Abstract
The human Nup107-160 nucleoporin complex plays a major role in formation of the nuclear pore complex and is localized to kinetochores in mitosis. Here we report that Seh1, a component of the Nup107-160 complex, functions in chromosome alignment and segregation by regulating the centromeric localization of Aurora B and other chromosome passenger complex proteins. Localization of CENP-E is not affected by Seh1 depletion and analysis by electron microscopy showed that microtubule kinetochore attachments are intact. Seh1-depleted cells show impaired Aurora B localization, which results in severe defects in biorientation and organization of the spindle midzone and midbody. Our results indicate that a major function of the Nup107 complex in mitosis is to ensure the proper localization of the CPC at the centromere.
INTRODUCTION
The nuclear envelope (NE) forms the interface between the nucleus and the cytoplasm of the interphase eukaryotic cell and is essential to maintain the unique identity of each compartment. Transport between the two compartments takes place via the nuclear pore complexes (NPCs) of which there are several thousand in vertebrate somatic cells (Allen et al., 2000; Conti and Izaurralde, 2001). Each NPC contains multiple subunits of ∼30 proteins called nucleoporins (Nups; Cronshaw et al., 2002). In higher eukaryotes NPCs are stable throughout interphase (Daigle et al., 2001), but during mitosis both the nuclear envelope and NPC undergo major structural reorganization. Starting early in prometaphase, breakdown of the nuclear envelope occurs, including disassembly of the nuclear lamina and NPCs. The nuclear envelope membrane proteins and the transmembrane nucleoporins relocalize to the endoplasmic reticulum (ER), whereas the rest of the nuclear envelope and NPC components become distributed throughout the mitotic cytoplasm (Antonin et al., 2008).
During mitosis, two complete sets of chromosomes are delivered to a pair of daughter cells. Segregation of each sister chromatid pair is achieved by a highly orchestrated process that requires attachment of the sister kinetochores of each chromosome to microtubules emanating from opposite spindle poles. Kinetochores are protein structures that assemble on chromosome regions known as centromeres and mediate microtubule attachment, mitotic checkpoint signaling, and force generation (Maiato et al., 2004; Tanaka et al., 2005; Cheeseman and Desai, 2008). Electron microscopy (EM) has provided information regarding the structure of vertebrate kinetochores and has led to the division of kinetochores into three distinct regions: the inner kinetochore that associates with chromatin, the outer kinetochore that interacts with spindle microtubules, and the less dense middle kinetochore region (McEwen et al., 2007). Multiple different MT-associated proteins function at kinetochores to form a core attachment site between kinetochore and MTs, with the KMN network (KNL–1/Mis12/NDC80 complex) being the major structural component (Cheeseman et al., 2006). Functional analysis of the NDC80 subcomplex both in yeast and mammalian cells has revealed its importance for kinetochore–microtubule (kMT) attachment. Cells that have impaired NDC80 complex function have elongated spindles and exhibit loss of tension at kinetochores as well as chromosome alignment problems (Wigge and Kilmartin, 2001; DeLuca et al., 2002; McCleland et al., 2003). Other proteins that function in parallel to the KMN network include the motor proteins CENP-E (McEwen et al., 2001), dynein (Howell et al., 2001; Yang et al., 2007), and the large coiled coil protein CENP-F (Feng et al., 2006).
Proper kinetochore attachment by MT bundles is achieved in a stepwise manner and misoriented kMT connections must be removed for biorientation to be established. The chromosome passenger complex (CPC), composed of the Aurora B kinase and its regulatory subunits INCENP, survivin, and borealin, is required for proper biorientation as well as affecting other important chromosomal and cytoskeletal events in mitosis. At the beginning of cell division, the CPC concentrates at the inner centromere (Ruchaud et al., 2007) but can also decorate the arms of chromosomes. At the metaphase-to-anaphase transition, CPC localization changes from centromeres to the central spindle and cell cortex (at the site of cleavage furrow formation) before eventually associating with the midbody in cytokinesis (Vagnarelli and Earnshaw, 2004; Tanaka et al., 2005). The CPC core components interact together to create a single structural unit that is involved in chromosome condensation, spindle assembly, chromosome biorientation, signaling to the spindle checkpoint, and completion of cytokinesis (Jeyaprakash et al., 2007; Ruchaud et al., 2007).
The Nup107-160 complex (Nup107 complex) is an evolutionary conserved nucleoporin subcomplex that plays a crucial role in nuclear pore complex assembly (Boehmer et al., 2003; Harel et al., 2003; Walther et al., 2003). A small fraction of the Nup107 complex localizes to kinetochores from early prophase to late anaphase (Belgareh et al., 2001). Certain other nucleoporins and nuclear export factors (Nup358, Rae1, CRM1) have also been shown to exert a mitotic function and to be located at kinetochores and/or spindle poles during mitosis (Arnaoutov et al., 2005; Blower et al., 2005). Studies on the kinetochore localization of the Nup107 complex have shown that stable anchoring of the complex at the kinetochore requires the NDC80 complex and CENP-F (Zuccolo et al., 2007). Efficient depletion of the Nup107 complex causes chromosome congression defects and a spindle checkpoint-dependent delay in mitosis (Zuccolo et al., 2007). These results suggest a defect in attachment of MTs to kinetochores in cells depleted of the Nup107 complex, but exactly how these defects occur was not yet known.
We have examined in detail the consequences of perturbing the Nup107 complex in mitotic cells. We find that the Nup107 complex plays a critical role in chromosome alignment and segregation by regulating the centromeric localization of Aurora B and the CPC proteins.
MATERIALS AND METHODS
Cell Culture
HeLa Kyoto and U20S cells were grown in DMEM, supplemented with 10% fetal calf serum, 0.2 mM l-glutamine, 100 U/ml penicillin, and 100 μg/ml streptomycin. HeLa KyotoEGFP-seh1 cell line was generated by transfection of EGFP-seh1 cDNA to HeLa Kyoto cells and maintained in G418. HeLa KyotoEGFP-H2B cell line was used for analysis of mitotic progression.
RNAi and Transfection Experiments
cDNA transfections were performed using Fugene (Roche, Indianapolis, IN). RNAi experiments were performed using annealed siRNA oligos (Ambion, Austin, TX, and Qiagen, Chatsworth, CA) using HiPerFect reagent (Qiagen). Full sequences of siRNA oligos are provided in Supplemental Table S1.
Electron Microscopy
HeLa cells grown on etched grid coverslips (Bellco, Vineland, NJ) were fixed with 2.5% glutaraldehyde in cacodylate buffer (pH 7.2) for 30 min and stained with 1% osmium tetroxide for 40 min followed by 0.5% uranyl acetate staining. After dehydration in EtOH coverslips were dipped in propylene oxide and quickly placed on top of a BEEM capsule filled with Epon (Roth, Karlsruhe, Germany). Blocks were incubated at 60°C for 48 h. Serial sections were cut 60 nm thick and placed on a copper palladium slot grid coated with 1% Formvar (Serva Biochemicals, Paramus, NJ). Imaging was done on a Biotwin (Philips Electronic Instruments, Mahwah, NJ) 120-kV electron microscope. EM analysis was carried out in six cells both from control and Seh1-depleted cells.
Microscopy and Image analysis
For immunofluorescence, three-dimensional (3D) data sets were acquired using a CoolSnap HQ cooled CCD camera (Photometrics, Tucson, AZ), on a DeltaVision Spectris microscope (Applied Precision, Issaquah, WA). Optical sections were acquired every 0.2 μm, and 3D data sets were deconvolved using the constrained iterative algorithm (Swedlow et al., 1997; Wallace et al., 2001) implemented in the SoftWoRx software (Applied Precision). For live cell imaging HeLa cells and HeLaEGFPH2B cells grown on glass-bottomed Lab-Tek dishes (Nunc, Rochester, NY) were transfected with appropriate siRNAs and maintained in a humidified 37°C chamber in a CO2-independent Phenol Red–free DMEM medium (Invitrogen). Images were collected using a 100×/1.4 NA PlanApo objective lens and a 20×/0.4 PlanFL Ph1 objective. Images were loaded into Photoshop (Adobe, San Jose, CA) or OMERO (www.openmicroscopy.org) and adjusted for display.
Quantification of Aurora B, survivin, MKLP2 and P-MKLP1, MCAK, P-MCAK, Nup107, and Mel28 was carried out using 2D Polygon Finder and 3D Object Builder implemented in the SoftWoRx software. Briefly for quantification of the signal at the metaphase plate the DAPI signal was used to define the chromosome area and create a 3D model from the 2D polygons in each z section. At anaphase the tubulin signal of MTs was used to create a 3D polygon of the central spindle, whereas at cytokinesis the midbody signal was used. The kinetochore signal was determined relative to an ACA reference channel. The fluorescence Aurora B, MKLP2, and P-MKLP1, MCAK, P-MCAK, and survivin signal within that volume was quantified, background corrected and represented as fluorescent intensity per pixel.
cDNAs, Antibodies, and Immunofluorescence
All fixation, permeabilization and immunostaining were performed at room temperature (RT). Cells grown on coverslips were fixed in a 3.7% formaldehyde/PBS solution for 10 min and permeabilized in PBS-0.5% Triton X-100 (Tx100) for 10 min. Cells were blocked in 10% normal donkey serum for 1 h at RT before antibody incubations. Antibodies used in this study: Aurora B (Ab2254, Abcam, Cambridge, MA), Hec1 (9G3, Abcam), survivin (Ab469, Abcam), BuBR1(Ab4637, Abcam), ACA (Abcam), CRM1 (H-300, Santa-Cruz Biotechnology), RanGAP1 (N-19, Santa Cruz Biotechnology), tubulin (DM1A; Sigma, St. Louis, MO), Mel28 (Franz et al., 2007), MKLP1, MKLP2, and P-MKLP1 (Neef et al., 2006), MCAK and P-MCAK (Andrews et al., 2004), RanBP2 (Hutten et al., 2008), Plk 1(35-206) (Upstate Biotechnology, Lake Placid, NY), phospho-histone H3 (Ser10; Upstate), pericentrin (Ab 448, Abcam), borealin (Gassmann et al., 2004), and INCENP (Ab36453, Abcam). Antibodies against human Nup43 and human Seh1 were raised in rabbits against the full-length proteins. Antibody to Human MEL-28 (FGj4) was generated in rabbits against residues Arg 1208–Gln 1800. All affinity-purified donkey secondary antibodies (labeled either with FITC, TRITC, or CY5) were purchased from Jackson ImmunoResearch (West Grove, PA).
