transfected with eIF2α expression vector (pETFVA−-2α) by Lipofactamine (Invitrogen) according to the manufacturer's procedure. After 36 h the transfected cells were UVB-irradiated (50 mJ/cm2) and lysed in the Nonidet P-40 lysis buffer containing proteinase inhibitor mixture (CompleteTM, Roche Molecular Biochemicals) at the indicated post irradiation time. The eIF2α and phosphorylated eIF2α were determined by Western blot analysis as previously reported (14) with modified conditions. Equal amounts of cell lysate (5 μg of total protein) were run on a 10% SDS-PAGE with 150 V using a Mini-Gel Apparatus II (Bio-Rad) until the 38-kDa prestained protein marker (Invitrogen) was close to the bottom of the gel. The proteins were electroblotted to a nitrocellulose membrane, and eIF2α was probed with antibodies against eIF2α (Cell Signaling) as described above. The intensities of the bands of eIF2α and phosphorylated eIF2α were quantitated using ImageJ (v1.42k, National Institutes of Health).
Analysis of Protein Synthesis
After treatment, the cells were labeled with Redivue Pro Mix [35S]Met/Cys (100 μCi/ml, 1000 Ci/mmol, Amersham Biosciences) for 20 min in methionine/cysteine-free minimal essential medium (Invitrogen). After washing with PBS, cell extracts were prepared by lysing the cells in Nonidet P-40 lysis buffer (2% Nonidet P-40, 80 mm NaCl, 100 mm Tris-HCl, 0.1% SDS). The protein concentration of the cell lysate was measured using the Bio-Rad protein DC assay kit (Bio-Rad). Equal amounts of protein were loaded onto a glass microfiber disk (Whatman GF/C). The disks were air-dried, washed once with ice-cold 20% trichloroacetate and twice with ice-cold 10% trichloroacetate. The disks were dried, and the radioactivity was measured in a scintillation counter (Packard).
PERK and GCN2 Silencing Using the RNA Interference Method
HaCaT cells were seeded in a 6-well plate and incubated at 37 °C with 5% CO2 until 60–80% confluent. The culture medium was replaced with 2 ml of fresh Dulbecco's modified Eagle's medium without antibiotics, and then the cells were transfected with scrambled small interfering RNA (siRNA) (10 nm, sc-37007), PERK siRNA (10 nm, sc-36213), or GCN2 siRNA (10 nm, sc-45644) in 50 μl of Opti-MEM (Invitrogen) containing 3 μl of Lipofectamine 2000 (Invitrogen). All the siRNAs were obtained from Santa Cruz Biotechnology. The cells were incubated at 37 °C with 5% CO2 for 24 h before treatment.
Immunoprecipitation
The lysates prepared from [35S]Met/Cys metabolic-labeled cells were incubated with rabbit anti-PERK (sc-13073, Santa Cruz Biotechnology) and rabbit anti-GCN2 (AP7130a, Abgent) polyclonal antibodies. The antibody-protein complexes were then precipitated by incubating with protein A-agarose beads (Vector Laboratories) at 4 °C overnight. The immunoprecipitates were resolved by 12% SDS-PAGE gel. The gel was stained with Coomassie Blue (R-250) for 20 min, destained overnight, treated with En3Hance (PerkinElmer) for 20 min, dried, and autoradiographed.
Semi-quantitative Analysis of l-Arg by Mass Spectrometry
Electrospray mass spectrometry analysis was carried out on an LCQ DECA mass spectrometer (ThermoFinnigan). The cells were harvested by trypsin digestion, washed two times with PBS, and lysed in four times the volume of lysis buffer (2% Nonidet P-40, 100 mm NH4OAc). The lysate was further diluted by 5-fold with MeOH/H2O/HOAc (50:50:1 by volume) and then introduced into the mass spectrometer at the flow rate of 5 μl/min for ionization by electrospray. The heated capillary temperature was kept at 150 °C. Data acquisition was performed using the software Xcalibur (version 2.0.7). Considering the possible matrix effect, the quantitative analysis of l-Arg in total lysate of HaCaT cells treated with 50 mm l-Arg was performed using a standard addition method (15).
Fluorescent Detection of ONOO−
Dihydrorhodamine 123 (DHR; Sigma-Aldrich) was dissolved in DMSO as a 5 mm stock solution. The non-fluorescent DHR is oxidized by peroxides, and notably ONOO− (16, 17), to form the green fluorescent product rhodamine 123, which can be detected with emission at 488 nm and excitation at 525 nm (18). HaCaT cells were treated with LNMMA, LNAC, or l-Arg as mentioned before. At 4 h post-UVB (50 mJ/cm2), DHR was added to the medium at a working concentration of 10 μm and incubated at 37 °C and 10% CO2 for 5 min. Medium was replaced by PBS before measuring the fluorescence using a multidetection fluorescence plate reader (SPECTRA MAX M2, MDS Analytical Technologies). The fluorescent images were captured by using an inverted fluorescence microscope (IX70, Olympus).
Statistical Analysis
Student's t test was used to analyze the significance of data. p < 0.05 was considered significant.
