Abstract
Rod-derived Cone Viability Factor (RdCVF) is a trophic factor with therapeutic potential for the treatment of retinitis pigmentosa, a retinal disease that commonly results in blindness. RdCVF is encoded by Nucleoredoxin-like 1 (Nxnl1), a gene homologous with the family of thioredoxins that participate in the defense against oxidative stress. RdCVF expression is lost after rod degeneration in the first phase of retinitis pigmentosa, and this loss has been implicated in the more clinically significant secondary cone degeneration that often occurs. Here, we describe a study of the Nxnl1 promoter using an approach that combines promoter and transcriptomic analysis. By transfection of selected candidate transcription factors, chosen based upon their expression pattern, we identified the homeodomain proteins CHX10/VSX2, VSX1 and PAX4, as well as the zinc finger protein SP3, as factors that can stimulate both the mouse and human Nxnl1 promoter. In addition, CHX10/VSX2 binds to the Nxnl1 promoter in vivo. Since CHX10/VSX2 is expressed predominantly in the inner retina, this finding motivated us to demonstrate that RdCVF is expressed in the inner as well as the outer retina. Interestingly, the loss of rods in the rd1 mouse, a model of retinitis pigmentosa, is associated with decreased expression of RdCVF by inner retinal cells as well as by rods. Based upon these results, we propose an alternative therapeutic strategy aimed at recapitulating RdCVF expression in the inner retina, where cell loss is not significant, to prevent secondary cone death and central vision loss in patients suffering from retinitis pigmentosa.
INTRODUCTION
Retinitis pigmentosa (RP) is an inherited photoreceptor degeneration leading to progressive visual loss that affects approximately 1 in 3000 individuals worldwide (1). RP is characterized by an initial loss of night and peripheral vision caused by the degeneration of rod photoreceptors, and this is generally followed by the loss of central visual acuity due to secondary degeneration of cone photoreceptors (2–4). Forty loci responsible for RP have been mapped and 31 of the responsible genes have been identified (http://www.sph.uth.tmc.edu/RetNet/). Most of the mutations identified to date have been found in genes encoding proteins that are expressed specifically or preferentially by rods (2,5–7), and they generally act by triggering photoreceptor death by apoptosis (8,9).
The rd1 mouse model of RP carries a recessive mutation in the beta subunit of rod phosphodiesterase (Pde6b). The mutation leads first to the rapid degeneration of rods, and this is followed by the death of cones (6,10–12). The fact that the Pde6b gene is expressed solely by rods implies that the cones degenerate through a non-cell autonomous mechanism. Clinically, in humans, the secondary loss of cones is the event that represents the major visual handicap, since cones are necessary for color and high acuity vision.
To study the mechanisms underlying the secondary cone death, we transplanted the photoreceptor layer from normal mice into the eyes of rd1 mice just after rod loss and showed that these grafts resulted in the partial rescue of the host animals’ cones (13,14). In vitro, the trophic effect was shown to be mediated by protein(s) secreted by rods (13,14). A cone viability activity was previously described (15), but it was the identification of rod-derived cone viability factor (RdCVF), a novel trophic factor specifically expressed by rods that was identified by expression cloning, that led to a therapeutic strategy aimed at preventing cone death and central vision loss in patients suffering from RP (16). This strategy, in theory, should apply regardless of the nature of the mutation causing the primary rod degeneration (1). In this model, the death of rods in the first phase of the disease results in the loss of expression of RdCVF, which in turn leads to cone death through a mechanism related to the loss of trophic support (17). Consistent with this model is the intriguing finding that the sequences of RdCVF and its paralogue RdCVF2 (18) are homologous to the family of thioredoxins (19,20), proteins involved in the defense mechanisms against oxidative stress (21–23). RdCVF and RdCVF2 correspond to the unspliced products of the nucleoredoxin-like genes Nxnl1 (Txnl6) and Nxnl2, respectively, and encode enzymatically inactive proteins with a truncation within the thioredoxin fold (18), similar to TRX80 (24). Interestingly, the spliced products of the nucleoredoxin-like genes, the longer isoforms RdCVFL and RdCVF2L, which encode putative thiol-oxidoreductase enzymes, may provide a way to link photo-oxidative stress that affects cones particularly (25) to an adaptive trophic response (26).
We hypothesize that protection of cones in RP could be achieved by recapitulating normal RdCVF expression in patients whose rods have been lost. One way to accomplish this would be through expression of exogenous RdCVF, for example by viral vector delivery (27), encapsulated cell technology (28,29) or by protein injection (30). Another approach would be to increase retinal levels of RdCVF by modulating expression of the endogenous gene. The restricted expression of RdCVF to the retina (16) (see also Txnl6 at http://biogps.gnf.org) suggested that it might be possible to upregulate its promoter without significantly affecting expression in other tissues. With this goal in mind, we have undertaken an effort to define the Nxnl1 promoter and the transcription factors (TFs) that regulate its activity. We have taken advantage of the opportunities provided by analysis of the transcriptome of the outer retina to develop a bioinformatic approach to identify candidate TFs that could regulate Nxnl1 expression. This analysis identified 44 candidate TFs, and these were screened in a transient transfection assay for the ability to transactivate the Nxnl1 promoter. We demonstrate here that the homeoproteins CHX10/VSX2, VSX1 and PAX4, and the zinc finger protein SP3 can activate both the mouse and human RdCVF promoters. The identification of CHX10/VSX2, VSX1, which have been reported elsewhere to be involved in bipolar cells differentiation (31,32), led us to reexamine the expression of RdCVF in the retina. This reexamination demonstrated that RdCVF is not only expressed by photoreceptors as previously thought, but is also expressed in the inner retina, most likely by bipolar cells.
RESULTS
Selection of TFs candidates
In order to identify TFs that were candidate regulators of the nucleoredoxin-like 1 (Nxnl1) gene, two successive incremented filters were applied. First, the Transfac® database was used to identify TFs that are predicted to bind to the 4.2 kb region upstream of the mouse Nxnl1 gene. This analysis yielded 133 predicted TFs (Supplementary Material, Table S1). The second filter, which retained from this set only those TFs expressed in the photoreceptor-containing outer region of the retina (402 TFs), reduced the candidate list to 44 TFs (Table 1). This selection utilized Affymetrix-based transcriptome analysis of the outer retina of 35-day-old mice. The expression profiles of this subset of 44 TF candidates can be classified into two patterns based upon relative expression in the photoreceptor region of the retina versus the whole (neural) retina: 27 TFs have an outer retina/neural retina ratio above 1, indicating preferential expression in the photoreceptor layer (e.g. Crx) and 32 TFs whose retina/brain ratio is higher that 1, potentially indicating involvement in a function that is particularly important in the retina (e.g. Chx10/Vsx2). As deduced from the pattern of expressed sequence tags (ESTs) available in the public databases, some of the 44 candidate factors are preferentially expressed in the retina, whereas others are more widely expressed (Table 1).
Table 1.
