Abstract
By combining biochemical purification and mass spectrometry, we identified proteins associated with human heterochromatin protein 1α (HP1α) both in the nucleoplasm and in chromatin. Some of these are RNA-binding proteins, and among them is the protein heterogeneous nuclear ribonucleoprotein U (hnRNP U)/SAF-A, which is linked to chromatin organization and transcriptional regulation. Here, we demonstrate that hnRNP U is a bona fide HP1α-interacting molecule. More importantly, hnRNP U depletion reduces HP1α-dependent gene silencing and disturbs HP1α subcellular localization. Thus, our data demonstrate that hnRNP U is involved in HP1α function, shedding new light on the mode of action of HP1α and on the function of hnRNP U.
Chromatin condensation, an essential component of transcriptional repression, is controlled by various enzymes modifying histones or DNA and by proteins binding to modified histones. Thus, enzymes methylating histone H3 lysine 9 (H3K9) are key elements of transcriptional regulation (1). Methylated H3K9, a hallmark of condensed and repressed heterochromatin, is specifically recognized by HP13 proteins (2, 3). HP1 proteins are well conserved (from Schizosaccharomyces pombe to mammals), and three isoforms (HP1α, HP1β, and HP1γ) have been characterized in higher eukaryotes. Mammalian HP1α interacts with a wide range of transcriptional regulators and chromatin-remodeling factors, as well as with DNA and RNA (4). It has thus been proposed that HP1α serves as an adaptor that assembles multimeric complexes on chromatin (5). Indeed, in addition to its role in chromatin compaction, there is evidence to support the idea that mammalian HP1α plays a major role in chromosomal segregation during mitosis (6, 7). It has also been shown, in fission yeast and in Drosophila, that HP1 couples the RNA interference machinery to chromatin condensation (8–10). Thus, HP1α participates in various aspects of chromatin organization and nuclear functions by interacting with a wide range of proteins and nucleic acids in specific nuclear territories.
Here, we characterized novel HP1α-interacting proteins both in the nucleoplasm and in chromatin. Among identified partners, we found RNA-binding proteins and, in particular, hnRNP U/SAF-A, an RNA- and DNA-binding protein (11). hnRNP U is a protein of the nuclear matrix that is tightly associated with chromatin and also found in soluble hnRNP particles. In addition, this protein plays a major role in chromatin organization and transcriptional regulation (12–18). We show here that hnRNP U is a bona fide HP1α-interacting protein and that depletion of hnRNP U affects HP1α nuclear localization and reduces its transcriptional repression. Thus, our data provide a functional link between hnRNP U and HP1α.
EXPERIMENTAL PROCEDURES
Cell Culture and Stable Cell Line
HeLa S3 cells were grown in spinner cultures in Dulbecco's modified Eagle's medium (PAA) supplemented with 10% fetal calf serum, and C2C12 mouse myoblasts were cultured in Dulbecco's modified Eagle's medium supplemented with 15% fetal calf serum. The HeLa S3 cell line stably expressing FLAG-HA-HP1α was obtained as described previously (19). A cell line transduced with the empty pREV vector was used as a control.
Affinity Purification of Human HP1α-containing Nuclear Soluble and Insoluble Complexes
20 g of cells were used per experiment. Cells were lysed in hypotonic buffer (10 mm Tris-HCl (pH 7.65), 1.5 mm MgCl2, and 10 mm KCl) and disrupted by 18 strokes of a tight-fitting Dounce homogenizer. The cytosolic fraction was separated from the pellet by centrifugation at 4 °C. The nuclear soluble fraction was obtained by incubation of the nuclear pellet in high salt buffer for 30 min at 4 °C and by centrifugation. The chromatin fraction was obtained by digestion of the nuclear pellet with micrococcal nuclease (Sigma). Fractions were ultracentrifuged at 32,000 rpm for 30 min at 4 °C in a Beckman SW-T32 rotor. HP1α was immunoprecipitated with anti-FLAG M2-agarose (Sigma), eluted with FLAG peptide, further affinity-purified with anti-HA antibody-conjugated agarose, and eluted with HA peptide. Complexes were resolved by SDS-PAGE and stained using the Silver Quest kit (Invitrogen). Electrospray ionization tandem mass spectrometry was done in the Taplin Biological Mass Spectrometry Facility (Harvard Medical School) or in the mass spectrometry facility of the Institute André Lwoff.
Immunoprecipitations and Western Blotting
For co-immunoprecipitation experiments, ∼5 mg of HeLa S3 nuclear extracts were precleared with protein G-agarose (Pierce) and submitted to immunoprecipitation using standard procedures.