RESULTS
PERK and GCN2 Both Phosphorylate eIF2α in Keratinocytes upon UVB Irradiation
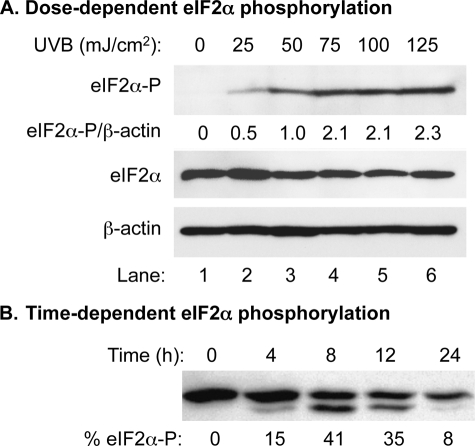
Previously, we, as well as others, reported that UVC induced eIF2α phosphorylation through activation of PERK and GCN2 (1, 3). However, there is no report indicating that the more physiological UVB also induces eIF2α phosphorylation in mammalian cells. Because keratinocytes comprise >90% of total skin cells, we first determined the dose-dependent effect of UVB on a human keratinocyte cell line: HaCaT cells. The cells were treated with UVB in a physiological dose ranging from 0 to 125 mJ/cm2 in 25 mJ/cm2 intervals. The phosphorylation of eIF2α was increased in a dose-dependent manner from 0 to 125 mJ/cm2 (Fig. 1A). Our results demonstrate that eIF2α phosphorylation is dependent on the UVB dose in the range of 25–75 mJ/cm2 (Fig. 1A), which is much lower that the human minimal erythema dose (19). The middle point dose of 50 mJ/cm2 was chosen for our following experiments.
FIGURE 1.
Dose-dependent analysis of UVB-induced phosphorylation of eIF2α in HaCaT cells. A, Western blot analysis using cell extracts prepared from UVB-irradiated HaCaT cells. The amounts of total eIF2α, phosphorylated eIF2α, and β-actin were probed with corresponding antibodies. The levels of phosphorylated eIF2α were normalized by the levels of β-actin and expressed as a percentage of phosphorylated eIF2α. Lane 3, with an UVB dose of 50 mJ/cm2, was set to 1. The rest of the lanes were normalized accordingly. B, Western blot analysis using cell extracts prepared from UVB-irradiated and pETFVA−-eIF2α transfected COS-1 cells. The amounts of both eIF2α and phosphorylated eIF2α were probed with an antibody against eIF2α. The percentage of phosphorylated eIF2α was calculated as eIF2α-P/(eIF2α + eIF2α-P).
Previous reports indicated that an estimated increase of 12% phosphorylated eIF2α is enough to significantly inhibit the activity of guanine nucleotide exchange factor and reduce 60–80% global protein synthesis (20, 21). To further assess the extent of the effect of UVB on translation, we estimated the degree of eIF2α phosphorylation in a time-dependent manner. COS-1 cells were transiently transfected with eIF2α expression vector to increase the total amount of eIF2α. Our data showed that 15% of eIF2α was phosphorylated at 4 h post-UVB (50 mJ/cm2) (Fig. 1B). The phosphorylation increased to 42% at 8 h post-UV and then decreased to 35 and 8% at 12 and 24 h post-UVB, respectively (Fig. 1B).
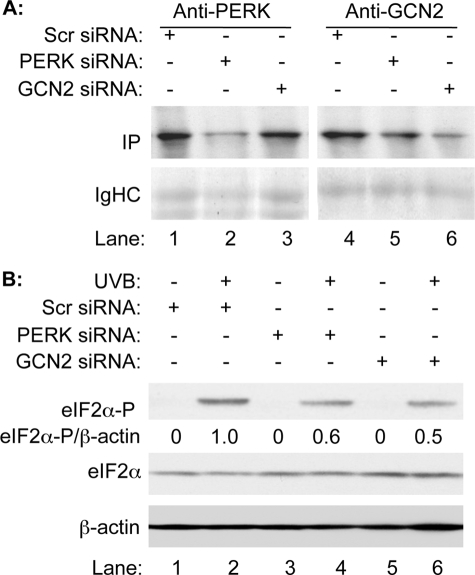
To assess the contributions of eIF2α kinases in UVB-induced translational inhibition, we used siRNA to reduce the expression of PERK or GCN2. Scrambled siRNA was also used as a control. Because the expression levels of PERK and GCN2 are low, [35S]Met/Cys metabolic labeling followed by immunoprecipitation was used to detect the efficiency of synthesis of PERK and GCN2 after siRNA treatment. Our data showed that the translation of PERK or GCN2 is significantly reduced in cells that were transfected with the corresponding siRNA (Fig. 2A, lane 2 versus 1 and lane 6 versus 4) at 24 h post-transfection. With reduced expression of PERK or GCN2, the levels of eIF2α phosphorylation upon UVB irradiation were also reduced almost half in the cells that were transfected with PERK siRNA (Fig. 2B, lane 4 versus 2) or GCN2 siRNA (Fig. 2B, lane 6 versus 2), respectively. The results indicate that both PERK and GCN2 contribute to the phosphorylation of eIF2α in human keratinocytes upon UVB irradiation.
FIGURE 2.
Both PERK and GCN2 contribute to UVB-induced phosphorylation of eIF2α. HaCaT cells were transiently transfected with either PERK or GCN2 siRNA (10 nm). A scrambled siRNA (10 nm) was also transfected into HaCaT cells as control. A, the transfected cells were metabolically labeled with [35S]Met/Cys and immunoprecipitated using anti-PERK or anti-GCN2 antibodies. The immunoprecipitates were resolved on SDS-PAGE. The newly synthesized PERK and GCN2 were visualized by autoradiograph. B, the transfected cells were treated or not treated with UVB (50 mJ/cm2). The amounts of total eIF2α, phosphorylated eIF2α, and β-actin were measured by Western blot analysis. The levels of phosphorylated eIF2α were normalized by the levels of β-actin and expressed as a percentage of phosphorylated eIF2α.