Expression level of the TFs selected in the study
| Gene ID NCBI | Gene name | Gene symbol | Probe set ID | Signal intensity (RMA) |
OR/NR | Tissue |
Clone number | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| OR | NR | Brain | ≤5 | >5 | ND | ||||||
| 1406 | Cone-rod homeobox containing gene | Crx | 99900_at | 1496.55 | 1028.57 | 138.36 | 1.45 | • | LA0AAA31YM07 | ||
| 11911 | Activating transcription factor 4 | Atf4 | 100599_at | 1162.04 | 834.98 | 526.77 | 1.39 | • | LA0AAA82YD23 | ||
| 20787 | Sterol regulatory element binding transcription factor 1 | Srebfl | 93264_at | 603.15 | 620.27 | 344.61 | 0.97 | • | 3030000054066890 | ||
| 22278 | Upstream transcription factor 1 | Usfl | 93655_at | 509.79 | 502.43 | 565.12 | 1.01 | • | LA0AAA18YG17 | ||
| 14886 | General transcription factor II I | Gtf2i | 94295_at | 490.59 | 657.20 | 443.33 | 0.75 | • | O-MC203628 | ||
| 17258 | Myocyte enhancer factor 2A | Mef2a | 93852_at | 467.42 | 256.30 | 334.67 | 1.82 | • | 1-4979487 | ||
| 12677 | Visual system homeobox 2/C. elegans ceh-10 homeodomain containing homolog | Chxl0/Vsx2 | 101127_at | 422.56 | 676.39 | 174.46 | 0.62 | • | LA0AAA60YM14 | ||
| 22282 | Upstream transcription factor 2 | Usf2 | 103013_at | 403.89 | 381.70 | 336.32 | 1.06 | • | LAOAAA14YJO6 | ||
| 20850 | Signal transducer and activator of transcription 5A | Stat5a | 100422_i_at | 348.98 | 393.99 | 275.96 | 0.89 | • | LAOAAA118YN05 | ||
| 18519 | p300/CBP-associated factor | Pcaf/Kat2b | 104070_at | 347.69 | 270.53 | 107.61 | 1.29 | • | O-MC206206 | ||
| 15110 | Heart and neural crest derivatives expressed transcript 1 | Hand1 | 92766_at | 298.25 | 272.02 | 275.05 | 1.10 | • | O-MC201881 | ||
| 18986 | POU domain, class 2, transcription factor 1 | Pou2f1 | 102894_g_at | 287.05 | 198.01 | 105.69 | 1.45 | • | O-SC321473 | ||
| 19883 | RAR-related orphan receptor alpha | Rora | 101889_s_at | 250.18 | 246.88 | 130.27 | 1.01 | • | 1-3592667 | ||
| 17187 | Max protein | Max | 99095_at | 249.53 | 168.31 | 128.92 | 1.48 | • | LA0AAA9YP20 | ||
| 13047 | Cut-like 1 | Cutl1/Cux1 | 98073_at | 244.28 | 247.43 | 245.71 | 0.99 | • | LAOAAA19YG15 | ||
| 22130 | Transcription termination factor 1 | Ttf1 | 96435_at | 242.04 | 200.37 | 165.17 | 1.21 | • | pEGFP-C1-TTF1 | ||
| 21406 | Transcription factor 12 | Tcf12 | 9898l_s_at | 230.07 | 183.68 | 127.60 | 1.25 | • | 1-5345693 | ||
| 18506 | Paired box gene 4 | Pax4 | 99908_at | 223.70 | 212.43 | 236.84 | 1.05 | • | pCMV5-Pax4 | ||
| 13653 | Early growth response 1 | Egr1 | 98579_at | 222.67 | 147.48 | 210.74 | 1.51 | • | LA0AAA78YH20 | ||
| 14463 | GATA binding protein 4 | Gata4 | 102713_at | 212.37 | 206.01 | 222.56 | 1.03 | • | pCDNA3-GATA4 | ||
| 22632 | YY1 transcription factor | Yy1 | 98767_at | 212.22 | 207.68 | 170.07 | 1.02 | • | LAOAAA124YE13 | ||
| 20687 | Trans-acting transcription factor 3 | Sp3 | 96192_at | 207.29 | 155.13 | 103.29 | 1.34 | • | 1-5323205 | ||
| 11695 | Aristaless 4 | Alx4 | 101622_at | 205.97 | 207.49 | 210.68 | 0.99 | 1-6506755 | |||
| 17128 | MAD homolog 4 (Drosophila) | Smad4 | 160440_at | 193.88 | 170.61 | 112.18 | 1.14 | • | 1-6313280 | ||
| 20181 | Retinoid X receptor alpha | Rxra | 92235_g_at | 178.40 | 153.92 | 167.35 | 1.16 | • | LAOAAA26YEO1 | ||
| 22772 | Zic finger protein of the cerebellum 2 | Zic2 | 98844_at | 172.96 | 194.99 | 186.02 | 0.89 | • | pcDNA-Zic2 | ||
| 20683 | Trans-acting transcription factor 1 | Sp1 | 100032_at | 163.35 | 123.95 | 100.51 | 1.32 | • | O-SC101137 | ||
| 11835 | Androgen receptor | Ar | 92667_at | 150.68 | 134.52 | 145.60 | 1.12 | • | pSG5-Ar | ||
| 17428 | Max binding protein | Mnt | 92300_at | 149.80 | 176.99 | 154.56 | 0.85 | • | pRc/CMV-Mnt | ||
| 18033 | Nuclear factor of kappa light chain gene enhancer in B-cells 1, p105 | Nfkb1 | 98427_s_at | 149.05 | 166.08 | 155.34 | 0.90 | • | LAOAAA79YO10 | ||
| 19724 | Regulatory factor X, 1 (influences HLA class II expression) | Rfx1 | 99880_at | 146.38 | 117.93 | 105.17 | 1.24 | • | LAOAAAl11Y022 | ||
| 21909 | T-cell leukemia, homeobox 2 | Tlx2 | 97679_at | 146.02 | 159.19 | 136.67 | 0.92 | • | O-SC123839 | ||
| 17764 | Metal response element binding transcription factor 1 | Mtf1 | 100018_at | 144.31 | 161.30 | 138.83 | 0.89 | • | 1-4210829 | ||
| 15499 | Heat shock factor 1 | Hsf1 | 10045 l_at | 144.31 | 129.06 | 108.95 | 1.12 | • | LAOAAA40YJ15 | ||
| 13712 | ELK1, member of ETS oncogene family | Elk1 | 96593_at | 139.54 | 128.23 | 146.58 | 1.09 | • | O-MC201893 | ||
| 71702 | Cell division cycle 5-like (S. pombe) | Cdc5l | 97846_at | 126.01 | 139.73 | 103.93 | 0.90 | • | 3030000053137993 | ||
| 11634 | Autoimmune regulator (autoimmune polyendocrinopathy candidiasis ectodermal dystrophy) | Aire | 97159_at | 122.46 | 113.97 | 111.15 | 1.07 | • | LA0AAA87YA17 | ||
| 18030 | Nuclear factor, interleukin 3, regulated | Nfil3 | 101805_f_at | 119.72 | 121.57 | 117.12 | 0.98 | • | LA0AAA21YP14 | ||
| 16871 | LIM homeobox protein 3 | Lhx3 | 102902_at | 116.83 | 123.03 | 74.42 | 0.95 | • | pCDNA3-Lhx3 | ||
| 18771 | Pbx/knotted 1 homeobox | Pknox1 | 102257_at | 109.04 | 104.19 | 94.44 | 1.05 | • | LA0AAA33YP05 | ||
| 21417 | Zinc finger E-box binding homeobox 1 | Zeb1 | 99052_at | 106.50 | 127.10 | 151.06 | 0.84 | • | pcl-Flag-Zebl | ||
| 54123 | Interferon regulatory factor 7 | Irf7 | 104669_at | 103.14 | 106.28 | 83.20 | 0.97 | • | LA0AAA120YH12 | ||
| 54006 | Deformed epidermal autoregulatory factor 1 (Drosophila) | Deaf1 | 96171_at | 101.56 | 105.33 | 99.68 | 0.96 | • | 1-5063995 | ||
| 16476 | Jun oncogene | Jun | 100130_at | 99.73 | 87.21 | 105.48 | 1.14 | • | LAOAAA20YJO1 | ||
OR, outer retina; NR, neural retina. The tissue distribution of ESTs within the public database (≤5 tissues with eyes), large (>5 tissues with eyes) or no determined (ND) of each TFs in mouse (•) tissues (according to UniGene, NCBI). Clone number refers to the sequenced library or to the plasmids used.
Screening of candidate TFs for their activity to transactivate the Nxnl1 promoter
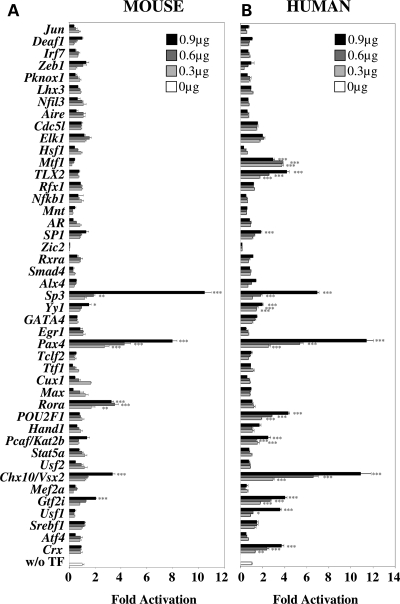
Although the mouse and human Nxnl1 5′ upstream regions do not show any obvious sequence conservation (Supplementary Material, Fig. S1), which we found to be a surprising finding given their similar expression patterns, we nevertheless screened the 44 candidate TFs on both mouse and human promoter constructs because we felt that the finding of common activity between the two species would likely indicate functionally important conservation that might not be evident from simple linear sequence comparison. For the transactivation assays, 4.2 and 2.1 kb of the mouse and human 5′ Nxnl1 upstream regions, respectively, were cloned upstream of the luciferase reporter plasmid pGL4. The 44 candidate TFs were tested by transient transfection of HEK 293 cells with increasing dose (0.3, 0.6 or 0.9 µg) of the respective expression plasmid, and reporter activity was tested by dual luciferase assay 48 h post-transfection. Chx10/Vsx2, Gtf2i, Rora, Pax4, Yy1 and Sp3 were all found to activate the mouse promoter in a dose dependant manner (Fig. 1A). At the highest transfected dose (0.9 µg), Chx10/Vsx2 demonstrated 3.3 ± 0.2-fold activation (P < 0.001), Gtf2i 2.1 ± 0.1 (P < 0.001); Rora 3.3 ± 0.2 (P < 0.001), Pax4 8.0 ± 0.4 (P < 0.001), Yy1 1.5 ± 0.2 (P < 0.001) and Sp3 demonstrated 10.5 ± 0.5-fold (P < 0.001) activation. We also noticed that some TFs, as Cux1 and Max reduce the activity of the promoter in a dose-dependent manner. The potential role of homeogenes as Chx10/Vsx2 and Pax4 in the regulation of RdCVF expression was further highlighted when the 44 TFs were assayed on the human NXNL1 5′ upstream region. Chx10/Vsx2 demonstrated 10.9 ± 0.9-fold activation (P < 0.001) at the highest dose (0.9 µg), Pax4 11.4 ± 0.6 (P < 0.001), Crx 3.7 ± 0.2 (P < 0.001), TLX2 4.2 ± 0.2 (P < 0.001), Sp3 6.9 ± 0.2 (P < 0.001), SP1 1.8 ± 0.1 (P < 0.001), Usf1 3.6 ± 0.1 (P < 0.001), Gtf2i 4.0 ± 0.2 (P < 0.001), Pcaf/Kat2b 2.5 ± 0.2 (P < 0.001), POU2F1 4.3 ± 0.1 (P < 0.001) and Yy1 2.0 ± 0.1-fold (P < 0.001) activation (Fig. 1B).
Figure 1.
Transcription factors (TFs) activity on both mouse and human Nxnl1 promoter constructs. (A) Assay with the murine 4.2 kb Nxnl1 5′ upstream region. (B) Assay with the human 2.1 kb NXNL1 5′ upstream region. The TFs were tested by transient transfection of 0.3, 0.6 and 0.9 µg of plasmid coding for each candidate TFs and 0.1 µg of reporter plasmid in HEK 293 cells and assessed by dual luciferase assays. The results were normalized by using an internal control plasmid (Renilla) driven by the thymidin kinase promoter. Each experiment was repeated in triplicate. P < 0.001 (***); P < 0.01 (**); P < 0.05 (*).