Plasmids and Recombinant Proteins
GST-HP1α fusion protein was purified using glutathione-agarose beads (Sigma) according to the manufacturer's instruction. The concentration of purified proteins was estimated on Coomassie stains after SDS-PAGE separation (supplemental Fig. S4). Deletion mutants of hnRNP U were described previously (17). TnT reactions were performed with the TnT in vitro lysate system (Promega) under standard conditions. TnT reactions were controlled by autoradiography after SDS-PAGE separation (supplemental Fig. S4). Beads coated with equal amounts of GST fusion protein (1 μg) were incubated with 10 μl of radioactive TnT reaction in reaction buffer (50 mm Tris (pH 7.6), 150 mm NaCl, and 0.1% Triton X-100) for 2 h at 4 °C. Beads were washed five times with wash buffer (50 mm Tris (pH 7.6), 300 mm NaCl, and 0.5% Triton X-100), and proteins were resolved by SDS-PAGE and revealed by autoradiography.
Glycerol Gradient Analysis
Nuclear soluble and chromatin complexes (obtained after two steps of immunoprecipitation) were fractionated on 19–41% glycerol gradients as described previously (20).
Small Interfering RNA Knockdown and Luciferase Assay
Control or hnRNP U siRNA (Qiagen) was transfected into 70% confluent HeLa S3 cells using HiPerFect (Qiagen) or Lipofectamine 2000 (Invitrogen) at a final siRNA concentration of 20 nm. For luciferase assays, C2C12 cells were first transfected with hnRNP U or scrambled siRNA (Qiagen). Following 24 h of incubation, cells were retransfected with 100 ng of Gal4 DBD or Gal4 DBD-HP1α vector together with 500 ng of Gal4-luciferase plasmid and 50 ng of firefly Renilla plasmid using Lipofectamine 2000 (Invitrogen). Luciferase activity was measured using the Dual-Luciferase® reporter assay system (Promega).
Immunofluorescence
Cells were fixed with 4% paraformaldehyde for 10 min at room temperature and stained using standard procedures.
Quantitative Reverse Transcription-PCR
RNA was extracted with RNeasy minikits (Qiagen) according to the manufacturer's instructions and analyzed by quantitative reverse transcription-PCR on a LightCycler®.
RESULTS
Identification of New HP1α-interacting Proteins
To identify new HP1α-associated proteins, we established a HeLa S3 cell line that stably expresses FLAG-HA tandem epitope-tagged HP1α (as verified by immunofluorescence and Western blotting) (supplemental Fig. S1). Nuclear fractionation was used to discriminate partner proteins associated with nuclear soluble HP1α from those associated with chromatin HP1α. Silver staining of immunoprecipitates revealed several polypeptides co-purifying specifically with tagged HP1α protein (Fig. 1A). Nuclear soluble and chromatin HP1α profiles showed some similarities but were not entirely identical. HP1α partners were characterized by mass spectrometry. The vast majority of HP1α-associated proteins were chromatin components, proteins involved in chromatin remodeling or associated with DNA repair (Table 1). Some have been previously shown to interact with HP1α, such as histones, TIF1β, HP1β, HP1γ, and CAF1. These proteins were found in the two nuclear fractions. Histone methyltransferases, components of the BAF53 complex, and members of the SWI/SNF complex were also specifically detected in the chromatin fraction (Table 1). We also identified, as described previously (7), HP1α protein partners involved in the formation of the mitotic spindle and proteins of the internal nuclear matrix, such as lamins A and C and matrin 3 (Table 1). Interestingly, mass spectrometry analysis revealed the presence of new potential HP1α protein partners (Table 1), among them RNA-binding proteins, such as RNA helicases and hnRNP, that have not been previously identified in HP1α complexes (Table 2). Among the RNA-binding proteins identified was hnRNP U. As this nuclear protein was previously shown to be involved in the organization of chromatin structure and transcriptional repression (12, 16), we decided to focus this study on the potential physical and functional link between hnRNP U and HP1α.
FIGURE 1.