NOS Mediates UVB-induced eIF2α Phosphorylation and Translation Inhibition
UV activates NOS and induces NO• production (12, 22, 23). Elevation of NO• and ONOO− has been shown to induce ER stress, which activates PERK and results in cell dysfunction and apoptosis in cultured cells (6–9). NOS-catalyzed NO• production depletes l-Arg and activates GCN2, which phosphorylates eIF2α (10). To determine whether UVB-induced phosphorylation of eIF2α is via activation of NOS signaling pathways, we pretreated the cells with a broad NOS inhibitor, an N-substituted l-arginine analog LNMMA (100 μm) for 1 h before UVB irradiation (50 mJ/cm2). Western blot analysis indicated that the UV-induced phosphorylation of eIF2α was reduced by 64% in the presence of LNMMA (Fig. 3, lane 4 versus 2) at 4 h post-UVB irradiation. This result suggests that NOS activation plays a role in UVB-induced phosphorylation of eIF2α.
FIGURE 3.
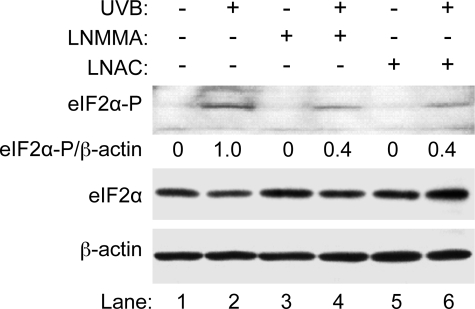
The effects of LNMMA and LNAC on UV-induced phosphorylation of eIF2α. HaCaT cells were treated with LNMMA (100 μm) or l-NAC (25 mm) and then UVB (50 mJ/cm2) as indicated. The amounts of total eIF2α, phosphorylated eIF2α, and β-actin in the total cell lysate were measured by Western blot analysis. The intensities of the bands were quantified using ImageJ (v1.42k, NIH). The levels of phosphorylated eIF2α were normalized by the levels of β-actin and expressed as a percentage of phosphorylated eIF2α.
Besides increasing the production of NO•, UV also induces an elevation of superoxide (O2˙̄) (23), which reacts rapidly with NO• to form ONOO− (24–26). Because O2˙̄ and ONOO− induce oxidative-stress, which leads to PERK activation (9, 27), we determined their roles in UVB-induced phosphorylation of eIF2α. A glutathione (GSH) synthesis precursor LNAC (25 mm) was used to reduce oxidative stress. Our data showed that the UV-induced phosphorylation of eIF2α is reduced 59% in the presence of LNAC (Fig. 3, lane 6 versus 2). Because the NOS inhibitor also inhibits UV-induced eIF2α phosphorylation (Fig. 3, lane 4 versus 2), these results indicate that UV-induced eIF2α phosphorylation could be a result of oxidative stress.
UV irradiation also activates GCN2 (1, 2, 28), which is regulated by amino acid starvation (29, 30). l-Arg is the precursor for NO• synthesis in vivo. Because NOS activation resulted in depletion of l-Arg and activation of GCN2 (10), we analyzed the impact of l-Arg on UVB-induced eIF2α phosphorylation. At 4 h post-UVB treatment, the phosphorylation of eIF2α decreased dramatically within the concentration range of 10 to 50 mm of l-Arg (Fig. 4, lanes 4, 6, and 8 versus 2). Treating with higher concentrations of l-Arg (100 mm) did not further reduce the UVB-induced phosphorylation of eIF2α. Our results suggest that UVB-induced production of NO• lead to l-Arg depletion and sequentially activated GCN2.
FIGURE 4.
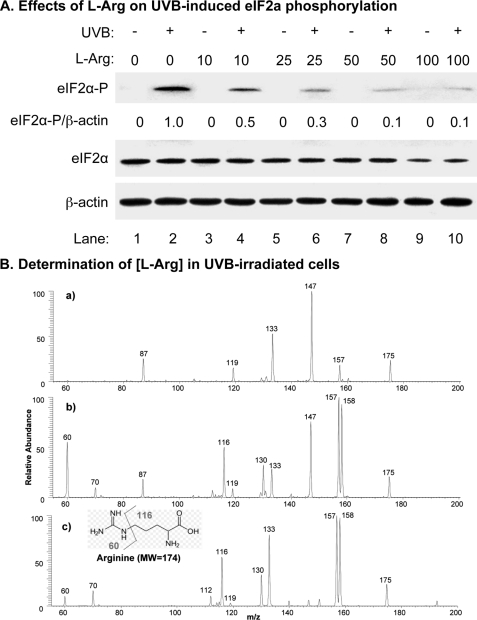
The effects of l-Arg concentration on UV-induced phosphorylation of eIF2α treatment. HaCaT cells were treated with different concentrations of l-Arg (0–100 mm) and then UVB (50 mJ/cm2) as indicated. A, the amounts of total eIF2α, phosphorylated eIF2α, and β-actin in total cell lysate were measured by Western blot analysis. The intensities of the bands were quantified using ImageJ (v1.42k, NIH). The levels of phosphorylated eIF2α were normalized by the levels of β-actin and expressed as a percentage of phosphorylated eIF2α. B, MS/MS spectra of m/z 175 ion generated from electrospray ionization of: a, total lysate of HaCaT cells without treatment; b, total lysate of HaCaT cells treated with 50 mm l-Arg; and c, a standard l-Arg solution (1.0 μm).