Activity of other homeodomain proteins on the mouse and human Nxnl1 promoters
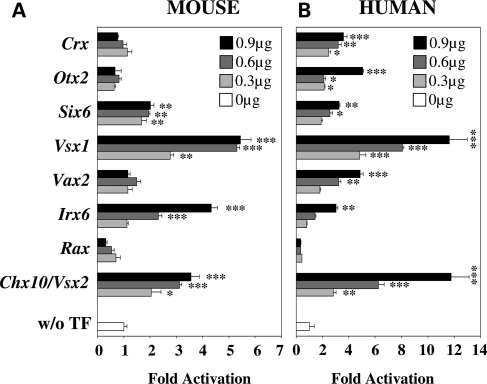
Since our bioinformatics analysis of the Nxnl1 promoter could have missed some important regulators, and because of the high frequency of homeodomain (HD) factors identified in our transient transfection analysis (Fig. 1) together with the known importance of HD proteins as important regulators of retinal gene expression (33), we evaluated the ability of additional HD proteins expressed in the retina (RAX, IRX6, VAX2, VSX1, SIX6 and OTX2) for their ability to activate the Nxnl1 promoter. On the mouse promoter, of the total of eight HD factors tested, four showed dose-dependent activation activity (Fig. 2A). At the highest dose, Chx10/Vsx2 activated the promoter 3.6 ± 0.3-fold (P < 0.001), Irx6 4.3 ± 0.2-fold (P < 0.001), Vsx1 5.4 ± 0.4-fold (P < 0.001) and Six6 2.0 ± 0.1-fold (P < 0.01). On the human promoter, seven of the eight HDs tested showed dose-dependent activation (Fig. 2B). At the highest dose, Chx10/Vsx2 activated the promoter 11.8 ± 1.4-fold (P < 0.001), Irx6 3.0 ± 0.1-fold (P < 0.01), Vax2 4.8 ± 0.2-fold (P < 0.001), Vsx1 11.6 ± 1.4-fold (P < 0.001), Six6 3.3 ± 0.1-fold (P < 0.01), Otx2 5.1 ± 0.1-fold (P < 0.001) and Crx 3.6 ± 0.3-fold (P < 0.001). Chx101/Vsx2, Vsx1, Six6 and Irx6 regulated both the mouse and human constructs. These results highlight the potential importance of HD proteins as regulators of RdCVF expression. Furthermore, the fact that only a subset of the tested HD proteins were able to active the RdCVF promoter, even though they all recognize a common TAAT core sequence, demonstrates substantial specificity in the reporter assay used in our analysis.
Figure 2.
Activity of retinally-expressed homeogenes on mouse and human Nxnl1 promoters constructs (A) Activity of homeogenes on mouse Nxnl1 promoter, and (B) on human NXNL1 promoter. The HD TFs were tested by transient transfection of 0.3, 0.6 and 0.9 µg of plasmid coding for each candidate HDs TF and 0.1 µg of reporter plasmid in HEK 293 cells and assessed by dual luciferase assays. The results were normalized by using an internal control plasmid (Renilla) driven by the thymidin kinase promoter. Each experiment was repeated in triplicate. P < 0.001 (***); P < 0.01 (**); P < 0.05 (*).
RdCVF is expressed by cells in the inner retina
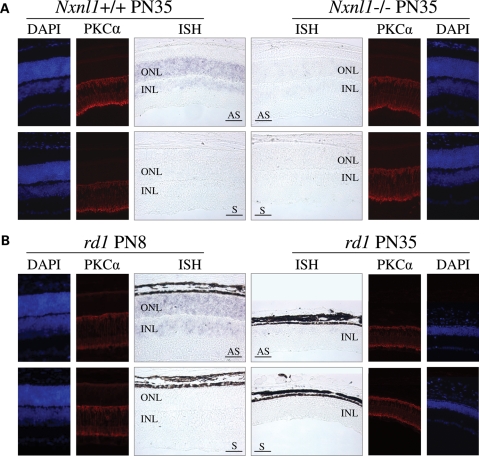
From our combined studies, the TFs that showed activity with both the mouse and human promoters were Chx10/Vsx2, Vsx1, Pax4, Sp3, Six6 and Irx6. As noted above, we felt that these factors that were active on both the mouse and human Nxnl1 promoter constructs were the strongest candidates to be biologically important regulators of RdCVF expression in vivo. We were therefore surprised to see Chx10/Vsx2 and Vsx1 in this group because both have been reported to be expressed in the inner nuclear layer (INL) of the retina but not in photoreceptors, and RdCVF was thought to be predominantly expressed in photoreceptors, although a possible signal was present in the INL (16). To resolve this uncertainty, we reexamined the expression pattern of RdCVF by in situ hybridization. Using an exon 1 riboprobe that should detect mRNA for both RdCVF and RdCVFL, we observed expression of RdCVF in the outer nuclear layer (ONL) of wild-type (wt) mice, which is consistent with what was observed previously (16). However, we also detected a weaker but reproducibly present in situ signal in the INL. As a control to test whether this INL signal was truly due to Nxnl1 expression or reflected background signal or perhaps cross-hybridization with another gene, we tested retinal sections from a Nxnl1 null mouse (Fig. 3A) (Cronin et al., in revision). The absence of signal in both the ONL and INL of the Nxnl1−/− retina demonstrates that the observed signal in the wt retina is real, indicating that in addition to the ONL, RdCVF is also expressed in the INL of the mouse retina.
Figure 3.
Nxnl1 is expressed in the outer and inner retina. (A) In situ hybridization and immunolocalization of PKCα (red) on Nxnl1+/+ or Nxnl1−/− retinal sections from mice at PN35. (B) In situ hybridization and immunolocalization of PKCα (red) on rd1 mice at PN8 and PN35 aligned at the INL level. Nuclei stained with DAPI (blue). ONL, Outer Nuclear Layer; INL, Inner Nuclear Layer. AS, antisense and S, sense riboprobes. Bar scale 50 µm.
According to our previous observations (16), RdCVF is expressed in a rod-dependant manner since it is easily detectable in the retina of PN8 rd1 mice, but it is not detectable at PN35, after rod degeneration (Fig. 3B). The loss of ONL expression of RdCVF seems explained by the loss of rods. However, the loss of RdCVF expression observed in the INL seemed surprising since there is no obvious loss of INL cells. Since Chx10/Vsx2 is involved in the differentiation of rod-bipolar cells, we examined the rod-bipolar cells in the retina at PN35 using protein kinase Cα (PKCα, red) as a marker (34). We did not observe any difference in PKCα labeling between rd1 mice at PN8 and PN35 (Fig. 3B) or between the Nxnl1+/+ and −/− retinas (Fig. 3A). These results suggest that the INL cells that were producing RdCVF are still present, but have turned off their expression of the gene.
Increased expression of Chx10/Vsx2 in the inner retina of rd1 mice
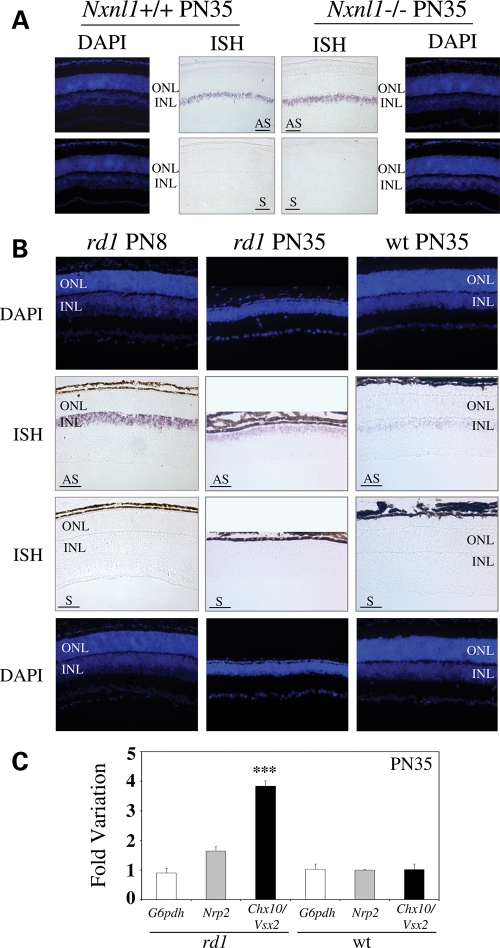
The expression profile of Chx10/Vsx2 was also studied by in situ hybridization in wt and mutant mice. The expression of Chx10/Vsx2 was localized in the INL of wt mice, consistent with a report (35), and was found to be expressed at a similar level in the retina of Nxnl1−/− mice (Fig. 4A). In the rd1 retina, we observed that expression of Chx10/Vsx2 was down regulated between PN8 and PN35 (Fig. 4B), as reported in the wt mouse (36). However, at PN35, the expression of Chx10/Vsx2 is higher in rd1 compared with wt retina (Fig. 4B), an observation that we confirmed by real time PCR (Fig. 4C). One trivial technical explanation for the observed increased expression of Chx10/Vsx2 is that the lack of ONL cells at P35 causes the INL cells to be a higher proportion of the remaining total retinal cells, and hence RNA from bipolar cells, for example, would appear to represent a higher fraction of total retinal RNA. To take into account this potential ‘artifact’, we analyzed the expression of Nrp2, a gene specifically expressed by ganglion cells (37) (Supplementary Material, Fig. S2), since enrichment of ganglion cell RNA should parallel that of INL cells. The expression of Nrp2 was indeed increased in wt compared with rd1 retina (1.7 ± 0.1-fold), in a proportion that corresponded to the loss of rod photoreceptors in the rd1 retina. Using this value for normalization, the observed 3.8 ± 0.2-fold increase in Chx10/Vsx2 expression would represent an actual increase of 2.4 ± 0.1-fold, indicating that there is indeed a real increase in Chx10/Vsx2 expression at the cellular level in the rd1 retina after rod degeneration.