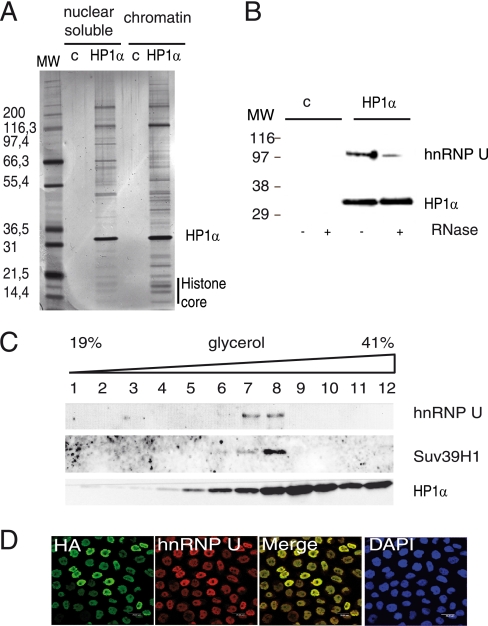
hnRNP U physically interacts with HP1α. A, shown is the characterization of FLAG-HA-HP1α complexes: silver staining of nuclear soluble and insoluble fractions isolated from either HeLa control cells (c) or stably expressing HP1α-tagged protein after two steps of immunoprecipitation. MW, protein molecular weight marker. B, nuclear soluble immunoprecipitates from either HeLa cells (c) or HeLa cells expressing FLAG-HA-HP1α were treated with RNase A prior to incubation with HA-Agarose beads and analyzed by Western blotting using anti-hnRNP U or anti-HP1α (2HP-2G9-AS, Euromedex) antibodies. C, HP1α immunoprecipitates were fractionated on glycerol gradients, and fractions were analyzed by Western blotting using anti-hnRNP U, anti-HP1α, or anti-Suv39h1 antibodies. D, shown is confocal microscopy of HeLa cells expressing tagged HP1α labeled with anti-HA and anti-hnRNP U antibodies. Cells were counterstained with 4′,6-diamidino-2-phenylindole (DAPI).
TABLE 1.
Proteins detected by mass spectrometry in HP1α complexes
| Gene ontology | Peptidea |
Molecular mass | Previously characterized | |
|---|---|---|---|---|
| C | S | |||
| kDa | ||||
| Establishment and/or maintenance of chromatin architecture | ||||
| HP1α | 12 | 10 | 22 | + |
| RUVB1 | 3 | 4 | 50 | − |
| HP1β | 5 | 4 | 21.5 | + |
| HP1γ | 4 | 2 | 21 | + |
| H4 | 8 | 5 | 11 | + |
| H2BA | 7 | 5 | 14 | + |
| H3B | 3 | 4 | 15 | + |
| H2AA | 3 | 3 | 14 | + |
| SSRP/FACT80 | 3 | 7 | 81 | − |
| SP100 | 4 | 1 | 100 | + |
| H2AX | 1 | 1 | 15 | + |
| SMARCA4/Brg1 | 13 | 184 | + | |
| ATRX | 11 | 282 | + | |
| SMARCA2/Brm | 10 | 180 | − | |
| ACTL6A | 7 | 47 | − | |
| CBX8/PC3 | 5 | 43 | − | |
| BAT8/G9a | 4 | 132 | + | |
| DAXX | 4 | 81 | + | |
| SUV39H1 | 4 | 48 | + | |
| HBXAP/RSF1 | 3 | 163 | − | |
| ZN644 | 3 | 150 | − | |
| SMARCA1/SNF2L | 2 | 114 | − | |
| SMARCA5/SNF2H | 2 | 122 | − | |
| HDAC2 | 1 | 55 | + | |
| BAF53 | 1 | 14 | + | |
| H2BF | 1 | 14 | + | |
| H2AY/macroH2A | 1 | 39 | + | |
| FACT complex subunit | ||||
| SPT16 | 3 | 140 | − | |
| RUVB2 | 2 | 50 | − | |
| Mitotic spindle/nuclear architecture | 19 | 7 | 316 | + |
| NIPBL | 5 | 2 | 32 | + |
| DC8 | 2 | 4 | 45 | − |
| STK6/aurora A | 8 | 40 | + | |
| CT172/C20orf172 | 8 | 32 | − | |
| CA048/C1orf48 | 8 | 74 | − | |
| LAMA | 11 | 119 | − | |
| KIF11/Eg5 | 2 | 225 | + | |
| CHTOG | 1 | 36 | + | |
| KI67 | 7 | 17 | 105 | + |
| DNA replication and repair | 12 | 9 | 61 | + |
| CAF1A | 5 | 5 | 47 | + |
| CAF1B | 11 | 155 | − | |
| CAF1C | 36 | 469 | − | |
| POGZ | 19 | 70 | + | |
| PRKDC | 14 | 86 | − | |
| KU70 | 7 | 170 | − | |
| KU86/XRCC5 | 3 | 68 | − | |
| TOP2A | 1 | 128 | − | |
| RAF1/RPA1 | 2 | 56 | − | |
| RFC1 | 1 | 58 | − | |
| Transport | ||||
| IMA3 | ||||
| IMA4 | ||||
a Values indicate the number of different peptides detected by mass spectrometry (nuclear soluble (S) and chromatin (C)). Proteins detected with less than one peptide are considered as non-significant.
TABLE 2.