To further demonstrate that lack of l-Arg could be a factor for UVB-induced eIF2α phosphorylation, we estimated the intracellular amount of l-Arg (Mr = 174) in HaCaT cells using mass spectrometry. Our data showed that collision-induced dissociation (CID) of the ion of m/z 175, the protonated l-Arg ion generated from electrospray ionization of standard l-Arg (Fig. 4B, panel c), gives rise to the fragments of m/z 158 and 130 by losses of NH3 and HCOO•, respectively, and the fragments ions at m/z 60 and 116 due to the side-chain cleavage (Fig. 4B, panel c). Our data also showed that the ion of m/z 175 generated from ionization of the l-Arg-treated HaCaT cell lysate gave similar characteristic fragment ions (i.e. m/z 60, 116, 130, and 158) as that of the standard l-Arg (Fig. 4B, panel b versus c), which indicate the presence of l-Arg in the lysate. Quantitative analysis of l-Arg in total lysate of HaCaT cells treated with 50 mm l-Arg was performed using the standard-addition method, and the result showed that the lysate contains 105.0 μm l-Arg after considering the dilution factor of 25. However, the characteristic CID fragments of m/z 175 were not detected in the HaCaT cell lysate without treatment (Fig. 4B, panel a); instead, the fragments from the background peak of m/z 175 was observed. For comparison, we tested CID experiments with low concentrations of l-Arg standard solutions. The data showed that the characteristic CID fragments are well observed even for the l-Arg standard in MeOH/H2O/HOAc (50:50:1 by volume) with concentration as low as 0.1 μm. These results suggest that l-Arg in the HaCaT cell lysate without treatment is lower than 2.5 μm after considering the dilution factor. These results suggest that lack of l-Arg could be the cause of UVB-induced GCN2 activation due to its low intracellular concentration.
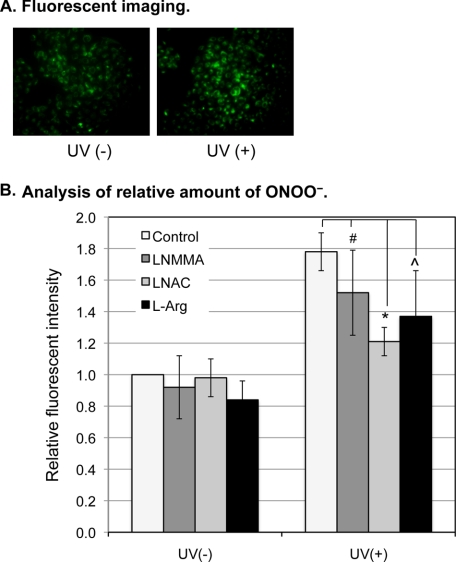
To more quantitatively analyze the oxidative stress and the generation of ONOO− after UVB irradiation, we determined the relative amount of ONOO− in the irradiated cells using the DHR fluorescence method (16, 17). The data showed that, compared with the control cells, an increased fluorescence was detected in the UVB-treated cells (Fig. 5A). Quantitative fluorescent analysis showed that ONOO− increased 0.8-fold at 4 h post irradiation (Fig. 5B). Although the UVB-induced elevation of ONOO− was reduced by a non-statistically significant 0.3- and 0.4-fold after LNMMA or l-Arg treatment, respectively, the ONOO− was statistically significantly reduced 0.6-fold after LNAC treatment (Fig. 5B). These results demonstrate that the generation of ONOO− was the result of, at least partially, to UVB-induced oxidative stress.
FIGURE 5.
The effects of UVB on ONOO− production and oxidative stress. A, HaCaT cells were treated with DHR for 5 min at 4 h post-UVB. The cells without receiving UVB were used as control. Fluorescent images were captured by an inverted fluorescence microscope at 200× (IX70 Olympus). B, HaCaT cells were not treated (control) or treated with LNMMA (100 μm), LNAC (25 mm), or l-Arg (50 mm) as indicated. The cells were then not exposed or exposed to UVB. At 4 h post-UVB, the cells were treated with DHR for 5 min, and the fluorescence intensities were recorded by a multidetection plate reader (Spectra Max M2, MDS Analytical Technologies). The error bars represent the standard deviation of three independent experiments. *, p ≤ 0.005; ⋀, p ≤ 0.1; and #, p ≤ 0.2.
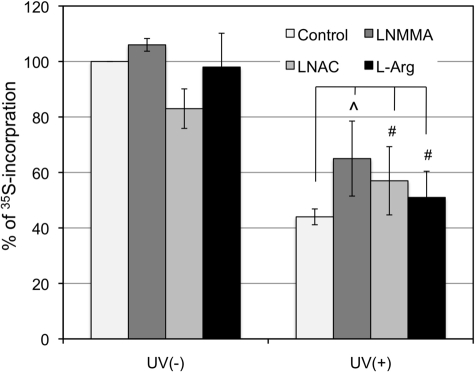
After elucidating the role of NOS in UVB-induced phosphorylation of eIF2α, we determined whether NOS-mediated eIF2α phosphorylation correlates with translation inhibition. [35S]Met/Cys metabolic labeling and trichloroacetate precipitation methods were used to quantitatively analyze the efficiency of nascent protein synthesis. The extent of translation was reduced to 44% at 4 h post-UVB irradiation (Fig. 6). Treating the irradiated cells with LNMMA, LNAC, or l-Arg partially restored protein synthesis to 65%, 57 and 51%, respectively (Fig. 6). However, the affects were not statistically significant. These results indicate that, although NOS plays a role in UVB-induced eIF2α phosphorylation, its role in translation regulation is still not conclusive.
FIGURE 6.