Figure 4.
Chx10/Vsx2 expression profile on Nxnl1+/+, Nxnl1−/− and rd1 mouse retinas at post-natal 8 and 35. (A) In situ hybridization on Nxnl1+/+ or Nxnl1−/− retinas at PN35. (B) In situ hybridization on rd1 mice at PN35 or PN8 and wild-type (wt) at PN35, aligned at the INL level. (C) Relative gene expression level in wt mice and rd1 at PN35. Nrp2, neuropilin 2 (Mm.266341). Nuclei are stained with DAPI (blue). ONL, Outer Nuclear Layer; INL, Inner Nuclear Layer; AS, antisense and S, sense riboprobes. Bar scale 50 µm. P < 0.001 (***).
Chx10/Vsx2 binds the Nxnl1 promoter in vivo
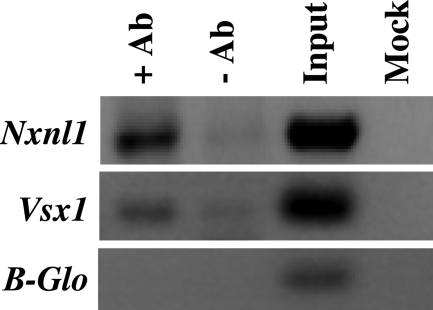
The above data are consistent with the possibility that Chx10/Vsx2 could regulate RdCVF expression in INL cells in vivo. In order to assess this hypothesis, we performed chromatin immunoprecipitation (ChIP) with mouse retina and anti-CHX10/VSX2 antibodies. The immunoprecipitate was used as a template for PCR with primers specific to the Nxnl1, Vsx1 and β-globin promoter regions (negative control). As a positive control, we used the known interaction between CHX10/VSX2 protein and Vsx1 promoter (38). Compared with the no antibody control, amplification of the Nxnl1 promoter region was significantly enhanced by the presence of the anti-CHX10/VSX2 antibody (Fig. 5). This shows that in the retina in vivo, a fraction of CHX10/VSX2 is bound to the Nxnl1 promoter, supporting a model in which the activation of expression of RdCVF by CHX10/VSX2 is mediated by binding to an element within the promoter.
Figure 5.
CHX10/VSX2 binds to the Nxnl1 promoter in vivo shown by chromosome immunoprecipitation assay (ChIP). ChIP was done with wt mouse retina. +Ab, anti-CHX10/VSX2 antibodies; –Ab, no antibody; Input, mouse genomic DNA; Mock, without mouse genomic DNA; Vsx1 and β-Globin (β-Glo) promoters amplification were used as positive and negative controls, respectively.
Synergistic activation of the Nxnl1 promoters
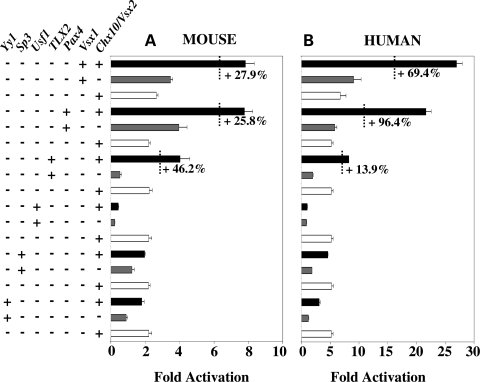
To explore possible interactions and co-operativity between Chx10/Vsx2 and other selected TFs, we tested Chx10/Vsx2 activity individually or in combination with a subset of TFs (Fig. 6A and B). On the mouse promoter, the results showed that significant enhancement of transcriptional activation could be obtained upon co-expression of Chx10/Vsx2 and Vsx1 (+27.9% more than a simple additive effect), Chx10/Vsx2-Pax4 (+25.8%), Chx10/Vsx2-TLX2 (+46.2%). On the human promoter, the synergistic activity was observed for the same couple of TFs: Chx10/Vsx2-Vsx1 (+69.4%), Chx10/Vsx2-Pax4 (+96.4%), Chx10/Vsx2-TLX2 (+13.9%). The fact than 0.9 µg of each TF transfected individually (Figs 1 and 2) does not activate the RdCVF 5′ region to the same extent as the combination of 0.5 µg of Chx10/Vsx2 + 0.5 µg of the other TF (Fig. 6) supports the synergetic effect observed here. Although Usf1 can activate the human Nxnl1 Promoter, its co-transfection with Chx10/Vsx2 led to a reduction of activity with both the mouse and human Nxnl1 promoter constructs.
Figure 6.
Chx10/Vsx2 acts with other TFs to synergistically activate the Nxnl1 promoter. Activity of the Chx10/Vsx2 with Vsx1, Pax4, TLX2, Usf1, Sp3 and Yy1 TFs on the (A) mouse and (B) on human Nxnl1 promoter. Activity tested by transient transfection in HEK 293 cells with dual luciferase assay. About 0.5 µg of each TF was tested individually or paired with 0.5 µg of Chx10/Vsx2 in co-transfection assays with 0.1 µg of reporter plasmid. The results are expressed as percentage of synergistic activation over the sum of both TFs when tested individually. The results were normalized by internal control (Renilla) driven by the thymidine kinase promoter. Each experiment was repeated in triplicate.
DISCUSSION
A bioinformatic approach to study the Nxnl1 promoter
The methodologies employed in the study of gene regulation at the promoter level classically involve gene reporter assays using transient transfection of luciferase, or other, reporters aimed at mapping the promoter sequences necessary and sufficient to drive the activity of the reporter constructs. Thereafter, this mapping information is used to identify, by diverse approaches including one-hybrid screening, the TFs that bind to these DNA elements (39,40). The development of functional genomic approaches, such as transcriptomics, and bioinformatics tools allowed us to employ an alternative strategy. The sequences of DNA that define the binding sites of TFs are generally quite degenerate, and as a result programs aimed at identifying putative TF binding sites within promoter regions are often imprecise and give variable results depending upon the binding thresholds that are chosen, and result in considerable false positive and false negative predictions. For example, use of the Transfac® database identified 13 242 putative DNA binding elements within the 4.2 kb sequence 5′ upstream of the translation initiation site of the Nxnl1 gene. As an alternative approach, since the absence of phylogenic conservation of the Nxnl1 promoter region prevented us from using sequence conservation as a method to aid in the identification of functionally important sites, we reasoned that the TFs regulating the Nxnl1 gene should be expressed in the outer retina. We further reasoned that those TFs that could transactivate both mouse and human Nxnl1 promoter constructs would be particularly strong candidates as biologically relevant regulators. We have defined the transcriptome of the outer retina of the mouse using Affymetrix GeneChip®-based microarray analysis, work that took advantage of the layered structure of the retina which allows the purification of the outer retina by sectioning of flat-mounted retinas using a vibratome (41). Signal intensity was used to filter among the TFs that possess a predicted DNA binding element in the Nxnl1 promoter, i.e. those whose expression was detected over a given threshold were chosen (see Material and Methods). The threshold used corresponds to the value above which the expression is unambiguous. This led us to establish a list of 44 candidate TFs that was tested in a transient transfection reporter assay. This novel approach was successfully developed and led to the identification of three TFs, two homeoproteins (CHX10/VSX2 and PAX4)) and one zinc finger (SP3) protein that could regulate Nxnl1 expression.
Chx10/Vsx2 regulates the activity of the Nxnl1 promoter
By screening the list of candidate TFs selected through these bioinformatics and expression criteria, we identified three factors (Pax4, Chx10/Vsx2 and Sp3) that in a transient transfection-based assay demonstrated reproducible activity on both the mouse and human Nxnl1 promoters. This result was further substantiated by the observation that another Chx10-like factor, Vsx1, also demonstrates a similar activity, and by the finding of synergistic activation of Chx10/Vsx2 with Vsx1, Pax4 and TLX2 on both mouse and human Nxnl1 promoters. The proposed biological role of Chx10/Vsx2 in the regulation of RdCVF expression in vivo is supported by ChIP studies (Fig. 5).
In our transcriptomic data, the TFs were selected through their expression in the outer retina. Thus, the identification of Pax4 and Sp3 seemed reasonable given that Pax4 is expressed by photoreceptor cells (42) and the Sp3-related factor Sp4 regulates the photoreceptor gene Pde6b (43). However, given their exclusive expression in the inner retina (31,32,44,45), the presence of Chx10/Vsx2 and Vsx1 in the outer retina preparation was unexpected and most likely results from a contamination during vibratome sectioning. Considering that RdCVF was originally described to be expressed by rod photoreceptors, its regulation by TFs normally present in the inner part of the retina was surprising. The data presented in this article show that RdCVF is expressed in inner retinal cells in addition to photoreceptors (Fig. 3A). The exon 1 probe used detects mRNA encoding both the short and the long isoforms of RdCVF. Additionally, the two other homeoproteins, SIX6 and OTX2, that activate the Nxnl1 promoter (Fig. 2), are expressed in the INL (46,47), further supporting the role of these HD proteins in the expression of RdCVF in the INL. The Nxnl1 signal revealed by in situ hybridization most likely arises from bipolar cells since we have previously demonstrated by RT–PCR, using freshly dissociated cells, that Müller glial cells do not express RdCVF (16). Although the role of the expression of RdCVF by bipolar cells is unclear, it is perhaps significant that CRX, a TF expressed in photoreceptors that controls photoreceptor differentiation (48), is also expressed in the INL (36,49). Additionally, since the Nxnl1 gene encodes secreted proteins, its expression by bipolar cells may be related to the trophic support of neighboring cells. The data presented here are consistent with what we have previously reported in 2004, with a lower Nxnl1 signal in the inner retina (16).