RNA-binding proteins detected in HP1α immunoprecipitates
| RNA-binding protein | Peptidea |
Molecular mass | Previously characterized | |
|---|---|---|---|---|
| C | S | |||
| kDa | ||||
| HNRPU | 4 | 7 | 90 | − |
| HNRPK | 2 | 6 | 50 | − |
| NPM/nucleophosmin | 1 | 4 | 32 | − |
| SF3B3 splicing factor 3b | 2 | 3 | 135 | − |
| DDX39 | 2 | 2 | 49 | − |
| HNRH1 | 2 | 1 | 49 | − |
| MATR3 | 2 | 2 | 94 | − |
| HNRPC | 2 | 33 | − | |
| DHX21 | 4 | 87 | − | |
| DHX15 | 4 | 90 | − | |
| Prp31 | 2 | 61 | − | |
| PTBP1 | 1 | 55 | − | |
| HNRPR | 1 | 31 | − | |
| DDX17 | 1 | 71 | − | |
| PCBP2 | 1 | 70 | − | |
a Values indicate the number of different peptides detected by mass spectrometry (nuclear soluble (S) and chromatin (C)). Proteins detected with less than one peptide are considered as non-significant.
Biochemical Analysis of hnRNP U-HP1α Interaction
hnRNP U was detected in both the nuclear soluble and chromatin-associated HP1α complexes (Table 2 and supplemental Table S1). The interaction between HP1α and hnRNP U was partially sensitive to treatment with RNase A (Fig. 1B), although this treatment did not modify dramatically the protein profile of the HP1α complex (supplemental Fig. S2). Similarly, less hnRNP U peptides were observed by mass spectrometry after RNase treatment, but the protein was still detected (data not shown). These data suggest that this interaction is, at least in part, RNA-dependent. To verify that hnRNP U is indeed part of a multimolecular complex that includes HP1α, nuclear soluble eluate (after tandem immunoprecipitation) was further purified on glycerol gradients (Fig. 1C). Tagged HP1α was detected over a range of fractions (fractions 5–12), suggesting the co-existence of several complexes. hnRNP U was detected in fractions 6–8 (corresponding to an estimated size of 1 MDa by reference to the ISWI complex) (data not shown), demonstrating that hnRNP U is part of a multimolecular complex. Suv39h1, a well characterized partner of HP1α, was also detected in the same fractions. Mass spectrometry analysis of these fractions indicated that HP1α co-fractionated with well known HP1α protein partners such as TIF1β, HP1β, and HP1γ, as well as with some of the RNA-binding proteins described in Table 2 (hnRNP U, hnRNP K, DHX15, and DDX17), suggesting that hnRNP U could be associated with chromatin proteins linked to transcriptional gene regulation. A similar treatment of the chromatin-associated complex showed that hnRNP U was found in the same fractions as histones H2A and H2AZ, HP1β, HP1γ, and some RNA-binding proteins (supplemental Fig. S3). To further confirm the association between HP1α and hnRNP U, cells expressing tagged HP1α were analyzed by two-color immunofluorescence using a confocal microscope. The results show a significant level of co-localization of hnRNP U and HP1α (Fig. 1D).
hnRNP U Interacts with Endogenous HP1α
To determine whether the interaction between ectopically expressed HP1α and hnRNP U has physiological relevance, we next evaluated whether endogenous hnRNP U and HP1α interact and co-localize in non-transduced HeLa S3 cells. Confocal microscopy images clearly showed a significant co-localization of the two proteins in the nucleus (Fig. 2A). Co-immunoprecipitation experiments showed that endogenous HP1α was detected in anti-hnRNP U immunoprecipitates (Fig. 2B), arguing for the interaction of the two proteins in live cells. The hnRNP U domain of interaction with HP1α was identified in vitro by GST pulldown assays (Fig. 2C) using various portions of the protein. The results demonstrate the involvement of the C-terminal part of hnRNP U, whereas the other domains were not sufficient to mediate the interaction. Taken together, these results indicate that hnRNP U and HP1α are found in association in live cells as well as in vitro.
FIGURE 2.
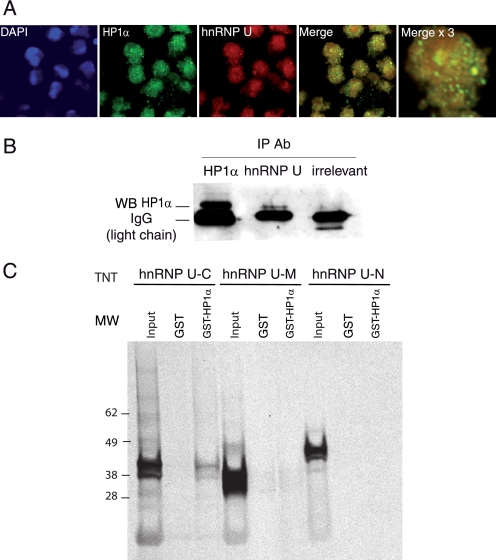
Interaction of endogenous HP1α with hnRNP U. A, HeLa cells were labeled with anti-HP1α and anti-hnRNP U antibodies, counterstained with 4′,6-diamidino-2-phenylindole (DAPI), and observed on a confocal microscope. Images were analyzed using ImageJ software (overlap coefficient, 0.7; Pearson coefficient, 0.68). The right panel is a ×3 magnification. B, nuclear HeLa cell extracts were immunoprecipitated (IP) with anti-HP1α, anti-hnRNP U, or control (irrelevant) antibodies (Ab) and analyzed by Western blotting (WB) using anti-HP1α antibody. C, equivalent amounts (supplemental Fig. S4) of in vitro translated and radiolabeled hnRNP U constructs (C-terminal domain (C), middle domain (M), and N-terminal domain (N)) were incubated with equivalent amounts (supplemental Fig. S4) of GST (negative control) or GST-HP1α. Bound proteins were detected by autoradiography (first three lanes). Input, 10% of the radiolabeled protein used in GST pulldown assays.