The effects of LNMMA, LNAC, and l-Arg on UV-induced protein synthesis inhibition. HaCaT cells were treated with LNMMA (100 μm), LNAC (25 mm), or l-Arg (50 mm) and then irradiated with UVB (50 mJ/cm2). The cells were pulse-labeled with [35S]Met/Cys, and 35S incorporation into proteins was determined by trichloroacetic acid precipitation and expressed as a percentage of 35S incorporation into the sample without treatment. The error bars represent the deviation of two sets of data. ⋀, p ≤ 0.1; #, p ≤ 0.2.
NOS Mediates UVB-induced eIF2α Phosphorylation through PERK and GCN2
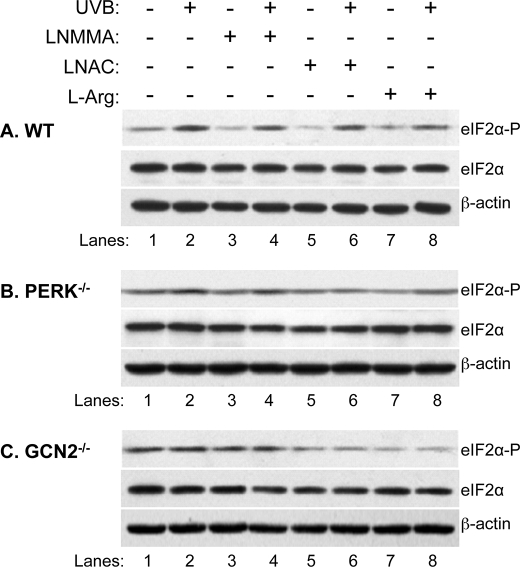
To further confirm that NOS coordinates the activation of PERK and GCN2, we analyzed the effect of LNMMA, LNAC, and l-Arg on the UVB-induced eIF2α phosphorylation in MEFwt, MEFPERK−/−, and MEFGCN2−/− cells. Western blot analysis demonstrated that the UVB-induced eIF2α phosphorylation increased >2-fold in MEFwt cells (Fig. 7A, lane 2 versus 1, and Table 1). However, the eIF2α phosphorylation increased <0.5-fold in MEFPERK−/− (Fig. 7B, lane 2 versus 1, and Table 1), whereas there was no increase in MEFGCN2−/− cells (Fig. 7C, lane 2 versus 1, and Table 1) after UVB irradiation. The results agree with previous reports that PERK or GCN2 mediate UV-induced eIF2α phosphorylation (1, 3). Treatment of LNMMA, LNAC, or l-Arg partially reduced eIF2α phosphorylation with differential percentages in all cell lines in the absence or presence of UVB irradiation (Table 1), which indicates that NOS/oxidative stress is playing a role in the maintenance of the basal level of eIF2α phosphorylation in MEF cells. These results also suggest that NOS, reactive oxygen species, and l-Arg depletion differentially coordinate the activation of PERK and GCN2 upon UVB irradiation. The NOS inhibitor LNMMA is more effective in the regulation of eIF2α phosphorylation in MEFwt cells than MEFPERK−/− and MEFGCN2−/− cells (Fig. 7 and Table 1), indicating that NOS coordinates the activation of both PERK and GCN2.
FIGURE 7.
The effects of LNMMA, LNAC, and l-Arg on UV-induced eIF2α phosphorylation in PERK and GCN2 knock-out MEF cells. MEFwt (A), MEFPERK−/− (B), and MEFGCN2−/− (C) cells were treated with LNMMA (200 μm), LNAC (25 mm), or l-Arg (25 mm) and then irradiated with UVB (50 mJ/cm2) as indicated. At 4 h post irradiation, the amounts of total eIF2α, phosphorylated eIF2α, and β-actin in the total cell lysate were determined by Western blot analysis.
TABLE 1.
Relative eIF2α phosphorylation
The levels of eIF2α phosphorylation in Fig. 7 were quantitated using ImageJ (Version 1.43k, NIH). The levels of phosphorylated eIF2α were normalized by the levels of β-actin and expressed as a percentage of phosphorylated eIF2α in control cells. Numbers represent the averages ± S.D. values of three sets of data.
| Control | UV | LNMMA | UV + LNMMA | LNAC | UV + LNAC | l-Arg | UV + l-Arg | |
|---|---|---|---|---|---|---|---|---|
| A: WT | 100 | 219.7 ± 19.3 | 57.3 ± 6.7 | 161.9 ± 4.5 | 49.5 ± 17.7 | 183.4 ± 28.6 | 92.6 ± 23.4 | 152.6 ± 41.3 |
| B: PERK−/− | 100 | 140.6 ± 11.2 | 87.5 ± 13.7 | 125.2 ± 11.1 | 81.1 ± 15.5 | 78.4 ± 10.6 | 67.5 ± 11.4 | 117.8 ± 3.2 |
| C: CN2−/− | 100 | 96.5 ± 5.0 | 77.5 ± 1.8 | 90.0 ± 3.5 | 48.0 ± 1.3 | 35.2 ± 8.3 | 28.1 ± 5.0 | 23.8 ± 5.6 |
| Column: | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
DISCUSSION
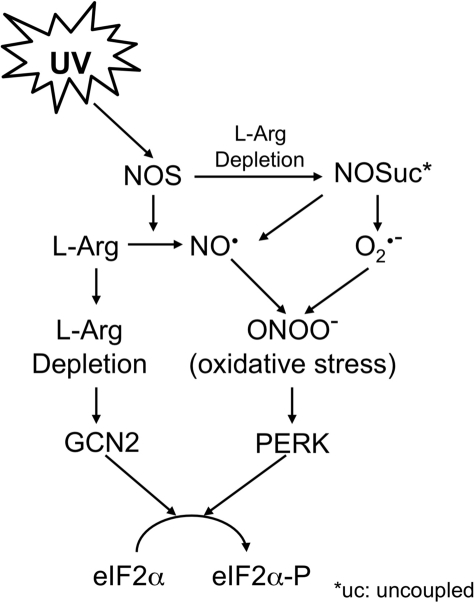
UV irradiation inhibits translation via activation of two serine-threonine kinases, PERK and GCN2, which phosphorylate the Ser-51 of eIF2α (1, 3). PERK is an ER membrane-localized kinase (31–33), which in its inactive monomer state is stabilized by an ER chaperone Ig heavy chain-binding protein. GCN2 is an amino acid abundance-controlled eIF2α kinase, which is activated during amino acid starvation (29, 34). Although the roles of PERK and GCN2 in UV-induced translation inhibition and downstream signal transduction have been previously elucidated (2, 4, 28), their key activators have not been identified. We now demonstrate that UVB-induced translational inhibition in cultured human skin cells is, at least partially, mediated by NOS-catalyzed NO• production in combination with superoxide production. Based on our results, we propose a pathway for the UVB-induced eIF2α phosphorylation (Fig. 8). In this model, UVB irradiation induces NOS activation, which rapidly generates NO• from l-Arg. The reduced availability of l-Arg leads to NOS uncoupling and GCN2 activation. The uncoupled NOS then produces O2˙̄, which reacts with NO• to form ONOO−. The elevated ONOO− level leads to the generation of oxidative stress and activation of PERK. Our findings not only significantly advance our understanding of the pathway for UV-induced eIF2α phosphorylation but also elucidate a role of NOS in coordinative regulation of two activator-specific eIF2α kinases.