An alternative strategy for preventing central vision loss
In the rd1 mouse, a model of RP, the expression by rods of the trophic factor RdCVF is lost through their degeneration, but the reduction in its expression by bipolar cells results from a downregulation of its expression (Fig. 3B). This phenomenon does not result from a decrease in Chx10/Vsx2 expression (Fig. 4B and C), nor of Vsx1 (data not shown). This observation indicates that Nxnl1 expression is rod-dependant, and suggests that cell-to-cell communication between photoreceptor cells and bipolar cells is needed to maintain Nxnl1 expression. As shown in Figure 4B and C, it is possible that the increase of Chx10/Vsx2 expression by degenerating retina could represent a compensatory response for decreased photoreceptor-derived RdCVF, perhaps mediated by increasing the factors responsible for its expression by bipolar cells. This creates by extension an alternative therapeutic strategy aimed at recapitulation of RdCVF expression in the retina after rod degeneration by increasing its expression by bipolar cells. Such increased expression by bipolar cells could potentially sustain cones and thereby maintain central vision. This strategy is further substantiated by the finding that the injection of RdCVF protein in the P23H transgenic rat, a dominant model of rod-cone degeneration, not only protects the cones but preserves the cone ERG to an even greater extent (30). What could be the reason for the loss of expression of RdCVF by bipolar cells in the degenerating retina and what could be the best means to drive its expression after rod loss are matters for future research.
MATERIALS AND METHODS
Animals
Care and handling of mice in these studies conformed to the rules established by the Association for Research in Vision and Ophthalmology. Retinal tissues were obtained from post-natal (PN) 8 or 35 day C3H wt/wt, C3H rd1/rd1 (50), BALB/c wt/wt or BALB/c Nxnl1−/− mice. Nxnl1−/− mice were created by homologous recombination of the entire exon 1, which encodes the short isoform of RdCVF and the N-terminal part of RdCVFL, on a pure BALB/c background (Cronin et al., in revision). Animals were sacrificed and eyes were removed for in situ hybridization as described.
Promoter reporter plasmids construction
The sequence upstream to ATG of both mouse and human Nxnl1 genes were cloned into the luciferase reporter plasmid pGL4.17 (Promega) using blunt-end cloning at the EcoRV site for the mouse 4.136 kb (chr8:74090479 + 74094614 at UCSC genome browser) 5′ sequence of and with the gateway technology (Invitrogen) for 2.133 kb (chr19:17432676–17434808 at UCSC genome browser) of the 5′ human NXNL1 sequence. Mouse genomic DNA was amplified by PCR with Pfx High fidelity polymerase (Invitrogen) using conditions provided by the supplier (Primers: Reverse 5′-GGTAGCAGTATGCAAGGAGCTG-3′/Forward 5′-GAGCACACAGCTGAGACCGCAGGTACCGC-3′). For the human promoter construct, the pGL4.17 vector was converted into a gateway destination vector by introducing a cassette containing the attR sites flanking the ccdB gene into the multiple cloning sites. A DNA fragment corresponding to the human 5′ sequence was obtained by PCR with Platinium Taq DNA Polymerase High Fidelity (Invitrogen) with human genomic DNA as template. Primers were designed to amplify the 5′upstream region of the NXNL1 gene, and the attB sites were incorporated into the forward (attB site 5′-GGGGACAAGTTTGTACAAAAAAGCAGGCT-3′) and the reverse (attB site 5′-GGGGACCACTTTGTACAAGAAAGCTGGGT-3′) primers. The PCR reactions were subjected to a hot start of 5 min at 94°C prior to the addition of the polymerase followed by seven cycles of amplification: denaturing for 25 s at 94°C, annealing for 7 min at 66°C, 35 cycles amplification: denaturing for 25 s at 94°C, annealing for 30 s to 7 min at 60°C and a final extension at 66°C for 10 min (Primers: Reverse 5′-GGCGGTAACCTGGGTTGGGTGCTGGGGAC-3′/Forward 5′-CCTTGAAGTAACTAGTAGAGTCCATGTGA-3′). Positive clones were identified by restriction profiling and verified by DNA sequencing. DNA used for transient transfection was prepared using Qiagen plasmid maxi-prep according to the manufacturer's protocol.
Expression plasmids for TFs
Twenty one to 44 TFs come from a normalized neural retinal library. This mouse library was constructed using mRNA purified from neural retina of C57BL/6@N mice at 8 and 35 days PN by Cesium Chloride ultracentrifugation (51), followed by oligodT purification on oligoTex (Qiagen). The normalization was performed by Invitrogen and the normalized libraries in pCMVsport6 were swapped altogether into pEXP-cDNA3.1 (5.6 kb), by Gateway technology (Invitrogen). Large scale sequencing of this library was performed at the Genoscope (www.genoscope.cns.fr/spip/Mus-musculus-degeneration-of.htm) using the 5′ Sequencing primer: 5′-TTAATACGACTCACTATAGGG-3′. Plasmids from neural retinal normalized library are identified by their identifying number in the Table 1. Fifteen to 44 expression vectors were obtained from commercial sources. Gtf2i, Pcaf/Kat2b, Hand1, Elk1, SP1, TLX2 and POU2F1 (Origene); Mtf1, Deaf1, Mef2a, Tcf12, Sp3, Alx4, Smad4 and Rora (Invitrogen). The remaining eight expression plasmids were generously provided by Ttf1 (I. Grummt), GATA4 (C. Asselin and W.L. Miller), Lhx3 (I. Bach), AR (H. Gronemeyer), ZEB1 (T. Brabletz), Mnt (R.N. Eisenman), Pax4 (A. Mansouri) and Zic2 (Y. Yang). All expression vectors are driven by a CMV promoter except for AR (SV40 promoter). All plasmids were constructed from mouse sequences except SP1, TLX2, POU2F1, AR and GATA4 derived from human sequence and ZEB1 from chicken.
In silico identification of potential regulatory elements
The analysis was performed on the Nxnl1 5′ genomic region of 4.2 kb upstream of the translation initiation site to predict TF binding elements potentially involved in the regulation of the expression of this gene. The TRANSFAC® professional database release 9.1 (http://www.biobase-international.com) (52,53) and the MATCH program (54,55) were used with the profile containing the high quality matrices allowing the minimization of the false positive and false negative predictions.
RNA purification
The Cesium chloride (CsCl2) method used here was based on the protocol described by Glisin et al. (51). RNA integrity was assessed by denaturant gel electrophoresis, using a method adapted from published procedures (56–58).
Transcriptomic analysis
The neural retina was dissected as in (59) and the brain sample corresponds to the whole brain hemisphere including the cerebellum. The outer retinal cells of the C57/BL6@N at PN35 (n = 5) were isolated using vibratome sectioning as described previously (41). The RNA was purified by CsCl2 ultracentrifugation as previously described and used to generate double-stranded cDNA, then transcribed in vitro to form biotin-labeled cRNA, fragmented and hybridized to the mouse Affymetrix U74v2 GeneChips® Array for 16 h at 45°C. Arrays were washed and stained using standard protocols (60). Scanned data (.DAT file) were captured by the Affymetrix GeneChip®. Laboratory Information Management System that allows for the average intensities for all probes cells (.CEL file). Quality control performed using RReportGenerator (61) confirmed that all arrays used in the study were of good and consistent quality (available on request). To select the TFs, a filter was used to retain 402 TFs with a signal intensity value superior to 40 units in the sample prepared from the outer retina at PN40 after normalization by Robust Multi-array Average (RMA) (62).
Retinobase utilization
Retinobase provides efficient access to the global expression profiles of genes from different tissues (Outer Retina, Neural Retina and Brain) of C57/BL6@N mouse at PN35 (62). After connection on the site http://genoret.igbmc.fr/RetinoBase/TranscriptomicGraphManager.php using either Internet explorer or Mozilla firefox enter the user name [guest] and password [star128], use the following instructions: In ‘Experiment’ select Exp-34:subPG161-NeuraleRetina-OuterRetina-Brain-PN35, In ‘related Arraytype’ select the Mouse Genome U74ABv2, In the new link, select ‘Search Probesets’, click on ‘Term search’, enter a keyword or when known, the code of the gene and press ‘Search’, In the table, select first the gene which possesses an RMA profile and then select the RMA column (alternatively dChip), In the new link, select ‘Graphes’ and change the ‘graph type’ radar to histogram, Press ‘Get graph’ to validate and see the results.