Nocodazole was used at 100 ng/ml for 12 h, taxol at 10 μM for 1 h, and monastrol at 100 μM (Calbiochem, La Jolla, CA) for 2 h. ZM447439 (Tocris Bioscience, Ballwin, MO) was used at 3 μM and hesperadin (Dr. Peter Lenart, EMBL) at 100 nM, leptomycin B (L2913, Sigma) at 20 nM. For analysis of cold stable MTs, cells were incubated in ice-cold medium for 10 min, before incubation in ice-cold PHEM buffer (60 mM PIPES, 25 mM HEPES, pH 6.9, 10 mM EGTA, and 4 mM MgSO4)/0.5% Tx100 for 2 min. Fixation was carried out in ice-cold methanol for 2 min, followed by 2× PBS wash and final rehydration in fresh PBS for 15 min. Inter-KT distances were measured using the kinetochore marker Hec1 on sections of deconvolved 3D data sets at the same focal plane. At least 60 kinetochore pairs from a total of 10 cells per condition were used. For spindle length measurements cells were stained with centromeric marker (ACA) to identify clear metaphase plates and centrosome marker (pericentrin). Pole-to-pole measurements were made from single sections of 3D data sets using OMERO. Only spindles parallel to the plane of focus were used to avoid artifacts that can be created by spindle rotation into the plane of focus.
For analysis of CRM1, RanGAP and RanBP2 localization, cells were rinsed in PHEM buffer incubated for 45 s in 0.1% Tx100 in PHEM buffer before fixation in 3.7% formaldehyde in PHEM buffer for 10 min, followed by 2× PBS wash and postfixation in ice-cold methanol for 5 min.
siRNA-resistant GFP-Seh1 cDNA was generated by site-directed mutagenesis carrying the following silent nucleotide mutations (bold letters) at positions 168-171-173, AAG ACA CAC AGC GAA TCT.
Fluorescence Recovery after Photobleaching
Fluorescence recovery after photobleaching (FRAP) was performed as described in Andrews et al. (2004). HeLaGFP-Aurora B stable cell line was maintained at 37°C in a CO2-independent medium. Experiments were carried out using the FRAP-enabled Delta Vision Spectris (Applied Precision) fitted with a 10-MW 488-nm solid-state laser. Recovery models and half-lives were generated using the method of Axelrod (Axelrod et al., 1976) implemented in the SoftWoRx software.
Statistical Analysis
To assess statistical significance, we determined the Student's t test value for equal or unequal variances, and p < 0.05 was considered to be statistically significant.
RESULTS
Kinetochore Localization of Nup107 Complex Is Required for Completion of Mitosis and Cytokinesis in Human Cells
Depletion of Nup107 from human cells has previously been shown to cause defects in mitosis (Zuccolo et al., 2007). Nup107 complex composed of 10 different nucleoporins (Nup160, Nup133, Nup107, Nup96, Nup85, Nup43, Nup37, Sec13, Seh1, and Mel28) has also been involved in spindle assembly in the Xenopus system (Orjalo et al., 2006). At the same time down-regulation of the Nup107 complex components Nup107 and Nup133 results in obvious nuclear transport defects when assayed by nuclear accumulation of polyA+ RNA (Walther et al., 2003) but also gives rise to an increase in midbody MTs (Rasala et al., 2006).
We decided to analyze further the functions of the different subunits of the Nup107 complex in live human cells during the cell cycle and specifically in mitosis in order to characterize the mechanism of mitotic defects observed. For this we generated a stable cell line expressing the Nup107 complex component Seh1 fused to enhanced green fluorescent protein (EGFP). In this cell line, EGFP-Seh1 is properly localized to nuclear pores and kinetochores and immunoprecipitates show association to Nup107 complex (data not shown). This allowed us to study the properties of the Nup107 complex using EGFP-Seh1 as a marker after depletion of any of the other Nup107 complex constituents by siRNA.
The Nup107 complex is composed in vertebrates of 10 different nucleoporins, and therefore we worked to establish the effects of depleting one of the components of the complex on the rest of the complex's members. Western blotting and immunofluorescence against members of the Nup107 complex confirmed the specificity of the siRNAs used (Supplemental Figure S1, A and E). Depletion of Nup107, Mel28, and Seh1 decreased the expression of these proteins, whereas scrambled control siRNAs had no effect. The level of expression of other members of the Nup107 complex, such as the nucleoporins Nup43 and Nup37 (Supplemental Figure S1E), was not substantially decreased. We decided to concentrate on the nucleoporin Seh1, whose depletion showed a clear inhibition of cell proliferation (Supplemental Figure S1D) and interfered with targeting of the Nup107 complex to the kinetochore, as assayed by Mel28 immunolocalization (Supplemental Figure S1, B and C; Franz et al., 2007). On Seh1 depletion the protein levels of Mel28 remain unchanged (Supplemental Figure S1E), whereas kinetochore signals of Mel28 was reduced to 9% of the residual staining and of Nup107 to 12% (Supplemental Figure S1C).
Depletion of Nup107 resulted in an increase in misaligned chromosomes in metaphase and lagging chromosomes in anaphase (Figure 1A). Dual wavelength time-lapse microscopy of HeLaEGFP-seh1 stable cell line depleted of Nup107 and transfected with a mRFP-tagged version of Histone H2B showed misaligned chromosomes that failed to align with the majority of chromosomes on the metaphase plate and remained close to the spindle poles. In control cells the metaphase-to-anaphase transition was completed in 25 min, whereas Nup107-depleted cells remained in metaphase for at least 35 min without completing correct chromosome alignment before anaphase onset.
Figure 1.
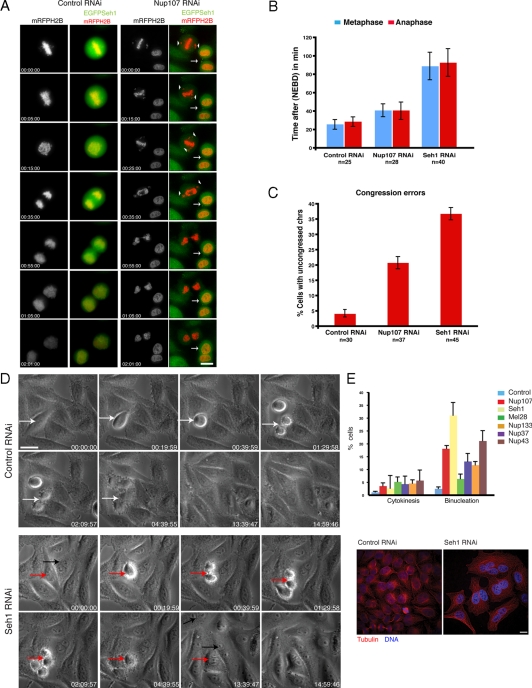
Seh1 and the Nup107 complex are required for chromosome alignment and completion of cytokinesis in human cells. (A) Depletion of Nup107 by siRNA causes major chromosome alignment defects and binucleation. Selected maximum intensity projections from time-lapse images show the mitotic and cytokinesis defects. HeLaEGFP-Seh1 cells (in green) were transfected with mRFP-H2B to label chromatin (in red) and either control or Nup107 siRNAs. Images were collected at 60 h for a period of 120 min. Numbers indicate time in h:min:s. White arrow points to the cleavage furrow ingression and formation of binucleated cell. Scale bar, 10 μm. (B) Mitotic progression plots of metaphase onset (blue bars) and anaphase onset (red bars) from live HeLaEGFPH2B cells treated with control, Nup107, and Seh1 siRNAs. NEBD is set as T = 0. Data were collected after 50 h of RNAi treatment for a period of 2 h every 2 min. (C) Percentage of HeLaEGFP-H2B cells treated with control, Nup107 and Seh1 siRNAs having uncongressed chromosomes at T = 24 min (time of anaphase onset of control cells). Images were collected after 50 h of RNAi treatment for a period of 2 h every 2 min. (D) HeLa cells transfected with control or Seh1 siRNAs were incubated for 48 h before phase-contrast imaging for further 16 h. Images were acquired every 20 min. Note the normal cell division of control cells (white arrow) and the binucleated cells forming (red arrow) or already existing (black arrow) in Seh1-depleted cells. Scale bar, 10 μm. (E) The percentage of cells with a clear cytokinesis bridge and two nuclei, in control and Nup107 complex members depleted cells (siRNAs against Nup107, Seh1, Mel28, Nup133, Nup37, Nup43) were determined 72 h after transfection. At least 300 cells were counted in each category. Cells were stained with α-tubulin, α-Nup37, α-Nup43, α-Mel28, α-Nup107, α-Seh1, α-mAb414, and DNA (DAPI). Corresponding immunofluorescence images of control and Seh1-depleted cells stained with antibodies to tubulin (red) and DNA (blue) are shown. Scale bar, 10 μm.
We next examined the details of mitotic progression upon Seh1 depletion. Chromosome movement in both Nup107 and Seh1-depleted cells were examined in detail by imaging HeLaEGFPH2B living cells after 50 h of RNAi treatment for a period of 2 h every 2 min. Nuclear envelope breakdown (NEBD) was set as time point zero T = 0 and the times to chromosome congression at the metaphase plate and anaphase onset were determined. NEBD was defined as the point that early prophase cells showing condensed chromosomes still had intact nuclear shape and morphology similar to neighboring interphase cells with uncondensed chromosomes. Control cells completed chromosome congression by T = 24 min and normally initiated anaphase by T = 35 min without evidence of lagging or unaligned chromosomes in anaphase. HeLaEGFPH2B cells transfected with Nup107 siRNAs had the majority of chromosomes reaching congression at the metaphase plate by T = 40 min and initiated anaphase by T = 42 min. In contrast, HeLaEGFPH2B cells transfected with Seh1 siRNAs had the majority of chromosomes reaching congression plate at T = 87 min followed by anaphase onset at T = 94 min (Figure 1B). The above suggest that Nup107 and especially Seh1-depleted cells have problems aligning chromosomes at the metaphase plate but once a reasonable metaphase plate is reached (even if not all chromosomes are aligned) progression to anaphase occurs without a full mitotic arrest. We quantified the percentage of cells with uncongressed chromosomes at the time point that control cells normally enter anaphase, which is T = 24 min. On Nup107 and Seh1 depletion, 20 and 37% of cells respectively, had uncongressed chromosomes versus 4% in control cells (Figure 1C). At the same time the spindle assembly checkpoint was activated as detected by the presence of BubR1 staining at unattached kinetochores (Supplemental Figure S2D).