FIGURE 8.
Proposed model for UVB-induced phosphorylation of eIF2α.
Our results also unify previously controversial conclusions regarding the role of PERK and GCN2 in UV-induced eIF2α phosphorylation (1, 3). Using the siRNA method, we demonstrated that both PERK and GCN2 contribute to the UVB-induced eIF2α phosphorylation at the same time frames and under the same conditions in human keratinocytes HaCaT (Fig. 2). Keratinocytes have been shown to contain NOS that can be activated by UVB exposure (12, 22, 36). Our data showed that eIF2α phosphorylation was reduced after treating the cells with an NOS inhibitor, LNMMA, the GSH precursor LNAC or l-Arg (Figs. 3 and 4). l-Arg is the only precursor of NO• generated by NOS. Generation of NO• could lead to l-Arg depletion, which could in turn lead to NOS uncoupling and O2˙̄ production (37). O2˙̄ rapidly reacts with NO• to form ONOO− (38, 39), which is a strong oxidant (40, 41) and induces ER stress (27). The effects of the antioxidant and l-Arg on UVB-induced eIF2α phosphorylation indicate that both NOS and oxidative stress play roles in UVB-induced eIF2α phosphorylation. The generation of the strong oxidant ONOO− was further substantiated by the increased fluorescence of oxidized DHR after UVB irradiation (Fig. 5A), which is significantly reduced by treating the cells with LNAC (Fig. 5B). The UVB-induced translation inhibition was also partially reversed, with less effectiveness and non-statistically significant, by treating the cells with LNMMA, LNAC, or l-Arg (Fig. 6). This could be due to the fact that only a very low percentage of phosphorylated eIF2α is required to reduce global protein synthesis (21). At 4 h post-UVB, ∼15% of the total eIF2α is phosphorylated (Fig. 1B). Even with a 60% reduction of the eIF2α phosphorylation, there are still 6% of phosphorylated eIF2α, which may reduce guanine nucleotide exchange factor activity and inhibit translation. The reduced effectiveness of l-Arg on reversing the translation inhibition is not clear. One possible cause is that supplement of l-Arg leads to the production of a large amount of NO•, which inhibits protein synthesis through an eIF2α-independent pathway as previously reported (42).
The extent of the effects of LNMMA, LNAC, or l-Arg on UVB-induced eIF2α phosphorylation in parent and mutated MEF cells is very interesting. Distinctive from HaCaT cells, a background eIF2α phosphorylation was observed in all three MEF cell lines, which was the same as we observed before in MEFwt cells (4). In MEFwt cells, NOS and oxidative stress is required, whereas supplement of l-Arg is not required, for maintaining basal eIF2α phosphorylation in MEFwt cells (Table 1, A3, 5, and 7 versus A1). UVB-induced eIF2α phosphorylation mainly results from l-Arg depletion, which could be mediated by NOS (Table 1, A4 and 8 versus A2). Interestingly, reducing oxidative stress had less impact on eIF2α phosphorylation in the UVB-treated cells than non-treated cells (Table 1, A6 versus A5). This could be due to the generation of the strong oxidant peroxynitrite by uncoupled NOS, which induces the growth arrest and DNA damage-inducible protein (43) and sequentially dephosphorylates eIF2α. In MEFPERK−/− cells, l-Arg shortage-mediated GCN2 activation plays a more significant role for maintaining basal eIF2α phosphorylation (Table 1, B7 versus B1, 3, and 5). However, UVB-induced eIF2α phosphorylation mainly resulted from oxidative stress (Table 1, B6 versus B2, 4, and 8). One possible pathway is that induced expression of activating transcription factor 4 is needed for biosynthesis of glutathione (44), which reduces oxidative stress induced by UVB irradiation (45). Reduced inducibility of activating transcription factor 4 in response to UVB-induced ER stress may lead to an increased level of oxidative-stress, which activates GCN2 as recently reported in yeast (46). The MEFGCN2−/− cells were under oxidative stress and l-Arg starvation, because the LNAC or l-Arg supplement significantly reduced eIF2α phosphorylation without or with UVB irradiation (Table 1, C5–8 versus C1 and 2). Because l-Arg biosynthesis is up-regulated by general control nonderepressible protein kinase 4, whose activity is positively controlled by GCN2 (47, 48), it is reasonable to assume that the concentration of l-Arg is lower in the cells. The lack of l-Arg could lead to NOS uncoupling and generation of NO• and O2˙̄, which rapidly form ONOO− and therefore oxidative stress (35, 38, 39). Because l-Arg is depleted and oxidative stress levels are high in the cells, UVB irradiation is unable to further induce them and sequentially eIF2α phosphorylation. Treatment of LNMMA inhibits NOS and stops further generation of ONOO− but does not reduce the existing oxidative stress. Treatment of l-Arg leads to NOS coupling, which increases the NO•/ONOO− ratio and reduces the oxidative stress. This pathway explains the effectiveness of LNAC or l-Arg in reduction of eIF2α phosphorylation without or with UVB irradiation. It could also explain the correlation of oxidative stress with l-Arg depletion in reducing eIF2α phosphorylation in MEFGCN2−/− cells.