Luciferase assay
Griptite 293 MSR cells (Invitrogen) were cultivated with DMEM (Gibco) with 10% BSA (Gibco), 600 µg/ml geneticin and subcultured a minimum of three passages after thawing before used. For transient transfection assays cells were seeded to 80–90% confluency in 96-wells plates, then after overnight incubation were transfected with Lipofectamine 2000 (Invitrogen) following manufacturer instructions. Before transfection, the medium was replaced by 100 µl of OPTI-MEM (Invitrogen) without serum. For all transfection, the total quantity of DNA and the volume of reagents and medium added to each well were equalized. For testing candidate TFs, a reaction mixture containing 1 µg of DNA in 100 µl (Opti MEM/lipofectamine 2000) corresponding to of 0.1 µg of luciferase reporter, 0.0–0.9 µg of plasmids encoding for candidate TF, 0.0–0.9 µg of pcDNA3.1 + plasmid (empty vector) and 1 ng of internal control pRL-TK (thymidine kinase promoter, Promega). We divided this transfection reaction in triplicates of 20 µl that were added to the cells in three wells. To assay for synergetic effect, a reaction mixture of 1.1 µg DNA in 100 µl (Opti MEM/lipofectamine 2000) containing 0.5 µg of individual TF and 0.5 µg pcDNA3.1+ or, alternatively 0.5 µg TF#1 + 0.5 µg TF#2 was mixed with 0.1 µg of reporter plasmid and 1 ng of internal control plasmid pRL-TK. This reaction was used to perform triplicates. The cells were lysed with 45 µl of 1× passive lysis buffer (Promega) after 48 h of culture. To measure luciferase activity using Dual Luciferase assay (Promega), 50 µl of firefly luciferase reagent and 50 µl of Renilla luciferase reagent were subsequently injected to 20 µl of cell lysate in each well and recorded using a luminometer (MicroLumat Plus LB 96V, Berthold) after 2 s delay for 10 s. Each experiment was repeated independently three times.
In situ hybridization and immunostaining
In situ hybridization was performed as described in Chotteau-Lelièvre et al. (63). Mouse eyes were fixed overnight in 4% paraformaldehyde in PBS at 4°C, incubate 1 h in 10% sucrose/PBS and incubate overnight in again 20% sucrose/PBS at 4°C. The eyes were orientated and flash frozen in O.C.T (Shandon Cryomatrix, Thermo) using liquid nitrogen steam. In situ hybridization analysis was performed with digoxigenin-labeled riboprobes on 8 µm cryostat sagital section through the optic nerve. The Nxnl1 probe was generated from pGEM-T plasmid (Promega) into which a 332 bp fragment of mouse Nxnl1 exon 1 (Primers, 5′-ATGGCATCTCTCTTCTCTGGACG-3′/5′-CCTCACCTCCTCAGTTCATCATGG-3′). The plasmids were linearized with either SacII (sense probe) or SacI (antisense probe). Chx10/Vsx2 probes (Generously gift by Sandro Banfi, TIGEM) corresponding to the ORF of Chx10/Vsx2, was linearized with NotI (sens probe) or EcoRI (antisense probe). Purification of the riboprobes was performed by size exclusion on a G-50 column (ProbeQuant, Amersham).
Chromatin immunoprecipitation
The ChIP was performed as described previously (64,65). Briefly, retinas from PN35 BALB/c wt mice were frozen on liquid nitrogen and kept at −80°C until use. Retina was cross-linked with ice-cold 4% formaldehyde in phosphate-buffered saline (PBS) for 30 min, rinsed in PBS, and sonicated in lysis buffer (1% SDS, 10 mm EDTA, 50 mm Tris–HCl pH 8.0) plus protease inhibitors (Sigma) to an average DNA size of 1 kb (Vibra Cell, Sonics and Materials Inc., Danbury, CT, USA). The sonicated sample was centrifuged at 15 000g for 10 min at 4°C, the supernatant, precleared with G-sepharose beads at room temperature (RT), was aliquoted to 100 µl (equivalent to 1 whole retina) and diluted to 1 ml with dilution buffer (1% Triton X-100, 2 mm EDTA, 150 mm NaCl, 20 mm Tris–HCl pH 8.0). Each diluted sample was incubated for 1 h at RT with 5 µl of anti-CHX10/VSX2 N-terminal antibody (Exalpha Biologicals). Samples were centrifuged at 15 000g for 10 min at 20°C, the supernatant mixed with 40 µl of protein G-Sepharose (Sigma), 200 µg of sonicated salmon sperm DNA and 2 mg of yeast tRNA (Invitrogen), and incubated for 1 h at RT. Precipitates were washed sequentially for 10 min at RT in 1× TSEI (0.1% SDS, 1% Triton X-100, 2 mm EDTA, 20 mm Tris–HCl pH 8.0, 150 mm NaCl), 4× TSEII (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mm Tris–HCl pH 8.0, 500 mm NaCl), 1× buffer III (0.25 m LiCl, 1% Nonidet P-40, 1% deoxycholate, 1 mm EDTA, 10 mm Tris–HCl pH 8.0) and 3× TE (10 mm Tris–HCl pH 8.0, 1 mm EDTA). Samples were then eluted and cross-links reversed by overnight incubation at 65°C in 100 µl of elution buffer (1% SDS, 0.1 m NaHCOB3B) and DNA fragments were purified by phenol-chloroform and resuspended in 100 µl de TE’ (10 mm Tris–HCl pH 8.0, 0.1 mm EDTA). PCR was used to amplify 2 µl of the final immunoprecipited material. The Primers used was 5′-CTGGTGGTGATGAAGATCTCGGCC-3′/5′-GGGGAGCAGGTGCAGAGAATCTGAT-3′ for the Nxnl1 promoter, 5′-GCCGAAATTTGGATTTACGA-3′/5′-TGGATGAGTGGGGAGAAATC-3′ for the Vsx1 promoter and 5′-TTACTTGAGAGATCCTGACTCAACAATAA-3′/5′-TCAATAACTGCCTTCAGAGAATCG-3′ for the β-Globin promoter. PCR reaction was performed in 25 µl for 94°C 3 min, 35× (94°C 30 s-60°C 30 s-72°C 1 min), 72°C 3 min.
Reverse transcription and real-time PCR
One microgram of RNA was used for the retro transcription with Superscript II RNase H- (Invitrogen) following manufacturer instructions. The cDNA were purified by phenol-chloroform extraction and ethanol precipitation and dissolved in 50 µl of TE (10 mm Tris–HCl pH 8.0, 1 mm EDTA). For the real-time PCR (Light cycler, Roche), 2 µl of 1/10 cDNA were used in a 25 µl total volume containing H2O, 2.5 µm of each primer, 5× FastStart LCDNA master SYBER (Roche) and amplified for 1 × 95°C 10 min, 35× (2 s at 95°C; 5 s at 60°C, 10 s at 72°C). The melting temperature was measured after 2 min at 95°C by increasing the temperature from 65 to 95°C at a rate of +0.1°C/s. The primers used was 5′-TGCTGGCCTCGGAACTGCTCTT-3′/5′-AGGACGCTGGATCGGGAGTATGTC-3′ for the Chx10/Vsx2 and 5′-CCCCATCCCGAATCTCAATGAAGT-3′/5′-CAGGACACGAAGTGAGAAGCCAGC-3′ for the Neuropilin 2 (Nrp2).
Tissues expression profile bioinformatic analysis
The expression profiles on the candidate TFs in Tables 1 and 2 were analyzed with data available at NCBI. The count of EST for each gene was calculated for all tissues reported in unigene (e.g. Chx10/Vsx2, http://www.ncbi.nlm.nih.gov/UniGene/ESTProfileViewer.cgi?uglist=Mm.4405).
Table 2.
Expression level of the homeogenes selected in the study
| Gene ID NCBI | Gene title | Gene symbol | Probe set ID | Signal intensity (RMA) |
OR/NR | Tissue |
Clone number | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| OR | NR | Brain | ≤5 | >5 | ND | ||||||
| 19434 | Retina and anterior neural fold homeobox | Rax | 97149_at | 447.86 | 431.18 | 130.36 | 1.04 | • | LA0AAA52YA10 | ||
| 64379 | Iroquois related homeobox | Irx6 | • | LA0AAA16YO24 | |||||||
| 24113 | Ventral anterior homeobox containing gene 2 | Vax2 | 99817_at | 263.54 | 259.22 | 204.13 | 1.02 | • | LA0AAA102YH09 | ||
| 114889 | Visual system homeobox 1 homolog (zebrafish) | Vsx1 | • | LA0AAA53YE15 | |||||||
| 20476 | Sine oculis-related homeobox 6 homolog | Six6 | 101797_at | 62.90 | 56.44 | 57.46 | 1.1 | • | LA0AAA107YP10 | ||
| 18424 | Orthodenticle homolog 2 (Drosophila) | Otx2 | • | LA0AAA28YF06 | |||||||
OR, outer retina; NR, neural retina. The tissue distribution of ESTs within the public database (≤5 tissues with eyes), large (>5 tissues with eyes) or no determined (ND) of each HDs in mouse (•) tissues (according to UniGene, NCBI). Clone number refers to the sequenced library.
SUPPLEMENTARY MATERIAL
FUNDING
This work was supported by Inserm, CIFRE, ANR Chaire d'excellence, NIH 5R01EY009769 and EVI-GENORET.
Supplementary Material
ACKNOWLEDGEMENTS
We thank E. Clerin, N. Berdugo, M.-L. Niepon, W. Raffelsberger, B. Kinzel, C. DaSilva, P. Winker, I. Grummt, C. Asselin, W.L. Miller, I. Bach, H. Gronemeyer, T. Brabletz, R.N. Eisenman, A. Mansouri, Y. Yang, M. Mouradian, A. Chédotal, S. Banfi, T. van Veen and Fovea Pharmaceuticals.
Conflict of Interest statement. None declared.