Functional Analysis of hnRNP U-HP1α Interaction
We next addressed the involvement of hnRNP U in HP1α function. HP1α is a transcriptional repressor that has been shown previously to severely repress transcription when it is artificially tethered to a promoter. We used such an assay to address the functional link between hnRNP U and HP1α. HP1α was fused to the DBD of the yeast Gal4 protein. Expression vectors bearing either Gal4 DBD or Gal4 DBD-HP1α were transfected into cells along with a Gal4-responsive luciferase reporter (and a Renilla plasmid as an internal control). Down-regulation of hnRNP U did not affect the basal activity of the promoter (supplemental Table S2). Gal4 DBD-HP1α strongly repressed luciferase (supplemental Table S2). This repressive effect was partially reduced by hnRNP U siRNAs (50 and 30% with two different siRNAs) (Fig. 3A). Repression was not fully released, in good concordance with partial hnRNP U down-regulation (Fig. 3B). hnRNP U seems to be an abundant protein and/or a protein important for cell viability. The level of down-regulation of hnRNP U with siRNAs varied from one experiment to another and was often partial, even though mRNA levels were generally dramatically decreased (see, for example, Fig. 4C). In any case, the results from the reporter assay indicate that the hnRNP U protein is required for full repression by HP1α.
FIGURE 3.
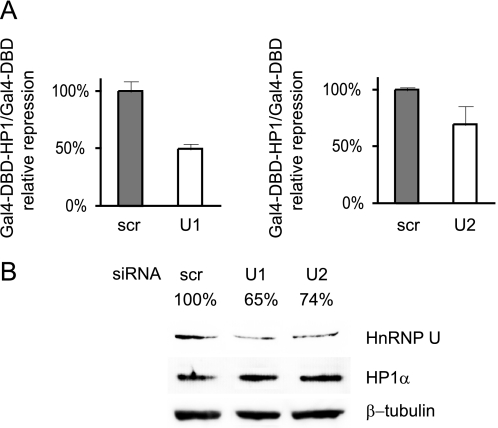
HP1α function depends on hnRNP U. A, C2C12 cells were transfected with Gal4 DBD (control) or Gal4 DBD-HP1α expression vector along with a Gal4-responsive luciferase reporter (and a Renilla expression vector as an internal control) and scrambled (scr) or hnRNP U (U1 and U2) siRNA. B, the Western blot shows down-regulation of hnRNP U with siRNAs U1 and U2 as in A. Bands were quantified using ImageJ software, and results are shown above each lane.
FIGURE 4.
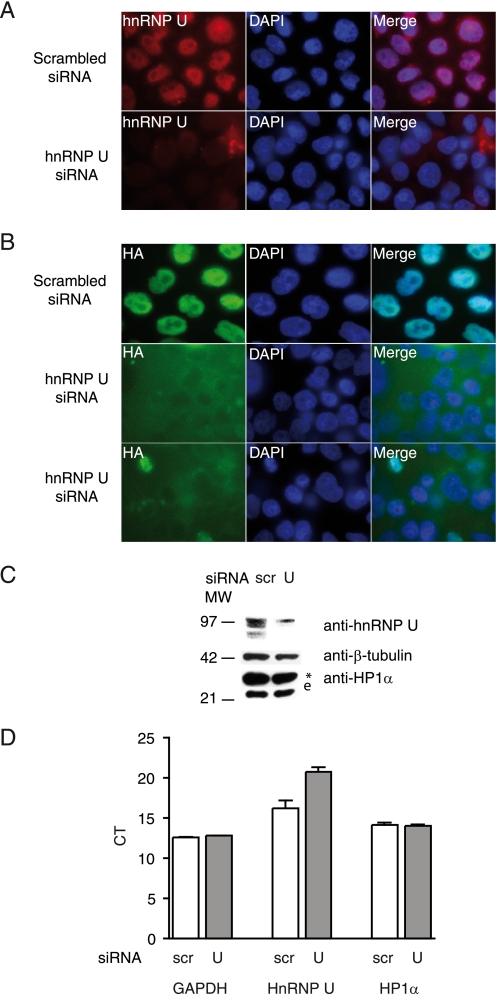
hnRNP U is required for HP1α localization in the nucleus. A and B, HeLa cells stably expressing HP1α-tagged protein were transfected with scrambled or hnRNP U siRNA and labeled with anti-hnRNP U (A) or anti-HA (B) antibodies. DAPI, 4′,6-diamidino-2-phenylindole. C, whole cell extracts of HeLa cells transfected with scrambled (scr) or hnRNP U siRNA were analyzed by Western blotting with the indicated antibodies. *, tagged HP1α; e, endogenous HP1α. D, quantitative reverse transcription-PCR shows the levels of hnRNP U, HP1α, and glyceraldehyde-3-phosphate dehydrogenase (GAPDH; used as an internal control) mRNA expression in cells treated as described for A and B; shown are the cycles (CT; the 4-fold difference for hnRNP U indicates a decrease in expression of ∼20-fold).