Acknowledgments
We thank Oliver L. Carpenter and Dr. Roxanne Male-Brune for editorial assistance.
This work was supported, in whole or in part, by National Institutes of Health Grants RO1 CA86928 (to S. W.) and R56 CA086928 (to S. W.).
- eIF2
- eukaryotic initiation factor 2
- ER
- endoplasmic reticulum
- PERK
- double strand RNA-dependent protein kinase-like ER kinase
- GCN2
- general control nonderepressible protein kinase 2
- HaCaT
- human keratinocytes
- MEF
- mouse embryonic fibroblast
- siRNA
- small interfere RNA
- NOS
- nitric-oxide synthase
- LNMMA
- NG-methyl-l-arginine
- LNAC
- N-acetyl-l-cysteine
- DHR
- dihydrorhodamine 123
- PBS
- phosphate-buffered saline
- CID
- collision-induced dissociation.
REFERENCES
- 1.Deng J., Harding H. P., Raught B., Gingras A. C., Berlanga J. J., Scheuner D., Kaufman R. J., Ron D., Sonenberg N. (2002) Curr. Biol. 12, 1279–1286 [DOI] [PubMed] [Google Scholar]
- 2.Jiang H. Y., Wek R. C. (2005) Biochem. J. 385, 371–380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wu S., Hu Y., Wang J. L., Chatterjee M., Shi Y., Kaufman R. J. (2002) J. Biol. Chem. 277, 18077–18083 [DOI] [PubMed] [Google Scholar]
- 4.Wu S., Tan M., Hu Y., Wang J. L., Scheuner D., Kaufman R. J. (2004) J. Biol. Chem. 279, 34898–34902 [DOI] [PubMed] [Google Scholar]
- 5.Kim Y. M., Son K., Hong S. J., Green A., Chen J. J., Tzeng E., Hierholzer C., Billiar T. R. (1998) Mol. Med. 4, 179–190 [PMC free article] [PubMed] [Google Scholar]
- 6.Oyadomari S., Takeda K., Takiguchi M., Gotoh T., Matsumoto M., Wada I., Akira S., Araki E., Mori M. (2001) Proc. Natl. Acad. Sci. U.S.A. 98, 10845–10850 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Oyadomari S., Araki E., Mori M. (2002) Apoptosis 7, 335–345 [DOI] [PubMed] [Google Scholar]
- 8.Oliver B. L., Cronin C. G., Zhang-Benoit Y., Goldring M. B., Tanzer M. L. (2005) J. Cell. Physiol. 204, 45–50 [DOI] [PubMed] [Google Scholar]
- 9.Kawahara K., Oyadomari S., Gotoh T., Kohsaka S., Nakayama H., Mori M. (2001) FEBS Lett. 506, 135–139 [DOI] [PubMed] [Google Scholar]
- 10.Lee J., Ryu H., Ferrante R. J., Morris S. M., Jr., Ratan R. R. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 4843–4848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Deliconstantinos G., Villiotou V., Fassitsas C. (1992) J. Cardiovasc. Pharmacol. 20, Suppl. 12, S63–S65 [DOI] [PubMed] [Google Scholar]
- 12.Kubaszewski E., Peters A., McClain S., Bohr D., Malinski T. (1994) Biochem. Biophys. Res. Commun. 200, 213–218 [DOI] [PubMed] [Google Scholar]
- 13.Kuchel J. M., Barnetson R. S., Halliday G. M. (2003) J. Invest. Dermatol. 121, 587–593 [DOI] [PubMed] [Google Scholar]
- 14.Brostrom C. O., Prostko C. R., Kaufman R. J., Brostrom M. A. (1996) J. Biol. Chem. 271, 24995–25002 [DOI] [PubMed] [Google Scholar]
- 15.Skoog D. A., Holler F. J., Crouch S. R. (2007) Principles of Instrumental Analysis, 6th Ed., pp. 13–17, Thomson Brooks/Cole, Belmont, CA [Google Scholar]
- 16.Rhyu D. Y., Yang Y., Ha H., Lee G. T., Song J. S., Uh S. T., Lee H. B. (2005) J. Am. Soc. Nephrol. 16, 667–675 [DOI] [PubMed] [Google Scholar]
- 17.Allen D. A., Harwood S., Varagunam M., Raftery M. J., Yaqoob M. M. (2003) FASEB J. 17, 908–910 [DOI] [PubMed] [Google Scholar]
- 18.Henderson L. M., Chappell J. B. (1993) Eur. J. Biochem. 217, 973–980 [DOI] [PubMed] [Google Scholar]
- 19.Beattie P. E., Finlan L. E., Kernohan N. M., Thomson G., Hupp T. R., Ibbotson S. H. (2005) Br. J. Dermatol. 152, 1001–1008 [DOI] [PubMed] [Google Scholar]
- 20.Clemens M. J., Galpine A., Austin S. A., Panniers R., Henshaw E. C., Duncan R., Hershey J. W., Pollard J. W. (1987) J. Biol. Chem. 262, 767–771 [PubMed] [Google Scholar]