Depletion of Nup133 or ELYS causes an increase in the appearance of midbodies (Rasala et al., 2006), suggesting a defect in cytokinesis. We therefore wanted to determine if the same phenotype occurs after down-regulation of other members of the Nup107 complex. Indeed, down-regulation of the Nup43, MEL28, and in particular Seh1 resulted in an increase in binucleated and multinucleated cells (Figure 1E, graph), both in HeLa and in U2OS cells. More specifically the number of binucleated cells varied depending on the member of the Nup107 complex that was targeted but increased dramatically upon depletion of Seh1, from 3 to ∼37% (Figure 1E, graph and immunofluorescence images). At the same time the number of cells with a clear cytokinesis bridge increased from 2% to almost 7% for Seh1 and up to 12% for Nup43 (Figure 1E, graph). Time-lapse phase-contrast microscopy of live cells confirmed the phenotype observed in fixed cells. HeLa cells transfected with Seh1 siRNA were incubated for 48 h before imaging for a further 16 h. At 48 h Seh1-depleted cells already had started showing signs of binucleation (Figure 1D, black arrow). Live cell imaging revealed that although the cytokinetic furrow ingressed efficiently, it subsequently regressed and no abscission occurred, resulting in binucleation (Figure 1D, red arrows). Control cells divided normally and cytokinesis was completed within 2 h (Figure 1D, white arrow) relative to Seh1-depleted cells that remained in cytokinesis before furrow regression for much longer periods of time (e.g., Figure 1D, red arrows; see also Figure 1A, arrow). Transfection of a HeLa cell line stably expressing a siRNA resistant form of GFP-Seh1 with a siRNA oligo targeting Seh1 gave no significant increase in binucleation, suggesting the siRNA depletion is specific for Seh1 (data not shown). Seh1 therefore appears to be required for normal cleavage furrow ingression and cytokinesis in HeLa cells.
To rule out the possibility that the observed phenotype was a byproduct of a general defect in protein import, we took advantage of a fluorescent import construct, a nuclear protein that carries a nuclear localization signal fused to GFP (GFP-NP–NLS). This construct is imported into the nucleus, whereas a mutant version of the same protein where the NLS signal has been mutated, the GFP-NP-ΔNLS construct, is not. HeLa cells, 24 h after treatment with Seh1 RNAi, were transfected with either of the two constructs. Seventy-two hours after RNAi treatment, GFP-NP–NLS protein accumulated in the nucleus of Seh1-depleted cells, whereas GFP-NP-ΔNLS remained cytoplasmic (Supplemental Figure S2, A and B). Our data suggest that although Seh1 is a component of the Nup107 complex, its depletion does not cause a major defect in nuclear import but plays a critical role in chromosome alignment and segregation.
The chromosome congression defects observed in Nup107- and Seh1-depleted cells might be caused by a decreased number of kMT associations (Zuccolo et al., 2007), or defects in targeting chromosome alignment proteins like CENP-E to kinetochores or the mislocalization of chromosome passenger proteins. We therefore investigated the effects of Seh1 depletion on the localization of individual centromere and kinetochore associated proteins by immunofluorescence and EM. We analyzed control and Seh1-depleted cells by EM after 60 h of RNAi treatment (n = 6). Similar to control cells, kinetochores of Seh1-depleted cells showed clear association with MTs. (Figure 2A, arrowheads). At the same time, in all images of Seh1-depleted cells and control cells, we observed no fibrous corona extending from the distal side of the kinetochore outer plate, which suggests the formation of attachments to MTs (Cassimeris et al., 1990). Quantification of the number of MTs associated per kinetochore/EM section resulted in the same mean number of 2–3 MTs per kinetochore/EM section in both control and Seh1-depleted cells (Figure 2B).
Figure 2.
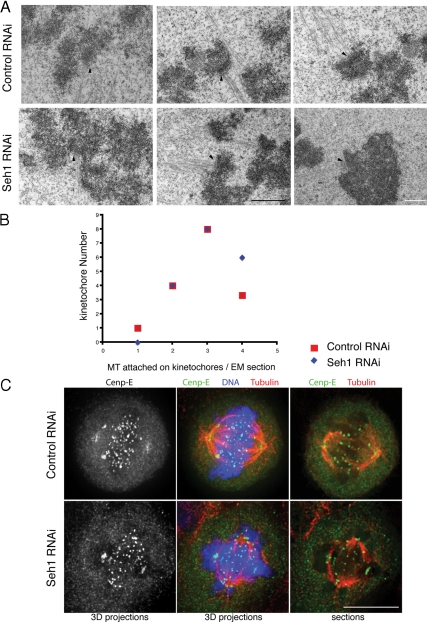
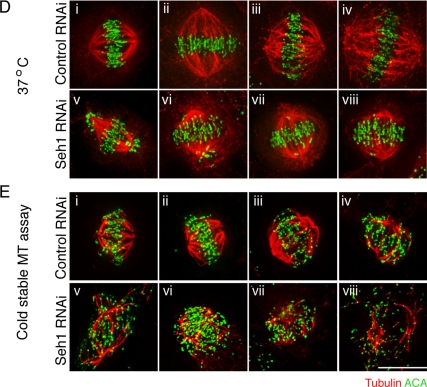
kMT attachments are maintained in Seh1-depleted cells. (A) Selected 60-nm EM sections from control or Seh1 siRNA-treated cells 65 h after transfection. Note the prominent kinetochore fibers (black arrowheads) that terminated within the trilaminar plate both in control and Seh1-depleted cells. Black scale bar, 1 μm; white scale bar, 500 nm. (B) Quantification of number of kMT fibers attached per kinetochore in EM sections of control (red squares) and Seh1-depleted (blue square) cells. Eighteen kinetochores were counted from six different cells in three different experiments per condition. (C) CENP-E localization is unaffected upon Seh1 depletion. Immunofluorescence images of control and Seh1-depleted cells are shown as maximum intensity projections or individual 3D sections. Microtubules are shown in red, chromosomes in blue, and CENP-E in green. Scale bar, 10 μm. (D and E) Immunofluorescence analysis of control (i–iv) and Seh1-depleted cells (v–viii) after incubation at 37°C (D), or cold stable microtubule assay at 4°C (E), stained with α-tubulin (red) and α-ACA (green). Scale bar, 10 μm.
The presence of stable kMT interaction in Seh1-depleted cells was also confirmed by the presence of cold stable MTs and inter-KT distances similar to control cells (Figures 2, D and E, and Supplemental Figure S3A). An anticipated decrease in the density of spindle MTs was observed upon Seh1 depletion, as the Nup107 complex has been described to affect spindle stability in the Xenopus system (Orjalo et al., 2006), but kinetochores in Seh1-depleted cells were attached to MTs after cold treatment, as in control cells (Figure 2E). Inter-KT distances on control cells had an average of 1.55 ± 0.2 μm on aligned chromosomes and 0.64 ± 0.1 μm on unaligned chromosomes, as expected for bioriented and unattached chromosomes, respectively (Supplemental Figure S3A). On Seh1 depletion the inter-KT distances of aligned chromosomes only slightly decreased to 1.45 ± 0.2 μm, suggesting normal MT attachment but with the generation of slightly less pulling forces, while unattached chromosomes had inter-KT distances (0.68 ± 0.1 μm) and remained closely associated with the spindle. Mitotic spindle length in Seh1-depleted cells showed a small but statistically significant reduction (Supplemental Figure S3B). These results suggest that in Seh1-depleted cells, bioriented kinetochores are attached to MTs and can generate significant tension across sister kinetochore pairs and that these cells might have a significant defect in chromosome alignment or biorientation.
Because CENP-E is required for proper chromosome alignment, we next examined whether the misalignment phenotype was caused by defects in CENP-E targeting (Schaar et al., 1997; McEwen et al., 2001; Weaver et al., 2003). In both control and Seh1-depleted cells the kinetochore localization of CENP-E was similar (Figure 2C). This suggests that the chromosome congression defect observed upon Seh1 depletion is not due to inability of misaligned or monooriented chromosomes to migrate toward the metaphase plate using CENP-E.
Seh1 Regulates Localization of the CPC Complex and Is Required for Correct Chromosome Alignment and Segregation
The phenotype of Seh1-depleted cells was strikingly similar to the phenotypes observed in cells after inhibition or depletion of members of the CPC (Giet and Glover, 2001; Carvalho et al., 2003; Ditchfield et al., 2003; Hauf et al., 2003; Honda et al., 2003; Lens et al., 2003). In control cells, Aurora B was detected as bright focused centromeric dots (Figure 3A), whereas in Seh1-depleted cells the centromeric Aurora B signal was dramatically reduced. Interestingly, Aurora B signal was retained on the centromeres of the misaligned chromosomes (Figure 3A). The same phenotype was observed upon depletion of another member of Nup107 complex, Mel28 (Figure 3A).
Figure 3.