hnRNP U Influences HP1α Localization in the Nucleus
We next investigated by which mechanism hnRNP U impacts HP1 activity. hnRNP U is known to act as a scaffold, attaching proteins to chromatin and nuclear lamina. An siRNA-based loss-of-function assay was used to investigate whether hnRNP U might be involved in the subcellular localization of the HP1α protein (Fig. 4). Immunofluorescence experiments on cells ectopically expressing HP1α revealed that, in hnRNP U-depleted cells, as documented by immunofluorescence (Fig. 4A), Western blotting (Fig. 4C), and quantitative reverse transcription-PCR (Fig. 4D), HP1α was translocated out of the nucleus (Fig. 4B), suggesting that hnRNP U influences HP1α cellular localization.
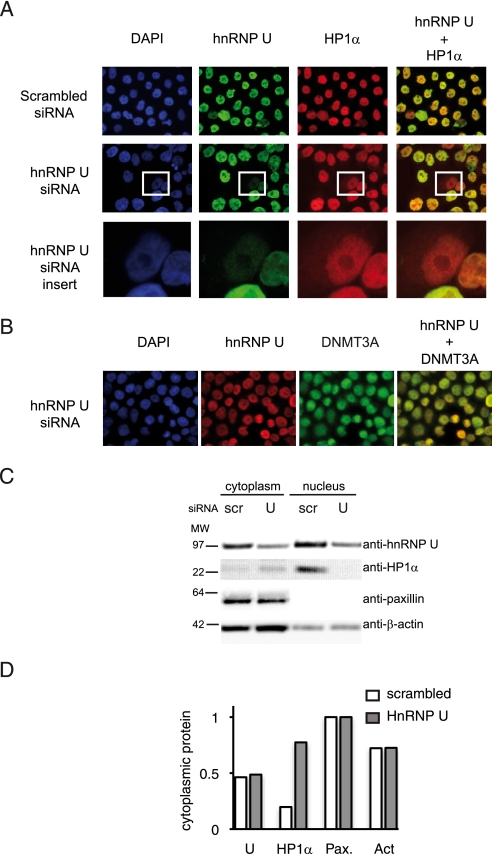
We next explored the effect of hnRNP U depletion on endogenous HP1α. In the experiment shown in Fig. 5, ∼30% of the cells transfected with an hnRNP U siRNA showed a significant reduction in hnRNP U by immunofluorescence. In the vast majority (77%) of these hnRNP U-low cells, the subcellular localization of HP1α was affected, as documented by immunofluorescence (Fig. 5A). The localization of the nuclear protein DNMT3A (DNA (cytosine-5)-methyltransferase 3A), an irrelevant control, was not modified by hnRNP U down-regulation (Fig. 5B). Biochemical cell fractionation was used to further document hnRNP U importance for HP1α subcellular localization. Cell extracts of siRNA-transfected cells were fractionated into nuclear and cytoplasmic fractions. As shown in Fig. 5C (quantitative analysis in Fig. 5D), a reduction in hnRNP U protein (50%) induced a translocation of HP1α from the nucleus into the cytosol (from 20% of total HP1 being cytoplasmic with the control scrambled siRNA to 70% with hnRNP U siRNA). No effect was observed on control proteins such as β-actin and paxillin (a cytoplasmic marker), which remained exclusively or mostly cytoplasmic. These data indicate that hnRNP U is required for HP1α subcellular localization.
FIGURE 5.