- 21.Rowlands A. G., Panniers R., Henshaw E. C. (1988) J. Biol. Chem. 263, 5526–5533 [PubMed] [Google Scholar]
- 22.Deliconstantinos G., Villiotou V., Stavrides J. C. (1996) Exp. Physiol. 81, 1021–1033 [DOI] [PubMed] [Google Scholar]
- 23.Wiswedel I., Keilhoff G., Dörner L., Navarro A., Böckelmann R., Bonnekoh B., Gardemann A., Gollnick H. (2007) Free Radic. Res. 41, 1017–1027 [DOI] [PubMed] [Google Scholar]
- 24.Crow J. P., Beckman J. S. (1995) Adv. Pharmacol. 34, 17–43 [DOI] [PubMed] [Google Scholar]
- 25.Malinski T. (2005) Am. J. Cardiol. 96, 13i–24i [DOI] [PubMed] [Google Scholar]
- 26.Weller R., Billiar T., Vodovotz Y. (2002) Skin Pharmacol. Appl. Skin Physiol. 15, 348–352 [DOI] [PubMed] [Google Scholar]
- 27.Dickhout J. G., Hossain G. S., Pozza L. M., Zhou J., Lhoták S., Austin R. C. (2005) Arterioscler. Thromb. Vasc. Biol. 25, 2623–2629 [DOI] [PubMed] [Google Scholar]
- 28.Parker S. H., Parker T. A., George K. S., Wu S. (2006) Mol. Cell Biochem. 293, 173–181 [DOI] [PubMed] [Google Scholar]
- 29.Sood R., Porter A. C., Olsen D. A., Cavener D. R., Wek R. C. (2000) Genetics 154, 787–801 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wek S. A., Zhu S., Wek R. C. (1995) Mol. Cell. Biol. 15, 4497–4506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Harding H. P., Zhang Y., Ron D. (1999) Nature 397, 271–274 [DOI] [PubMed] [Google Scholar]
- 32.Kaufman R. J. (1999) Genes Dev. 13, 1211–1233 [DOI] [PubMed] [Google Scholar]
- 33.Sonenberg N., Hershey J. W. B., Mathews M. (2000) Translational Control of Gene Expression, 2nd Ed., pp. 547–560, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- 34.Berlanga J. J., Santoyo J., De Haro C. (1999) Eur. J. Biochem. 265, 754–762 [DOI] [PubMed] [Google Scholar]
- 35.Pou S., Keaton L., Surichamorn W., Rosen G. M. (1999) J. Biol. Chem. 274, 9573–9580 [DOI] [PubMed] [Google Scholar]
- 36.Deliconstantinos G., Villiotou V., Stravrides J. C. (1995) Br. J. Pharmacol. 114, 1257–1265 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Huk I., Nanobashvili J., Neumayer C., Punz A., Mueller M., Afkhampour K., Mittlboeck M., Losert U., Polterauer P., Roth E., Patton S., Malinski T. (1997) Circulation 96, 667–675 [DOI] [PubMed] [Google Scholar]
- 38.Beckman J. S., Koppenol W. H. (1996) Am. J. Physiol. 271, C1424–C1437 [DOI] [PubMed] [Google Scholar]
- 39.Groves J. T. (1999) Curr. Opin. Chem. Biol. 3, 226–235 [DOI] [PubMed] [Google Scholar]
- 40.Zou M. H., Hou X. Y., Shi C. M., Nagata D., Walsh K., Cohen R. A. (2002) J. Biol. Chem. 277, 32552–32557 [DOI] [PubMed] [Google Scholar]
- 41.Zou M. H., Shi C., Cohen R. A. (2002) J. Clin. Invest. 109, 817–826 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Curran R. D., Ferrari F. K., Kispert P. H., Stadler J., Stuehr D. J., Simmons R. L., Billiar T. R. (1991) FASEB J. 5, 2085–2092 [DOI] [PubMed] [Google Scholar]
- 43.Oh-Hashi K., Maruyama W., Isobe K. (2001) Free Radic. Biol. Med. 30, 213–221 [DOI] [PubMed] [Google Scholar]
- 44.Ogawa Y., Saito Y., Nishio K., Yoshida Y., Ashida H., Niki E. (2008) Free Radic. Res. 42, 674–687 [DOI] [PubMed] [Google Scholar]
- 45.Tyrrell R. M., Pidoux M. (1986) Photochem. Photobiol. 44, 561–564 [DOI] [PubMed] [Google Scholar]
- 46.Mascarenhas C., Edwards-Ingram L. C., Zeef L., Shenton D., Ashe M. P., Grant C. M. (2008) Mol. Biol. Cell 19, 2995–3007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hinnebusch A. G. (1988) Microbiol. Rev. 52, 248–273 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Crabeel M., Huygen R., Verschueren K., Messenguy F., Tinel K., Cunin R., Glansdorff N. (1985) Mol. Cell. Biol. 5, 3139–3148 [DOI] [PMC free article] [PubMed] [Google Scholar]