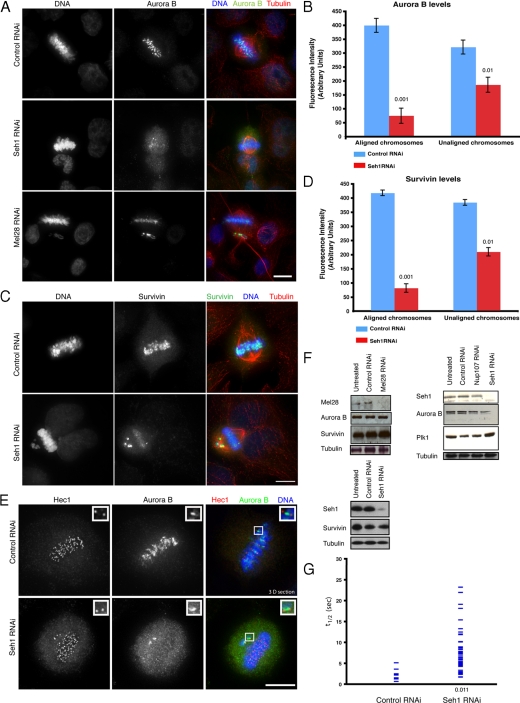
Seh1 regulates the localization of Aurora B and the chromosome passenger proteins on mitotic chromosomes and centromeres. (A) Aurora B is delocalized in Seh1- and Mel28-depleted cells. Control, Mel28, and Seh1-depleted cells were fixed and immunostained with α-Aurora B (green), α-tubulin (red), and DNA (blue). Note the staining of Aurora B on misaligned chromosomes. Scale bar, 10 μm. (B) Quantification of Aurora B levels from control (blue bars) and Seh1-depleted cells (red bars) on aligned and unaligned chromosomes. Quantification was performed on 3D data sets. Significance: p values from Student's t test are shown; n = 40 from two different experiments. (C) Survivin is delocalized in Seh1-depleted cells. Control and Seh1-depleted cells were immunostained with α-survivin (green), α-tubulin (red), and DNA (blue). (D) Quantification of survivin levels from control (blue bars) and Seh1-depleted cells (red bars) on aligned and unaligned chromosomes. Quantification was performed on 3D data sets. Significance: p values from Student's t test are shown; n = 32 from two different experiments. (E) Hec1 localization is not affected upon Seh1 depletion. Control and Seh1-depleted cells were immunostained with α-Aurora B (green) and α-Hec1 (red) to visualize kinetochores and DNA (blue). Scale bar, 10 μm. (F) Immunoblots of HeLa cell lysates treated with siRNAs corresponding to negative control, Seh1, Nup107, and Mel28 (using α-Seh1, α-Mel28 antibodies), shows efficient depletion of Seh1 and Mel28 but no down-regulation of Aurora B, survivin, or Plk1 (using α-Aurora B, α-survivin, α-Plk1). (G) FRAP analysis of centromeric association of Aurora B. Scatter plot of t1/2 (horizontal lines) for Aurora B in control and Seh1-depleted cells on unaligned centromeres. Aurora B recovery times on centromeres of Seh1-depleted cells are significantly longer than on centromeres of control cells.
In Xenopus cell-free extracts, inhibition of Aurora B kinase activity causes loss of Ndc80/Hec1 from kinetochores (Emanuele et al., 2008). However, upon depletion of Seh1 both aligned and unaligned chromosomes had strong Hec1 signals marking the kinetochores (Figure 3E), indicating that this critical kinetochore component is still targeted properly after Seh1 depletion despite the loss of Aurora B targeting.
We examined if other nonenzymatic members of the CPC showed the same phenotype upon Seh1 depletion. Survivin was also delocalized from the centromeres of aligned chromosomes but was retained on the centromeres of misaligned chromosomes upon Seh1 depletion (Figure 3C). The same was observed for borealin and INCENP (Supplemental Figure S3C). Quantification of Aurora B and survivin signal both on aligned and misaligned chromosomes between control and Seh1-depleted cells confirmed the immunofluorescence data (Figure 3, B and D). Aurora B and survivin levels are reduced on aligned chromosomes while being mostly retained on unaligned chromosomes. Although decreased expression of Seh1, Nup107 and Mel28 affected the localization of Aurora B and survivin, we observed no decrease in the total protein level of either by Western blotting (Figure 3F).
We wanted to validate the Aurora B result in live cells and confirm that the loss of Aurora B was due to change in its association with its targets. For this reason we used FRAP to measure the binding of Aurora B to centromeres of misaligned chromosomes, taking advantage of a HeLaEGFP-Aurora B cell line (Andrews, 2003). In both control and Seh1-depleted cells Aurora B fluorescence recovered rapidly, but the recovery time for Seh1-depleted cells was significantly increased, suggesting a change in the affinity of Aurora B for the centromere after Seh1 depletion (Figure 3G).
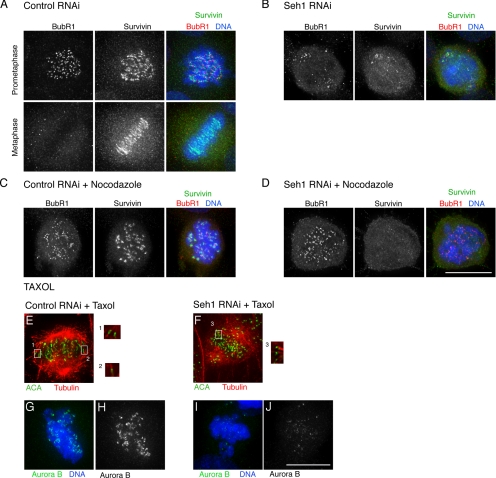
The observation that the CPC is selectively lost on aligned centromeres, but not on misaligned ones prompted us to investigate the connection to MT attachment and spindle checkpoint activation. We performed a 12-h nocodazole treatment in control and Seh1-depleted cells (Figure 4, A–D). At 64 h of treatment (52 h RNAi + nocodazole) control cells were arrested with condensed chromosomes, and the CPC components Aurora B and survivin were loaded on centromeres (Figure 4, A, Prometaphase, and C). BubR1 staining of kinetochores is reduced upon checkpoint silencing (Figure 4A, Metaphase). Seh1-depleted cells were also arrested, but no loading of CPC components at centromeres was observed (Figure 4, B and D). In both control and Seh1-depleted cells treated with nocodazole, the mitotic checkpoint was activated as judged by BubR1 staining of kinetochores (Figure 4, C and D).
Figure 4.
Nocodazole and taxol treatment do not suppress the Seh1-depletion phenotype. Control and Seh1-depleted cells, without nocodazole treatment (A and B) or with 12-h nocodazole treatment (C and D) were stained for survivin (green), BubR1 (red), and DNA (blue). Control (E) and Seh1-depleted cells (F) were treated with taxol and stained for tubulin (red) and ACA (green). Insets (optical sections), kinetochore-associated microtubules. Control (G and H) and Seh1-depleted cells (I and J) were treated with taxol and stained for Aurora B (green) and DNA (blue). Scale bar, 10 μm.
We then investigated the effect of CPC loading on centromeres after taxol treatment. Although, we did observe stabilization of MTs, both in control and Seh1-depleted cells (Figure 4, E–J), we observed no recruitment of either Aurora B or survivin on centromeres of aligned chromosomes. Restoration of CPC localization was also not observed upon checkpoint inactivation by codepletion of BubR1 and Seh1 (data not shown).
Interestingly, we observed no changes in the localization or expression of Plk1, another kinase that plays an important role in mitotic progression and cytokinesis (Figure 3F and Supplemental Figure S4). Thus Seh1 and the Nup107 complex play a specific role in the centromeric localization not only of Aurora B but also of the whole CPC complex, and this is not coupled to MT attachment.
Previously it was shown that the presence of Nup107 complex at the kinetochore is required for the recruitment of a fraction of CRM1 to the outer kinetochore structure (Zuccolo et al., 2007). CRM1 has been suggested to be required for proper localization of survivin at the centromere (Knauer et al., 2006). To assess the role of CRM1 in Aurora B localization, we treated cells with leptomycin B (a CRM1-specific drug that inhibits formation of the ternary complex among CRM1, RanGTP, and NES proteins; Fornerod et al., 1997) for various lengths of time (Supplemental Figure S5). We observed no changes in the localization of Aurora B (Supplemental Figure S5) or survivin (data not shown) at centromeres in early prophase, prometaphase, or metaphase.
It is also known that once at the kinetochore CRM1 is required to recruit the RanBP2- RanGAP-SUMO complex (Arnaoutov et al., 2005). Absence of the RanBP2-RanGAP-SUMO complex leads to defective kinetochore fibers that are sufficient to release some checkpoint proteins but are insufficient for K-fiber stability or full silencing of the spindle assembly checkpoint (Joseph et al., 2004; Arnaoutov and Dasso, 2005). We observed a small reduction of CRM1 and RanBP2-RanGAP recruitment at kinetochores upon Seh1 depletion (Supplemental Figures S6 and S7). However, although we reproducibly observed Aurora B and the CPC concentrated at all unaligned chromosomes in Seh1-depleted cells, the localization of CRM1 and RanBP2-RanGAP was more variable (Supplemental Figures S6 and S7). We propose that although the RANBP2-RanGAP-SUMO complex may play a role in K-fiber stability the chromosome segregation errors observed upon depletion of Seh1 are more likely explained by the loss of centromeric localization of the CPC, and especially Aurora B, and not by a failure to localize CRM1 and RanBP2-RanGAP.
Seh1 Is Required for Proper Aurora B Substrate Phosphorylation
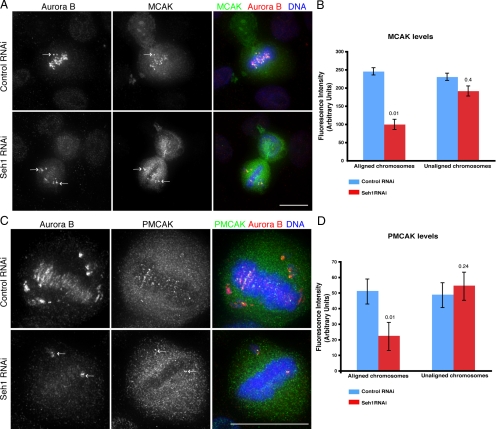
We next examined the phosphorylation of known Aurora B substrates that play a role in mitosis and cytokinesis. Phosphorylation of the mitotic centromere-associated kinesin (MCAK) by Aurora B inhibits MCAK's MT depolymerization activity and is required for the localization of MCAK at the inner centromere and MT ends (Andrews et al., 2004; Kline-Smith et al., 2004; Lan et al., 2004; Knowlton et al., 2006). Immunofluorescence against MCAK and phosphorylated MCAK with an antibody that specifically recognizes Aurora B–phosphorylated MCAK at S92 (P-MCAK) showed that although MCAK was localized both at centromeres and spindle poles in control cells, in Seh1-depleted cells MCAK signal was lost from metaphase aligned chromosomes and was retained only on the centromeres of the misaligned chromosomes (Figure 5A, white arrows). At the same time P-MCAK accumulation was observed only on misaligned centromeres in Seh1-depleted cells (Figure 5C, white arrow). Quantification of the fluorescence signal of MCAK and PMCAK on both aligned and unaligned chromosomes in control and Seh1-depleted cells confirmed the phenotypes observed by immunofluorescence (Figure 5, B and D). Phosphorylated Histone H3 staining verified that the ability of Aurora B to phosphorylate at least this substrate was not compromised (Supplemental Figure S2C). Thus, Seh1 depletion results in mislocalization of Aurora B, which in turn regulates phosphorylation and localization of MCAK on the centromeres. In this way Aurora B on misaligned chromosomes on Seh1-depleted cells affects phosphorylation of MCAK and may prevent MCAK performing its function of correcting kinetochore MT misattachments (Kline-Smith et al., 2004; Lan et al., 2004).