HP1α subcellular localization is dependent on hnRNP U. A and B, HeLa cells transfected with scrambled or hnRNP U siRNA were labeled with anti-hnRNP U, anti-HP1α (A; enlarged view in the bottom row), or anti-DNMT3A (B) antibodies as indicated. DAPI, 4′,6-diamidino-2-phenylindole. C, nuclear extracts of HeLa cells transfected with scrambled (scr) or hnRNP U siRNA were fractionated and analyzed by Western blotting with the indicated antibodies. Paxillin was used as a marker of the cytoplasmic fraction. D, the graph represents the proportions of cytoplasmic proteins (ratio between cytoplasmic and total proteins) after quantitative analysis of the gel showed in C using Image J. Pax., paxillin; Act, β-actin.
DISCUSSION
To explore HP1α functions, we have characterized new proteins associated with nuclear soluble and chromatin-associated HP1α. In both complexes, we detected transcriptional repressors, chromatin-remodeling factors, DNA repair proteins, mitotic spindle-associated proteins, and RNA-binding proteins. Among the RNA-binding proteins, we focused our attention on hnRNP U, as this protein participates in transcriptional regulation and chromatin organization (12). We have shown here that hnRNP U co-purifies with HP1α in both nucleoplasmic and chromatin-associated complexes. Interestingly, another member of the family, HP1γ, was also reported to be associated with hnRNP U and matrin 3 (21). By co-immunoprecipitation, two-color immunofluorescence, and GST pulldown, we confirmed that the two proteins interact in vitro and in live cells. The interaction is sensitive to RNase treatment. Moreover, the hnRNP U region of interaction with HP1α is the C-terminal portion, which includes the RNA-binding domain. It is thus likely that the interaction is not direct but rather mediated by some uncharacterized RNA.
We next addressed the functional interaction between HP1α and hnRNP U and showed that down-regulating hnRNP U affected transcriptional repression by HP1α in a reporter assay. This result is in good concordance with the observation that various transcriptional repressors (Suv39h1, HP1β, HP1γ, and TIF1β) are found in the hnRNP U-containing, high molecular weight complex detected in glycerol gradients. Moreover, a repressive function was previously shown for hnRNP U in X chromosome silencing and in transcriptional repression (16, 22).
We next explored the mechanism through which hnRNP U affects HP1α function. hnRNP U is known to be located at the nuclear matrix and binds to scaffold-associated region DNA. The nuclear matrix plays a major role in the structural organization of chromatin, in the integrity of the nucleus, and in transcriptional regulation via the scaffold/matrix attachment regions. Nuclear lamina and heterochromatin are attached to the inner nuclear membrane at the nuclear envelope, and HP1α is tightly associated with the inner nuclear membrane protein lamin B receptor at the nuclear envelope (23). We thus tested the hypothesis that hnRNP U influences HP1α proper subcellular localization. We observed by immunofluorescence and biochemical cell fractionation experiments that down-regulating hnRNP U using specific siRNAs indeed perturbed HP1α localization.
HP1α is a small protein (<40 kDa) that could, in theory, diffuse passively through the nuclear pore and that requires nuclear retention signals. We believe that hnRNP U provides such a signal. Moreover, the localization of HP1α at the nuclear periphery seems to be highly dependent on hnRNP U: in hnRNP U-low cells, HP1α is more severely depleted at the nuclear periphery than in the inner nucleus (Fig. 5). We therefore conclude that hnRNP U is necessary to anchor HP1α to peripheral chromatin territories.
Previously published data obtained by cell treatment with RNase support the hypothesis that an RNA component plays a major role in targeting HP1α to chromatin in mammalian cells (24). Our finding that an RNA-binding protein is associated with HP1α, in an RNA-dependent manner, reinforces this idea and raises the possibility that hnRNP U provides an additional molecular link between HP1 and RNA. In this model, it would be partly through hnRNP U, and maybe other RNA-binding proteins, that RNA tethers HP1α to chromatin. Taken together, our data clearly show that hnRNP U and HP1α interact in human cells and that hnRNP U is required for HP1α repressive activity and subcellular localization, shedding new light on the mode of action of HP1 and on the function of hnRNP U.
Supplementary Material
Acknowledgments
We thank Michael Carey and Régine Losson for the gift of plasmids; Khalid Ouararhni, Alexandra Garancher, and Arnaud Legrand for technical help; and Linda L. Pritchard, Anna Polesskaya, Pascal Drané, Nora Nonne, and Jacques Matthieu for critical reading of the manuscript.
This work was supported by CNRS and by European Union 6th Framework Programme TRANS-REG Grant LSHG-CT-2004-502950 and SIROCCO Grant LSHG-CT-2006-037900.

The on-line version of this article (available at http://www.jbc.org) contains supplemental “Experimental Procedures,” Figs. S1–S4, and Tables S1 and S2.
