Abstract
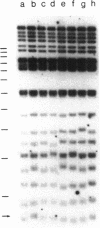
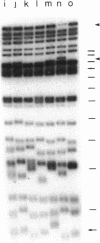
Chromosome ends in the lower eukaryotes terminate in variable numbers of tandem, simple DNA repeats. We tested predictions of a model in which these telomeric repeats provide a substrate for the addition of more repeats by a terminal transferase-like mechanism that, in concert with DNA polymerase and primase, effectively counterbalances the loss of DNA due to degradation or incomplete replication. For individual chromosome ends in yeast, the mean length of any given telomere was shown to vary between different clonal populations of the same strain and to be determined by the initial length of that telomere in the single cell giving rise to the clone. This type of variation was independent of the major yeast recombination pathway. The length heterogeneity at each telomeric end increased with additional rounds of cell division or DNA replication. Lengths of individual telomeres within a single clone varied independently of each other. Thus, this clonal variability is distinct from genetic regulation of chromosome length, which acts on all chromosome ends coordinately. These in vivo phenomena suggest that lengthening and shortening activities act on yeast telomeres during each round of replication.
Full text
PDF



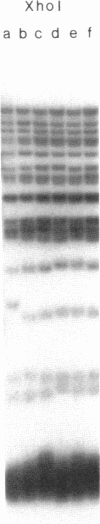
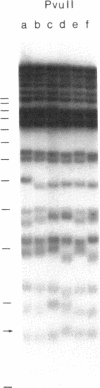
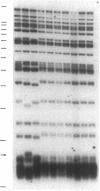
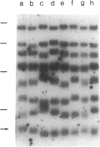
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bernards A., Michels P. A., Lincke C. R., Borst P. Growth of chromosome ends in multiplying trypanosomes. Nature. 1983 Jun 16;303(5918):592–597. doi: 10.1038/303592a0. [DOI] [PubMed] [Google Scholar]
- Blackburn E. H., Gall J. G. A tandemly repeated sequence at the termini of the extrachromosomal ribosomal RNA genes in Tetrahymena. J Mol Biol. 1978 Mar 25;120(1):33–53. doi: 10.1016/0022-2836(78)90294-2. [DOI] [PubMed] [Google Scholar]
- Blackburn E. H. The molecular structure of centromeres and telomeres. Annu Rev Biochem. 1984;53:163–194. doi: 10.1146/annurev.bi.53.070184.001115. [DOI] [PubMed] [Google Scholar]
- Carlson M., Celenza J. L., Eng F. J. Evolution of the dispersed SUC gene family of Saccharomyces by rearrangements of chromosome telomeres. Mol Cell Biol. 1985 Nov;5(11):2894–2902. doi: 10.1128/mcb.5.11.2894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carson M. J., Hartwell L. CDC17: an essential gene that prevents telomere elongation in yeast. Cell. 1985 Aug;42(1):249–257. doi: 10.1016/s0092-8674(85)80120-3. [DOI] [PubMed] [Google Scholar]
- Chan C. S., Tye B. K. Organization of DNA sequences and replication origins at yeast telomeres. Cell. 1983 Jun;33(2):563–573. doi: 10.1016/0092-8674(83)90437-3. [DOI] [PubMed] [Google Scholar]
- Dunn B., Szauter P., Pardue M. L., Szostak J. W. Transfer of yeast telomeres to linear plasmids by recombination. Cell. 1984 Nov;39(1):191–201. doi: 10.1016/0092-8674(84)90205-8. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. "A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity". Addendum. Anal Biochem. 1984 Feb;137(1):266–267. doi: 10.1016/0003-2697(84)90381-6. [DOI] [PubMed] [Google Scholar]
- Greider C. W., Blackburn E. H. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell. 1985 Dec;43(2 Pt 1):405–413. doi: 10.1016/0092-8674(85)90170-9. [DOI] [PubMed] [Google Scholar]
- Horowitz H., Thorburn P., Haber J. E. Rearrangements of highly polymorphic regions near telomeres of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Nov;4(11):2509–2517. doi: 10.1128/mcb.4.11.2509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klobutcher L. A., Swanton M. T., Donini P., Prescott D. M. All gene-sized DNA molecules in four species of hypotrichs have the same terminal sequence and an unusual 3' terminus. Proc Natl Acad Sci U S A. 1981 May;78(5):3015–3019. doi: 10.1073/pnas.78.5.3015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larson D. D., Spangler E. A., Blackburn E. H. Dynamics of telomere length variation in Tetrahymena thermophila. Cell. 1987 Jul 31;50(3):477–483. doi: 10.1016/0092-8674(87)90501-0. [DOI] [PubMed] [Google Scholar]
- Lustig A. J., Petes T. D. Identification of yeast mutants with altered telomere structure. Proc Natl Acad Sci U S A. 1986 Mar;83(5):1398–1402. doi: 10.1073/pnas.83.5.1398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morin G. B., Cech T. R. The telomeres of the linear mitochondrial DNA of Tetrahymena thermophila consist of 53 bp tandem repeats. Cell. 1986 Sep 12;46(6):873–883. doi: 10.1016/0092-8674(86)90069-3. [DOI] [PubMed] [Google Scholar]
- Pluta A. F., Dani G. M., Spear B. B., Zakian V. A. Elaboration of telomeres in yeast: recognition and modification of termini from Oxytricha macronuclear DNA. Proc Natl Acad Sci U S A. 1984 Mar;81(5):1475–1479. doi: 10.1073/pnas.81.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Shampay J., Szostak J. W., Blackburn E. H. DNA sequences of telomeres maintained in yeast. Nature. 1984 Jul 12;310(5973):154–157. doi: 10.1038/310154a0. [DOI] [PubMed] [Google Scholar]
- Smith G. E., Summers M. D. The bidirectional transfer of DNA and RNA to nitrocellulose or diazobenzyloxymethyl-paper. Anal Biochem. 1980 Nov 15;109(1):123–129. doi: 10.1016/0003-2697(80)90019-6. [DOI] [PubMed] [Google Scholar]
- Szostak J. W., Blackburn E. H. Cloning yeast telomeres on linear plasmid vectors. Cell. 1982 May;29(1):245–255. doi: 10.1016/0092-8674(82)90109-x. [DOI] [PubMed] [Google Scholar]
- Treco D., Arnheim N. The evolutionarily conserved repetitive sequence d(TG.AC)n promotes reciprocal exchange and generates unusual recombinant tetrads during yeast meiosis. Mol Cell Biol. 1986 Nov;6(11):3934–3947. doi: 10.1128/mcb.6.11.3934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walmsley R. M., Petes T. D. Genetic control of chromosome length in yeast. Proc Natl Acad Sci U S A. 1985 Jan;82(2):506–510. doi: 10.1073/pnas.82.2.506. [DOI] [PMC free article] [PubMed] [Google Scholar]