Figure 5.
MCAK localization and phosphorylation is affected in Seh1-depleted cells. Control and Seh1-depleted cells immunostained with α-Aurora B (red) and α-MCAK (green) for total MCAK population and DNA (blue) are shown in A. Cells stained for phospho-Ser92-MCAK levels shown in C. Aurora B phosphorylation of MCAK is seen on misaligned chromosomes after Seh1 depletion. Arrows, misaligned chromosomes both in control and Seh1-depleted cells. Quantification of MCAK (B) and P-MCAK (D) levels from control (blue bars) and Seh1-depleted cells (red bars) on aligned and unaligned chromosomes. Quantification was performed on 3D data sets. Significance: p values from Student's t test are shown; n = 20 from two different experiments.
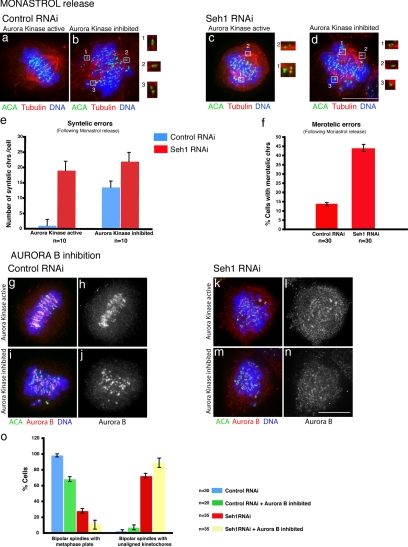
To test if Seh1 and the Nup107 complex affect CPC function before anaphase onset and to investigate the ability of Seh1-depleted cells to correct faulty kMT attachments, a process that requires Aurora B at centromeres, we performed a monastrol washout experiment. Monastrol treatment causes monopolar spindles with chromosomes that have syntelic or monotelic attachments, but upon washout Aurora B activity enables correction of syntelic attachments and formation of bipolar spindle with fully aligned chromosomes (Khodjakov et al., 2003; Lampson and Kapoor, 2005). Monastrol treatment for 2 h of control and Seh1-depleted cells was followed by release in either normal medium or medium containing Aurora B inhibitor (Figure 6, a–d). Ninety minutes after monastrol release, control cells were able to form a bipolar spindle with chromosomes correctly aligned in the middle when Aurora B kinase is active (Figure 6a) but retained multiple syntelic chromosomes upon Aurora B inhibition (Figure 6b). The average number of syntelic kinetochores per cell increased from 2 to 19 in control cells upon Aurora B inhibition (Figure 6e). Seh1-depleted cells were also able to form a bipolar spindle, but a large number of syntelic chromosomes remained near the poles even when Aurora B Kinase was active (Figure 6c). Once Aurora B kinase was inhibited in Seh1-depleted cells the average number of syntelic kinetochores per cell increased from 13.5 to 22 (Figure 6, d and e). The above results suggest that the attachment error correction by Aurora B is compromised upon Seh1 depletion. At the same time, the increased number of merotelic chromosomes in Seh1-depleted cells (Figure 6f) is in agreement with the loss of Aurora B activity as chromosomes congress.
Figure 6.
Seh1-depleted cells are able to achieve bipolarization after monastrol release. Control (a and b) and Seh1-depleted cells (c and d) after removal of monastrol were incubated for 1 h with (b and d) or without ZM447439 (a and c) and then processed for immunofluorescence. Tubulin is shown in red, ACA in green, and DNA in blue. Insets (optical sections), individual kinetochores with syntelic attachments. Quantification of syntelic errors after monastrol release is shown in panel e; quantification of cells with merotelic chromosomes after monastrol release in control and Seh1-depleted cells is shown in panel f. Inhibition of Aurora B does not suppress the Seh1 depletion phenotype. Control (g–j) and Seh1-depleted cells (k–n) were incubated with (i, j, m, and n) or without ZM447439 (g, h, k, and l) and stained for Aurora B (red), ACA (green), and DNA (blue). The percentage of control and Seh1-depleted cells containing a bipolar spindle with a recognizable metaphase plate in the presence or absence of ZM447439 was calculated (o). Cells were stained for tubulin, ACA, and DNA. Scale bar, 10 μm.
Finally, to test if potential hyper-activation of Aurora B activity on misaligned chromosomes was responsible for the phenotypes observed, we carried out an Aurora B inhibition experiment. Control and Seh1-depleted cells were incubated with either ZM447439 or hesperadin (Lampson and Kapoor, 2005) for 1 h before analysis. We observed no suppression of the Seh1 depletion chromosome misalignment phenotype nor restoration of centromeric chromatin CPC loading on Seh1-depleted cells (Figure 6, g–n). At the same time, quantification of the number of cells containing bipolar spindles with a clear metaphase plate and the number of bipolar spindles with unaligned kinetochores in control, Seh1-depleted cells and Seh1-depleted cells in the presence of the Aurora B inhibitor ZM447439 (Figure 6o) showed an increase in the number of cells with unaligned chromosomes, suggesting that the phenotypes observed were not due to hyperactivation of Aurora B kinase.
Organization of the Spindle Midzone Region Depends on Seh1
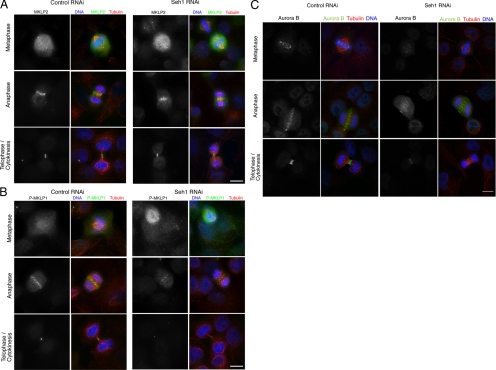
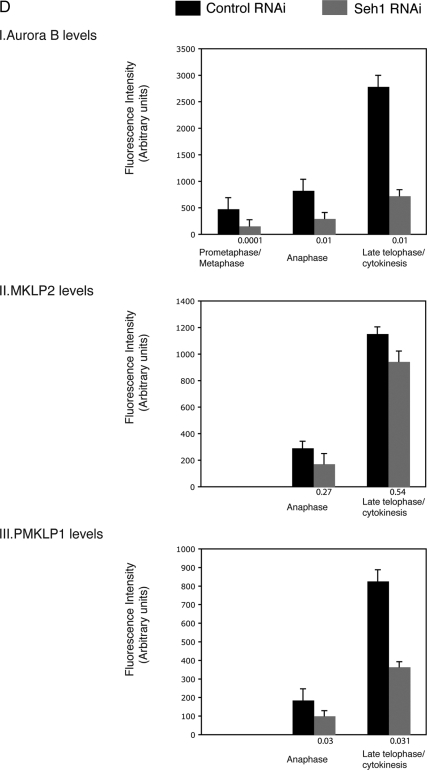
Having shown that the Nup107 complex is responsible for the centromeric localization of CPC members, the well-established cytokinesis defect might also be due to mislocalization of the CPC on the spindle midzone. Hence to understand the mechanism of cytokinesis failure in Seh1-depleted cells we examined the organization of the central spindle and midbody components in anaphase and telophase cells. MKLP2 is a mitotic kinesin-like protein responsible for the spatial restriction of Aurora B to the central spindle (Gruneberg et al., 2004; Neef et al., 2005). MKLP1 is a kinesin-6 family motor protein that becomes activated in anaphase by Aurora B phosphorylation, when cleavage furrow ingression occurs, and is essential for cytokinesis (Neef et al., 2006). Immunofluorescence experiments in Seh1-depleted cells showed no difference relative to control cells in the localization and intensity levels of MKLP2 at the spindle midzone or midbody (Figure 7A). Interestingly P-MKLP1 (detected with a phospho specific antibody that recognizes MKLP1 phosphorylated specifically by Aurora B; Neef et al., 2006) failed to localize properly and the intensity levels of P-MKLP1 at both midzone and midbody were dramatically reduced (Figure 7B). Localization and intensity levels of MKLP1 remained unchanged (data not shown). Moreover Seh1-depleted cells failed to assemble an organized midzone and midbody as detected by the loss of localized Aurora B signal during anaphase/telophase (Figure 7C). Quantification of the fluorescence signal of Aurora B, MKLP2 and P-MKLP1 from 3D deconvolved images confirmed the phenotypes observed by immunofluorescence (Figure 7D). We also examined the localization of Kif14, a kinesin required for the later stages of cytokinesis, and PRC1, an MT-bundling protein that is critical to the formation of the central spindle, but we observed no changes in their localization patterns in Seh1 RNAi-treated cells (data not shown). The above data suggest that Seh1 either directly or indirectly regulates both the chromosome segregation function and the cytokinesis function of Aurora B and the chromosome passenger proteins.
Figure 7.
The organization of the anaphase spindle midzone and midbody depends on Seh1. Localization of MKLP2 (A) and P-MKLP1 (B) were analyzed by immunofluorescence on control and Seh1-depleted cells. MKLP2 signal was visualized with α-MKLP2 antibody (green, in A), P-MKLP1 with α-PMKLP1 antibody (green, in B), microtubules with α-tubulin (red) and DNA (blue). (C) Control and Seh1-depleted cells at different stages of mitosis were immunostained with α-Aurora B (green), α-tubulin (red), and DNA (blue). (D) Quantification of Aurora B (I), MKLP2 (II), and P-MKLP1 (III) levels from control ■) and Seh1-depleted cells ( ). Significance: p values from Student's t test are shown; n = 40 from two different experiments. Note the reduction in metaphase, anaphase, and midbody of Aurora B signal and corresponding reduction P-MKLP1. Scale bar, 10 μm.
). Significance: p values from Student's t test are shown; n = 40 from two different experiments. Note the reduction in metaphase, anaphase, and midbody of Aurora B signal and corresponding reduction P-MKLP1. Scale bar, 10 μm.