- HP1
- heterochromatin protein 1
- hnRNP
- heterogeneous nuclear ribonucleoprotein
- HA
- hemagglutinin
- GST
- glutathione S-transferase
- siRNA
- small interfering RNA
- DBD
- DNA-binding domain.
REFERENCES
- 1.Nakayama J., Rice J. C., Strahl B. D., Allis C. D., Grewal S. I. (2001) Science 292, 110–113 [DOI] [PubMed] [Google Scholar]
- 2.Lachner M., O'Carroll D., Rea S., Mechtler K., Jenuwein T. (2001) Nature 410, 116–120 [DOI] [PubMed] [Google Scholar]
- 3.Peters A. H., O'Carroll D., Scherthan H., Mechtler K., Sauer S., Schöfer C., Weipoltshammer K., Pagani M., Lachner M., Kohlmaier A., Opravil S., Doyle M., Sibilia M., Jenuwein T. (2001) Cell 107, 323–337 [DOI] [PubMed] [Google Scholar]
- 4.Maison C., Almouzni G. (2004) Nat. Rev. Mol. Cell Biol. 5, 296–304 [DOI] [PubMed] [Google Scholar]
- 5.Lechner M. S., Schultz D. C., Negorev D., Maul G. G., Rauscher F. J., 3rd (2005) Biochem. Biophys. Res. Commun. 331, 929–937 [DOI] [PubMed] [Google Scholar]
- 6.Sugimoto K., Tasaka H., Dotsu M. (2001) Cell Struct. Funct. 26, 705–718 [DOI] [PubMed] [Google Scholar]
- 7.Obuse C., Iwasaki O., Kiyomitsu T., Goshima G., Toyoda Y., Yanagida M. (2004) Nat. Cell Biol. 6, 1135–1141 [DOI] [PubMed] [Google Scholar]
- 8.Brower-Toland B., Findley S. D., Jiang L., Liu L., Yin H., Dus M., Zhou P., Elgin S. C., Lin H. (2007) Genes Dev. 21, 2300–2311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pal-Bhadra M., Leibovitch B. A., Gandhi S. G., Rao M., Bhadra U., Birchler J. A., Elgin S. C. (2004) Science 303, 669–672 [DOI] [PubMed] [Google Scholar]
- 10.Zofall M., Grewal S. I. (2006) Cold Spring Harbor Symp. Quant. Biol. 71, 487–496 [DOI] [PubMed] [Google Scholar]
- 11.Fackelmayer F. O., Richter A. (1994) Biochim. Biophys. Acta 1217, 232–234 [DOI] [PubMed] [Google Scholar]
- 12.Fackelmayer F. O., Dahm K., Renz A., Ramsperger U., Richter A. (1994) Eur. J. Biochem. 221, 749–757 [DOI] [PubMed] [Google Scholar]
- 13.Romig H., Fackelmayer F. O., Renz A., Ramsperger U., Richter A. (1992) EMBO J. 11, 3431–3440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Göhring F., Fackelmayer F. O. (1997) Biochemistry 36, 8276–8283 [DOI] [PubMed] [Google Scholar]
- 15.Helbig R., Fackelmayer F. O. (2003) Chromosoma 112, 173–182 [DOI] [PubMed] [Google Scholar]
- 16.Kim M. K., Nikodem V. M. (1999) Mol. Cell. Biol. 19, 6833–6844 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kukalev A., Nord Y., Palmberg C., Bergman T., Percipalle P. (2005) Nat. Struct. Mol. Biol. 12, 238–244 [DOI] [PubMed] [Google Scholar]
- 18.Yugami M., Kabe Y., Yamaguchi Y., Wada T., Handa H. (2007) FEBS Lett. 581, 1–7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ouararhni K., Hadj-Slimane R., Ait-Si-Ali S., Robin P., Mietton F., Harel-Bellan A., Dimitrov S., Hamiche A. (2006) Genes Dev. 20, 3324–3336 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tagami H., Ray-Gallet D., Almouzni G., Nakatani Y. (2004) Cell 116, 51–61 [DOI] [PubMed] [Google Scholar]
- 21.Malyavantham K. S., Bhattacharya S., Barbeitos M., Mukherjee L., Xu J., Fackelmayer F. O., Berezney R. (2008) J. Cell. Biochem. 105, 391–403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fackelmayer F. O. (2005) J. Biol. Chem. 280, 1720–1723 [DOI] [PubMed] [Google Scholar]
- 23.Kourmouli N., Theodoropoulos P. A., Dialynas G., Bakou A., Politou A. S., Cowell I. G., Singh P. B., Georgatos S. D. (2000) EMBO J. 19, 6558–6568 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maison C., Bailly D., Peters A. H., Quivy J. P., Roche D., Taddei A., Lachner M., Jenuwein T., Almouzni G. (2002) Nat. Genet. 30, 329–334 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.