REFERENCES
- 1.Delyfer M.N., Leveillard T., Mohand-Said S., Hicks D., Picaud S., Sahel J.A. Inherited retinal degenerations: therapeutic prospects. Biol. Cell. 2004;96:261–269. doi: 10.1016/j.biolcel.2004.01.006. [DOI] [PubMed] [Google Scholar]
- 2.Kajiwara K., Berson E.L., Dryja T.P. Digenic retinitis pigmentosa due to mutations at the unlinked peripherin/RDS and ROM1 loci. Science. 1994;264:1604–1608. doi: 10.1126/science.8202715. [DOI] [PubMed] [Google Scholar]
- 3.McLaughlin M.E., Sandberg M.A., Berson E.L., Dryja T.P. Recessive mutations in the gene encoding the beta-subunit of rod phosphodiesterase in patients with retinitis pigmentosa. Nat. Genet. 1993;4:130–134. doi: 10.1038/ng0693-130. [DOI] [PubMed] [Google Scholar]
- 4.Rosenfeld P.J., Cowley G.S., McGee T.L., Sandberg M.A., Berson E.L., Dryja T.P. A null mutation in the rhodopsin gene causes rod photoreceptor dysfunction and autosomal recessive retinitis pigmentosa. Nat. Genet. 1992;1:209–213. doi: 10.1038/ng0692-209. [DOI] [PubMed] [Google Scholar]
- 5.al-Maghtheh M., Gregory C., Inglehearn C., Hardcastle A., Bhattacharya S. Rhodopsin mutations in autosomal dominant retinitis pigmentosa. Hum. Mutat. 1993;2:249–255. doi: 10.1002/humu.1380020403. [DOI] [PubMed] [Google Scholar]
- 6.McLaughlin M.E., Ehrhart T.L., Berson E.L., Dryja T.P. Mutation spectrum of the gene encoding the beta subunit of rod phosphodiesterase among patients with autosomal recessive retinitis pigmentosa. Proc. Natl. Acad. Sci. USA. 1995;92:3249–3253. doi: 10.1073/pnas.92.8.3249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rosenfeld P.J., Hahn L.B., Sandberg M.A., Dryja T.P., Berson E.L. Low incidence of retinitis pigmentosa among heterozygous carriers of a specific rhodopsin splice site mutation. Invest. Ophthalmol. Vis. Sci. 1995;36:2186–2192. [PubMed] [Google Scholar]
- 8.Chang G.Q., Hao Y., Wong F. Apoptosis: final common pathway of photoreceptor death in rd, rds, and rhodopsin mutant mice. Neuron. 1993;11:595–605. doi: 10.1016/0896-6273(93)90072-y. [DOI] [PubMed] [Google Scholar]
- 9.Travis G.H. Mechanisms of cell death in the inherited retinal degenerations. Am. J. Hum. Genet. 1998;62:503–508. doi: 10.1086/301772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bowes C., Li T., Danciger M., Baxter L.C., Applebury M.L., Farber D.B. Retinal degeneration in the rd mouse is caused by a defect in the beta subunit of rod cGMP-phosphodiesterase. Nature. 1990;347:677–680. doi: 10.1038/347677a0. [DOI] [PubMed] [Google Scholar]
- 11.Carter-Dawson L.D., LaVail M.M., Sidman R.L. Differential effect of the rd mutation on rods and cones in the mouse retina. Invest. Ophthalmol. Vis. Sci. 1978;17:489–498. [PubMed] [Google Scholar]
- 12.Farber D.B. From mice to men: the cyclic GMP phosphodiesterase gene in vision and disease. The Proctor Lecture. Invest. Ophthalmol. Vis. Sci. 1995;36:263–275. [PubMed] [Google Scholar]
- 13.Mohand-Said S., Hicks D., Dreyfus H., Sahel J.A. Selective transplantation of rods delays cone loss in a retinitis pigmentosa model. Arch. Ophthalmol. 2000;118:807–811. doi: 10.1001/archopht.118.6.807. [DOI] [PubMed] [Google Scholar]
- 14.Mohand-Said S., Hicks D., Simonutti M., Tran-Minh D., Deudon-Combe A., Dreyfus H., Silverman M.S., Ogilvie J.M., Tenkova T., Sahel J. Photoreceptor transplants increase host cone survival in the retinal degeneration (rd) mouse. Ophthalmic Res. 1997;29:290–297. doi: 10.1159/000268027. [DOI] [PubMed] [Google Scholar]
- 15.Hewitt A.T., Lindsey J.D., Carbott D., Adler R. Photoreceptor survival-promoting activity in interphotoreceptor matrix preparations: characterization and partial purification. Exp. Eye Res. 1990;50:79–88. doi: 10.1016/0014-4835(90)90013-k. [DOI] [PubMed] [Google Scholar]
- 16.Leveillard T., Mohand-Said S., Lorentz O., Hicks D., Fintz A.C., Clerin E., Simonutti M., Forster V., Cavusoglu N., Chalmel F., et al. Identification and characterization of rod-derived cone viability factor. Nat. Genet. 2004;36:755–759. doi: 10.1038/ng1386. [DOI] [PubMed] [Google Scholar]
- 17.Oppenheim R.W. The neurotrophic theory and naturally occurring motoneuron death. Trends Neurosci. 1989;12:252–255. doi: 10.1016/0166-2236(89)90021-0. [DOI] [PubMed] [Google Scholar]
- 18.Chalmel F., Leveillard T., Jaillard C., Lardenois A., Berdugo N., Morel E., Koehl P., Lambrou G., Holmgren A., Sahel J.A., et al. Rod-derived Cone Viability Factor-2 is a novel bifunctional-thioredoxin-like protein with therapeutic potential. BMC Mol. Biol. 2007;8:74. doi: 10.1186/1471-2199-8-74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Funato Y., Miki H. Nucleoredoxin, a novel thioredoxin family member involved in cell growth and differentiation. Antioxid. Redox. Signal. 2007;9:1035–1057. doi: 10.1089/ars.2007.1550. [DOI] [PubMed] [Google Scholar]
- 20.Lillig C.H., Holmgren A. Thioredoxin and related molecules–from biology to health and disease. Antioxid. Redox. Signal. 2007;9:25–47. doi: 10.1089/ars.2007.9.25. [DOI] [PubMed] [Google Scholar]
- 21.Koharyova M., Kolarova M. Oxidative stress and thioredoxin system. Gen. Physiol. Biophys. 2008;27:71–84. [PubMed] [Google Scholar]
- 22.Masutani H., Bai J., Kim Y.C., Yodoi J. Thioredoxin as a neurotrophic cofactor and an important regulator of neuroprotection. Mol. Neurobiol. 2004;29:229–242. doi: 10.1385/MN:29:3:229. [DOI] [PubMed] [Google Scholar]
- 23.Nakamura H. Thioredoxin as a key molecule in redox signaling. Antioxid. Redox. Signal. 2004;6:15–17. doi: 10.1089/152308604771978309. [DOI] [PubMed] [Google Scholar]
- 24.Pekkari K., Holmgren A. Truncated thioredoxin: physiological functions and mechanism. Antioxid. Redox. Signal. 2004;6:53–61. doi: 10.1089/152308604771978345. [DOI] [PubMed] [Google Scholar]
- 25.Komeima K., Rogers B.S., Lu L., Campochiaro P.A. Antioxidants reduce cone cell death in a model of retinitis pigmentosa. Proc. Natl. Acad. Sci. USA. 2006;103:11300–11305. doi: 10.1073/pnas.0604056103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fridlich R., Delalande F., Jaillard C., Lu J., Poidevin L., Cronin T., Perrocheau L., Millet-Puel G., Niepon M.L., Poch O., et al. The thioredoxin-like protein RdCVFL interacts with Tau and inhibits its phosphorylation in the retina. Mol. Cell. Proteomics. 2009;8:1206–1218. doi: 10.1074/mcp.M800406-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bennett J. Strategies for delivery of rod-derived cone viability factor. Retina. 2005;25:S47. doi: 10.1097/00006982-200512001-00020. [DOI] [PubMed] [Google Scholar]
- 28.Tao W., Wen R., Goddard M.B., Sherman S.D., O'Rourke P.J., Stabila P.F., Bell W.J., Dean B.J., Kauper K.A., Budz V.A., et al. Encapsulated cell-based delivery of CNTF reduces photoreceptor degeneration in animal models of retinitis pigmentosa. Invest. Ophthalmol. Vis. Sci. 2002;43:3292–3298. [PubMed] [Google Scholar]
- 29.Sieving P.A., Caruso R.C., Tao W., Coleman H.R., Thompson D.J., Fullmer K.R., Bush R.A. Ciliary neurotrophic factor (CNTF) for human retinal degeneration: phase I trial of CNTF delivered by encapsulated cell intraocular implants. Proc. Natl. Acad. Sci. USA. 2006;103:3896–3901. doi: 10.1073/pnas.0600236103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yang Y., Mohand-Said S., Danan A., Simonutti M., Fontaine V., Clerin E., Picaud S., Leveillard T., Sahel J.A. Functional Cone Rescue by RdCVF Protein in a Dominant Model of Retinitis Pigmentosa. Mol. Ther. 2009;17:787–795. doi: 10.1038/mt.2009.28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Burmeister M., Novak J., Liang M.Y., Basu S., Ploder L., Hawes N.L., Vidgen D., Hoover F., Goldman D., Kalnins V.I., et al. Ocular retardation mouse caused by Chx10 homeobox null allele: impaired retinal progenitor proliferation and bipolar cell differentiation. Nat. Genet. 1996;12:376–384. doi: 10.1038/ng0496-376. [DOI] [PubMed] [Google Scholar]
- 32.Chow R.L., Snow B., Novak J., Looser J., Freund C., Vidgen D., Ploder L., McInnes R.R. Vsx1, a rapidly evolving paired-like homeobox gene expressed in cone bipolar cells. Mech. Dev. 2001;109:315–322. doi: 10.1016/s0925-4773(01)00585-8. [DOI] [PubMed] [Google Scholar]
- 33.Lupo G., Andreazzoli M., Gestri G., Liu Y., He R.Q., Barsacchi G. Homeobox genes in the genetic control of eye development. Int. J. Dev. Biol. 2000;44:627–636. [PubMed] [Google Scholar]