DISCUSSION
In this study we have identified a link between the Nup107 nucleoporin complex and the function of the CPC. Building on previous work showing that depletion of the Nup107 complex is required for correct chromosome congression and timely progression through mitosis (Zuccolo et al., 2007), we have demonstrated that the complex, and specifically Seh1, is essential for chromosome alignment, segregation, and completion of cytokinesis in human cells and plays a critical role in the association of the CPC with the inner centromere and the central spindle. Although we observed significant numbers of misaligned chromosomes in Seh1-depleted cells, analysis of kMT interaction at the ultrastructural level showed no direct attachment defect. As discussed bellow, chromosome misalignment maybe related to a critical defect in the targeting of Aurora B and the CPC.
Seh1 Down-Regulation Results in Mislocalization of the CPC Proteins and Specifically Aurora B
Depletion of Seh1 from mammalian cells and the subsequent loss of the Nup107 complex from the kinetochore results in multiple mitotic defects, including chromosome–MT attachment errors, chromosome misalignments, spindle midzone defects, and cytokinesis failure (Zuccolo et al., 2007 and this work). All of the above events are regulated by the CPC and Aurora B protein kinase (Vagnarelli and Earnshaw, 2004). Early in mitosis, during prometaphase the CPC concentrates at centromeres when chromosomes start attaching to MTs; upon anaphase onset the CPC complex then moves to the central spindle before accumulating at the equatorial cortex just before cleavage furrow assembly. We observed severe defects in the localization of Aurora B and survivin throughout the cell cycle upon Seh1 depletion.
The finding that Seh1 depletion does not affect either the ability of kinetochores to capture MTs, either at the metaphase plate or on unattached polar chromosomes, or the presence of the CENP-E motor protein at kinetochores excludes the possibility that the chromosome congression defect observed is due to loss of MT–kinetochore attachments. We observed no major differences in the overall spindle morphology but we did observe a reduction in the density of cold stable kMTs. We do not know if Seh1 has another yet unidentified function in spindle assembly but the Nup107 complex in Xenopus system has been described to affect spindle stability. Microtubule nucleation around centrosomes was normal, but MTs quickly disassembled, reminiscent of CPC depletion in the Xenopus system (Orjalo et al., 2006).
Survivin's localization to centromeres has been suggested to be controlled by CRM1 (Knauer et al., 2006). It is unknown whether the localization of the entire CPC to the centromere is controlled by Ran via CRM1-survivin interaction in mammalian cells or whether additional mechanisms also exist, but CPC loading on chromatin in Xenopus laevis egg extracts occurs independently of Ran (Kelly et al., 2007). On Seh1 depletion we observed no defect in the early prophase targeting of survivin or the rest of the CPC (data not shown). This could also clearly be concluded from the strong survivin and Aurora B staining on misaligned metaphase chromosomes, suggesting that the defects observed after Seh1 depletion involve the stable binding of the CPC to the centromere and not its initial recruitment.
The kinesin-13 family MT depolymerase MCAK associates with centromeres from prophase to anaphase and is required to release kMT misattachments. MCAK localization at the centromere and its depolymerase activity are regulated by Aurora B. More precisely, phosphorylation of MCAK by Aurora B inhibits MCAK's ability to depolymerize MTs at the kinetochore (Andrews et al., 2004; Kline-Smith et al., 2004; Lan et al., 2004; Knowlton et al., 2006). In Seh1-depleted cells we observed an excess of Aurora B and P-MCAK on misaligned chromosomes but neither Aurora B nor MCAK were retained on the centromeres of metaphase plate chromosomes. This suggests that the error-correction system that eliminates kMT misattachments, so that correct end-on attachments can be made, is compromised. This would explain the misaligned chromosome phenotype observed upon depletion of Seh1.
During metaphase to anaphase transition relocalization of the CPC from the centromere to the central spindle requires dynamic MTs, protein phosphatase cdc14, and MKLP2, a mitotic kinesin that binds and relocates both Aurora B and cdc14 (Gruneberg et al., 2004; Vader et al., 2006). Human cells depleted of MKLP2 or treated with MT-stabilizing drugs such as taxol have impaired CPC localization (Wheatley et al., 2001; Gruneberg et al., 2004). On Seh1 depletion MKLP2 localization is largely unaffected suggesting that known “upstream” factor(s) required for the relocation of the CPC are functional and present but that the CPC complex cannot respond and bind to central spindle MTs. This leads to loss of Aurora B and its enzymatic activity, consistent with the observed reduction of phosphorylation of one of its central spindle substrates MKLP1 (Figure 7, B and D).
Proper targeting of MKLP1 to the central spindle requires PRC1, the MT-bundling protein (Kurasawa et al., 2004). We observed no changes in PRC1 localization upon Seh1 depletion (data not shown), despite observing delocalization of P-MKLP1. Phosphorylation of the nuclear localization signal sequence (NLS) of MKLP1 is required to allow P-MKLP1 to remain at the midzone and midbody in order to promote efficient completion of cytokinesis (Guse et al., 2005; Neef et al., 2006). This suggests that the Nup107 complex functions downstream of PRC1 to mediate the phosphorylation and localization of MKLP-1.
Although it is possible that the binucleation observed in Seh1-depleted cells may result from lagging chromosomes blocking the cytokinesis furrow, the loss of CPC complex from both central spindle in anaphase and midbody in telophase together with the loss of phosphorylated MKLP1 point to the idea that Seh1 and the Nup107 complex are required, either directly or indirectly, for both the centromeric and cytokinesis function of the CPC.
We believe that although a threshold activity of Aurora B kinase still remains at the midzone it is not sufficient to promote efficient cell division. Perhaps cytokinesis can proceed with low level activity of Aurora B, whereas for error correction that activity needs to be enhanced.
Seh1 Is Required for the Maintenance of Aurora B and Survivin on Centromeric Chromatin
FRAP analysis of Aurora B and survivin has shown that both proteins are highly dynamic while at centromeres but less so once associated with the central spindle (Murata-Hori and Wang, 2002; Beardmore et al., 2004; Wheatley et al., 2007). Our FRAP analysis of Aurora B upon Seh1 depletion suggests that Seh1 affects the dynamic association of Aurora B with centromeres and hence its subsequent association with the central spindle and midbody. The data show an increase in the half-life of the kinase at the centromere of unaligned chromosomes, confirming the change in the affinity of the CPC for the centromere/kinetochore as chromosomes align. The dynamic exchange of CPC members at the centromere provides a mechanism that regulates the interaction of the kinase with its substrates.
Binding of CPC to unaligned chromosomes is perturbed in Seh1-depleted cells treated with taxol or nocodazole (Figure 4). In taxol-treated cells, CPC is still observed at centromeres, although at a reduced level, whereas nocodazole treatment completely removes CPC. This suggests that CPC targeting is sensitive to tension or MT dynamics but requires MT attachment. The partial loss of CPC in taxol suggests either separate populations of the CPC complex that may be differentially regulated, or still uncharacterized signaling and modification events may affect targeting and activation of the CPC. Alternatively other adapter proteins might be required to retain CPC and are lost upon alignment in Seh1-depleted cells.
To maintain a stable CPC structure all of the CPC component proteins (survivin, Aurora B, INCENP, and borealin) are needed, but survivin plays a special role in mediating the localization of the CPC to its proper mitotic locations. Different domains within survivin dictate either the centromere targeting or the central spindle targeting and although the centromeric localization of the CPC is not a prerequisite for central spindle localization the two events are linked (Lens et al., 2006; Jeyaprakash et al., 2007; Yue et al., 2008). It appears that the survivin-centromere interaction is regulated by phosphorylation and ubiquitination (Vong et al., 2005; Wheatley et al., 2007).
Finally recent work has highlighted the importance of modification by SUMO or ubiquitin of key kinetochore and centromere associated proteins (Ramadan et al., 2007; Sumara et al., 2007; Dawlaty et al., 2008; Zhang et al., 2008). These modifications regulate the localization of important mitotic regulators in space and time, allowing proper completion of mitosis and reformation of the nuclear envelope. Our data suggest that nucleoporins and their interacting partners function in these pathways to mediate proper function of kinetochores and the cytokinetic furrow.
Supplementary Material
ACKNOWLEDGMENTS
We thank Francis Barr (University of Liverpool, Liverpool, United Kingdom) for MKLP1, P-MKLP1, and MKLP2 antibodies; Ulrike Kutay (ETH, Zürich, Switzerland) for Nup107 antibody; Peter Sorger (Harvard Medical School, Boston, MA) for CENP-E antibody; Jan Ellenberg (EMBL, Heidelberg, Germany) for EGFP-Seh1, mRFP-H2B plasmids, and HeLaGFPH2B cell line; and Frauke Melchior (ZMBH, Heidelberg, Germany) for RanBP2 antibody. We are especially grateful to members of the Advanced Light Microscopy Facility of EMBL, members of the Mattaj lab for valuable discussions, and Sevil Yavuz for wild-type and mutant GFP-NP–NLS constructs. M.P. was supported by Human Frontier Science Programme Long-Term fellowship LT00519/2003 and Wellcome Trust Project Grant (Ref R41067). C.R.W. was supported by an EMBL Predoctoral Fellowship, and J.R.S. was supported by a Wellcome Trust Senior Research Fellowship (Ref 067433).
Footnotes
This article was published online ahead of print in MBC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E09-05-0377) on October 28, 2009.
REFERENCES
- Allen T. D., Cronshaw J. M., Bagley S., Kiseleva E., Goldberg M. W. The nuclear pore complex: mediator of translocation between nucleus and cytoplasm. J. Cell Sci. 2000;113(Pt 10):1651–1659. doi: 10.1242/jcs.113.10.1651. [DOI] [PubMed] [Google Scholar]
- Andrews P. D. Cell of the month: two telophase HeLa cells expressing GFP-tagged human Aurora B. Nat. Rev. Mol. Cell. Biol. 2003;4:92–93. [Google Scholar]
- Andrews P. D., Ovechkina Y., Morrice N., Wagenbach M., Duncan K., Wordeman L., Swedlow J. R. Aurora B regulates MCAK at the mitotic centromere. Dev. Cell. 2004;6:253–268. doi: 10.1016/s1534-5807(04)00025-5. [DOI] [PubMed] [Google Scholar]
- Antonin W., Ellenberg J., Dultz E. Nuclear pore complex assembly through the cell cycle: regulation and membrane organization. FEBS Lett. 2008;582:2004–2016. doi: 10.1016/j.febslet.2008.02.067. [DOI] [PubMed] [Google Scholar]
- Arnaoutov A., Azuma Y., Ribbeck K., Joseph J., Boyarchuk Y., Karpova T., McNally J., Dasso M. Crm1 is a mitotic effector of Ran-GTP in somatic cells. Nat. Cell Biol. 2005;7:626–632. doi: 10.1038/ncb1263. [DOI] [PubMed] [Google Scholar]
- Arnaoutov A., Dasso M. Ran-GTP regulates kinetochore attachment in somatic cells. Cell Cycle. 2005;4:1161–1165. doi: 10.4161/cc.4.9.1979. [DOI] [PubMed] [Google Scholar]
- Axelrod D., Koppel D. E., Schlessinger J., Elson E., Webb W. W. Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys. J. 1976;16:1055–1069. doi: 10.1016/S0006-3495(76)85755-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beardmore V. A., Ahonen L. J., Gorbsky G. J., Kallio M. J. Survivin dynamics increases at centromeres during G2/M phase transition and is regulated by microtubule-attachment and Aurora B kinase activity. J. Cell Sci. 2004;117:4033–4042. doi: 10.1242/jcs.01242. [DOI] [PubMed] [Google Scholar]
- Belgareh N., et al. An evolutionarily conserved NPC subcomplex, which redistributes in part to kinetochores in mammalian cells. J. Cell Biol. 2001;154:1147–1160. doi: 10.1083/jcb.200101081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blower M. D., Nachury M., Heald R., Weis K. A Rae1-containing ribonucleoprotein complex is required for mitotic spindle assembly. Cell. 2005;121:223–234. doi: 10.1016/j.cell.2005.02.016. [DOI] [PubMed] [Google Scholar]
- Boehmer T., Enninga J., Dales S., Blobel G., Zhong H. Depletion of a single nucleoporin, Nup107, prevents the assembly of a subset of nucleoporins into the nuclear pore complex. Proc. Natl. Acad. Sci. USA. 2003;100:981–985. doi: 10.1073/pnas.252749899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carvalho A., Carmena M., Sambade C., Earnshaw W. C., Wheatley S. P. Survivin is required for stable checkpoint activation in taxol-treated HeLa cells. J. Cell Sci. 2003;116:2987–2998. doi: 10.1242/jcs.00612. [DOI] [PubMed] [Google Scholar]
- Cassimeris L., Rieder C. L., Rupp G., Salmon E. D. Stability of microtubule attachment to metaphase kinetochores in PtK1 cells. J. Cell Sci. 1990;96(Pt 1):9–15. doi: 10.1242/jcs.96.1.9. [DOI] [PubMed] [Google Scholar]
- Cheeseman I. M., Chappie J. S., Wilson-Kubalek E. M., Desai A. The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell. 2006;127:983–997. doi: 10.1016/j.cell.2006.09.039. [DOI] [PubMed] [Google Scholar]
- Cheeseman I. M., Desai A. Molecular architecture of the kinetochore-microtubule interface. Nat. Rev. 2008;9:33–46. doi: 10.1038/nrm2310. [DOI] [PubMed] [Google Scholar]
- Conti E., Izaurralde E. Nucleocytoplasmic transport enters the atomic age. Curr. Opin. Cell Biol. 2001;13:310–319. doi: 10.1016/s0955-0674(00)00213-1. [DOI] [PubMed] [Google Scholar]
- Cronshaw J. M., Krutchinsky A. N., Zhang W., Chait B. T., Matunis M. J. Proteomic analysis of the mammalian nuclear pore complex. J. Cell Biol. 2002;158:915–927. doi: 10.1083/jcb.200206106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daigle N., Beaudouin J., Hartnell L., Imreh G., Hallberg E., Lippincott-Schwartz J., Ellenberg J. Nuclear pore complexes form immobile networks and have a very low turnover in live mammalian cells. J. Cell Biol. 2001;154:71–84. doi: 10.1083/jcb.200101089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dawlaty M. M., Malureanu L., Jeganathan K. B., Kao E., Sustmann C., Tahk S., Shuai K., Grosschedl R., van Deursen J. M. Resolution of sister centromeres requires RanBP2-mediated SUMOylation of topoisomerase IIalpha. Cell. 2008;133:103–115. doi: 10.1016/j.cell.2008.01.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeLuca J. G., Moree B., Hickey J. M., Kilmartin J. V., Salmon E. D. hNuf2 inhibition blocks stable kinetochore-microtubule attachment and induces mitotic cell death in HeLa cells. J. Cell Biol. 2002;159:549–555. doi: 10.1083/jcb.200208159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditchfield C., Johnson V. L., Tighe A., Ellston R., Haworth C., Johnson T., Mortlock A., Keen N., Taylor S. S. Aurora B couples chromosome alignment with anaphase by targeting BubR1, Mad2, and Cenp-E to kinetochores. J. Cell Biol. 2003;161:267–280. doi: 10.1083/jcb.200208091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emanuele M. J., Lan W., Jwa M., Miller S. A., Chan C. S., Stukenberg P. T. Aurora B kinase and protein phosphatase 1 have opposing roles in modulating kinetochore assembly. J. Cell Biol. 2008;181:241–254. doi: 10.1083/jcb.200710019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng J., Huang H., Yen T. J. CENP-F is a novel microtubule-binding protein that is essential for kinetochore attachments and affects the duration of the mitotic checkpoint delay. Chromosoma. 2006;115:320–329. doi: 10.1007/s00412-006-0049-5. [DOI] [PubMed] [Google Scholar]
- Fornerod M., Ohno M., Yoshida M., Mattaj I. W. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell. 1997;90:1051–1060. doi: 10.1016/s0092-8674(00)80371-2. [DOI] [PubMed] [Google Scholar]
- Franz C., Walczak R., Yavuz S., Santarella R., Gentzel M., Askjaer P., Galy V., Hetzer M., Mattaj I. W., Antonin W. MEL-28/ELYS is required for the recruitment of nucleoporins to chromatin and postmitotic nuclear pore complex assembly. EMBO Rep. 2007;8:165–172. doi: 10.1038/sj.embor.7400889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gassmann R., Carvalho A., Henzing A. J., Ruchaud S., Hudson D. F., Honda R., Nigg E. A., Gerloff D. L., Earnshaw W. C. Borealin: a novel chromosomal passenger required for stability of the bipolar mitotic spindle. J. Cell Biol. 2004;166:179–191. doi: 10.1083/jcb.200404001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giet R., Glover D. M. Drosophila aurora B kinase is required for histone H3 phosphorylation and condensin recruitment during chromosome condensation and to organize the central spindle during cytokinesis. J. Cell Biol. 2001;152:669–682. doi: 10.1083/jcb.152.4.669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruneberg U., Neef R., Honda R., Nigg E. A., Barr F. A. Relocation of Aurora B from centromeres to the central spindle at the metaphase to anaphase transition requires MKlp2. J. Cell Biol. 2004;166:167–172. doi: 10.1083/jcb.200403084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guse A., Mishima M., Glotzer M. Phosphorylation of ZEN-4/MKLP1 by aurora B regulates completion of cytokinesis. Curr. Biol. 2005;15:778–786. doi: 10.1016/j.cub.2005.03.041. [DOI] [PubMed] [Google Scholar]
- Harel A., Orjalo A. V., Vincent T., Lachish-Zalait A., Vasu S., Shah S., Zimmerman E., Elbaum M., Forbes D. J. Removal of a single pore subcomplex results in vertebrate nuclei devoid of nuclear pores. Mol. Cell. 2003;11:853–864. doi: 10.1016/s1097-2765(03)00116-3. [DOI] [PubMed] [Google Scholar]
- Hauf S., Cole R. W., LaTerra S., Zimmer C., Schnapp G., Walter R., Heckel A., van Meel J., Rieder C. L., Peters J. M. The small molecule Hesperadin reveals a role for Aurora B in correcting kinetochore-microtubule attachment and in maintaining the spindle assembly checkpoint. J. Cell Biol. 2003;161:281–294. doi: 10.1083/jcb.200208092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Honda R., Korner R., Nigg E. A. Exploring the functional interactions between Aurora B, INCENP, and survivin in mitosis. Mol. Biol. Cell. 2003;14:3325–3341. doi: 10.1091/mbc.E02-11-0769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howell B. J., McEwen B. F., Canman J. C., Hoffman D. B., Farrar E. M., Rieder C. L., Salmon E. D. Cytoplasmic dynein/dynactin drives kinetochore protein transport to the spindle poles and has a role in mitotic spindle checkpoint inactivation. J. Cell Biol. 2001;155:1159–1172. doi: 10.1083/jcb.200105093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hutten S., Flotho A., Melchior F., Kehlenbach R. H. The Nup358-RanGAP complex is required for efficient importin alpha/beta-dependent nuclear import. Mol. Biol. Cell. 2008;19:2300–2310. doi: 10.1091/mbc.E07-12-1279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeyaprakash A. A., Klein U. R., Lindner D., Ebert J., Nigg E. A., Conti E. Structure of a Survivin-Borealin-INCENP core complex reveals how chromosomal passengers travel together. Cell. 2007;131:271–285. doi: 10.1016/j.cell.2007.07.045. [DOI] [PubMed] [Google Scholar]
- Joseph J., Liu S. T., Jablonski S. A., Yen T. J., Dasso M. The RanGAP1-RanBP2 complex is essential for microtubule-kinetochore interactions in vivo. Curr. Biol. 2004;14:611–617. doi: 10.1016/j.cub.2004.03.031. [DOI] [PubMed] [Google Scholar]