- 34.Greferath U., Grunert U., Wassle H. Rod bipolar cells in the mammalian retina show protein kinase C-like immunoreactivity. J. Comp. Neurol. 1990;301:433–442. doi: 10.1002/cne.903010308. [DOI] [PubMed] [Google Scholar]
- 35.Liang L., Sandell J.H. Focus on molecules: homeobox protein Chx10. Exp. Eye Res. 2008;86:541–542. doi: 10.1016/j.exer.2007.03.004. [DOI] [PubMed] [Google Scholar]
- 36.Liu I.S., Chen J.D., Ploder L., Vidgen D., van der Kooy D., Kalnins V.I., McInnes R.R. Developmental expression of a novel murine homeobox gene (Chx10): evidence for roles in determination of the neuroretina and inner nuclear layer. Neuron. 1994;13:377–393. doi: 10.1016/0896-6273(94)90354-9. [DOI] [PubMed] [Google Scholar]
- 37.Shen J., Samul R., Zimmer J., Liu H., Liang X., Hackett S., Campochiaro P.A. Deficiency of neuropilin 2 suppresses VEGF-induced retinal neovascularization. Mol. Med. 2004;10:12–18. doi: 10.2119/2004-00017.campochiaro. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Clark A.M., Yun S., Veien E.S., Wu Y.Y., Chow R.L., Dorsky R.I., Levine E.M. Negative regulation of Vsx1 by its paralog Chx10/Vsx2 is conserved in the vertebrate retina. Brain Res. 2008;1192:99–113. doi: 10.1016/j.brainres.2007.06.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hayashi T., Huang J., Deeb S.S. RINX(VSX1), a novel homeobox gene expressed in the inner nuclear layer of the adult retina. Genomics. 2000;67:128–139. doi: 10.1006/geno.2000.6248. [DOI] [PubMed] [Google Scholar]
- 40.Kumar R., Chen S., Scheurer D., Wang Q.L., Duh E., Sung C.H., Rehemtulla A., Swaroop A., Adler R., Zack D.J. The bZIP transcription factor Nrl stimulates rhodopsin promoter activity in primary retinal cell cultures. J. Biol. Chem. 1996;271:29612–29618. doi: 10.1074/jbc.271.47.29612. [DOI] [PubMed] [Google Scholar]
- 41.Fontaine V., Kinkl N., Sahel J., Dreyfus H., Hicks D. Survival of purified rat photoreceptors in vitro is stimulated directly by fibroblast growth factor-2. J. Neurosci. 1998;18:9662–9672. doi: 10.1523/JNEUROSCI.18-23-09662.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Rath M.F., Bailey M.J., Kim J.S., Coon S.L., Klein D.C., Moller M. Developmental and daily expression of the Pax4 and Pax6 homeobox genes in the rat retina: localization of Pax4 in photoreceptor cells. J. Neurochem. 2009;108:285–294. doi: 10.1111/j.1471-4159.2008.05765.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lerner L.E., Gribanova Y.E., Whitaker L., Knox B.E., Farber D.B. The rod cGMP-phosphodiesterase beta-subunit promoter is a specific target for Sp4 and is not activated by other Sp proteins or CRX. J. Biol. Chem. 2002;277:25877–25883. doi: 10.1074/jbc.M201407200. [DOI] [PubMed] [Google Scholar]
- 44.Hatakeyama J., Tomita K., Inoue T., Kageyama R. Roles of homeobox and bHLH genes in specification of a retinal cell type. Development. 2001;128:1313–1322. doi: 10.1242/dev.128.8.1313. [DOI] [PubMed] [Google Scholar]
- 45.Rowan S., Cepko C.L. Genetic analysis of the homeodomain transcription factor Chx10 in the retina using a novel multifunctional BAC transgenic mouse reporter. Dev. Biol. 2004;271:388–402. doi: 10.1016/j.ydbio.2004.03.039. [DOI] [PubMed] [Google Scholar]
- 46.Toy J., Sundin O.H. Expression of the optx2 homeobox gene during mouse development. Mech. Dev. 1999;83:183–186. doi: 10.1016/s0925-4773(99)00049-0. [DOI] [PubMed] [Google Scholar]
- 47.Alfano G., Vitiello C., Caccioppoli C., Caramico T., Carola A., Szego M.J., McInnes R.R., Auricchio A., Banfi S. Natural antisense transcripts associated with genes involved in eye development. Hum. Mol. Genet. 2005;14:913–923. doi: 10.1093/hmg/ddi084. [DOI] [PubMed] [Google Scholar]
- 48.Furukawa T., Morrow E.M., Cepko C.L. Crx, a novel otx-like homeobox gene, shows photoreceptor-specific expression and regulates photoreceptor differentiation. Cell. 1997;91:531–541. doi: 10.1016/s0092-8674(00)80439-0. [DOI] [PubMed] [Google Scholar]
- 49.Bibb L.C., Holt J.K., Tarttelin E.E., Hodges M.D., Gregory-Evans K., Rutherford A., Lucas R.J., Sowden J.C., Gregory-Evans C.Y. Temporal and spatial expression patterns of the CRX transcription factor and its downstream targets. Critical differences during human and mouse eye development. Hum. Mol. Genet. 2001;10:1571–1579. doi: 10.1093/hmg/10.15.1571. [DOI] [PubMed] [Google Scholar]
- 50.Viczian A., Sanyal S., Toffenetti J., Chader G.J., Farber D.B. Photoreceptor-specific mRNAs in mice carrying different allelic combinations at the rd and rds loci. Exp. Eye Res. 1992;54:853–860. doi: 10.1016/0014-4835(92)90148-l. [DOI] [PubMed] [Google Scholar]
- 51.Glisin V., Crkvenjakov R., Byus C. Ribonucleic acid isolated by cesium chloride centrifugation. Biochemistry. 1974;13:2633–2637. doi: 10.1021/bi00709a025. [DOI] [PubMed] [Google Scholar]
- 52.Knuppel R., Dietze P., Lehnberg W., Frech K., Wingender E. TRANSFAC retrieval program: a network model database of eukaryotic transcription regulating sequences and proteins. J. Comput. Biol. 1994;1:191–198. doi: 10.1089/cmb.1994.1.191. [DOI] [PubMed] [Google Scholar]
- 53.Wingender E., Dietze P., Karas H., Knuppel R. TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic Acids Res. 1996;24:238–241. doi: 10.1093/nar/24.1.238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kel A.E., Gossling E., Reuter I., Cheremushkin E., Kel-Margoulis O.V., Wingender E. MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 2003;31:3576–3579. doi: 10.1093/nar/gkg585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Lardenois A., Chalmel F., Bianchetti L., Sahel J.A., Leveillard T., Poch O. PromAn: an integrated knowledge-based web server dedicated to promoter analysis. Nucleic Acids Res. 2006;34:W578–W583. doi: 10.1093/nar/gkl193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Goldberg D.A. Isolation and partial characterization of the Drosophila alcohol dehydrogenase gene. Proc. Natl. Acad. Sci. USA. 1980;77:5794–5798. doi: 10.1073/pnas.77.10.5794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lehrach H., Diamond D., Wozney J.M., Boedtker H. RNA molecular weight determinations by gel electrophoresis under denaturing conditions, a critical reexamination. Biochemistry. 1977;16:4743–4751. doi: 10.1021/bi00640a033. [DOI] [PubMed] [Google Scholar]
- 58.Rosen K.M., Lamperti E.D., Villa-Komaroff L. Optimizing the northern blot procedure. Biotechniques. 1990;8:398–403. [PubMed] [Google Scholar]
- 59.Mohand-Said S., Deudon-Combe A., Hicks D., Simonutti M., Forster V., Fintz A.C., Leveillard T., Dreyfus H., Sahel J.A. Normal retina releases a diffusible factor stimulating cone survival in the retinal degeneration mouse. Proc. Natl. Acad. Sci. USA. 1998;95:8357–8362. doi: 10.1073/pnas.95.14.8357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Lockhart D.J., Dong H., Byrne M.C., Follettie M.T., Gallo M.V., Chee M.S., Mittmann M., Wang C., Kobayashi M., Horton H., et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nat. Biotechnol. 1996;14:1675–1680. doi: 10.1038/nbt1296-1675. [DOI] [PubMed] [Google Scholar]
- 61.Raffelsberger W., Krause Y., Moulinier L., Kieffer D., Morand A.L., Brino L., Poch O. RReportGenerator: automatic reports from routine statistical analysis using R. Bioinformatics. 2008;24:276–278. doi: 10.1093/bioinformatics/btm556. [DOI] [PubMed] [Google Scholar]
- 62.Kalathur R.K., Gagniere N., Berthommier G., Poidevin L., Raffelsberger W., Ripp R., Leveillard T., Poch O. RETINOBASE: a web database, data mining and analysis platform for gene expression data on retina. BMC Genomics. 2008;9:208. doi: 10.1186/1471-2164-9-208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Chotteau-Lelievre A., Dolle P., Gofflot F. Expression analysis of murine genes using in situ hybridization with radioactive and nonradioactively labeled RNA probes. Methods Mol. Biol. 2006;326:61–87. doi: 10.1385/1-59745-007-3:61. [DOI] [PubMed] [Google Scholar]
- 64.Pattenden S.G., Klose R., Karaskov E., Bremner R. Interferon-gamma-induced chromatin remodeling at the CIITA locus is BRG1 dependent. Embo. J. 2002;21:1978–1986. doi: 10.1093/emboj/21.8.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Dorval K.M., Bobechko B.P., Fujieda H., Chen S., Zack D.J., Bremner R. CHX10 targets a subset of photoreceptor genes. J. Biol. Chem. 2006;281:744–751. doi: 10.1074/jbc.M509470200. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.