- Kelly A. E., Sampath S. C., Maniar T. A., Woo E. M., Chait B. T., Funabiki H. Chromosomal enrichment and activation of the aurora B pathway are coupled to spatially regulate spindle assembly. Dev. Cell. 2007;12:31–43. doi: 10.1016/j.devcel.2006.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khodjakov A., Copenagle L., Gordon M. B., Compton D. A., Kapoor T. M. Minus-end capture of preformed kinetochore fibers contributes to spindle morphogenesis. J. Cell Biol. 2003;160:671–683. doi: 10.1083/jcb.200208143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kline-Smith S. L., Khodjakov A., Hergert P., Walczak C. E. Depletion of centromeric MCAK leads to chromosome congression and segregation defects due to improper kinetochore attachments. Mol. Biol. Cell. 2004;15:1146–1159. doi: 10.1091/mbc.E03-08-0581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knauer S. K., Bier C., Habtemichael N., Stauber R. H. The Survivin-Crm1 interaction is essential for chromosomal passenger complex localization and function. EMBO Rep. 2006;7:1259–1265. doi: 10.1038/sj.embor.7400824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knowlton A. L., Lan W., Stukenberg P. T. Aurora B is enriched at merotelic attachment sites, where it regulates MCAK. Curr. Biol. 2006;16:1705–1710. doi: 10.1016/j.cub.2006.07.057. [DOI] [PubMed] [Google Scholar]
- Kurasawa Y., Earnshaw W. C., Mochizuki Y., Dohmae N., Todokoro K. Essential roles of KIF4 and its binding partner PRC1 in organized central spindle midzone formation. EMBO J. 2004;23:3237–3248. doi: 10.1038/sj.emboj.7600347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lampson M. A., Kapoor T. M. The human mitotic checkpoint protein BubR1 regulates chromosome-spindle attachments. Nat. Cell Biol. 2005;7:93–98. doi: 10.1038/ncb1208. [DOI] [PubMed] [Google Scholar]
- Lan W., Zhang X., Kline-Smith S. L., Rosasco S. E., Barrett-Wilt G. A., Shabanowitz J., Hunt D. F., Walczak C. E., Stukenberg P. T. Aurora B phosphorylates centromeric MCAK and regulates its localization and microtubule depolymerization activity. Curr. Biol. 2004;14:273–286. doi: 10.1016/j.cub.2004.01.055. [DOI] [PubMed] [Google Scholar]
- Lens S. M., Rodriguez J. A., Vader G., Span S. W., Giaccone G., Medema R. H. Uncoupling the central spindle-associated function of the chromosomal passenger complex from its role at centromeres. Mol. Biol. Cell. 2006;17:1897–1909. doi: 10.1091/mbc.E05-08-0727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lens S. M., Wolthuis R. M., Klompmaker R., Kauw J., Agami R., Brummelkamp T., Kops G., Medema R. H. Survivin is required for a sustained spindle checkpoint arrest in response to lack of tension. EMBO J. 2003;22:2934–2947. doi: 10.1093/emboj/cdg307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maiato H., DeLuca J., Salmon E. D., Earnshaw W. C. The dynamic kinetochore-microtubule interface. J. Cell Sci. 2004;117:5461–5477. doi: 10.1242/jcs.01536. [DOI] [PubMed] [Google Scholar]
- McCleland M. L., Gardner R. D., Kallio M. J., Daum J. R., Gorbsky G. J., Burke D. J., Stukenberg P. T. The highly conserved Ndc80 complex is required for kinetochore assembly, chromosome congression, and spindle checkpoint activity. Genes Dev. 2003;17:101–114. doi: 10.1101/gad.1040903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McEwen B. F., Chan G. K., Zubrowski B., Savoian M. S., Sauer M. T., Yen T. J. CENP-E is essential for reliable bioriented spindle attachment, but chromosome alignment can be achieved via redundant mechanisms in mammalian cells. Mol. Biol. Cell. 2001;12:2776–2789. doi: 10.1091/mbc.12.9.2776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McEwen B. F., Dong Y., VandenBeldt K. J. Using electron microscopy to understand functional mechanisms of chromosome alignment on the mitotic spindle. Methods Cell Biol. 2007;79:259–293. doi: 10.1016/S0091-679X(06)79011-2. [DOI] [PubMed] [Google Scholar]
- Murata-Hori M., Wang Y. L. Both midzone and astral microtubules are involved in the delivery of cytokinesis signals: insights from the mobility of aurora B. J. Cell Biol. 2002;159:45–53. doi: 10.1083/jcb.200207014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neef R., Gruneberg U., Barr F. A. Assay and functional properties of Rabkinesin-6/Rab6-KIFL/MKlp2 in cytokinesis. Methods Enzymol. 2005;403:618–628. doi: 10.1016/S0076-6879(05)03054-5. [DOI] [PubMed] [Google Scholar]
- Neef R., Klein U. R., Kopajtich R., Barr F. A. Cooperation between mitotic kinesins controls the late stages of cytokinesis. Curr. Biol. 2006;16:301–307. doi: 10.1016/j.cub.2005.12.030. [DOI] [PubMed] [Google Scholar]
- Orjalo A. V., Arnaoutov A., Shen Z., Boyarchuk Y., Zeitlin S. G., Fontoura B., Briggs S., Dasso M., Forbes D. J. The Nup107-160 nucleoporin complex is required for correct bipolar spindle assembly. Mol. Biol. Cell. 2006;17:3806–3818. doi: 10.1091/mbc.E05-11-1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramadan K., Bruderer R., Spiga F. M., Popp O., Baur T., Gotta M., Meyer H. H. Cdc48/p97 promotes reformation of the nucleus by extracting the kinase Aurora B from chromatin. Nature. 2007;450:1258–1262. doi: 10.1038/nature06388. [DOI] [PubMed] [Google Scholar]
- Rasala B. A., Orjalo A. V., Shen Z., Briggs S., Forbes D. J. ELYS is a dual nucleoporin/kinetochore protein required for nuclear pore assembly and proper cell division. Proc. Natl. Acad. Sci. USA. 2006;103:17801–17806. doi: 10.1073/pnas.0608484103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruchaud S., Carmena M., Earnshaw W. C. Chromosomal passengers: conducting cell division. Nat. Rev. 2007;8:798–812. doi: 10.1038/nrm2257. [DOI] [PubMed] [Google Scholar]
- Schaar B. T., Chan G. K., Maddox P., Salmon E. D., Yen T. J. CENP-E function at kinetochores is essential for chromosome alignment. J. Cell Biol. 1997;139:1373–1382. doi: 10.1083/jcb.139.6.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumara I., Quadroni M., Frei C., Olma M. H., Sumara G., Ricci R., Peter M. A Cul3-based E3 ligase removes Aurora B from mitotic chromosomes, regulating mitotic progression and completion of cytokinesis in human cells. Dev. Cell. 2007;12:887–900. doi: 10.1016/j.devcel.2007.03.019. [DOI] [PubMed] [Google Scholar]
- Swedlow J. R., Sedat J. W., Agard D. A. Deconvolution in optical microscopy. In: Jansson P. A., editor. Deconvolution of Images and Spectra. New York: Academic Press; 1997. pp. 284–309. [Google Scholar]
- Tanaka T. U., Stark M. J., Tanaka K. Kinetochore capture and bi-orientation on the mitotic spindle. Nat. Rev. 2005;6:929–942. doi: 10.1038/nrm1764. [DOI] [PubMed] [Google Scholar]
- Vader G., Medema R. H., Lens S. M. The chromosomal passenger complex: guiding Aurora-B through mitosis. J. Cell Biol. 2006;173:833–837. doi: 10.1083/jcb.200604032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vagnarelli P., Earnshaw W. C. Chromosomal passengers: the four-dimensional regulation of mitotic events. Chromosoma. 2004;113:211–222. doi: 10.1007/s00412-004-0307-3. [DOI] [PubMed] [Google Scholar]
- Vong Q. P., Cao K., Li H. Y., Iglesias P. A., Zheng Y. Chromosome alignment and segregation regulated by ubiquitination of survivin. Science. 2005;310:1499–1504. doi: 10.1126/science.1120160. [DOI] [PubMed] [Google Scholar]
- Wallace W., Schaefer L. H., Swedlow J. R. A workingperson's guide to deconvolution in light microscopy. Biotechniques. 2001;31:1076–1080. doi: 10.2144/01315bi01. [DOI] [PubMed] [Google Scholar]
- Walther T. C., et al. The conserved Nup107-160 complex is critical for nuclear pore complex assembly. Cell. 2003;113:195–206. doi: 10.1016/s0092-8674(03)00235-6. [DOI] [PubMed] [Google Scholar]
- Weaver B. A., Bonday Z. Q., Putkey F. R., Kops G. J., Silk A. D., Cleveland D. W. Centromere-associated protein-E is essential for the mammalian mitotic checkpoint to prevent aneuploidy due to single chromosome loss. J. Cell Biol. 2003;162:551–563. doi: 10.1083/jcb.200303167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wheatley S. P., Barrett R. M., Andrews P. D., Medema R. H., Morley S. J., Swedlow J. R., Lens S. M. Phosphorylation by aurora-B negatively regulates survivin function during mitosis. Cell Cycle. 2007;6:1220–1230. doi: 10.4161/cc.6.10.4179. [DOI] [PubMed] [Google Scholar]
- Wheatley S. P., Carvalho A., Vagnarelli P., Earnshaw W. C. INCENP is required for proper targeting of Survivin to the centromeres and the anaphase spindle during mitosis. Curr. Biol. 2001;11:886–890. doi: 10.1016/s0960-9822(01)00238-x. [DOI] [PubMed] [Google Scholar]
- Wigge P. A., Kilmartin J. V. The Ndc80p complex from Saccharomyces cerevisiae contains conserved centromere components and has a function in chromosome segregation. J. Cell Biol. 2001;152:349–360. doi: 10.1083/jcb.152.2.349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Z., Tulu U. S., Wadsworth P., Rieder C. L. Kinetochore dynein is required for chromosome motion and congression independent of the spindle checkpoint. Curr. Biol. 2007;17:973–980. doi: 10.1016/j.cub.2007.04.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yue Z., et al. Deconstructing Survivin: comprehensive genetic analysis of Survivin function by conditional knockout in a vertebrate cell line. J. Cell Biol. 2008;183:279–296. doi: 10.1083/jcb.200806118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X. D., Goeres J., Zhang H., Yen T. J., Porter A. C., Matunis M. J. SUMO-2/3 modification and binding regulate the association of CENP-E with kinetochores and progression through mitosis. Mol. Cell. 2008;29:729–741. doi: 10.1016/j.molcel.2008.01.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuccolo M., et al. The human Nup107-160 nuclear pore subcomplex contributes to proper kinetochore functions. EMBO J. 2007;26:1853–1864. doi: 10.1038/sj.emboj.7601642. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.